Ryanodane diterpenes: occurrence, structural diversity, bioactivities, and synthesis
Bodou
Zhang
ab,
Jingwen
Zhao
c,
Sheng
Li
a,
Hong
Liang
a,
Xiaojiang
Hao
 a and
Yu
Zhang
a and
Yu
Zhang
 *a
*a
aState Key Laboratory of Phytochemistry and Natural Medicines, Kunming Institute of Botany, Chinese Academy of Sciences, Kunming, 650201, China. E-mail: zhangyu@mail.kib.ac.cn
bUniversity of Chinese Academy of Sciences, Beijing, 100049, China
cSchool of Traditional Chinese Materia Medica, Shenyang Pharmaceutical University, Shenyang, 110016, China
First published on 1st October 2025
Abstract
Covering: 1948 to 2025
Ryanodane diterpenes (RDs) are a unique class of plant-derived natural products characterized by their complex, polyoxygenated pentacyclic frameworks. They have been primarily identified in plants from the Salicaceae and Lauraceae families. In recent years, RDs have garnered significant interest due to their notable bioactivities, particularly their modulation of ryanodine receptors (RyRs) and their insecticidal properties. Since the initial isolation of ryanodine from the shrub Ryania speciosa Vahl in 1948, a total of 135 natural RDs across nine subtypes have been discovered. These compounds exhibit a range of biological activities, including insecticidal, cardiac activity, and immunomodulatory effects. However, the limited natural abundance of RDs has posed challenges for their comprehensive biological evaluation. Fascinated by their high affinity for RyRs and their intricate polycyclic structures, synthetic chemists have pursued the total synthesis of RDs since the 1990s, with notable progress in recent decades. Advances in synthetic methodology have enabled the successful construction of key RD scaffolds, facilitating further exploration of their biological potential. This review provides a comprehensive overview of RDs from 1948 to May of 2025, highlighting their significance in drug discovery and development. It also emphasizes the need for interdisciplinary collaboration to fully harness the therapeutic potential of these complex natural products.
1 Introduction
Diterpenes, typically composed of four isoprene units (C5H8), constitute a significant class of natural products characterized by remarkable structural diversity and potent biological activities.1,2 Plant-derived diterpenes, in particular, have garnered substantial interest in drug discovery due to their distinguished pharmacological profiles.3,4 Among them, ryanodane diterpenes (RDs) stand out as a structurally unique and biologically significant subclass. Structurally, RDs are characterized by a polycyclic cage-like framework bridged by a hemiketal unit, highly oxidized functional groups, and multiple stereocenters. This intricate architecture renders RDs a compelling yet underexplored area within natural product chemistry. Ryanodine, one of the most well-known RDs, was first isolated from Ryania speciosa Vahl in 1948.5 It exhibits nanomolar affinity for its namesake receptors, the ryanodine receptors (RyRs),6 which are intracellular Ca2+ release channels located on the sarcoplasmic/endoplasmic reticulum (SR/ER) membranes. These channels play critical roles in cellular signal transduction processes in neurons, skeletal and cardiac muscles, and immune cells.7,8 As one of the few natural small-molecule modulators of RyRs, ryanodine has served as an important pharmacological tool for elucidating the mechanisms of intracellular Ca2+ release, particularly in the context of excitation–contraction coupling in skeletal and cardiac muscles.9RDs gained widespread recognition in the 1990s for their potent antifeedant and insecticidal activities.6,10 The insect RyR has since been validated as a target for commercial insecticides, as exemplified by the development of phthalic acid diamides and anthranilic diamides.11 Over the past three decades, significant advances in structural biology, particularly through cryo-electron microscopy (cryo-EM), have enabled the determination of RyR structures at atomic resolution. These studies have provided critical insights into the molecular mechanisms underlying RyR dysfunction in various human diseases.12–15 Notably, stress-induced RyR-mediated Ca2+ leak has been implicated in the pathogenesis of heart failure, cardiac arrhythmias, malignant hyperthermia, age-related muscle loss, and cognitive dysfunction.16 To date, several FDA approved drugs targeting RyR2 have been identified, some of which are currently used clinically or are under investigation in clinical trials for the treatment of arrhythmogenic heart diseases.17 In 2016, a landmark cryo-EM study mapped the binding sites of key RyR modulators, including ATP, Ca2+, caffeine, and ryanodine.18 Ryanodine was found to stabilize the channel in an open conformation by binding within the pore near residue Q4933, thereby reducing Ca2+ conductance to a sub-conductance state.18
Given the biological importance of RyRs, a detailed understanding of the structural basis for RD binding to RyRs is highly valuable for rational drug design. However, the limited natural abundance of RDs has hindered their biological exploration, necessitating the investigation of chemical diversity and synthetic approaches to enable comprehensive structural and pharmacological studies. Since the 1980s, 135 RDs have been identified exclusively from plant sources, yet their structural diversity and distribution across different species remain largely unexplored. Over 80% of the reported RDs share a common molecular framework, differing mainly in the positions and orientations of oxygen-containing functional groups. These compounds have been primarily isolated from R. speciosa, Spigelia anthelmia, Cinnamomum cassia, and Persea indica. Harnessing this structural diversity offers unprecedented opportunities for the development of next-generation therapeutics targeting RyR-mediated pathologies, as well as for the design of eco-selective insecticides. Thus, RDs represent compelling candidates for interdisciplinary drug discovery efforts.
Beyond their classic insecticidal action via RyR modulation, recent studies have revealed promising additional pharmacological activities for certain RDs. These include potent immunomodulatory effects,19 anti-inflammatory properties,20 and antiviral activity.21 The intricate, highly oxidized polycyclic framework of RDs provides a robust scaffold for designing novel drug candidates targeting RyRs or other pathways. This review aims to consolidate current knowledge on RDs, covering their occurrence and distribution, structural characteristics and classification, diverse pharmacological activities and underlying mechanisms, and progress in chemical synthesis. By providing this integrated overview, we aim to establish a foundation for the rational development and utilization of RDs as pharmacological tools and potential therapeutic agents.
2 Occurrence and botanical distribution
RDs represent a rare group of naturally occurring plant secondary metabolites, found in nine plant species mainly from the Lauraceae and Salicaceae families. As shown in Fig. 1 and Table S1, a total of 135 natural and structurally diverse RDs from these species have been summarized in this review. Due to their rare and unique polycyclic frameworks restricted to these species, RDs hold significant potential as chemotaxonomic markers for elucidating evolutionary relationships in plant science.2.1 RDs from the Lauraceae family
Lauraceae, a large family within the order Laurales in Magnoliids, comprises approximately 2500–3000 species across 45 genera.22 Within this family, RDs are primarily concentrated in three species: C. cassia, C. zeylanicum, and P. indica. Notably, 65 RDs (48.1% of the total) have been identified from Lauraceae sources, including 52 from C. cassia, two from C. zeylanicum, and 22 from P. indica. The genus Cinnamomum, comprising approximately 250 species distributed across the tropical and subtropical regions of Asia, Australia, and the Pacific islands,23 holds significant economic importance due to its applications in food spices, traditional medicine, and the fragrance industry.24C. cassia, an evergreen tree of medium height, is widely cultivated in South China and Southeast Asia. It is renowned for its bark, which is used globally as a spice and in traditional medicine for treating rheumatoid arthritis, gastrointestinal disorders, and microbial infections.25,26 Phytochemical studies confirm that its bark and leaves are a major reservoir of RD diversity, accounting for 38.5% of all reported RDs. C. zeylanicum, native to Sri Lanka, Myanmar, and Southeast Asia, has bark that is a cornerstone of Ayurvedic medicine, prescribed for arthralgia, respiratory diseases, and gynecological conditions.27 Despite its medicinal prominence, it yields fewer characterized RDs (only two compounds), suggesting either a lower biosynthetic capacity or underexplored chemical diversity.As a member of the Lauraceae family, the genus Persea comprises approximately 200 species, primarily distributed in the subtropical and tropical regions of Asia and the American.28,29P. indica, a perennial tree and one of the dominant species in the Canarian laurel forest of the Canary islands, likely originated from tropical American lineages.30 This plant exhibits adaptations to humid, thermally stable environments.31 Its stems and leaves contain high levels of RDs, which sever as defense compounds and are toxic to both insects and mammals.10,32 These RDs demonstrate potent broad-spectrum insecticidal activity against Spodoptera litura, Heliothis armigera, and other pests.33
The high concentration of RDs in the genera Cinnamomum and Persea underscores their role as evolutionary hotspots for RDs biosynthesis. This distribution pattern aligns with the proposed function of diterpenes in defending against herbivory and pathogen stress in tropical ecosystems,34 where Lauraceae species face intense biotic pressure. Future phytochemical surveys of understudied genera within Lauraceae may reveal additional RD structural diversity and further support chemotaxonomic hypotheses.
2.2 RDs from the Salicaceae family
Complementing the Lauraceae sources, the Salicaceae family contributes 46 documented RDs, accounting for 34.1% of all known RDs. Salicaceae comprises dioecious woody trees and shrubs primarily distributed in the Northern Hemisphere. This family exhibits close molecular affinities to the former Flacourtiaceae (now subsumed within Salicaceae under the APG IV system),35,36 highlighting evolutionary drivers for specialized metabolite diversification. Species-specific distribution reveals R. speciosa as the dominant reported source, yielding 38 RDs (82.6% of Salicaceae-derived RDs), followed by Itoa orientalis with seven RDs (15.2%), and Populus davidiana with one RD (2.2%). The genus Ryania comprises shrubs or slender trees distributed across various tropical regions of Central and South America, including Trinidad, and the Amazon basin.6 Its most notable species, R. speciosa, has been utilized indigenous communities for centuries. The stem and roots of R. speciosa, which contain toxic alkaloidal RDs, were applied to arrowheads for hunting and used as insecticides against cockroaches.6 Since the 1940s, extracts of R. speciosa have been commercially employed to control citrus thrips on citrus crops, codling moth (Cydia pomonella) on apples and pears, and European corn borer (Ostrinia nubilalis) on corn.37I. orientalis, once classified under the now-defunct Flacourtiaceae family, is a small tree native to southern and southwest China. Its barks and roots have been utilized in traditional Chinese medicine to treat conditions such as rheumatism, injuries from falls, hepatitis, and anemia.38 Additionally, its seeds have been employed to control the black cutworm, Agrotis ipsilon (Rott).38 This pesticidal use strongly suggest the involvement of RDs, although only seven RDs have been isolated from this species to date. I. orientalis represents an underexplored resource for novel RD scaffolds with potential agrochemical applications. The single RD identified from P. davidiana highlights an unexpected phylogenetic breadth in RD distribution.39 This temperate species differs ecologically from the tropical genera Ryania and Itoa, suggesting possible convergent evolution of RD biosynthesis or horizontal gene transfer events. Although only one RD has been reported in P. davidiana, it remains unclear whether RDs are widely distributed throughout the genus Populus. Further systematic phytochemical investigations are therefore needed to determine the prevalence of RDs in different Populus species and to clarify both their taxonomic significance and their potential medicinal value.
2.3 RDs from other families
Beyond Lauraceae and Salicaceae, an additional 27 RDs have been identified from three phylogenetically distinct families: Loganiaceae (22 RDs from S. anthelmia), Erythroxylaceae (two RDs from Erythroxylum passerinum), and Lecythidaceae (three RDs from Barringtonia macrocarpa). S. anthelmia, also known as arapabaca or erva-lombrigueira (meaning “anthelmintic herb”), is native to Asia and tropical America. It is widely used as an anthelmintic in the folk medicine of Brazil, Panama, and the Virgin Islands.40,41 In the German Homeopathic Pharmacopoeia, an extract from the aerial parts of S. anthelmia is recognized as an official treatment for neuralgic and cardiac disorders,42 suggesting that the extract may possess RyR-modulating activity, likely associated with its high RD content. The genus Erythroxylum, comprising approximately 97% of the species in the Erythroxylaceae family, is predominantly found in tropical and subtropical regions.43E. passerinum, commonly known as “Bomnome”, is a shrub or small tree native to humid forests, dry forests, and scrublands of Colombia, Venezuela, Peru, and Brazil.44B. macrocarpa, an evergreen tree or shrub native to Southeast Asia,45 has been reported to contain three RDs.46 These plants represent an underexplored botanical source of RDs, with their compounds exhibiting structural novelty.Notably, ryanodine is a shared metabolite across three families and three species: P. indica, R. speciosa, and S. anthelmia. This tri-taxonomic convergence may reflect adaptive gene recruitment events in response to analogous herbivore pressures in tropical forests.47 The high RD content in these plants is strongly associated with their insecticidal activities, reflecting an evolved plant defense mechanism against insect herbivores. Despite their functional significance, RDs exhibit extreme phylogenetic restriction, with only nine species across five families confirmed producers. This scarcity contrasts sharply with more ubiquitous diterpenes (e.g., labdanes found in > 200 genera) and positions RDs as high-priority chemotaxonomic markers. Conservation assessments indicate that four RD-producing species (S. anthelmia, E. passerinum, B. macrocarpa, and R. speciosa) are facing habitat loss, emphasizing the urgent need for in situ protection to preserve these “living libraries” of RyR-targeting scaffolds.
3 Structural diversity
3.1 Structural classification
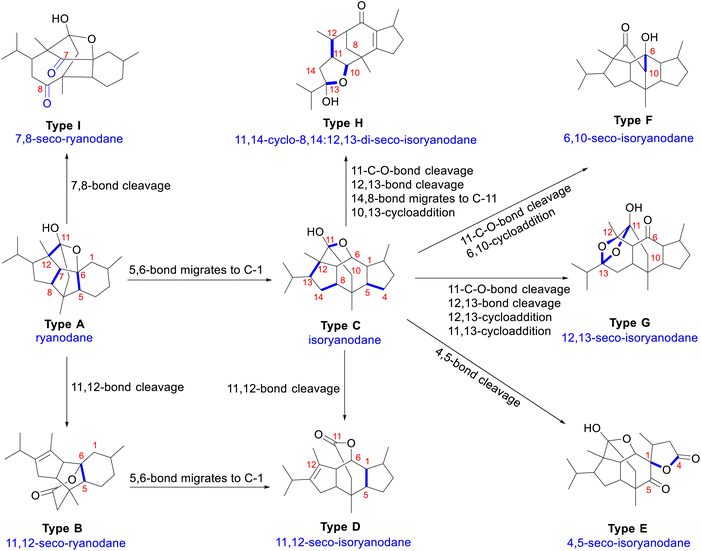
Although 135 naturally occurring RDs across different structural subtypes have been reported, a clear and rational structural classification remains elusive. Current categorization primarily employs three orthogonal criteria: (1) core scaffold topology (ryanodane vs. isoryanodane); (2) incorporation of nitrogen atoms (alkaloidal vs. nonalkaloidal); and (3) oxidation pattern (hemiketal, ketal, lactone, or diketone). Alkaloidal RDs (compounds 1–35, 65–70) dominate in R. speciosa and S. anthelmia, whereas non-alkaloidal types prevail in P. indica and C. cassia. Herein, we propose a biogenetically informed classification based on ring fusion patterns and plausible biosynthetic relationships, with plant distribution, structural details, and bioactivities cataloged in SI. For consistency, a unified numbering system for RDs is used throughout this review. Here, RDs are classified into nine subtypes across three structural types: ryanodane core, isoryanodane core, and others (Fig. 2).The ryanodane core typically features a 6/5/5 ring system (A–B–C rings) with contiguous stereocenters and multiple hydroxyl substitution, and includes two structural subtypes: ryanodane (type A) and 11,12-seco-ryanodane (type B). Type A is characterized by a C-6/C-10/C-12 hemiketal bridge, a rare structural motif in plant-derived diterpenes. This hemiketal confers pH-sensitive lability and can undergo acid-catalyzed of the 11,12-bond to form Type B. Type B contains a C-6/C-11 δ-lactone resulting from hemiketal rearrangement. Both subtypes exhibit high oxidation states and are prone to oxidation at C-1, C-5, C-7, and C-19, often bearing hydroxyl groups at these positions. In the presence of biosynthetic enzymes in plants, the 5,6 bond in the ryanodane core could readily migrate to the C-1 position, forming the isoryanodane core (Scheme 1). The isoryanodane core is structurally characterized by a 5/6/5 A–B–C tricyclic core and is divided into four subtypes: isoryanodane (type C), 11,12-seco-isoryanodane (types D),23 4,5-seco-isoryanodane (type E),48 and 6,11-seco-isoryanodane (type F).48 Types C and D are structurally analogous to types A and B, distinguished by a six-membered ring containing a hemiketal (type C) or a six-membered lactone ring (type D), respectively. Type C serves as a key precursor to a variety of structural scaffolds that contribute to the diversity of RDs. Type E arises via oxidative cleavage of the 4,5-carbon, generating a C-5 ketal functionality and a 1,4-lactone moiety through enzyme-mediated reactions.48 Additionally, type C undergoes ester hydrolysis between C-11 and C-6, followed by formation of a new carbon bond between C-6 and C-10, resulting in the formation of type F.48 The scaffold diversity outlined above not only underpins the broad bioactivity of RDs but also presents significant synthetic challenges.
Complex rearranged scaffolds give rise to other types of RDs, including 12,13-seco-isoryanodane (type G),19,49 11,14-cyclo-8,14:12,13-di-seco-isoryanodane (type H),50 and 7,8-seco-isoryanodane (type I).51 Type G features a unique 11,13:12,13-diepoxy-6,11-epoxy:12,13-disecoisoryanodane bearing hemiketal functionalities.19,49 Structurally, the C ring in type G is reformed into a distinctive furan ring via cleavage of the 6,11-ether linkage, 12,13-bond cleavage, and recyclization through intramolecular nucleophilic addition of the C-13 hydroxyl group to the C-11 carbonyl moiety. Additionally, when the C-6 hydroxyl group is oxidized to a ketone, a carbon–carbon bond between C-6 and C-10 can be formed via an aldol reaction catalyzed by an alkaline enzyme.49 Type H possesses a rare tetracyclic 11,14-cyclo-8,14:12,13-di-seco-isoryanodane diterpene carbon skeleton, derived from type C through 11,14-carbon–carbon bond formation and 8,14:12,13-carbon–carbon bond cleavage, featuring a unique pentadecane bridged system.50 Type I contains a rare structural skeleton derived from type A, first identified in 1981.51 It arises from oxidative cleavage of the C-11/C-6 bond in type A, forming a distinctive eight-membered ring with a diketone functional group. Detailed proposed biosynthetic pathways for types E–I will be presented in Sections 3.2.2 and 3.2.3.
3.2 Structures of RDs
3.2.1.1 Ryanodane (type A). Ryanodine (1) remains the most biologically significant type A RD since its isolation from R. speciosa in 1948.5,52,53 Its challenging structure was elucidated through classic chemical degradation studies in the 1970s.54,55 Compounds 2–22 are structurally related ryanodine derivatives isolated from the aerial parts of S. anthelmia, exhibiting significant cardiac activities (Fig. 3).42,53 Most RDs exhibit variable oxidation states and diverse substituent patterns across their scaffolds. For instance, 20-deoxyspiganthine (2) lacks the C-8 hydroxyl group present in ryanodine (1). Spiganthine (3), 8α-hydroxyspiganthine (4), and 20-hydroxyryanodine (6) display oxidation of the C-16 methyl group to hydroxymethyl moieties, while 20-norspiganthine-5-carboxylic acid (5) contains a carboxyl group. Compounds 7–10 are characterized by isomerization and oxidation at the C-1 and C-2 positions. Specifically, 9,18-dihydroxy-9-epi-10-epi-ryanodine (9) and 9,20-dihydroxy-9-epi-10-epi-ryanodine (10) undergo oxidation at C-19 and C-16, respectively, forming hydroxymethyl groups. The common structural feature of compounds 11–16 is the presence of unsaturation at the Δ2,3-position and additional hydroxyl groups within the A ring system. The structures of 17–20 feature an α-oriented three-membered epoxy ring, while epoxy-10-epi-ryanodine (21) exhibits a β-oriented epoxy ring in the A-ring. Notably, 6-deoxy-6β,9β-epoxy-8β-hydroxy-10-epi-ryanodine (22) undergoes protonation of the oxirane oxygen, facilitating a nucleophilic attack by the axial hydroxyl at C-2 on C-5, thereby forming two unique oxygen-bridged rings.
Compounds 23–46 are natural ryanodine derivatives reported from R. speciosa (Fig. 3 and 4).52,56–61 These compounds commonly undergo reductive cleavage and dehydrogenation at C-2/C-15 and C-2/C-3, introducing unsaturation at these sites. The C-3 position is particularly susceptible to oxidation, resulting in the formation of hydroxyl (27–30) or ketone functional groups (25). Furthermore, C-2, C-15, and C-19 are also prone to oxidation, as evidenced by the presence of hydroxyl groups at these positions. Ester G (31) is the first alkaloidal type RD found to possess a 3-formylpyridine moiety at the C-14 position,52 while 9a,21-epoxyryanodine (32) uniquely features a three-membered oxygen ring bridging C-2 and C-15.58 Additionally, the C-1 hydroxyl group is liable to undergo methylation or isomerization, giving rise to a variety of structural types. Notably, 10-O-pyrrolecarbonylryanodine (37) is the only ryanodine derivative characterized by two pyrrole-2-carboxylate ester groups at the C-1 and C-14 positions reported to date.59
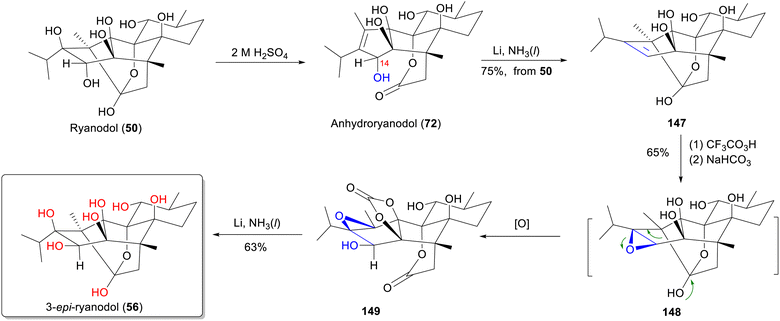
Compounds 47–64 constitute the remaining nonalkaloidal type A RDs, which lack the C-14 pyrrole-2-carboxylate group. Compounds 47–50 were isolated from R. speciosa58,61 while 51–64 were primarily obtained from C. cassia and P. indica (Fig. 5).62–65 Ryanodol (50)60 and 3-epi-ryanodol (56)66 are a pair of isomers differing only in the configuration of the C-14 hydroxyl group. Ryanodol (50) is produced by basic hydrolysis of ryanodine (1), involving elimination of the pyrrole-2-carboxylic acid moiety.67 As a key synthetic precursor to ryanodine (1), ryanodol (50) has been a prominent target for total synthesis in recent years. Notably, Inoue's group achieved the total synthesis of 3-epi-ryanodol (56) and used it to revise the reported structure of natural ryanodol (50) from P. indica.68 Consequently, the actual structure of epi-ryanodol 14-monoacetate (57),62 isolated from P. indica, is the C-14 acetylated derivative of 3-epi-ryanodol (56). Compounds 14-O-methyl-ryanodanol (58) and ryanodanol (59) differ from 56 primarily by the absence of the C-8 hydroxyl group.69 Garajonone (61) featuring an uncommon C-3 ketone carbonyl group,32 was recently synthesized by Zhao group.70
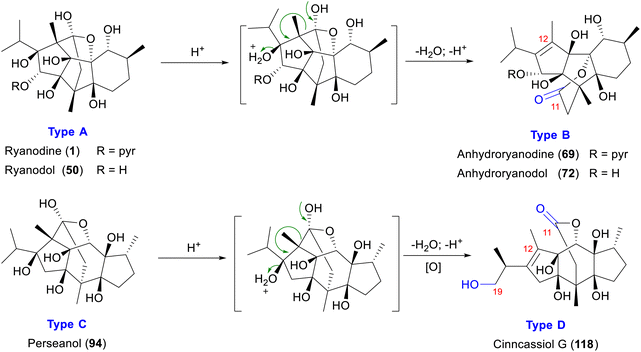
3.2.1.2 11,12-Seco-ryanodane (type B). Natural RDs isolated from R. speciosa are predominantly type A, with only a small amount of type B compounds (66–71) reported (Fig. 6).57,59 These type B compounds are acid hydrolysis products of type A, formed through chemical synthesis. In acidic media, ryanodine (1) and ryanodol (51) readily dehydrate to yield the degradation products anhydroryanodine (70) and anhydroryanodol (72), respectively,55 forming the 11,12-seco-ryanodane (type B) skeleton (Scheme 2). The loss of a H2O is accompanied by a significant modification of the molecular framework. Notably, compounds 67–71, which are alkaloidal RDs bearing a C-14 pyrrole-2-carboxylate ester group, have also been reported exclusively from R. speciosa.6 Compounds 73–79 are primarily isolated from P. indica and C. cassia,31,32,71 and are characterized by the absence of the C-14 hydroxyl group, a key structural difference between type B RDs from R. speciosa and C. cassia. Cinncassiol H (78) is the only known structure with an unsaturated double bond between C-12 and C-17.72 2,3-dehydroanhydrocinnzeylanine (79) undergoes reductive dehydrogenation at C-2 and C-3, resulting in unsaturated at these positions.73 Compounds 18S-cinncassiol A 19-O-β-D-glucopyranoside (81) and 18R-cinncassiol A 19-O-β-D-glucopyranoside (82) are a pair of epimers at the C-18,73 and feature glycosylation of the C-19 hydroxyl group through the addition of a D-glucose moiety, a common glycosylation pattern observed in RDs.
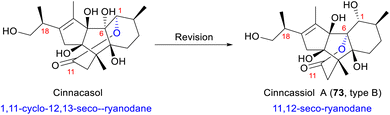
The complex oxidation patterns and continuous chiral centers in RDs often lead to structural misassignment. Cinnacasol, isolated from C. cassia in 2009, was initially assigned a 1,11-linked 12,13-seco-isoryanodane skeleton (Scheme 3).74 This misassignment likely stemmed from the misinterpretation of key HMBC correlations, particularly the incorrect attribution of connectivity between H-1 and C-11. Inoue and co-workers later revised the structure of cinnacasol to a 6,11-linked 12,13-seco-isoryanodane framework through the total synthesis of cinncassiol A (73) and density functional theory (DFT) calculations.68 Comparative analysis of synthetic cinncassiol A and natural cinnacasol revealed identical spectroscopic data, supporting the revised structure of cinnacasol as a highly strained seven-membered ring with an unfavorable C6–C11 anti conformation. DFT calculations further demonstrated that this conformation is 95.7 kcal mol−1 less stable than cinncassiol A (0 kcal mol−1), primarily due to the distorted tetrasubstituted carbon at C-11. Collectively, synthetic validation combined with computational stability analysis invalidated the original structural assignment. Consequently, the structure of cinnacasol was revised to match that of cinncassiol A, with 18S stereochemistry.68
3.2.2.1 Isoryanodane (type C). Type C readily undergoes skeletal cleavage and cyclization, leading to diverse rearranged skeletons and contributing to the structural diversity of RDs. Although fewer type C compounds have been reported compared to type A RDs, type C derivatives give rise to five distinct skeleton subtypes (types D, E, F, G, and H), highlighting their significant potential for structural exploration. Compounds 83–104 are exclusively isolated from P. indica and C. cassia (Fig. 7),10,75,76 serving as characteristic components and chemotaxonomic markers for these two species. Cinncassiol D1 (83) contains hydroxyl groups at C-2, C-5, and C-19;77 while its glucoside derivative (84) retains these hydroxyl groups and features additional glycosylation at C-19.77 Compounds 85–91 are predominantly characterized by hydroxyl groups at C-2 and C-5.75,77 Compounds 83–86 and 97–98 share hydroxylation at C-2 and C-19.76 Compounds 87–91 mainly feature oxidation at C-3, forming a β-configured hydroxyl or hydroxymethyl group. Compounds 94–96 undergo multiple oxidations at C-1, C-5, C-7, and C-8 on the basic isoryanodane core.19,76 Highly oxidized perseanol (94) serves as an important precursor for the biosynthesis of various skeletons.10,19 18-Hydroxyperseanol (95) features uncommon oxidation at C-18,19 while cinnacassin H (96) is oxidized at C-16,76 representing the highest oxidation levels observed in type C RDs. Cinnacassin E (97) and cinnacassin F (98) are a pair of epimers differing in configuration at C-18. Cinnacassin G (103) is a rare compound featuring an isomerized hydroxyl group at C-13, and cinnacassin I (104) has a rare double bond between C-13 and C-18.76
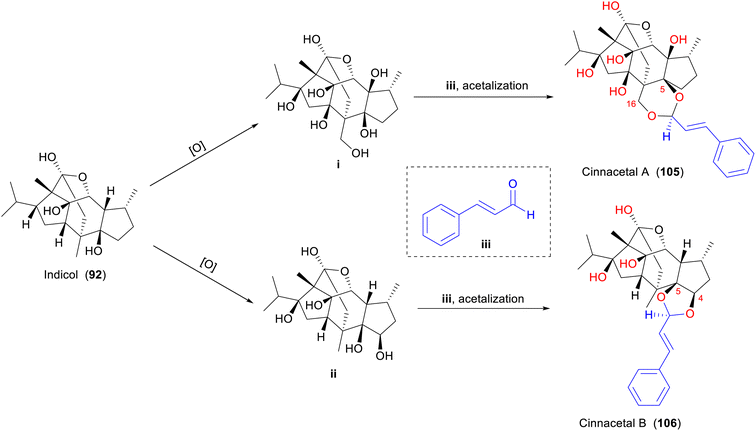
Indicol (92) may serve as a biosynthetic precursor for cinnacetal A (105) and cinnacetal B (106) in C. cassia.10,78 Compounds 105 and 106 are two unprecedented conjugates featuring a highly oxidated and modified isoryanodane core, with cinnamaldehyde-derived 1,3-dioxane and 1,3-dioxolane moieties, respectively. The hypothetical biosynthetic pathways were proposed by Huang, et al. in Scheme 4.78 Indicol (92) is readily produced through cascade oxidations by assembling intermediates i and ii with cinnamaldehyde (iii). Following the acetalization reactions at 5-OH and 16-OH in i and at 4-OH and 5-OH in ii, cinnacetal A (105) and cinnacetal B (106) are finally formed.
Itols A–D (107–110), along with the monoglucosides itol A 14-O-β-D-glucopyranoside (111) and itol B 20-O-β-D-glucopyranoside (112), were isolated from the bark and twigs of I. orientalis (Fig. 8),20 while 13-deoxyitol A (113) was isolated from the seeds of I. orientalis.79 These compounds exhibit a defining structural feature not observed in Lauraceae-derived type C RDs: a β-configured hydroxyl group at the C-14 position. Compounds 114–116 were isolated from B. macrocarpa. Barringisol (114) is characterized by the oxidation of a methyl group at the C-15 position to form a hydroxyl group, which is further glycosylated to form barringisosides A and B (115–116).46 Populusol A (117) is the sole RD isolated from the stem and bark of P. davidiana.39
3.2.2.2 11,12-Seco-ryanodane (type D). To date, type D RDs (118–124) have been exclusively isolated from C. cassia,19,23 establishing this species as the sole botanical source of this structural subclass. Analogous to the acid-catalyzed conversion of type A to type B RDs, perseanol (94) undergoes hydrolytic ring opening under acidic conditions to generate the corresponding 11,12-seco derivative (Scheme 2). Subsequent C-19 oxidation of 94 yields cinncassiol G (118), whereas sequential oxidation at C-16 and glycosylation affords 16-O-β-D-glucopyranosyl-19-deoxycinncassiol G (119). Compounds 120–122 share C-19 oxygenation, with cinnamomol C (120) and cinnamomol E (122) additionally featuring C-17 oxidation. Notably, cinnamomol F (123) exhibits a rare skeletal modification involving elimination of the C-13 isopropyl group, resulting in loss of a C3 unit and formation of a quaternary center, while its C-13 hydroxylated analogue undergoes O-glycosylation to form cinnamomol F glucoside (124) (Fig. 9).
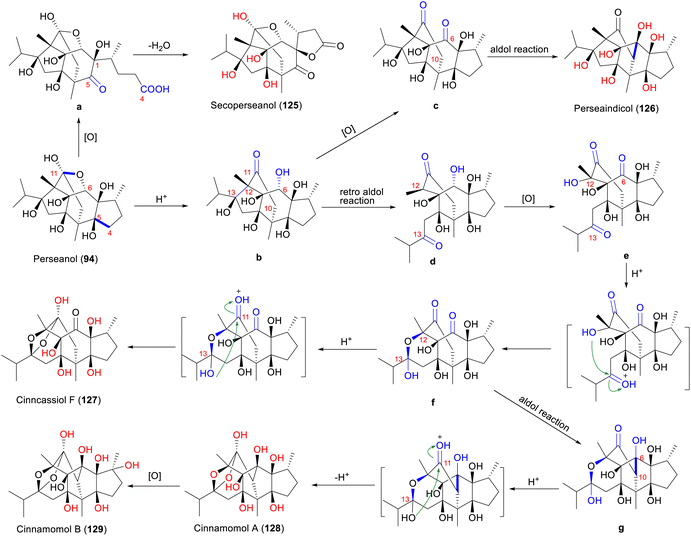
3.2.2.3 4,5-Seco-isoryanodane (type E) and 6,11-seco-isoryanodane (type F). Only secoperseanol (125) (type E) and perseaindicol (126) (type F) have each been isolated from P. indica.48 The biosynthetic hypothesis for types E and F, derived from perseanol (94, type C) with an isoryanodane skeleton, is proposed for the first time in this review (Scheme 5). Perseanol (94) first undergoes oxidative ring opening at the C-4 and C-5 positions to form a probable intermediate a. Subsequent intramolecular esterification occurs between the C-4 carboxyl group and the C-1 hydroxyl group, yielding 1,4-recyclized product secoperseanol (125). Conversely, perseanol (94) undergoes cleavage of the ether bond between C-11 and C-6, forming intermediate b under the catalysis of acid. Further oxidation of b leads to the di-ketone c at the C-6 and C-11 positions. Finally, an intramolecular aldol condensation of intermediate c affords perseaindicol (126).
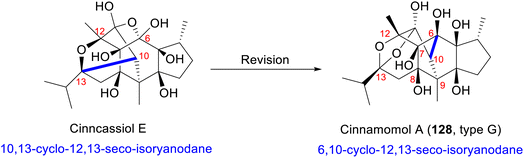
Cinncassiol E was first isolated from C. cassia in 1985 and was initially assigned as a 10,13-cyclo-12,13-seco-isoryanodane due to the misinterpretation of the chemical shifts of C-13 and C-6 (Scheme 6).80 Re-evaluation by Fraga et al.48 confirmed the correct 6,10-cyclo-12,13-seco-isoryanodane (type G) framework through critical HMBC correlations: H-10 showed unambiguous couplings to C-6, C-7, C-8, and C-9, while the downfield C-13 chemical shift (δC 107.5) was attributed to C-13 hemiketal group. Single-crystal X-ray diffraction ultimately resolved the ambiguity, proving that cinnamomol A (128) is structurally identical to cinncassiol E and establishing its absolute configuration.49
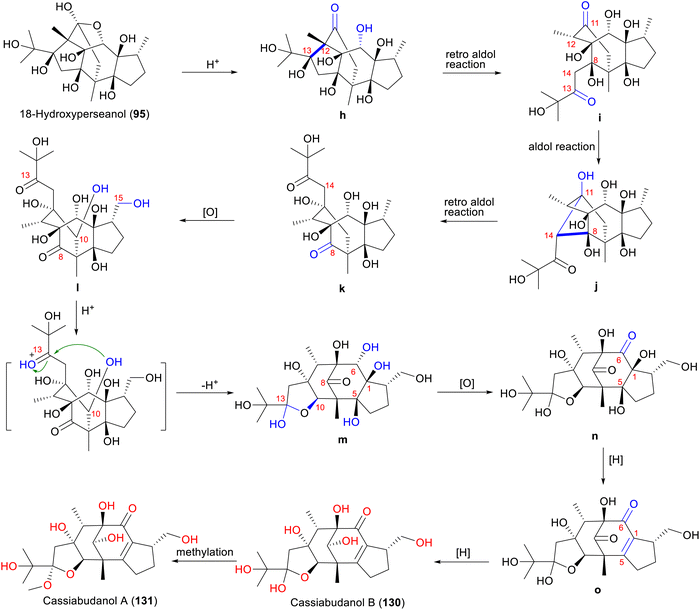
Type H compounds cassiabudanols B and A (130 and 131) are speculated to be biosynthesized from 18-hydroxyperseanol (95, type C) through a series of cascade reactions in C. cassia buds (Scheme 7).50 Similar to the formation of type G compounds, 18-hydroxyperseanol (95) undergoes cleavage of the ether linkage between C-6 and C-11 and a retro-aldol reaction between C-12 and C-13 to form ketone i. An aldol reaction in i constructs the bond between C-11 and C-14, generating j, which is the key step in forming the type H skeleton. Intermediate j undergoes a retro-aldol reaction and bond cleavage between C-14 and C-8, forming the diketone k, which undergoes a series of oxidation to produce the C-10 and C-15 hydroxylated l. Subsequently, the C-10 hydroxyl group of l nucleophilically attacks the C-13 ketone carbonyl group, re-forming the hemiketal m. Intermediate m undergoes a series of oxidation and reduction reactions and finally forms cassiabudanol B (130).49 Cassiabudanol A (131) is generated from 130 by enzyme-mediated methylation.49
Type I compounds 132–135 are derived from the oxidation and cleavage of the C-7 and C-8 positions of type A, resulting in a unique 7,8-diketone structure (Fig. 10).51 Precursor cinnzeylanol (52) undergoes oxidative cleavage at the C-7 and C-8 positions, forming cinncassiol C3 (135) with an eight-membered ring and a 7,8-diketone group. Compound 135 undergoes dehydration under acidic conditions to form cinncassiol C2 (134), which has an unsaturated bond between C-13 and C-14. Compound 134 is oxidized at the C-19 position to form cinncassiol C1 (132). Subsequent glycosylation of the C-19 hydroxyl group forms cinncassiol C1 (133).
4 Biological activities
In previous studies, RDs were renowned for their distinctive modulation of RyRs and potent insecticidal activity. Radiolabeled ryanodine ([3H] ryanodine) serves as a pivotal pharmacological tool for investigating RyR-mediated Ca2+ release dynamics in muscular and non-muscular cells.81,82 However, the toxicity of ryanodine (1), stemming from its marked effects on neural and muscle systems, can induce spasms, arrhythmias, or death in experimental animals at high doses,83,84 which limits therapeutic utility. In contrast, nonalkaloidal RDs exhibited better safety profiles and diverse bioactivities, such as insecticidal, immunomodulatory, and anti-inflammatory effects.4.1 RyRs modulating activity
Advancing insights into RyR2 dysfunction in cardiac muscle disorders have established RyR2 as a promising therapeutic target. To date, several FDA-approved drugs, including dantrolene, flecainide, propafenone, and carvedilol, have been found to inhibit RyR2.17 However, no RyR2-selective inhibitors are available for clinical use. In a recent study involving preclinical and phase I drug experiments, a novel RyR2 inhibitor, M201-A,96 was found to inhibit diastolic Ca2+ leak, improve cardiac lusitropic effects in rats, and enhance natriuresis and renal function in humans. Rycals are 1,4-benzothiazepine derivatives and structurally related to diltiazem but with higher specificity for RyR. They promote FKBP12.6 binding to RyR1 and RyR2,93,97 improving muscle function and reducing arrhythmias. In preclinical studies, rycal® ARM210 confirmed its safety and tolerability in treating RyR1-related myopathy in a Phase Ib trial, and showed first clinical efficacy by improving fatigue and proximal muscle strength in patients.98 Rycal restored muscle function by repairing the mutated “leaky” RyR1 channels. Moreover, the crystal structure of RyR2 bound to ARM210 was solved using cryo-EM and revealed the binding site of the drug, providing further insight into the mechanism of rycals in stabilizing leaky RyR2.87
The cryo-EM structure reveals that ryanodine binds directly within the transmembrane pore of RyR1, specifically at a site adjacent to residue Q4933 (Q4863 in RyR2) that lines the pore lumen.18 This binding site, identified through localized difference density maps between ryanodine-bound and ligand-free states, is located within the ion permeation pathway near the cytoplasmic gate (I4937). Ryanodine binding stabilizes the pore in a dilated conformation characteristic of the open state, thereby directly reducing Ca2+ permeation (Fig. 11). Due to its physical occupancy within the pore, a single ryanodine molecule partially obstructs ion flux, resulting in the characteristic sub-conductance state. At higher concentrations, ryanodine induces complete channel blockade. Current cryo-EM evidence suggests that multiple ryanodine molecules may bind within the pore lumen, with higher occupancy leading to full channel occlusion. The reduced binding affinity for secondary ryanodine ligands is attributable to steric hindrance imposed by the initially bound molecule.105 Mutational analysis confirmed the functional significance of Q4933, as its substitution drastically reduces ryanodine binding affinity without abolishing channel function,81 highlighting this residue's critical role in forming the high-affinity ryanodine-binding pocket conserved among RyRs. The structure of ryanodine-modified RyR1 provides insights into the mechanism of ryanodine binding and its modulation of RyR channel activity.
 | ||
| Fig. 11 Schematic representation of RyR1 activation and pore opening, which datasets in particular conditions: (A) Ca2+-free state, (B) Ca2+, (C) ryanodine-activated, (D) ryanodine-blocked. Figure adapted from ref. 18, with permission from Cell. | ||
 | ||
| Fig. 12 Structures of 1, 23, 50, and 3-epi-ryanodine (C-14 epimer), and a brief SAR diagram for RyRs binding potency. | ||
Critically, while the interaction of ryanodine with intracellular Ca2+ release channels is well investigated, little is known about the modes of action of other RDs as potential molecule targeting RyRs. Additionally, the minimally effective structure capable of high affinity binding to RyRs remains to be defined. Whether other RDs modulated channel function by virtue of their effects on Ca2+ release as (ant)agonists still need systematic investigation.
4.2 Insecticidal and antifeedant activity
The SAR of RDs indicates that substituents at both C-1 and C-14 substituents are crucial for their antifeedant and toxic activity against S. litura, S. littoralis, and L. decemlineata (Fig. 13).33 For S. littoralis, type A RDs with the C-1 β-stereochemistry and O-acetylation at this position show increased toxic and/or antifeedant activity. Hydroxylation, O-acetylation, and pyrrole-2-carboxylate-esterification at the C-14 position, along with the introduction of a C-1 ketone carbonyl group on the A ring, result in intermediate activities. Additionally, the presence of the C-2/C-3 epoxide enhances toxicity. For L. decemlineata, the presence of a C-1 ketone group and the acetylation or hydroxylation of C-1 or C-14 produce strong antifeedant and toxic effects. However, the C-14 pyrrole-2-carboxylate ester does not enhance these effects. Additionally, the C-2 hydroxylation and the presence of an epoxide (C-2/C-3 and C-2/C-5) enhance the antifeedant and postingestive effects, respectively.33
Type C RDs are recognized as potent botanical insecticides, exhibiting strong and selective toxicity towards insects compared to mammals.31 Among the type C RDs, indicol (92), vignaticol (93), and perseanol (94) demonstrate antifeedant effects against S. litura, with perseanol (94) showing the strongest effect (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) EC50 2.23 ppm) similar to ryanodol (50).10 The C-1 hydroxylation and the polarity of the C ring (C-8 and C-13 hydroxyl groups) are key determinants of their antifeedant activity against S. littoralis, L. decemlineata, and S. litura.10 Conversely, type B compounds anhydrocinnzeylanine (74) and anhydrocinnzeylanone (76) are less potent antifeedants and toxicants against S. littoralis and L. decemlineata than their type A counterparts, indicating that the C-11 hemiketal group is important for the antifeedant effects of RDs.32 These results suggest the potential for developing non-alkaloidal RDs as a new generation of target-oriented, natural botanical insecticides, though further research is needed to characterize their mechanisms of action and systematically assess their mammalian toxicity.
EC50 2.23 ppm) similar to ryanodol (50).10 The C-1 hydroxylation and the polarity of the C ring (C-8 and C-13 hydroxyl groups) are key determinants of their antifeedant activity against S. littoralis, L. decemlineata, and S. litura.10 Conversely, type B compounds anhydrocinnzeylanine (74) and anhydrocinnzeylanone (76) are less potent antifeedants and toxicants against S. littoralis and L. decemlineata than their type A counterparts, indicating that the C-11 hemiketal group is important for the antifeedant effects of RDs.32 These results suggest the potential for developing non-alkaloidal RDs as a new generation of target-oriented, natural botanical insecticides, though further research is needed to characterize their mechanisms of action and systematically assess their mammalian toxicity.
Unlike ryanodine, which exerts insecticidal effects by causing Ca2+ to flood into the cytoplasm via insect RyRs, itol A (type C RDs) does not act on RyR.129 Itol A adversely affects development by decreasing JH levels and disrupting the JH signaling pathway on Spodoptera frugiperda.130 It causes severe growth impairment, extends larval duration, and reduces mean larval weight and body length growth rates. Exposure to a sublethal concentration (500 mg L−1) significantly reduced larval JH levels by 36.59% and 22.70% at three and seven days post-treatment, respectively. qPCR analysis showed corresponding significant increases (6.58-fold and 2.12-fold) in the mRNA expression levels of key JH metabolism enzymes, S. frugiperda juvenile hormone esterase (sfJHE) and S. frugiperda juvenile hormone epoxide hydrolase (sfJHEH), at day three compared to controls.
Further research indicates that itol A significantly downregulates JHBP expression (Fig. 14).129 Under normal conditions, the JH-JHBP complex facilitates JH transport and protects it from metabolism.131 Itol A perturbs this JH transport pathway, likely through direct or indirect effects on JHBP and its signaling pathways. This dependence on JHBP was demonstrated by silencing the S. frugiperda JHBP gene (sfJHBP), which blocks its protein production, significantly reduces the efficacy of itol A. Furthermore, parallel reaction monitoring confirmed that itol A affects the expression abundance of JHBP. While JHBP may not be the sole or direct target of itol A, these findings indicate that itol A's insecticidal effect largely depends on JHBP and that it inhibits primarily the growth and development of S. frugiperda by disrupting JHBP function.129
 | ||
| Fig. 14 Hypothetical model for the mechanism of the associations between JH transport signaling pathways and itol A in S. frugiperda. Figure adapted from ref. 129, with permission from Journal of Agricultural and Food Chemistry. | ||
4.3 Cardiac activity
Spiganthine (3) significantly affected cardiac muscle by delaying contraction development.42 It produced a concentration-dependent delay in the contraction of guinea-pig heart ventricular papillary muscle, doubling the period between stimulation and the attainment of 10% of the force of contraction. The EC50 of spiganthine (3) is 56.7 ± 2.5 nM, which is slightly higher than that of ryanodine (1) at 16.5 ± 1.5 nM at the same contraction frequency.53 In addition, 12 natural ryanodine derivatives were systematically evaluated for cardiac activity and SAR (Fig. 15).53 Each of the tested compounds induced a delay in cardiac contraction development similar to spiganthine (3) and ryanodine (1), through with varying potencies. Ryanodine (1), 20-deoxyspiganthine (2), spiganthine (3), and 8,9-dehydro-10-epi-ryanodine (11) showed the most effective activities with EC50 14–54 nM. SAR studies revealed that the absence of a hydroxyl group at position C-8 or C-16, with the rest of the structure unchanged, caused only a slight reduction in the cardiac activity. Moving the hydroxyl from C-8 to C-16 did not significantly affect the activity. 8,9-dehydro-10-epi-ryanodine (11), which structurally differs by Δ2,3 unsaturation and C-1 epimerization, is similarly active as ryanodine (1). However, 1-epimerisation combined with 2,3-epoxidation significantly reduced cardiac activity to less than 10%. The presence of hydroxy groups at both C-8 and C-16 led to an even more pronounced reduction in activity. These results suggest that the polarity of natural ryanodine derivatives plays a crucial role in their cardiac activity.4.4 Immunomodulatory activity
Diterpenes are an important source of natural immunomodulators132,133 and play a crucial role in the prevention and treatment of autoimmune diseases, tumors, and chronic inflammatory diseases.134,135 Some RDs, isolated from C. cassia, exhibited dual immunomodulatory activities, encompassing both immunostimulation and immunosuppression.19 The ConA/LPS-induced splenocyte proliferation assay is a sensitive method for assessing the immunomodulatory effects of natural products.136 Specifically, cinncassiol A (73, type B) and cinncassiol G (118, type D) exhibit potent, dose-dependent suppression of ConA-induced T-cell proliferation, with 58.8% and 86.1% inhibition at 50 μM, respectively, which matches the positive control cyclosporine A without cytotoxicity.19 Conversely, type G RDs (cinnamomols A and B, 127 and 128) enhanced ConA-induced T-cell proliferation by 29–64% (0.391–100 μM), exceeding the potency of the clinical first-line immunostimulator thymosin α1.49 Type H compounds (cassiabudanols A and B, 131 and 130) augment T/B-cell responses at nanomolar-to-micromolar concentrations, mechanistically linked to CD4+/CD8+ T-cell upregulation and Treg downregulation.50 Type C RDs exhibit exceptional activity:76 cinncassiol D1 (83) enhances LPS-induced B-cell proliferation by 116.1% at 1.5625 μM (doubling thymosin α1 efficacy), and cinnacassin A (99) significantly enhances T-cell responses. Notably, cinnacassin F (98) displays concentration-dependent biphasic effects, inhibiting B-cell proliferation at low concentrations of 0.391–1.5625 μM, while stimulating it at higher doses of 6.25–100 μM.76Collectively, types C, G, and H RDs showed potent immunomodulatory effects. These findings reveal RDs from C. cassia as structurally diverse modulators with potentials for immune therapeutics, evidenced by in vitro potency surpassing clinical benchmarks in specific contexts and favorable cytotoxicity profiles. However, their translational promise remains provisional, necessitating rigorous mechanistic studies to define molecular targets (e.g., Treg modulation pathways and biphasic signaling switches), in vivo validation of efficacy/safety in disease models, and systematic SAR analyses to rationalize the design of RD-based immunomodulators. Future research should prioritize these gaps to objectively evaluate their applicability in treating immune dysregulation.
4.5 Anti-inflammatory activity
The development of effective anti-inflammatory therapeutics remains challenging due to the inherent complexity of inflammatory cascades. Microglia are immune cells in the central nervous system that, when activated, produce NO, a mediator that plays a key role in neuroinflammation. In LPS-stimulated BV-2 microglia, nonalkaloidal type A RDs cinnzeylanol (52) and cinnzeylanine (55), along with type B RDs cinncassiol A (73), anhydrocinnzeylanine (74), anhydrocinnzeylanol (75), epi-anhydrocinnzeylanol (77), and cinncassiol H (78), exhibited moderate inhibition of NO production, with IC50 values of 68.6–81.8 μM, indicating modest anti-neuroinflammatory potential.72Complementarily, type C RDs itols A–D (107–110), itol A 14-O-β-D-glucopyranoside (111), and itol B 20-O-β-D-glucopyranoside (112) demonstrated significant COX-2 inhibitory activity in LPS-induced A549 cells at 10 μM.20 Among these, itol D (110) showed the highest inhibitory rate (78.3%), followed by itol A (106, 61.0%), itol B 20-O-β-D-glucopyranoside (112, 57.0%), and itol A 14-O-β-D-glucopyranoside (111, 54.7%), collectively approaching the efficacy of the selective COX-2 inhibitor NS398 (85.3%). These findings suggest that type C RDs, particularly itols with aglycone structures like itol D, may offer targeted anti-inflammatory applications, warranting further investigation into their structure–activity relationships and in vivo efficacy for inflammation-associated pathologies.
4.6 Antiviral activity
Using native mass spectrometry to screen a 1614-compound natural product library against the SARS-CoV-2 nucleocapsid protein Nsp9, ryanodine (1) was identified as a binder with a protein binding ratio of 0.3 and a KD of 48.05 ± 5.03 μM, as determined through titration studies.137 This study represents the first report of ryanodine as a ligand for SARS-CoV-2 Nsp9, binding at a site distinct from the known inhibitor oridonin, as evidenced by the formation of a ternary complex (Nsp9:oridonin:ryanodine) in competition assays. Additionally, cinnzeylanine (55, type A) has demonstrated significant inhibition of herpes simplex virus type 1 (HSV-1) proliferation in Vero cells, which are derived from monkeys. The effect of various concentrations of 55 on HSV-1 proliferation was assessed using a vacuole reduction assay in the Vero cell line.21 Cinnzeylanine reduced the number of empty plaques formed by the virus, with an IC50 value of 230 μg mL−1 and achieving complete inhibition at 320 μg mL−1 without cytotoxicity. These findings highlight the potential of 1 and 55 as promising antiviral agents against SARS-CoV-2 and HSV-1, respectively.4.7 Other activities
Cinnzeylanol (52, type A) has been found to possess analgesic properties.138 In a paclitaxel-induced cold allodynia mouse model, 52 significantly reduced the number of responses to cold stimuli in mice when administered orally at a dose of 10 mg kg−1, demonstrating its analgesic effect against paclitaxel-induced cold allodynia, compared with the positive control drug gabapentin (30 mg kg−1).138 A study examined the cytotoxic effects of 14-O-methyl-ryanodanol (58, type A) isolated from E. passerinum on human astrocytic GL-15 cells.139 Results showed that 58 exerted dose-dependent cytotoxicity, with a significant reduction in cell viability at 500 μg mL−1 14-O-methyl-ryanodanol could induce apoptosis, causing nuclear changes and DNA fragmentation. Subtoxic exposure (250 μg mL−1) induced cytoplasmic contraction and projection expansion, indicating disruption of astrocytic cytoskeletal proteins including glial fibrillary acidic protein. These findings indicate that 58 targets astrocytic cells and may be linked to psychoactive symptoms observed after consumption of Erythroxylum fruits.Overall, these preliminary evaluations reveal the multifaceted bioactivities of RDs, including analgesic and cytotoxic effects, positioning them as structurally diverse candidates for therapeutic development. However, their translational potential requires rigorous validation. Future research must prioritize elucidating molecular mechanisms, establishing dose–response relationships across in vivo models, assessing in vivo pharmacokinetics and safety profiles, and confirming target specificity to advance these natural leads toward clinical or agricultural applications.
5 Synthesis of RDs
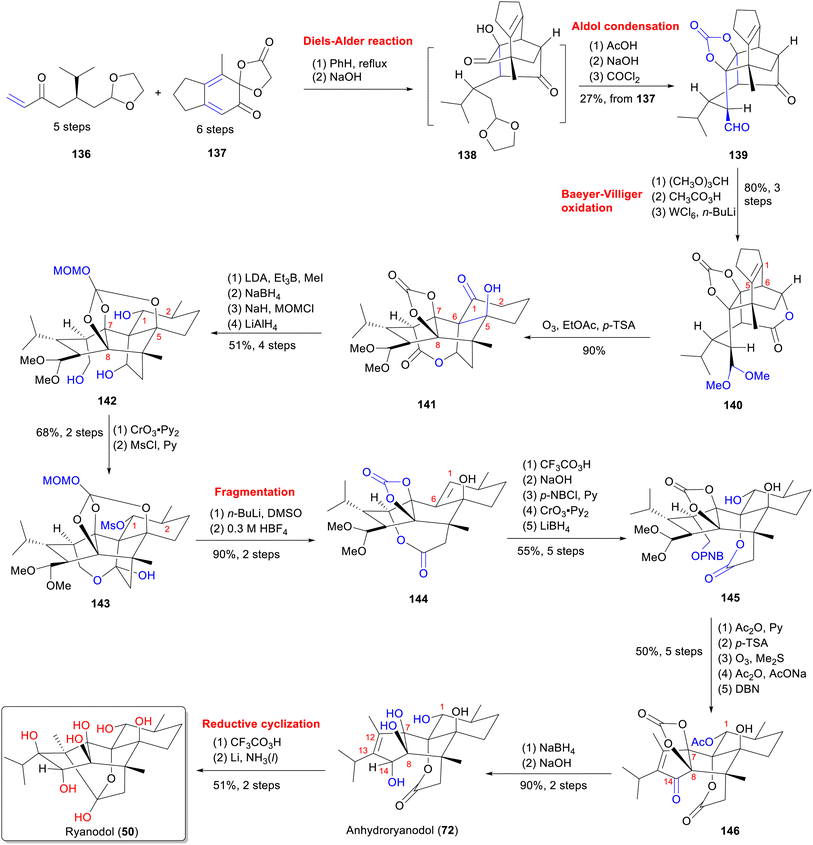
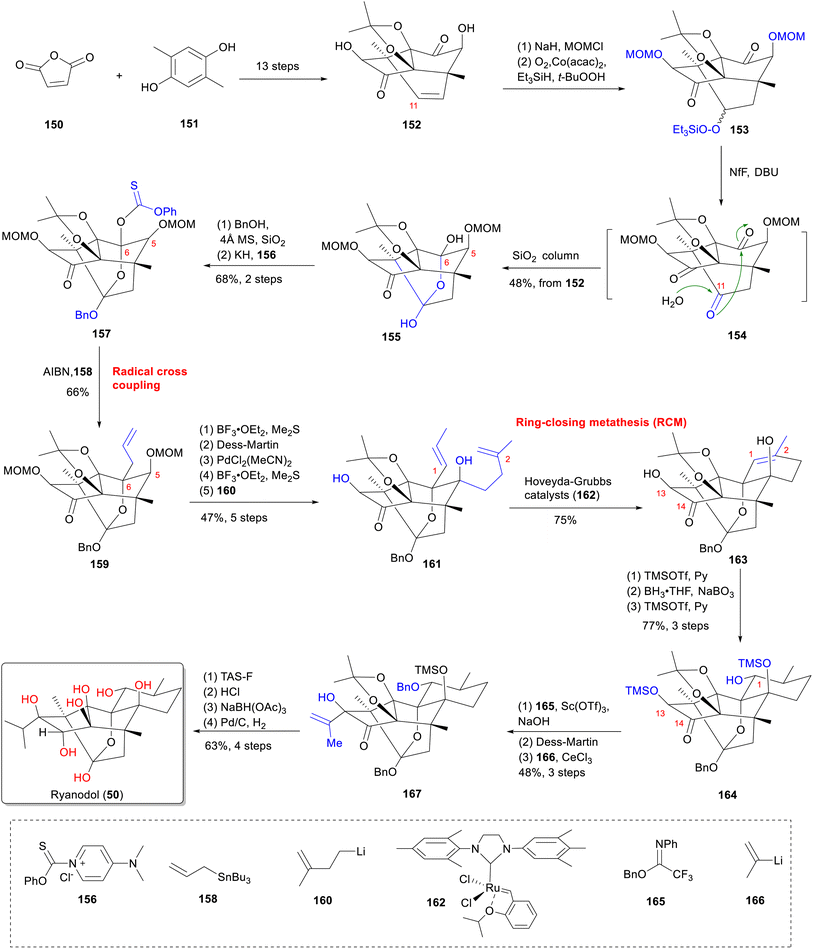
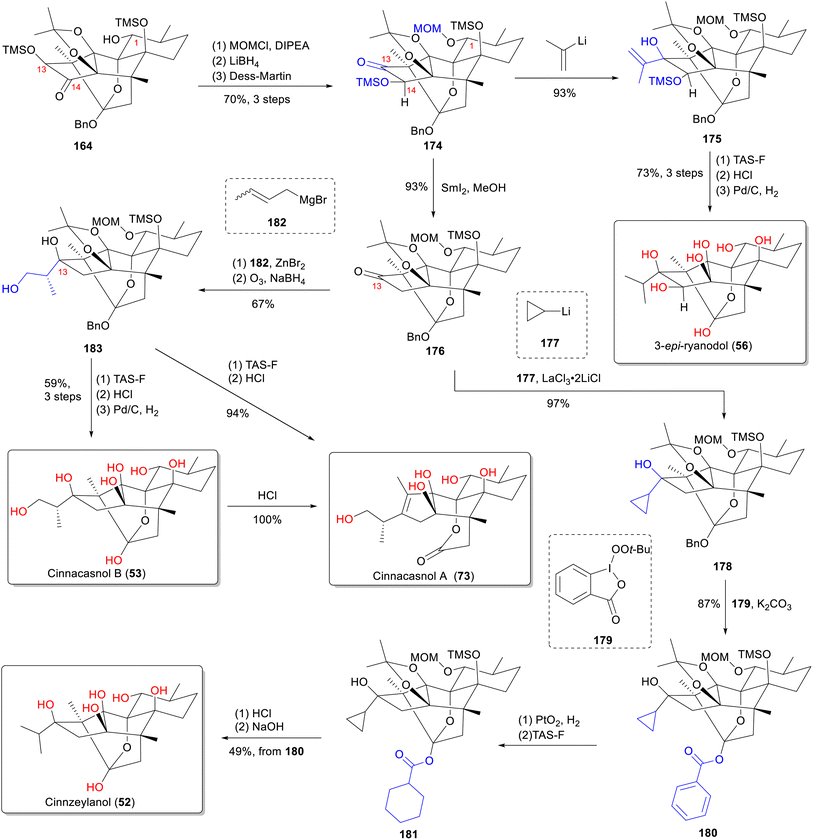
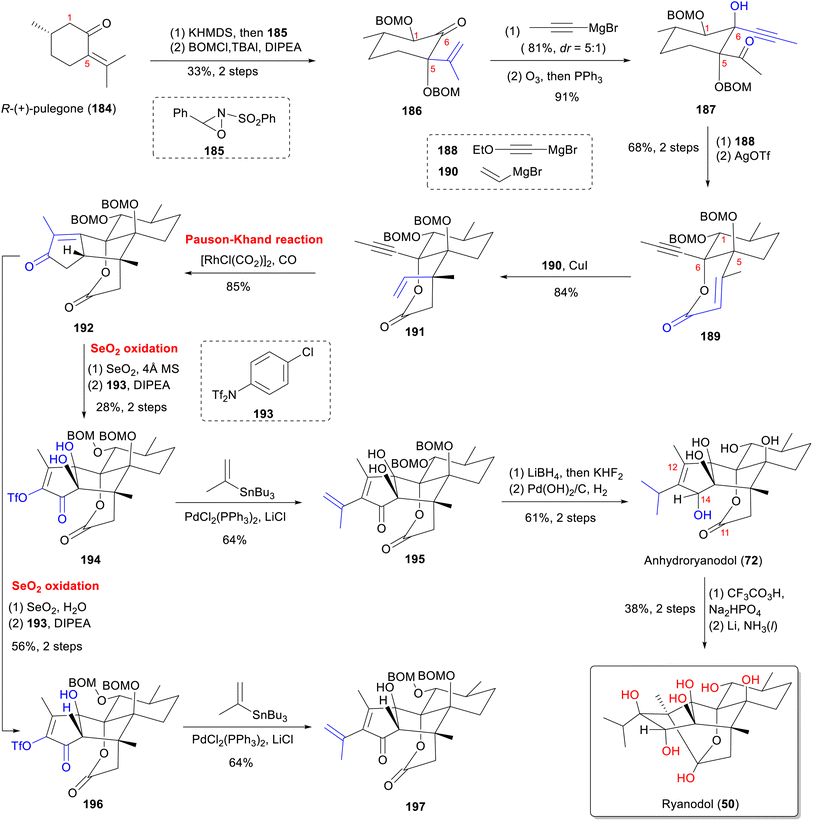
From a structural perspective, ryanodine (1) possesses a fundamental skeleton of 6/5/5/6/6 fused rings, 11 contiguous stereocenters (8 of which are quaternary carbon centers), and a labile hemiketal unit. This extreme oxidation state, marked by multiple C–O bonds, renders it one of the most complex known diterpenes. The structure poses formidable synthetic challenges, particularly for installing tetrasubstituted carbon centers within oxygen-dense frameworks.140 In this part, we summarize the total synthesis work of ten different RDs (belonging to types A–C) (Fig. 16) from five research groups, along with the key synthetic strategies and reactions according to the timeline (Table S3).Since Deslongchamps group pioneering synthesis of racemic anhydroryanodol (72) in a 41-steps linear sequence in 1979,141 later extended to ryanodol (50) and 3-epi-ryanodol (56), advances in ring-construction methodologies have accelerated RDs synthesis.142–144 This achievement was made possible by utilizing intermolecular Diels–Alder reactions and oxidative cleavage/transannular aldol cascades as the principal steps. Subsequently, Inoue's group achieved total syntheses of ryanodine (1), ryanodol (50), cinnzeylanol (52), cinncassiol B (53), 3-epi-ryanodol (56), and cinncassiol A (73) by applying creative strategies of radical-mediated C–O/C–N bond formations and ring-closing metathesis (RCM) reactions.68,145,146 A transformative breakthrough came from Reisman's 15-step enantioselective synthesis of (+)-ryanodol (50),147 ryanodine (1),148 and 20-deoxyspiganthine (2),148 which leveraged a strategic Pauson–Khand reaction149 to build the ABCDE rings and late-stage SeO2-mediated oxidation to install three oxygen atoms convergently, representing a drastic reduction in step count and a milestone in skeletal efficiency. Complementary approaches include Micalizio's synthesis of anhydroryanodol (72) by employing a key step involving titanium-mediated intramolecular stereoselective coupling of an alkyne and a 1,3-diketone.150 More recently, Zhao's synthesis routes to garajonone (61) using a two-stage synthetic strategy by employing an early-stage palladium-catalyzed Heck/carbonylative esterification cascade annulation, followed by successive late-stage selective redox manipulation to rapidly assemble the core skeleton.70,151
Total synthesis of RDs persists as a frontier challenge in organic synthesis due to their architecturally congested scaffolds bearing multiple oxygenated quaternary centers. Despite these achievements, persistent challenges include scalability of multi-step sequences and lack of modular strategies for diverse RDs analogues to support biological activity screening. These molecules serve as ideal testing grounds for devising innovative bond-forming tactics and strategic ring-construction approaches, driving ongoing efforts to achieve step-economical and stereocontrolled RDs syntheses.
5.1 Total synthesis of ryanodane core by Deslongchamps group
5.2 Total synthesis of ryanodane core by Inoue group
Radical-mediated homolytic cleavage of the C–O bond at C-6 in 157 abstracted an allyl group from allyltributyltin, establishing the C-6 tetrasubstituted stereocenter in 159. Subsequent isomerization of the C-6 allyl group's double bond and C-5-isobutylation over four steps yielded diene 161. RCM between the C-2 and isomerized C-1 olefins using second-generation Hoveyda-Grubbs catalyst (162) formed pentacycle 163, thereby constructing the framework of ryanodol. Final installation of the C-13 isopropyl group yielded 167, and adjustment of oxidation states at C-1 and C-14 of 167 completed the synthesis of ryanodol (50).
5.3 Total synthesis of ryanodane and isoryanodane core by Reisman group
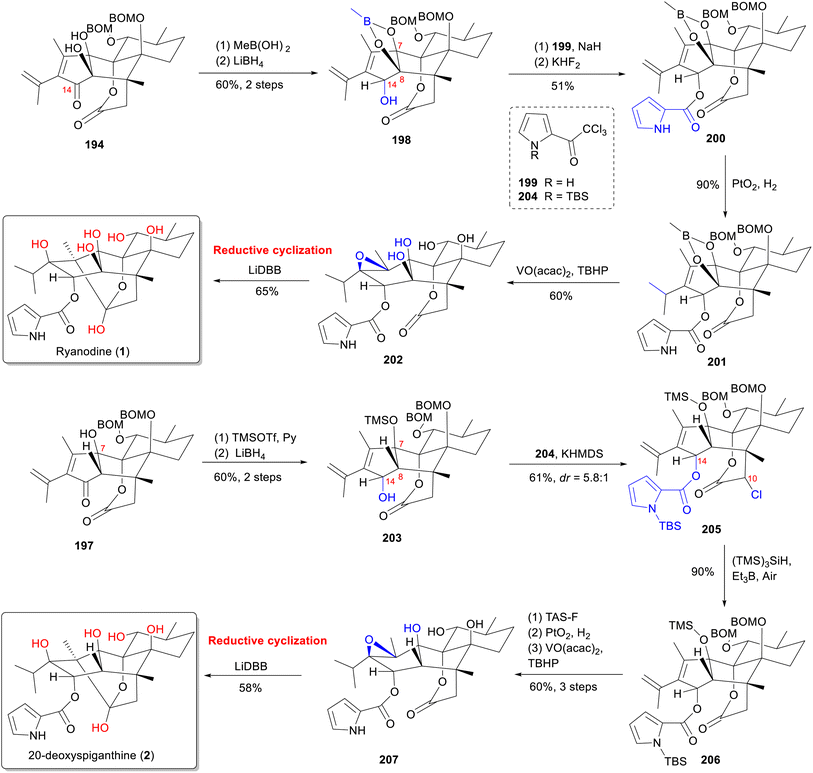
A similar sequence148 was used to convert alcohol 197 into (+)-20-deoxyspiganthine (2) (Scheme 14) in eight steps. Initially, the C-7 hydroxyl group of 197 was protected as a TMS ether with TMSOTf. Subsequent reduction of the C-14 ketone yielded 203. Treatment of 203 with potassium hexamethyldisilazide (KHMDS) and 204 enolized the lactone and prevented translactonization, facilitating acylation of the C-14 alcohol with pyrrole. However, this also resulted in the formation of the α-chlorination product 205 at the C-10 position. Subsequent radical dechlorination of 205 yielded lactone 206. Final hydrogenation, epoxidation, and reductive cyclization of 206 with four-steps sequence was uneventfully advanced to the first asymmetric synthesis of (+)-20-deoxyspiganthine (2). This work demonstrates that strategical formation of C–C and C–O bonds at the correct oxidation states minimizes protecting group manipulations and leads to concise syntheses.157
5.4 Total synthesis of anhydroryanodol (72) by Micalizio group
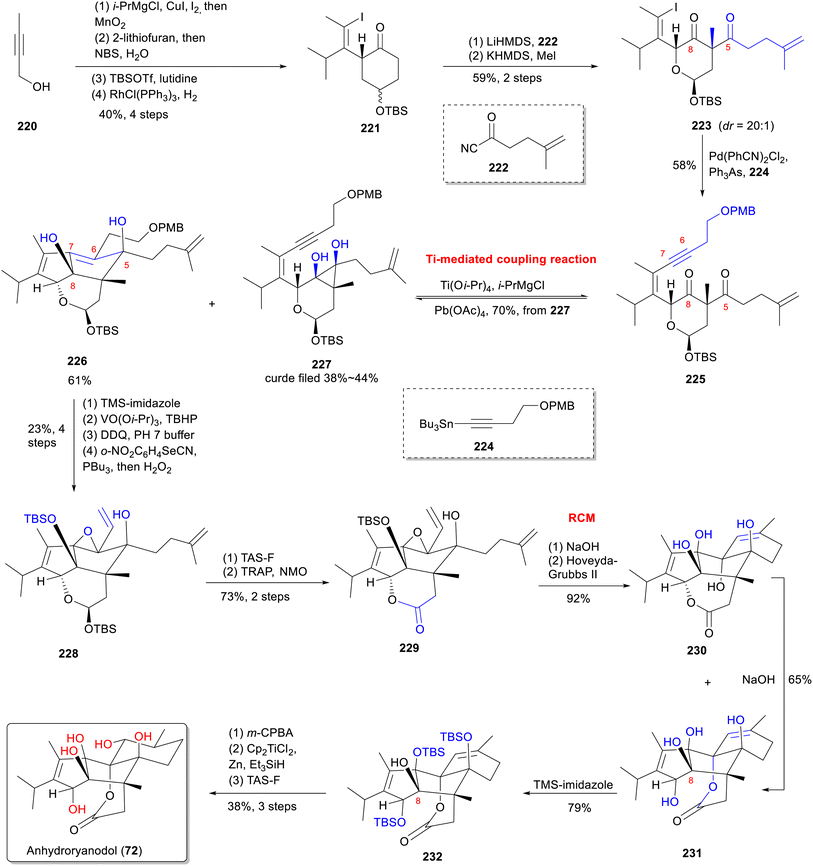
Micalizio and co-workers achieved the first total synthesis of anhydroryanodol (72)150 through a concise, stereoselective route that hinges on a metallacycle-mediated intramolecular coupling between an alkyne and a 1,3-diketone (Scheme 16).161 The synthesis initiated from commercially available alkyne 220, advanced to enyne 221, involving copper-mediated addition of isopropylmagnesium chloride (i-PrMgCl), 1,2-addition of 2-lithiofuran, Achmatowicz rearrangement,162 silylation, and selective hydrogenation with Wilkinson's catalyst163 over four steps. Kinetic enolate acylation of 221 with acyl cyanide 222 delivered a 1,3-diketone that was methylated to give 223. Stille coupling with stannane 224 then provided enyne 225. The key transformation involved treating 225 with titanium isopropoxide (Ti(Oi-Pr)4) and i-PrMgCl. This generated a mixture without oxidative termination to deliver the desired product 226 in 61% yield alongside byproduct 227. A six-step sequence converted 226 into divinyl epoxide 229. Ring-closing metathesis of 229 with Hoveyda-Grubbs II catalyst delivered tetracycle 230, which underwent base-promoted isomerization and lactonization (NaOH) to afford 231. Final stage manipulations selective partial silylation (leaving C-8 unprotected), chemo- and stereoselective epoxidation with m-CPBA, reductive epoxide opening (Cp2TiCl2/Zn/EtSiH), and final silyl ether deprotection-completed the synthesis of anhydroryanodol (72). The first application of a metallacycle-mediated intramolecular alkyne-diketone coupling in natural product synthesis underscores the enduring value of complex total synthesis.5.5 Total synthesis of garajonone (61) by Zhao group
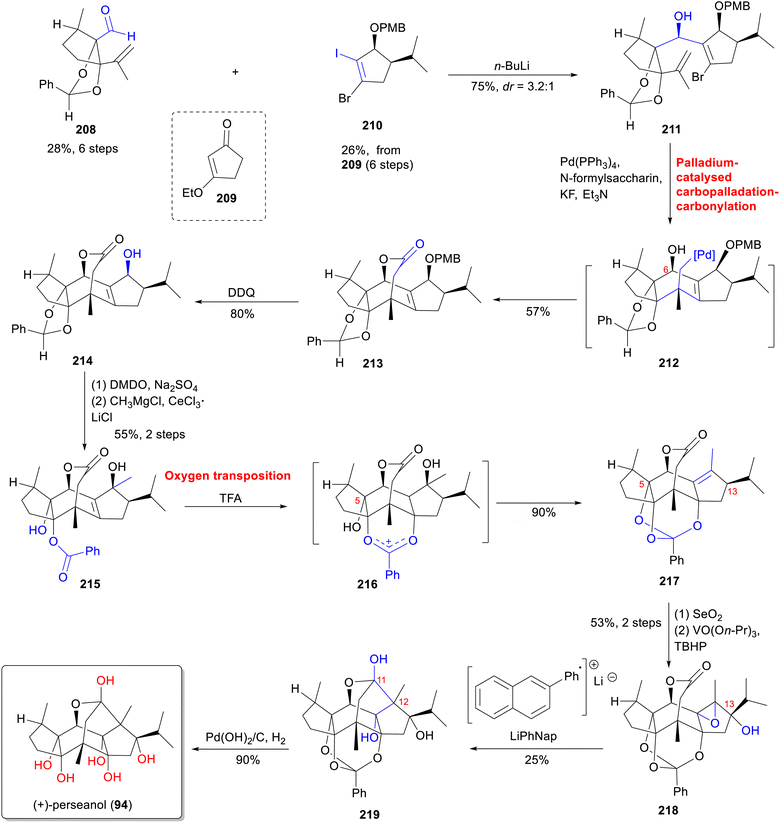
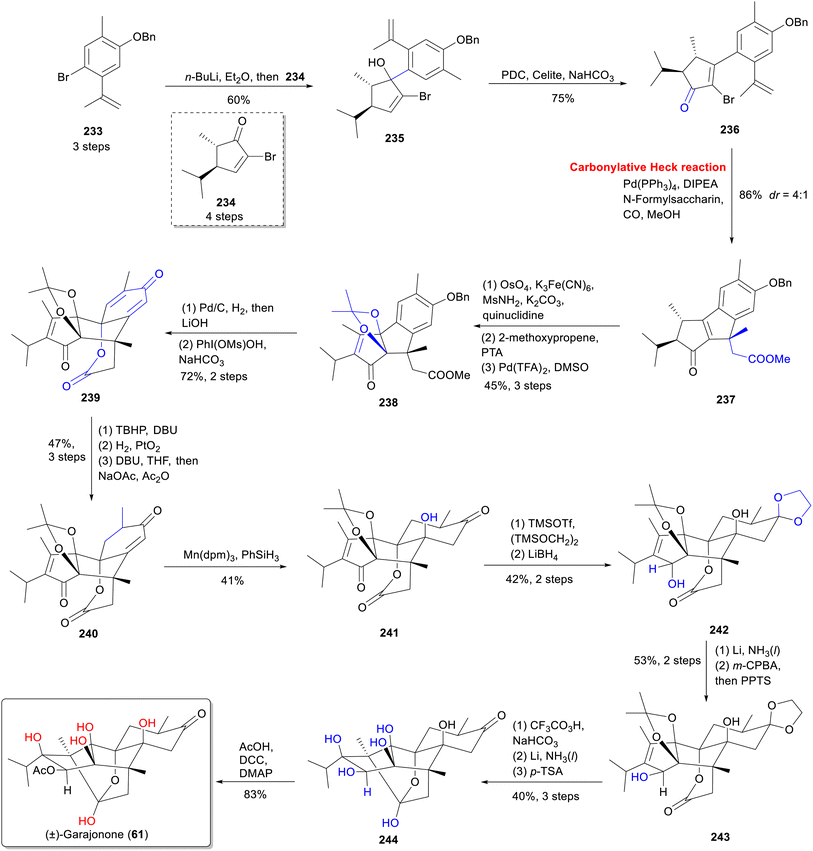
Inspired by terpenes biosynthesis, the total synthesis of garajonone (61) was achieved through a two-phase strategy involving carbon framework assembly followed by oxidation state adjustment (Scheme 17).70 In the first stage, the carbon skeleton was constructed starting with the 1,2-addition of an aryl lithium reagent, prepared in situ from 233 and n-butyllithium with enone 234,164 to form allylic alcohol 235. The resulting 235 was then oxidized with pyridinium dichromate (PDC) to yield vinyl bromide 236. A palladium-catalyzed Heck/carbonylative esterification cascade annulation,165 utilizing N-formylsaccharin, Pd(PPh3)4, and diisopropylethylamine (DIPEA) in methanol under a CO atmosphere, produced desired tricyclic 237 as the major diastereomer. Subsequent oxidation with OsO4 and K3Fe(CN)6, alcohols protection, and dehydrogenative oxidation, smoothly led to enone 238. Catalytic hydrogenation removed the benzyl groups, followed by saponification, and the resulting product was oxidized with PhIOMs(OH) to form dienone 239. In the second stage, Weitz-Scheffer epoxidation166 of 239 was used to mask the bridging olefin and enable regioselective hydrogenation with concomitant epoxide opening. Subsequent hydrogenation using Adam's catalyst, and a retro-Michael reaction under basic conditions to yield desired monoenone 240. Mn(dpm)3-catalyzed hydration of 240 was inverted through elimination and Mukaiyama hydration furnishing tertiary alcohol 241. Followed by ketal protection, 1,2-reduction (using LiBH4), and lithium-ammonia reductive annulation, the lactone 243 was established. Finally, acidic removal of the ethylene glycol protection and esterification with acetic acid using dicyclohexylcarbodiimide/4-methylbenzene (DCC/DMAP) furnished garajonone (61). This synthesis highlights a strategic combination of Pd-catalyzed cascades and selective oxidative manipulations to assemble the highly complex diterpenoid.Despite significant advances in the total synthesis of RDs, much remains to be done to achieve synthetic diversity and enable comprehensive exploration. Continued development of efficient synthetic methods and a deeper understanding of biosynthetic pathways should further unlock the medicinal potential of these complex molecules.
6 Conclusion and future perspectives
The expanding catalogue of naturally occurring RDs, now 135 structures spanning nine subtypes from nine plant species, reveals unprecedented chemical diversity with profound biological implications. RDs are mainly distributed in tropical and subtropical plants, which is closely related to the insecticidal activities, yet most remain chemically underexplored. They are comparatively rare across five plant families and could serve as valuable chemotaxonomic markers. Systematic phytochemical investigations of neglected Lauraceae and Salicaceae lineages may unveil missing scaffolds that bridge the biosynthetic gap between ryanodane and isoryanodane cores. NMR- and MS-guided dereplication and in silico prediction tools will accelerate the targeted isolation of high-content RDs for biological evaluation.167–169The intricate, polyoxygenated structures of RDs, coupled with their capacity to modulate RyRs, position them as privileged scaffolds for treating Ca2+ dysregulation disorders such as cardiac arrhythmias and neurodegenerative disorders.170 Rational design of next-generation RyR therapeutics demands deeper mechanistic insight into isoform-specific channel dysfunction and the molecular basis of existing ligand efficacy.6 Selective insect toxicity paired with low mammalian toxicity also makes RDs attractive leads for eco-friendly insecticides, although their structural complexity and limited supply render development more costly than commercial phthalic or anthranilic diamide RyR modulators.171 No RD-based drugs are currently in clinical use or trials, but continued structural diversification and mechanistic dissection guided by ligands may be achieved in the future. Future studies could map ligand-RyR interaction hotspots and computational docking to design subtype-selective RD derivatives.
Efficient synthetic access remains the critical bottleneck. Despite landmark total synthesis of ten RDs, the dense arrays of quaternary stereocenters and labile hemiketals still demand lengthy sequences; Gram-scale production has not been realized. Convergent synthetic strategies must be integrated with biosynthetic discovery. To date, only two speculative biosynthetic precursors, geranylgeraniol and (R)-mevalonate, have been proposed,77 and the enzymatic mechanisms responsible for the structural diversification of these compounds remain entirely unknown. Their intricate polycyclic architectures, featuring a hemiketal group, introduce substantial complexity and pose formidable challenges for biosynthetic studies. Elucidating the complete pathway from geranylgeranyl diphosphate to the RD skeleton is therefore an urgent objective. The biosynthesis is anticipated to begin with cyclization and skeletal rearrangement mediated by diterpene synthases, followed by extensive oxidative modifications that refine the core structure. Genomics-driven discovery of biosynthetic gene clusters in key species such as Ryania and Cinnamomum, combined with heterologous expression in engineered microbial systems, offers a promising strategy to establish sustainable production platforms and uncover novel bioactive scaffolds.
Comprehensive toxicity and environmental-fate studies are equally essential. Preliminary metabolism data show that anhydrocinnzeylanine (74) is rapidly converted (to 3% remaining) by human liver microsomes via a CYP3A4-dependent pathway in the presence of β-NADPH.172 Interdisciplinary collaboration among synthetic chemists, structural biologists, chemical ecologist, and pharmacologists-augmented by systems biology, environmental toxicology, and AI-driven structure prediction will be required to translate RD research into sustainable agrochemicals and precision therapeutics.
In summary, RDs offer exceptional promise as novel RyR ligands, insecticidal agents, and anti-inflammatory or cardioprotective leads. Their unique architectures and biological profiles merit intensified investigation across chemistry, biology, and environmental science.
7 Author contributions
Bodou Zhang: investigation, data curation, visualization, and writing original draft. Jingwen Zhao: investigation, visualization, and data curation. Sheng Li: data curation and visualization. Hong Liang: data curation and visualization. Xiaojiang Hao: organization, supervision, and revision. Yu Zhang: conceptualization, resources, supervision, revision, and funding acquisition. All authors agree to be accountable for all aspects of work ensuring integrity and accuracy.8 Conflicts of interest
There are no conflicts of interest to declare.9 Data availability
Supplementary information: distribution, bioactivity, and progress in total synthesis of ryanodane diterpenes. See DOI: https://doi.org/10.1039/d5np00052a.10 Acknowledgements
This research obtained financial support from the National Key Research and Development Program of China (2022YFF1100301), Major Science and Technology Project of Henan Province (231100310200), Strategic Priority Research Program of the Chinese Academy of Sciences (XDB1230000), Key Research and Development Program of Yunnan Province (202403AC100010 and 202203AC100009), and CAS Interdisciplinary Team of “Light of West China” Program (xbzg-zdsys-202405). The authors thank Prof. Bo Xu for his advice and polishing on this review.11 References
- Y. Wang, P. Tang, W. Tu, Q. Gao, C. Wang, L. Tan, L. Zhao, H. Han, L. Ma, K. Otsuki, W. Xiao, W. Wang, J. Liu, Y. Li, Z. Zhan, W. Li, X. Zhou and N. Li, Chin. Chem. Lett., 2025, 36, 109955 CrossRef CAS.
- J. R. Hanson, Nat. Prod. Rep., 2003, 20, 70–78 RSC.
- R. Li, S. L. Morris-Natschke and K.-H. Lee, Nat. Prod. Rep., 2016, 33, 1166–1226 RSC.
- J. R. Hanson, Nat. Prod. Rep., 2017, 34, 1233–1243 RSC.
- E. F. Rogers and F. R. Koniuszy, et al. , J. Am. Chem. Soc., 1948, 70, 3086–3088 CrossRef CAS PubMed.
- J. L. Sutko, J. A. Airey, W. Welch and L. Ruest, Pharmacol. Rev., 1997, 49, 53–98 CrossRef CAS.
- G. Santulli, D. Lewis, A. des Georges, A. R. Marks and J. Frank, in Membrane protein complexes: structure and function, 2018, pp. 329–352 Search PubMed.
- G. Meissner, J. Gen. Physiol., 2017, 149, 1065–1089 CrossRef CAS.
- T. Murayama, N. Kurebayashi, R. Ishida and H. Kagechika, Curr. Opin. Pharmacol., 2023, 69, 102356 CrossRef CAS.
- B. M. Fraga, A. Gonzalez-Coloma, C. Gutierrez and D. Terrero, J. Nat. Prod., 1997, 60, 880–883 CrossRef CAS.
- R. Ma, O. Haji-Ghassemi, D. Ma, H. Jiang, L. Lin, L. Yao, A. Samurkas, Y. Li, Y. Wang, P. Cao, S. Wu, Y. Zhang, T. Murayama, B. Moussian, F. Van Petegem and Z. Yuchi, Nat. Chem. Biol., 2020, 16, 1246–1254 CrossRef CAS.
- R. Zalk, O. B. Clarke, A. des Georges, R. A. Grassucci, S. Reiken, F. Mancia, W. A. Hendrickson, J. Frank and A. R. Marks, Nature, 2015, 517, 44–49 CrossRef CAS.
- Z. Yan, X. Bai, C. Yan, J. Wu, Z. Li, T. Xie, W. Peng, C. Yin, X. Li, S. H. W. Scheres, Y. Shi and N. Yan, Nature, 2015, 517, 50–55 CrossRef CAS PubMed.
- L. Lin, C. Wang, W. Wang, H. Jiang, T. Murayama, T. Kobayashi, H. Hadiatullah, Y. S. Chen, S. Wu, Y. Wang, H. Korza, Y. Gu, Y. Zhang, J. Du, F. Van Petegem and Z. Yuchi, Nat. Commun., 2024, 15, 9056 CrossRef CAS.
- Y. S. Chen, M. Garcia-Castañeda, M. Charalambous, D. Rossi, V. Sorrentino and F. Van Petegem, Nat. Commun., 2024, 15, 8630 CrossRef CAS.
- A. R. Marks, J. Clin. Invest., 2023, 133, e162891 CrossRef CAS.
- T. Q. Do and B. C. Knollmann, Annu. Rev. Pharmacol. Toxicol., 2025, 65, 443–463 CrossRef CAS.
- A. des Georges, O. B. Clarke, R. Zalk, Q. Yuan, K. J. Condon, R. A. Grassucci, W. A. Hendrickson, A. R. Marks and J. Frank, Cell, 2016, 167, 145–157 CrossRef CAS PubMed.
- J. Zeng, Y. Xue, P. Shu, H. Qian, R. Sa, M. Xiang, X. N. Li, Z. Luo, G. Yao and Y. Zhang, J. Nat. Prod., 2014, 77, 1948–1954 CrossRef CAS.
- X. Y. Chai, C. C. Bai, H. M. Shi, Z. R. Xu, H. Y. Ren, F. F. Li, Y. N. Lu, Y. L. Song and P. F. Tu, Tetrahedron, 2008, 64, 5743–5747 CrossRef CAS.
- Y. Orihara, H. Hamamoto, H. Kasuga, T. Shimada, Y. Kawaguchi and K. Sekimizu, J. Gen. Virol., 2008, 89, 188–194 CrossRef CAS.
- Y. Zhang, J. Zhou, D. Y. P. Tng, S. Wang, Y. Wang, Y. Peng, H. Liu and Z. Wang, Plants, 2023, 12, 1419 CrossRef CAS.
- J. Wang, B. Su, H. Jiang, N. Cui, Z. Yu, Y. Yang and Y. Sun, Fitoterapia, 2020, 146, 104675 CrossRef CAS PubMed.
- J. Sharifi-Rad, A. Dey, N. Koirala, S. Shaheen, N. El Omari, B. Salehi, T. Goloshvili, N. C. Cirone Silva, A. Bouyahya, S. Vitalini, E. M. Varoni, M. Martorell, A. Abdolshahi, A. O. Docea, M. Iriti, D. Calina, F. Les, V. López and C. Caruntu, Front. Pharmacol., 2021, 12, 600139 CrossRef CAS.
- X. Li, H. Y. Lu, X. W. Jiang, Y. Yang, B. Xing, D. Yao, Q. Wu, Z. H. Xu and Q. C. Zhao, J. Ethnopharmacol., 2021, 266, 113413 CrossRef CAS.
- H. Fan, G. Huang, Q. Guo, J. Ma, Y. Huang, S. Huang, M. Wei, C. Xie, B. Yan, S. Zhao, G. Chen, J. Zheng, Z. Zhou and H. Gao, J. Agric. Food Chem., 2024, 72, 16263–16275 CrossRef CAS PubMed.
- D. Mohanty, S. Padhee, C. Sahoo, S. Jena, A. Sahoo, P. Chandra Panda, S. Nayak and A. Ray, Heliyon, 2024, 10, e24120 CrossRef CAS.
- C. H. Totini, E. Umehara, I. M. A. Reis, J. H. G. Lago and A. Branco, Chem. Biodiversity, 2023, 20, e202300947 CrossRef CAS PubMed.
- L. Martín, A. González-Coloma, R. Adami, M. Scognamiglio, E. Reverchon, G. D. Porta, J. S. Urieta and A. M. Mainar, J. Supercrit. Fluids, 2011, 60, 16–20 CrossRef.
- P. Kondraskov, N. Schütz, C. Schüßler, M. M. de Sequeira, A. S. Guerra, J. Caujapé-Castells, R. Jaén-Molina, Á. Marrero-Rodríguez, M. A. Koch, P. Linder, J. Kovar-Eder and M. Thiv, PLoS One, 2015, 10, e0132091 CrossRef.
- B. M. Fraga, C. E. Diaz, P. Bolanos, M. Bailen, M. F. Andres and A. Gonzalez-Coloma, Phytochemistry, 2020, 176, 112398 CrossRef CAS.
- B. M. Fraga, D. Terrero, C. Gutierrez and A. Gonzalez-Coloma, Phytochemistry, 2001, 56, 315–320 CrossRef CAS.
- A. Gonzalez-Coloma, M. Reina, C. Gutierrez and B. M. Fraga, in Studies in Natural Products Chemistry, ed. R. Attaur, Elsevier, 2002, vol. 26, pp. 849–879 Search PubMed.
- J. Li, R. Halitschke, D. Li, C. Paetz, H. Su, S. Heiling, S. Xu and I. T. Baldwin, Science, 2021, 371, 255–260 CrossRef CAS PubMed.
- E. Leskinen and C. Alstrom-Rapaport, Plant Syst. Evol., 1999, 215, 209–227 CrossRef.
- T. A. P. Group, M. W. Chase, M. J. M. Christenhusz, M. F. Fay, J. W. Byng, W. S. Judd, D. E. Soltis, D. J. Mabberley, A. N. Sennikov, P. S. Soltis and P. F. Stevens, Bot. J. Linn. Soc., 2016, 181, 1–20 CrossRef.
- P. Cabras, P. Caboni and M. Cabras, J. Agric. Food Chem., 2001, 49, 3161–3163 CrossRef CAS PubMed.
- W. Tang, H. Xu, D. Zeng and L. Yu, Fitoterapia, 2012, 83, 513–517 CrossRef CAS PubMed.
- X. F. Zhang, X. Li, B. S. Min and K. H. Bae, Chin. Chem. Lett., 2008, 19, 1080–1082 CrossRef CAS.
- A. L. F. Camurça-Vasconcelos, N. R. F. Nascimento, C. M. Sousa, L. M. Melo, S. M. Morais, C. M. L. Bevilaqua and M. F. G. Rocha, J. Ethnopharmacol., 2004, 92, 257–261 CrossRef PubMed.
- W. L. Ribeiro, W. P. Andre, G. S. Cavalcante, J. V. de Araújo-Filho, J. M. Santos, I. T. Macedo, J. V. de Melo, S. M. de Morais and C. M. Bevilaqua, Small Rumin. Res., 2017, 153, 146–152 CrossRef.
- H. Achenbach, H. Hubner, W. Vierling, W. Brandt and M. Reiter, J. Nat. Prod., 1995, 58, 1092–1096 CrossRef CAS PubMed.
- Y. Lv, T. Tian, Y. J. Wang, J. P. Huang and S. X. Huang, Nat. Prod. Bioprospect., 2022, 12, 15 CrossRef PubMed.
- J. F. Araújo Neto, E. M. Ribeiro, A. E. do Vale, J. M. David and J. P. David, Mini-Rev. Med. Chem., 2021, 21, 2458–2480 CrossRef.
- G. T. Prance, Allertonia, 2012, 12, 1–164 CrossRef.
- P. T. M. Huong, C. P. Thi, V. L. Thi, H. T. T. Hong, C. N. The, V. L. Dang, H. Q. Tran, X. C. Nguyen, N. N. Hoai and C. V. Minh, Nat. Prod. Res., 2024, 38, 3140–3145 CrossRef CAS.
- B. Negin and G. Jander, Curr. Opin. Plant Biol., 2023, 73, 102368 CrossRef CAS.
- B. M. Fraga, D. Terrero, P. Bolanos and C. E. Diaz, Tetrahedron Lett., 2017, 58, 2261–2263 CrossRef CAS.
- L. Zhou, Y. Tuo, Y. Hao, X. Guo, W. Tang, Y. Xue, J. Zeng, Y. Zhou, M. Xiang, J. Zuo, G. Yao and Y. Zhang, Org. Lett., 2017, 19, 3029–3032 CrossRef CAS.
- H. Zhou, Y. Guoruoluo, Y. Tuo, J. Zhou, H. Zhang, W. Wang, M. Xiang, H. A. Aisa and G. Yao, Org. Lett., 2019, 21, 549–553 CrossRef CAS.
- Y. Kashiwada, T. Nohara, T. Tomimatsu and I. Nishioka, Chem. Pharm. Bull., 1981, 29, 2686–2688 CrossRef CAS.
- P. R. Jefferies, R. F. Toia and J. E. Casida, J. Nat. Prod., 1991, 54, 1147–1149 CrossRef CAS.
- H. Hubner, W. Vierling, W. Brandt, M. Reiter and H. Achenbach, Phytochemistry, 2001, 57, 285–296 CrossRef CAS PubMed.
- K. Wiesner, Collect. Czech. Chem. Commun., 1968, 33, 2656–2665 CrossRef CAS.
- K. Wlesner, Adv. Org. Chem., 1972, 8, 295–316 Search PubMed.
- P. R. Jefferies, W. W. Lam, R. F. Toia and J. E. Casida, J. Agric. Food Chem., 1992, 40, 509–512 CrossRef CAS.
- P. R. Jefferies, R. F. Toia, B. Brannigan, I. Pessah and J. E. Casida, J. Agric. Food Chem., 1992, 40, 142–146 CrossRef CAS.
- L. Ruest and M. Dodier, Can. J. Chem., 1996, 74, 2424–2433 CrossRef CAS.
- L. Ruest, M. Dodier, H. De Seve, C. Lessard and P. Mongrain, Can. J. Chem., 2002, 80, 483–488 CrossRef CAS.
- L. Ruest, D. R. Taylor and P. Deslongchamps, Can. J. Chem., 1985, 63, 2840–2843 CrossRef CAS.
- L. Ruest, C. Berthelette, M. Dodier, L. Dube and D. St-Martin, Can. J. Chem., 1999, 77, 12–15 CrossRef CAS.
- A. Gonzalez-Coloma, D. Terrero, A. Perales, P. Escoubas and B. M. Fraga, J. Agric. Food Chem., 1996, 44, 296–300 CrossRef CAS.
- A. Isogai, S. Murakoshi, A. Suzuki and S. Tamura, Agric. Biol. Chem., 1977, 41, 1779–1784 CAS.
- T. Nohara, N. Tokubuchi, M. Kuroiwa and I. Nishioka, Chem. Pharm. Bull., 1980, 28, 2682–2686 CrossRef CAS.
- A. Gonzalez-Coloma, R. Cabrera, P. Castanera, C. Gutierrez and B. M. Fraga, Phytochemistry, 1992, 31, 1549–1552 CrossRef CAS.
- A. Gonzalez-Coloma, M. F. Andres, R. Contreras, G. E. Zuniga and C. E. Diaz, Plants, 2022, 11, 418 CrossRef CAS.
- R. B. Kelly, D. J. Whittingham and K. Wiesner, Can. J. Chem., 1951, 29, 905–910 CrossRef.
- M. Koshimizu, M. Nagatomo and M. Inoue, Angew Chem., Int. Ed. Engl., 2016, 55, 2493–2497 CrossRef CAS PubMed.
- M. L. Barreiros, J. P. David, J. M. David, L. M. Xavier Lopes, M. S. de Sa, J. F. Costa, M. Z. Almeida, L. P. de Queiroz and A. E. Sant'ana, Phytochemistry, 2007, 68, 1735–1739 CrossRef CAS.
- J. B. Qiao, L. Meng, J. Y. Pei, H. Shao and Y. M. Zhao, Angew Chem., Int. Ed. Engl., 2025, 64, e202417647 CrossRef CAS PubMed.
- T. N. Yagi A, T. Nohara, G. Nonaka, I. Nishioka and A. Koda, Chem. Pharm. Bull., 1980, 28, 1432–1436 CrossRef.
- S. He, Y. Jiang and P. F. Tu, J. Asian Nat. Prod. Res., 2016, 18, 134–140 CrossRef CAS.
- V. C. Pham, T. T. A. Nguyen, T. O. Vu, T. Q. Cao, B. S. Min and J. A. Kim, Phytochem. Lett., 2019, 32, 23–28 CrossRef CAS.
- T. M. Ngoc, D. T. Ha, I. S. Lee, B. S. Min, M. K. Na, H. J. Jung, S. M. Lee and K. H. Bae, Helv. Chim. Acta, 2009, 92, 2058–2062 CrossRef CAS.
- T. Nohara, Y. Kashiwada, T. Tomimatsu and I. Nishioka, Phytochemistry, 1982, 21, 2130–2132 CrossRef CAS.
- L. Zhou, G. Zheng, H. Li, B. Gao, Y. Guoruoluo, W. Tang, G. Yao and Y. Zhang, Phytochemistry, 2022, 196, 113077 CrossRef CAS PubMed.
- T. Nohara, Y. Kashiwada, K. Murakami, T. Tomimatsu, M. Kido, A. Yagi and I. Nishioka, Chem. Pharm. Bull., 1981, 29, 2451–2459 CrossRef CAS.
- Y. Q. Huang, B. Zhou, Y. R. Yuan, Y. H. Ren, D. L. Li, K. Zhang and J. M. Yue, Tetrahedron Lett., 2021, 73, 153110 CrossRef CAS.
- W. Tang, X. Wei, H. Xu, D. Zeng and L. Long, Fitoterapia, 2009, 80, 286–289 CrossRef CAS PubMed.
- T. Nohara, Y. Kashiwada and I. Nishioka, Phytochemistry, 1985, 24, 1849–1850 CrossRef CAS.
- J. D. Fessenden, L. Chen, Y. Wang, C. Paolini, C. Franzini-Armstrong, P. D. Allen and I. N. Pessah, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 2865–2870 CrossRef CAS PubMed.
- A. K. Isaacs, S. Qi, R. Sarpong and J. E. Casida, Chem. Res. Toxicol., 2012, 25, 1571–1573 Search PubMed.
- G. Meissner, J. Biol. Chem., 1986, 261, 6300–6306 CrossRef CAS PubMed.
- M. W. Fryer, G. D. Lamb and I. R. Neering, J. Physiol., 1989, 414, 399–413 CrossRef CAS.
- W. Peng, H. Shen, J. Wu, W. Guo, X. Pan, R. Wang, S. R. W. Chen and N. Yan, Science, 2016, 354, aah5324 CrossRef.
- H. Takeshima, S. Nishimura, T. Matsumoto, H. Ishida, K. Kangawa, N. Minamino, H. Matsuo, M. Ueda, M. Hanaoka and T. Hirose, Nature, 1989, 339, 439–445 CrossRef CAS PubMed.
- M. C. Miotto, G. Weninger, H. Dridi, Q. Yuan, Y. Liu, A. Wronska, Z. Melville, L. Sittenfeld, S. Reiken and A. R. Marks, Sci. Adv., 2022, 8, eabo1272 CrossRef CAS.
- A. Kushnir, B. Wajsberg and A. R. Marks, Biochim. Biophys. Acta, Mol. Cell Res., 2018, 1865, 1687–1697 CrossRef CAS PubMed.
- S. O. Marx, S. Reiken, Y. Hisamatsu, T. Jayaraman, D. Burkhoff, N. Rosemblit and A. R. Marks, Cell, 2000, 101, 365–376 CrossRef CAS PubMed.
- X. H. Wehrens, S. E. Lehnart, F. Huang, J. A. Vest, S. R. Reiken, P. J. Mohler, J. Sun, S. Guatimosim, L. S. Song, N. Rosemblit, J. M. D'Armiento, C. Napolitano, M. Memmi, S. G. Priori, W. J. Lederer and A. R. Marks, Cell, 2003, 113, 829–840 CrossRef CAS PubMed.
- X. Liu, M. J. Betzenhauser, S. Reiken, A. C. Meli, W. Xie, B. X. Chen, O. Arancio and A. R. Marks, Cell, 2012, 150, 1055–1067 CrossRef CAS PubMed.
- D. C. Andersson, M. J. Betzenhauser, S. Reiken, A. C. Meli, A. Umanskaya, W. Xie, T. Shiomi, R. Zalk, A. Lacampagne and A. R. Marks, Cell Metab., 2011, 14, 196–207 CrossRef CAS.
- A. M. Bellinger, S. Reiken, C. Carlson, M. Mongillo, X. Liu, L. Rothman, S. Matecki, A. Lacampagne and A. R. Marks, Nat. Med., 2009, 15, 325–330 CrossRef CAS PubMed.
- D. C. Andersson and A. R. Marks, Drug Discovery Today: Dis. Mech., 2010, 7, e151–e157 CrossRef CAS.
- N. L. Thomas and A. J. Williams, Wiley Interdiscip. Rev.: Membr. Transp. Signaling, 2012, 1, 383–397 CAS.
- N. Kaneko, C. M. Loughrey, G. Smith, R. Matsuda, T. Hasunuma, P. B. Mark, M. Toda, M. Shinozaki, N. Otani, S. Kayley, A. Da Silva Costa, T. P. Martin, S. Dobi, P. Saxena, K. Shimamoto, T. Ishikawa, R. Kambayashi, A. Riddell, E. B. Elliott, C. S. McCarroll, T. Sakai, Y. Mitsuhisa, S. Hirano, T. Kitai, K. Kusano, Y. Inoue, M. Nakamura, M. Kikuchi, S. Toyoda, I. Taguchi, T. Fujiwara, A. Sugiyama, Y. Kumagai and K. Iwata, Br. J. Pharmacol., 2024, 181, 3401–3419 CrossRef CAS.
- A. M. Bellinger, S. Reiken, M. Dura, P. W. Murphy, S. X. Deng, D. W. Landry, D. Nieman, S. E. Lehnart, M. Samaru, A. LaCampagne and A. R. Marks, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 2198–2202 CrossRef PubMed.
- J. J. Todd, T. A. Lawal, I. C. Chrismer, A. Kokkinis, C. Grunseich, M. S. Jain, M. R. Waite, V. Biancavilla, S. Pocock, K. Brooks, C. J. Mendoza, G. Norato, K. Cheung, W. Riekhof, P. Varma, C. Colina-Prisco, M. Emile-Backer, K. G. Meilleur, A. R. Marks, Y. Webb, E. E. Marcantonio, A. R. Foley, C. G. Bönnemann and P. Mohassel, EClinicalMedicine, 2024, 68, 102433 CrossRef PubMed.
- K. R. Bidasee and H. R. Besch, J. Biol. Chem., 1998, 273, 12176–12186 CrossRef CAS PubMed.
- E. Buck, I. Zimanyi, J. J. Abramson and I. N. Pessah, J. Biol. Chem., 1992, 267, 23560–23567 CrossRef CAS PubMed.
- K. M. Ranatunga, C. S. R. Wayne, R. Luc, W. William and A. J. Williams, Mol. Membr. Biol., 2007, 24, 185–193 CrossRef CAS.
- B. Tanna, W. Welch, L. Ruest, J. L. Sutko and A. J. Williams, Mol. Pharmacol., 2006, 69, 1990–1997 CrossRef CAS.
- V. A. Ngo, L. L. Perissinotti, W. Miranda, S. R. W. Chen and S. Y. Noskov, Biophys. J., 2017, 112, 1645–1653 CrossRef CAS PubMed.
- B. Tanna, W. Welch, L. Ruest, J. L. Sutko and A. J. Williams, J. Gen. Physiol, 2000, 116, 1–9 CrossRef CAS PubMed.
- F. Van Petegem, Cell Calcium, 2017, 61, 50–52 CrossRef CAS PubMed.
- P. R. Jefferies, P. Yu and J. E. Casida, Pestic. Sci., 1997, 51, 33–38 CrossRef CAS.
- A. L. Waterhouse, I. N. Pessah, A. O. Francini and J. E. Casida, J. Med. Chem., 1987, 30, 710–716 CrossRef CAS.
- W. Welch, A. J. Williams, A. Tinker, K. E. Mitchell, P. Deslongchamps, J. Lamothe, K. Gerzon, K. R. Bidasee, H. R. Besch, J. A. Airey, J. L. Sutko and L. Ruest, Biochemistry, 1997, 36, 2939–2950 CrossRef CAS PubMed.
- W. Welch, S. Ahmad, J. A. Airey, K. Gerzon, R. A. Humerickhouse, H. R. J. Besch, L. Ruest, P. Deslongchamps and J. L. Sutko, Biochemistry, 1994, 33, 6074–6085 CrossRef CAS PubMed.
- P. R. Jefferies, T. A. Blumenkopf, P. J. Gengo, L. C. Cole and J. E. Casida, J. Med. Chem., 1996, 39, 2331–2338 CrossRef CAS.
- P. R. Jefferies, P. J. Gengo, M. J. Watson and J. E. Casida, J. Med. Chem., 1996, 39, 2339–2346 CrossRef CAS.
- W. Welch, J. L. Sutko, K. E. Mitchell, J. Airey and L. Ruest, Biochemistry, 1996, 35, 7165–7173 CrossRef CAS.
- J. Ramos-Franco, A. M. Gomez, A. Nani, Y. Liu, J. A. Copello and M. Fill, Pflug. Arch. Eur. J. Phys., 2010, 460, 767–776 CrossRef CAS PubMed.
- M. Wu, L. Ni, H. Lu, H. Xu, S. Zou and X. Zou, J. Chem., 2020, 2020, 5097542 Search PubMed.
- D. B. Sattelle, D. Cordova and T. R. Cheek, Invertebr. Neurosci., 2008, 8, 107 CrossRef CAS PubMed.
- Z. Sun and H. Xu, Mini-Rev. Med. Chem., 2019, 19, 22–33 CrossRef CAS.
- P. N. R. Usherwood and H. Vais, Toxicol. Lett., 1995, 82–83, 247–254 CrossRef CAS.
- A. Gonzalez-Coloma, C. Gutierrez, H. Hubner, H. Achenbach, D. Terrero and B. M. Fraga, J. Agric. Food Chem., 1999, 47, 4419–4424 CrossRef CAS PubMed.
- S. Q. Ling, Y. N. Xu, Y. P. Gu, S. Y. Liu and W. W. Tang, Pestic. Biochem. Physiol., 2018, 152, 90–97 CrossRef CAS PubMed.
- S. Q. Ling, B. He, D. Q. Zeng and W. W. Tang, Environ. Sci. Pollut. Res., 2020, 27, 12181–12191 CrossRef CAS PubMed.
- J. E. Casida, Annu. Rev. Entomol., 2018, 63, 125–144 CrossRef CAS.
- A. Samurkas, L. Yao, H. Hadiatullah, R. Ma, Y. Xie, R. Sundarraj, H. Zuilhof and Z. Yuchi, Curr. Pharm. Des., 2022, 28, 26–35 CrossRef CAS.
- M. Jindra, S. Tumova, L. Bittova, R. Tuma and D. Sedlak, Curr. Opin. Insect Sci., 2024, 101234 CrossRef PubMed.
- N. Okamoto and N. Yamanaka, J. Pestic. Sci., 2021, 46, 23–28 CrossRef CAS.
- S. Liu, K. Li, Y. Gao, X. Liu, W. Chen, W. Ge, Q. Feng, S. R. Palli and S. Li, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, 139–144 CrossRef CAS.
- W. Li, T. Cheng, W. Hu, Z. Peng, C. Liu and Q. Xia, Mol. Genet. Genomics, 2016, 291, 2159–2171 CrossRef CAS PubMed.
- D. Bystranowska, Z. Szewczuk, M. Lisowski, E. Sitkiewicz, P. Dobryszycki, A. Ożyhar and M. Kochman, Arch. Biochem. Biophys., 2012, 517, 12–19 CrossRef CAS PubMed.
- W. Zhang, G. Liang, L. Ma, T. Jiang and H. Xiao, J. Econ. Entomol., 2019, 112, 1411–1417 CrossRef CAS.
- L. Xu, S. C. Yi, J. Y. Li, Y. Tong, C. Xie, D. Q. Zeng and W. W. Tang, J. Agric. Food Chem., 2022, 70, 3151–3161 CrossRef CAS PubMed.
- L. Xu, X. L. Meng, S. H. Bangash, F. Zhang, D. Q. Zeng and W. W. Tang, Pest Manage. Sci., 2022, 78, 134–142 CrossRef CAS PubMed.
- T. Kayukawa, C. Minakuchi, T. Namiki, T. Togawa, M. Yoshiyama, M. Kamimura, K. Mita, S. Imanishi, M. Kiuchi, Y. Ishikawa and T. Shinoda, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 11729–11734 CrossRef CAS PubMed.
- P. V. Licciardi and J. R. Underwood, Int. Immunopharmacol., 2011, 11, 390–398 CrossRef CAS PubMed.
- M. T. Islam, S. K. Bardaweel, M. S. Mubarak, W. Koch, K. Gaweł-Beben, B. Antosiewicz and J. Sharifi-Rad, Front. Immunol., 2021, 11, 572136 CrossRef.
- J. E. Almeida, A. C. de Oliveira, C. E. de Castro Alves, S. M. C. Filho, E. C. P. de Oliveira, J. P. Zuliani and G. S. Pontes, Int. J. Mol. Sci., 2025, 26, 2250 CrossRef CAS PubMed.
- Q. H. Wei, X. X. Cao, D. F. Xu, S. T. Wang, J. S. Zhang and H. Zhang, Phytochemistry, 2023, 210, 113646 CrossRef CAS.
- H. E. Lee, G. Yang, J. S. Choi and J. Y. Lee, Toxicol. Res., 2017, 33, 283–290 CrossRef CAS PubMed.
- R. J. Quinn, T. Mak, D. R. Littler, J. Rossjohn and M. Liu, J. Nat. Prod., 2023, 86, 2630–2637 CrossRef CAS PubMed.
- S. W. Chang, J. S. Lee, J. H. Lee, J. Y. Kim, J. Hong, S. K. Kim, D. Lee and D. S. Jang, J. Nat. Prod., 2021, 84, 553–561 CrossRef CAS PubMed.
- N. de Jesus Menezes-Filho, C. d. S. Souza, T. C. S. Costa, V. D. A. Da Silva, C. S. de Oliveira Ribeiro, M. L. Barreiros, J. F. O. Costa, J. M. David, J. P. David and S. L. Costa, Nat. Prod. Commun., 2014, 9, 1245–1248 CrossRef CAS.
- L. Meng, J. B. Qiao and Y. M. Zhao, Chin. J. Org. Chem., 2025, 45, 804–813 CrossRef CAS.
- A. Belanger, D. J. Berney, H. J. Borschberg, R. Brousseau, A. Doutheau, R. Durand, H. Katayama, R. Lapalme, D. M. Leturc and C. C. Liao, Can. J. Chem., 1979, 57, 3348–3354 CrossRef CAS.
- P. Deslongchamps, A. Belanger, D. J. Berney, H. J. Borschberg, R. Brousseau, A. Doutheau, R. Durand, H. Katayama, R. Lapalme and D. M. Leturc, Can. J. Chem., 1990, 68, 127–152 CrossRef CAS.
- P. Deslongchamps, A. Belanger, D. J. Berney, H. J. Borschberg, R. Brousseau, A. Doutheau, R. Durand, H. Katayama, R. Lapalme and D. M. Leturc, Can. J. Chem., 1990, 68, 186–192 CrossRef CAS.
- P. Deslongchamps, A. Belanger, D. J. Berney, H. J. Borschberg, R. Brousseau, A. Doutheau, R. Durand, H. Katayama, R. Lapalme and D. M. Leturc, Can. J. Chem., 1990, 68, 115–126 CrossRef CAS.
- M. Nagatomo, M. Koshimizu, K. Masuda, T. Tabuchi, D. Urabe and M. Inoue, J. Am. Chem. Soc., 2014, 136, 5916–5919 CrossRef CAS.
- D. Urabe, M. Nagatomo, K. Hagiwara, K. Masuda and M. Inoue, Chem. Sci., 2013, 4, 1615–1619 RSC.
- K. V. Chuang, C. Xu and S. E. Reisman, Science, 2016, 353, 912–915 CrossRef CAS PubMed.
- C. Xu, A. Han, S. C. Virgil and S. E. Reisman, ACS Cent. Sci., 2017, 3, 278–282 CrossRef CAS PubMed.
- M. M. Heravi and L. Mohammadi, RSC Adv., 2021, 11, 38325–38373 RSC.
- K. Du, M. J. Kier, Z. D. Stempel, V. Jeso, A. L. Rheingold and G. C. Micalizio, J. Am. Chem. Soc., 2020, 142, 12937–12941 CrossRef CAS PubMed.
- L. Meng, J. Y. Pei, J. B. Qiao and Y. M. Zhao, Org. Lett., 2025, 27, 2521–2525 CrossRef CAS.
- S. Fatima, A. F. Zahoor, S. G. Khan, S. A. R. Naqvi, S. M. Hussain, U. Nazeer, A. Mansha, H. Ahmad, A. R. Chaudhry and A. Irfan, RSC Adv., 2024, 14, 23423–23458 RSC.
- K. Hagiwara, M. Himuro, M. Hirama and M. Inoue, Tetrahedron Lett., 2009, 50, 1035–1037 CrossRef CAS.
- K. Masuda, M. Nagatomo and M. Inoue, Chem. Pharm. Bull., 2016, 64, 874–879 CrossRef CAS PubMed.
- M. Nagatomo, K. Hagiwara, K. Masuda, M. Koshimizu, T. Kawamata, Y. Matsui, D. Urabe and M. Inoue, Chem.–Eur. J., 2016, 22, 222–229 CrossRef CAS.
- K. Masuda, M. Koshimizu, M. Nagatomo and M. Inoue, Chem.–Eur. J., 2016, 22, 230–236 CrossRef CAS.
- S. E. Dibrell, Y. Tao and S. E. Reisman, Acc. Chem. Res., 2021, 54, 1360–1373 CrossRef CAS PubMed.
- A. Han, Y. Tao and S. E. Reisman, Nature, 2019, 573, 563–567 CrossRef CAS.
- C. Eschmann, L. Song and P. R. Schreiner, Angew Chem. Int. Ed. Engl., 2021, 133, 4873–4882 CrossRef.
- T. Ueda, H. Konishi and K. Manabe, Org. Lett., 2013, 15, 5370–5373 CrossRef CAS.
- K. Du, M. J. Kier, A. L. Rheingold and G. C. Micalizio, Org. Lett., 2018, 20, 6457–6461 CrossRef CAS PubMed.
- L. Liang, L. D. Guo and R. Tong, Acc. Chem. Res., 2022, 55, 2326–2340 CrossRef CAS PubMed.
- S. K. Tanielyan, R. L. Augustine, N. Marin and G. Alvez, ACS Catal., 2011, 1, 159–169 CrossRef CAS.
- Y. Wang, M. Zarca, L. Z. Gong and L. Zhang, J. Am. Chem. Soc., 2016, 138, 7516–7519 CrossRef CAS PubMed.
- H. S. Sims and M. Dai, J. Org. Chem., 2023, 88, 4925–4941 CrossRef CAS PubMed.
- W. Adam, P. B. Rao, H.-G. Degen, A. Levai, T. Patonay and C. R. Saha-Moller, J. Org. Chem., 2002, 67, 259–264 CrossRef CAS.
- B. Zhang, S. Li, Z. Liang, Y. Wei, J. Dong, H. Wen, L. Guo, X. Hao and Y. Zhang, Food Sci. Hum. Wellness, 2025, 14, 9250003 CrossRef.
- M. Meunier, A. Schinkovitz and S. Derbré, Nat. Prod. Rep., 2024, 41, 1766–1786 RSC.
- M. A. Beniddir, K. B. Kang, G. Genta-Jouve, F. Huber, S. Rogers and J. J. J. van der Hooft, Nat. Prod. Rep., 2021, 38, 1967–1993 RSC.
- L. Sun and H. Wei, Cell. Mol. Neurobiol., 2021, 41, 1613–1624 CrossRef CAS PubMed.
- L. Lin, H. Jiang, H. Hadiatullah, R. Ma, H. Korza, Y. Gu and Z. Yuchi, J. Agric. Food Chem., 2022, 70, 16156–16163 CrossRef CAS PubMed.
- S. M. Choi, V. C. Pham, S. Lee and J. A. Kim, Pharmaceutics, 2021, 13, 1316 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |