Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Concomitant polymorphism in oxime-bridged pyrazole–tetrazole energetic materials
Vikranth
Thaltiri
 a,
Richard J.
Staples
a,
Richard J.
Staples
 b and
Jean'ne M.
Shreeve
b and
Jean'ne M.
Shreeve
 *a
*a
aDepartment of Chemistry, University of Idaho, Moscow, Idaho 83844-2343, USA. E-mail: jshreeve@uidaho.edu
bDepartment of Chemistry, Michigan State University, East Lansing, Michigan 48824, USA
First published on 31st December 2025
Abstract
A one-step synthesis of an oxime-bridged nitropyrazole–tetrazole yields two concomitant polymorphs, planar 3-α and twisted 3-β. Their contrasting solid-state geometries give rise to distinct packing, π-delocalization, and thermal stability. Both exhibit RDX-like detonation performance, demonstrating oxime bridging as an effective strategy for thermally stable, safer high-performance energetic materials.
Polymorphism is a well-recognized phenomenon in energetic materials, arising when a single compound crystallizes into multiple packing arrangements with distinct intermolecular interactions.1–5 Such differences often translate into measurable variations in density, thermal stability, mechanical sensitivity, and detonation performance. Classical explosives including RDX, HMX, and FOX-7 exhibit several polymorphic forms in which subtle changes in layer stacking or molecular orientation led to markedly different energetic behavior.6–12 As illustrated in Fig. 1a, the α-, β-, and ε-phases of RDX highlight how packing diversity strongly governs macroscopic properties. Polymorph generation is typically influenced by crystallization parameters such as solvent, temperature, anti-solvent addition, pressure, and seeding. However, polymorph formation arising directly from intrinsic molecular geometry and directional intermolecular interactions, rather than deliberate crystallization control, remains uncommon and is often discovered only serendipitously. In such cases, the solid-state outcome is governed by how a given molecular geometry propagates into distinct packing motifs within the crystal lattice.
Pyrazole–tetrazole hybrids represent a compelling platform for high-energy-density materials owing to their nitrogen-rich frameworks, high heats of formation, and favorable heterocycle-driven packing characteristics.13–20 Previous studies have largely focused on modifying the bridging fragment connecting the two rings (Fig. 1b). C–C linkages typically confer moderate thermal stability yet remain relatively sensitive, whereas N-methylene and N–H bridges offer improved stability and safety.21–23 More recently, carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) bridging strategies have been shown to enhance thermal stability and reduce sensitivity in energetic heterocycles by promoting favourable intermolecular interactions and solid-state organization.24 Among these, N–H linkages have delivered the best balance of high energy and reduced sensitivity. Collectively, these findings underscore the pivotal role of the bridging unit in governing molecular geometry, intermolecular interactions, and solid-state organization, thereby dictating energetic performance.
O) bridging strategies have been shown to enhance thermal stability and reduce sensitivity in energetic heterocycles by promoting favourable intermolecular interactions and solid-state organization.24 Among these, N–H linkages have delivered the best balance of high energy and reduced sensitivity. Collectively, these findings underscore the pivotal role of the bridging unit in governing molecular geometry, intermolecular interactions, and solid-state organization, thereby dictating energetic performance.
The oxime bridge provides an additional design handle through its C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–OH functionality, which can adopt distinct configurational (E/Z) orientations about the C
N–OH functionality, which can adopt distinct configurational (E/Z) orientations about the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond, influencing intermolecular contacts and packing preferences. In densely functionalized heterocycles, even minor changes in planarity or torsion can translate into measurable differences in packing and thermal behavior. Oxime-containing heterocycles also benefit from strong O–H⋯N/O hydrogen bonding and partial C
N bond, influencing intermolecular contacts and packing preferences. In densely functionalized heterocycles, even minor changes in planarity or torsion can translate into measurable differences in packing and thermal behavior. Oxime-containing heterocycles also benefit from strong O–H⋯N/O hydrogen bonding and partial C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–OH conjugation, contributing to enhanced stability. Reported mono-oxime derivatives—including tetrazoles and oxadiazoles—exhibit high decomposition temperatures (Td = 265–288 °C) and are extremely insensitive (IS > 40 J, FS > 360 N), although their detonation performance is generally moderate (Fig. 1c).25,26 Despite these advances, the influence of oxime-linked molecular geometry and directional hydrogen bonding on solid-state organization within highly nitrated energetic frameworks has not been thoroughly examined. Motivated by this, we introduced an oxime bridge into a nitropyrazole–tetrazole scaffold to evaluate how the C
N–OH conjugation, contributing to enhanced stability. Reported mono-oxime derivatives—including tetrazoles and oxadiazoles—exhibit high decomposition temperatures (Td = 265–288 °C) and are extremely insensitive (IS > 40 J, FS > 360 N), although their detonation performance is generally moderate (Fig. 1c).25,26 Despite these advances, the influence of oxime-linked molecular geometry and directional hydrogen bonding on solid-state organization within highly nitrated energetic frameworks has not been thoroughly examined. Motivated by this, we introduced an oxime bridge into a nitropyrazole–tetrazole scaffold to evaluate how the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–OH bridging affects molecular geometry, intermolecular interactions, and thermal behaviour in an oxygen-balanced energetic system, while maintaining strong energetic performance.
N–OH bridging affects molecular geometry, intermolecular interactions, and thermal behaviour in an oxygen-balanced energetic system, while maintaining strong energetic performance.
Now we describe an oxime-bridged nitropyrazole–tetrazole system synthesized under ZnCl2-mediated conditions, which resulted in the formation of two concomitant polymorphs, 3-α and 3-β (Fig. 1d). The two solids exhibit identical molecular connectivity but distinct solid-state geometries arising from different E/Z orientations of the oxime group, which in turn generate different hydrogen-bonding networks and packing arrangements. Polymorph 3-α adopts a nearly planar arrangement that supports tighter packing, whereas 3-β displays a twisted solid-state geometry that reduces planarity and leads to more expanded lattice. These crystallographically distinct packing motifs provide a clear framework for assessing how oxime-orientation-dependent molecular geometry governs polymorphism and energetic properties within nitropyrazole–tetrazole frameworks.
The synthetic route to the oxime-bridged pyrazole–tetrazole and its concomitant polymorphs is outlined in Scheme 1. The sequence begins with (E)-N-hydroxy-3,4-dinitro-1H-pyrazole-5-carbimidoyl chloride (1), prepared based on a reported procedure.27 Nucleophilic substitution of the chloro group with sodium cyanide in a H2O/EtOAc biphasic system gave the nitrile intermediate (2) in 77% yield. Subsequent [3 + 2] cycloaddition of 2 with NaN3 in the presence of ZnCl2 directly generated the oxime-bridged pyrazole–tetrazole scaffold as two concomitant polymorphs. Upon acidification with 2 N HCl, the planar form 3-α precipitated from the reaction mixture, whereas the twisted solid-state geometry 3-β remained in the mother liquor and was obtained by ethyl acetate extraction. No recrystallization or solvent-induced transformation was required, confirming that both polymorphs originate directly from the reaction medium.
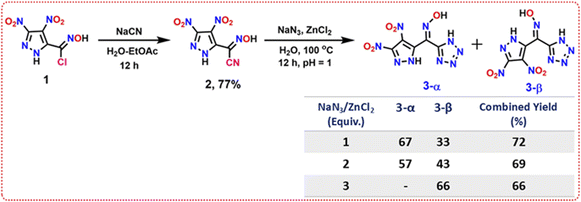 | ||
| Scheme 1 Synthesis of oxime-bridged pyrazole–tetrazole derivatives 3-α and 3-β, showing the effect of NaN3/ZnCl2 equivalents on polymorph distribution and combined yields. | ||
Varying the NaN3/ZnCl2 loading exerted a clear influence on the relative formation of the two polymorphs. With 1 equiv of NaN3/ZnCl2, both solid forms were obtained in 72% combined yield with a 3-α/3-β ratio of 67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 33. Increasing the loading to 2 equiv afforded a 57
33. Increasing the loading to 2 equiv afforded a 57![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 43 mixture in 69% combined yield, whereas 3 equiv furnished exclusively the β-form in 66% yield, with no detectable 3-α upon acidification. These results indicate that lower azide/Zn2+ loading preferentially nucleates the planar α-form, whereas higher loading shifts the product distribution toward the twisted β-form. All compounds were comprehensively characterized by multinuclear NMR (1H, 13C), IR spectroscopy, elemental analysis, and single-crystal X-ray diffraction.
43 mixture in 69% combined yield, whereas 3 equiv furnished exclusively the β-form in 66% yield, with no detectable 3-α upon acidification. These results indicate that lower azide/Zn2+ loading preferentially nucleates the planar α-form, whereas higher loading shifts the product distribution toward the twisted β-form. All compounds were comprehensively characterized by multinuclear NMR (1H, 13C), IR spectroscopy, elemental analysis, and single-crystal X-ray diffraction.
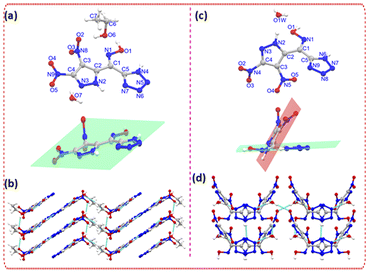
Single-crystal X-ray diffraction showed that 3-α crystallizes in the triclinic space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) with one molecule in the asymmetric unit, accompanied by lattice water and ethanol (Fig. 2). The molecular framework adopts a nearly coplanar arrangement across the pyrazole–oxime–tetrazole axis, with minimal torsional distortion. Representative dihedral angles—O1–N1–C1–C2 = −178.7°, O1–N1–C1–C5 = −0.4°, N1–C1–C2–N2 = −3.0°, and N3–N2–C3–C4 = −1.9°—confirm a planar, π-extended geometry. This alignment promotes efficient π-overlap and organizes the molecules into densely stacked sheets. The lattice is further stabilized by a network of directional hydrogen bonds, including O1–H1⋯O6 = 2.596 Å, N2–H2⋯N6 = 2.926 Å, and N4–H4⋯O7 = 2.647 Å, supplemented by interactions involving the co-crystallized solvent molecules. Collectively, these contacts generate a tightly packed structure with minimal free volume and strong intermolecular cohesion.
with one molecule in the asymmetric unit, accompanied by lattice water and ethanol (Fig. 2). The molecular framework adopts a nearly coplanar arrangement across the pyrazole–oxime–tetrazole axis, with minimal torsional distortion. Representative dihedral angles—O1–N1–C1–C2 = −178.7°, O1–N1–C1–C5 = −0.4°, N1–C1–C2–N2 = −3.0°, and N3–N2–C3–C4 = −1.9°—confirm a planar, π-extended geometry. This alignment promotes efficient π-overlap and organizes the molecules into densely stacked sheets. The lattice is further stabilized by a network of directional hydrogen bonds, including O1–H1⋯O6 = 2.596 Å, N2–H2⋯N6 = 2.926 Å, and N4–H4⋯O7 = 2.647 Å, supplemented by interactions involving the co-crystallized solvent molecules. Collectively, these contacts generate a tightly packed structure with minimal free volume and strong intermolecular cohesion.
In contrast, 3-β crystallizes in the orthorhombic Aea2 space group and adopts a distinctly twisted solid-state geometry. Several torsion angles deviate substantially from coplanarity—N1–C1–C2–N2 = −53.8°, N1–C1–C5–N6 = −15.6°, C2–C3–N5–O5 = 153.1°, and C5–C1–C2–C3 = −25.7°—indicating significant rotation within the heterocyclic fragments that disrupts π-conjugation across the oxime bridge. The solid-state arrangement consists of zig–zag molecular chains along the a-axis rather than planar stacks, and intermolecular stabilization is limited to a few weaker contacts such as O1–H1⋯O1W = 2.685 Å and N2–H2⋯O1W = 2.725 Å. The absence of extended hydrogen-bonding networks and the reduced face-to-face contact area result in a more open and less densely interconnected lattice.
Taken together, the two solid forms illustrate how the oxime bridge can impose distinct soli-state geometries within an identical molecular framework. The planar geometry of 3-α promotes extended conjugation, strong hydrogen-bonding networks, and dense packing, whereas the twisted geometry of 3-β disrupts π-alignment and generates a more-weakly connected, spatially expanded lattice. These contrasting structural features provide a clear basis for interpreting the subsequent differences in their thermal behavior and energetic properties.
The physicochemical and energetic properties of 3-α and 3-β are summarized in Table 1, with RDX included for reference. Density is a key determinant of detonation performance, as both detonation velocity (Dv) and detonation pressure (P) scale directly with ρ. Pycnometric measurements show that 3-α attains a high density of 1.831 g cm−3—slightly above that of RDX (1.80 g cm−3)—whereas 3-β reaches 1.802 g cm−3, essentially matching the benchmark material. Heats of formation calculated using isodesmic reactions (Gaussian 09)28 are highly positive for both polymorphs (ΔHf = 698.3 kJ mol−1 for 3-α and 710.3 kJ mol−1 for 3-β). Incorporation of these parameters into EXPLO5 (v7.01.01) produced RDX-class detonation performance: 3-α has Dv = 8739 m s−1 and P = 32.7 GPa, while 3-β has Dv = 8645 m s−1 and P = 31.9 GPa. Thus, both polymorphs exhibit strong energetic output despite their distinct solid-state geometries.
| Compound | 3-α | 3-β | RDXh |
|---|---|---|---|
| a Density determined by gas pycnometer at 25 °C. b Thermal decomposition temperature (5 °C min−1). c Heat of formation. d Detonation velocity. e Detonation pressure. f Impact sensitivity (BAM drophammer). g Friction sensitivity (BAM friction tester). h Ref. 29. | |||
| ρ (g cm−3) | 1.831 | 1.802 | 1.80 |
| T d (°C) | 241 | 219 | 204 |
| ΔHfc(kJ mol−1) | 698.3 | 710.3 | 92.6 |
| D v (m s−1) | 8739 | 8645 | 8795 |
| P (GPa) | 32.7 | 31.9 | 34.5 |
| ISf (J) | >40 J | >40 J | 7.4 |
| FSg (N) | >360 N | >360 N | 120 |
Thermal stability and sensitivity are critical parameters for assessing the practical suitability of energetic materials. Differential scanning calorimetry (5 °C min−1) shows that 3-α decomposes at 241 °C—substantially higher than RDX (204 °C)—while 3-β exhibits a still-elevated Td of 219 °C. Both values exceed those of previously reported C–C-, N-methylene-, and N–H-linked pyrazole–tetrazole systems, underscoring the stabilizing effect of the oxime bridge. These trends align with the distinct structural and electronic features of the two polymorphs: the extended π-conjugation and dense packing in 3-α support its superior thermal stability, whereas the torsionally distorted geometry of 3-β accounts for its slightly lower yet still enhanced performance relative to traditional linkages. Sensitivity data further highlight the excellent safety characteristics of both materials, with both polymorphs classified as insensitive energetic materials (IS: >40 J; FS: >360 N) according to BAM criteria—markedly safer than RDX (IS = 7.4 J; FS = 120 N). The combination of high thermal stability, exceptional insensitivity, and RDX-level detonation performance positions 3-α and 3-β as promising candidates for next-generation energetic materials with improved safety margins.
Hirshfeld surface analysis provides quantitative insight into the intermolecular contacts that define the solid-state organization of the two polymorphs.30 For 3-α, O⋯H/H⋯O (35.7%) and N⋯H/H⋯N (25.9%) interactions dominate, consistent with its extensive hydrogen-bonding network and densely packed lattice. In contrast, 3-β exhibits markedly lower contributions from O⋯H/H⋯O (22.4%) and N⋯H/H⋯N (18.4%) contacts, reflecting fewer strong intermolecular interactions and a more open packing arrangement. The corresponding 2D fingerprint plots reinforce this distinction: 3-α shows sharper, deeper spikes characteristic of short, well-defined contacts, whereas 3-β displays broader, more diffuse features. These quantitative trends are fully consistent with the crystallographic observations and underscore the structural differences between the planar and twisted forms (Fig. 3). The localized orbital locator (LOL-π) surfaces provide a clear visualization of the π-communication differences between the two polymorphs (Fig. 4).31 In 3-α, a continuous π-density pathway extends across the pyrazole–oxime–tetrazole backbone, consistent with its nearly planar geometry and uninterrupted conjugation. This electronic continuity aligns with the dense packing and strong intermolecular interactions observed crystallographically. In contrast, 3-β exhibits distinct interruptions in π-density at the oxime–tetrazole junction, matching the torsional distortions revealed in its solid-state structure. The disrupted π-framework limits through-bond and through-space overlap between the heterocycles and rationalizes the weaker packing interactions in the twisted form.
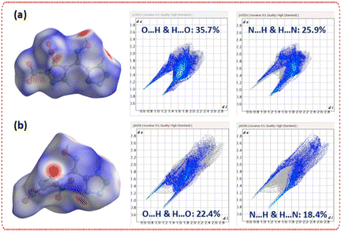 | ||
| Fig. 3 (a) Hirshfeld surface of polymorph 3-α with corresponding 2D fingerprint plots. (b) Hirshfeld surface of polymorph 3-β with corresponding 2D fingerprint plots. | ||
 | ||
| Fig. 4 Localized orbital locator (LOL-π) isosurfaces for polymorphs 3-α (a) and 3-β (b), illustrating continuous versus disrupted π-electron pathways. | ||
Electrostatic potential (ESP) mapping further highlights how molecular geometry dictates charge distribution (Fig. 5).32 The planar 3-α displays a smoother potential along its conjugated backbone, with ESP values ranging from +80.80 to −33.68 kcal mol−1, indicative of more even charge delocalization. In contrast, 3-β shows a broader ESP range (+82.31 to −35.30 kcal mol−1) and more localized high-intensity regions, reflecting charge accumulation around its bent junctions and reduced delocalization. Frontier molecular orbital analysis reinforces these trends. In 3-α, the HOMO and LUMO are distributed continuously across the heterocyclic chain (−8.112 eV and −3.653 eV; ΔE = 4.459 eV), consistent with its extended π-system. For 3-β, the HOMO and LUMO (−8.35 eV and −3.66 eV; ΔE = 4.69 eV) appear more fragmented, in line with its twisted geometry and interrupted π-communication. Together, the ESP and FMO analyses corroborate the intrinsic electronic distinctions between the two polymorphs and rationalize their divergent solid-state behavior.
Conclusions
In conclusion, the oxime bridge emerges as a versatile structural element capable of stabilizing distinct solid-state arrangements within a single pyrazole–tetrazole energetic framework. A one-step transformation unexpectedly yielded two concomitant polymorphs—3-α and 3-β—that differ markedly in solid-state geometry, intermolecular interactions, and electronic structure despite identical connectivity. Structural analyses (SCXRD, Hirshfeld, LOL-π) show that 3-α adopts a planar, π-delocalized geometry supported by strong directional contacts and dense packing, whereas 3-β features significant torsional distortion, disrupted π-communication, and reduced intermolecular cohesion. ESP mapping and frontier-orbital analysis further indicate smoother charge distribution and greater electronic coherence in 3-α relative to the more localized features observed for 3-β. Overall, these results demonstrate that subtle oxime-driven differences in solid-state geometry can directly tune π-delocalization, packing efficiency, and solid-state behaviour, providing valuable design principles for developing next-generation, thermally stable, insensitive, and safer high-energy heterocyclic materials.Author contributions
V. T. investigation, methodology, conceptualization and manuscript writing. R. J. S. X-ray data collection and structures solving. V. T. and J. M. S. conceptualization, manuscript writing – review and editing, supervision.Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting findings of this study are available within the article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ta09791f.CCDC 2511981 and 2511983 contain the supplementary crystallographic data for this paper.33
Acknowledgements
The Rigaku Synergy S Diffractometer was purchased with support from the National Science Foundation MRI program (1919565). We are grateful for the support of the Fluorine-19 fund.References
- J.-P. Brog, C.-L. Chanez, A. Crochet and K. M. Fromm, RSC Adv., 2013, 3, 16905 RSC.
- A. J. Cruz-Cabeza, S. M. Reutzel-Edens and J. Bernstein, Chem. Soc. Rev., 2015, 44, 8619 RSC.
- J. Bernstein, Polymorphism in Molecular Crystals, Oxford University Press, Oxford, 2002 Search PubMed.
- A. J. Cruz-Cabeza and J. Bernstein, Chem. Rev., 2014, 114, 2170 CrossRef CAS PubMed.
- F. Sánchez-Férez, X. Solans-Monfort, T. Calvet, M. Font-Bardia and J. Pons, Inorg. Chem., 2022, 61, 4965 Search PubMed.
- J. Bernstein, Polymorphism of High Energy Materials, Polymorphism in Molecular Crystals, IUCr Monographs on Crystallography, Oxford University Press, New York, 2002, ch. 9 Search PubMed.
- J. P. Agrawal, High Energy Materials: Propellants, Explosives and Pyrotechnics, Wiley-VCH, Weinheim, 2010 Search PubMed.
- G. Liu, R. Gou, H. Li and C. Zhang, Cryst. Growth Des., 2018, 18, 4174 CrossRef CAS.
- J. Chang, Q. Sun, N. Ding, Y. Jiang, N. Yao, X. Xu, B. Tian, S. Li and S. Pang, Org. Lett., 2015, 17, 1074 CrossRef.
- I. G. Goldberg and J. A. Swift, Cryst. Growth Des., 2012, 12, 1040 CrossRef CAS.
- E. D. Emmons, M. E. Farrell, E. L. Holthoff, A. Tripathi, N. Green, R. P. Moon, J. A. Guicheteau, S. D. Christesen, P. M. Pellegrino and A. W. Fountain III, Appl. Spectrosc., 2012, 66, 628 CrossRef CAS.
- R. Infante-Castillo, L. C. Pacheco-Londoño and S. P. Hernández-Rivera, J. Mol. Struct., 2010, 970, 51 CrossRef CAS.
- T. M. Klapötke, Chemistry of High-Energy Materials, 6th edn, de Gruyter, Berlin, 2022 Search PubMed.
- D. M. Badgujar, M. B. Talawar, S. N. Asthana and P. P. Mahulikar, J. Hazard. Mater., 2008, 151, 289 CrossRef CAS.
- P. F. Pagoria, G. S. Lee, A. R. Mitchell and R. D. Schmidt, Thermochim. Acta, 2002, 384, 187 CrossRef CAS.
- M. B. Talawar, R. Sivabalan, T. Mukundan, H. Muthurajan, A. K. Sikder, B. R. Gandhe and A. S. Rao, J. Hazard. Mater., 2009, 161, 589 Search PubMed.
- P. Yin and J. M. Shreeve, Adv. Heterocycl. Chem., 2017, 121, 89 Search PubMed.
- S. Zhang, Z. Gao, D. Lan, Q. Jia, N. Liu, J. Zhang and K. Kou, Molecules, 2020, 25, 3475 CrossRef CAS.
- B. Chen, H. Lu, J. Chen, Z. Chen, S.-F. Yin, L. Peng and R. Qiu, Top. Curr. Chem., 2023, 381, 25 CrossRef CAS PubMed.
- J. Liu, M. Liao, J. Tang, W. Huang, G. Cheng, Y. Liu, C. Xiao and Y. Tang, Org. Lett., 2025, 27, 12851 CrossRef CAS.
- V. Thaltiri, R. J. Staples and J. M. Shreeve, J. Mater. Chem. A, 2024, 12, 16729 RSC.
- D. Kumar, G. H. Imler, D. A. Parrish and J. M. Shreeve, Chem.–Eur. J., 2017, 23, 7876 CrossRef CAS.
- V. Thaltiri, R. J. Staples and J. M. Shreeve, ACS Appl. Mater. Interfaces, 2025, 17, 38122 CrossRef PubMed.
- Y. Yang, W. Zhang, H. Huang and C. Sun, ACS Appl. Mater. Interfaces, 2025, 17, 66716 CrossRef CAS.
- D. Chand, D. A. Parrish and J. M. Shreeve, J. Mater. Chem. A, 2013, 1, 15383 RSC.
- P. Saini, J. Singh, V. Thaltiri, R. J. Staples and J. M. Shreeve, Mater. Adv., 2025, 6, 3338 RSC.
- V. Thaltiri, R. J. Staples and J. M. Shreeve, J. Org. Chem., 2024, 89, 18612 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, T. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian 09, Revision E.01, Gaussian, Inc., Wallingford, CT, 2013 Search PubMed.
- V. Thaltiri, K. Chavva, B. S. Kumar and P. K. Panda, New J. Chem., 2019, 43, 12318 RSC.
- M. Wolff, D. J. Grimwood, J. J. McKinnon, M. J. Turner, D. Jayatilaka and M. A. Spackman, CrystalExplorer 17.5, 2012 Search PubMed.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580 CrossRef CAS PubMed.
- J. Zhang and T. Lu, Phys. Chem. Chem. Phys., 2021, 23, 20323 RSC.
- (a) CCDC 2511981: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2q9xnm; (b) CCDC 2511983: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2q9xqp.
| This journal is © The Royal Society of Chemistry 2026 |