Themed collection Glycomics & Glycoproteomics: From Analytics to Function

Glycomics & Glycoproteomics: From Analytics to Function
Morten Thaysen-Andersen, Daniel Kolarich and Nicolle H. Packer introduce the Molecular Omics themed issue on Glycomics & Glycoproteomics: From Analytics to Function.

Mol. Omics, 2021,17, 8-10
https://doi.org/10.1039/D0MO90019B
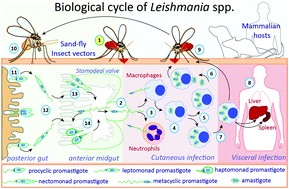
Protein glycosylation in Leishmania spp.
Protein glycosylation is a co- and post-translational modification that, in Leishmania parasites, plays key roles in vector–parasite–vertebrate host interaction.
Mol. Omics, 2020,16, 407-424
https://doi.org/10.1039/D0MO00043D
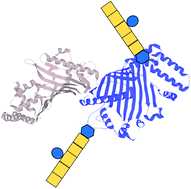
Identifying the targets and functions of N-linked protein glycosylation in Campylobacter jejuni
Virulence of Campylobacter jejuni is dependent on the ability to glycosylate membrane-associated proteins.
Mol. Omics, 2020,16, 287-304
https://doi.org/10.1039/D0MO00032A
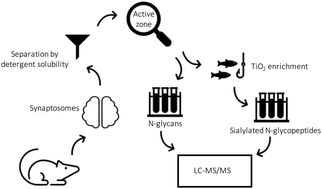
N-Glycosylation in isolated rat nerve terminals
Glycomics and sialiomics of isolated synaptosomes reveal distinct glycosylation of surface proteins localized in the active zone of synapses.
Mol. Omics, 2021,17, 517-532
https://doi.org/10.1039/D0MO00044B
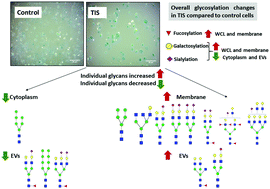
N-Linked glycosylation profiles of therapeutic induced senescent (TIS) triple negative breast cancer cells (TNBC) and their extracellular vesicle (EV) progeny
Therapeutic-induced-senescent (TIS) Cal51 TNBC cells display differential N-glycan moieties compared to non-senescent cells, depending on cellular location and EV progeny.
Mol. Omics, 2021,17, 72-85
https://doi.org/10.1039/D0MO00017E
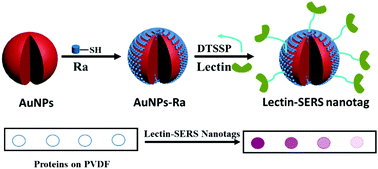
Rapid and sensitive glycan targeting by lectin-SERS assay
The fabrication of lectin-SERS nanotags and the assay designed for rapid glycoprotein identification and quantification.
Mol. Omics, 2020,16, 339-344
https://doi.org/10.1039/C9MO00181F
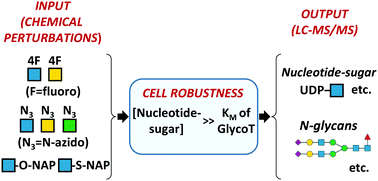
Robustness in glycosylation systems: effect of modified monosaccharides, acceptor decoys and azido sugars on cellular nucleotide-sugar levels and pattern of N-linked glycosylation
Chemical perturbation studies reveal robustness in glycosylation systems, based on comparison of LC-MS/MS quantification of cellular nucleotide-sugar levels with the observed N-linked glycan patterns.
Mol. Omics, 2020,16, 377-386
https://doi.org/10.1039/D0MO00023J
Development of a 96-well plate sample preparation method for integrated N- and O-glycomics using porous graphitized carbon liquid chromatography-mass spectrometry
The developed workflow allows high throughput sample preparation for glycomics analysis.

Mol. Omics, 2020,16, 355-363
https://doi.org/10.1039/C9MO00180H
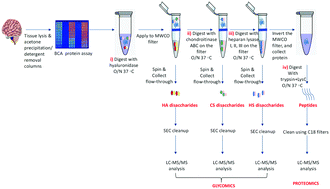
Serial in-solution digestion protocol for mass spectrometry-based glycomics and proteomics analysis
A high-throughput & efficient protocol for mass spectrometry-based glycomic and proteomic analysis.
Mol. Omics, 2020,16, 364-376
https://doi.org/10.1039/D0MO00019A
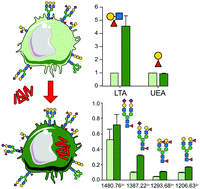
Mycobacterium bovis BCG infection alters the macrophage N-glycome
Macrophage glycosylation that is essential to the host-immune defense may be modulated by pathogens infection.
Mol. Omics, 2020,16, 345-354
https://doi.org/10.1039/C9MO00173E
Recombinant mucin-type proteins carrying LacdiNAc on different O-glycan core chains fail to support H. pylori binding
The β4-N-acetylgalactosaminyltransferase 3 (B4GALNT3) transfers GalNAc in a β1,4-linkage to GlcNAc forming the LacdiNAc (LDN) determinant on oligosaccharides.
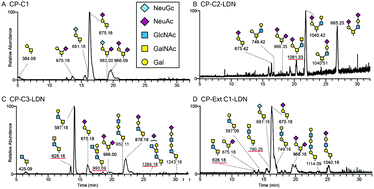
Mol. Omics, 2020,16, 243-257
https://doi.org/10.1039/C9MO00175A
β-Catenin/CBP inhibition alters epidermal growth factor receptor fucosylation status in oral squamous cell carcinoma
Genomic and structural analyses reveal that β-catenin/CBP signaling represses epidermal growth factor receptor (EGFR) N-glycan antennary fucosylation in oral cancer.
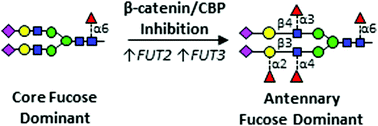
Mol. Omics, 2020,16, 195-209
https://doi.org/10.1039/D0MO00009D
Choosing proper normalization is essential for discovery of sparse glycan biomarkers
In this work we assess the effect of different normalization methods on variable selection in an emerging field of glycomics.
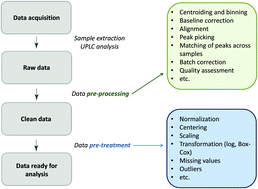
Mol. Omics, 2020,16, 231-242
https://doi.org/10.1039/C9MO00174C
Mass spectrometric analysis of core fucosylation and sequence variation in a human–camelid monoclonal antibody
ESI-MS fucosylation studies on an intact EG2-hFc monoclonal antibody reveal the presence of fucose on both Fc N-glycans.
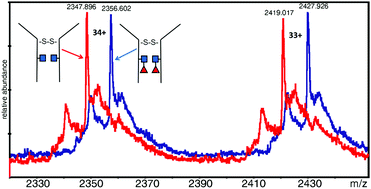
Mol. Omics, 2020,16, 221-230
https://doi.org/10.1039/C9MO00168A
Quantitative capillary zone electrophoresis-mass spectrometry reveals the N-glycome developmental plan during vertebrate embryogenesis
The Xenopus laevis N-glycome undergoes massive reprogramming after the midblastula transition and the onset of neuronal development.
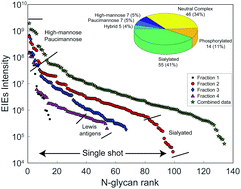
Mol. Omics, 2020,16, 210-220
https://doi.org/10.1039/D0MO00005A
DIALib: an automated ion library generator for data independent acquisition mass spectrometry analysis of peptides and glycopeptides
Data Independent Acquisition (DIA) Mass Spectrometry (MS) workflows allow unbiased measurement of all detectable peptides from complex proteomes, but require ion libraries for interrogation of peptides of interest.
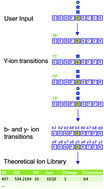
Mol. Omics, 2020,16, 100-112
https://doi.org/10.1039/C9MO00125E
The effectiveness of filtering glycopeptide peak list files for Y ions
Novel software workflow for identifying additional glycopeptides in complex datasets.
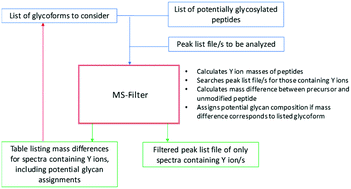
Mol. Omics, 2020,16, 147-155
https://doi.org/10.1039/C9MO00178F
Novel O-linked sialoglycan structures in human urinary glycoproteins
Identification of new glycoforms for glycopeptides confidently assigned from primary database searches permitting the most common O-glycans only.
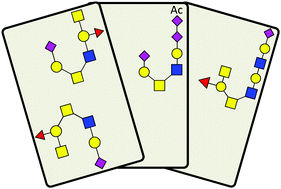
Mol. Omics, 2020,16, 156-164
https://doi.org/10.1039/C9MO00160C
Reduced sialyl-Lewisx on salivary MUC7 from patients with burning mouth syndrome
We analyse and compare MUC7 O-glycosylation and inflammatory biomarkers in saliva from female patients with burning mouth syndrome (BMS) and gender/age-matched controls.
Mol. Omics, 2019,15, 331-339
https://doi.org/10.1039/C9MO00061E