Themed collection Accelerating Chemistry Symposium Collection

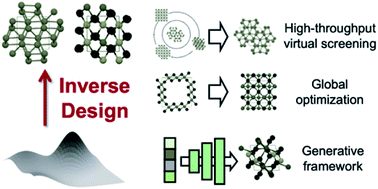
Machine-enabled inverse design of inorganic solid materials: promises and challenges
The grand challenge of materials science, discovery of novel materials with target properties, can be greatly accelerated by machine-learned inverse design strategies.
Chem. Sci., 2020,11, 4871-4881
https://doi.org/10.1039/D0SC00594K
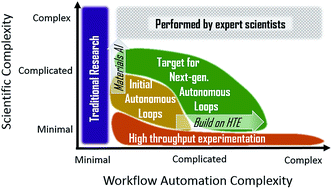
Progress and prospects for accelerating materials science with automated and autonomous workflows
Integrating automation with artificial intelligence will enable scientists to spend more time identifying important problems and communicating critical insights, accelerating discovery and development of materials for emerging and future technologies.
Chem. Sci., 2019,10, 9640-9649
https://doi.org/10.1039/C9SC03766G
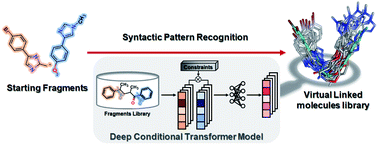
SyntaLinker: automatic fragment linking with deep conditional transformer neural networks
Linking fragments to generate a focused compound library for a specific drug target is one of the challenges in fragment-based drug design (FBDD).
Chem. Sci., 2020,11, 8312-8322
https://doi.org/10.1039/D0SC03126G
Predicting the chemical reactivity of organic materials using a machine-learning approach
Stability and compatibility between chemical components are essential parameters that need to be considered in the selection of functional materials in configuring a system.

Chem. Sci., 2020,11, 7813-7822
https://doi.org/10.1039/D0SC01328E
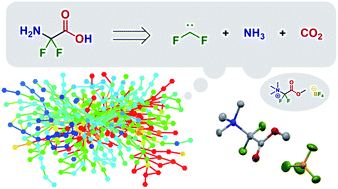
Discovery of a synthesis method for a difluoroglycine derivative based on a path generated by quantum chemical calculations
QCaRA successfully predicted a new synthetic path based on the reaction path network produced by quantum chemical calculation.
Chem. Sci., 2020,11, 7569-7577
https://doi.org/10.1039/D0SC02089C
Pushing property limits in materials discovery via boundless objective-free exploration
Our developed algorithm, BLOX (BoundLess Objective-free eXploration), successfully found “out-of-trend” molecules potentially useful for photofunctional materials from a drug database.
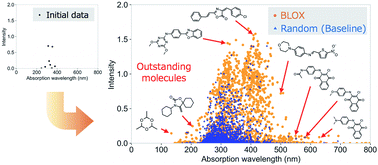
Chem. Sci., 2020,11, 5959-5968
https://doi.org/10.1039/D0SC00982B
Geometric landscapes for material discovery within energy–structure–function maps
We introduce a representation for the geometric features of the pores of porous molecular crystals. This representation provides a good basis for supervised (predict adsorption properties) and unsupervised (polymorph classification) tasks.
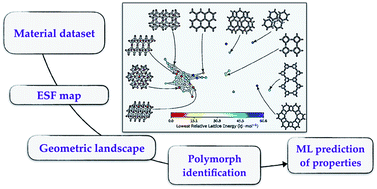
Chem. Sci., 2020,11, 5423-5433
https://doi.org/10.1039/D0SC00049C
Structure-mechanics statistical learning unravels the linkage between local rigidity and global flexibility in nucleic acids
The mechanical properties of nucleic acids underlie biological processes ranging from genome packaging to gene expression. We devise structural mechanics statistical learning method to reveal their molecular origin in terms of chemical interactions.
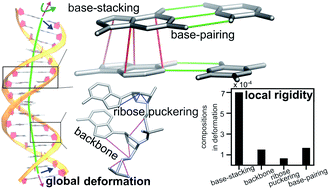
Chem. Sci., 2020,11, 4969-4979
https://doi.org/10.1039/D0SC00480D
Evolutionary chemical space exploration for functional materials: computational organic semiconductor discovery
Evolutionary optimisation and crystal structure prediction are used to explore chemical space for molecular organic semiconductors.

Chem. Sci., 2020,11, 4922-4933
https://doi.org/10.1039/D0SC00554A
Machine learning dihydrogen activation in the chemical space surrounding Vaska's complex
A machine learning exploration of the chemical space surrounding Vaska's complex.
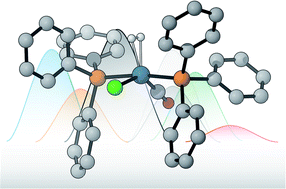
Chem. Sci., 2020,11, 4584-4601
https://doi.org/10.1039/D0SC00445F
Spectral deep learning for prediction and prospective validation of functional groups
A new multi-label deep neural network architecture is used to combine Infrared and mass spectra, trained on single compounds to predict functional groups, and experimentally validated on complex mixtures.

Chem. Sci., 2020,11, 4618-4630
https://doi.org/10.1039/C9SC06240H
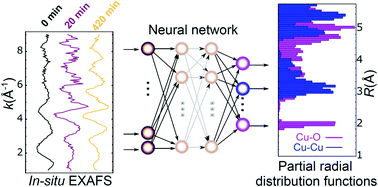
Linking the evolution of catalytic properties and structural changes in copper–zinc nanocatalysts using operando EXAFS and neural-networks
A neural network is used to reveal composition-dependent structural evolution under operando conditions in CuZn nanocatalysts for CO2 electroreduction.
Chem. Sci., 2020,11, 3727-3736
https://doi.org/10.1039/D0SC00382D
Automatic retrosynthetic route planning using template-free models
Retrosynthetic pathway planning using a template-free model coupled with heuristic Monte Carlo tree search.

Chem. Sci., 2020,11, 3355-3364
https://doi.org/10.1039/C9SC03666K
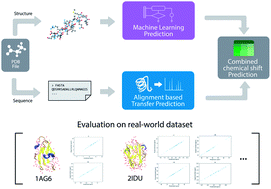
Accurate prediction of chemical shifts for aqueous protein structure on “Real World” data
UCBShift predicts NMR chemical shifts of proteins that exceeds accuracy of other popular chemical shift predictors on real-world data sets.
Chem. Sci., 2020,11, 3180-3191
https://doi.org/10.1039/C9SC06561J
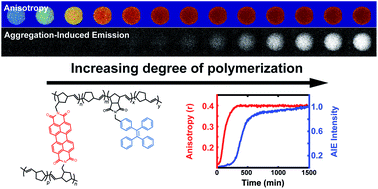
Optical monitoring of polymerizations in droplets with high temporal dynamic range
Two complementary measurements, fluorescence polarization anisotropy and aggregation-induced emission, allow for in situ optical monitoring of polymerization reaction progress in droplets across varying temporal regimes of the reaction.
Chem. Sci., 2020,11, 2647-2656
https://doi.org/10.1039/C9SC05559B
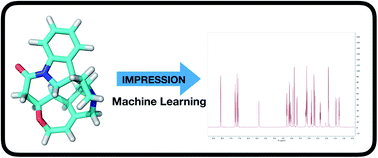
IMPRESSION – prediction of NMR parameters for 3-dimensional chemical structures using machine learning with near quantum chemical accuracy
The IMPRESSION machine learning system can predict NMR parameters for 3D structures with similar results to DFT but in seconds rather than hours.
Chem. Sci., 2020,11, 508-515
https://doi.org/10.1039/C9SC03854J
Datasets and their influence on the development of computer assisted synthesis planning tools in the pharmaceutical domain
Computer Assisted Synthesis Planning (CASP), datasets and their performance.
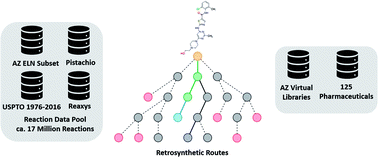
Chem. Sci., 2020,11, 154-168
https://doi.org/10.1039/C9SC04944D
Mining predicted crystal structure landscapes with high throughput crystallisation: old molecules, new insights
New crystal forms of two well-studied organic molecules are identified in a computationally targeted way, by combining structure prediction with a robotic crystallisation screen, including a ‘hidden’ porous polymorph of trimesic acid.
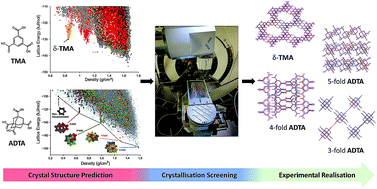
Chem. Sci., 2019,10, 9988-9997
https://doi.org/10.1039/C9SC02832C
Accelerated robotic discovery of type II porous liquids
High-throughput automation was used to streamline the synthesis, characterisation, and solubility testing, of new Type II porous liquids, accelerating their discovery.
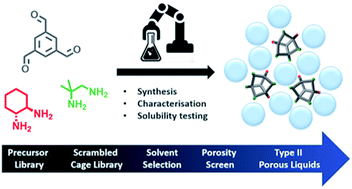
Chem. Sci., 2019,10, 9454-9465
https://doi.org/10.1039/C9SC03316E
Delfos: deep learning model for prediction of solvation free energies in generic organic solvents
We introduce Delfos, a novel, machine-learning-based QSPR method which predicts solvation free energies for generic organic solutions.
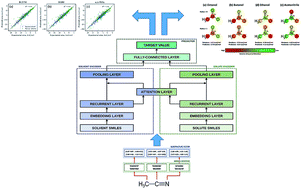
Chem. Sci., 2019,10, 8306-8315
https://doi.org/10.1039/C9SC02452B
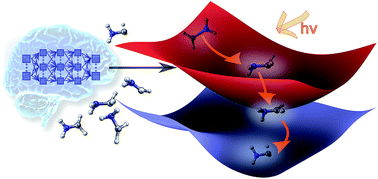
Machine learning enables long time scale molecular photodynamics simulations
Machine learning enables excited-state molecular dynamics simulations including nonadiabatic couplings on nanosecond time scales.
Chem. Sci., 2019,10, 8100-8107
https://doi.org/10.1039/C9SC01742A
Efficient multi-objective molecular optimization in a continuous latent space
We utilize Particle Swarm Optimization to optimize molecules in a machine-learned continuous chemical representation with respect to multiple objectives such as biological activity, structural constrains or ADMET properties.

Chem. Sci., 2019,10, 8016-8024
https://doi.org/10.1039/C9SC01928F
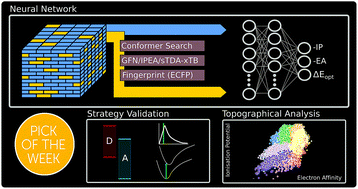
Mapping binary copolymer property space with neural networks
We map the property space of binary copolymers to understand how copolymerisation can be used to tune the optoelectronic properties of polymers.
Chem. Sci., 2019,10, 4973-4984
https://doi.org/10.1039/C8SC05710A
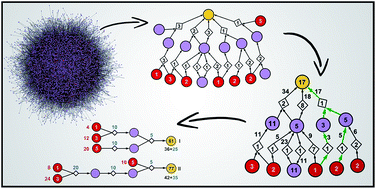
Selection of cost-effective yet chemically diverse pathways from the networks of computer-generated retrosynthetic plans
A family of network algorithms allows the Chematica retrosynthetic platform to plan both cost-effective and chemically diverse syntheses.
Chem. Sci., 2019,10, 4640-4651
https://doi.org/10.1039/C8SC05611K
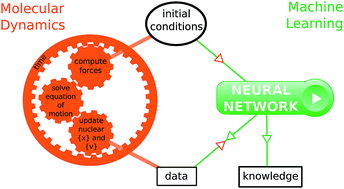
How machine learning can assist the interpretation of ab initio molecular dynamics simulations and conceptual understanding of chemistry
Machine learning models, trained to reproduce molecular dynamics results, help interpreting simulations and extracting new understanding of chemistry.
Chem. Sci., 2019,10, 2298-2307
https://doi.org/10.1039/C8SC04516J
Learning continuous and data-driven molecular descriptors by translating equivalent chemical representations
Translation between semantically equivalent but syntactically different line notations of molecular structures compresses meaningful information into a continuous molecular descriptor.
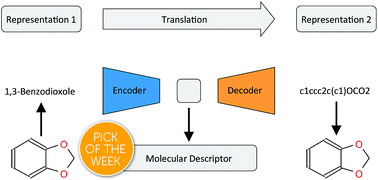
Chem. Sci., 2019,10, 1692-1701
https://doi.org/10.1039/C8SC04175J
A graph-convolutional neural network model for the prediction of chemical reactivity
We present a supervised learning approach to predict the products of organic reactions given their reactants, reagents, and solvent(s).
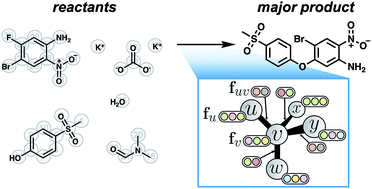
Chem. Sci., 2019,10, 370-377
https://doi.org/10.1039/C8SC04228D
An evolutionary algorithm for the discovery of porous organic cages
An evolutionary algorithm is developed and used to search for shape persistent porous organic cages.

Chem. Sci., 2018,9, 8513-8527
https://doi.org/10.1039/C8SC03560A
Chimera: enabling hierarchy based multi-objective optimization for self-driving laboratories
Chimera enables multi-target optimization for experimentation or expensive computations, where evaluations are the limiting factor.
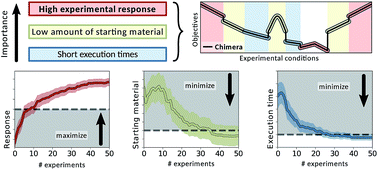
Chem. Sci., 2018,9, 7642-7655
https://doi.org/10.1039/C8SC02239A
Computer-aided design of metal chalcohalide semiconductors: from chemical composition to crystal structure
The standard paradigm in computational materials science is INPUT: STRUCTURE; OUTPUT: PROPERTIES, which has yielded many successes but is ill-suited for exploring large areas of chemical and configurational hyperspace.
Chem. Sci., 2018,9, 1022-1030
https://doi.org/10.1039/C7SC03961A
About this collection
In honour of our second Chemical Science symposium titled ‘How can machine learning and autonomy accelerate chemistry?’, being held virtually in September 2020, we have put together a special collection of some of our favourite articles in this area, including a few by some of the speakers and scientific committee of the symposium.