Themed collection Single Cell Analysis

The next frontier in single cell analysis: multimodal studies and clinical translation
Thought leaders Pratip Chattopadhyay and Daniel Chiu introduce the Lab on a Chip single cell analysis thematic collection.

Lab Chip, 2019,19, 3573-3574
https://doi.org/10.1039/C9LC90109D
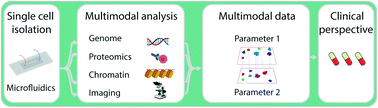
Frontiers in single cell analysis: multimodal technologies and their clinical perspectives
Multimodal single cell analysis provides insights in cellular processes such as cell fate decisions, physiological heterogeneity or genotype–phenotype linkages. This review presents an overview of recent multimodal microfluidic platforms with potential in biomedical research.
Lab Chip, 2022,22, 2403-2422
https://doi.org/10.1039/D2LC00220E
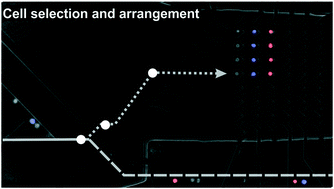
Combining dielectrophoresis and computer vision for precise and fully automated single-cell handling and analysis
We employ real-time image processing in the active control of dielectrophoretic actuation to select, isolate and arrange individual cells in a microfluidic channel.
Lab Chip, 2019,19, 4016-4020
https://doi.org/10.1039/C9LC00800D
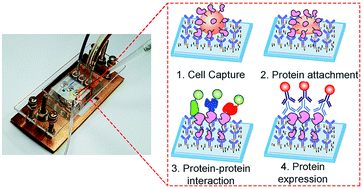
Profiling protein–protein interactions of single cancer cells with in situ lysis and co-immunoprecipitation
We developed a single-cell version of the co-immunoprecipitation (co-IP) analysis that examines the amount and protein–protein interactions of target proteins immunoprecipitated from individual cells.
Lab Chip, 2019,19, 1922-1928
https://doi.org/10.1039/C9LC00139E
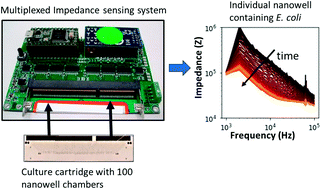
Digital electrical impedance analysis for single bacterium sensing and antimicrobial susceptibility testing
We report on a hand-held multiplexed impedance sensor system and show evidence for impedance-based monitoring of the growth of a single bacterium.
Lab Chip, 2021,21, 1073-1083
https://doi.org/10.1039/D0LC00937G
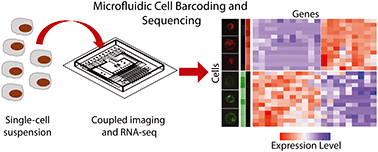
μCB-seq: microfluidic cell barcoding and sequencing for high-resolution imaging and sequencing of single cells
We present a platform for on-chip molecular barcoding that combines high-resolution imaging with genomic analysis, enabling multi-modal phenotypic measurements in single cells.
Lab Chip, 2020,20, 3899-3913
https://doi.org/10.1039/D0LC00169D
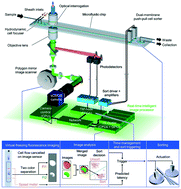
Intelligent image-activated cell sorting 2.0
The upgraded version of intelligent image-activated cell sorting (iIACS) has enabled higher-throughput and more sensitive intelligent image-based sorting of single live cells from heterogeneous populations.
Lab Chip, 2020,20, 2263-2273
https://doi.org/10.1039/D0LC00080A
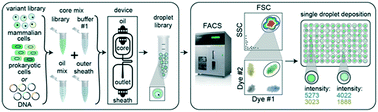
Double emulsion flow cytometry with high-throughput single droplet isolation and nucleic acid recovery
We have developed a novel workflow (sdDE-FACS, ![[s with combining low line]](https://www.rsc.org/images/entities/char_0073_0332.gif) ingle
ingle ![[d with combining low line]](https://www.rsc.org/images/entities/char_0064_0332.gif) roplet
roplet ![[D with combining low line]](https://www.rsc.org/images/entities/char_0044_0332.gif) ouble
ouble ![[E with combining low line]](https://www.rsc.org/images/entities/char_0045_0332.gif) mulsion FACS) that allows robust production, screening, and sorting of single double emulsion droplets with complete nucleic acid recovery.
mulsion FACS) that allows robust production, screening, and sorting of single double emulsion droplets with complete nucleic acid recovery.
Lab Chip, 2020,20, 2062-2074
https://doi.org/10.1039/D0LC00261E
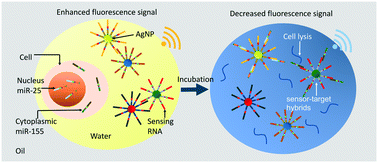
Nanoplasmon-enhanced drop-screen for high throughput single-cell nucleocytoplasmic miRNA profiling
Nanoplasmon-enhanced drop screen is developed to measure single-cell multiple miRNAs with high sensitivity of 0.1 nM, addressing cell nucleocytoplasmic profile in a throughput ∼100 cells per minute.
Lab Chip, 2020,20, 1939-1946
https://doi.org/10.1039/C9LC01226E
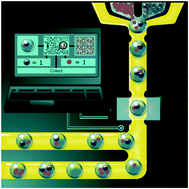
Deep learning guided image-based droplet sorting for on-demand selection and analysis of single cells and 3D cell cultures
To uncover the heterogeneity of cellular populations and multicellular constructs we show on-demand isolation of single mammalian cells and 3D cell cultures by coupling bright-field microdroplet imaging with real-time classification and sorting using convolutional neural networks.
Lab Chip, 2020,20, 889-900
https://doi.org/10.1039/D0LC00055H
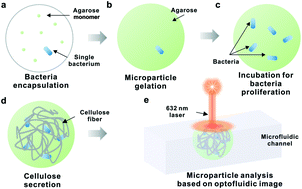
Intelligent optofluidic analysis for ultrafast single bacterium profiling of cellulose production and morphology
A continuous-flow intelligent optofluidic device using a convolutional neural network (CNN) computational method was developed to enable high-throughput single-bacterium profiling of bacteria cellulose (BC) with a throughput of ∼35 bacteria per second.
Lab Chip, 2020,20, 626-633
https://doi.org/10.1039/C9LC01105F
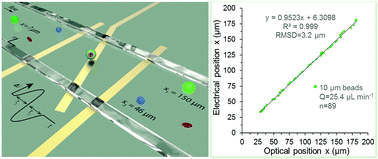
Microfluidic impedance cytometry device with N-shaped electrodes for lateral position measurement of single cells/particles
In this paper, we present an N-shaped electrode-based microfluidic impedance cytometry for the measurement of the lateral position of single cells and particles in continuous flows.
Lab Chip, 2019,19, 3609-3617
https://doi.org/10.1039/C9LC00819E
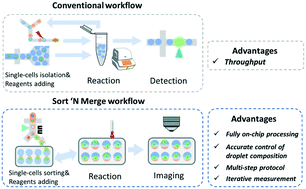
Single-cell RT-LAMP mRNA detection by integrated droplet sorting and merging
We present a droplet-based microfluidic platform that permits seamless on-chip droplet sorting and merging, which enables completing multi-step reaction assays within a short time, and demonstrate detection of specific single-cell mRNA expressions.
Lab Chip, 2019,19, 2425-2434
https://doi.org/10.1039/C9LC00161A
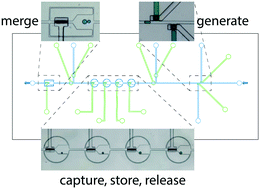
Microfluidic on-demand droplet generation, storage, retrieval, and merging for single-cell pairing
A multifunctional microfluidic platform combining on-demand aqueous-phase droplet generation, multi-droplet storage, and controlled merging of droplets selected from a storage library in a single integrated microfluidic device is described.
Lab Chip, 2019,19, 493-502
https://doi.org/10.1039/C8LC01178H
About this collection
A collection of papers focussed on multimodal single cell analysis. There is an array of tools that have the potential to allow us to comprehensively and accurately characterize the cells involved in a biological process, and we are a step away from using these tools widely and efficiently to impact clinical care, but there are large obstacles we must break down first. The papers in this collection suggest cutting edge solutions to the obstacles that we face in the advancement of multimodal single cell analysis.