Open Access Article
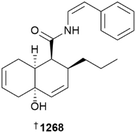
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Marine natural products
Anthony R.
Carroll
 *a,
Brent R.
Copp
*a,
Brent R.
Copp
 b,
Tanja
Grkovic
b,
Tanja
Grkovic
 c,
Robert A.
Keyzers
c,
Robert A.
Keyzers
 d and
Michèle R.
Prinsep
d and
Michèle R.
Prinsep
 e
e
aSchool of Environment and Science, Griffith University, Gold Coast, Australia. E-mail: A.Carroll@griffith.edu.au
bSchool of Chemical Sciences, University of Auckland, Auckland, New Zealand
cNatural Products Branch, Developmental Therapeutics Program, Division of Cancer Treatment and Diagnosis, Molecular Targets Program, Center for Cancer Research, National Cancer Institute, Frederick, MD, USA
dCentre for Biodiscovery, School of Chemical and Physical Sciences, Victoria University of Wellington, Wellington, New Zealand
eSchool of Science, University of Waikato, Hamilton, New Zealand
First published on 19th January 2026
Abstract
Covering: January to the end of December 2024
This review covers the literature published in 2024 for marine natural products (MNPs), with 617 citations (578 for the period January to December 2024) referring to compounds isolated from marine microorganisms and phytoplankton, green, brown and red algae, sponges, cnidarians, bryozoans, molluscs, tunicates, echinoderms, the submerged parts of mangroves and other intertidal plants. The emphasis is on new compounds (1256 in 336 papers for 2024), together with the relevant biological activities, source organisms and country of origin. Pertinent reviews, biosynthetic studies, first syntheses, and syntheses that led to the revision of structures or stereochemistries, have been included. An analysis of the role of artificial intelligence in marine natural products research is discussed.
1 Introduction
This review is of the literature for 2024 and describes 1256 new compounds from 336 papers, compared to 1220 new compounds in 340 papers reported for 2023.1 In addition, 30 known NPs were reported from a marine source for the first time, three artefacts were identified, and 48 known MNPs had their structures revised. Only new MNP structures or previously reported compounds where there has been a structural revision, or a newly established stereochemistry are shown in this review. The review also covers previously reported MNPs with significant new bioactivities or ones that have been synthesised for the first time, but their structures are generally not shown. A † symbol on the identifying diagram number is used to distinguish structures where the absolute configuration has been determined for all stereogenic centres, axes and/or planes in a compound. Reports of new MNPs that were identified based solely on a combination of biosynthetic gene cluster (BGC) information, MS/MS data and/or Global Natural Products Social (GNPS)-based molecular networking, with compounds not isolated and no NMR data recorded, are excluded from the review. Only a selection of highlighted structures (54) is shown. Compound numbers for structures not highlighted in the review are italicised, and all structures are available for viewing, along with their names, taxonomic origins, collection locations, and biological activities, in an associated SI document. Over the past 20 years we have discussed a variety of topics relevant to MNP research in the conclusion section. Given that these treatises introduce new data derived from analysis of historical trends, we feel that it is more appropriate to include this discussion in a stand-alone section rather than the conclusion section. Therefore, this year we introduce a new “Perspective” section to the review, leaving the conclusion section to sum-up the overall insights gained from documenting the years MNP research outputs. Access to the curated MNP data held in the Marinlit database2 provides all the structural and literature data used to prepare this review.A pioneer of research on MNPs from Red Sea organisms, Professor Yoel Kashman from Tel Aviv University, passed away in mid-2025. Over his long career, Prof Kashman made significant discoveries, mostly associated with sponges, ascidians and soft corals. He will be sadly missed.
Trends in the number of new MNPs reported annually over the decade show that numbers of new sponge MNPs dropped again to a new low, while numbers of mollusc, (2) tunicate (6) and red algae MNPs (3) also dropped to decadal lows and numbers of reported cnidarian MNPs dropped to a near decadal low. No new bryozoan NPs were reported for the second year in a row and the number of new brown algae MNPs was lower than the decadal average. In contrast, numbers of MNPs isolated from fungi continue to rise and based on our changed reporting criteria, that now only includes fungi isolated from submerged environments, it is likely that 2024 had the highest number of new fungal MNPs reported over the 42-year history of this review. New MNPs reported from bacteria resurged to a decadal average level (Fig. 1).
 | ||
| Fig. 1 Trends in new MNPs. The bars represent the total number of new MNPs reported each year over the last 10 years. Fungi MNPs only includes those isolated from submerged sources. | ||
2 Marine microorganisms and phytoplankton
2.1 Marine-sourced bacteria
Actinobacteria continue to be the most prolific source of MNPs reported from bacteria (with 165 new structures), even though they represent approximately 10% of total marine bacterial biodiversity. A sediment-derived Actinomadura sp. yielded five phenyl polyenes, maduraflavacins A–E 1–5.3 Two rare genera of bacteria yielded MNPs for the first time in 2024, linear peptides cellulamides A 6 and B 7 were reported from Cellulosimicrobium funkei,4 and glycoglycerolipids testacosides A–D 8–11 were the first MNPs reported from Microbacterium testaceum.5 A sponge-derived Microbispora sp. strain yielded okichromanone, a chromanone NP isolated as a mixture of two interconverting epimers 12 and 13 due to hemiketal ring opening and recyclisation.6The genus Micromonospora yielded two anthracyclinones 14 and 15 and one anthraquinone 16,7 three nona-2,7-dienoic acid derivatives apocimycins A–C 17–19,8 as well as three spiroketal-containing macrolides 20–22, and a diketopiperazine 23.9 X-ray diffraction (XRD) analysis was used to define the absolute configuration of the co-occurring known MNP IB96212 24 for the first time.9 A large scale (96 L) fermentation of Nocardiopsis maritima yielded two cyclic hexapeptides, maritiamides A 25 and B 26, which showed moderate activity against S. epidermidis and weak activity against E. coli and P. fluorescens.10 Two α-pyrone polyketides, nocardiopyrones D 27 and E 28, as well as an alkaloid, nocarterphenyl I 29, were reported from a sediment-derived Nocardiopsis sp.11 Nocarterphenyl I 29 possesses a rare 2,2′-bithiazole-p-terphenyl scaffold, the structure of which was confirmed via XRD, and showed weak to moderate activity against four bacterial strains with MIC values ranging from 0.8 to 1.6 µM. Nocarterphenyl I showed no cytotoxicity against methicillin-resistant S. aureus (MRSA) or four human tumour cell lines (HTCLs).
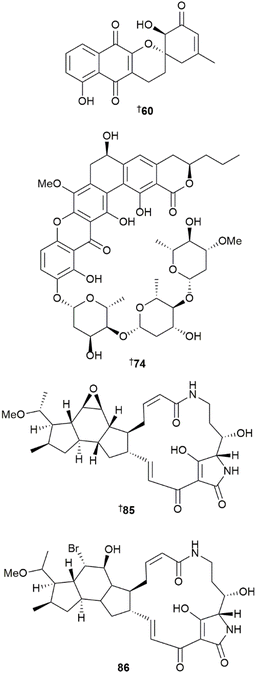
The genus Saccharomonospora yielded four pyridine-, thiazole-, and chloroquinoline-containing alkaloids lodopyridones D–G 30–33,12 and two prenylated indole alkaloids, penipaline D 34 and N-acetyl-6-dimethylallyl-L-tryptophan 35.13 Three sulfur-bridged angucycline dimers, spongisulfins A–C 36, 37 and 38, and one monomer, rubiginone A3 39, were reported from a sponge-derived Spongiactinospora rosea, with the structures of 37 and 39 confirmed via XRD.14
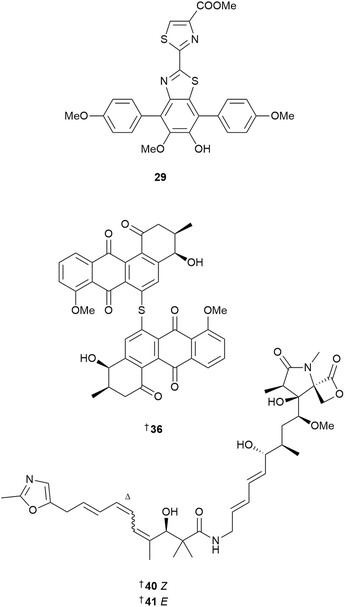
As for previous years, the genus Streptomyces was the most studied, with 125 new MNPs reported. A discovery campaign targeting the identification of terminal oxazole-bearing NPs using PCR screening for oxazole cyclase encoding genes coupled with NMR-based screening measuring 1JCH one-bond coupling constants specific to oxazole rings, identified two polyketide MNPs, methyl-oxazolomycins A 40 and B 41.15 The two compounds showed moderate cytotoxicity against two estrogen receptor (ER) positive breast cancer cell lines MCF7 and T-47D, but no activity against ER-negative cell line MDA-MB-231.
Two linear polyketides, alpiniamides H 42 and I 43 were isolated from a sediment-derived collection of Streptomyces sp.,16 and a coral-derived Streptomyces sp. yielded polyketide chromone 44, polyketide 45 and fatty acids 46 and 47, and 48 and 49, each as racemates but purified to separate each enantiomeric pair.17
A series of methylene-bridged, dimeric, 4-hydroxy-2H-pyran-2-one (HPO)-containing pyrones, the phaeochromycins M–U 50–59 were produced by a large-scale culture of sediment-derived Streptomyces sp.18 The authors showed that simple HPO-containing molecules can react with formaldehyde to generate methylene-bridged dimers, proposing a chemical defence mechanism where HPO-producing bacteria can sequester, and effectively neutralise formaldehyde from the environment for competitive gain. Cloning and heterologous expression of a type II polyketide BGC spi1 sourced from a marine sediment-derived Streptomyces sp. into a S. coelicolor host, led to the isolation and identification of six angucyclines 60, 61–65, including spirocyclinone A 60 with a rare oxaspiro[5,5]undecane motif.19 Madeirone 66, a new mixed terpene- and polyketide-derived MNP, was reported from a sediment-derived Streptomyces aculeoletus.20
Glycosylated angucyclines reported from Streptomyces sp. included seven C-glycosides, chrysomycins F–J 67–71,21 and grincamycins V 72 and U 73.22 A mangrove rhizosphere-derived Streptomyces sp. yielded three glycosylated polycyclic xanthones, kebanmycins A–C 74, 75 and 76, and the aglycone kebanmycin D 77.23 Kebanmycin A showed potent activity against S. aureus, including four MRSA isolates, with MIC values ranging from 0.125 to 0.5 µg mL−1, but did not inhibit B. subtilis growth. Based on the annotations from the genome mining tool antiSMASH, the candidate BGC keb was proposed to be responsible for assembly of the compounds.
Sediment-derived collections of Streptomyces sp. yielded four ansamycin-type macrolides 78–81,24 five tetramate-containing macrolactams, aburatubolactams D–E 82–84,25 as well as hydroxycapsimycin 85 and brokamycin 86.26 Two 6,6-spiroketal-containing MNPs, streptospirodienoic acids D 87 and E 88 were reported from a coral-derived S. cavourensis together with two known compounds bafilomycins P 89, and Q 90 whose absolute configurations were revised based on nOe correlation analysis, DP4+ calculations, and XRD data.27 Compounds 89 and 90 showed moderate activity against the A-549 and HCT-116 HTCLs. Five oligomycins 91–95 were reported from a sediment-derived Streptomyces sp.,28 four polyene macrolides 96–99, were isolated from a mangrove-derived S. hiroshimensis,29 and a sponge-derived Streptomyces sp. yielded a tricyclic polyene macrolactam, weddellamycin 100.30
Albusamides A–G, 101–107, a series of new long-chain hydroxylated acetamide MNPs, were reported from S. albus,31 and a sediment-derived collection of Streptomyces sp. yielded additional acetamide-containing MNPs, streptothiomycin F 108 and N-(5-nitropentyl)acetamide 109, the latter reported as a first time MNP.32 Albusamide C 103 displayed moderate cytotoxicity against a panel of fourteen HTCLs, but with no selectivity margin against the normal B lymphocyte cell line RPMI-1788. A genome-based mining strategy targeting the presence of the carpatamide BCG ctd, identified five acylated arylamines, carpatamides I–M 110–114 from S. parvus.33
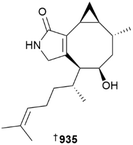
A series of shorter chain aryl amines 115–119, a phenylpropanoid 120, unsaturated fatty acids 121, 122, and two thiazinone-containing MNPs 123, 124 were reported from a sediment-derived Streptomyces sp., with 119 and 121–124 representing new MNPs and others, known compounds reported from the marine environment for the first time.34 A mudflat-derived Streptomyces sp. yielded a phenylthiazole anithiactin D 125,35 and heterologous expression of an NRPS BGC grsc, sourced from a marine sediment-derived Streptomyces sp. into three different Streptomyces surrogate hosts, yielded eight new thiazole-containing MNPs, grisechelins E 126, F 127, and I–N 128–133, as well as two quinoline derivatives, grisechelins G 134 and H 135.36 Notably, grisechelin E 127 showed moderate antimycobacterial activity, with an MIC of 8 µg mL−1 against M. tuberculosis.
A pyrrolo-pyrido-benzoquinone, streptoquinoneazaindole A 136, a urea-containing pyrrole alkaloid, streptoureipyrrole A 137 and 6-hydroxybenzothiazole 138, reported as a MNP for the first time, were isolated from a deep-sea sediment-derived Streptomyces sp.37 A thioether-linked quinoline-quinazoline, quinosumycin 139 and a chromone, chromycone 140, were reported from a sponge-derived S. diastaticus.38 A dithiolopyrrolone, 141 was isolated from a tunicate-derived Streptomyces sp.39 and a staurosporine analogue, streptomholyrine A 142 was reported from a sediment-derived Streptomyces sp.40
Pattern-based genome mining, combining mass spectrometry (MS) isotopic signature and biosynthetic sequence data, led to the identification of two chlorinated pyrroloketoindanes, indanopyrroles A 143 and B 144, with their candidate BGC idp identified and biosynthetic assembly proposed.41 The trichlorinated indanopyrrole A showed moderate to potent activity against several Gram-positive and -negative bacteria, while the dichlorinated analogue indanopyrrole A was inactive.
In a high-throughput screening (HTS) discovery campaign against Mycobacterium avium complex (MAC) pathogens, seven disaccharide pyrimidine nucleosides, mavintramycins A–G 145, 146–151 were identified from a sediment-derived Streptomyces sp.42 Mavintramycin A 145 showed potent activity against two different MAC strains, with only moderate activity against two TB-associated mycobacterial strains and no cytotoxicity against other bacterial and fungal pathogens and one mammalian cell line. Moreover, 145 was found to be active against 40 clinically isolated M. avium strains, including those that were multidrug-resistant. Mechanism of action studies found that mavintramycin A binds to 23S ribosomal DNA and inhibits protein synthesis. The structure of 145 was later unequivocally confirmed via total synthesis.43
A Korean sediment-derived Streptomyces sp. yielded six linear depsipeptides, homiamides A–C 152–154,44 and cavomycins A–C 155–157,45 all of which contained a core amino acid sequence of α-hydroxyisovaleric acid (Hiv)-valine (Val)-lactic acid (Lac)-valine (Val) residues like the valinomycin family of cyclic peptides. Three linear lipopeptides, albubactins A–C 158–160 were isolated from a tunicate-derived S. albidoflavus,46 with the structure of 158 confirmed via total synthesis. A sediment-derived Streptomyces sp. yielded N-acylated cyclic octapeptides, acyl-surugamides A1–A4 161–164,47 with the structure of acyl-surugamide A2 162 also reported, but not fully characterised, from a tunicate-derived S. albidoflavus.48 A sponge-derived Streptomyces sp. yielded four glycosylated cyclopeptides, pyridapeptides F–I 165–168,49 and three cyclic lipopeptides olenamidonins A–C 169–171 were reported from a deep-sea derived S. olivaceus.50 Six terpenoid structures reported from Streptomyces spp. included the iridoid lucknolide A 172, reported as a MNP for the first time,51 and five labdanes, chlorolabdans A–C 173–175, and epoxylabdans A 176 and B 177.52
Only one MNP from the phylum Bacteroidota was reported in 2024; an alga-derived Algoriphagus sp. strain yielded a new carotenoid, 2-hydroxyflexixanthin 178.53 Five MNPs were identified from the phylum Fimicutes, which included three glycosylated, 24-membered polyene macrolides, amylomacrolactines A–C 179–181 sourced from a sediment-derived Bacillus amyloliquefaciens,54 and a new 25-membered polyene macrolide, macrolactin XY 182 and a new fatty alcohol 183 from a sponge-derived B. subtilis.55
Fourteen MNPs were reported from the phylum Proteobacteria in 2024. A deep-sea sediment-derived Alcanivorax dieselolei strain yielded two p-aminoacetophenonic acid analogues, mohangic acids E 184 and F 185,56 and four lipopeptides, bokeelamides A–D 186–189 were isolated from snail egg mass-derived Ectopseudomonas khazarica.57 Two ureido-containing linear peptides, alteropeptilides A 190 and B 191 were reported from an arthropod-derived Pseudoalteromonas flavipulchra.58 Structures of sulfur-containing amino lipids cysteinolides A 192 and B 193 were reported from bacterial species Ruegeria pomeroyi and R. meonggei and represent the first members of the sulfonolipid structural class of MNPs that have been fully characterised by NMR spectroscopy and their structures confirmed by total synthesis.59 A seawater-derived Vibrio ruber yielded a new pyrrolidine, vibripyrrolidine A 194, a piperazine vibripiperazine A 195, and two diazinanes, vibridiazinanes A 196 and B 197, with 196 showing potent activity against S. aureus with a MIC value of 0.98 µg mL−1 and no inhibitory activity against E. coli, K. pneumoniae, P aeruginosa, or a panel of three HTCLs.60
As in previous years, some MNPs reported from marine bacteria did not have adequate spectrometric or NMR spectroscopic data in full support of the proposed structures and were omitted from this review.61–64 Total synthesis of the reported structure of marinoaziridine B65 revealed significant inconsistencies in the 1H and 13C NMR spectroscopic data, suggesting the structure of this MNP should be revised. Total syntheses of other bacteria-sourced MNPs reported in 2024 included bacillimidazole B and discolins A, B, and E,66 bacilotetrin C,67 dassonmycins A and B,68 caerulomycin K,69 cihunamide B,70 cyclomarin,71 discoipyrroles A–C,72 (+)-dixiamycin,73 homoseongomycin,74 ieodomycins A and B,75 incarnatapeptins A and B,76 iturin A,77 lorneic acids C and D,78 marformycins A and D,79 mavintramycin A,43 moiramide B,80 octalactins A and B,81 rufomycins 1, 2, 4, 11, 22, and 24,82 salimabromide,83 and sealutomicin C.84
Reviews focused on bacterial MNPs published during 2024 included a summary of metagenomic strategies for the discovery of MNPs,85 and actinomycete promoter engineering approaches.86 The bacterium B. subtilis was reviewed as a versatile host for heterologous expression of NP BGCs,87 as was the regulation of acyl homoserine lactone-based quorum sensing effects on the biosynthesis of bioactive molecules in bacteria.88
Structural classes of bacterial NPs reviewed included polyacteylenes,89 alkaloids,90 and γ-pyrones.91 Groups of marine bacteria reviewed for biosynthetic potential and the diversity and activity of MNPs produced included the MAR4 clade of the genus Streptomyces,92 and marine-sourced Pseudomonas sp.93 Biological activity studies of previously reported bacteria-sourced MNP structures included in vitro and in vivo anti-Toxoplasma gondii activity assessment of a series of marinoquinolines,94 mechanism of action (MoA) studies of marinopyrrole, pentachloropseudilin, and pentabromopseudilin,95 tartrolon D,96 and tunicamycins,97 and structure–activity relationship (SAR) reports of thiomarinol A,98 and sunshinamide.99
2.2 Cyanobacteria
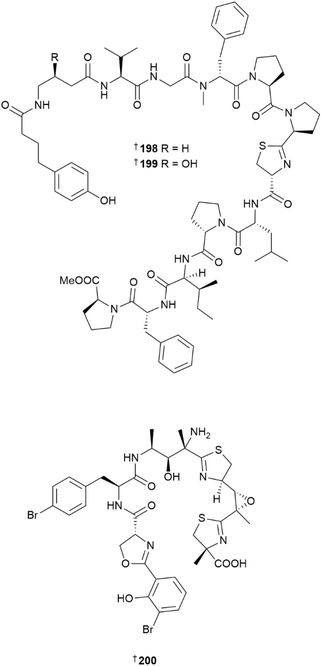
Two linear lipopeptides, pemuchiamides A 198 and B 199 were isolated from a relatively large-scale (3 kg) Japanese collection of the filamentous cyanobacterium Hormoscilla sp.100 The compounds contain 13 amino acid residues including four prolines and several unusual ones such as 4-aminobutanoic acid, 4-(4-hydroxyphenyl)butanoic acid and N-methyl-D-phenylalanine. Pemuchiamide A showed moderate activity against Tryponosoma brucei rhodesiense (IC50 = 0.63 µM), while the hydroxylated analogue pemuchiamide B was an order of magnitude less active. This indicated that the 4-aminobutanoic acid moiety may be an important pharmacophore feature involved in the antitrypanosomal activity for this peptide series. Laboratory cultures (some up to six months in length) of three geographically dispersed collections of Lepthotoe strains yielded three new PKS/NRPS hybrid linear peptide metallophones, leptochelins A–C 200, 201 and 202.101 The leptochelin structures were challenging to elucidate since they contain modified amino acids and isolated stereogenic centres. Their structures were deduced using a combination of tandem mass spectrometry, NMR spectroscopy, and bioinformatic analyses of the putative leptochelin BGC lec. The leptochelins bind to zinc, copper, iron, and cobalt and were moderately cytotoxic against four HTCLs. They are also the first reported MNPs from the genus Lepthotoe.Studies on Moorena spp. yielded seven MNPs. A glycosylated macrolide, moorenaside 203 was reported from a Florida collection of Moorena sp.102 and a Guam-sourced collection yielded two polyketides 204 and 205.103 A co-culture of M. producens with Candida albicans led to significant upregulation of the production of a linear depsipeptide hectoramide B 206. Its putative BGC hca was identified and its biosynthetic assembly annotated.104 Even though the presence of a fungal competitor was crucial to produce hectoramide in liquid culture, when tested against two Candida strains, 206 did not show any antifungal activity. M. bouillonii collected in India, yielded a linear lipodepsipeptide, kavaratamide A 207,105 and a cyclic depsipeptide, wajeepeptin 208 was isolated from a Japanese collection of Moorena sp.106 A benthic collection of Moorena sp. (annotated as Lyngbya sp.) in Indonesia yielded a cyclic peptide PM170453 209, the structure and absolute configuration of which was unequivocally confirmed via total synthesis.107 Six lipopeptides, noducyclamides A1–A4 210–213 and noducyclamides B1 214 and B2 215, were reported from a laboratory culture of Nodularia sp.108 The dodecapeptide noducyclamide B1 was weakly cytotoxic against the MCF7 breast cancer cell line.
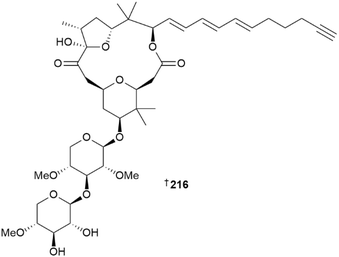
The genus Okeania yielded six new MNPs. A glycosylated, 16-membered macrolide, polycavernoside E 216, was reported from a coastal Okinawan collection of Okeania sp.109 Related analogues of this MNP, the polycavernosides A–C were originally reported from the red alga Polycavernosa tsudaii110–112 and were implicated in human illness and death from consumption of this edible seaweed in Guam. An additional congener, polycavernoside D, was later identified from a Caribbean collection of the cyanobacterium Okeania sp.,113 and this latest report of 216 provides increasing evidence that polycavernosides are produced by cyanobacteria rather than red algae. Polycavernoside D did not show significant in vitro cytotoxicity against normal human fibroblast WI-38 cells, was weakly active against T. b. rhodesiense,109 but demonstrated potent in vivo toxicity in mice with an ip. lethal dose of 0.81 mg kg−1.114 Additional Okeania sp. collections yielded a lipopeptide, N-desmethylmajusculamide B 217,115 cyclic peptide okeaniazole A 218,116 and a linear lipopeptide, amantamide C 219.117 Terpenoids continue to be reported from cyanobacterial sources, with kagimminols A 220 and B 221 representing the first examples of cembrene diterpenoids isolated from the genus Okeania.118
An unidentified cyanobacterium with morphological resemblance to Symploca sp. collected in Guam, yielded three enyne-containing chlorinated fatty acid amides, taveuniamides L–N 222–224 and the absolute configuration of the C-8 stereogenic centre was reported for this MNP class for the first time.119 Using genome-mining and metabolomic strain prioritization strategies, two dialkylresorcinol polyketides, 29-dechloro-29-hydroxybartoloside A 225 and 17-dechloro-17-hydroxybartoloside A 226, were isolated from Synechocystis salina,120 with hydroxylation along the aliphatic side-chain in both compounds representing a novel structural modification of the bartoloside family of polyketides. Finally, a methyl-branched fatty acid, jobosic acid 227 was reported from an unidentified bacterial biomass collected in Puerto Rico.121 Cyanobacteria remain one of the few marine taxa where reports of new MNPs from mixed assemblages or incomplete taxonomic identifications continue to be accepted for publication.
The total synthesis of the cyclic depsipeptide lagunamide C was achieved,122 resulting in the revision of the proposed structure from a 27-membered macrocycle originally published123 to the 26-membered odoamide,124 a known cyanobacterial MNP. Other cyanobacterial MNPs with reported total syntheses included dragocins A–C,125 irijimaside A,126 jamaicamide B,127 majusculamide D,128 rivularin A,129 serinolamides A and B,130 and trichophycin-I.131
Reviews included summaries of anti-infective,132 antiprotozoal,133 and cancer-related134 activities of cyanobacterial MNPs, as well as a comprehensive review of the diversity of NP structures and biological activities reported from the genus Oscillatoria.135 Notable work on the biosynthesis of cyanobacterial MNPs included the identification and characterisation of the putative BGC for the assembly of dolastatin, reported from Caldora penicillata,136 and SAR studies of known cyanobacterial compounds biselyngbyolide B,137 carmaphycin B138 iezoside,139 and majusculamide D.140 Environmental reports of cyanobacterial MNPs with broad implications to human health included a review on impacts of cyanobacterial blooms on fish farming,141 and the first account of bioaccumulation of lyngbyatoxin A in edible shellfish along the coast of New Zealand.142 This, together with a report of acute toxicity of a new polycavernoside analogue 216 in mice, accentuates the importance of accurate annotation, full structural characterisation, and monitoring of the occurrence of cyanobacterial MNPs that can impact marine ecosystems and human health.
2.3 Marine-sourced fungi
Co-culture of an Acremonium species with heat-killed Pseudomonas aeruginosa, resulted in isolation of acremosides A–G 228–234, linear sesquiterpenoids with a 2-phenyl acetoxy substituent conjugated to a sugar alcohol.143 Other Acremonium strains were the source of the meroterpenoids, acremochlorins A 235 and B 236, pyridine alkaloid acremopyridone A 237 and cyclopentenone derivative acremoketene A 238,144 sorbicillin-like compound 239 and acremonilactone 240,145 and ascochlorin glycoside 241.146Fermentation of Albifimbria verrucaria yielded the nonenes, verrucanonenes A–C,147242–244 and tetrahydrofuran-containing polyketide derivatives148245–248, the butanolide derivative, alterbutenolide 249 was obtained from Alternaria alternata,149 and Arthrinium arundinis cultures, respectively derived from a sea anemone and a sponge, yielded arthrinoic acid 250, hexylaconitic anhydride methyl ester 251, nonanoic acid derivative 252, arthripenoids G 253 and H 254,150 and alternapyrone derivatives, alternapyrones G 255 and H 256. The BGC for alternapyrone biosynthesis was identified and heterologous expression led to isolation of an additional alternapyrone, alterpyranone I 257.151
As has been the case in many previous years, the genus Aspergillus was the most common source of fungal metabolites. Co-culture of deep-sea-derived Aspergillus aculeatinus with a mangrove-derived Penicillium strain yielded the paraherquamides, aculeaquamides A 258 and B 259,152 while addition of sodium bromide to growth media of A. alliaceus resulted in production of brominated anthraquinone monomer 260 and dimers 261–263.153p-Terphenyl 264, 265 and diphenyl 266, 267 ether derivatives were obtained from A. candidus,154 whilst the benzophenone derivatives, carneusones A–F 268–273 were isolated from a culture of A. carneus.155A. chevalieri and A. fischeri respectively yielded prenylated indole diketopiperazine 274 and prenylated indole derivative 275,156,157 ascidian-derived A. clavatus was the source of 12S-deoxynortryptoquivaline 276,158 co-culture of A. insulacola with a mangrove-derived Alternaria strain led to isolation of cyclic tetrapeptides, violaceotides B–E, 277–280,159 miniaturised cultivation profiling enabled isolation of 2,6-diketopiperazines, noonazines A–C 281–283, and azaphilone, noonaphilone A 284,160 and a nitrobenzoyl sesquiterpenoid 285 was obtained from culture of A ochraceopetaliformis.161
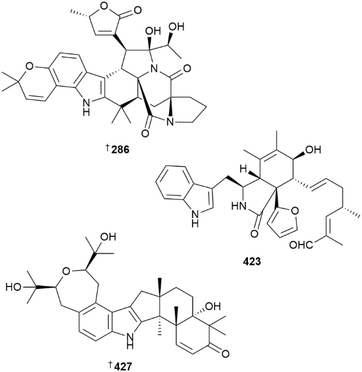
Stephaochratidin A 286, an unusual stephacidin-asperochratide hybrid, was isolated from culture of deep-sea-derived A. ochraceus.162 Several deep-sea sediment-derived Aspergillus strains yielded a range of metabolites. The sesquiterpenoid malfilanol C 287 was isolated from a culture of A. puniceus, along with known terrestrial but new MNP, malifilanol B 288,163 phenol derivatives 289–298, (the last a known synthetic compound but new NP), were obtained from A. subversicolor,164 and separate cultures of A. sydowii yielded sesquiterpenoid 299 and sydowimide A 300,165,166 the latter by epigenetic modification. Another culture of A. sydowii was the source of xanthone-alkaloids sydoxanthones F–M 301–311,167 and a prenylated indole alkaloid, aspertaichamide A 312 was obtained from A. taichungensis.168
Cultures of A. terreus were the source of a range of metabolites including caryophyllene-type sesquiterpenoids, asperporonins A 313 and B 314,169 ergone derivatives 315–324,170 (all deep-sea-derived), sulfur containing metabolites asperteretals L 325 and M 326 and terreins A 327 and B 328,171 brominated aromatic butenolides 329–342,172 meroterpenoids 343–348,173 butenolide derivatives 349 and 350 (via epigenetic manipulation and the latter a known synthetic compound but new NP)174 and a suite of azaphilones, 351–361, which were produced through activation of a cryptic BGC.175
Some polyketides 362–365, and a nucleoside 366 were isolated from a culture of A. versicolor,176 while a range of metabolites were obtained from Aspergillus strains from a variety of sources. Sponge-derived Aspergillus strains yielded ergostane-type steroid 367,177 terminal olefin-containing unsaturated fatty acid 368, 2-carboxy-2′-methyl-azobenzene 369 and 1,3,9-trimethyluric acid 370 (the last two known synthetic compounds but new NPs).178 Sediment-derived Aspergillus strains yielded aspergilalkaloid A 371,179 asperindopiperazines A–C 372–374, 5-methoxy-8,9-dihydroxy-8,9-deoxyaspyrone 375, and 12S-aspertetranone D 376,180 emericelactones F 377 and G 378, preshamixanthones 379 and 380, 25-O-methylarugosin A enantiomers 381 and 382, furanones, 9-hydroxymicroperfuranone 383 and microperfuranone 384, (these last four known terrestrial but new MNPs), phthalimidinic acids 385 and 386, aspergilol G 387, anthraquinone 388, and propionic acid lactone derivative 389,181 (the last a known synthetic compound but new NP). A seawater strain of Aspergillus yielded ceramide 390 while a coral-derived Aspergillus strain yielded megastigmanones A–C 391–393 and prenylterphenyllin H 394 and a starfish-derived strain yielded brominated isocoumarin 395.182–184 The production of 395 resulted from addition of bromide salts to the culture medium.184Aspergillus species from mangrove sediments have yielded a nor-diterpenoid acid, asperbrunneo acid 396,185 indole alkaloids asperdinones A–H 397–404,186 isoprenyl phenol 405,187 and alkyl resorcinols 406–408.188
Culture of Aureobasidium melanogenum yielded aliphatic δ-lactones 409–411 and fatty acid methyl esters 412–414,189 sterol 415 was isolated from a culture of Beauveria sp.190 and triterpenoid 416 was obtained from shellfish-derived Ceriporia lacerata.191 Sesterterpenoids with a 5/8/6/5 tetracyclic ring system, sesterchaetins A 417 and B 418 and diepoxide polyketides 419 and 420, were isolated from deep sea sediment-derived Chaetomium globosum,192 while sponge-derived C. globosum yielded six new cyctochalasans, marcytoglobosins A 421 and B 422 and marchaetoglobins A–D 423, 424–426, of which 423 is the first furan-containing cytochalasin.193,194 Asepterpenedol A 427, an indole sesquiterpenoid obtained from culture of a Chloridium strain, contains a seven-membered ether ring as part of a 7/6/5/5/6/6 hexacyclic framework.195
Cladosporium strains were the source of a range of new metabolites. Dodecanoic acid derivatives 428–431 (the last two known as semi-synthetic compounds) were isolated from a culture of C. cladosporioides,196C. halotolerans yielded pyrone derivatives 432 and 433,197 drimane sesquiterpenoids 434, isocoumarin 435, coumarin 436 and known synthetic compound, 9-formylfluorene 437 as a new NP,198 and a further Cladosporium strain was the source of isochromanone 438.199Curvularia aeria and C. lunata respectively yielded tricyclic aromatic polyketide 439 (as a racemic mixture) and resorcylic acid lactones 440–443 and isolation of curdepsidone A from a Curvularia strain led to revision of the structure to 444.200–202
A one strain many compound (OSMAC) approach to cultivation of Emericellopsis maritima resulted in isolation of eremophilane sesquiterpenoids 445–447,203 6/7/6 tricyclic diterpenoid eutyditerpenoid A 448 and cytochalasin 449 were obtained from a culture of Eutypella scoparia,204 whilst another Eutypella strain yielded fifteen new sesquiterpenoids, eutypelides A–O 450–464.205 Sesquiterpenoids, including eremophilane derivatives fureremophilanes A–D 465–468 and acorane analogues furacoranes A 469 and B 470, were isolated from culture of Furcasterigmium furcatum.206Fusarium strains were the source of fusarochromanone derivatives 471 and 472, hydroxyphenylacetic acid 473–476 and hydroxyphenylethanol derivatives 477 and polyketide 478.207–209 The cyclic pentapeptides, avellanins D–O 479–490 were obtained from culture of Hamigera ingelheimenensis.210 An Irpex strain produced the heptaketides irpetones A 491 and B 492,211 catechol derivatives, meirols A–C 493–495 were obtained by fermentation of a Meira strain,212 whilst a Microascus strain yielded eight decahydrofluorene alkaloids, 496–503.213 Azaphilones 504–506 and a dihydroisocoumarin 507 were isolated from a strain of Neopestalotiopsis,214 mangrove sediment-derived Nigrospora oryzae contained four aniline derivatives 508–511,215 spiro trione compound 512 and asparvenone derivatives 513–516 were obtained from a culture of Paraconiothyrium sporulosum,216 and a culture of Parengyodontium album was the source of aromatic polyketide alternaphenol B2 517.217
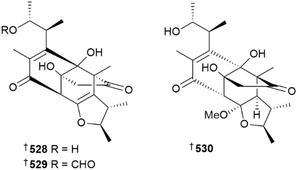
The Penicillium genus has proven once again to be a prolific source of new metabolites. A phenyl 6,7-dihydroxygeranyl ether derivative 518 was obtained from a P. arabicum culture,218P. brasilianum yielded diketopiperazine alkaloids 519 and 520,219 soft coral-derived strains of P. chrysogenum were the source of aromatic heterocycles 521–525, and a sorbicillinoid 526,220 and of anthranilic acid-peptide derivative 527.221 Dicitrinols A–C 528–530, citrinin derivatives with a 6/5/7/5 core, were obtained from P. citrinum derived from hydrothermal vent sediment,222 a culture of P. corylophilum yielded enantiomeric hydroxyphenylacetic acid derivatives 531–534 and α-pyrone analogue 535,223 and co-culture of deep-sea sediment-derived P. crustosum with mangrove-derived P. citrinum resulted in production of the dihydropyrones, rhytismatones C 536, and D 537.224
The relative and absolute configuration of the known indolone, notoamide X 538 was assigned for the first time. It was obtained from the mangrove sediment-derived P. janthinellum along with three other indolones with proposed new structures. However, the structures of these additional compounds were only proposed based on MS data and so are not included here.225 Culture of P. oxalicum derived from a red alga yielded prenylated indole alkaloids 539 and 540 and polyoxygenated steroid 541,226,227 sediment-derived strains, P. pancosmium and P. rubens respectively yielded meroterpenoids 542 and 543 (P. pancosmium)228 and polyketide derivatives 544 and 545 and amino-bis-tetrahydrofuran derivatives 546 and 547 (P. rubens),229 while a coral-derived P. sclerotium culture was the source of azaphilone pigments 548–559.230 Highly oxygenated meroterpenoids, penisimplinoids A–K 560–570, were isolated from a P. simplicissimum culture,231 steroid 571 and polyketides 572 and 573 were obtained from P. variabile232 and seagrass-derived cultures of P. velutinum and P. yezoense respectively yielded piperazine derivative 574,233 and decalin polyketides 575–584.234 Meroterpenoids, including 585–591 and penicianstinoids F–K 592–597, indole diterpenoids, including 598–604 and penpaxilloids F 605 and G 606 and diketopiperazine alkaloid 607 (the last as a racemate), were isolated from cultures of various Penicillium strains.235–239 Sediment-derived strains were the source of azaphilones 608–612 and clavatol derivative 613,240,241 and Penicillium strains derived from seawater yielded racemic polyketides 614 and 615 and enantiomeric hydroxyphenylacetic acid derivatives 616 and 617,242,243 which were separated by chiral chromatography. Cultures of various Penicillium strains also resulted in production of racemic phenalenone 618,244 quinolone-citrinin hybrids 619 and 620 and speradines I 621 and J 622.245,246
Mangrove sediment or root-derived Penicillium spp. have also yielded pyridyl derivatives 623 and 624, indole 625,247 isobenzofuranone derivative 626, α-pyrone 627, quinone 628, phenol derivative 629,248 steroids 630 and 631, benzopyran 632,249 diterpenoid indole alkaloid derivatives 633 and 634,250 sesquiterpenoids eupenicisirenins C–O 635–647,251 aryl and decalin polyketide acids 648–669,252 and alkyl resorcinols 670–673.253
Peniotrinins A–F, comprising cytochalasin derivatives 674 and 675, citrinin derivatives 676–678 and tetramic acid derivative 679, were isolated from a culture of a Peniophora strain254 and merocytochalasans, perochalasin A 680, and inseparable C20 epimeric pairs 681 and 682 were obtained from a Peroneutypa species. The BGC responsible for cytochalasin biosynthesis was identified through full genome sequencing and the biosynthesis elucidated through feeding experiments utilising 15N and deuterium labelled hydroxylamine.255Pestalotiopsis strains were the source of rearranged pimarane diterpenoid 683 and known terrestrial but new MNPs, nodulisporenones A 684 and B 685,256 and polyketide derivatives, pestalotiopols E–J 686–691 (the last two as a racemic mixture).257 Cultures of deep-sea sediment-derived Phomopsis strains yielded benzoic acid derivative 692 and 3-carboxy-indole derivatives 693–695.258,259 Co-culture of two Phomopsis strains obtained from the roots of the Chinese mangrove Rhizophora mangle yielded pyrazine 696, steroid 697 and α-pyrone 698 (from P. asparagi)260 and bis-chromones phomoxanthones L–N 699–701 (from Phomopsis sp.),261 none of which were produced in monoculture of the individual strains.
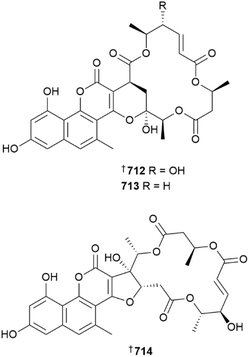
Sediment-derived fungal strains have yielded the sesquiterpenoids 702–711 (from Pseudallescheria boydii)262 naphthopyrone-macrolide hybrids, gymnoasins A–C 712–714 and glucolipid 715 (from Pseudogymnoascus strains) and sesquiterpenoids 716 and 717 (from Pyrrhoderma noxium).263–265 Isolated from the Antarctic, the gymnoasins A–C 712–714 are the first naturally occurring compounds of their type and a biomimetic synthesis of 712 was achieved.263 Deep-sea sediment-derived fungal strains were the source of bisabolene sesquiterpenoid 718 (from Retroconis fusiformis),266 sesquiterpenoid-hydroquinone hybrids, saromacrophorins A–C 719–721 (from Sarocladium terricola)267 and 4-hydroxyphenyl derivative 722 (from Scedosporium apiospermum), which displayed moderate to potent activity against a panel of Candida auris strains,268 while mangrove sediment-derived Roussoella sp. contained two sesquiterpenoids 723 and 724 and a shikimate 725.269
Dihydroisocoumarins, exserolides L 726 and M 727 were obtained from culture of a Setosphaeria species,270 while various Simplicillium strains yielded a range of metabolites including nortriterpenoids 728–730,271 peptaibiotics 731–741,272 dipeptide 742 and acylated valine derivative, siamysin 743 (the last discovered via heterologous expression of a cryptic BGC).273,274 Eleven brominated depsidones 744–754 were isolated from a culture of Spiromastix sp.,275Stachybotrys strains yielded atranones V–Z 755–759 and dolabellane diterpenoids 760–762,276 and phenylspirodrimanes, stachybotrins K 763 and L, 764,277 and cyclic tetrapeptides, endolides E 765 and F 766 were obtained from a Stachylidium strain. The endolides contain the unusual amino acid, N-methyl-3-(3-furyl)-alanine which was used in fragment pattern searches of tandem mass spectrometry data, in conjunction with molecular networking, to assist both isolation and structural elucidation of these metabolites.278
Culture of Talaromyces strains isolated from sediment have yielded a diverse array of metabolites, including oligophenalenone dimers 767–775,279 phenylhydrazone alkaloids 776–779,280 highly oxygenated seco-terpenoids 780–785,281 macrolactin 786,282 highly oxygenated phenol derivatives 787–789, isocoumarin glycosides 790 and 791 and naphthalene glycoside 792.283 Culture of T. purpureogenus yielded six prenylated indole diketopiperazine alkaloids, talaromyines A–F 793–798,284 cyclic heptapeptides 799 and 800, nor-diterpenoids 801 and 802, diterpenoid acids 803 and 804 and triterpenoid 805 were obtained from cultivation of T. scorteus,285T. siglerae yielded cyclic heptapeptides, talaromides A–C 806–808 and T. variabilis was the source of hybrid pentaketide sesquiterpenoids 809–811.286,287 Seawater-derived Talaromyces strains yielded meroterpenoids 812–814 and benzoquinone 815 and furopyridinone 816 derivatives,288,289 a strain derived from sediment was the source of ring-opened azaphilone derivative 817,290 thirteen new polyketide derivatives 818–830, (6-hydroxy-4-methoxycoumarin 829 a known synthetic compound but new NP) were obtained from a deep-sea mussel-derived strain,291 and duclauxin analogues, taladuxins A–N 831–844 were isolated from a culture of a coral-derived Talaromyces strain from which verruculosin B was also isolated and the configuration revised to 1S as in 845.292 Investigations of mangrove sediment-derived Talaromyces spp. have resulted in isolation of γ-lactams, talaroilactams A–D 846–849,293 sesquiterpenoids, talaroterpenoids A–D 850–853,294p-terphenyl derivatives, talaroterphenyls A–D 854–857,295 diketopiperazines, 858–860, a cyclopentenone 861, an α-pyrone 862,296 and linear polyketides 863–868.297 Two of the talaroterphenyls 855 and 856 potently inhibited the enzyme phosphodiesterase-4 (PDE4) linked to inflammation.
Tetramic acid alkaloids, tolypyridones I 869 and J 870 were isolated from a culture of Tolypocladium cylindrosporum,298Trichoderma strains were the source of sesquiterpenoid derivatives 871–874,299 bergamotene-derived sesquiterpenoids 875–882, (through heterologous expression of a cryptic BGC),300 cyclonerane sesquiterpenoid 883,301 γ-butyrolactones 884–889 and monomeric sorbicillinoids 890–893.302,303 Steroid derivatives 894 and 895 and linear polyketide 896 have been isolated from mangrove sediment-derived Trichoderma spp.304,305 and culture of Xylaria acuta yielded a range of metabolites; lactones 897, 898, 899, glucopyranosides 900 and 901 and mannopyranoside 902, epoxycytochalasins 903 and 904 and butylitaconic acid 905.306
Total synthesis of acremolides A and B has been achieved and the absolute configurations established as 906 and 907,307 while asymmetric syntheses of aspilactonol F and aspiketolactonol were achieved in eight and ten steps respectively and their absolute configurations determined as 908 and 909.308 (±)-Notoamide N was prepared in a biomimetic approach via synthesis of (±)-stephacidin A,309 and several prenylated indole alkaloids of the asperversiamide family, namely asperversiamides B, D, E, G and J and dihydrocarneamide A, were prepared in biogenetically patterned total syntheses. The structure originally proposed for asperversiamide A was shown to be incorrect with a revised structure proposed but not yet proven.310 A unified approach was utilised to prepare cyclic tetrapeptide JM-47,311 cyclocondensation of an electrophilic homoserine lactone was employed in the total syntheses of polonimides A and C312 and synthesis of bipenicilisorin was achieved through initial preparation of its monomer penicilisorin and the absolute configuration was established as 910.313 Penostatin D was prepared in an asymmetric synthesis from L-ascorbic acid,314 12β-hydroxy conidiogenone C was also obtained in an asymmetric total synthesis315 and total synthesis of the cyclic peptide, talarolide A showed that it (and by inference, talarolides C and D), occur naturally as atropisomers.316 Racemic total synthesis of phomopsol B has been reported317 and syntheses of asperisocoumarin A and fusarimatin C were achieved from commercially available starting materials.318
Alternariol monomethyl ether was shown to relieve food allergy symptoms in mice and reduce histamine release into blood serum,319 the verticillin analogue, leptosin A showed potent inhibition of the parasite Toxoplasma gondii,320 brevianamide F exhibited antithrombotic activity in a zebrafish model321 and agonodepside B showed potential as a photoprotection agent.322 Fumetremorgin derivatives 12R,13S-dihydroxyfumitremorgin C and tryprostatin A displayed anti-hypercholesterolemia activity in an in vivo model,323 terphenyl HN-001 was shown to attenuate obesity in mice through induction of thermogenesis in adipose tissue324 and to suppress lipotoxicity from metabolic-associated fatty liver disease through inhibition of phospholipase A2 (PLA2)325 and linoleic acid ameliorated metabolic dysfunction-associated steatotic liver disease in mice by targeting fatty acid binding protein 4.326 The mechanism underlying the antithrombotic activity of the peptide isaridin E was shown to be associated with inhibition of von Willebrand Factor secretion from activated platelets and this leads to protection against sepsis,327 butyrolactone-I alleviated intestinal heat stress in vivo through inhibition of two key enzymes which are upregulated in heat stress328 and pannorin was shown to be a selective, if relatively weak (IC50 1.73 µM) monoamine oxidase inhibitor.329
The BGC responsible for the biosynthesis of the quinolone alkaloids, the asperalins in Aspergillus alabamensis, was identified and the biosynthesis elucidated through heterologous expression.330
‘Omics’-based methods for isolation and cultivation of marine fungi have been reviewed331 as have genome mining and biosynthetic pathways of biologically active marine fungal metabolites.332
2.4 Dinoflagellates
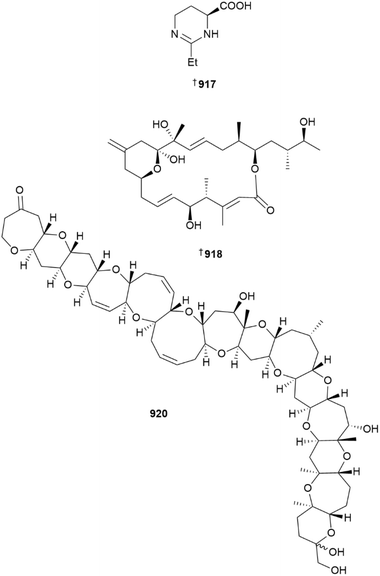
The number of new compounds reported from dinoflagellates and other microalgae is slightly lower than the decadal average.1Prorocentrum lima was the source of polyketides prorocentin-5 911 and three okadaic acid derivatives 912–914, although surprisingly, the latter compounds were not assessed for bioactivity.333,334 An indole methyl ether 915 and a biaryl ester 916 were reported from a Vietnamese Aurantiochytrium strain, although 915 is a likely artefact of isolation.335Ectoine plays an important role in controlling osmotic responses in marine planktonic species. Using a hydrophilic interaction liquid chromatography (HILIC) based molecular networking approach, new analogue 2-homoectoine 917 was detected in multiple harmful algal bloom genera including Alexandrium, Karenia, and Prorocentrum. 2-Homoectoine plays a similar osmoadaptive role as archetype ectoine, for the producing strains. The synthesis of 917 was also achieved.336
Other structures were suggested from MS and transcriptomics data but are not shown here.337,338
Although only isolated in 2023, the total synthesis of diatom sex pheromone peptide SIP+ has been accomplished,339 as has the production of seco-steroid gibbosterol A.340 The original structure of iriomoteolide-1a was disproven by synthesis in 2010, but efforts have not revealed the correct constitution of the molecule until now. The corrected structures of iriomoteolides-1a 918 and 1b 919 have now been determined following an extensive integration of DFT calculations, NMR analysis, and total syntheses.341
A summary of the biomedical application of Alexandrium spp. NPs has been published.342 Compound specific reviews focused on amphidinolides,343 azaspiracids,344 and ciguatera poisoning, especially related to ciguatoxins.345 The structure of ciguatoxin-5 920, originally proposed based solely upon MS data, has been confirmed by NMR spectroscopic analysis of only 40 µg of isolated material.346 The ambiguity between synthetic and naturally occurring portimine B has been resolved, in part by use of the new i-HMBC NMR experiment that can differentiate 2JC,Hvs.3JC,H correlations. Differences in the NMR spectra observed were attributed to the formation of a transient hemiaminal hydrate, facilitated by acid modifiers added to aid HPLC separation. The authors advise researchers to be careful when adding acid or base to their chromatography solvents, especially when comparing spectroscopic data to literature values.347
The structure of 44-methylgambierone 921 was revised following extensive NMR analysis. The compound was also found to be unstable in DMSO-d6 used for spectroscopic analysis hence a microstudy was undertaken to determine its stability towards various solvents over the course of eight months. The effects of the solvent chosen for NMR analysis, and potential storage, should be considered before committing an entire sample.348 A study of the accumulation of domoic acid (DA), the causative agent of amnesic shellfish poisoning, in five species of invertebrate including commercially valuable shellfish, has shown that DA concentrates in scallops (Pecten maximus) at levels 20-fold higher than in clams, slippersnails and sea squirts. The Pectenid species showed a distinctive ability to accumulate, transform and distribute DA within their body tissue.349 Transcriptomics analysis of saxitoxin and non-saxitoxin producing species of the prymnesins, super-carbon chain ladder polyethers produced by Prymnesium parvus, are some of the largest known natural products yet their biosynthetic origins have remained elusive until now. The megadalton PKZILLA-1 and -2 polyketide BGC's encode for 140 and 99 enzyme domains, respectively, that match the predicted “pre-prymnesin” linear backbone. A smaller PKZILLA-B1 encodes for a smaller analogue, prymnesin-B1. This works expands the state of knowledge of large biosynthetic genes and enzymatic pathways to important biomolecules.350
3 Green algae
A Chinese Caulerpa racemosa var. turbinata yielded caulespiros A 922 and B 923, with neither being active against various bacterial strains.351 Mebamamide C 924 was discovered from a Japanese specimen of Bryopsis, but also from the algal gastropod predator Elysia marginata.352Two notable reviews of green algal metabolites were published in 2024. One focused on the allelopathic compounds from Ulva prolifera, the algal species that creates “green tides”,353 while the other summarised the metabolites and bioactivities of compounds from Chlorella vulgaris.354
4 Brown algae
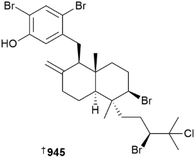
Glyceroether 925 and polyunsaturated glycolipid 926 metabolites were reported from Sargassum sp. (China) and Hizikia fujiformis (South Korea), respectively.355,356 Macrocarquinoid J 927 and sargasilols J–N 928–931 are meroterpenoid chromanes from Sargassum spp., although all are weak or inactive as anti-inflammatory agents.357,358 An Indian Stoechospermum marginatum was the source of three new spatane diterpenoids 932, 933, 934 that showed weak cytotoxicity against four HTCLs.359Coriaceumins A–D 935, 936–938 are rare examples of nitrogenous xenicane diterpenoids. Two diterpenoid lactams, dictyolactams C 939 and D 940 were also isolated from the same algal species, Dictyota coriacea, collected at Nanji Island, Zhejiang Province, China, although the lactams are artefacts of isolation as proven by extraction using different alcoholic solvents.360
The total syntheses of ecklonialactones C and D, and of eiseniachloride A, have been achieved for the first time.361 Notable reviews include a summary of the phytochemicals produced by Halopteris scoparia,362 and those from the Sargassaceae between 1975 and August 2023.363 Marine carotenoid fucoxanthin has been shown to reduce the effects of autism spectrum disorder induced by valproate treatment in a mouse model in a dose dependent manner. This reduction is induced by alterations to AKT/GSK-3β signalling.364
5 Red algae
An Okinawan Portieria sample yielded a new halogenated monoterpenoid 941, which also necessitated the reassignment of related compound 942. Both metabolites exhibited antifouling activity at 0.16 µmol cm−2.365 The only other report of new red algal metabolites was of diastereomeric sesquiterpenoids laurenolides A 943 and B 944 from a Vietnamese sample of Palisada intermedia.366The same group has reported the total synthesis of pigments borolithochromes A, D, G, H1, H2, I1 and I2, metabolites of fossilised putative red alga Solenopora jurassica, via two separate publications.367,368
A review of the metabolites of Gelidium corneum including mycosporine-like amino acids, carotenoids and polyphenols, along with their associated bioactivities, has been published.369 The current status of Japanese Laurencia complex focusing on chemical diversity and morphological characters for identification has been reviewed.370 A comprehensive review of the metabolites of Asparagopsis and their use for chemotaxonomic analysis of the genus has also been released.371
A study has analysed the metabolic profile of different races of Laurencia nipponica considering their cytotoxicity, genotoxicity and their antifungal properties as it all relates to chemotaxonomy. Ultimately, with the aid of multivariate statistics, three different races can be differentiated using four different compounds, laurencin, laurallene, prepacifenol and laureatin.372 An innovative approach to establishing the absolute configuration of metabolites with vicinal bromo-chloro substituents using chiroptical methods has been published. By applying van't Hoff's principle of optical superposition, and with the aid of chiroptical measurements of suitably well characterised model compounds for comparison, the contributions of different chromophores within a compound to the overall molar rotations can be mathematically disentangled to provide the absolute configuration of the metabolites in question. Use of this method has necessitated the revision of the structures of the halogenated meroditerpenoids callyphycols A 945 and B 946 and suggested that the structures of other metabolites within the class should be reinvestigated.373
6 Sponges
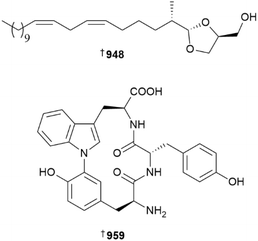
Aurantoside L 947 is an anti-leishmanial tetramic acid glycoside from Siliquariaspongia japonica.374 A Leucetta sponge from Bohol Province, the Philippines, was the source of five dioxolane ether lipids 948, 949–952. To probe the relative and absolute configurations of their dioxolane cores, model compounds were prepared and their NMR data compared with that of the isolated metabolites. Even then, some compounds were indistinguishable, so the authors utilised europium-based lanthanide chiral shift reagents to ultimately discriminate all the compounds. None of the metabolites exhibited antibacterial activity.375Endoperoxides, plakortides V 953 and W 954, and related gracilioether M 955, 11,12-dihydrogracilioether M 956, and 9,10-dihydrogracilioether G 957, were reported from Plakinastrella and Plakortis sponges, respectively.376,377
Somewhat surprisingly, only two new peptides from sponges were reported. Halichondamide A 958 was isolated from a USA-sourced Halichondria bowerbanki, with molecular dynamics simulation used to help delineate the biosynthetic formation of its disulfide bonds; note the name halichondamide should not be confused with that of the much more commonly found halichondramide macrolide family.378 Neopetromin 959 is a cyclic tripeptide that contains a rare C–N biaryl link. Although inactive against two HTCLs, various enzyme targets, and as an antimicrobial, it does cause vacuole fragmentation in an actin-independent manner in tobacco BY-2 cells, like other actin polymerisation inhibitors but without disrupting cellular actin-dynamics. Studies of the impact of this unique bioactivity upon plant physiology are underway.379
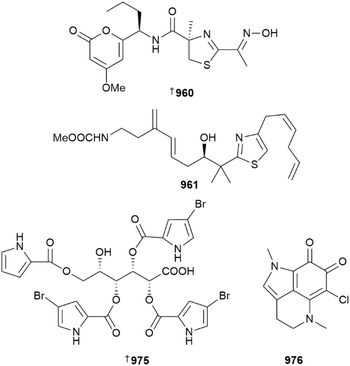
PM742 960 is the first example of a polyketide α-pyrone coupled to an oxime-containing, non-ribosomal peptide. Isolated from a Discodermia sponge, PM742 is potently cytotoxic against eight cancer cell lines. Mode of action studies indicate it binds at the colchicine site, causing mitotic arrest via inhibition of microtubule dynamics at the G2/M checkpoint.380 A synthetic SAR study identified an analogue, PM534, that covers 80% of the centres within the pharmacophore model of the colchicine binding site. PM534 has demonstrated a potent ability to reduce tumour growth in mouse xenograft models and has entered phase 1 clinical trials for the treatment of non-small cell lung cancer.381
A new mycothiazole congener 961 was reported from a sample of Cacospongia mycofijiensis collected in Vanuatu. Although a potent cytotoxin, the new analogue is still 400- to 700-fold less active than mycothiazole itself which is cytotoxic in the pM-range. Nematodes (Caenorhabditis elegans) treated with sub-lethal doses of 961 exhibited an extended mitochondrial lifespan. SAR analysis shows that Z-Δ5 alkene geometry is critical for pM levels of cytotoxicity but does not impact the mitochondrial life extension bioactivity.382
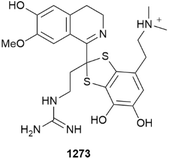
Polybrominated 962 and lipidated 963 phenols have been reported from Lendenfeldia and Hemimycale sponges, respectively.383,384 Several lactam- (964–967) and pyridine-based (968 and 969) alkaloids have been reported from a Chinese (Xisha Is.) Pseudospongosorites suberitoides. All are likely artefacts of extraction with methanol and are inactive as antivirals.385 A pair of enantiomeric indolinone alkaloids, 970 and 971, were sourced from a Tedania sponge.386 Neopetrotaurines A–C 972–974 are taurine-bridged isoquinoline dimer alkaloids that are weak suppressors of oncogenic fusion protein PAX3-FOXO-1 driven transcription. Neopetrotaurine C is a likely artefact of isolation.387
A new bromopyrrole glycoside has been reported from a deep sea Lissodendoryx papillosa specimen. Named in honour of the major contributions of Prof. Valentin Stonik to the field of MNPs, stonikacidin A 975 contains the unusual sugar, L-idonic acid, at its core. Although inactive against numerous mammalian cell lines, 975 has some antibacterial activity and it inhibits biofilm formation in S. aureus.388
Pyrroloiminoquinone alkaloids are commonly encountered in Latrunculid sponges. A dredged Latrunculia specimen yielded 6-chlorodamirone A 976 and its brominated analogue 977; over time the latter converts to the former when exposed to chloride in seawater in a non-enzymatic reaction. The total synthesis of both compounds was also achieved.389
The PIP-HSQCMBC-IPAP NMR pulse sequence was instrumental in helping identify a series of discorhabdin D congeners 978–982 reported from New Zealand Latrunculia spp.390
Multiple pyrrololactam alkaloids and their related metabolites (983–1007) have been reported from Phakellia, Stylissa, and Pseudospongosorites sponges collected around Xisha Is. in the South China Sea.391–393 A New Zealand Dictyodendrilla c.f. dendyi yielded five new sulfated denigrin metabolites 1008–1012 with one rearranging spontaneously upon short term storage in an NMR tube.394
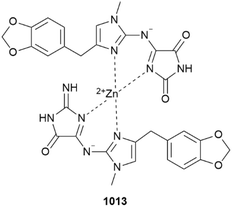
A diverse series of metabolites, including multiple 2-aminoimidazolone derivatives, were isolated from calcareous sponge Pericharax heteroraphis collected from a lagoon in Wallis and Futuna Is.; a new Zn2+-containing, heterodimeric clathridine A – clathridimine complex 1013 was one of the isolates. Although inactive as a stimulant of endochondrial ossification, the structure of the compound was confirmed by total synthesis as were those of several other congeners.395
New bromotyrosine derivatives were isolated from Suberea1014–1017, Ircinia1018–1021, and Pseudoceratina 1022 sponges collected in Australia, the Philippines, and Okinawa, respectively.396–398 Merosesquiterpenoids have been reported from Pseudoceratina purpurea (1023, 1024, and 1025–1036) and Dysidea arenaria (1037–1042), although all the isolates were only weakly active or inactive altogether in a variety of bioassays.399,400 Further meroterpenoids were isolated from Hyrtios (1043 and 1044) and Hippospongia (1045–1052) sponges,401,402 while a number of norditerpenoid cyclic peroxides 1053–1062 were reported from a Bohol Is. (the Philippines) specimen of Diacarnus spinipoculum.403 A Hemimycale sponge was the source of seco-diterpenoid 1063, which was not assessed for bioactivity.404 Isolation of ircinialactams 1064–1066 from a Mexican Ircinia felix also provided an opportunity to revise the structure of related variabilin metabolite 1067; the total synthesis of 1064 was also achieved.405 An Antarctic Suberites sponge yielded suberitenone sesterterpenoids 1068–1072 although none showed activity as antivirals.406 Given how common scalarane sesterterpenoids are in marine sponges, surprisingly only one, lendenfeldarane V 1073, was reported in 2024,407 while two nor-sterol epoxide saponins 1074 and 1075 were obtained from a Petrosia sponge collected in the Philippines.408Seco-Sterols were reported from Cliona (1076), Hippospongia (1077, 1078) and Spongia (1079) sponges, respectively.409–411 A series of 12 nor-terpenoid peroxides, 1080–1091, all derived from oxidative cleavage of larger metabolites, were isolated from a Diacarnus sponge and all with varying levels of antibacterial and cytotoxic activities.412 Other structures have been claimed but are inconsistent with the spectroscopic data provided, or were only detected by GNPS-based molecular networking, and hence are not shown.384,404,413
Numerous first total syntheses were reported and these included halisphingosine A 1092, which necessitated a revision of structure,414 cyclic peroxide ethyl plakortide Z,415 peptide neopetromin,416 and macrolide salarin C, which also validated a biosynthetic oxidation – Wasserman rearrangement to give salarin A in a biomimetic approach.417 A multiplexed strategy using 96-well plates to optimise conditions for reactions between fatty dialdehydes, a nitrogen source and acrolein has been used to probe and verify the Whitehead – Baldwin biosynthetic proposal for the manzamines.418 Indole alkaloids dragmacidins G and H, and tulongicin A, have been prepared for the first time,419,420 as has aminophenol erebusinone, leading to its revised structure 1093.421 The first total synthesis of spirodactylone has been accompanied by the synthesis of several related denigrin metabolites that has indicated the published structure of denigrin C is incorrect, although no alternative was proposed.422 Guanidine alkaloids (−)-monanchoradin A and (−)-crambescin A2 393 have been prepared for the first time,423 as have makaluvamines H, L, M and N.424,425 A series of BF2-complexes of ageladine A have been prepared as BODIPY mimics, with their fluorescence being heavily dependent on both pH and halogenation of the imidazole ring.426 Psammaplysins M, O and Q, along with ceratinamide A, have been synthesised as racemates,427 as have sesquiterpenoids halichonadins A, B and C.428
A review of sponge-derived MNPs with activity against lung cancer has been published.429 Taxa-based reviews have focused on compounds from the Spongia and Neopetrosia genera, respectively,430,431 while compound specific reviews have summarised the literature pertaining to the convoluted structure elucidation, total synthesis and biological evaluation of PUFA-derivative (−)-mucosin,432 and also of the immunomodulatory effects of halichondrin and its derivatives.433 Other compound specific reviews have concentrated on aaptamine and its derivatives,434 the anti-cancer potential of marine bis-indoles such the dragmacidins and topsentins,435 and of bromotyrosine-derivatives, mainly from Verongiid sponges.436 A compendium of 413 sponge-derived alkaloids published between 2000–2023 has been released,437 with a similar summary of diterpenoids isolated from sponges in 2009–2022 covering 228 structures.438
The inseparable endoperoxide pair, plakortinic acids C and D, isolated from the mutualistic symbiotic sponge pair Plakortis symbiotica and Xestospongia deweerdtae, have antiplasmodial activity without causing haemolysis. They exhibit promising absorption, distribution, metabolism, excretion, and toxicity (ADMET) properties making them potential antimalarial leads.439 Chemoproteomic analysis has indicated that ubiquitin carboxyl-terminal lyase 5 (USP5) is the molecular target of endoperoxide gracilioether A (Plakinastrella mamillaris). USP5 is heavily involved in chromatin maintenance, catalyses degradation of abnormal proteins, and is over-expressed in multiple cancer types. Inhibitors of USP5 therefore have promise as anticancer agents.440
Peptide stylissatin A and its analogues inhibit membranous neuraminidase-1 functioning by modulating the chaperone activity of cathepsin A, thereby having application in preventing the formation of adipose tissue.441 Chemoproteomics has also been used to detect agents that protect against inflammatory acute lung injury. Naamidine J interacts with His745 and Phe903 in chromosome segregation 1-like protein (CSE1L) which inhibits nuclear translocation and transcriptional activity of SP1, suppressing inflammation. This study also provided more naturally occurring material which facilitated the reassignment of naamidines A, G and J 1094–1096 through X-ray crystallographic analyses.442 Although only weak inhibitors of M. tuberculosis under standard culturing conditions, axinellamines A and B have moderate to potent activity against the pathogen when grown with limited protein supplementation, mimicking in vivo conditions. Both compounds are inactive against the RAW264.7 cell line as a model host.443 Several bromotyrosine metabolites sourced from Verongiid sponges were found to be weak to moderate inhibitors of prion PrPSC protein, which in turn reduces ER stress and also the spread of the pathogen in eukaryotic cells. This work provides a lead series of structures for development against a currently untreatable, and often fatal, disease.444
A nematocidal screen of over ten thousand fractions from approximately two thousand extracts held in the NatureBank repository highlighted seven extracts for further investigation. Deeper study of two chemotypes with activity against the sheep nematode Haemonchus contortus led to halaminol A and several agelasines as the dominant bioactive metabolites, but only agelasine B showed weak activity against the model roundworm Caenorhabditis elegans, suggesting either a minor component is responsible for the broader activity profile, or there is some instability in the isolated compounds.445 Sterol sokotrasterol sulfate (Topsentia sp.) was found to inhibit IFN-γ-induced expression of programmed cell death – ligand 1 (PD-L1) protein via targeting Janus kinase with a downstream activation of signal transducer and activator of transcription (STAT) signalling, making it a new immunomodulatory lead.446
Assignment of the absolute configuration of antifungal depsipeptide cyclolithistide A 1097 has been completed by the synthesis of all four stereoisomers of the rare amino acid, 4-chloroisoleucine, and then use of Marfey's method to establish the configuration of the natural residue within the metabolite.447
An important cautionary observation regarding isolation of artefacts has been made. Pyrrole-containing MNPs including makaluvamines, discorhabdins, tambjamines and brominated tryptamines, were found to be trideuteromethylated when exposed to DMSO-d6 NMR solvent in the presence of water, methanol, TFA and trace amounts of iron. Mechanistic investigation indicates that the reaction is caused by the presence of trideuteromethyl radical species generated by Fenton-type chemistry catalysed by the presence of iron or TFA.448
Finally, a study has explored the state of knowledge of the currently underexplored terpenoid biosynthesis in Dictyoceratid sponges, which, for example, are well known producers of spongian diterpenoids. While bioinformatic analysis detected analogues of known terrestrial prenyltransferases for one sesterterpenoid and multiple diterpenoid, steroid and carotenoid biosyntheses across various species, a lack of relatives to known terpenoid cyclase enzymes suggests the existence of divergent and unknown biosynthetic pathways in sponges. This study provides a start-point for exploring these currently cryptic biosynthetic genes and enzymes.449
7 Cnidarians
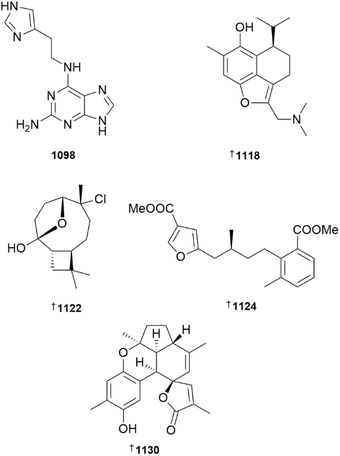
Tubastrine, sourced from the hard coral Tubastraea tagusensis, acts as an antioxidant to reduce levels of ROS, which is able to, in turn prevent neuron-like cell death caused by the oligomeric amyloid-beta peptide oAβ42.450 While acroamine A 1098, a 2-amino adenine alkaloid reported from the soft coral Acrozoanthus australiae, was essentially inactive as an inhibitor of cAMP-dependent protein kinase A, several semi-synthetic analogues including di- and tri-bromo derivatives were active.451 One of the more common family of alkaloids reported from cnidaria are the zoanthamines, isolated from Zoanthus sp. A new, efficient synthetic route to norzoanthamine has been reported, notable for including three photochemical reactions used at key steps.452 Of a set of thirteen prostanoids isolated from Clavularia sp., two (1099, 1100) were new, with the former demonstrating weak in vitro cytotoxicity towards the Ca9-22 oral cancer cell line.453 A new stereoselective approach to the synthesis of prostaglandin A2 epimers (15S)-PGA2 and (15R)-PGA2 has been reported that makes use of only catalytic amounts of heavy metals.454 The study also revealed that (15R)-PGA2 is a weak inhibitor of GABAA receptors in Xenopus laevis oocytes, while the (15S)-epimer was inactive.While the structures of sesquiterpenoids 1101–1103, isolated from Lemnalia sp., were established using a combination of NMR and MS techniques,455 the structures of six of seven new sesquiterpenoids, bellissinanes A–G 1104–1110 (Stereonephthya bellissima) and each of lemneolemnanes A–D 1111–1114 (Lemnalia sp.) were secured using XRD.456,457 Two new examples of isoledene-type sesquiterpenoids, 1115 and 1116 were isolated from the soft coral Heteroxenia sp., with both MNPs being inactive against SAR-CoV-2 and Leishmania major.458 A deep-sea collection of Anthothela grandiflora afforded unusual dimethylamino substituted cadinene-type sesquiterpenoids, anthoteibinenes A–E 1117, 1118, 1119–1121.459 Anthoteibinene B exhibited weak activity towards respiratory syncytial virus in vitro. Asymmetric syntheses of the sesquiterpenoids (−)-lemnalemnane A and (+)-lemnardosinane A, with both routes starting from (S)-carvone,460,461 and a racemic synthesis of paralemnolin A have been reported.462 Using a synthetic route starting from (−)-β-caryophyllene oxide, the structure of the chlorinated norsesquiterpenoid rumphellatin B has been confirmed, while those of rumphellatins A and C have been revised to 1122 and 1123 respectively.463 During the study, it was observed that reaction of rumphelloilide C with HCl in dioxane afforded rumphellatin B, with the authors speculating that the simple transformation may occur in soft coral species.
The structures of meroterpenoids sinudenoids F 1124 and G 1125, isolated from Sinularia densa, are unusual due to the presence of a methylbenzoate core464 while more classically functionalized meroterpenoids bearing hydroquinol or naphthoquinone cores 1126–1129 were isolated from Litophyton brassicum.465 An asymmetric total synthesis of the pentacyclic benzosesquiterpenoid (+)-verrubenzospirolactone 1130 made use of a late-stage, spontaneous, intramolecular Diels–Alder reaction.466
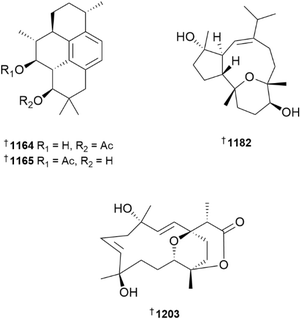
Four examples of diterpenoids embodying three different skeletons, herpetopanone B 1131 bellissimain A 1132 elisabethadienol B 1133 and elisabethadienol C 1134 were reported from extracts of Stereonephthya bellissima, with none showing activity in a pro-angiogenesis assay in zebrafish.456 Further examples of biflorane-type diterpenoids, biofloranates E–I 1135–1139, in addition to two glycosides lemnaboursides H 1140 and I 1141, were reported from a South China Sea collection of Lemnalia bournei.467 Of the set, only lemnabourside H exhibited antibacterial activity, weakly inhibiting the growth of E. coli. The structures and relative configurations of seco-diterpenoids kallopterolides A–I 1142–1150, isolated from the sea plume Antillogorgia kallos, were secured by combinations of NMR analysis and DFT DU8ML calculations.468,469 Structures were assigned to polyoxygenated casbane-type diterpenoids sinueracasbanones E–O 1151–1161, isolated from Sinularia erecta, by extensive use of NMR data in conjunction with DFT DP4+ analysis, while absolute configurations were assigned by the use of Mosher's method and TDDFT-ECD calculations.470 Sinueracasbanones I and O exhibited weak anti-inflammatory activity. The capnosane-type diterpenoid 1162, isolated from Lobophytum pauciflorum, was inactive in a zebrafish model of inflammation.471 Three cycloamphilectane-type diterpenoids, sinucycloamtins A–C 1163, 1164 and 1165 were isolated from a South China Sea collection of Sinularia brassica.472 Purification of sinucycloamtins B and C was hampered by the apparent ability of the two compounds to undergo intramolecular ester exchange. Subsequent saponification afforded a diol product, the structure and absolute configuration of which were established by XRD. Pseudopteroxazole and pseudopterosin G are antibacterial diterpenoids previously reported from the Caribbean soft coral Antillogorgia (formerly Pseudopterogorgia) elisabethae. Using an unbiased proteomics approach, the antibacterial mechanism of action of both MNPs appears to be associated with disruption of membrane-associated steps involved in cell wall biosynthesis.473
With over sixty new examples, cembranoids remain the dominant sub-structural class of diterpenoids reported from soft corals. Of a set of eleven cembranoids isolated from a South China Sea collection of Litophyton amentaceum, litoamentenes A–K 1166–1176, only litoamentene I exhibited cytotoxicity, being weakly active against the NCI-H446 cell line.474 In addition to sterols (see later), an extract of Lobophytum catalai afforded the cembrane-type diterpenoids lobocalines A–E 1177–1181.475 While XRD analysis provided the structure and absolute configuration of lobocaline A, interpretation of NMR data, including DP4+ probability analysis, and TDDFT-ECD calculations, secured the structures and configuration of the remaining examples. The structure of co-metabolite lobophytrol B, previously reported as a bicyclic terpenoid, was revised on the basis of XRD analysis to 1182.
The 3D structure of 7S,8R-dihydrodeepoxysarcophytoxide 1183 has been secured by XRD analysis.476 The findings call into doubt the structure and/or stereochemistry previously proposed for (+)-7β,8β-dihydroxydeepoxysarcophytoxide. Tetrahydropyrane ring-containing cembranoid 1184 and a 4-hydroperoxy derivative of sarcophine 1185 were reported from an extract of Sarcophyton glaucum, collected off the coast of Egypt.477 Four sarcophine analogues, sarcoeleganolides H–K 1186–1189 were isolated from a South China Sea collection of Sarcophyton elegans,478 while S. crassocaule was the source of meijicrassolins A–E 1190–1194.479
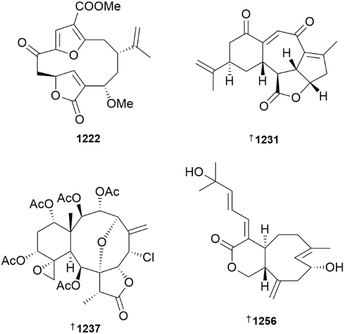
A series of capnosane-type cembranoids, sarcocinerenoids A–H 1195–1202 and two unusual variants, sarcocinerenoids I 1203 and J 1204, were reported from extracts of a Xisha Is., South China Sea collection of Sarcophyton cinereum.480 XRD analysis was used to secure the structure and absolute configuration of sarcocinerenoids A–C, F, G and I. In a separate publication, details of the characterisation of a further nine cembranoid-type diterpenoids, sarcocinerenolides A–I 1205–1213 from the same S. cinereum biomass were reported.481 Amongst these structures, sarcocinerenolide A is unusual in being a ring-opened variant, while the remaining congeners are more typical of cembranoids, displaying variation in the location of ether linkages. A structurally-related sarcophine derivative bearing a C-1 to C-12 peroxy linkage, sarcacutumolid A 1214 was reported from the extract of a Red Sea collection of Sarcophyton acutum.482
A Ximao Is., South China Sea collection of Sinularia pedunculata afforded a structurally-diverse set of cembranoid diterpenoids including sinupedunolides A–E 1215–1219 and sinupedunols A 1220 and B 1221.483,484 The structures of sinupedunolide A and sinupedunols A and B were secured by XRD analysis. Of the five cembrane diterpenoids, sinudenoids H–L 1222, 1223–1226 isolated from Sinularia densa, sinudenoid H is an unusual example of a bisnorfuranocembrenolide skeleton.464 The tricyclic structures of 4α-hydroxy-chatancin 1227 and sarcotoroid 1228, isolated from Sarcophyton tortuosum, were proposed to be derived from cembranoid precursors that had undergone transannular Diels–Alder cycloaddition reactions.485
The possible role of photochemistry in the biosynthesis of the cyclobutane ring of the cembranoid diterpenoid (+)-providencin 1229 has been supported by its semisynthesis from (−)-bipinnatin E 1230, which was isolated from a Floridian specimen of Antillogorgia kallos.486 The Norrish-Yang cyclisation step also afforded a C-14 regioisomeric product, which is likely an anticipated NP. Absolute configuration was assigned to all three compounds based upon XRD analysis of (−)-bipinnatin E. Two accounts of the synthesis of the revised structure of scabrolide B were reported in 2024. In the first publication, (−)-scabrolide B was synthesised in 19 steps (longest linear sequence), and in a simple set of reactions was converted into (−)-sinuscalide C 1231, (+)-ineleganolide and (+)-horiolide 1232.487 A second route to (−)-scabrolide B was reported by another research group, with simple conversions again affording related diterpenoids (−)-sinuscalide C and (+)-ineleganolide, as well as a third MNP, (+)-fragilolide A 1233.488 A full account of the synthesis of (−)-scabrolide F has been reported, with weak anti-fouling activity observed for the NP and synthetic intermediates.489
New biological activities were reported for cnidarian-derived cembranoids including the finding that 8-epi-sarcophinone and ent-sarcophine exhibit anti-fouling activity when included in marine paint preparations,490 sinulariolide exhibits anti-inflammatory activity in a rheumatoid arthritis model,491 7,8-epoxy-11-sinulariolide acetate also exhibits anti-inflammatory as well as inhibition of osteoclastogenesis,492 sinularin induces apoptosis in prostate cancer cells via a mechanism related to up-regulation of FOXO3 forkhead transcription factor,493 and 4-carbomethoxyl-10-epi-gyrosanolide E inhibits oral cancer cell migration and invasion by inhibition of the MAPL pathway.494 Sarcotrocheliol exhibits weak cytotoxicity, antimicrobial and larvicidal activities.495 An aquaculture specimen of Briareum stechei afforded the epoxybriarane briastecholide O 1234, while wild-type specimens of the same organism yielded the related analogue briastecholide P 1235.496
XRD analysis was used to secure the structure and absolute configuration of not only briastecholide P, but also of the known co-metabolite briaexcavatin X 1236. In two separate accounts, a large array of new and known polyoxygenated briaranes were isolated from a collection of Junceella fragilis collected off the coast of Southern Taiwan. In addition to two new chlorine-containing congeners, 1237 and 1238, two previously reported analogues gemmacolide X 1239 and frajunolide I 1240 were re-isolated.497 Absolute configurations of the latter two diterpenoids were determined for the first time. In the second publication, one new briarane example was reported, fragilide Z 1241, while the structures and absolute configurations of known co-metabolites 12-epi-fragilide G 1242, fragilide P 1243, junceellolide D 1244, junceellonoid A 1245 and juncin ZI 1246 were secured by XRD analysis.498 Excavatolide B and two semisynthetic analogues, one being an ester linked piperidine salt and the second being an oxime-linked terminal alkyne, inhibit inflammation and angiogenesis in models of atopic dermatitis.499
As part of a SAR study of transient receptor potential melastatin 7 (TRPM7) ion channel inhibitors, six new xenicane diterpenoids 7S,8S-epoxywaixenicin A 1247, 7S,8S-epoxywaixenicin B 1248, 12-deacetylwaixenicin A 1249, waixenicin E 1250, waixenicin F 1251 and 20-acetoxyxeniafaraunol B 1252 were isolated from a Hawaiian collection of Sarcothelia edmondsoni.500 The authors suggested that the epoxide-containing analogues 7S,8S-epoxywaixenicin A and 7S,8S-epoxywaixenicin B may be artefactual. Preliminary elements of an SAR were proposed, indicating the requirement of the 12-acetoxy group in combination with a dihydropyran ring for activity, and that bioactive compounds may act as latent electrophiles. Three examples of xenicane diterpenoids, miolenol 1253, epoxymiolenol 1254 and epoxycoraxeniolide A 1255, were reported from a deep-sea collection of the soft coral Paragorgia arborea.501
The first asymmetric total synthesis of the xenicane isoxeniolide A 1256 has been reported, confirming the structure and defining the absolute configuration.502 Ten new sterols were reported from cnidarians in 2024, including lobocaloids A–D 1257–1260 from Lobophytum catalai,475 cholesterols 1261 and 1262 from Litophyton mollis,503 capnesterones A 1263 and B 1264 from Capnella imbricata,504 and cholestene-7-ones 1265 and 1266 from Lobophytum durum and Capnella imbricata, respectively.505,506 The study of Lobophytum durum also led to the re-isolation of the known ergosterol derivative 1267, the relative configuration of which was determined by XRD analysis, providing clarification on the configuration of the 24-methyl group.505
A secogorgostenone steroid, originally sourced from Sinularia leptoclados, exerts anti-inflammatory activity and brain-protective effects by attenuating matrix metalloproteinase-mediated events.507 The MNP chemistry of cnidarians belonging to the genus Capnella, covering the period 1974 to May 2024, has been reviewed.508 The background history of the palytoxins, originally isolated from the zoanthid Palythoa sp., as well as their biological activities and mechanism of action, has also been reviewed.509 The chemistry and biological activities of ketosteroids, a number of which are isolated from soft corals, have been reviewed.510 A recombinant variant of Tst2, a 38-residue peptide identified by transcriptomic and proteomic analysis of the Australian sea anemone Telmatactis stephensoni, is an inhibitor of the transient receptor potential subfamily V member 1 (TRPV1) ion channel.511 HCIQ2c1, a Kunitz-peptide identified in cDNA analysis of the sea anemone Heteractis crispa, binds to the transient potential ankyrin 1 (TRPA1) calcium-permeable channel, appearing to stabilise the channel in the open conformation, and reduces pain and inflammation in vivo.512 Bioassay-directed fractionation of toxins from a Japanese collection of the sea anemone Heteractis aurora led to the identification of one novel class of toxin, denoted Hau I, with as-yet undetermined target(s), two new examples of toxins from the boundless β-hairpin family (Hau II and III) and a new example of a sea anemone type 1 sodium channel toxin (Hau IV).513
8 Bryozoans
Brominated indole alkaloids, alternatamides B and D have been synthesised from 3,4-dibromoaniline via the intermediate 5,6-dibromotryptamine514 and bryostatin 4 has been shown to inhibit the transforming growth factor (TGF)-β signalling pathway, responsible for disease progression in acute erthroleukaemia.5159 Molluscs
Compared to previous years, there was a significant drop in new MNPs reported from molluscs in 2024, with the two examples being bursatamide A 1268 and bursatellin B 1269 isolated from the sea hare Bursatella leachii.516 A general review covering the nutraceutical and medical potential of marine molluscs has been published.517 The potential of dolastatin MNPs and synthetic analogues as anticancer agents has been reviewed.518 Exploration of the SAR of (−)-doliculide A led to the identification of some analogues with similar levels of cytotoxicity as the MNP towards the HCT-116 cell line.519 Furodysinin, isolated from the nudibranch Hypselodoris tryoni, exhibited weak in vitro activity against wild-type Plasmodium berghei, but was considerable less active against an atovaquone-resistant strain.520Tambjamine analogues show interesting results as antileishmanials, with potent intracellular activity against Leishmania mexicana and L. donovani, and with one analogue demonstrating partial protection in a rodent in vivo model.521 The structures, synthetic approaches and SARs of the lamellarin alkaloids have been reviewed.522–524
The global occurrence, toxicity and risk assessment of cyclic imine shellfish toxins have been reviewed.525 Analysis of scallops Argopecten irradians exposed to paralytic shellfish toxin producing dinoflagellate Gymnodinium catenatum identified that the toxins accumulate in the adductor muscle, leading to levels exceeding regulatory limits.526 Diatoms of the genus Pseudo-nitzschia produce the neurotoxin (DA), which in the case of king scallop contamination, can take many months to clear from the mollusc tissue. DA immunodetection has been used to examine the time course of subcellular location of the toxin, finding preliminary evidence of accumulation in autophagosomes in the digestive gland, which may go some way to explain the toxin's persistence.527 Two fatty acid desaturase genes were identified in the dwarf surf clam Mulinia lateralis, the expression levels of which varied during embryonic development.528
Investigation of the transcriptome of the carnivorous snail Raphitoma purpurea has identified putative conotoxin-like peptides in both salivary gland and venom duct tissue, proving further implication for venom evolution in molluscs of the family Conidae.529 The predatory and defensive venoms associated with 16 different cone snail species have been reviewed,530 while another review has focused more on the effects of κ-conotoxins on potassium channels.531 A cyclic octapeptide Am931, identified from analysis of the venom duct transcriptome of Conus amadis, appears to act as a catalyst promoting formation of the natively folded disulfide form of α-conotoxin ImI.532 Replacement of L-lysine or L-arginine residues in α-conotoxin RgIA with their corresponding D-enantiomers has afforded weakly antifungal analogues.533
Six synthetic α-conotoxins, sequences of which were derived from transcriptome sequencing of venom of C. quercinus, exhibited potent cytotoxicity towards fall army worm cells as well as anti-larval toxicity.534 In two separate studies by the same research group, α-O-conotoxin GeXIVA[1,2] was shown to both reduce cancer chemotherapy induced neuropathic pain in a mouse model and to exhibit in vivo tumour growth suppression of triple-negative breast cancer.535,536 A disulfide bond deleted and truncated conotoxin analogue, based on the sequences of α-O-conotoxins GeXIVA and GeXXVIIA, demonstrated retained potency of human α9α10 nAChR inhibitory activity, inhibited CaV2.2 channels and alleviated pain in a mouse model.537 Conotoxin kM-RIIIJ has been used to explore the physiological functions of hetereodimeric KV1.2 channels, revealing that the receptors are highly expressed in proprioceptive (sensory motor) neurons in the dorsal root ganglion.538
10 Tunicates (ascidians)
Further investigation of an extract of an unidentified Didemnidae ascidian collected in Palau, afforded guanidine alkaloids 1-carboxydopargimine 1270, mellpaladines D–F 1271, 1272 and 1273, serodopalgimine 1274 and a 4,4′-dimer of serotonin 1275.539 Sulfoxide-containing mellpaladines D and E were isolated in equal amounts, with the authors suggesting they may be products of autoxidation. Mellpaladines D–F were evaluated for effects on behavioural activities in mice, with mellpaladine F being found to induce whole body convulsion.The original proposed disulfide, homocoupled, cyclic hexapeptide structures of antatollamides A and B have been revised by total synthesis to be simpler monomeric cyclic peptides 1276 and 1277, respectively.540 The first enantioselective synthesis of (+)-cylindricine B has been reported.541 The synthesis and multidrug resistance reversing activities of the pyrrole-containing ningalin MNPs have been reviewed,542 as have the cytotoxic properties of cephalostatin and ritterazine dimeric steroidal pyrazine alkaloids.543 The human triggering receptor expressed on myeloid cells 2 (TREM2) is an important regulator of immunological responses. N-Deacetylshermilamine B, as well as a partially characterised mixture of glucosylceramides, isolated from the ascidian Cystodytes dellechiajei, bind to TREM2, with the pyridoacridine alkaloid also exhibiting weak in vitro cytotoxicity towards two multiple myeloma cell lines.544
Phosphoeleganin and one semisynthetic analogue decrease interleukin 6 secretion, suggestive of this compound class having potential in the management of diabetes.545 A further SAR study of phosphoeleganin has identified the importance of the phosphate and glycine groups, in combination with backbone carbinol stereochemistry, for inhibition of h-15-LOX-1 enzyme.546 Of a library of synthetic analogues of rubrolide-related γ-butenolides, a non-halogenated example exhibited weak to moderate cytotoxicity towards a panel of HTCLs, induced ROS production leading to apoptosis, arrested cell cycle at G2/M and exhibited in vivo activity in a HeLa xenograft assay.547
A structurally simplified analogue of diazonamide A exhibited potent antiproliferative activity towards a panel of HTCLs which appeared to be related to inhibition of tubulin assembly, and the simplified analogue was less toxic to noncancerous cell lines.548 Screening a library of marine bacterium Tistrella strains revealed the majority were capable of producing didemnin B.549 The introduction of a second copy of the didemnin B BGC afforded a modified strain with excellent production capability (75 mg L−1), enabling semisynthesis of a series of new analogues. A cytotoxic photoaffinity probe successfully enriched a large number of potential protein targets, including the previously noted depalmitoylating enzyme PPT1. Before being discontinued due to a lack of COVID-19 related hospitalisations, a Phase III trial of plitidepsin showed it to be generally well tolerated and suggested a two-day improvement in the median time to withdrawal of supplementary patient oxygen support.550 Screening a library of marine-derived compounds identified polycarpine as a potent inhibitor of the main protease Mpro of coronaviruses, also inhibiting SARS-CoV-2 and PEDV (porcine epidemic diarrhoea virus) viral replication in Vero-E6 cells.551 The uses of the tetrahydroquinoline alkaloid trabectedin and its related analogue lurbinectedin in oncology, mechanisms of action, and uses in combination therapies have been reviewed.552
11 Echinoderms
Extracts of the arms of the starfish Thromidia catalai afforded three indole derivatives, catalindoles A–C 1278, 1279 and 1280, the structures of which were confirmed by synthesis.553 When collected, the starfish were found feeding on the sponge Theonella swinohei – LCMS analysis of extracts of the starfish digestive gland identified the presence of Theonella-sourced NPs while none were located in the arms, suggesting that the starfish does not accumulate sponge metabolites.Steroidal disulfates 1281–1284 were isolated from a Far Eastern collection of the slime sea-star Pteraster marsippus.554 The biological activities of saponins isolated from sea-stars has been reviewed.555 Several examples of polyhydroxylated steroidal saccharides were also reported from starfish, including pectiniferosides A–J 1285–1294 from Patiria (= Asterina) pectinifera,556 spiculiferosides A–D 1295–1298 from Henricia leviuscula spiculifera,557 and ceramasterosides A, B, D and E 1299–1302, from Ceramaster patagonicus.558 Of these MNPs, only the latter, ceramasteroside D, exhibited a notable biological activity, of moderate ability to inhibit the release of nitrite from LPS stimulated microglial cells.
Ten examples of triterpenoid glycosides were reported from sea cucumber species, including pacificusosides L–Q 1303–1308 from Solaster pacificus,559 peronioside A 1309 from Psolus peronii,560 conicospermiumosides A3-1, A3-2, A3-3, A7-1 and A7-2 1310–1314 from Cucumaria conicospermium,561 and 1315–1318 from Apostichopus japonicus.562,563 Investigation of extracts of the sea urchin Clypeaster humilis afforded sulfate 1319, methane sulfonate 1320 and boldine 1321, the latter two compounds being reported from a marine organism for the first time,564 while bibenzochromenones phanogracilins A–C 1322–1324 were isolated from the crinoid Phanogenia gracilis.565 Additional biological properties have been reported for echinochrome A, including anti-asthmatic activity566 and the ability to modulate migration of non-small cell lung cancer cells.567 2-Butoxytetrahydrofuran, previously reported from Holothuria scabra, attenuates the aggregation and oxidative properties of α-synuclein, lessening its toxicity in a C. elegans model of Parkinson's disease.568
The effect of the saponin frondoside A on inflammation associated with tooth replantation has been investigated.569 Holothurin B and related saponins are able to prevent mast cell degranulation, as measured by the release of β-hexosaminidase, and to alleviate the action of a number of pro-inflammatory cytokines.570 The chemistry and biological activities of MNPs reported from the sea cucumber Apostichopus japonicus (2000 to late 2023) have been reviewed.571 Holotoxin A1, isolated from A. japonicus, inhibits candidiasis by a wide range of mechanisms including increased intracellular accumulation of ROS, mitochondrial membrane depolarisation, and inhibition of mitochondrial complex I.572 Further examination of the triterpenoid glycoside cucumarioside A2-2 has identified it to be active against prostate cancer cell lines, while cucumarioside A0-1 and djakonovioside A are active against triple-negative breast cancer cell lines.573,574
12 Miscellaneous
The structure of the bicycloheptane derivative (−)-raikovenal 1325, originally isolated from the marine ciliate Euplotes raikovi, has been confirmed by total synthesis, with the study also assigning absolute configuration.575 The potential of seagrass meadows as a source of bioactive molecules has been reviewed,576 while species specific reviews of Halophila stipulacea577 and Halodule uninervis578 covered the phytochemistry and pharmacological activities of metabolites. Further investigation of the diarylheptanoid heterodimer zosterabisphenone B determined it can induce apoptosis in HCT116 cells and reduce the growth of human colon tumour xenografts in mice.579Five pyridyl and dihydroisoquinoline alkaloids, 1326–1330, were reported from the nemertean worm Amphiporus angulatus.580 Collections of the intertidal sipunculid worm Phascolosoma granulatum afforded phascolosomines A–F 1331–1336, characterised as linear guanidine amides.581 A SAR study of the antimicrobial peptide capitellacin, originally isolated from the annelid worm Capitella teleta, identified the importance of one of the disulfide linkages for activity, that the linear peptide was devoid of activity, the D-enantiomer had reduced activity and that most analogues only exhibited weak, at best, activity towards E. coli.582 Bioinformatic searching of the transcriptome of the lugworm Arenicola marina identified a new BRICHOS-related antimicrobial peptide, AmBRI-aaa, that was prepared using heterologous expression in E. coli.583 The 44 residue peptide exhibited weak to moderate activity towards some Gram-positive bacterial strains and whole genome sequencing of a resistant mutant identified targets associated with cell wall biosynthesis.
The ribbon worm Cephalothrix simula, known to contain tetrodotoxin, may be a source of toxin in British farmed Pacific oysters Magallana gigas.584 A truncated variant of a peptide, bolespleenin,334–347 derived from a gene product of the mudskipper fish Boleophthalmus pectinirostris, exhibited broad-spectrum antibacterial activity, inhibited biofilm formation and was active in a mouse model of skin infection.585 Further investigation of the peptide epinecidin-1, originally isolated from the orange-spotted grouper Epinephelus coioides, has determined its ability to suppress lipoteichoic acid-enhanced proliferation of non-small cell lung cancer cells and induce necrotic cell death via a mechanism involving mitochondrial damage and elevated levels of ROS.586 The 22-residue peptide piscidin-1, derived from mast cells of the hybrid striped bass Morone saxatilis × M. chrysops, induces apoptosis in oral squamous cell carcinoma cells, and also increases ROS levels leading to mitochondrial dysfunction.587
GC/electron capture negative ion-MS analysis of seventeen species of fish from the Seychelles in the Western Indian Ocean identified the predominant polyhalogenated compounds present were halogenated NPs of the methoxylated diphenyl ether class, rather than PCB or DDT anthropogenic persistent pollutants.588
13 Perspective – the emerging role of artificial intelligence in MNP research
The rapid uptake of artificial intelligence (AI) in commerce, health, transportation and communications is stunning. In the sciences, AI is equally transformative. AI is used to analyse large amounts of different kinds of data, to find patterns and relationships, and use those to make predictions, decisions or generate “new” content. Machine learning (ML) is the most widely used approach, and it identifies patterns in data without relying on rules-based programming, allowing tasks to be performed automatically. When dealing with more complex, high-dimensional, unstructured data, a specialized subset of ML called deep-learning (DL) employs multilayered neural networks to identify intricate patterns.Given that computer assisted analysis have been used for decades in MNP research, how much has the introduction of AI improved productivity in MNP research and increased the discovery of novel and/or potently active MNPs in the last decade?
Two main applications of AI are generative (GAI) and predictive (PAI). GAI is based on large language models (LLN) mostly trained on online data and analysed using ML or DL algorithms to produce content copied from existing data but stitched together in different ways to construct a “new” amalgam as text, image or video that mimics the training data. GAI can aid researchers to write code for software packages such as Python and R, produce images for graphical abstracts or presentations, help construct grammatically correct text, and provide suggestions for areas to investigate when initiating research on an unfamiliar topic. But GAI needs to be used with caution because it has a propensity to generate wildly inaccurate results. For instance, creating images of chemical structures using GAI is particularly unreliable (Fig. 2).589
PAI by contrast can analyse large amounts of orthogonal data to identify patterns that are often hidden or difficult to discern and predict new findings based on this pattern recognition.
The application of AI in the molecular sciences is mostly predictive rather than generative and this approach has resulted in important breakthroughs. The development of AlphaFold, an AI model to predict 3D protein structures based on amino acid sequence, for example, is transformative. Using a transformer neural network architecture trained on ∼180![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 experimentally determined 3D protein structures, AlphaFold can now predict the 3D structures of over 200 million proteins.590 These predictions are now being incorporated into the drug discovery process.
000 experimentally determined 3D protein structures, AlphaFold can now predict the 3D structures of over 200 million proteins.590 These predictions are now being incorporated into the drug discovery process.
Computational methods such as TDDFT calculations to assign absolute configurations to molecules, spectroscopic simulations, computer-assisted structure elucidation using rules-based algorithms, compound similarity and substructure searches, dereplication using spectral database matching tools and virtual screening of compound libraries through in silico protein docking are all computer assisted (but not AI) methods developed since the 1990s. Many of these methods are still widely used in NP research and most involve complex calculations that require supercomputers to do time-consuming analyses. The arrival of ML has created opportunities to massively reduce the time required to generate similar results without using supercomputers.591–594
ML is now used as an aid in MS-based dereplication by predicting MS fragmentation patterns for NPs found in databases to augment the annotation of molecules in molecular networks that were previously only identified through matching experimental spectrometric data.595 Augmenting lower level DFT calculations with ML generated parametric corrections has resulted in more accurate predictions of NMR data in minutes rather than days.596,597 ML assisted genome mining has resulted in the discovery of BGCs and the prediction of their products.598,599
MNP researchers still face many bottlenecks that impede productivity. These include access to biota, accurate taxonomic identification of biota, identification of “interesting” NPs especially those present in complex mixtures, separation and structure identification of NPs, identifying a NP's bioactivity and understanding its mode of action, identifying the genes and proteins responsible for NP biosynthesis and synthesis of the NP for sustainable production or to generate analogues.
We analysed 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 582 papers and reviews published in the field of MNP (obtained from MarinLit) over the last 10 years (January 2015–December 2024) to help understand how AI has contributed to solving these MNP research bottlenecks. Keyword searches (Table S2) of all abstracts allowed a connectivity map (using VOSviewer)600 to be generated by creating a thesaurus of keywords that we grouped into ontologies based on relatedness of the words. The limitation of this approach is that some abstracts do not necessarily include information regarding the use of AI, but the keyword connectivity maps can help to infer AI usage based on a combination of ontologies. A network from genome mining to bacteria, BGC, molecular network, predict, MS, bioinformatic, model and database for example infers AI is being used in microbial MNP research even though there is no explicit association between the use of AI terms and genome mining in the abstracts (Fig. 3).
582 papers and reviews published in the field of MNP (obtained from MarinLit) over the last 10 years (January 2015–December 2024) to help understand how AI has contributed to solving these MNP research bottlenecks. Keyword searches (Table S2) of all abstracts allowed a connectivity map (using VOSviewer)600 to be generated by creating a thesaurus of keywords that we grouped into ontologies based on relatedness of the words. The limitation of this approach is that some abstracts do not necessarily include information regarding the use of AI, but the keyword connectivity maps can help to infer AI usage based on a combination of ontologies. A network from genome mining to bacteria, BGC, molecular network, predict, MS, bioinformatic, model and database for example infers AI is being used in microbial MNP research even though there is no explicit association between the use of AI terms and genome mining in the abstracts (Fig. 3).
 | ||
| Fig. 3 Connectivity map (created in VOSviewer)600 of network linkages between keywords extracted from MNP paper abstracts. The size of a keyword node is proportional to the number of papers that contain that ontology and the width of the linkage to another node is proportional to the number of instances that the linkage occurs. Each of the top 21 nodes has a individual colour. (A) Total network; (B) highlighted network from “genome mining”. | ||
We found that very few studies have used AI to aid in the discovery of new MNPs or bioactivities over the last decade. Overall, the rate of discovery of new MNPs (Fig. 1) and potently active MNPs has remained static over the decade.601
AI has only been explicitly referred to in 82 MNP papers over this period (Fig. 4 and 5) and is most highly associated with the prediction of bioactivity through either in silico docking of MNPs to proteins or inferred bioactivity based on structural similarities. ML augmentation of DFT calculations for more accurate NMR chemical shift prediction (DU8ML),596 pattern recognition within 2D NMR spectra to predict structures (SMART2.0/DeepSAT),602,603 pattern recognition within structures to predict NP structure classes (NPclassifier),604 prediction of peptide quaternary structures (conotoxins),605 and genome mining to generate hypothetical virtual libraries of potential structures of undiscovered molecules for MS molecular network matching (HypoRiPPAtlas)599 are examples of other uses. To date, none of these methods have been widely adopted.
 | ||
| Fig. 5 Connectivity map (created in VOSviewer)600 of network linkages between keywords extracted from MNP paper abstracts. (A) Highlighted network from “artificial intelligence” ontology; (B) highlighted network from “in silico” ontology. | ||
In silico screening methods (233 papers) have steadily increased over the decade. There have been several studies that have used databases to predict MNPs active against specific targets (such as the SARS-COV 19 virus) through virtual screening. This approach previously used the knowledge of the protein structure (obtained mostly by XRD) and in silico docking techniques to identify molecules that interacted with the protein. However, AI methods now tend to use the knowledge gained from molecules that have been shown experimentally (or virtually through in silico methods) to interact with a specific target protein (or a structurally related protein) and these interactions are used to train a model to identify other compounds that will also interact. This pattern recognition approach is potentially quicker to implement and can be used to analyse large datasets. Unfortunately, where this approach has been adopted using MNPs, the verification of “hits” obtained mostly relies on virtual screening of the “hit” molecules through molecular docking experiments. This seems to be a circular approach given that in some cases, the training data was originally obtained through virtual screening. There has been limited examples where the virtual “hit” compounds have been tested in real laboratory assays.
One of the key advantages of studying marine biodiversity is the propensity to discover unprecedented chemistry. PAI learns from previous discoveries to identify known molecules and their close unknown analogues and to predict new bioactivities for these compounds based on their structural similarity to other molecules with demonstrated bioactivity against a known target. This has the potential to “de-orphan” known MNPs lacking any bioactivity data, but to date this has not occurred.
Linking biosynthetic genes with known NPs or predicting new NPs from bioinformatics associated with BGCs are also effective outcomes from PAI assisted research (BGC is mentioned 561 times over the decade). MNP research increasingly uses ML augmented molecular networking of LCMS data as an aid in the selection of biota to investigate or to link molecular network clusters to BGC predictions. Molecular networking has been discussed in 294 papers over the last 10 years, but invariably the selection of clusters has resulted in the isolation of closely related analogues of known compounds. So, based on this, one might expect that greater effort would now be focused on the unknowns with AI doing the heavy lifting in dereplication by sifting through the large volumes of “known” chemistry to identify the uniquely unknown metabolites that can then become the focus of new research. There is little evidence that this approach is being used yet but hopefully, the future use of this powerful tool can be tailored to highlight the unknowns rather than the knowns.
There is no evidence of an increase in new MNP novelty (i.e. structures possessing new ring systems or features not reported before; information from previous MNP reviews). The term “novel” has been used 2255 times and “unprecedented” 344 times, but virtually never in relation to either truly novel scaffolds or bioactivities. The lack of increased structural novelty might be related to diminishing returns that are anticipated when specific biota resources have been extensively studied.606 However, it could also reflect the sustained use of standard purification, biota selection, culture methods, or bioassay methods that unintentionally exclude certain structures. An example of this bias can be seen when normal-phase silica gel separation methods which were almost exclusively used until the early 1990s started to be supplemented with reversed-phase separation methods, thus expanding the latent chemical diversity accessible within well studied species (Fig. 6). This resulted in a significant increase in average polar surface area of MNPs after the early 1990s that also reflected a difference in structural novelty. If a ML approach were adopted, it could potentially identify specific purification methods that have a higher chance of yielding novel chemistry.
 | ||
| Fig. 6 Box plot of calculated polar surface area (PSA) for all new MNPs published each year between 1972 and 2024. Black horizontal lines represent median PSA. | ||
There are still very significant pockets of marine biodiversity that remain unstudied. Small animals and plants (species with individuals occupying 1 cm3 or less) are rarely sampled but potentially account for most biodiversity in an area.607,608 They likely occupy different niches where environmental pressures could drive differences in MNP expression. ML from ecological data could prove insightful. The propensity to isolate and culture microorganisms sourced from marine environments using media suited to fast growing, terrestrial soil-derived microbes, does not help to exploit truly marine biodiversity and their potentially unique chemical diversity. The choice of isolation media and culture media used to access truly marine microbes might benefit from an AI approach. Combining data from microbial ecology, biogeochemistry, aquaculture, food sciences, ecological studies and metagenomics for ML might help to uncover specific nutrient requirements that could enhance the isolation and cultivability of truly marine species.
MNP discovery continues to rely on serendipity. There is no evidence so far that AI has been applied to the biota selection process. At its most basic, MNP researchers have used prior knowledge to target specific species for study (soft bodied marine invertebrates, algae, fungi, bacteria, cyanobacteria). AI could aid in this process. However, we know from experience that the random collection of an invertebrate that was collected simply because it looked different or was found in an unusual place, has led to some of the most unusual molecules we have discovered. Perhaps an AI assistant on a SCUBA dive could help find that elusive organism.
Genome mining helps to identify the vast potential of BGCs (most of which are silent) in microorganisms, but what about the NPs that are not made by enzymes encoded by clustered genes? Pertinent examples are many types of alkaloids which represent a substantial proportion of MNPs and drugs derived from NPs. ML methods using Alphafold could provide insights to predict proteins from genomic data for non-clustered biosynthesis.
The structure determination of NPs has always been challenging, and continued reports of MNP structure revisions show that improved methods to aid in structure determination are always welcomed. Computationally demanding DFT methods to predict NMR chemical shifts have been augmented with parametric rules obtained through ML approaches using NMR data obtained from large libraries of published structures. This has resulted in much faster predictions and resulted in two MNP structure revisions.596 The tool can also be used to establish if a proposed structure is correct prior to publication. SMART2.0 is an image-based AI tool that predicts the structure of a compound based on the similarity of its HSQC image to those generated from a training set of over 53![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 experimental HSQC images or HSQC images generated through predictions of H and C chemical shifts for compounds lacking experimental data.602 The four MNP studies that have used SMART2.0 to guide prioritisation all resulted in the isolation of either close analogues of known compounds or a chimeric molecule containing features from two known molecules. Overall, these ML NMR methods can improve the structure determination workflow, but do not necessarily result in greater novelty of structures being published. Hopefully, it will reduce the number of incorrect structures appearing in the literature (if used judiciously at the time of peer review prior to publication).
000 experimental HSQC images or HSQC images generated through predictions of H and C chemical shifts for compounds lacking experimental data.602 The four MNP studies that have used SMART2.0 to guide prioritisation all resulted in the isolation of either close analogues of known compounds or a chimeric molecule containing features from two known molecules. Overall, these ML NMR methods can improve the structure determination workflow, but do not necessarily result in greater novelty of structures being published. Hopefully, it will reduce the number of incorrect structures appearing in the literature (if used judiciously at the time of peer review prior to publication).
Open Molecules 2025 (OMol25), released in mid-2025 by Meta and the US Department of Energy's Lawrence Berkeley National Laboratory, contains 100 million DFT calculations covering a huge variety of molecular systems that can now be used in ML applications.609 This has the potential to massively reduce the time required to predict NMR and ECD data with estimates suggesting a 10-thousand-fold reduction in cpu time to generate equivalent quality predictions to those generated by DFT calculations. This resource has the potential, through ML algorithms, to predict NMR and ECD data for all known MNPs with comparable accuracy to quantum mechanical calculations virtually instantaneously on a laptop or smart phone. Linking this potential capability with other AI platforms such as 2D NMR image recognition, MS2 data recognition and NP structure prediction from genomic data and NP bioactivity predictions could provide a paradigm shift in NP discovery efforts.
Since ML methods are trained on known data (such as molecules, NMR spectra, MS2 data, genes) there is an obvious bias to match unknowns that mimic knowns. “Rare” structures might not feature in a training set, and undiscovered novel molecules clearly cannot be there at all. Therefore, important potential matches might be missing in the model. This is analogous to the inherent bias used by Amazon to recruit IT staff where female applicants were culled early in the automated selection process because the data used to train the algorithm did not include applications from women.610 Training models to highlight and prioritise the unknowns will help to counter these biases.
There is a danger when AI is used as a “black box” method by non-experts who accept the results as gospel without any understanding or questioning because of a lack informed knowledge. If these unverified AI results are subsequently used to train new models, a misinformation loop is exacerbated, rendering future models even less accurate than their predecessors. There is, therefore, an ongoing need for orthogonal testing of results obtained from AI tools in MNP research. In structure determination, this means that XRD and total synthesis will continue to be important verification tools. Laboratory-based bioassays need to be used to confirm or reject an AI computational result and MS2 fragmentation matching for structure annotation will still rely on real MS data from standards.
There have been highly innovative AI methods developed, and some have been widely adopted while others have not. It is somewhat baffling to know why this is the case. For instance, molecular networking is now widely adopted with platforms such as GNPS and applications such as MZmine providing a simple computational engine for upload of raw data and download of clustered and annotated results.611,612 The barrier to adopting this technology is low and labs using these platforms benefit from their simplicity to access. Equally as innovative is the SMART2.0/DeepSAT platform for interpretation of 2D NMR data to match known compounds and to predict structure classes for unknown compounds. However, the adoption of these methods by the MNP community is low (only four papers refer to SMART2.0 in their abstract). One reason may be that many MNP research groups have NMR “gurus” who can identify a known structure from limited experimental data extremely quickly and do not need to rely on computational methods. However, the most likely reason for slow uptake is that these methods still do not work well with NMR of mixtures (apart from matching major components). The time involved to purify an extract and then obtain HSQC data on individual molecules compared to injecting a minute quantity of an extract into an HPLC column and generating MS2 data for thousands of compounds in minutes is likely to be a serious impediment.
Negative results are a major gap in published knowledge. For example, many MNP groups have large inventories of marine species collected or cultured randomly. These collections invariably contain individual samples that have been studied but either produce no MNPs or only known MNPs or the isolated MNPs showed low or no bioactivity in a range of bioassays. These results do not generally make it into research papers but are a gold mine of information that could be used in AI analysis. To make the most of all research outputs therefore requires an acceptable mechanism to capture open access, validated negative research results as well as positive ones.
Linking datasets will be key to seeing the most productive uptake and application of AI in MNP research. The World Register of Marine Species (WoRMs) contains up-to-date taxonomic classifications, collection records and distributions for over 500![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 species,613 the Ocean Biodiversity Information System contains 167 million geolocation records for over 200
000 species,613 the Ocean Biodiversity Information System contains 167 million geolocation records for over 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 marine species and 27 million gene sequences,614 citizen science initiatives such as iNaturalist have images of species linked to taxonomic data and geolocation with close to 280 million observations linked to 536
000 marine species and 27 million gene sequences,614 citizen science initiatives such as iNaturalist have images of species linked to taxonomic data and geolocation with close to 280 million observations linked to 536![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 species (including 4041 observations from 1280 marine invertebrate species),615 the Lotus initiative provides taxonomic links to over 750
000 species (including 4041 observations from 1280 marine invertebrate species),615 the Lotus initiative provides taxonomic links to over 750![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 NP structure/organism pairs,616 the newly released Minimum Information about a Tailoring Enzyme (MITE) links biosynthetic enzymes with reaction type and precursor and product structures,617 SMART2.0 uses a database of over 50
000 NP structure/organism pairs,616 the newly released Minimum Information about a Tailoring Enzyme (MITE) links biosynthetic enzymes with reaction type and precursor and product structures,617 SMART2.0 uses a database of over 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 HSQC images to link with structures, GNPS and other MS based platforms link MS2 data to structures. Linking these disparate datasets could provide opportunities to guide new discoveries and to unlock novelty in plain sight.
000 HSQC images to link with structures, GNPS and other MS based platforms link MS2 data to structures. Linking these disparate datasets could provide opportunities to guide new discoveries and to unlock novelty in plain sight.
This perspective has highlighted that MNP researchers are only beginning to embrace the power of AI and it will be fascinating to see where MNPs research in the AI era evolves over the coming decade.
14 Conclusion
New MNP discoveries in 2024 have shown an increasing prevalence of MNPs compounds derived from marine fungi and a decreasing trend in macro-organism studies. Although close to half of all new MNPs were reported from marine fungi in 2024, the majority are minor variations on already well represented structure classes. Actinomycetes are a minor contributor to overall marine procaryote biodiversity, but they continue to be the major source of new bacterial MNPs.BCGs associated with MNPs obtained from bacteria, cyanobacteria and to a lesser extent fungi continue to be characterised, and genome mining is increasingly being used to find new bacterial MNPs. In some cases, these genomic approaches have resulted in proposed new MNP structures without comprehensive and convincing spectroscopic evidence and from our perspective these compounds have not been structurally verified. Rigorous spectroscopic characterisation including NMR spectroscopy and or XRD analysis should complement MS-based evidence in future studies.
The bioactivities reported for select MNPs continue to be a highlight, with the clinical development of a synthetic derivative of the Indonesian sponge-derived alkaloid PM742 holding promise for lung cancer patients. Synthetic studies continue to be an important technique to support or refute MNP structures and in 2024, over 120 first total syntheses of MNPs were accomplished, leading to several structure revisions and over 10 absolute configurational assignments being established.
New sponge-derived MNPs had fewer than expected examples of peptides and alkaloids based on previous trends. Cnidarian studies were predominantly associated with organisms collected from Chinese, Vietnamese and Taiwanese waters in the South China Sea pointing to reduced emphasis on cnidarian biodiversity hotspots. Molluscs, ascidians, bryozoans, red, green and brown algae were, in total, only represented by a handful of studies, while echinoderms were mainly studied from Russian collections. These observations point to diminished geospatial and phylogenetic diversity of MNP discoveries. If these biodiversity trends continue, the overall relevance of MNP research will be diminished. That notwithstanding, the contributions of new MNPs to the structural diversity of organic chemistry remains high and they represent a valuable resource for biodiscovery.
15 Conflicts of interest
There are no conflicts to declare.16 Data availability
Supplementary information (SI): all structures along with their names, taxonomic origins, collection locations, and biological activities. See DOI: https://doi.org/10.1039/d5np00080g.17 Acknowledgements
We thank the Royal Society of Chemistry for the provision of data used in this review, adapted from the MarinLit database with permission from the Royal Society of Chemistry.2 The contributions of the NIH author(s) were made as part of their official duties as NIH federal employees, are in compliance with agency policy requirements, and are considered Works of the United States Government. However, the findings and conclusions presented in this paper are those of the author(s) and do not necessarily reflect the views of the NIH or the U.S. Department of Health and Human Services. We thank Santiago Bueno Horcajadas from PharmaMar for permission to use his photograph of the Indonesian sponge Discodermia sp. in the table of contents graphic.18 References
- A. R. Carroll, B. R. Copp, T. Grkovic, R. A. Keyzers and M. R. Prinsep, Nat. Prod. Rep., 2025, 42, 257–297, 10.1039/D4NP00067F.
- https://pubs.rsc.org/marinlit, accessed July 2025.
- Y. Zou, J. Shi, J. L. Sun, L. Y. Li, Z. Y. Yan, Z. K. Guo and R. H. Jiao, J. Nat. Prod., 2024, 87, 2530–2536, DOI:10.1021/acs.jnatprod.4c00836.
- M. Girão, J. Murillo-Alba, J. Martín, I. Pérez-Victoria, R. B. Leite, R. Urbatzka, P. N. Leão, M. F. Carvalho and F. Reyes, Mar. Drugs, 2024, 22, 268, DOI:10.3390/md22060268.
- J. I. Quintana-Bulla, L. A. C. Tonon, L. F. Michaliski, E. Hajdu, A. G. Ferreira and R. G. S. Berlinck, Appl. Microbiol. Biotechnol., 2024, 108, 112, DOI:10.1007/s00253-023-12870-0.
- M. Elsbaey, T. Jomori, J. Tanaka, N. Oku and Y. Igarashi, J. Antibiot., 2024, 77, 389–392, DOI:10.1038/s41429-024-00718-2.
- H. P. S. de Freitas Cesário, F. das Chagas Lima Pinto, K. M. Canuto, E. R. Silveira, D. V. Wilke, E. G. Ferreira, A. M. da Fonseca, M. A. de Vasconcelos, E. H. Teixeira and O. D. L. Pessoa, Chem. Biodiversity, 2024, 21, e202301771, DOI:10.1002/cbdv.202301771.
- M. Wang, Z. Zheng, Z. Hu, B. Fan, J. Liu and Q. Xu, Nat. Prod. Res., 2024, 38, 2681–2687, DOI:10.1080/14786419.2023.2200183.
- W. Zhao, H. Jiang, Y. Ge, C. Zhou, Y. Ma, J. Zhou, Y. Xie, Y. Wang and B. Wu, Fitoterapia, 2024, 175, 105946, DOI:10.1016/j.fitote.2024.105946.
- J. Lee, S. Um, E.-H. Kim and S. H. Kim, J. Nat. Prod., 2024, 87, 733–742, DOI:10.1021/acs.jnatprod.3c00843.
- L. Zhou, Y. Chang, S. Yang, X. Huang, J. Wang, C. Jiang, T. Zhu, D. Li and Q. Che, J. Antibiot., 2024, 77, 201, DOI:10.1038/s41429-023-00698-9.
- P. F. Hillman, J. Y. Lee, J. Lee, J. Lee, D.-C. Oh, W. Fenical and S.-J. Nam, Nat. Prod. Res., 2024, 38, 2357–2362, DOI:10.1080/14786419.2023.2173192.
- S. Um, H. Jeong, J.-E. Park, J. Seo, S. H. Jung, M. Bae, K.-T. Lee and K. Moon, Mar. Drugs, 2024, 22, 539, DOI:10.3390/md22120539.
- W. Tang, D. Zhang, J. Xu, S. Wang, B. Wei and L. Li, Arab. J. Chem., 2024, 17, 105687, DOI:10.1016/j.arabjc.2024.105687.
- J. Park, Y. Shin, S. Hwang, J. Kim, D. H. Moon, I. Kang, Y. Ko, B. Chung, H. Nam, S. Kim, K. Moon, K. Oh, J. Cho, S. K. Lee and D. Oh, Angew. Chem., Int. Ed., 2024, 63, e202402465, DOI:10.1002/anie.202402465.
- F. Pu, J. Fang, W. Li, B. Zhang, X. Hong, L. Xu, X. Li and Y. Jiang, Chem. Biodiversity, 2024, 21, e202400029, DOI:10.1002/cbdv.202400029.
- W. Ding, Y. Li, X. Li, J. Yin, S. Shi, X. Tian, S. Zhang and H. Yin, Mar. Drugs, 2024, 22, 181, DOI:10.3390/md22040181.
- W. Wang, J. Shang, S. Yin, K. Xia, R. Ma, W. Zhu and P. Fu, J. Nat. Prod., 2024, 87, 2580–2591, DOI:10.1021/acs.jnatprod.4c00702.
- X. Xu, Y. Chang, Y. Chen, L. Zhou, F. Zhang, C. Ma, Q. Che, T. Zhu, B. A. Pfeifer, G. Zhang and D. Li, Org. Lett., 2024, 26, 7489–7494, DOI:10.1021/acs.orglett.4c02074.
- J. L. Wissner, J. R. Almeida, I. R. Grilo, J. F. Oliveira, C. Brízida, W. Escobedo-Hinojosa, P. Pissaridou, M. I. Vasquez, I. Cunha, R. G. Sobral, V. Vasconcelos and S. P. Gaudêncio, Front. Chem., 2024, 12, 1425953, DOI:10.3389/fchem.2024.1425953.
- J. Yu, H. Guo, J. Zhang, J. Hu, H. He, C. Chen, N. Yang, F. Yang, Z. Lin, H. Dai, L. Ouyang, C. Liu, X. Lei, L. Zhang, G. Zhu and F. Song, Mar. Drugs, 2024, 22, 259, DOI:10.3390/md22060259.
- H. Huang, Y. Zhang, Y. Song, C. Ling, S. Peng, B. Ding, Y. Tao and J. Ju, Mar. Drugs, 2024, 22, 32, DOI:10.3390/md22010032.
- M. Zhao, W. Zhang, C. Yang, L. Zhang, H. Huang, Y. Zhu, D. Ratnasekera and C. Zhang, J. Nat. Prod., 2024, 87, 1591–1600, DOI:10.1021/acs.jnatprod.4c00232.
- K.-X. Yi, Q.-Y. Xie, Q.-Y. Ma, L. Yang, H.-F. Dai, Y.-X. Zhao and Y.-E. Hao, Fitoterapia, 2024, 173, 105814, DOI:10.1016/j.fitote.2023.105814.
- Y. Chen, H. Jin, W. Xiong, Z. Fang, W. Sun, Y. Zhu, L. Zhang, Y. Zhang, W. Zhang and C. Zhang, Org. Lett., 2024, 26, 1677–1682, DOI:10.1021/acs.orglett.4c00168.
- S. Shigeno, M. Kadowaki, K. Nagai, K. Hosoda, T. Terahara, T. Nishimura, N. Hasegawa, H. Tomoda and T. Ohshiro, J. Antibiot., 2024, 77, 265–271, DOI:10.1038/s41429-024-00710-w.
- W. Ding, Y. Li, X. Li, S. Shi, J. Yin, X. Tian, M. Xiao, S. Zhang and H. Yin, Phytochemistry, 2024, 222, 114101, DOI:10.1016/j.phytochem.2024.114101.
- X.-Y. Feng, J.-H. Li, R.-J. Li, S.-Z. Yuan, Y.-J. Sun, X.-P. Peng, H. Dong, H.-X. Lou and G. Li, J. Agric. Food Chem., 2024, 72, 1082–1095, DOI:10.1021/acs.jafc.3c06307.
- Z. Wang, J. Yin, M. Bai, J. Yang, C. Jiang, X. Yi, Y. Liu and C. Gao, Mar. Drugs, 2024, 22, 38, DOI:10.3390/md22010038.
- L. Chen, K. Liu, J. Hong, Z. Cui, W. He, Y. Wang, Z. Deng and M. Tao, Mar. Drugs, 2024, 22, 189, DOI:10.3390/md22040189.
- H. J. Shin, S.-Y. Jung, J. S. Kang, C.-S. Heo and S. J. Park, J. Nat. Prod., 2024, 87, 2432–2440, DOI:10.1021/acs.jnatprod.4c00683.
- Y. Xu, F. Pu, Y. Cui, Y. Wu, Z. Lei and Y. Jiang, Chem. Biodiversity, 2024, 21, e202401154, DOI:10.1002/cbdv.202401154.
- S.-M. Shen, Y.-C. Xie, L.-R. Tu, M.-E. Wu, Y.-M. Wang, C.-H. Song, Y.-H. Sun and M.-H. Luo, Mar. Drugs, 2024, 22, 521, DOI:10.3390/md22110521.
- Q. Dai, M. Ma, N. Wang, Y. Zhou and Z. Zhang, Fitoterapia, 2024, 178, 106176, DOI:10.1016/j.fitote.2024.106176.
- S. Pulat, I. Yang, J. Lee, S. Hwang, R. Zhou, C. D. B. Gamage, M. Varlı, İ. Taş, Y. Yang, S.-Y. Park, A. Hong, J.-H. Kim, D.-C. Oh, H. Kim, S.-J. Nam and H. Kang, Mar. Drugs, 2024, 22, 88, DOI:10.3390/md22020088.
- W. Liu, S. Zhai, L. Zhang, Y. Chen, Z. Liu, W. Ma, T. Zhang, W. Zhang, L. Ma, C. Zhang and W. Zhang, J. Nat. Prod., 2024, 87, 371–380, DOI:10.1021/acs.jnatprod.3c01132.
- S. Liu, C. Sun, Y. Ha, M. Ma, N. Wang, Y. Zhou and Z. Zhang, Tetrahedron Lett., 2024, 137, 154935, DOI:10.1016/j.tetlet.2024.154935.
- S. Tan, Y. Liu, H. Fu, Y. Xue, F. Feng, J. Li, X. Yan, S. Wu, S. He and L. Ding, J. Oceanol. Limnol., 2024, 42, 1991–1997, DOI:10.1007/s00343-024-3195-9.
- H. P. S. d. F. Cesário, F. C. O. Silva, M. K. A. Ferreira, J. E. S. A. de Menezes, H. S. dos Santos, A. M. da Fonseca, C. E. S. Nogueira, M. M. Marinho, E. S. Marinho, A. M. R. Teixeira, E. R. Silveira and O. D. L. Pessoa, J. Biomol. Struct. Dyn., 2024, 42, 445–460, DOI:10.1080/07391102.2023.2193988.
- L. Xu, R. Liu, J. Fang, N. Zhang, F. Pu, Z. Lei, W. Ding and Y. Jiang, Chem. Biodiversity, 2024, 21, e202301712, DOI:10.1002/cbdv.202301712.
- D. Sweeney, A. Bogdanov, A. B. Chase, G. Castro-Falcón, A. Trinidad-Javier, S. Dahesh, V. Nizet and P. R. Jensen, J. Nat. Prod., 2024, 87, 2768–2778, DOI:10.1021/acs.jnatprod.4c00934.
- K. Hosoda, N. Koyama, S. Shigeno, T. Nishimura, N. Hasegawa, A. Kanamoto, T. Ohshiro, H. Tomoda and C. A. Arias, Antimicrob. Agents Chemother., 2024, 68, e00917–e00923, DOI:10.1128/aac.00917-23.
- H. Satoh, D. Lee, S. Arima, K. Hosoda, S. Shigeno, H. Tomoda, T. Ohshiro and T. Nagamitsu, Tetrahedron, 2024, 168, 134299, DOI:10.1016/j.tet.2024.134299.
- J.-H. Kim, J. Y. Lee, J. Lee, P. F. Hillman, J. Lee, B. Choi, M.-J. Paik, S. Lee and S.-J. Nam, Molecules, 2024, 29, 5539, DOI:10.3390/molecules29235539.
- W. Wang, J. Lee, E. Roh, G. Shetye, J. Cao, J. McAlpine, G. Pauli, S. Franzblau, T. Hanh Nguyen Vu, N. T. Quach, E. Oh, K.-H. Park, C. Park, Y. Cho, H. Jang, S. Han, H. Kim, S. Cho, Q.-T. Phi and H. Kang, J. Nat. Prod., 2024, 87, 976–983, DOI:10.1021/acs.jnatprod.3c01275.
- Z. A. Maw, A. L. Grunwald, B. A. Haltli, C. Cartmell and R. G. Kerr, J. Nat. Prod., 2024, 87, 1682–1693, DOI:10.1021/acs.jnatprod.3c01234.
- T. Wu, W. M. Hussein, K. Samarasekera, Y. Zhu, Z. G. Khalil, S. Jin, D. F. Bruhn, Y. Moreno, A. A. Salim and R. J. Capon, Mar. Drugs, 2024, 22, 312, DOI:10.3390/md22070312.
- Z. A. Maw, B. Haltli, J. J. Guo, D. M. Baldisseri, C. Cartmell and R. G. Kerr, Molecules, 2024, 29, 1482, DOI:10.3390/molecules29071482.
- S. Zhao, P. Yan, P. Liu, H. Liu, N. Li, P. Fu and W. Zhu, Chin. Chem. Lett., 2024, 35, 108950, DOI:10.1016/j.cclet.2023.108950.
- Q. Sun, D. Yu, X. Zhang, F. Xiao and W. Li, Mar. Drugs, 2024, 22, 262, DOI:10.3390/md22060262.
- J. H. Lee, B.-K. Choi, M. Kim, H. J. Shin and S. J. Park, Mar. Drugs, 2024, 22, 533, DOI:10.3390/md22120533.
- C.-S. Heo, J. S. Kang, J.-W. Yang, M. A. Lee, H.-S. Lee and H. J. Shin, Int. J. Mol. Sci., 2024, 25, 3311, DOI:10.3390/ijms25063311.
- N. Takatani, T. Maoka, T. Sawabe, F. Beppu and M. Hosokawa, Appl. Microbiol. Biotechnol., 2024, 108, 102, DOI:10.1007/s00253-023-12995-2.
- Y. Song, Y. Zhou, M. Cong, S. Deng, Y. Chen, X. Pang, Y. Liu, L. Liao, L. Yang and J. Wang, Mar. Drugs, 2024, 22, 484, DOI:10.3390/md22110484.
- Y. Xu, Y. Song, Y. Ning, S. Li, Y. Qu, B. Jiao and X. Lu, Mar. Drugs, 2024, 22, 331, DOI:10.3390/md22080331.
- H.-W. Zhao, Y. Yi, J.-Q. Li and J.-H. Wang, Nat. Prod. Res., 2024, 38, 206–210, DOI:10.1080/14786419.2022.2112578.
- R. Campbell, L. Kyei, K. Piedl, Z. Zhang, M. Chen and E. Mevers, Org. Lett., 2024, 26, 9693–9697, DOI:10.1021/acs.orglett.4c03470.
- R. Feng, Q. Dai, M. Ma, S. Zhao, Z. Wang, H. Wang, Y. Zhang, L. Huo and F. Yan, J. Nat. Prod., 2024, 87, 2801–2809, DOI:10.1021/acs.jnatprod.4c00965.
- D. Roman, P. Meisinger, R. Guillonneau, C. Peng, L. K. Peltner, P. M. Jordan, V. Haensch, S. Götze, O. Werz, C. Hertweck, Y. Chen and C. Beemelmanns, Angew. Chem., Int. Ed., 2024, 63, e202401195, DOI:10.1002/anie.202401195.
- X. Zha, Y. Li, H. Zhao, Y. Tan and S. Zhou, Molecules, 2024, 29, 4446, DOI:10.3390/molecules29184446.
- N. Sermkaew, A. Atipairin, S. Krobthong, C. Aonbangkhen, Y. Yingchutrakul, J. Uchiyama and N. Songnaka, Antibiotics, 2024, 13, 846, DOI:10.3390/antibiotics13090846.
- M. Girão, S. Freitas, T. P. Martins, R. Urbatzka, M. F. Carvalho and P. N. Leão, Front. Pharmacol, 2024, 15, 1347485, DOI:10.3389/fphar.2024.1347485.
- P. Li, Y. Wu, S. He, H. Lu, S. He and L. Liu, Chem. Nat. Compd., 2024, 60, 831–834, DOI:10.1007/s10600-024-04458-5.
- R. Ahmad, M. I. Tousif, M. Nazir, A. Yaqoob, S. A. Ali Shah, G. Zengin, A. I. Uba and S. Muhammad, Food Biosci., 2024, 57, 103600, DOI:10.1016/j.fbio.2024.103600.
- A. Buljan, V. Stepanić, A. Čikoš, S. B. Brčić, K. Bojanić and M. Roje, Mar. Drugs, 2024, 22, 310, DOI:10.3390/md22070310.
- R. R. Khatua, S. A. Z. Ahmad, S. Das, R. Anindya and F. A. Khan, Synlett, 2024, 35, 1872–1876, DOI:10.1055/a-2256-5509.
- S. S. Auddy, S. Gupta, S. Mandi, H. Sharma, S. Sinha and R. K. Goswami, ACS Med. Chem. Lett., 2024, 15, 1340–1350, DOI:10.1021/acsmedchemlett.4c00237.
- T. C. Murgatroyd, T. Söhnel and J. Sperry, Org. Lett., 2024, 26, 10041–10045, DOI:10.1021/acs.orglett.4c03937.
- A. Dimasi, M. Failla, A. Montoli, A. Citarella, P. Ronchi, D. Passarella and V. Fasano, RSC Adv., 2024, 14, 5542–5546, 10.1039/D4RA00589A.
- L. Yu, Y. Nagata and H. Nakamura, J. Am. Chem. Soc., 2024, 146, 2549–2555, DOI:10.1021/jacs.3c11016.
- F. Fei, S. Lun, A. Saxena, M. Raghavan, J. L. DeRisi, W. R. Bishai and J. K. Sello, Org. Lett., 2024, 26, 9698–9703, DOI:10.1021/acs.orglett.4c03473.
- G. G. Dake and K. P. Kaliappan, J. Org. Chem., 2024, 89, 5825–5832, DOI:10.1021/acs.joc.4c00462.
- S. Niyogi, A. Mondal, M. Nandy, S. Pal, A. Khatua and A. Bisai, Org. Lett., 2024, 26, 8643–8647, DOI:10.1021/acs.orglett.4c02436.
- G. Petruncio, Z. Shellnutt, L. L. Young, M. Girgis, W. K. Strangman, R. T. Williamson, K. Kehn-Hall and M. Paige, ACS Omega, 2024, 9, 30993–30997, DOI:10.1021/acsomega.4c04249.
- D.-B. Sung, D.-B. Choi, J. H. Seol, N. Kang, E.-A. Kim, S.-Y. Heo, S.-J. Heo and J. S. Lee, ACS Omega, 2024, 9, 27592–27609, DOI:10.1021/acsomega.4c03241.
- F. Han, J. Li, S. Li, Z. Wang, Y. Guo and T. Ye, Angew. Chem., Int. Ed., 2024, 63, e202317636, DOI:10.1002/anie.202317636.
- P. Karamanis, J. Muldoon, C. D. Murphy and M. Rubini, J. Pept. Sci., 2024, 30, e3569, DOI:10.1002/psc.3569.
- S. Kreeger, B. Sarrels and C. Lucero, Tetrahedron, 2024, 163, 134139, DOI:10.1016/j.tet.2024.134139.
- A. F. Anwar, Y. M. Elbatrawi, D. Degen, R. H. Ebright and J. R. Del Valle, Org. Lett., 2024, 26, 10056–10060, DOI:10.1021/acs.orglett.4c04013.
- C. Prudel and U. Kazmaier, Synlett, 2024, 35, 2133–2137, DOI:10.1055/s-0042-1751578.
- J. Wu and M. J. Krische, Org. Lett., 2024, 26, 4830–4834, DOI:10.1021/acs.orglett.4c01644.
- M. J. Bedding, B. C. Forster, A. M. Giltrap, M. T. Stevens, L. Corcilius, W. J. Britton and R. J. Payne, Org. Lett., 2024, 26, 10993–10998, DOI:10.1021/acs.orglett.4c04163.
- K. Gan, Y. Zhu, G. Shi, C. Wu, F. Ni, L. Zhao, X. Dou, Z. Zhang and H. Lu, Chin. J. Chem., 2024, 43, 79–89, DOI:10.1002/cjoc.202400751.
- S. M. Astle, S. Guggiari, J. R. Frost, H. B. Hepburn, D. J. Klauber, K. E. Christensen and J. W. Burton, J. Am. Chem. Soc., 2024, 146, 17757–17764, DOI:10.1021/jacs.4c02969.
- S. Wang, X. Li, W. Yang and R. Huang, Microb. Biotechnol., 2024, 17, e14533, DOI:10.1111/1751-7915.14533.
- C.-H. Ji, H.-W. Je, H. Kim and H.-S. Kang, Nat. Prod. Rep., 2024, 41, 672–699, 10.1039/D3NP00049D.
- H. Put, H. Gerstmans, H. V. Capelle, M. Fauvart, J. Michiels and J. Masschelein, Nat. Prod. Rep., 2024, 41, 1113–1151, 10.1039/D3NP00065F.
- M. Gonzales, P. Jacquet, F. Gaucher, É. Chabrière, L. Plener and D. Daudé, J. Nat. Prod., 2024, 87, 1268–1284, DOI:10.1021/acs.jnatprod.3c00672.
- W.-C. Chin, Y.-Z. Zhou, H.-Y. Wang, Y.-T. Feng, R.-Y. Yang, Z.-F. Huang and Y.-L. Yang, Nat. Prod. Rep., 2024, 41, 977–989, 10.1039/D3NP00059A.
- Z. Liu, W. Sun, Z. Hu, W. Wang and H. Zhang, Mar. Drugs, 2024, 22, 51, DOI:10.3390/md22010051.
- R. T. Magar and J. K. Sohng, Appl. Microbiol. Biotechnol., 2024, 108, 471, DOI:10.1007/s00253-024-13296-y.
- D. Sweeney, A. B. Chase, A. Bogdanov and P. R. Jensen, J. Nat. Prod., 2024, 87, 439–452, DOI:10.1021/acs.jnatprod.3c01007.
- A. L. B. Canellas and M. S. Laport, Lett. Appl. Microbiol., 2024, 77, ovae099, DOI:10.1093/lambio/ovae099.
- L. T. H. Diethelm, A. B. da Silva Bellini Ramos, G. B. de Lorena, B. I. Trajano, R. D. do Espírito Santo, R. P. B. de Menezes, M. T. Scotti, F. A. Colombo, M. J. Marques, C. R. D. Correia and J. Q. Reimão, Pharmaceutics, 2024, 16, 262, DOI:10.3390/pharmaceutics16020262.
- G. Castro-Falcón, J. Straetener, J. Bornikoel, D. Reimer, T. N. Purdy, A. Berscheid, F. M. Schempp, D. Y. Liu, R. G. Linington, H. Brötz-Oesterhelt and C. C. Hughes, ACS Chem. Biol., 2024, 19, 743–752, DOI:10.1021/acschembio.3c00773.
- T. L. de Brito, E. A. Edson, K. G. D. Florêncio, J. A. Machado-Neto, A. D. M. B. Garnique, J. P. M. Luiz, F. de Queiroz Cunha, J. C. Alves-Filho, M. Haygood and D. V. Wilke, Chem.-Biol. Interact., 2024, 400, 111177, DOI:10.1016/j.cbi.2024.111177.
- J. Lee, J.-Y. Hwang, D. Oh, D.-C. Oh, H.-G. Park, J. Shin and K.-B. Oh, Mar. Drugs, 2024, 22, 293, DOI:10.3390/md22070293.
- R. M. Johnson, K. Li, X. Chen, G. L. Morgan, J. Aubé and B. Li, ACS Infect. Dis., 2024, 10, 582–593, DOI:10.1021/acsinfecdis.3c00504.
- A. Chatterjee, J. Mondal, S. Paul, H. Sharma, R. K. Goswami and P. Sen, J. Med. Chem., 2024, 67, 21952–21974, DOI:10.1021/acs.jmedchem.4c01902.
- K. Irie, G. Jeelani, T. Nozaki and A. Iwasaki, J. Nat. Prod., 2024, 87, 2292–2301, DOI:10.1021/acs.jnatprod.4c00733.
- N. E. Avalon, M. A. Reis, C. C. Thornburg, R. T. Williamson, D. Petras, A. T. Aron, G. F. Neuhaus, M. Al-Hindy, J. Mitrevska, L. Ferreira, J. Morais, Y. E. Abiead, E. Glukhov, K. L. Alexander, F. A. Vulpanovici, M. J. Bertin, S. Whitner, H. Choi, G. Spengler, K. Blinov, A. M. Almohammadi, L. A. Shaala, W. R. Kew, L. Paša-Tolić, D. T. A. Youssef, P. C. Dorrestein, V. Vasconcelos, L. Gerwick, K. L. McPhail and W. H. Gerwick, J. Am. Chem. Soc., 2024, 146, 18626–18638, DOI:10.1021/jacs.4c05399.
- F. H. Al-Awadhi, S. Kokkaliari, R. Ratnayake, V. J. Paul and H. Luesch, J. Nat. Prod., 2024, 87, 2355–2365, DOI:10.1021/acs.jnatprod.4c00420.
- S. Kokkaliari, L. Grauso, A. Mangoni, G. Seabra, V. J. Paul and H. Luesch, Chemistry, 2024, 30, e202401393, DOI:10.1002/chem.202401393.
- T.-E. Ngo, A. Ecker, B. Ryu, A. Guild, A. Remmel, P. D. Boudreau, K. L. Alexander, C. B. Naman, E. Glukhov, N. E. Avalon, V. V. Shende, L. Thomas, S. Dahesh, V. Nizet, L. Gerwick and W. H. Gerwick, ACS Chem. Biol., 2024, 19, 619–628, DOI:10.1021/acschembio.3c00391.
- B. Ryu, E. Glukhov, T. R. Teixeira, C. R. Caffrey, S. Madiyan, V. Joseph, N. E. Avalon, C. A. Leber, C. B. Naman and W. H. Gerwick, J. Nat. Prod., 2024, 87, 1601–1610, DOI:10.1021/acs.jnatprod.4c00242.
- N. Kurisawa, G. Jeelani, T. Nozaki, A. L. Agusta, K. Suenaga and A. Iwasaki, J. Nat. Prod., 2024, 87, 1838–1843, DOI:10.1021/acs.jnatprod.4c00499.
- R. Fernández, M. Pérez, A. Losada, S. Reboredo, A. G.-S. Juan, M. J. Martín, A. Francesch, S. Munt and C. Cuevas, Mar. Drugs, 2024, 22, 303, DOI:10.3390/md22070303.
- J. J. Mehjabin, C.-S. Phan and T. Okino, J. Nat. Prod., 2024, 87, 984–993, DOI:10.1021/acs.jnatprod.3c01272.
- K. Umeda, N. Kurisawa, G. Jeelani, T. Nozaki, K. Suenaga and A. Iwasaki, Beilstein J. Org. Chem., 2024, 20, 645–652, DOI:10.3762/bjoc.20.57.
- M. Yotsu-Yamashita, R. L. Haddock and T. Yasumoto, J. Am. Chem. Soc., 2002, 115, 1147–1148, DOI:10.1021/ja00056a048.
- M. Yotsu-Yamashita, T. Seki, V. J. Paul, H. Naoki and T. Yasumoto, Tetrahedron Lett., 1995, 36, 5563–5566, DOI:10.1016/0040-4039(95)01052-J.
- M. Yotsu-Yamashita, K. Abe, T. Seki, K. Fujiwara and T. Yasumoto, Tetrahedron Lett., 2007, 48, 2255–2259, DOI:10.1016/j.tetlet.2007.02.003.
- G. Navarro, M. E. Cummings, J. Lee, N. Moss, E. Glukhov, F. A. Valeriote, L. Gerwick and W. H. Gerwick, Environ. Sci. Technol. Lett., 2015, 2, 166–170, DOI:10.1021/acs.estlett.5b00116.
- M. Yotsu-Yamashita, K. Umeda, A. Iwasaki and K. Suenaga, Toxicon, 2024, 250, 108122, DOI:10.1016/j.toxicon.2024.108122.
- B.-T. Zhang, H. Nishino, R. Kawabe, M. Kamio, R. Watanabe, H. Uchida, M. Satake and H. Nagai, Biosci. Biotechnol. Biochem., 2024, 88, 517–521, DOI:10.1093/bbb/zbae015.
- B. Zhang, R. Kawabe, M. Kamio, H. Uchida, N. Matsuura, K. Mori-Yasumoto, M. Satake and H. Nagai, Tetrahedron Lett., 2024, 150, 155284, DOI:10.1016/j.tetlet.2024.155284.
- N. Watanabe, G. Jeelani, T. Nozaki and A. Iwasaki, ACS Omega, 2024, 9, 36795–36801, DOI:10.1021/acsomega.4c05909.
- A. Ebihara, R. Taguchi, G. Jeelani, T. Nozaki, K. Suenaga and A. Iwasaki, J. Nat. Prod., 2024, 87, 1116–1123, DOI:10.1021/acs.jnatprod.4c00056.
- L. A. Elsadek, E. K. Ellis, G. Seabra, V. J. Paul and H. Luesch, Mar. Drugs, 2023, 22, 28, DOI:10.3390/md22010028.
- J. P. A. Reis, S. Freitas, T. Prochazkova and P. N. Leão, J. Nat. Prod., 2024, 87, 2709–2715, DOI:10.1021/acs.jnatprod.4c00832.
- M. L. Matos-Hernández, R. Samples, G. Dyer, V. M. Casimir Montán, C. A. Morales-Colón, J. M. Salvino, L. J. Montaner, J. A. Cassel, T. E. Messick, I. Tietjen and E. J. E. Caro-Diaz, J. Nat. Prod., 2024, 87, 1513–1520, DOI:10.1021/acs.jnatprod.3c01071.
- K. Hagimoto, S. Tojo, T. Teruya, M. Yoshida and H. Kigoshi, Tetrahedron, 2024, 154, 133871, DOI:10.1016/j.tet.2024.133871.
- A. Tripathi, J. Puddick, M. R. Prinsep, M. Rottmann, K. P. Chan, D. Y. Chen and L. T. Tan, Phytochemistry, 2011, 72, 2369–2375, DOI:10.1016/j.phytochem.2011.08.019.
- K. Sueyoshi, M. Kaneda, S. Sumimoto, S. Oishi, N. Fujii, K. Suenaga and T. Teruya, Tetrahedron, 2016, 72, 5472–5478, DOI:10.1016/j.tet.2016.07.031.
- B. P. Smith, N. J. Truax, A. S. Pollatos, M. Meanwell, P. Bedekar, A. F. Garrido-Castro and P. S. Baran, Angew. Chem., Int. Ed., 2024, 63, e202401107, DOI:10.1002/anie.202401107.
- T. Suwa, M. Sasaki and A. Umehara, Org. Lett., 2024, 26, 4377–4382, DOI:10.1021/acs.orglett.4c01367.
- R. Shigeta, T. Suzuki, K. Kaneko, H. Tanaka, I. Haishima, K. Norio, A. Tanaka-Yanuma and T. Usuki, Org. Biomol. Chem., 2024, 22, 4637–4640, 10.1039/d4ob00580e.
- X. Zhao, M. Lv, X. Xi, Y. Lu, L. Wang and Y. Chen, Chin. J. Chem., 2023, 42, 605–610, DOI:10.1002/cjoc.202300526.
- K. Tokushige and T. Abe, Chem.–Eur. J., 2024, 30, e202302963, DOI:10.1002/chem.202302963.
- S. B. Waghmode and A. R. Jadhao, Synthesis, 2023, 56, 1407–1414, DOI:10.1055/a-2222-3822.
- K. Annapurna and A. Venkat Narsaiah, Tetrahedron Lett., 2024, 142, 155081, DOI:10.1016/j.tetlet.2024.155081.
- L. T. Tan and N. F. Salleh, Molecules, 2024, 29, 5307, DOI:10.3390/molecules29225307.
- Â. P. Matos, F. M. P. Saldanha-Corrêa, R. D. S. Gomes and G. R. Hurtado, Acta Trop., 2024, 251, 107116, DOI:10.1016/j.actatropica.2023.107116.
- A. Bouyahya, S. Bakrim, I. Chamkhi, D. Taha, N. El Omari, N. El Mneyiy, N. El Hachlafi, M. El-Shazly, A. Khalid, A. N. Abdalla, K. W. Goh, L. C. Ming, B. H. Goh and T. Aanniz, Biomed. Pharmacother., 2024, 170, 115989, DOI:10.1016/j.biopha.2023.115989.
- A. K. Bishoyi, C. P. Mandhata, C. R. Sahoo, S. K. Paidesetty and R. N. Padhy, Naunyn-Schmiedeberg's Arch. Pharmacol., 2024, 397, 1347–1375, DOI:10.1007/s00210-023-02719-8.
- D. Kallifidas, D. Dhakal, M. Chen, Q. Y. Chen, S. Kokkaliari, N. A. Colon Rosa, R. Ratnayake, S. D. Bruner, V. J. Paul, Y. Ding and H. Luesch, Org. Lett., 2024, 26, 1321–1325, DOI:10.1021/acs.orglett.3c04083.
- P. Mandal, D. Paul, H. Sharma, S. Saha, P. Chakrabarti and R. K. Goswami, ACS Med. Chem. Lett., 2024, 15, 696–705, DOI:10.1021/acsmedchemlett.4c00094.
- L. J. Liu, K. R. Francisco, Y. U. Sun, M. S. M. Serafim, D. K. Amarasinghe, T. R. Teixeira, B. Lucero, T. Kronenberger, W. Elsayed, H. Elwakeel, M. Al-Hindy, J. Almaliti, W. H. Gerwick, A. J. O'Donoghue and C. R. Caffrey, ACS Infect. Dis., 2024, 10, 4182–4193, DOI:10.1021/acsinfecdis.4c00441.
- N. Kurisawa, K. Teranuma, A. Noto, A. Iwasaki, Y. Kabashima, R. Nakajima, C. Toyoshima and K. Suenaga, Bull. Chem. Soc. Jpn., 2024, 97, uoae070, DOI:10.1093/bulcsj/uoae070.
- X. Zhao, X. Xi, M. Zhang, M. Lv, X. Zhang, Y. Lu, L. Wang and Y. Chen, Mar. Drugs, 2024, 22, 537, DOI:10.3390/md22120537.
- É. C. de Almeida, L. S. Passos, P. N. N. de Freitas, A. O. de Souza and E. Pinto, Aquac. Res., 2024, 16, 1186–1211, DOI:10.1111/raq.12891.
- L. Biessy, J. Puddick, S. A. Wood, A. I. Selwood, M. Carbines and K. F. Smith, Toxins, 2024, 16, 522, DOI:10.3390/toxins16120522.
- X. Hao, S. Li, J. Li, G. Wang, J. Li, Z. Peng and M. Gan, J. Nat. Prod., 2024, 87, 1059–1066, DOI:10.1021/acs.jnatprod.4c00015.
- Y. Zhang, J.-B. Zhou, S.-T. Yang, X. Liu, W. Cao, P.-H. Li, H. Chen and Y.-Q. Fan, Mar. Drugs, 2024, 22, 25, DOI:10.3390/md22010025.
- C.-L. An, Q.-Y. Ma, Q.-Y. Xie, L. Yang, J.-Z. Yuan, H.-F. Dai and Y.-X. Zhao, J. Asian Nat. Prod. Res., 2024, 26, 489–496, DOI:10.1080/10286020.2023.2251114.
- Z. Luo, K. Liu, Z. Tang, L. Peng, C. Gao, C. Xia, Y. Liu and X. Chen, Rec. Nat. Prod., 2024, 18, 136–142, DOI:10.25135/rnp.438.12310.2940.
- Y.-H. Kim, D.-C. Choi, D.-W. Ki, Y.-S. Won, S.-J. Lee, J.-Y. Kim, I.-K. Lee and B.-S. Yun, J. Antibiot., 2024, 77, 466–470, DOI:10.1038/s41429-024-00735-1.
- S.-T. Fang, Y.-P. Song, Z.-Z. Shi and N.-Y. Ji, Phytochem. Lett., 2024, 61, 218–221, DOI:10.1016/j.phytol.2024.05.006.
- J.-J. Xue, F.-P. Miao and S.-T. Fang, Nat. Prod. Res., 2024, 38, 2480–2485, DOI:10.1080/14786419.2023.2183500.
- Y.-S. Lee, H.-C. Wu, S.-J. Huang, G. Hsiao, W.-C. Chi and T.-H. Lee, Phytochemistry, 2024, 219, 113998, DOI:10.1016/j.phytochem.2024.113998.
- Y. Hu, X. Zhao, Y. Song, J. Jiang, T. Long, M. Cong, Y. Miao, Y. Liu, Z. Yang, Y. Zhu and J. Wang, J. Nat. Prod., 2024, 87, 1975–1982, DOI:10.1021/acs.jnatprod.4c00393.
- P. P. Edjah, M.-F. Lu, W.-C. Chen, J. Wu, Y.-T. Zhong, M. Li, Q. Chen, K.-K. Zhu, R.-Y. Yuan, W.-X. Tao, K.-J. Wu and Y.-S. Cai, Fitoterapia, 2024, 179, 106251, DOI:10.1016/j.fitote.2024.106251.
- P. E. Mandelare, S. S. Yee, D. G. Icenhour, E. N. Kaweesa, M. McCauley, A. L. Risinger, J. D. Rudolf and S. Loesgen, ChemBioChem, 2024, 25, e202400398, DOI:10.1002/cbic.202400398.
- Y. Zeng, S. Wang, H. Peng, W. Zhao, W. Chang, H. Wang, H. Chen and H. Dai, Mar. Drugs, 2024, 22, 13, DOI:10.3390/md22010013.
- C.-J. Lu, L.-F. Liang, G.-S. Zhang, H.-Y. Li, C.-Q. Fu, Q. Yu, D.-M. Zhou, Z.-W. Su, K. Liu, C.-H. Gao, X.-Y. Xu and Y.-H. Liu, Mar. Drugs, 2024, 22, 63, DOI:10.3390/md22020063.
- D. H. El-Kashef, D. D. Obidake, K. Schiedlauske, A. Deipenbrock, S. Scharf, H. Wang, D. Naumann, D. Friedrich, S. Miljanovic, T. Haj Hassani Sohi, C. Janiak, K. Pfeffer and N. Teusch, Mar. Drugs, 2024, 22, 5, DOI:10.3390/md22010005.
- H. R. A. Cardona, B. C. da Silva, F. O. de Lima, F. H. A. Leite, B. C. de Souza, H. N. Brandão, J. M. David, C. Q. Alves and A. Kijjoa, Mar. Drugs, 2024, 22, 97, DOI:10.3390/md22030097.
- E. Doro-Goldsmith, Q. Song, X.-L. Li, X.-M. Li, X.-Y. Hu, H.-L. Li, H.-R. Liu, B.-G. Wang and H. Sun, J. Nat. Prod., 2024, 87, 381–387, DOI:10.1021/acs.jnatprod.3c01157.
- Q. Li, S. Li, S. Li, X. Hao, A. Wang, S. Si, Y. Xu, J. Shu and M. Gan, J. Nat. Prod., 2024, 87, 365–370, DOI:10.1021/acs.jnatprod.3c01115.
- S. Kankanamge, P. V. Bernhardt, Z. G. Khalil and R. J. Capon, Mar. Drugs, 2024, 22, 243, DOI:10.3390/md22060243.
- Y. He, T. Yang, J. Li, K. Li, C. Zhuang, M. Zhang, R. Li, Y. Zhao, Q. Song, M. Jiang, S. Mao, X. Song, Y. Guo, X. Li, F. Tan, S. Jitkaew, W. Zhang and Z. Cai, Br. J. Pharmacol., 2024, 181, 2443–2458, DOI:10.1111/bph.16364.
- Z.-B. Zou, Y. Li, Y. Wang, C.-L. Xie, Z.-Q. Li, S.-S. Nie, Y. Li, S.-Y. Fang, T.-H. Zhong, L.-S. Li and X.-W. Yang, Org. Lett., 2024, 26, 5695–5699, DOI:10.1021/acs.orglett.4c01745.
- S. Niu, S. Huang, J. Wang, J. He, M. Chen, G. Zhang and B. Hong, Nat. Prod. Res., 2024, 38, 1362–1368, DOI:10.1080/14786419.2022.2144299.
- Y.-H. Che, W.-P. Ding, Z.-H. Xiao, J.-M. Wu, H. Yin, F.-Z. Wang and S. Zhang, Mar. Drugs, 2024, 22, 117, DOI:10.3390/md22030117.
- X. Zhang, Y. Dong, X. Liu, R. Wang, J. Lu and F. Song, Nat. Prod. Res., 2024, 38, 2792–2799, DOI:10.1080/14786419.2023.2236764.
- X. Shi, X. Li, X. He, D. Zhang, C. Quan, Z. Xiu and Y. Dong, Molecules, 2024, 29, 670, DOI:10.3390/molecules29030670.
- R. Ge, X. Guo, H. Jia, J. Zhang, A. Fan, D. Liu, J. Huang and W. Lin, J. Org. Chem., 2024, 89, 7692–7704, DOI:10.1021/acs.joc.4c00367.
- Y. Chen, S.-P. Wang, L.-C. Xu, C. Liang, G.-D. Liu, X. Ji, W.-H. Luo, S. Liu, Z.-X. Zhang and G.-Y. Cao, Fitoterapia, 2024, 172, 105763, DOI:10.1016/j.fitote.2023.105763.
- J. Zhang, L. Gao, H. Lin, Y. Liang, M. You, L. Ding, F. Feng, B. Yang and Y. Liu, J. Agric. Food Chem., 2024, 72, 12596–12606, DOI:10.1021/acs.jafc.4c02769.
- Z. Zhang, Y. Li, H. Wang, W. Xu, C. Wang, H. Ma, F. Zhong, J. Ou, Z. Luo, H.-B. Luo and Z. Cheng, J. Nat. Prod., 2024, 87, 1563–1573, DOI:10.1021/acs.jnatprod.4c00154.
- G.-Y. Zhang, B.-J. Liu, H.-L. Pan, H.-J. Li, Z.-F. Huang, T. Mahmud, W.-Z. Ma and W.-J. Lan, Fitoterapia, 2024, 175, 105967, DOI:10.1016/j.fitote.2024.105967.
- H. Fan, X.-H. Shao, P. Wu, A.-L. Hao, Z.-W. Luo, M.-D. Zhang, J. Xie, B. Peng and C.-X. Zhang, J. Agric. Food Chem., 2024, 72, 19869–19882, DOI:10.1021/acs.jafc.4c04728.
- H. Cui, Y. Tang, C. Yang, H. Deng, L. Chen, X. Fan, L. Zhu, Y. Liu, Z. Zhao and T. Su, Phytochemistry, 2024, 219, 113983, DOI:10.1016/j.phytochem.2024.113983.
- Y. Zhou, X. Chen, Y. Tang, Y. Liu, Z. Zhao and H. Cui, Nat. Prod. Res., 2024, 38, 1334–1340, DOI:10.1080/14786419.2022.2140338.
- Y.-Y. Zheng, J.-Q. Mao, Y. Liu, N. Han, L. Lv, Y.-H. Zhang, M. Chen, Z.-Q. Liu, C.-L. Shao, G.-S. Yao and C.-Y. Wang, Bioorg. Chem., 2024, 153, 107832, DOI:10.1016/j.bioorg.2024.107832.
- Y.-P. He, Z.-K. Zhang, Z.-J. Li, P.-P. Wu, J.-S. Hu, H. Fan and C.-X. Zhang, Fitoterapia, 2024, 177, 106136, DOI:10.1016/j.fitote.2024.106136.
- H.-M. Wen, Y.-W. Zhang, F.-J. Feng, G.-B. Huang, Y.-H. Lv, Z.-Y. Zhang and L.-J. Ding, J. Asian Nat. Prod. Res., 2024, 26, 548–554, DOI:10.1080/10286020.2023.2259317.
- N. T. Ngoc, T. H. Quang, T. T. H. Hanh, N. X. Cuong, V. T. Quyen, N. T. T. Ngan, D. V. Ha, N. H. Nam and C. V. Minh, Phytochem. Lett., 2024, 61, 29–34, DOI:10.1016/j.phytol.2024.03.010.
- K. Anjum, X. Huang, L. Zhou, T. Zhu, Q. Che, G. Zhang and D. Li, Nat. Prod. Res., 2024, 38, 3231–3236, DOI:10.1080/14786419.2023.2227754.
- C. Sun, Y. Ha, X. Liu, N. Wang, X.-Y. Lian and Z. Zhang, Molecules, 2024, 29, 459, DOI:10.3390/molecules29020459.
- Y. Ha, Y. Zhou, M. Ma, N. Wang, P. Wang and Z. Zhang, Phytochemistry, 2024, 224, 114164, DOI:10.1016/j.phytochem.2024.114164.
- Y.-H. Chen, Q. Zhu, J. Li, R. Yang, J. Zhang, M. You, L. Luo and B. Yang, Metabolites, 2024, 14, 99, DOI:10.3390/metabo14020099.
- A. Abulaizi, R. Wang, Z. Xiong, S. Zhang, Y. Li, H. Ge and Z. Guo, Mar. Drugs, 2024, 22, 270, DOI:10.3390/md22060270.
- C. Li, X. Lin, S. Wang, D. Guan, X. Wang, B. Yang, X. Zhou, J. Li, B. Xiong, Y. Liu and Y. Sun, Chem. Biodiversity, 2024, 21, e202301706, DOI:10.1002/cbdv.202301706.
- X. Xu, J. Han, X. Zhang, W. Xu, J. Yang and F. Song, Nat. Prod. Res., 2024, 38, 1369–1374, DOI:10.1080/14786419.2022.2144300.
- P. Lu, Y. Shi, J. Zhang, K. Hong, Y. Xue and L. Liu, Int. J. Biol. Macromol., 2024, 257, 128808, DOI:10.1016/j.ijbiomac.2023.128808.
- D. Tian, M. Chen, W. Ding, J. Li, S. Wu, K. Hong and J. Tang, Nat. Prod. J., 2024, 14, e031023221602, DOI:10.2174/0122103155274328230926160728.
- J. Xue, X. Guo, G. Xu, X. Chen, L. Jiao and X. Tang, J. Agric. Food Chem., 2024, 72, 2989–2996, DOI:10.1021/acs.jafc.3c07826.
- H.-Y. Dong, Y. Wang, X.-Y. Zhang, M. Zhang, L.-H. Yang, Z.-B. Zou, Y. Li, M.-M. Xie, X.-W. Yang and B. Wang, Chem. Biodiversity, 2024, 21, e202400507, DOI:10.1002/cbdv.202400507.
- G.-J. Ran, P. Wang, H.-K. Wang, T.-T. Wei, Y.-Y. Liu, H.-X. Jin, T.-T. Wang and S. He, Chem. Biodiversity, 2024, 21, e202401689, DOI:10.1002/cbdv.202401689.
- S.-T. Fang, F.-P. Miao, X.-L. Yin and N.-Y. Ji, Nat. Prod. Res., 2024, 38, 1510–1516, DOI:10.1080/14786419.2022.2154345.
- X.-D. Li, X.-M. Li, B.-G. Wang and X. Li, Org. Biomol. Chem., 2024, 22, 3979–3985, 10.1039/D4OB00449C.
- X.-X. Miao, L.-L. Hong, H.-Y. Liu, R.-Y. Shang, W.-H. Jiao, S.-H. Xu and H.-W. Lin, Chem. Biodiversity, 2024, 21, e202400832, DOI:10.1002/cbdv.202400832.
- X. Miao, L. Hong, Z. Ju, H. Liu, R. Shang, P. Li, K. Liu, B. Cheng, W. Jiao, S. Xu and H. Lin, ACS Omega, 2024, 9, 22450–22458, DOI:10.1021/acsomega.4c02488.
- Q. Gao, Y. Shi, H. Wen, D. Zhang, X. Yan, S. He and L. Ding, J. Mol. Struct., 2024, 1296, 136723, DOI:10.1016/j.molstruc.2023.136723.
- C.-P. Li, Y.-P. Song, X.-R. Liang and N.-Y. Ji, Nat. Prod. Res., 2024, 38, 3051–3056, DOI:10.1080/14786419.2023.2214291.
- X. Li, Y. Chen, Z. Liu, S. Li, H. Liu, Y. Wang, W. Zhang and H. Yan, Nat. Prod. Res., 2024, 38, 594–600, DOI:10.1080/14786419.2023.2187794.
- W. Zhang, Y. Yuan, X. Pang, Y. Liu, X. Zhang, Q. Liu and J. Wang, Chem. Biodiversity, 2024, 21, e202401751, DOI:10.1002/cbdv.202401751.
- L. Dai, Q. Xie, J. Guo, Q. Ma, L. Yang, J. Yuan, H. Dai, Z. Yu and Y. Zhao, J. Oceanol. Limnol., 2024, 42, 905–914, DOI:10.1007/s00343-023-3084-7.
- H. Kamauchi, M. Tanaka, M. Suzuki, M. Furukawa, A. Ikeda, C. Sasho, Y. Kiba, M. Kitamura, K. Takao and Y. Sugita, Chem. Pharm. Bull., 2024, 72, 98–101, DOI:10.1248/cpb.c23-00723.
- J. Wang, H. Lu, W. Fang, M. Lin, Y. Feng, X. Qi, C. Gao, Y. Liu, X. Wang and X. Luo, RSC Adv., 2024, 14, 38697–38705, 10.1039/D4RA06292B.
- S. Xu, Z. Li, X. Xin and F. An, Mar. Drugs, 2024, 22, 227, DOI:10.3390/md22050227.
- J. R. Virués-Segovia, C. Pinedo, D. Zorrilla, J. Sánchez-Márquez, P. Sánchez, M. C. Ramos, M. de la Cruz, J. Aleu and R. Durán-Patrón, Front. Mar. Sci., 2024, 11, 1386175, DOI:10.3389/fmars.2024.1386175.
- J. He, Q. Zou, H. Deng, S. He, D. Yan, K. Pan, Y. Zhou, Z. Zhao, H. Cui and Y. Liu, Fitoterapia, 2024, 173, 105804, DOI:10.1016/j.fitote.2023.105804.
- Z.-P. Jiang, R. Su, M.-T. Chen, J.-Y. Li, H.-Y. Chen, L. Yang, F.-F. Liu, J. Liu, C.-J. Xu, W.-S. Li, Y. Rao and L. Huang, Phytochemistry, 2024, 223, 114121, DOI:10.1016/j.phytochem.2024.114121.
- X.-D. Chen, X. Li, X.-M. Li, S.-Q. Yang and B.-G. Wang, Mar. Drugs, 2024, 22, 574, DOI:10.3390/md22120574.
- G. N. Pham, B. Josselin, A. Cousseau, B. Baratte, M. Dayras, C. Le Meur, S. Debaets, A. Weill, T. Robert, G. Burgaud, I. Probert, F. M. Abdoul-Latif, L. Boyer, S. Bach and M. Mehiri, Mar. Drugs, 2024, 22, 444, DOI:10.3390/md22100444.
- Z. Zhao, F. Cen, T. Mo, W. Li and G. Y. Chen, Chem. Biodiversity, 2024, 21, e202400118, DOI:10.1002/cbdv.202400118.
- Y. Zhang, W. Liu, G. Li, C. Wu, J. Yan, D. Feng, S. Yuan, R. Zhang, H. Lou and X. Peng, Nat. Prod. Res., 2024, 38, 2957–2963, DOI:10.1080/14786419.2023.2201885.
- H. Li, Y. Chen, B. Tang, Z. Liu, B. Peng, J. Li, H. Gao, S. Wang and Z. Li, J. Nat. Prod., 2024, 87, 2695–2708, DOI:10.1021/acs.jnatprod.4c00740.
- T. Wang, J. Tian, J. Huang, Y. Yuan, C. B. Naman, S. Wu, H. Wang, W. Lin, Z. Tong, L. Ding, W. Wang and S. He, J. Nat. Prod., 2024, 87, 1203–1208, DOI:10.1021/acs.jnatprod.3c01078.
- M. A. Lee, J. S. Kang, J.-W. Yang, H.-S. Lee, C.-S. Heo, S. J. Park and H. J. Shin, Mar. Drugs, 2024, 22, 87, DOI:10.3390/md22020087.
- F.-H. Yao, X. Liang, W.-B. Shen, X.-H. Lu, G.-C. Li and S.-H. Qi, J. Nat. Prod., 2024, 87, 810–819, DOI:10.1021/acs.jnatprod.3c00984.
- T. Feng, R. Wu, Y. Wang, P. Wang, L. Zhou, C. Wang and F. Kong, Mar. Drugs, 2024, 22, 241, DOI:10.3390/md22060241.
- X. Chen, S. Chen, H. Guo, X. Lu, H. Shen, L. Liu, L. Wang, B. Chen, Y. Zhang and Y. Liu, Mar. Drugs, 2024, 22, 214, DOI:10.3390/md22050214.
- M. T. Ngo, J. W. Han, M. V. Nguyen, Y. Choi, B. Kim, E. S. Gho, H. Kim and G. J. Choi, J. Agric. Food Chem., 2024, 72, 16359–16367, DOI:10.1021/acs.jafc.4c04706.
- X. Li, W. Shen, G. Li, Y. Song, X. Lu, N.-K. Wong and Y. Yan, Nat. Prod. Res., 2024, 38, 3917–3923, DOI:10.1080/14786419.2023.2269462.
- J. Yang, X. Zhang, W. Xu, X. Xu and F. Song, Nat. Prod. Res., 2024, 38, 2231–2236, DOI:10.1080/14786419.2023.2169917.
- Y.-H. Zhang, H.-F. Du, Y.-F. Liu, F. Cao, D.-Q. Luo and C.-Y. Wang, Appl. Microbiol. Biotechnol., 2024, 108, 194, DOI:10.1007/s00253-024-13026-4.
- J. Yang, Y. Zong, C. Wang, K. Li, Y. Zhang and P. Li, Mar. Drugs, 2024, 22, 517, DOI:10.3390/md22110517.
- N. M. Anh, L. T. H. Minh, N. T. Linh, P. T. Dao, D. T. Quynh, D. T. M. Huong, P. V. Cuong, V. T. T. Huyen and T. T. H. Dat, Biosci., Biotechnol., Biochem., 2024, 88, 1254–1260, DOI:10.1093/bbb/zbae113.
- J. Wei, X. Chen and B. Wu, J. Org. Chem., 2024, 89, 15264–15270, DOI:10.1021/acs.joc.4c02067.
- C.-S. Zhu, X.-M. Li, S.-Q. Yang, Y.-W. Liu, B.-G. Wang and X. Li, Chem. Biodiversity, 2024, 21, e202400584, DOI:10.1002/cbdv.202400584.
- G. Yu, L. Zhou, G. Wu and D. Li, J. Ocean Univ. China, 2024, 23, 216–220, DOI:10.1007/s11802-023-5676-4.
- B.-C. Hao, Y.-Y. Zheng, Z.-H. Li, C.-J. Zheng, C.-Y. Wang and M. Chen, Nat. Prod. Res., 2024, 38, 2252–2257, DOI:10.1080/14786419.2023.2171401.
- W. Song, L. Ji, Y. Zhang and L. Cao, Front. Microbiol., 2024, 15, 1400803, DOI:10.3389/fmicb.2024.1400803.
- F. Lu, W. Song, H. Li and L. Cao, Rec. Nat. Prod., 2024, 18, 699–704, DOI:10.25135/rnp.486.2410.3339.
- Z. Chen, Y. Lv, Z. Luo, B. Hong and S. Niu, Fitoterapia, 2024, 179, 106248, DOI:10.1016/j.fitote.2024.106248.
- X. Xu, Y. Dong, J. Yang, L. Wang, L. Ma, F. Song and X. Ma, J. Fungi, 2024, 10, 424, DOI:10.3390/jof10060424.
- J. Bao, Y.-F. Zhao, X.-X. Wang, K. Zhu, R. Ao, H. Liu, X.-X. Li, J.-S. Zhang and H. Zhang, Bioorg. Chem., 2024, 148, 107434, DOI:10.1016/j.bioorg.2024.107434.
- Y. Yang, X. Han, C. Jiang, Y. Chen, X. Tang and G. Li, Bioorg. Chem., 2024, 153, 107897, DOI:10.1016/j.bioorg.2024.107897.
- F.-Z. Zhang, X.-M. Li, L.-H. Meng and B.-G. Wang, J. Antibiot., 2024, 77, 13–20, DOI:10.1038/s41429-023-00666-3.
- G. V. Borkunov, E. V. Leshchenko, D. V. Berdyshev, R. S. Popov, E. A. Chingizova, N. P. Shlyk, A. V. Gerasimenko, N. N. Kirichuk, Y. V. Khudyakova, V. E. Chausova, A. S. Antonov, A. I. Kalinovsky, A. R. Chingizov, E. A. Yurchenko, M. P. Isaeva and A. N. Yurchenko, Nat. Prod. Bioprospect., 2024, 14, 32, DOI:10.1007/s13659-024-00449-9.
- E. V. Leshchenko, E. A. Chingizova, A. S. Antonov, N. P. Shlyk, G. V. Borkunov, D. V. Berdyshev, V. E. Chausova, N. N. Kirichuk, Y. V. Khudyakova, A. R. Chingizov, A. I. Kalinovsky, R. S. Popov, N. Y. Kim, K. A. Chadova, E. A. Yurchenko, M. P. Isaeva and A. N. Yurchenko, Mar. Drugs, 2024, 22, 317, DOI:10.3390/md22070317.
- Y.-Q. Tang, X. Liang, Q.-H. Zou, H. Cui, L.-X. Luo and S.-H. Qi, J. Nat. Prod., 2024, 87, 1209–1216, DOI:10.1021/acs.jnatprod.3c01121.
- F.-P. Miao, S.-T. Fang, Z.-Z. Shi, Y.-P. Song and N.-Y. Ji, Fitoterapia, 2024, 177, 106110, DOI:10.1016/j.fitote.2024.106110.
- L.-T. Dai, L. Yang, J.-C. Guo, Q.-Y. Ma, Q.-Y. Xie, L. Jiang, Z.-F. Yu, H.-F. Dai and Y.-X. Zhao, Bioorg. Chem., 2024, 145, 107205, DOI:10.1016/j.bioorg.2024.107205.
- L. Yang, M.-M. Quan, Q.-Y. Ma, Q.-Y. Xie, C.-J. Gai, H.-F. Dai, Y.-E. Hao and Y.-X. Zhao, Chem. Biodiversity, 2024, 21, e202401871, DOI:10.1002/cbdv.202401871.
- C. Wang, Y. Zhou, L. Yang, H. Hu, J. Chen, Y. Ying and H. Wang, Nat. Prod. Res., 2024, 38, 3605–3612, DOI:10.1080/14786419.2023.2258441.
- W. Chen, J. Jiang, X. Pang, Y. Song, Z. Yang, J. Wang and Y. Liu, Org. Biomol. Chem., 2024, 22, 4521–4527, 10.1039/D4OB00586D.
- Y.-Y. Sun, B. Hu, H.-B. Yu, J. Zhou, X.-C. Meng, Z. Ning, J.-F. Ding, M.-H. Cui and X.-Y. Liu, Mar. Drugs, 2024, 22, 236, DOI:10.3390/md22060236.
- C. Wang, C. Wang, Y. Liu, Y. Yue, X. Lu, H. Wang, Y. Ying and J. Chen, Appl. Environ. Microbiol., 2024, 90, e01603–e01624, DOI:10.1128/aem.01603-24.
- C. Wang, Y. Fan, C. Wang, J. Tang, Y. Qiu, K. Xu, Y. Ding, Y. Liu, Y. Ying and H. Wang, Mar. Drugs, 2024, 22, 296, DOI:10.3390/md22070296.
- N. J. Morehouse, K. M. Graham, S. L. Cox, J. A. Johnson and C. A. Gray, Can. J. Chem., 2024, 102, 86–90, DOI:10.1139/cjc-2023-0100.
- H. Fan, X.-H. Shao, Z.-K. Zhang, C. Ni, C. Feng, X. Wei, J.-Q. Zhu, X.-H. Li and C.-X. Zhang, J. Nat. Prod., 2024, 87, 705–712, DOI:10.1021/acs.jnatprod.3c00765.
- H. Li, J. Long, X. Wang, J. She, Y. Liu, Y. Li and B. Yang, Nat. Prod. Res., 2024, 38, 2996–3003, DOI:10.1080/14786419.2023.2207133.
- J.-W. Wang, Y.-Y. Liu, G.-Y. Chen, C.-J. Zheng and X.-H. Nong, Fitoterapia, 2024, 175, 105906, DOI:10.1016/j.fitote.2024.105906.
- Y. Chen, J. Cai, Z. Xia, C. Chen, Y. Liu, L. Jayasinghe, X. Wang and X. Zhou, Mar. Drugs, 2024, 22, 384, DOI:10.3390/md22090384.
- L. Huang, C. Chen, J. Cai, Y. Chen, Y. Zhu, B. Yang, X. Zhou, Y. Liu and H. Tao, Mar. Drugs, 2024, 22, 393, DOI:10.3390/md22090393.
- J. Cai, M. Li, C. Chen, B. Yang, C. Gao, Y. Liu, X. Luo, Y. Tan and X. Zhou, J. Nat. Prod., 2024, 87, 1401–1406, DOI:10.1021/acs.jnatprod.4c00116.
- J. Cai, L. Gao, Y. Wang, Y. Zheng, X. Lin, P. Zhou, C. Chen, K. Liu, L. Tang, Y. Liu, Y. Tan, M. Jin and X. Zhou, Eur. J. Med. Chem., 2024, 265, 116068, DOI:10.1016/j.ejmech.2023.116068.
- C. Chen, L. Xiao, X. Luo, J. Cai, L. Huang, H. Tao, X. Zhou, Y. Tan and Y. Liu, J. Med. Chem., 2024, 67, 2602–2618, DOI:10.1021/acs.jmedchem.3c01748.
- Y.-R. Wang, Y.-L. Dong, X.-M. Li, X.-S. Shi, H.-L. Li, L.-H. Meng, R. Xu and B.-G. Wang, Phytochemistry, 2024, 220, 114000, DOI:10.1016/j.phytochem.2024.114000.
- B. Yang, S. Shao, M. Nie, Q. Tie, X. Pang, X. Lin, X. Zhou, Y. Liu, X. Wang and Y. Li, Mar. Drugs, 2024, 22, 218, DOI:10.3390/md22050218.
- M. R. de Amorim, S. M. Schoellhorn, C. d. S. Barbosa, G. R. Mendes, K. de L. Macedo, A. G. Ferreira, T. Venâncio, R. V. C. Guido, A. N. L. Batista, J. M. Batista Jr, E. Skellam and R. G. S. Berlinck, J. Nat. Prod., 2024, 87, 2204–2215, DOI:10.1021/acs.jnatprod.4c00516.
- T. Wang, Y. Feng, J. Huang, S. Wu, K. Hu, J. Wu, C. B. Naman, H. Wang, W. Lin and S. He, J. Nat. Prod., 2024, 87, 160–165, DOI:10.1021/acs.jnatprod.3c00892.
- L. Jiang, B. Teng, M. Zhang, S. Chen, D. Zhang, L. Zhai, J. Lin and H. Lei, Mar. Drugs, 2024, 22, 15, DOI:10.3390/md22010015.
- Z.-L. Tan, Y.-C. Chen, J.-P. Zhang, H.-X. Liu, W.-M. Zhang and H.-J. Yan, J. Asian Nat. Prod. Res., 2024, 26, 534–540, DOI:10.1080/10286020.2023.2249832.
- X. Li, Y. Chen, S. Li, W. Zhang, H. Yan, H. Liu and W. Zhang, Fitoterapia, 2024, 172, 105772, DOI:10.1016/j.fitote.2023.105772.
- J. Wu, J. Ye, J. Cen, Y. Chen and J. Xu, Mar. Drugs, 2024, 22, 332, DOI:10.3390/md22080332.
- J. Wu, D. Chen, Q. Li, T. Feng and J. Xu, Mar. Drugs, 2024, 22, 102, DOI:10.3390/md22030102.
- Z. Ying, X.-M. Li, S.-Q. Yang, H.-L. Li, X. Li, B.-G. Wang and L.-H. Meng, J. Nat. Prod., 2024, 87, 1347–1357, DOI:10.1021/acs.jnatprod.3c01175.
- C. Sun, Y. Jiang, C. Li, S. Sun, J. Lin, W. Wang, L. Zhou, L. Li, M. Shah, Q. Che, G. Zhang, D. Wang, T. Zhu and D. Li, Angew. Chem., Int. Ed., 2024, 63, e202405860, DOI:10.1002/anie.202405860.
- L. Yang, Q. Zhang, D. Yu, W. Zhu and Y. Wang, Chem. Biodiversity, 2024, 21, e202400578, DOI:10.1002/cbdv.202400578.
- J.-W. Wang, D.-Y. Liu, H.-Z. Zhang, Z. Tan, C.-J. Zheng, G.-Y. Chen and X.-H. Nong, Nat. Prod. Res., 2024, 38, 1711–1718, DOI:10.1080/14786419.2023.2218008.
- X. Liu, Y. Lv, J. He, B. Hong, Z. Shao, M. Yu and S. Niu, Chem. Biodiversity, 2024, 21, e202400805, DOI:10.1002/cbdv.202400805.
- Y. Xie, Y. Wang, K. Zhang, Q.-M. Liu, Q.-X. Yan, Z.-B. Zou, Y.-Q. Xu, G.-M. Liu, X.-F. Huang and X.-W. Yang, Eur. J. Org Chem., 2024, 27, e202301250, DOI:10.1002/ejoc.202301250.
- H. Azami, Y. Watanabe, K. Sakai, H. Nakahara, H. Kojima, T. Tokiwa, K. Nonaka, Y. Noguchi, Y. Nagano, T. Hirose, T. Sunazuka, H. Matsui, N. Arima, K. Abe, H. Hanaki and M. Iwatsuki, J. Antibiot., 2024, 77, 156–162, DOI:10.1038/s41429-023-00696-x.
- Z. Xiao, J. Cai, T. Chen, Y. Wang, Y. Chen, Y. Zhu, C. Chen, B. Yang, X. Zhou and H. Tao, Mar. Drugs, 2024, 22, 103, DOI:10.3390/md22030103.
- X. Pang, B. Yang, X. Zhou, J. Wang, J. Yang and Y. Liu, Chem. Biodiversity, 2024, 21, e202302069, DOI:10.1002/cbdv.202302069.
- H. Kwon, B.-R. Jin, S. Yoo, H.-J. Kim, B. Y. Hwang, Y. Guo, J. H. Yim, I.-C. Kim, S. H. Shim, H. J. An and D. Lee, Bioorg. Chem., 2024, 143, 107070, DOI:10.1016/j.bioorg.2023.107070.
- X. Liang, J.-F. Yang, Z.-H. Huang, X. Ma, Y. Yan and S.-H. Qi, J. Agric. Food Chem., 2024, 72, 6402–6413, DOI:10.1021/acs.jafc.4c00493.
- Y.-C. He, M.-Q. Wang, Q.-Q. Tie, X.-W. Huang, Y.-H. Liu, Y.-Q. Li and B. Yang, J. Antibiot., 2024, 78, 64–67, DOI:10.1038/s41429-024-00780-w.
- Z. Wu, W. Wang, J. Li, C. Ma, L. Chen, Q. Che, G. Zhang, T. Zhu and D. Li, J. Nat. Prod., 2024, 87, 1222–1229, DOI:10.1021/acs.jnatprod.3c01202.
- Z. Huang, D. Liu, S. Chen, J. Ren, C. Gao, Z. Li, A. Fan and W. Lin, Mar. Drugs, 2024, 22, 78, DOI:10.3390/md22020078.
- S. Lin, Z. Chai, H. Zeng, B. Yang, J. Chi, Y. Zhang and Z. Hu, Phytochemistry, 2024, 225, 114199, DOI:10.1016/j.phytochem.2024.114199.
- X. Ma, M. Wu, Z. Chen, F. Cao, T. Zhong, Z. Luo, Z. Shao, Y. Zhang, L. Chen and Z. Zhang, Molecules, 2024, 29, 1685, DOI:10.3390/molecules29071685.
- T. Berger, J. Alenfelder, S. Steinmüller, D. Heimann, N. Gohain, D. Petras, M. Wang, R. Berger, E. Kostenis and R. Reher, J. Nat. Prod., 2024, 87, 692–704, DOI:10.1021/acs.jnatprod.3c00750.
- M. Zheng, Q. Li, H. Liao, Y. Li, C. Zhou, X. Zhao, C. Chen, W. Sun, Y. Zhang and H. Zhu, J. Nat. Prod., 2024, 87, 1921–1929, DOI:10.1021/acs.jnatprod.4c00330.
- J. Wu, W. Wang, Y. Yang, M. Shah, J. Peng, L. Zhou, G. Zhang, Q. Che, J. Li, T. Zhu and D. Li, J. Nat. Prod., 2024, 87, 1407–1415, DOI:10.1021/acs.jnatprod.4c00132.
- Z.-H. Peng, H. Jia, Y.-L. Luo, L.-J. Zhang, J.-T. Zhou, Y.-H. Xie, L.-J. Wang, J.-K. Qin, J. Li, G.-H. Zhang, R.-Y. Yang and W.-F. Xu, Mar. Drugs, 2024, 22, 475, DOI:10.3390/md22100475.
- K. Zhang, Y. Wang, T. Long, Y.-P. Xie, Z.-B. Zou, Y. Zhang, Y. Li, M.-M. Xie, G.-M. Liu and X.-W. Yang, Biochem. Syst. Ecol., 2024, 114, 104829, DOI:10.1016/j.bse.2024.104829.
- W. Wang, J. Wang, F. Song, R. Jia, L. Wang, X. Xu and N. Yang, Mar. Drugs, 2024, 22, 237, DOI:10.3390/md22060237.
- X. Liang, Z.-H. Huang, W.-B. Shen, X.-H. Lu, X.-X. Zhang, X. Ma and S.-H. Qi, Phytochemistry, 2024, 223, 114119, DOI:10.1016/j.phytochem.2024.114119.
- Y. Wang, X.-M. Li, N. Song, B.-G. Wang, H.-L. Li and L.-H. Meng, Bioorg. Chem., 2024, 147, 107417, DOI:10.1016/j.bioorg.2024.107417.
- Y. Cho, K.-H. Park, E. Kim, S. Kim, W. Wang, H. Choi and H. Kang, J. Nat. Prod., 2024, 87, 1230–1234, DOI:10.1021/acs.jnatprod.3c01227.
- L. Tang, J. Xia, Z. Chen, X. Wu, G. Li, Q. Lai, Z. Shao, W. Wang and X. Hong, Mar. Drugs, 2024, 22, 274, DOI:10.3390/md22060274.
- L. Tang, J. Xia, Z. Chen, F. Lin, Z. Shao, W. Wang and X. Hong, Mar. Drugs, 2024, 22, 186, DOI:10.3390/md22040186.
- X. Hong, L. Tang, Z. Chen, Q. Lai, B. Zhang, Y. Jiang, X. Wang, R. He, J. Lin, Z. Shao, S. Lin and W. Wang, Nat. Prod. Res., 2024, 38, 320–326, DOI:10.1080/14786419.2022.2121830.
- T. Y. Kim, C. Lee, J. W. Lee and S. H. Shim, Nat. Prod. Sci., 2024, 30, 59–63, DOI:10.20307/nps.2024.30.1.59.
- Z. Wu, X.-M. Li, S.-Q. Yang, B.-G. Wang and X. Li, Mar. Drugs, 2024, 22, 204, DOI:10.3390/md22050204.
- H. Zeng, Y. Wang, C. Wang, Y. Huang, S. Lin, J. Wei, W. Sun, F. Cao, Y. Zhang and Z. Hu, J. Nat. Prod., 2024, 87, 2592–2603, DOI:10.1021/acs.jnatprod.4c00709.
- Y. Shi, X.-Q. Sun, J.-X. Zhang, R.-H. Zhang, K. Hong, Y.-X. Xue, H. Qiu and L. Liu, J. Agric. Food Chem., 2024, 72, 17431–17443, DOI:10.1021/acs.jafc.4c03959.
- T. Gu, J. Cai, D. Xie, J. She, Y. Liu, X. Zhou and L. Tang, Mar. Drugs, 2024, 22, 403, DOI:10.3390/md22090403.
- J. Cai, Q. Zhou, X. Qi, F. Zhang, J. Yang, C. Chen, K. Zhang, Z. Chen, H.-B. Luo, Y. Liu, Y.-Y. Huang and X. Zhou, J. Nat. Prod., 2024, 87, 1808–1816, DOI:10.1021/acs.jnatprod.4c00422.
- Z. Zeng, J. Cai, Y. Chen, X. Li, C. Chen, Y. Liu, L. Jayasinghe and X. Zhou, Mar. Drugs, 2024, 22, 559, DOI:10.3390/md22120559.
- C. Chen, X. Wang, W. Fang, J. Liang, J. Cai, D. Yang, X. Luo, C. Gao, X. Yi, Y. Liu and X. Zhou, Chin. J. Nat. Med., 2024, 22, 1047–1056, DOI:10.1016/S1875-5364(24)60659-X.
- Y. Jung, C. Kwon, T. Kim, J. W. Lee, M.-K. Shin and S. H. Shim, Mar. Life Sci. & Technol., 2024, 6, 84–92, DOI:10.1007/s42995-023-00198-7.
- Y. Liu, L. Qi, M. Xu, W. Li, N. Liu, X. He and Y. Zhang, Front. Microbiol., 2024, 15, 1446283, DOI:10.3389/fmicb.2024.1446283.
- W. Yang, S. Tian, Y.-F. Du, X.-L. Zeng, J.-J. Liang, W.-J. Lan and H. Li, J. Nat. Prod., 2024, 87, 2746–2756, DOI:10.1021/acs.jnatprod.4c00905.
- X. Dong, L.-C. Jia, H.-Q. Chai, X.-X. Liu and L. Yang, Rec. Nat. Prod., 2024, 18, 352–356, DOI:10.25135/rnp.456.2403.3175.
- C. Zhou, Y. Ge, D. Lan, M. Zhao and B. Wu, Fitoterapia, 2024, 176, 106039, DOI:10.1016/j.fitote.2024.106039.
- Y. Zhang, Y. Zhang, G. Li, K. Dong, J. Wang, S. Xiao, H. Lou and X. Peng, Fitoterapia, 2024, 175, 105963, DOI:10.1016/j.fitote.2024.105963.
- C. Chen, L. Liu, S. Ye, J. Li, L. Wu, J. Li, H. Jia and Y. Long, Steroids, 2024, 208, 109449, DOI:10.1016/j.steroids.2024.109449.
- L. Huang, C. Chen, J. Cai, Y. Chen, Y. Zhu, B. Yang, X. Zhou, Y. Liu and H. Tao, Chem. Biodiversity, 2024, 21, e202400070, DOI:10.1002/cbdv.202400070.
- H.-Y. Hsi, S.-W. Wang, G. Hsiao, L.-K. Chang, Y.-C. Cheng, S.-J. Huang, Y.-S. Lu and T.-H. Lee, J. Food Drug Anal., 2024, 32, 155–167, DOI:10.38212/2224-6614.3501.
- Y. Xiao, J. Liu, Y. Jiang, Y. Guo and T. Ye, Molecules, 2024, 29, 3599, DOI:10.3390/molecules29153599.
- S. B. Khandekar and R. A. Fernandes, Org. Biomol. Chem., 2024, 22, 4508–4515, 10.1039/D4OB00549J.
- Z. Xu, X.-T. Liang, L. V. White, M. G. Banwell and S. Tan, J. Org. Chem., 2024, 89, 12639–12650, DOI:10.1021/acs.joc.4c01559.
- Z. Xu, X.-T. Liang, P. Lan and M. G. Banwell, Org. Chem. Front., 2024, 11, 2433–2441, 10.1039/D4QO00131A.
- R. Ghosh and S. K. Chattopadhyay, Eur. J. Org Chem., 2024, 27, e202400228, DOI:10.1002/ejoc.202400228.
- G. Han, W. Zhang, E. Acs, A. Paquin, Q. Ronzon, N. Casaretto and B. Nay, Org. Lett., 2024, 26, 2629–2634, DOI:10.1021/acs.orglett.4c00658.
- E. Fukuda, I. Fujiwara, S. Maruno, K. Motomura, S. Endo, A. Iwasaki, T. Fukuta, A. Nakayama and T. Shinada, J. Nat. Prod., 2024, 87, 1394–1400, DOI:10.1021/acs.jnatprod.4c00114.
- M. Wang, Y. Jia, F. Wu, Z. Li, L. Shao and X. Chen, Org. Biomol. Chem., 2024, 22, 7649–7658, 10.1039/D4OB01186D.
- T. Li, S. Jiang, Y. Dai, X. Wu, H. Guo, L. Shi, X. Sang, L. Ren, J. Wang, L. Shi, W. Zhou, H. Li and H.-D. Hao, Nat. Commun., 2024, 15, 10581, DOI:10.1038/s41467-024-55189-8.
- W. M. Hussein, Y. Zhu, A. A. Salim and R. J. Capon, Mar. Drugs, 2024, 22, 454, DOI:10.3390/md22100454.
- Y. Kawamoto, A. Nishitani, Y. Yoshimura, T. Kobayashi and H. Ito, Chem. Commun., 2024, 60, 95–97, 10.1039/D3CC05130G.
- G. Rajendar and M. Rehman, Tetrahedron, 2024, 168, 134343, DOI:10.1016/j.tet.2024.134343.
- Y. Liu, Z.-B. Zou, F.-D. Gu, Y.-F. Lin, Y. Li, H.-Y. Chen, H. Liu, X.-W. Yang, G.-M. Liu and Q.-M. Liu, J. Agric. Food Chem., 2024, 72, 5463–5476, DOI:10.1021/acs.jafc.3c08126.
- T. Jiang, K. P. Godinez-Macias, J. E. Collins, J. W. Lee, K. L. Wendt, K. Carolino, D. Chakrabarti, R. H. Cichewicz and E. A. Winzeler, Microbiol. Spectr., 2024, 12, e04142–23, DOI:10.1128/spectrum.04142-23.
- H. Zhang, C. Sun, Q. Xia, P. Li, K. Liu and Y. Zhang, Mar. Drugs, 2024, 22, 439, DOI:10.3390/md22100439.
- A. C. Jordão, G. S. dos Santos, T. R. Teixeira, A. J. P. Gluzezak, C. B. de Souza Azevedo, K. de Castro Pereira, L. Tonani, L. R. Gaspar, M. R. von Zeska Kress, P. Colepicolo and H. M. Debonsi, Arch. Microbiol., 2024, 206, 35, DOI:10.1007/s00203-023-03756-w.
- Y. Fang, J. She, X. Zhang, T. Gu, D. Xie, X. Luo, X. Yi, C. Gao, Y. Liu, C. Zhang, L. Tang and X. Zhou, J. Nat. Prod., 2024, 87, 322–331, DOI:10.1021/acs.jnatprod.3c01029.
- Y. Rao, R. Su, C. Wu, G. Yang, R. Fu, J. Wu, J. Liang, J. Liu, Z. Jiang, C. Xu and L. Huang, Front. Pharmacol, 2024, 15, 1320040, DOI:10.3389/fphar.2024.1320040.
- Y. Rao, R. Su, C. Wu, X. Chai, J. Li, G. Yang, J. Wu, T. Fu, Z. Jiang, Z. Guo, C. Xu and L. Huang, Acta Pharm. Sin. B, 2024, 14, 304–318, DOI:10.1016/j.apsb.2023.08.032.
- C. Wu, Y. Chen, M. Chen, T. Fu, J. Liu, F. Liu, C. Xu, W. Li, B. Li, Z. Jiang, Y. Rao and L. Huang, Free Radical Biol. Med., 2024, 224, 630–643, DOI:10.1016/j.freeradbiomed.2024.09.019.
- Y.-S. Liu, W.-L. Chen, Y.-W. Zeng, Z.-H. Li, H.-L. Zheng, N. Pan, L.-Y. Zhao, S. Wang, S.-H. Chen, M.-H. Jiang, C.-C. Jin, Y.-C. Mi, Z.-H. Cai, X.-Z. Fang, Y.-J. Liu, L. Liu and G.-L. Wang, Mar. Drugs, 2024, 22, 283, DOI:10.3390/md22060283.
- X. Niu, S. Chen, X. Wang, J. Wen, X. Liu, Y. Yong, Z. Yu, X. Ma, A. M. Abd El-Aty and X. Ju, Mar. Drugs, 2024, 22, 564, DOI:10.3390/md22120564.
- J. M. Oh, Q. Gao, W.-H. Shin, E.-Y. Lee, D. Chung, G. Choi, S.-J. Nam and H. Kim, Appl. Biol. Chem., 2024, 67, 26, DOI:10.1186/s13765-024-00878-7.
- Y. Zeng, T. Lu, S. Ren, Z. Hu, J. Fang, Z. Guan, J. Li, L. Liu and Z. Gao, Org. Lett., 2024, 26, 6692–6697, DOI:10.1021/acs.orglett.4c02372.
- Z.-F. Zhang, F. Liu, L.-R. Liu, M. Li, L. Cai, S. Liu and J. Mao, Fungal Biol. Rev., 2024, 48, 100353, DOI:10.1016/j.fbr.2023.100353.
- C. Han, A. Song, Y. He, L. Yang, L. Chen, W. Dai, Q. Wu and S. Yuan, Front. Microbiol., 2024, 15, 1520446, DOI:10.3389/fmicb.2024.1520446.
- Y. Choi, E. J. Jeong, Y. D. Yoo, J. Park and J.-R. Rho, J. Nat. Prod., 2024, 87, 2537–2542, DOI:10.1021/acs.jnatprod.4c00851.
- S. Moon, D. H. Choi, Y. Lee and J.-R. Rho, J. Korean Magn. Reson., 2024, 28, 25–31, DOI:10.6564/JKMRS.2024.28.3.025.
- N. T. T. Thuy, H. T. M. Hien, N. C. Ha, L. T. Thom, D. D. Hong, N. V. Thinh, N. T. Dan, N. D. Hoi, V. T. Loan, H. D. Quang, D. T. T. Hang, P. V. Kiem, B. H. Tai, N. T. M. Hang, N. T. Dat and N. X. Nhiem, Nat. Prod. Res., 2024, 38, 789–795, DOI:10.1080/14786419.2023.2201882.
- M. Azizah and G. Pohnert, J. Nat. Prod., 2024, 87, 50–57, DOI:10.1021/acs.jnatprod.3c00766.
- M. Hakamada, C. Tokairin, H. Ishizuka, K. Adachi, T. Osawa, S. Aonuma, R. Hirozumi, S. Tsuchiya, Y. Cho, Y. Kudo, K. Konoki, Y. Oshima, K. Nagasawa and M. Yotsu-Yamashita, Chem.–Eur. J., 2024, 30, e202304238, DOI:10.1002/chem.202304238.
- K. Zhang, X. Yin, Y. Huang, C. Liu, Q. Zhang, Q. Liu, S. Wang, W. Fei, Q. Shi and L. Qiu, Int. J. Mol. Sci., 2024, 25, 13736–13751, DOI:10.3390/ijms252413736.
- A. M. White, B. D. Schwartz, M. G. Gardiner and L. R. Malins, Org. Lett., 2024, 26, 6803–6808, DOI:10.1021/acs.orglett.4c02004.
- Y. Ning, Y. Wang and J. Gui, JACS Au, 2024, 4, 635–641, DOI:10.1021/jacsau.3c00698.
- T. Obana, M. Nakajima, K. Nakazato, H. Nakagawa, K. Murata, M. Tsuda and H. Fuwa, J. Am. Chem. Soc., 2024, 146, 29836–29846, DOI:10.1021/jacs.4c11714.
- E. Montuori, D. De Luca, A. Penna, D. Stalberga and C. Lauritano, Mar. Drugs, 2024, 22, 31–50, DOI:10.3390/md22010031.
- I. Ciss, M. Seck, B. Figadère and L. Ferrié, Chem.–Eur. J., 2024, 30, e202400471, DOI:10.1002/chem.202400471.
- J. Yang, W. Sun, M. Sun, Y. Cui and L. Wang, Mar. Drugs, 2024, 22, 79–95, DOI:10.3390/md22020079.
- J. C. Perkins, K. R. Zenger, Y. Liu and J. M. Strugnell, Harmful Algae, 2024, 139, 102735–102754, DOI:10.1016/j.hal.2024.102735.
- C. O. Miles, I. W. Burton, N. I. Lewis, A. Robertson, S. D. Giddings, P. McCarron and E. M. Mudge, Tetrahedron, 2024, 162, 134115–134124, DOI:10.1016/j.tet.2024.134115.
- J. S. Wood, J. Tang, W. K. Strangman and R. T. Williamson, J. Nat. Prod., 2024, 87, 2120–2125, DOI:10.1021/acs.jnatprod.4c00525.
- E. M. Mudge, A. L. Wilkins, J. S. Murray, F. Rise and C. O. Miles, Toxicon, 2024, 251, 108154–108165, DOI:10.1016/j.toxicon.2024.108154.
- J. L. García-Corona, H. Hegaret, M. Lassudrie, A. Derrien, A. Terre-Terrillon, T. Delaire and C. Fabioux, Aquat. Toxicol., 2024, 266, 106793, DOI:10.1016/j.aquatox.2023.106793.
- T. R. Fallon, V. V. Shende, I. H. Wierzbicki, A. L. Pendleton, N. F. Watervoort, R. P. Auber, D. J. Gonzalez, J. H. Wisecaver and B. S. Moore, Science, 2024, 385, 671–678, DOI:10.3390/md22050213.
- D. Li, X. Liao, S. Xu and B. Zhao, Nat. Prod. Res., 2024, 38, 1320–1325, DOI:10.1080/14786419.2022.2140152.
- S. Komori, S. Yamabe, R. Matsuta, Y. Yamazaki, M. Fukuoka, S. Sato and K. Takada, Biosci. Biotechnol. Biochem., 2024, 88, 399–404, DOI:10.1093/bbb/zbae007.
- Y. Zeng, X. Yang, Z. Xia, R. Chen, F. He, J. Zhang and P. He, Biology, 2024, 13, 456–471, DOI:10.3390/biology13060456.
- A. R. Mendes, M. P. Spínola, M. Lordelo and J. A. M. Prates, Appl. Sci., 2024, 14, 10810–10835, DOI:10.3390/app142310810.
- Y. Sun, H. Li, X. Ma, M. Pu, Y. Zhang, Z. Dong, P. He and S. Zheng, Mar. Drugs, 2024, 22, 279–294, DOI:10.3390/md22060279.
- H. S. Kim, J. W. Ahn, N. R. Ha, K. Damodar, S. K. Jang, Y.-M. Yoo, Y. S. Gyoung and S. S. Joo, Mar. Drugs, 2024, 22, 354–374, DOI:10.3390/md22080354.
- J.-Y. Kim, G.-W. Oh, D.-C. Choi, D.-W. Ki, S.-C. Ko, K. W. Kim, D. Yang, M.-J. Yim, J. M. Lee, K. Lee, D.-M. Jo, C. Grace and D.-S. Lee, J. Appl. Biol. Chem., 2024, 67, 459–463, DOI:10.3839/jabc.2024.063.
- J. Pan, H. Liu, J. Xu, X. Xu, Y. Qi, C. Wang, C. Wu, Z. Sun, Y. Lv and P. Yan, Phytochem. Lett., 2024, 64, 133–137, DOI:10.1016/j.phytol.2024.11.001.
- K. Ashwini, B. Siva, P. Poornima, S. D. Reddy, V. G. Sastry and K. S. Babu, Fitoterapia, 2024, 177, 106071–110677, DOI:10.1016/j.fitote.2024.106071.
- K. Wang, C.-H. Xu, L. Tong, B. Wang, A.-H. Liu, S.-W. Li and S.-C. Mao, J. Nat. Prod., 2024, 87, 121–131, DOI:10.1021/acs.jnatprod.3c00898.
- A. Guy, S. Delly, J. Galano, T. Durand and C. Oger, Chem.–Eur. J., 2024, 30, e202401632, DOI:10.1002/chem.202401632.
- Y. Aalilou, H. Moussa, L. H. Lee, A. Bouyahya, G. Zengin and M. El Abbes Faouzi, Fitoterapia, 2024, 176, 106016–106030, DOI:10.1016/j.fitote.2024.106016.
- Y. Peng, X. Yang, R. Huang, B. Ren, B. Chen, Y. Liu and H. Zhang, Mar. Drugs, 2024, 22, 59–78, DOI:10.3390/md22020059.
- P. Anand, A. Kaur and S. Singh, Eur. J. Pharmacol., 2024, 967, 176335–176348, DOI:10.1016/j.ejphar.2024.176335.
- S. Ishigami, R. Fukada, G. Nagasaka, T. Tsuruta, K. Nishikawa, Y. Sasaki, K. Nimura, I. Oshima, Y. Yamagishi, Y. Morimoto, T. Kamada and T. Ishii, Chem. Biodiversity, 2024, 21, e202400436, DOI:10.1002/cbdv.202400436.
- V. T. Doan, V. T. Pham, C. H. D. Le, T. T. N. Luu, T. U. Dam, H. K. H. Le, Q. T. That, K. P. P. Nguyen and T. T. D. Ngo, Nat. Prod. Res., 2024, 38, 2258–2263, DOI:10.1080/14786419.2023.2171407.
- K. Kirita, H. Matsumoto, G. Endo and S. Hosokawa, Angew. Chem., Int. Ed., 2024, 63, e202400586, DOI:10.1002/anie.202400586.
- K. Kirita, H. Matsumoto, G. Endo, K. Inoue and S. Hosokawa, Biosci., Biotechnol., Biochem., 2024, 88, 1144–1154, DOI:10.1093/bbb/zbae104.
- N. Castejón, A. Adrien, L. Spitzer and S. C. M. Fernandes, J. Appl. Phycol., 2024, 36, 291–311, DOI:10.1007/s10811-023-03115-8.
- Y. Yamagishi, T. Kamada, T. Ishii, H. Matsuura, N. Kikuchi, T. Abe and M. Suzuki, Chem. Biodiversity, 2024, 21, e202400833, DOI:10.1002/cbdv.202400833.
- K.-L. Cheong, G. Li, M. Wang, Y. Yang and S. Zhong, Front. Mar. Sci., 2024, 11, 1360425–1360435, DOI:10.3389/fmars.2024.1360425.
- K. Kaneko, M. Shinke, T. Kamada, T. Okino and A. Yamamoto, J. Appl. Phycol., 2024, 36, 3795–3808, DOI:10.1007/s10811-024-03327-6.
- T. F. Molinski, J. Org. Chem., 2024, 89, 10027–10036, DOI:10.1021/acs.joc.4c00905.
- Y. Oyadomari, Y. Goto, K. Suganuma, S.-I. Kawazu, L. E. Becking, N. Fusetani and Y. Nakao, Mar. Drugs, 2024, 22, 171, DOI:10.3390/md22040171.
- A.-Y. Shin and J. Lee, J. Nat. Prod., 2024, 87, 1872–1880, DOI:10.1021/acs.jnatprod.4c00692.
- R. Kumar, R. K. Akee, L. Martínez-Fructuoso, V. F. Freire, C. C. Thornburg, J. R. Evans, B. D. Peyser, S. Ensel, B. R. O'Keefe and T. Grkovic, Bioorg. Med. Chem. Lett., 2024, 113, 129963–129967, DOI:10.1016/j.bmcl.2024.129963.
- L. A. Amador, A. D. Rodríguez, L. Carmona-Sarabia, E. E. Colón-Lorenzo and A. E. Serrano, Appl. Sci., 2024, 14, 281–294, DOI:10.3390/app14010281.
- W. Zhong, J. O. Olugbami, P. Rathakrishnan, I. Mohanty, S. G. Moore, N. Garg, A. K. Oyelere, T. L. Turner, A. C. McShan and V. Agarwal, J. Org. Chem., 2024, 89, 12748–12752, DOI:10.1021/acs.joc.4c01104.
- Y. Hitora, A. H. El-Desoky, Y. Sadahiro, A. Sejiyama, A. Kinoshita, Y. Ise, E. D. Angkouw, R. E. P. Mangindaan, T. Higaki and S. Tsukamoto, J. Nat. Prod., 2024, 87, 1197–1202, DOI:10.1021/acs.jnatprod.4c00158.
- P. G. Cruz, R. Fernández, R. Rodríguez-Acebes, M. Martínez-Díez, G. Santamaría-Núñez, M. Pérez and C. Cuevas, Mar. Drugs, 2024, 22, 339–349, DOI:10.3390/md22080339.
- D. Lucena-Agell, M. J. Guillén, R. Matesanz, B. Álvarez-Bernad, R. Hortigüela, P. Avilés, M. Martínez-Díez, G. Santamaría-Núñez, J. Contreras, I. Plaza-Menacho, J. F. Giménez-Abián, M. A. Oliva, C. Cuevas and J. F. Díaz, J. Med. Chem., 2024, 67, 2619–2630, DOI:10.1021/acs.jmedchem.3c01775.
- J. A. Gerke, S. F. Odron, J. Kim, N. Dutta, J. G. Clarke, J. Media, D. A. Coppage, M. Oorloff, A. Alcala, G. Garcia, M. E. F. Kang, C. L. Gerke, J. C. Peterson, J. D. Morris, R. Higuchi-Sanabria, F. A. Valeriote, P. Crews and T. A. Johnson, J. Nat. Prod., 2024, 87, 2523–2529, DOI:10.1021/acs.jnatprod.4c00691.
- C. S. Hassane, F. Tintillier, P.-E. Campos, G. Herbette, N. J. de Voogd, J. Ouazzani, M. Fouillaud, L. Dufossé and A. Gauvin-Bialecki, Nat. Prod. Res., 2024, 38, 2973–2982, DOI:10.1080/14786419.2023.2204431.
- A. A. E. Said, E. Z. Attia, A. S. Wanas, M. N. Samy, U. R. Abdelmohsen, K. Matsunami and M. A. Fouad, Nat. Prod. Res., 2024, 38, 3581–3587, DOI:10.1080/14786419.2023.2256448.
- Z. Wang, G.-Q. Li and H.-T. Zhang, Chem. Biodiversity, 2024, 21, e202400939, DOI:10.1002/cbdv.202400939.
- Z.-J. Guo, H.-X. Liang, X.-Y. Lian, X.-J. Liao, X.-W. Xing, S.-H. Xu and B.-X. Zhao, J. Asian Nat. Prod. Res., 2024, 26, 328–333, DOI:10.1080/10286020.2023.2244432.
- D. Wang, W. Jiang, M. Churiwal, K. Jia, S. P. D. Senadeera, H. R. Bokesch, G. M. Woldemichael, Y. Kim, R. G. Hawley, J. S. Wei, J. Khan, B. R. O'Keefe, J. A. Beutler and K. R. Gustafson, J. Nat. Prod., 2024, 87, 332–339, DOI:10.1021/acs.jnatprod.3c01041.
- K. M. Tabakmakher, T. N. Makarieva, Y. E. Sabutski, M. S. Kokoulin, A. S. Menshov, R. S. Popov, A. G. Guzii, L. K. Shubina, E. A. Chingizova, A. R. Chingizov, E. A. Yurchenko, S. N. Fedorov, B. B. Grebnev, G. v. Amsberg, S. A. Dyshlovoy, N. V. Ivanchina and P. S. Dmitrenok, Mar. Drugs, 2024, 22, 396–413, DOI:10.3390/md22090396.
- S. Sala, M. Shimomura, L. Tham, J. Sakata, A. N. Sobolev, S. A. Moggach, J. Fromont, O. Gomez, M. J. Piggott, H. Tokuyama, S. G. Stewart and G. R. Flematti, J. Nat. Prod., 2024, 87, 2507–2514, DOI:10.1021/acs.jnatprod.4c00549.
- M. Orfanoudaki, M. Dalilian, L. Du, C. H. Chau, W. D. Figg, B. R. O'Keefe and T. Grkovic, J. Nat. Prod., 2024, 87, 2640–2648, DOI:10.1021/acs.jnatprod.4c01036.
- L.-P. He, X.-C. Luo, C.-X. Zhang and H.-W. Lin, Phytochemistry, 2024, 228, 114250–114256, DOI:10.1016/j.phytochem.2024.114250.
- D. Peng, X. Luo, R. Zhu, W. Tong, Y. Yang, G. Li and Q. Wang, Org. Lett., 2024, 26, 5794–5798, DOI:10.1021/acs.orglett.4c01976.
- X. Liu, Q. Wang, Y. Zhang and H. Zhang, Mar. Drugs, 2024, 22, 477–486, DOI:10.3390/md22100477.
- L. Gris, M. R. Prinsep, L. M. Peters and C. N. Battershill, Mar. Drugs, 2024, 22, 231–243, DOI:10.3390/md22050231.
- C. J. de Muizon, C. Moriou, M. Levasseur, D. Touboul, B. I. Iorga, H. Nedev, E. V. Elslande, P. Retailleau, S. Petek, E. Folcher, A. Bianchi, M. Thomas, S. Viallon, S. Peyroche, S. Nahle, M. Rousseau and A. Al-Mourabit, Mar. Drugs, 2024, 22, 196–215, DOI:10.3390/md22050196.
- S. Hayes, Y. Lu, B. H. A. Rehm and R. A. Davis, Beilstein J. Org. Chem., 2024, 20, 3205–3214, DOI:10.3762/bjoc.20.266.
- A. M. G. Ang, M. M. Uy, E. Ohta, H. Ômura and S. Ohta, Chem. Biodiversity, 2024, 21, e202400962, DOI:10.1002/cbdv.202400962.
- S.-I. Kurimoto, K. Inoue, T. Ohno and T. Kubota, Beilstein J. Org. Chem., 2024, 20, 3215–3220, DOI:10.3762/bjoc.20.267.
- X. Yu, X. Han, Y. Mi, Y. Cui, A. Fu, K. Liu, X. Li, X. Tang and G. Li, Phytochemistry, 2024, 226, 114220–114230, DOI:10.1016/j.phytochem.2024.114220.
- J.-X. Li, R.-Y. Shang, D.-D. Xie, X.-C. Luo, T.-Y. Hu, B.-H. Cheng, H.-W. Lin and W.-H. Jiao, J. Nat. Prod., 2024, 87, 396–403, DOI:10.1021/acs.jnatprod.3c01239.
- J. Wang, Y.-L. Yan, X.-Y. Yu, J.-Y. Pan, X.-L. Liu, L.-L. Hong and B. Wang, Mar. Drugs, 2024, 22, 183–193, DOI:10.3390/md22040183.
- D.-X. Fan, X.-C. Luo, Y.-F. Ding, L.-Y. Liu, X. Wang, J.-Y. Pan, Y.-Y. Ji, J. Wang, C. Li, L.-L. Hong and H.-W. Lin, Phytochemistry, 2024, 220, 114017–114025, DOI:10.1016/j.phytochem.2024.114017.
- Y. Cho, C. Bawkar, J. M. Hyun, M. J. Song, K. Jeong and Y.-J. Lee, J. Nat. Prod., 2024, 87, 358–364, DOI:10.1021/acs.jnatprod.3c01104.
- A. A. E. Said, I. M. Abdel-Rahman, Y. A. Mostafa, E. Z. Attia, M. N. Samy, U. R. Abdelmohsen, K. Matsunami, M. A. Fouad and Y. G. Gouda, BMC Chem., 2024, 18, 213–222, DOI:10.1186/s13065-024-01325-w.
- A. Ruiz-Molina, D. Pech-Puch, R. E. Millán, L. Ageitos, H. Villegas-Hernández, J. Pachón, J. P. Sestelo, J. Sánchez-Céspedes, J. Rodríguez and C. Jiménez, Chem.–Eur. J., 2024, 30, e202401844, DOI:10.1002/chem.202401844.
- S. S. H. Olsen, S. K. Morrow, J. L. Szabo, M. N. Teng, K. C. Tran, C. D. Amsler, J. B. McClintock and B. J. Baker, Mar. Drugs, 2024, 22, 551–561, DOI:10.3390/md22120551.
- L.-G. Zheng, Y.-Y. Chen, C.-C. Liaw, Y.-C. Lin, S.-Y. Chien, Y.-H. Lo, Y.-C. Tsai, H.-C. Hu, Y.-Y. Chen, T.-L. Hwang, T.-M. Ou and P.-J. Sung, Phytochem. Lett., 2024, 61, 177–181, DOI:10.1016/j.phytol.2024.04.013.
- R. L. Mama, C. D. Gelani, J. M. T. Daluz, M. M. Uy, E. Ohta and S. Ohta, Nat. Prod. Res., 2024, 38, 2395–2403, DOI:10.1080/14786419.2023.2175359.
- M.-C. Liu, X.-J. Liao, X.-W. Xing, S.-H. Xu and B.-X. Zhao, J. Asian Nat. Prod. Res., 2024, 26, 1254–1260, DOI:10.1080/10286020.2024.2360114.
- Y.-F. Ding, L.-Y. Liu, J. Tang, D.-X. Fan, Y.-Y. Ji, H.-W. Lin, J. Wang and L.-L. Hong, Nat. Prod. Res., 2024, 38, 2562–2568, DOI:10.1080/14786419.2023.2188588.
- J. Liu, Z.-Y. Liu, Y. Fu, Y.-C. Gu, S.-W. Li, H.-Y. Zhang and Y.-W. Guo, Chem. Biodiversity, 2024, 21, e202400519, DOI:10.1002/cbdv.202400519.
- X. Leng, H. He, J. E. H. Lazaro, X. Chen, H. Ouyang, T. Li, X. Yan and S. He, Phytochemistry, 2024, 223, 114097, DOI:10.1016/j.phytochem.2024.114097.
- P. Vlachou, N. Tsafantakis, N. Milic, A. Polyzois, E. Baira, A. Termentzi, G. L. Goff, J. Ouazzani and N. Fokialakis, Mar. Drugs, 2024, 22, 522–540, DOI:10.3390/md22110522.
- M. Sauer, A. M. Kany, S. Götze, R. Müller and C. Beemelmanns, Angew. Chem., Int. Ed., 2024, 63, e202416036, DOI:10.1002/anie.202416036.
- N. Jamey and L. Ferrié, Org. Lett., 2024, 26, 5741–5745, DOI:10.1021/acs.orglett.4c01893.
- H. Ogawa, Y. Nagata and H. Nakamura, Chin. J. Chem., 2024, 42, 3023–3028, DOI:10.1002/cjoc.202400637.
- D. M. Wilson and R. Britton, J. Am. Chem. Soc., 2024, 146, 8456–8463, DOI:10.1021/jacs.3c14553.
- A. Leblond, A. Nguyen, C. Alcover, K. Leblanc, J.-F. Gallard, D. Joseph, E. Poupon and M. A. Beniddir, Org. Lett., 2024, 26, 2163–2168, DOI:10.1021/acs.orglett.4c00194.
- K. Yamazaki, Y. Okuda, A. Takaya and T. Nemoto, Org. Lett., 2024, 26, 670–675, DOI:10.1021/acs.orglett.3c04039.
- S. Tsuruda, T. Sato, H. Aoyama, N. Sirimangkalakitti, S. Fujimura, M. Arisawa and K. Murai, J. Nat. Prod., 2024, 87, 1556–1562, DOI:10.1021/acs.jnatprod.4c00129.
- L. Tham, B. W. Bulcock, S. Sala, G. L. Nealon, G. R. Flematti, S. A. Moggach and M. J. Piggott, J. Nat. Prod., 2024, 87, 2499–2506, DOI:10.1021/acs.jnatprod.4c00973.
- L. Yi, S. Tan, L. V. White, M.-Y. Liang and M. G. Banwell, J. Nat. Prod., 2024, 87, 1521–1531, DOI:10.1021/acs.jnatprod.3c01177.
- T. Tsukamoto, K. Takahashi, N. Murase, K. Someya, F. Sakata, T. Yue, T. Kusakabe and K. Kato, Org. Lett., 2024, 26, 9011–9016, DOI:10.1021/acs.orglett.4c03158.
- G. L. Barnes, N. L. Magann, D. Perrotta, F. M. Hörmann, S. Fernandez, P. Vydyam, J.-Y. Choi, J. Prudhomme, A. Neal, K. G. Le Roch, C. B. Mamoun and C. D. Vanderwal, J. Am. Chem. Soc., 2024, 146, 29883–29894, DOI:10.1021/jacs.4c11897.
- S. P. Rezgui, J. Farhi, H. Yu, Z. P. Sercel, S. C. Virgil and B. M. Stoltz, Chem. Sci., 2024, 15, 12284–12290, 10.1039/D4SC02981J.
- C. Tolle, M. Fresia, P. G. Jones and T. Lindel, ChemPhotoChem, 2024, 8, e202400065, DOI:10.1002/cptc.202400065.
- A. P. Morrow and M. W. Smith, J. Am. Chem. Soc., 2024, 146, 2913–2918, DOI:10.1021/jacs.3c14120.
- R. Fujita, K. Ooka, H. Wasada, Y. Ichikawa and S. Hosokawa, Org. Biomol. Chem., 2024, 22, 1901–1906, 10.1039/D4OB00088A.
- A. Ortigosa-Palomo, F. Quiñonero, R. Ortiz, F. Sarabia, J. Prados and C. Melguizo, Mar. Drugs, 2024, 22, 101–122, DOI:10.3390/md22030101.
- Q.-B. Yang and L.-F. Liang, Mar. Drugs, 2024, 22, 213–234, DOI:10.3390/md22050213.
- N. Barzkar, S. Sukhikh and O. Babich, Int. J. Biol. Macromol., 2024, 280, 135823–135840, DOI:10.1016/j.ijbiomac.2024.135823.
- J. M. J. Nolsøe, J. Underhaug, Å. M. Sørskar, S. G. Antonsen, K. E. Malterud, O. Gani, Q. Fan, M. Hjorth, T. Sæther, T. V. Hansen and Y. H. Stenstrøm, Molecules, 2024, 29, 994–1008, DOI:10.3390/molecules29050994.
- D. S. Dissanayake, D. P. Nagahawatta, J.-S. Lee and Y.-J. Jeon, Mar. Drugs, 2024, 22, 426–447, DOI:10.3390/md22090426.
- N. K. Tailor, G. Deswal and A. S. Grewal, Chemistry, 2024, 6, 677–694, DOI:10.3390/chemistry6040040.
- M. Xu, Z. Bai, B. Xie, R. Peng, Z. Du, Y. Liu, G. Zhang, S. Yan, X. Xiao and S. Qin, Molecules, 2024, 29, 933–956, DOI:10.3390/molecules29050933.
- P. F. Montenegro, G. N. Pham, F. M. Abdoul-Latif, E. Taffin-de-Givenchy and M. Mehiri, Mar. Drugs, 2024, 22, 132–166, DOI:10.3390/md22030132.
- A. Khatun, K. Hazra, B. Mahato, R. Koley, R. Ghorai, A. Adhikary, B. Debnath and S. Das, Eur. J. Med. Chem. Rep., 2024, 11, 100158–100196, DOI:10.1016/j.ejmcr.2024.100158.
- J. Xia, X. Chen, G. Li, P. Qiu, W. Wang and Z. Shao, Mar. Drugs, 2024, 22, 447–482, DOI:10.3390/md22100447.
- L. A. Amador, E. E. Colón-Lorenzo, A. D. Rodríguez and A. E. Serrano, Life, 2024, 14, 684–696, DOI:10.3390/life14060684.
- A. Capuano, G. D'Urso, M. Aliberti, D. Ruggiero, S. Terracciano, C. Festa, A. Tosco, M. G. Chini, G. Lauro, G. Bifulco and A. Casapullo, Mar. Drugs, 2024, 22, 41–51, DOI:10.3390/md22010041.
- Y. Sun, A. Dakiiwa, M. Zhang, T. Shibata and M. Kita, Chem.–Eur. J., 2024, 30, e202402049, DOI:10.1002/chem.202402049.
- C.-L. Gao, J.-Q. Song, Z.-N. Yang, H. Wang, X.-Y. Wu, C. Shao, H.-X. Dai, K. Chen, Y.-W. Guo, T. Pang and X.-W. Li, J. Am. Chem. Soc., 2024, 146, 28384–28397, DOI:10.1021/jacs.4c09695.
- E. J. Strong, L. Tan, S. Hayes, H. Whyte, R. A. Davis and N. P. West, Mar. Drugs, 2024, 22, 298–306, DOI:10.3390/md22070298.
- M. Sinane, C. Grunberger, L. Gentile, C. Moriou, V. Chaker, P. Coutrot, A. Guenneguez, M.-A. Poullaouec, S. Connan, V. Stiger-Pouvreau, M. Zubia, Y. Fleury, S. Cérantola, N. Kervarec, A. Al-Mourabit, S. Petek and C. Voisset, Mar. Drugs, 2024, 22, 456–473, DOI:10.3390/md22100456.
- G. Risi, M. Liu, F. Vairoletti, R. J. Quinn and G. Salinas, J. Nat. Prod., 2024, 87, 1532–1539, DOI:10.1021/acs.jnatprod.3c01212.
- X. Wang, W. Xu, Z. Wang, Q. Yu, L. Yuan, Y. Liu, J. Sang, W. Li, S. Zhu, W. Jiang, Z. Li, W. Zhang and Y. Dang, J. Nat. Prod., 2024, 87, 713–721, DOI:10.1021/acs.jnatprod.3c00811.
- T. Tian, Y. Ogura, Y. Ise, H. Takikawa, S. Okada, S. Matsunaga and G.-L. Tang, J. Org. Chem., 2024, 89, 9135–9138, DOI:10.1021/acs.joc.4c00913.
- M. Orfanoudaki, R. K. Akee, L. Martínez-Fructuoso, D. Wang, J. A. Kelley, E. A. Smith, C. J. Henrich, M. J. Schnermann, B. R. O'Keefe and T. Grkovic, J. Nat. Prod., 2024, 87, 415–423, DOI:10.1021/acs.jnatprod.3c01113.
- A. Galitz, S. Vargas, O. P. Thomas, M. M. Reddy, G. Wörheide and D. Erpenbeck, Chem. Biodiversity, 2024, 21, e202400549, DOI:10.1002/cbdv.202400549.
- A. G. Silva, J. V. C. Rostirola, F. D. Speri, J. G. Pina, M. V. Kitahara, G. B. Longato and J. M. Sciani, J. Cell. Mol. Med., 2024, 28, e70165, DOI:10.1111/jcmm.70165.
- S. P. D. Senadeera, D. Wang, B. A. P. Wilson, E. A. Smith, A. Wamiru, J. A. Martinez Fiesco, L. Du, P. Zhang, B. R. O'Keefe and J. A. Beutler, J. Nat. Prod., 2024, 87, 2014–2020, DOI:10.1021/acs.jnatprod.4c00477.
- Y. Sun, X. Zhang, F. Jiang, M. Zhang, W. Wu and Y. Sun, J. Am. Chem. Soc., 2024, 146, 32305–32310, DOI:10.1021/jacs.4c14081.
- M.-Y. Cheng, I.-C. Hsu, S.-Y. Huang, Y.-T. Chuang, T.-Y. Ke, H.-W. Chang, T.-H. Chu, C.-Y. Chen and Y.-B. Cheng, Mar. Drugs, 2024, 22, 219, DOI:10.3390/md22050219.
- J. Lackner, C. Alberti, T. Bock, K. Neßmerak, E. Urban, S. Khom and N. Schützenmeister, Chem.–Eur. J., 2024, 30, e202401921, DOI:10.1002/chem.202401921.
- A. Koutsaviti, M. Kvasnicová, G. Gonzalez, T. Štenclová, S. Agusti, C. M. Duarte, L. Rarová, M. Strnad, V. Roussis and E. Ioannou, Chem. Biodiversity, 2024, 21, e202400235, DOI:10.1002/cbdv.202400235.
- X. Yu, X. Han, Y. Mi, Y. Cui, A. Fu, K. Liu, X. Tang and G. Li, J. Nat. Prod., 2024, 87, 1150–1158, DOI:10.1021/acs.jnatprod.4c00112.
- Y. Zong, T.-Y. Jin, J.-J. Yang, K.-Y. Wang, X. Shi, Y. Zhang and P.-L. Li, Mar. Drugs, 2024, 22, 145, DOI:10.3390/md22040145.
- V. Kurnianda, H.-C. Hu, P.-J. Sung, Y. Hayashi, K. Mori-Yasumoto, A. Suetake, H. Nakayama, M. Yasumoto-Hirose, Y. Tsutsumi, G. Koja, Y.-C. Tsai, T. Jomori and J. Tanaka, Tetrahedron Lett., 2024, 151, 155323, DOI:10.1016/j.tetlet.2024.155323.
- S. S. H. Olsen, S. Afoullouss, R. M. Young, M. Johnson, A. L. Allcock, M. N. Teng, K. C. Tran and B. J. Baker, Org. Lett., 2024, 26, 9419–9424, DOI:10.1021/acs.orglett.4c02549.
- T. Kobayashi, R. Sugitate, K. Uchida, Y. Kawamoto and H. Ito, Org. Lett., 2024, 26, 1803–1806, DOI:10.1021/acs.orglett.3c04314.
- T. Kobayashi, F. Hayano, A. Kamiya, Y. Kawamoto and H. Ito, Org. Lett., 2024, 26, 5105–5109, DOI:10.1021/acs.orglett.4c01384.
- H. Abe, N. Inaba, M. Nomura, Y. Ogura, N. Fukazawa, R. Saito-Moriya, Y. Kawamoto, T. Kobayashi and H. Ito, Tetrahedron Lett., 2024, 139, 154991, DOI:10.1016/j.tetlet.2024.154991.
- G. Stakanovs, A. Blazevica, D. Rasina, S. Belyakov and A. Jirgensons, Org. Lett., 2024, 26, 8074–8078, DOI:10.1021/acs.orglett.4c02942.
- C. Wang, J. Zhang, K. Li, J. Yang, L. Li, S. Wang, H. Hou and P. Li, Mar. Drugs, 2024, 22, 442, DOI:10.3390/md22100442.
- X. Chen, J. Zhang, J. Yang, B. Li, T. Li, H. Ouyang, W. Lin, H. Hu, X. Yan and S. He, Mar. Drugs, 2024, 22, 392, DOI:10.3390/md22090392.
- D. H. Dethe, S. Juyal, N. Sharma and U. Kundu, Org. Lett., 2024, 26, 3010–3013, DOI:10.1021/acs.orglett.4c00605.
- X. Han, H. Wang, B. Li, X. Chen, T. Li, X. Yan, H. Ouyang, W. Lin and S. He, Mar. Drugs, 2024, 22, 157, DOI:10.3390/md22040157.
- J. Marrero, L. A. Amador, I. M. Novitskiy, A. G. Kutateladze and A. D. Rodríguez, Molecules, 2024, 29, 2493, DOI:10.3390/molecules29112493.
- J. Marrero, L. A. Amador, I. M. Novitskiy, A. G. Kutateladze and A. D. Rodríguez, Molecules, 2025, 30, 2929, DOI:10.3390/molecules30142929.
- S.-W. Li, C. Cuadrado, J. Liu, H. Li, W. Tang, A. H. Daranas and Y.-W. Guo, J. Chromatogr., A, 2024, 1736, 465416, DOI:10.1016/j.chroma.2024.465416.
- D. Zhang, Y. Li, X. Li, X. Han, Z. Wang and G. Li, Nat. Prod. Res., 2024, 38, 97–102, DOI:10.1080/14786419.2022.2106568.
- Z.-Y. Xia, M.-M. Sun, Y. Jin, M.-Z. Su, S.-W. Li, H. Wang and Y.-W. Guo, Phytochemistry, 2024, 219, 113960, DOI:10.1016/j.phytochem.2023.113960.
- N. B. M. Janzing, C. H. R. Senges, P. Dietze, B. Haltli, D. H. Marchbank, R. G. Kerr and J. E. Bandow, Proteomics, 2024, 24, 2300390, DOI:10.1002/pmic.202300390.
- Y. Zong, J.-J. Yang, K. Li, Y.-F. Pei, H.-Y. Hou, Y. Zhang, C.-L. Wang and P.-L. Li, Phytochemistry, 2024, 223, 114113, DOI:10.1016/j.phytochem.2024.114113.
- S.-H. Zhu, Y.-M. Chang, M.-Z. Su, L.-G. Yao, S.-W. Li, H. Wang and Y.-W. Guo, Mar. Drugs, 2024, 22, 50, DOI:10.3390/md22010050.
- J.-H. Sheu, L.-G. Zheng, Y.-Y. Chen, S.-Y. Chien and P.-J. Sung, Phytochem. Lett., 2024, 63, 14–17, DOI:10.1016/j.phytol.2024.07.012.
- M. Shaaban and M. A. Ghani, Z. Naturforsch., C: J. Biosci., 2024, 79, 81–87, DOI:10.1515/znc-2024-0004.
- C. Wang, J. Zhang, T. Jin, X. Shi, M. Wang, W. Zhang, Y. Zong, G. Li and P. Li, Nat. Prod. Res., 2024, 38, 3597–3604, DOI:10.1080/14786419.2023.2258436.
- H. Peng, Y. Zeng, R. Zhang, L. Yang, F. Wu, C. Gai, J. Yuan, W. Chang, H. Dai and X. Wang, Mar. Drugs, 2024, 22, 536, DOI:10.3390/md22120536.
- Y. Mi, Y. Yang, X. Tang, X. Han and G. Li, Chin. J. Chem., 2024, 42, 815–822, DOI:10.1002/cjoc.202300592.
- Y. Mi, X. Han, X. Yu, L. Li, X. Tang and G. Li, Phytochemistry, 2024, 223, 114109, DOI:10.1016/j.phytochem.2024.114109.
- T. A. Mohamed, S. M. Abdelmawgoud, A. A. Hamdy, M. A. A. Ibrahim, A. I. Elshamy, M. A. M. Atia, H. A. Kassem, M.-E. F. Hegazy and N. M. Selim, Nat. Prod. Res., 2024, 38, 512–522, DOI:10.1080/14786419.2022.2130915.
- Y.-Y. Cui, Y. Jin, R.-N. Sun, X. Wang, C.-L. Gao, X.-Y. Cui, K.-X. Chen, Y.-L. Sun, Y.-W. Guo, J. Li and X.-W. Li, J. Med. Chem., 2024, 67, 12248–12260, DOI:10.1021/acs.jmedchem.4c00950.
- C. Huang, Y. Jin, R. Sun, K. Hu, L. Yao, Y. Guo, Z. Yuan and X. Li, Chem. Biodiversity, 2024, 21, e202401146, DOI:10.1002/cbdv.202401146.
- M. Sun, S. Li, J. Zeng, Y. Guo, C. Wang and M. Su, Mar. Drugs, 2024, 22, 553, DOI:10.3390/md22120553.
- P. D. Scesa, S. P. Roche and L. West, Org. Lett., 2024, 26, 1123–1127, DOI:10.1021/acs.orglett.3c03604.
- D. S. Lin, G. Späth, Z. Meng, L. H. E. Wieske, C. Farès and A. Fürstner, J. Am. Chem. Soc., 2024, 146, 24250–24256, DOI:10.1021/jacs.4c09467.
- E. J. Simmons, D. B. Ryffel, D. A. Lopez, Y. D. Boyko and D. Sarlah, J. Am. Chem. Soc., 2024, 147, 130–135, DOI:10.1021/jacs.4c16629.
- H. Takamura, Y. Sugitani, R. Morishita, T. Yorisue and I. Kadota, Org. Biomol. Chem., 2024, 22, 5739, 10.1039/D4OB00698D.
- E. M. Elkady, H. R. Tadros, Y. A. Soliman, M. Raafat and A. M. Abdel-Tawab, Egypt. J. Aquat. Res., 2024, 50, 234–240, DOI:10.1016/j.ejar.2024.06.004.
- S.-W. Tsai, Y.-C. Cheng, Y.-H. Chao and D.-H. Yang, J. Inflammation Res., 2024, 17, 8299–8311, DOI:10.2147/JIR.S476847.
- L.-M. Ke, D.-D. Yu, M.-Z. Su, L. Cui and Y.-W. Guo, Mar. Drugs, 2024, 22, 95, DOI:10.3390/md22020095.
- X.-Y. Meng, K.-J. Wang, S.-Z. Ye, J.-F. Chen, Z.-Y. Chen, Z.-Y. Zhang, W.-Q. Yin, X.-L. Jia, Y. Li, R. Yu and Q. Ma, Biochem. Pharmacol., 2024, 220, 116011, DOI:10.1016/j.bcp.2023.116011.
- R.-Y. Fang, Y.-W. Liu, Y.-G. Goan, J.-J. Lin, J.-H. Su, W.-T. Wu, Y.-J. Wu and N. Ismail, Adv. Pharmacol. Pharm. Sci., 2024, 2024, 6695837, DOI:10.1155/2024/6695837.
- M. A. Bashar, A. Hamed, M. A. M. El-Tabakh, A. H. Hashem, A. A. Zaki, A. A. Al-Askar, E. S. Abou-Amra, M. E. El-Beeh, A. B. M. Mehany, M. Shaaban, A. M. Abdelaziz, K. A. Shaaban and A. I. Hasaballah, Sci. Rep., 2024, 14, 28028, DOI:10.1038/s41598-024-75446-6.
- Y.-W. Chuang, W.-C. Ting, S.-Y. Chien, L.-G. Zheng, C.-Y. Huang, Z.-H. Wen, Y.-J. Wu, J. Tanaka, Y.-Y. Chen, J.-H. Su and P.-J. Sung, Phytochem. Lett., 2024, 64, 117–122, DOI:10.1016/j.phytol.2024.10.008.
- H. N. Do, Y.-T. Chen, S.-Y. Chien, Y.-Y. Chen, M. M. Zhang, L. K. Tsou, J.-J. Chen, Z.-H. Wen, Y.-H. Lo, L.-G. Zheng and P.-J. Sung, RSC Adv., 2024, 14, 17195–17201, 10.1039/D4RA03062A.
- H. N. Do, C.-Y. Huang, S.-Y. Chien, L.-G. Zheng, Y.-Y. Chen, Y.-T. Zeng, J.-H. Su, Z.-H. Wen, Y.-C. Chang and P.-J. Sung, Chem. Pharm. Bull., 2024, 72, 1110–1115, DOI:10.1248/cpb.c24-00574.
- H.-W. Chen, F.-C. Liu, H.-M. Kuo, S.-H. Tang, G.-H. Niu, M. M. Zhang, L. K. Tsou, P.-J. Sung and Z.-H. Wen, Biomed. Pharmacother., 2024, 172, 116279, DOI:10.1016/j.biopha.2024.116279.
- G. Yao, M. R. Parris, W. C. Kuo, P. Pörzgen, B. Castillo, E. S. Mason, A. Chinchilla, J. Huang, S. Suzuki, R. Ross, E. Akana, S. V. Schuit, S. P. Miller, R. Penner, H.-S. Sun, Z.-P. Feng, K. G. Hull, D. Romo, A. Fleig and F. D. Horgen, J. Nat. Prod., 2024, 87, 783–797, DOI:10.1021/acs.jnatprod.3c00942.
- S. Afoullouss, R. M. Young, L. K. Jennings, J. Doyle, K. Croke, D. Livorsi, J. H. Adams, M. P. Johnson, O. P. Thomas and A. L. Allcock, ACS Omega, 2024, 9, 41914–41922, DOI:10.1021/acsomega.4c06361.
- L. Betschart and K. Altmann, Angew. Chem., Int. Ed., 2024, 63, e202315423, DOI:10.1002/anie.202315423.
- A. H. Eissa, A. M. Abdel-Tawab, E. S. A. E. Hamed, F. Z. El-Ablack and S.-E. N. Ayyad, Nat. Prod. Res., 2024, 38, 4390–4398, DOI:10.1080/14786419.2023.2284259.
- T.-Y. Cheng, C.-J. Yang, P.-J. Chen, Y.-L. Chen, B.-R. Peng, T.-A. Kung, Z.-H. Wen, K.-H. Lai and H.-M. Chung, Phytochem. Lett., 2024, 61, 115–119, DOI:10.1016/j.phytol.2024.03.015.
- C.-L. Hung, L.-C. Huang, K.-Y. Shih, K.-H. Lai, B.-R. Peng, T.-L. Hwang, S.-Y. Chien, C.-Y. Li and H.-M. Chung, J. Mol. Struct., 2024, 1296, 136790, DOI:10.1016/j.molstruc.2023.136790.
- H.-M. Chung, Z.-Z. Tsai, C.-K. Hsu, P.-J. Chen, Y.-L. Chen, S.-Y. Chien, Z.-H. Wen, P.-J. Sung and K.-H. Lai, Rec. Nat. Prod., 2024, 18, 357, DOI:10.25135/rnp.455.2402.3141.
- T.-H. Lee, J.-L. Chen, C.-H. Chang, M.-M. Tsai, H.-C. Tseng, Y.-C. Chang, V. Shanmugam and H.-L. Hsieh, Biomedicines, 2024, 12, 226, DOI:10.3390/biomedicines12010226.
- C.-Q. Liu, Q.-B. Yang, L. Zhang and L.-F. Liang, Mar. Drugs, 2024, 22, 402, DOI:10.3390/md22090402.
- H. L. Hammond and C. J. Roy, Toxins, 2024, 16, 417, DOI:10.3390/toxins16100417.
- V. M. Dembitsky, Microbiol. Res., 2024, 15, 1516–1575, DOI:10.3390/microbiolres15030103.
- K. A. Elnahriry, D. C. Wai, L. M. Ashwood, M. U. Naseem, T. G. Szanto, S. Guo, G. Panyi, P. J. Prentis and R. S. Norton, Biochim. Biophys. Acta, Proteins Proteomics, 2024, 1872, 140952, DOI:10.1016/j.bbapap.2023.140952.
- A. N. Kvetkina, S. D. Oreshkov, P. A. Mironov, M. M. Zaigraev, A. A. Klimovich, Y. V. Deriavko, A. S. Menshov, D. S. Kulbatskii, Y. A. Logashina, Y. A. Andreev, A. O. Chugunov, M. P. Kirpichnikov, E. N. Lyukmanova, E. V. Leychenko and Z. O. Shenkarev, Mar. Drugs, 2024, 22, 542, DOI:10.3390/md22120542.
- T. Homma, M. Ishida, Y. Nagashima and K. Shiomi, J. Venom. Anim. Toxins Incl. Trop. Dis., 2024, 30, e20240019, DOI:10.1590/1678-9199-jvatitd-2024-0019.
- B.-Y. Wu and C.-K. Lin, Asian J. Org. Chem., 2024, 13, e202400074, DOI:10.1002/ajoc.202400074.
- Y.-Y. Kou, J. Liu, Y.-T. Chang, L.-Y. Liu, F. Sun, Y.-L. Li, J.-R. Leng, H.-W. Lin and F. Yang, Cell. Oncol., 2024, 47, 1863–1878, DOI:10.1007/s13402-024-00968-0.
- X. Zhang, M. Su, M. Zhu, S. Chen, Z. Gao, Y. Guo and X. Li, Chin. J. Nat. Med., 2024, 22, 1030–1039, DOI:10.1016/S1875-5364(24)60611-4.
- Y. A. T. Ngandjui, T. T. Kereeditse, I. Kamika, L. M. Madikizela and T. A. M. Msagati, Mar. Drugs, 2024, 22, 201, DOI:10.3390/md22050201.
- S. Nandi, S. Khatua, A. Nag, S. Sen, N. Chakraborty, A. Naskar, K. Acharya, R. H. Mekky, M. del Mar Contreras, D. Calina, I. Dini and J. Sharifi-Rad, Curr. Res. Biotechnol., 2024, 7, 100167, DOI:10.1016/j.crbiot.2023.100167.
- M. Tost and U. Kazmaier, Mar. Drugs, 2024, 22, 165, DOI:10.3390/md22040165.
- I. W. Mudianta, J. E. Siregar, A. F. M. Rizki, W. A. Azmi, N. E. Pravitasari, G. M. Gholam, F. R. Putri, R. Kristiana, N. K. D. Cahyani and I. M. Artika, Biochem. Biophys. Res. Commun., 2024, 737, 150921, DOI:10.1016/j.bbrc.2024.150921.
- P. Kancharla, D. Ortiz, C. M. Fargo, X. Zhang, Y. Li, M. Sanchez, A. Kumar, M. Yeluguri, R. A. Dodean, D. Caridha, M. S. Madejczyk, M. Martin, X. Jin, C. Blount, R. Chetree, K. Pannone, H. T. Dinh, J. DeLuca, M. Evans, R. Nadeau, C. Vuong, S. Leed, W. E. Dennis, N. Roncal, B. S. Pybus, P. J. Lee, A. Roth, K. A. Reynolds, J. X. Kelly and S. M. Landfear, J. Med. Chem., 2024, 67, 8323–8345, DOI:10.1021/acs.jmedchem.4c00517.
- M. Wei, J. Chen, Y. Song, J.-P. Monserrat, Y. Zhang and L. Shen, Eur. J. Med. Chem., 2024, 269, 116294, DOI:10.1016/j.ejmech.2024.116294.
- P. Sivaganesan, S. VL, A. Sahoo, C. Elanchezhian, G. Nataraj and S. Chaudhuri, ChemistrySelect, 2024, 9, e202403112, DOI:10.1002/slct.202403112.
- V.-P. M. Mitsiou, A.-M. N. Antonaki, M. D. Douka and K. E. Litinas, Molecules, 2024, 29, 4032, DOI:10.3390/molecules29174032.
- S. C. Finch, D. T. Harwood, M. J. Boundy and A. I. Selwood, Mar. Drugs, 2024, 22, 129, DOI:10.3390/md22030129.
- Z.-R. Lin, H.-X. Geng and R.-C. Yu, J. Hazard. Mater., 2024, 469, 133896, DOI:10.1016/j.jhazmat.2024.133896.
- J. L. García-Corona, C. Fabioux, J. Vanmaldergem, S. Petek, A. Derrien, A. Terre-Terrillon, L. Bressolier, F. Breton and H. Hegaret, Harmful Algae, 2024, 133, 102607, DOI:10.1016/j.hal.2024.102607.
- T. Teng, Z. Zheng, W. Jiao, N. Liu, A. Wang, M. Liu, L. Xie, Z. Yang, J. Hu and Z. Bao, Genes, 2024, 15, 365, DOI:10.3390/genes15030365.
- G. Chiappa, G. Fassio, M. V. Modica and M. Oliverio, Toxins, 2024, 16, 348, DOI:10.3390/toxins16080348.
- Z. Ratibou, N. Inguimbert and S. Dutertre, Toxins, 2024, 16, 94, DOI:10.3390/toxins16020094.
- L. Martínez-Hernández, E. López-Vera and M. B. Aguilar, Toxins, 2024, 16, 504, DOI:10.3390/toxins16120504.
- S. Shekh, S. Dhannura, P. Dhurjad, C. Ravali, S. M M, S. Kakkat, Vishwajyothi, M. Vijayasarathy, R. Sonti and K. H. Gowd, Toxicon, 2024, 250, 108087, DOI:10.1016/j.toxicon.2024.108087.
- M. Wang, Z. Liao, D. Zhangsun, Y. Wu and S. Luo, Molecules, 2024, 29, 1181, DOI:10.3390/molecules29051181.
- C. Lin, H. Qin, Y. Liao, J. Chen and B. Gao, Molecules, 2024, 29, 2846, DOI:10.3390/molecules29122846.
- H. Wang, X. Li, Y. Qiao, M. Wang, W. Wang, J. M. McIntosh, D. Zhangsun and S. Luo, Mar. Drugs, 2024, 22, 49, DOI:10.3390/md22010049.
- X. Guo, L. He, W. Xu, W. Wang, X. Feng, Y. Fu, X. Zhang, R.-B. Ding, X. Qi, J. Bao and S. Luo, Mar. Drugs, 2024, 22, 252, DOI:10.3390/md22060252.
- X. Li, H.-S. Tae, S. Chen, A. Yousuf, L. Huang, J. Zhang, T. Jiang, D. J. Adams and R. Yu, J. Med. Chem., 2024, 67, 971–987, DOI:10.1021/acs.jmedchem.3c00979.
- S. Raghuraman, J. Carter, M. Walter, M. Karthikeyan, K. Chase, M. L. Giglio, M. Giacobassi, R. W. Teichert, H. Terlau and B. M. Olivera, Sci. Rep., 2024, 14, 31001, DOI:10.1038/s41598-024-82165-5.
- R. Sakai, K. Matsumura, H. Uchimasu, K. Miyako, T. Taniguchi, V. Raghavendra Rao Kovvuri, A. D. Acharige, K. G. Hull, D. Romo, L. Thaveepornkul, S. Chimnaronk, H. Miyamoto, A. Takada, H. Watari, M. J. Fujita and J. Sakaue, J. Org. Chem., 2024, 89, 5977–5987, DOI:10.1021/acs.joc.3c02765.
- Y. Li, K. Takamatsu, T. Aota and H. Konno, Org. Lett., 2024, 26, 3375–3379, DOI:10.1021/acs.orglett.4c00808.
- D. M. Dukes, V. K. Atanassov and J. M. Smith, Chem. Sci., 2024, 15, 16554–16558, 10.1039/D4SC04910A.
- L. A. Segura-Quezada, E. D. Hernández-Velázquez, A. R. Corrales-Escobosa, C. de León-Solis and C. R. Solorio-Alvarado, Chem. Biodiversity, 2024, 21, e202300883, DOI:10.1002/cbdv.202300883.
- M. A. Tammam, M. I. Gamal El-Din, A. Aouidate and A. El-Demerdash, Bioorg. Chem., 2024, 151, 107654, DOI:10.1016/j.bioorg.2024.107654.
- P. J. Batista, G. Nuzzo, C. Gallo, D. Carbone, M. dell'Isola, M. Affuso, G. Barra, F. Albiani, F. Crocetta, R. Virgili, V. Mazzella, D. Castiglia, G. d'Ippolito, E. Manzo and A. Fontana, Mar. Drugs, 2024, 22, 75, DOI:10.3390/md22020075.
- A. L. Agognon, M. Casertano, A. Vito, S. Orso, S. Cabaro, F. Mormone, C. Morelli, G. Perruolo, P. Formisano, M. Menna, C. Imperatore and F. Oriente, Int. J. Mol. Sci., 2024, 25, 6039, DOI:10.3390/ijms25116039.
- N. Spacho, M. Casertano, C. Imperatore, C. Papadopoulos, M. Menna and N. Eleftheriadis, Chem.–Eur. J., 2024, 30, e202402279, DOI:10.1002/chem.202402279.
- H. Wu, G. Lv, L. Liu, R. Hu, F. Zhao, M. Song, S. Zhang, H. Fan, S. Dai, S. u. Rehman, H. Wang and X. Mou, J. Nat. Prod., 2024, 87, 2779–2789, DOI:10.1021/acs.jnatprod.4c00946.
- T. Kalnins, V. Vitkovska, M. Kazak, D. Zelencova-Gopejenko, M. Ozola, N. Narvaiss, M. Makrecka-Kuka, I. Domračeva, A. Kinens, B. Gukalova, N. Konrad, R. Aav, F. Bonato, D. Lucena-Agell, J. F. Díaz, E. Liepinsh and E. Suna, J. Med. Chem., 2024, 67, 9227–9259, DOI:10.1021/acs.jmedchem.4c00388.
- H. Zhang, X. Li, Z. Hui, S. Huang, M. Cai, W. Shi, Y. Lin, J. Shen, M. Sui, Q. Lai, Z. Shao, J. Dou, X. Luo, Y. Ge and X. Tang, Angew. Chem., Int. Ed., 2024, 63, e202318784, DOI:10.1002/anie.202318784.
- P. Landete, O.-A. Caliman-Sturdza, J. A. Lopez-Martin, L. Preotescu, M.-C. Luca, A. Kotanidou, P. Villares, S.-P. Iglesias, P. Guisado-Vasco, E.-M. Saiz-Lou, M. del Carmen Farinas-Alvarez, E. M. de Lucas, E. Perez-Alba, J.-M. Cisneros, V. Estrada, C. Hidalgo-Tenorio, G. Poulakou, M. Torralba, J. Fortun, P. Garcia-Ocana, A. Lemaignen, M. Marcos-Martin, M. Molina, R. Paredes, M. T. Perez-Rodriguez, D. Raev, P. Ryan, F. Meira, J. Gomez, N. Torres, D. Lopez-Mendoza, J. Jimeno and J.-F. Varona, Clin. Infect. Dis., 2024, 79, 910–919, DOI:10.1093/cid/ciae227.
- J. Zhang, L. Zhao, Y. Bai, S. Li, M. Zhang, B. Wei, X. Wang, Y. Xue, L. Li, G. Ma, Y. Tang and X. Wang, Bioorg. Med. Chem. Lett., 2024, 103, 129706, DOI:10.1016/j.bmcl.2024.129706.
- A. Povo-Retana, R. Landauro-Vera, C. Alvarez-Lucena, M. Cascante and L. Boscá, Molecules, 2024, 29, 331, DOI:10.3390/molecules29020331.
- R. Matsuta, S. Shigeno, T. Ohshiro, M. Ueda and K. Takada, Fish. Sci., 2024, 90, 787–793, DOI:10.1007/s12562-024-01803-z.
- A. A. Kicha, T. V. Malyarenko, A. S. Kuzmich, O. S. Malyarenko, A. I. Kalinovsky, R. S. Popov, D. K. Tolkanov and N. V. Ivanchina, Mar. Drugs, 2024, 22, 43, DOI:10.3390/md22010043.
- S. J. Smith, T. Wang and S. F. Cummins, Mar. Drugs, 2024, 22, 552, DOI:10.3390/md22120552.
- R. Zhang, Z. Lu, D. Wang, Z. Yan, X. Sun, X. Li, X. Yin, S. Wang and K. Li, Mar. Drugs, 2024, 22, 545, DOI:10.3390/md22120545.
- A. A. Kicha, D. K. Tolkanov, T. V. Malyarenko, O. S. Malyarenko, A. S. Kuzmich, A. I. Kalinovsky, R. S. Popov, V. A. Stonik, N. V. Ivanchina and P. S. Dmitrenok, Mar. Drugs, 2024, 22, 294, DOI:10.3390/md22070294.
- T. V. Malyarenko, V. M. Zakharenko, A. A. Kicha, A. I. Ponomarenko, I. V. Manzhulo, A. I. Kalinovsky, R. S. Popov, P. S. Dmitrenok and N. V. Ivanchina, Mar. Drugs, 2024, 22, 508, DOI:10.3390/md22110508.
- T. V. Malyarenko, A. A. Kicha, A. S. Kuzmich, O. S. Malyarenko, A. I. Kalinovsky, R. S. Popov, P. S. Dmitrenok, N. V. Ivanchina and V. A. Stonik, Mar. Drugs, 2024, 22, 19, DOI:10.3390/md22010019.
- A. S. Silchenko, A. I. Kalinovsky, S. A. Avilov, R. S. Popov, E. A. Chingizova, E. S. Menchinskaya, E. A. Zelepuga, K. M. Tabakmakher, V. G. Stepanov and V. I. Kalinin, Mar. Drugs, 2024, 22, 292, DOI:10.3390/md22070292.
- A. S. Silchenko, E. A. Chingizova, E. S. Menchinskaya, E. A. Zelepuga, A. I. Kalinovsky, S. A. Avilov, K. M. Tabakmakher, R. S. Popov, P. S. Dmitrenok, S. S. Dautov and V. I. Kalinin, Mar. Drugs, 2024, 22, 560, DOI:10.3390/md22120560.
- Y. Liu, Z. Lu, Z. Yan, X. Li, X. Yin, R. Zhang, Y. Li, S. Wang, R. Xie and K. Li, Chem. Biodiversity, 2024, 21, e202400335, DOI:10.1002/cbdv.202400335.
- Y. Liu, Z. Lu, Z. Yan, A. Lin, S. Han, Y. Li, X. Yang, X. Li, X. Yin, R. Zhang and K. Li, J. Chem. Ecol., 2024, 50, 185–196, DOI:10.1007/s10886-024-01483-0.
- F. M. Abdelkarem, H. K. Assaf, Y. A. Mostafa, A. Mahdy, M. F. Hussein, S. A. Ross and N. M. Mohamed, RSC Adv., 2024, 14, 14185–14193, 10.1039/D4RA01966K.
- E. A. Vasileva, D. V. Berdyshev, N. P. Mishchenko, A. V. Gerasimenko, E. S. Menchinskaya, E. A. Pislyagin, E. A. Chingizova, L. A. Kaluzhskiy, S. S. Dautov and S. A. Fedoreyev, Biomolecules, 2024, 14, 151, DOI:10.3390/biom14020151.
- I. A. Abdelmawgood, N. A. Mahana, A. M. Badr and A. S. Mohamed, Naunyn-Schmiedebergs Arch. Pharmakol., 2024, 397, 1803, DOI:10.1007/s00210-023-02678-0.
- D.-Y. Eum, C. Lee, C. S. Tran, J. Lee, S. Y. Park, M.-S. Jeong, Y. Jin, J. W. Shim, S. R. Lee, M. Koh, E. A. Vasileva, N. P. Mishchenko, S.-J. Park, S. H. Choi, Y. J. Choi, H. Yun and K. Heo, Toxicol. Res., 2024, 40, 409–419, DOI:10.1007/s43188-024-00232-5.
- S. Promtang, T. Sanguanphun, P. Chalorak, L. S. Pe, N. Niamnont, P. Sobhon and K. Meemon, ACS Chem. Neurosci., 2024, 15, 2182–2197, DOI:10.1021/acschemneuro.4c00008.
- L. Herr, J. R. Ye, S. W. Kang, S. T. Ro, Y. K. Chae, K. E. Lee, M. S. Kim, M. K. Jih, C. Lee, S. C. Choi and O. H. Nam, Dent. Traumatol., 2024, 40, 178–186, DOI:10.1111/edt.12891.
- A. Elkattan, M. Matsumoto, M. Nagata, Y. Mittraphab, G. A. Wahab, A. Ashour, A. A. Zaki, A. E. H. El-Sayed and K. Shimizu, Cytotechnology, 2024, 76, 721–734, DOI:10.1007/s10616-024-00649-8.
- C. Lu, X. Wang, J. Ma, M. Wang, W. Liu, G. Wang, Y. Ding, Z. Lin and Y. Li, Arch. Pharm., 2024, 357, 2300427, DOI:10.1002/ardp.202300427.
- M. Liao, X. Xia, Q. Meng, C. Zhu, B. Liao, J. Wang, L. Gou, X. Zhou, W. Yuan, L. Cheng and B. Ren, Br. J. Pharmacol., 2024, 181, 1857–1873, DOI:10.1111/bph.16333.
- E. S. Menchinskaya, S. A. Dyshlovoy, S. Venz, C. Jacobsen, J. Hauschild, T. Rohlfing, A. S. Silchenko, S. A. Avilov, S. Balabanov, C. Bokemeyer, D. L. Aminin, G. von Amsberg and F. Honecker, Mar. Drugs, 2024, 22, 20, DOI:10.3390/md22010020.
- E. S. Menchinskaya, E. A. Chingizova, E. A. Pislyagin, E. A. Yurchenko, A. A. Klimovich, E. A. Zelepuga, D. L. Aminin, S. A. Avilov and A. S. Silchenko, Mar. Drugs, 2024, 22, 474, DOI:10.3390/md22100474.
- S. Wang, C. Zhong, Y. Huang and P. Lu, Angew. Chem., Int. Ed., 2024, 63, e202400515, DOI:10.1002/anie.202400515.
- H. M. Ameen, A. Jayadev, G. Prasad and D. I. Nair, Molecules, 2024, 29, 4596, DOI:10.3390/molecules29194596.
- Z. Chebaro, J. E. Mesmar, A. Badran, A. Al-Sawalmih, M. Maresca and E. Baydoun, Biomolecules, 2024, 14, 991, DOI:10.3390/biom14080991.
- N. Wehbe, M. Bechelany, A. Badran, A. Al-Sawalmih, J. E. Mesmar and E. Baydoun, Pharmaceuticals, 2024, 17, 993, DOI:10.3390/ph17080993.
- N. A. Cacciola, P. De Cicco, R. Amico, F. Sepe, Y. Li, L. Grauso, M. F. Nanì, S. Scarpato, C. Zidorn, A. Mangoni and F. Borrelli, Phytother Res., 2024, 38, 4168–4176, DOI:10.1002/ptr.8269.
- W. R. Kem, F. Soti, J. R. Rocca and J. V. Johnson, Mar. Drugs, 2024, 22, 141, DOI:10.3390/md22040141.
- L. K. Jennings, N. Kaur, M. C. Ramos, F. Reyes, M. M. Reddy and O. P. Thomas, J. Nat. Prod., 2024, 87, 906–913, DOI:10.1021/acs.jnatprod.3c01186.
- O. A. Shepperson, P. W. R. Harris, M. A. Brimble and A. J. Cameron, Antibiotics, 2024, 13, 615, DOI:10.3390/antibiotics13070615.
- V. N. Safronova, P. V. Panteleev, R. N. Kruglikov, I. A. Bolosov, E. I. Finkina and T. V. Ovchinnikova, Russ. J. Bioorg. Chem., 2024, 50, 629–643, DOI:10.1134/S1068162024030087.
- M. Dhanji-Rapkova, R. G. Hatfield, D. I. Walker, C. Hooper, S. Alewijnse, C. Baker-Austin, A. D. Turner and J. M. Ritchie, Mar. Drugs, 2024, 22, 458, DOI:10.3390/md22100458.
- Y. Bai, W. Zhang, W. Zheng, X.-Z. Meng, Y. Duan, C. Zhang, F. Chen and K.-J. Wang, Biochem. Pharmacol., 2024, 226, 116344, DOI:10.1016/j.bcp.2024.116344.
- H.-H. Yu, L.-Y. Wu, P.-L. Hsu, C.-W. Lee and B.-C. Su, Probiotics Antimicrob. Proteins, 2024, 16, 1724, DOI:10.1007/s12602-023-10130-1.
- F.-C. Chiu, H.-M. Kuo, C.-L. Yu, P. Selvam, I.-L. Su, C.-C. Tseng, C.-H. Yuan and Z.-H. Wen, Free Radical Biol. Med., 2024, 220, 28–42, DOI:10.1016/j.freeradbiomed.2024.04.235.
- Q. Wu, C. Munschy, N. Bodin and W. Vetter, J. Agric. Food Chem., 2024, 72, 15643–15652, DOI:10.1021/acs.jafc.4c03503.
- A. Moores and V. G. Zuin Zeidler, Nat. Rev. Chem., 2025, 9, 649–650, DOI:10.1038/s41570-025-00757-9.
- M. Varadi, D. Bertoni, P. Magana, U. Paramval, I. Pidruchna, M. Radhakrishnan, M. Tsenkov, S. Nair, M. Mirdita, J. Yeo, O. Kovalevskiy, K. Tunyasuvunakool, A. Laydon, A. Zidek, H. Tomlinson, D. Hariharan, J. Abrahamson, T. Green, J. Jumper, E. Birney, M. Steinegger, D. Hassabis and S. Velankar, Nucleic Acids Res., 2024, 52, D368–D375, DOI:10.1093/nar/gkad1011.
- F. I. Saldivar-Gonzalez, V. D. Aldas-Bulos, J. L. Medina-Franco and F. Plisson, Chem. Sci., 2022, 13, 1526–1546, 10.1039/d1sc04471k.
- G. Hu and M. Qiu, Nat. Prod. Rep., 2023, 40, 1735–1753, 10.1039/d3np00025g.
- D. Meijer, M. A. Beniddir, C. W. Coley, Y. M. Mejri, M. Ozturk, J. J. J. van der Hooft, M. H. Medema and A. Skiredj, Nat. Prod. Rep., 2025, 42, 654–662, 10.1039/d4np00008k.
- A. Gangwal and A. Lavecchia, J. Med. Chem., 2025, 68, 3948–3969, DOI:10.1021/acs.jmedchem.4c01257.
- A. Rutz, W. Bittremieux, R. Schmid, O. Cailloux, J. J. J. van der Hooft and M. A. Beniddir, Nat. Prod. Rep., 2025 10.1039/d5np00034c.
- I. M. Novitskiy and A. G. Kutateladze, J. Org. Chem., 2022, 87, 4818–4828, DOI:10.1021/acs.joc.2c00169.
- A. Howarth, K. Ermanis and J. M. Goodman, Chem. Sci., 2020, 11, 4351–4359, 10.1039/d0sc00442a.
- G. D. Hannigan, D. Prihoda, A. Palicka, J. Soukup, O. Klempir, L. Rampula, J. Durcak, M. Wurst, J. Kotowski, D. Chang, R. Wang, G. Piizzi, G. Temesi, D. J. Hazuda, C. H. Woelk and D. A. Bitton, Nucleic Acids Res., 2019, 47, e110, DOI:10.1093/nar/gkz654.
- Y. Y. Lee, M. Guler, D. N. Chigumba, S. Wang, N. Mittal, C. Miller, B. Krummenacher, H. Liu, L. Cao, A. Kannan, K. Narayan, S. T. Slocum, B. L. Roth, A. Gurevich, B. Behsaz, R. D. Kersten and H. Mohimani, Nat. Commun., 2023, 14, 4219, DOI:10.1038/s41467-023-39905-4.
- N. J. van Eck and L. Waltman, Scientometrics, 2010, 84, 523–538, DOI:10.1007/s11192-009-0146-3.
- A. R. Carroll, B. R. Copp, R. A. Davis, R. A. Keyzers and M. R. Prinsep, Nat. Prod. Rep., 2022, 39, 1122–1171, 10.1039/d1np00076d.
- R. Reher, H. W. Kim, C. Zhang, H. H. Mao, M. Wang, L. F. Nothias, A. M. Caraballo-Rodriguez, E. Glukhov, B. Teke, T. Leao, K. L. Alexander, B. M. Duggan, E. L. Van Everbroeck, P. C. Dorrestein, G. W. Cottrell and W. H. Gerwick, J. Am. Chem. Soc., 2020, 142, 4114–4120, DOI:10.1021/jacs.9b13786.
- H. W. Kim, C. Zhang, R. Reher, M. Wang, K. L. Alexander, L. F. Nothias, Y. K. Han, H. Shin, K. Y. Lee, K. H. Lee, M. J. Kim, P. C. Dorrestein, W. H. Gerwick and G. W. Cottrell, J. Cheminf., 2023, 15, 71, DOI:10.1186/s13321-023-00738-4.
- H. W. Kim, M. Wang, C. A. Leber, L. F. Nothias, R. Reher, K. B. Kang, J. J. J. van der Hooft, P. C. Dorrestein, W. H. Gerwick and G. W. Cottrell, J. Nat. Prod., 2021, 84, 2795–2807, DOI:10.1021/acs.jnatprod.1c00399.
- J. R. Pardos-Blas, M. J. Tenorio, J. C. G. Galindo and R. Zardoya, Mar. Drugs, 2022, 20, 149, DOI:10.3390/md20020149.
- C. R. Pye, M. J. Bertin, R. S. Lokey, W. H. Gerwick and R. G. Linington, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 5601–5606, DOI:10.1073/pnas.1614680114.
- C. R. McClain, T. J. Webb, N. A. Heim, M. L. Knope, P. M. Monarrez and J. L. Payne, Oikos, 2025, e10828, DOI:10.1111/oik.10828.
- P. G. Albano, B. Sabelli and P. Bouchet, Biodivers. Conserv., 2011, 20, 3223–3237, DOI:10.1007/s10531-011-0117-x.
- D. Levine, M. Shuaibi, E. C. Spotte-Smith, M. G. Taylor, M. R. Hasyim, K. Michel, I. Batatia, G. Csányi, M. Dzamba, P. Eastman, N. C. Frey, X. Fu, V. Gharakhanyan, A. S. Krishnapriyan, J. A. Rackers, S. Raja, A. Rizvi, A. S. Rosen, Z. Ulissi, S. Vargas, C. L. Zitnick, S. M. Blau and B. M. Wood, arXiv, 2025, preprint, arXiv:2505.08762, DOI:10.48550/arXiv.2505.08762.
- X. Chang, Adv. Econ. Manag. Political Sci., 2023, 23, 134–140, DOI:10.54254/2754-1169/23/20230367.
- M. Wang, J. J. Carver, V. V. Phelan, L. M. Sanchez, N. Garg, Y. Peng, D. D. Nguyen, J. Watrous, C. A. Kapono, T. Luzzatto-Knaan, C. Porto, A. Bouslimani, A. V. Melnik, M. J. Meehan, W. T. Liu, M. Crusemann, P. D. Boudreau, E. Esquenazi, M. Sandoval-Calderon, R. D. Kersten, L. A. Pace, R. A. Quinn, K. R. Duncan, C. C. Hsu, D. J. Floros, R. G. Gavilan, K. Kleigrewe, T. Northen, R. J. Dutton, D. Parrot, E. E. Carlson, B. Aigle, C. F. Michelsen, L. Jelsbak, C. Sohlenkamp, P. Pevzner, A. Edlund, J. McLean, J. Piel, B. T. Murphy, L. Gerwick, C. C. Liaw, Y. L. Yang, H. U. Humpf, M. Maansson, R. A. Keyzers, A. C. Sims, A. R. Johnson, A. M. Sidebottom, B. E. Sedio, A. Klitgaard, C. B. Larson, C. A. B. P, D. Torres-Mendoza, D. J. Gonzalez, D. B. Silva, L. M. Marques, D. P. Demarque, E. Pociute, E. C. O'Neill, E. Briand, E. J. N. Helfrich, E. A. Granatosky, E. Glukhov, F. Ryffel, H. Houson, H. Mohimani, J. J. Kharbush, Y. Zeng, J. A. Vorholt, K. L. Kurita, P. Charusanti, K. L. McPhail, K. F. Nielsen, L. Vuong, M. Elfeki, M. F. Traxler, N. Engene, N. Koyama, O. B. Vining, R. Baric, R. R. Silva, S. J. Mascuch, S. Tomasi, S. Jenkins, V. Macherla, T. Hoffman, V. Agarwal, P. G. Williams, J. Dai, R. Neupane, J. Gurr, A. M. C. Rodriguez, A. Lamsa, C. Zhang, K. Dorrestein, B. M. Duggan, J. Almaliti, P. M. Allard, P. Phapale, L. F. Nothias, T. Alexandrov, M. Litaudon, J. L. Wolfender, J. E. Kyle, T. O. Metz, T. Peryea, D. T. Nguyen, D. VanLeer, P. Shinn, A. Jadhav, R. Muller, K. M. Waters, W. Shi, X. Liu, L. Zhang, R. Knight, P. R. Jensen, B. O. Palsson, K. Pogliano, R. G. Linington, M. Gutierrez, N. P. Lopes, W. H. Gerwick, B. S. Moore, P. C. Dorrestein and N. Bandeira, Nat. Biotechnol., 2016, 34, 828–837, DOI:10.1038/nbt.3597.
- R. Schmid, S. Heuckeroth, A. Korf, A. Smirnov, O. Myers, T. S. Dyrlund, R. Bushuiev, K. J. Murray, N. Hoffmann, M. Lu, A. Sarvepalli, Z. Zhang, M. Fleischauer, K. Duhrkop, M. Wesner, S. J. Hoogstra, E. Rudt, O. Mokshyna, C. Brungs, K. Ponomarov, L. Mutabdzija, T. Damiani, C. J. Pudney, M. Earll, P. O. Helmer, T. R. Fallon, T. Schulze, A. Rivas-Ubach, A. Bilbao, H. Richter, L. F. Nothias, M. Wang, M. Oresic, J. K. Weng, S. Bocker, A. Jeibmann, H. Hayen, U. Karst, P. C. Dorrestein, D. Petras, X. Du and T. Pluskal, Nat. Biotechnol., 2023, 41, 447–449, DOI:10.1038/s41587-023-01690-2.
- WoRMS Editorial Board, World Register of Marine Species (WoRMS), https://www.marinespecies.org, accessed 2025-11-06, DOI:10.14284/170.
- Intergovernmental Oceanographic Commission of UNESCO, Ocean Biodiversity Information System (OBIS), obis.org, accessed 2025-11-06.
- iNaturalist, https://www.inaturalist.org, accessed 2025-11-06.
- A. Rutz, M. Sorokina, J. Galgonek, D. Mietchen, E. Willighagen, A. Gaudry, J. G. Graham, R. Stephan, R. Page, J. Vondrasek, C. Steinbeck, G. F. Pauli, J. L. Wolfender, J. Bisson and P. M. Allard, eLife, 2022, 11, e70780, DOI:10.7554/eLife.70780.
- A. Rutz, D. Probst, C. Aguilar, D. Y. Akiyama, F. Alberti, H. E. Augustijn, N. E. Avalon, C. Beemelmanns, H. B. Barbieri, F. Biermann, A. J. Bridge, E. C. Giron, R. Cox, M. Crusemann, P. M. D'Agostino, M. Feuermann, J. Gerke, K. G. Garcia, J. E. Holme, J. Y. Hwang, R. Iacovelli, J. C. Jeronimo Barbosa, N. Kaur, M. Klapper, A. M. Kohler, A. Korenskaia, N. Kubach, B. T. Lee, C. Loureiro, S. Mantri, S. Narula, D. Meijer, J. C. Navarro-Munoz, G. S. Nguyen, S. Paliyal, M. Panghal, L. Rao, S. Sieber, N. Sokolova, S. T. Sowa, J. Szenei, B. R. Terlouw, H. G. Weddeling, J. Yu, N. Ziemert, T. Weber, K. Blin, J. J. J. van der Hooft, M. H. Medema and M. M. Zdouc, Nucleic Acids Res., 2025, gkaf969, DOI:10.1093/nar/gkaf969.
| This journal is © The Royal Society of Chemistry 2026 |