Open Access Article
Open Access ArticleChemical activation of native cytochrome P450s in soil-derived bacteria by external molecules enables biodegradation of aromatic pollutants
Fumiya
Ito
,
Masayuki
Karasawa
 and
Osami
Shoji
and
Osami
Shoji
 *
*
Department of Chemistry, School of Science, Nagoya University, Furo-cho, Chikusa-ku, Nagoya, Aichi 464-0802, Japan. E-mail: shoji.osami.w3@f.mail.nagoya-u.ac.jp
First published on 9th March 2026
Abstract
The use of genetically engineered microorganisms for pollutant degradation is strictly regulated, limiting their deployment in open environments. To address this challenge, we introduce a non-genetic strategy that activates native soil bacteria to degrade otherwise inert aromatic pollutants. This approach employs small molecules, termed decoy molecules, which mimic native ligands and bind within the active-site cavity of cytochrome P450 enzymes. By partially occupying the substrate-binding pocket, they redirect the enzyme's catalytic activity toward non-native substrates. Using whole-cell biotransformation assays with Priestia megaterium JCM 2506T (CYP102A1) and Bacillus subtilis JCM 1465T (CYP102A3), we show that decoy molecules enable hydroxylation of benzene, toluene, xylenes, naphthalene, and halobenzenes. Remarkably, in the presence of decoy molecules, B. subtilis achieved complete degradation of 2-chlorinated dibenzo-p-dioxin within 2 hours at 45 °C. These findings demonstrate an externally controllable, non-genetic means of repurposing native soil bacteria as biocatalysts, offering a promising basis for environmentally compatible bioremediation strategies.
1. Introduction
Aromatic hydrocarbons—including benzene, toluene, xylenes (BTX), and naphthalene (a polycyclic aromatic hydrocarbon)—and persistent halogenated compounds, such as dioxins, are major organic pollutants in soil.1–4 These compounds are highly stable under typical environmental conditions, showing strong resistance to microbial and chemical degradation, which results in bioaccumulation and pronounced toxicity to living organisms.5–8 Consequently, they represent a significant challenge for soil remediation. Reported soil concentrations at contaminated sites vary widely: benzene, toluene, xylene, and naphthalene are commonly detected at levels ranging from below 1 ppm to several hundred ppm, and in some cases exceeding 1000 ppm.9–11 Dioxin concentrations also exhibit extreme heterogeneity, with reported levels in contaminated soils and sediments spanning more than six orders of magnitude, from below 1 ppt TEQ (total toxic equivalency, a weighted measure of PCDD/Fs and dioxin-like PCBs) upward.12–16 The use of microbial approaches to biodegrade such persistent and recalcitrant aromatic pollutants offers a promising and sustainable strategy for environmental restoration.17,18 Although a number of microorganisms that inhabit polluted environments have been reported to degrade these compounds,19–30 the introduction of non-indigenous microorganisms into polluted sites (bioaugmentation) often fails to achieve effective degradation under natural environmental conditions.31–34Microbial enzymes involved in the degradation of aromatic hydrocarbons—such as dioxygenases (e.g., naphthalene dioxygenase) and monooxygenases (e.g., cytochrome P450)—have been extensively studied for potential use in bioremediation. These include both naturally occurring enzymes35–40 and artificially engineered variants obtained through site-directed mutagenesis41–45 or directed evolution46–48 to enhance their catalytic activity toward such compounds. Since hydroxylation of these molecules serves as a key initial step that increases their aqueous solubility and facilitates subsequent microbial degradation,49,50 several of these enzymes have been utilized in various recombinant host strains to degrade environmental contaminants.51 However, the use of genetically engineered microorganisms (GEMs) or transgenic bacteria in open environments faces strict regulatory hurdles due to potential risks of genetic pollution.52–57 These microorganisms cannot be used in open environments unless they are equipped with robust intrinsic biocontainment mechanisms,58–60 and their application is consequently limited.
Recent studies have shown that even advanced biocontainment systems can exhibit substantial performance loss under realistic environmental conditions. For example, a GEM with CRISPR-based kill switch displayed markedly elevated escape rates in natural surface waters compared with standard laboratory media, a trend potentially linked to pH-dependent inducer speciation and nutrient limitation.61 In parallel, horizontal gene transfer (HGT) has been explored as an alternative means to support bioremediation, as hydrocarbon-degrading plasmids can be transferred to indigenous bacteria and sustain pollutant degradation even after the donor strain disappears.62 However, the environmental fate of such transferred genes remains insufficiently understood; emerging evidence shows that the fitness effects and persistence of acquired traits differ substantially across environmental contexts, making HGT outcomes difficult to predict in open environments.63 Together, these limitations highlight the need for alternative strategies that do not rely on genetically engineered organisms. If the substrate specificity of endogenous enzymes in ubiquitous, naturally occurring bacteria could be altered without genetic manipulation to enable the degradation of environmental pollutants, such bacteria could be broadly applied for bioremediation in open environments without being subject to the regulatory constraints of the Cartagena Protocol on Biosafety.
We have previously demonstrated that the substrate specificity of a heme enzyme isolated from Priestia megaterium (formerly known as Bacillus megaterium64), cytochrome P450BM3 (P450BM3, CYP102A1), can be drastically altered by the addition of dummy substrates referred to as “decoy molecules”. P450BM3 is an NADPH-dependent monooxygenase that catalyzes the hydroxylation of subterminal C–H bonds in long-chain fatty acids with a high turnover rate.65–67 While it typically shows no activity toward substrates other than long-chain fatty acids, P450BM3 efficiently hydroxylates aromatic hydrocarbons and gaseous alkanes in the presence of decoy molecules.68–76 Decoy molecules possess shorter alkyl chains than natural substrates and, therefore, can position their molecular termini away from the active site upon binding to the enzyme (Fig. 1). The remaining small space serves as a surrogate binding pocket for smaller organic molecules such as benzene, thereby promoting the hydroxylation of non-native substrates. Importantly, the decoy molecule itself is not oxidized during the reaction and acts catalytically rather than being consumed. Unlike conventional mutagenesis-based approaches,42–46,77–80 this methodology does not require any modification to the amino acid sequence of wild-type CYP102A1; instead, it relies on a transient, decoy-induced activation mechanism.
We focused on the fact that CYP102A1 originates from P. megaterium, a ubiquitous and easily culturable Gram-positive soil bacterium widely used in both research and industrial applications owing to its genetic accessibility and lack of alkaline proteases and endotoxins.81–86 If we can alter the substrate specificity of endogenous CYP102A1 in P. megaterium by exploiting decoy molecules to hydroxylate BTX as well as dioxins, we could convert the common soil bacterium P. megaterium into a microorganism capable of degrading environmental pollutants. In this study, we first investigated whether the decoy molecules can also induce oxidation reactions of non-native substrates in the original host of the CYP102A1 gene, P. megaterium. Furthermore, in addition to CYP102A1, we examined other CYP102 family enzymes. Since many CYP102 family members have been identified, we selected nine readily available soil bacterial strains with CYP102 enzymes to test the applicability of the decoy molecule approach and to assess their degradation activities toward aromatic pollutants (Table 1). Herein, we report that decoy molecules can repurpose ubiquitous soil bacteria harboring CYP102 enzymes for the degradation of aromatic compounds. This approach establishes a fundamentally new bioremediation paradigm in which externally supplied small molecules reprogram the substrate specificity of endogenous enzymes. Unlike conventional strategies that require genetic engineering or engineered strains, this small-molecule-mediated modulation strategy activates latent catalytic functions in native bacteria and enables the use of readily available biological resources for pollutant degradation. By circumventing the biosafety and regulatory constraints associated with genetic modification, this strategy provides a viable foundation for developing safer and more scalable pollutant-removal technologies.
| Species | JCM | CYP102 | Decoy molecule | GC yield of phenol | |
|---|---|---|---|---|---|
| µM | % | ||||
| a GC yield of phenol was calculated as [phenol]/[initial benzene] × 100. The values represent the mean of three experiments (±standard deviation). Reaction conditions: bacterial cell density (OD600 = 6.3), decoy molecule (100 µM), benzene (10 mM), glucose (40 mM), at 25 °C for 4 h in phosphate buffer (pH 7.4). | |||||
| Priestia megaterium | 2506T | A1 | None | <1 | <0.1 |
| C6-Pro-Phe | 197 ± 8 | 2.0 | |||
| C7-Pro-Phe | 216 ± 6 | 2.2 | |||
| (S)-Ibu-Gly | 159 ± 1 | 1.6 | |||
| (S)-Ibu-Ala | 220 ± 6 | 2.2 | |||
| (S)-Ibu-Leu | 187 ± 12 | 1.9 | |||
| (S)-Ibu-Ile | 174 ± 14 | 1.7 | |||
| (S)-Ibu-Phe | 151 ± 3 | 1.5 | |||
| (R)-Ibu-Phe | 134 ± 3 | 1.3 | |||
| Bacillus subtilis subsp. subtilis | 1465T | A2, A3 | None | <1 | <0.1 |
| C7-Pro-Phe | 2 ± 0 | <0.1 | |||
| (S)-Ibu-Gly | 10 ± 1 | 0.1 | |||
| (S)-Ibu-Ala | 50 ± 1 | 0.5 | |||
| (S)-Ibu-Leu | 103 ± 2 | 1.0 | |||
| (S)-Ibu-Ile | 81 ± 11 | 0.8 | |||
| (S)-Ibu-Val | 51 ± 2 | 0.5 | |||
| (S)-Ibu-Phe | 76 ± 3 | 0.8 | |||
| (R)-Ibu-Phe | 18 ± 2 | 0.2 | |||
| PFC7-Trp | 20 ± 1 | 0.2 | |||
| Bacillus cereus | 2152T | A5 | — | — | — |
| Bacillus licheniformis | 2505T | A7 | None | <1 | <0.1 |
| PFC9-Ala | 25 ± 1 | 0.3 | |||
| PFC9-Met | 9 ± 0 | <0.1 | |||
| (S)-Ibu-Phe | 10 ± 0 | 0.1 | |||
| Bacillus thuringiensis | 20386T | A8 | — | — | — |
| Bacillus pumilus | 2508T | A15 | — | — | — |
| Streptomyces avermitilis | 5070T | B2, D1 | — | — | — |
| Rhodococcus erythropolis | 3201T | C2 | None | 4 ± 2 | <0.1 |
| C10-Phe | 12 ± 3 | 0.1 | |||
| Actinosynnema pretiosum subsp. auranticum | 7343T | F1 | — | — | — |
| Saccharopolyspora erythraea | 4748T | G2 | — | — | — |
2. Experimental
2.1. Chemicals and bacterial strains
All chemicals were purchased from commercial sources (e.g., FUJIFILM Wako Pure Chemical Corporation, Tokyo Chemical Industry, Sigma-Aldrich, Kanto Chemical) and used without further purification unless otherwise stated. Mono-chlorinated dibenzo-p-dioxins (CDDs) were purchased from AccuStandard Inc. (New Haven, CT, USA). Dibenzo-p-dioxin (DD) and BSTFA-TMCS (99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) were purchased from Tokyo Chemical Industry Co., Ltd (Tokyo, Japan). Decoy molecules used in this study were synthesized in a similar manner to that previously described.71,72,74 All bacterial type strains were provided by Japan Collection of Microorganisms, RIKEN BRC which is participating in the National Bio-Resource Project of the MEXT, Japan. Bacillus subtilis strain 168 (MGNA-A001) and its gene-knockout mutants87 (BKE07250 and BKE27160) were provided by National BioResource Project (NIG, Japan): B. subtilis.
1) were purchased from Tokyo Chemical Industry Co., Ltd (Tokyo, Japan). Decoy molecules used in this study were synthesized in a similar manner to that previously described.71,72,74 All bacterial type strains were provided by Japan Collection of Microorganisms, RIKEN BRC which is participating in the National Bio-Resource Project of the MEXT, Japan. Bacillus subtilis strain 168 (MGNA-A001) and its gene-knockout mutants87 (BKE07250 and BKE27160) were provided by National BioResource Project (NIG, Japan): B. subtilis.
2.2. Cell preparation for whole-cell hydroxylation
All bacterial strains were inoculated into appropriate growth media in Erlenmeyer flasks. Cultures were incubated at their optimal growth temperatures with shaking for the respective durations (Table S1). After incubation, the cells were harvested and resuspended in phosphate buffer (87 mM Na2HPO4, 16 mM KH2PO4, 7 mM MgSO4, 86 mM NaCl, 0.1 mM CaCl2, pH 7.4), which served as a solvent for the subsequent hydroxylation reactions. Cell density was adjusted based on optical density at 600 nm (OD600), except for Streptomyces avermitilis JCM 5070T, Actinosynnema pretiosum subsp. auranticum JCM 7343T, and Saccharopolyspora erythraea JCM 4748T for which wet cell weight (WCW) was used instead.2.3. Screening of bacterium-decoy molecule combinations
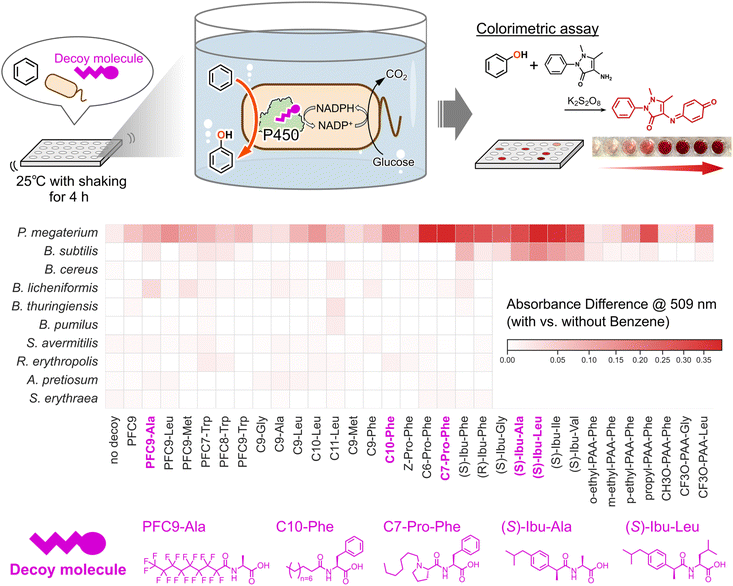
Benzene hydroxylation reactions were performed in 96-well microplates with a total reaction volume of 200 µL. Each reaction mixture contained bacterial cells (OD600 = 6.3 or WCW = 22.5 g L−1), decoy molecule (100 µM), glucose (40 mM), DMSO (1.5% [v/v]), and benzene (10 mM) in the phosphate buffer. Glucose was included in the screening medium to ensure sufficient NADPH supply for P450-catalyzed hydroxylation, which proceeds efficiently during active cellular metabolism. Since the hydroxylation reaction was performed using non-growing (static) cells harvested after cultivation, any potential effects of the decoy molecule on cell growth would not influence the assay outcome. In addition, DMSO was used to dissolve the substrate and decoy molecules, and the same final concentration of DMSO was included in all control experiments. After incubation for 4 h at 25 °C on a plate shaker (Biomixer PMA-001, Biotec), the amount of phenol in the supernatants was determined by a colorimetric assay with 4-aminoantipyrine (Fig. 2).74,75,88,89 Absorbance was measured using a TECAN Infinite M200 PRO Multimode Microplate Reader (Tecan Ltd, Männedorf, Switzerland). The capability of benzene hydroxylation was assessed using two distinct negative controls: one lacking decoy molecules, and another lacking benzene. The latter control was included to rule out false positives arising from unintended reactions, since 4-aminoantipyrine also detects phenolic compounds other than phenol. A heatmap was created using the absorbance differences between the benzene-present and benzene-absent conditions. The 76 decoy molecules used cover all three generations reported before; They are perfluorocarboxylic acids (PFCs),70N-perfluoroacyl amino acids (PFC-AAs),71 and amino acids or dipeptides with various N-substituent,72 respectively (Fig. S1).2.4. Quantification of whole-cell reactions
Reactions were carried out in 6 mL vials with a total reaction volume of 1 mL. After completion, reactions were quenched by adding dichloromethane for GC-MS analysis or acetonitrile for high performance liquid chromatography (HPLC) analysis. For benzene, toluene, xylenes, fluorobenzene, and chlorobenzene, hydroxylated products were quantified by GC-MS, whereas HPLC analysis was conducted for naphthalene and dibenzo-p-dioxin derivatives (DD and CDDs). Detailed GC-MS and HPLC analytical conditions are provided in the SI.2.5. Cloning and overexpression of P450s in E. coli
The CYP102A1 gene from P. megaterium JCM 2506T and the CYP102A2 and CYP102A3 genes from B. subtilis JCM 1465T were individually cloned into the pET28a(+) vector. Each constructed vector was transformed into E. coli BL21(DE3). To induce enzyme expression, 5-aminolevulinic acid (0.5 mM) and isopropyl-β-D-1-thiogalactopyranoside (IPTG, 0.1 mM) were added during culture incubation.2.6. Modeling of protein–ligand complexes
For CYP102A2 (UniProt ID: O08394) and CYP102A3(UniProt ID: O08336), modeled structures using AlphaFold2 (ref. 90) were downloaded from AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk/), followed by truncation to the heme domains. Since AlphaFold2 models do not include cofactors, the heme b was subsequently inserted via structural alignment with a crystal structure of CYP102A1 (PDB: 3WSP).In order to refine the structures, three independent 50 ns MD simulations were performed for each protein using GROMACS 2024.2.91 For each protein, the most average-like structure was extracted from the combined last 40 ns of all three trajectories. The substrate (1- or 2-CDD) and the decoy molecule ((S)-Ibu-Leu) were docked stepwise to the extracted structures using AutoDock Vina v1.1.2.92 Finally, the top-ranked docking poses were analyzed using the Protein–Ligand Interaction Profiler (PLIP, v2.3.1).93
3. Results and discussion
3.1. Repurposing of natural soil bacteria as hydroxylation catalysts
Whole-cell hydroxylation of benzene was performed by adding benzene to the suspension of the native bacterial cells in the presence of each of the 76 decoy molecules, followed by stirring at 25 °C for 4 h. The reaction was initially evaluated by a colorimetric assay to detect phenol formation, and samples showing detectable activity were subsequently analyzed by GC-MS for accurate quantification of phenol. The structures and activities of representative decoy molecules are summarized in Fig. 2, while those of all 76 tested decoy molecules are provided in the SI (Fig. S1 and S2). The type strain P. megaterium JCM 2506T alone exhibited no detectable benzene-hydroxylating activity, indicating that it cannot be directly applied to the bioremediation of aromatic pollutants (Table 1). In contrast, the addition of decoy molecule, such as C7-Pro-Phe, to the reaction mixture resulted in clear benzene hydroxylation, indicating that C7-Pro-Phe activates endogenous CYP102A1 to catalyze this reaction through a substrate-misrecognition mechanism previously reported to be induced by decoy molecule binding. Based on the screening, several decoy molecules were found to be effective in promoting benzene hydroxylation by P. megaterium. Among them, decoy molecules such as (S)-Ibuprofen-fused amino acids ((S)-Ibu-AAs) and Cn-Pro-Phe (n = 6, 7) were particularly effective in accelerating the reaction. The observed preference for these decoy molecules is consistent with that of purified CYP102A1,72 suggesting that benzene hydroxylation is catalyzed by CYP102A1 in P. megaterium. Phenol formation in the presence of the most effective decoy molecule, (S)-Ibu-Ala, reached 220 ± 6 µM (2.2% yield). Encouraged by the finding that even the native bacterium P. megaterium became capable of hydroxylating benzene, which is normally an inert substrate toward oxidation, simply by the addition of decoy molecule, we envisioned that other native bacteria possessing CYP102 family enzymes might also acquire benzene-hydroxylating activity if appropriate decoy molecules could interact with their intracellular CYP102 family enzymes, thereby repurposing these bacteria as potential bioremediation agents. We selected nine representative bacterial type strains, each harboring at least one innate CYP102 enzyme highly similar to CYP102A1 (Table S2), and performed screening of decoy molecules. As expected, the addition of decoy molecules enabled several bacteria to hydroxylate benzene.B. subtilis was demonstrated to hydroxylate benzene in response to the addition of several decoy molecules, whereas Bacillus licheniformis and Rhodococcus erythropolis showed weak but measurable benzene-hydroxylating activity in the presence of some decoy molecules as determined by the colorimetric assay (Fig. 2 and Table 1). B. subtilis produced 103 ± 2 µM phenol in the presence of (S)-Ibu-Leu, whereas B. licheniformis and R. erythropolis produced 25 ± 1 µM and 12 ± 3 µM phenol, respectively, in the presence of PFC9-Ala and C10-Phe as decoy molecules. The variation in the decoy molecules that yielded the highest activity among different bacterial strains is likely attributable to the preferences of the P450 enzymes for specific decoy molecules in each strain. Indeed, the correlation of decoy molecule dependence for benzene hydroxylation observed with E. coli BL21(DE3) overexpressing CYP102A1 as whole-cell biocatalysts was similar to that of P. megaterium (Fig. 3a, S3, and Table S3). Because B. subtilis possesses CYP102A2 and CYP102A3 (Table S4), benzene hydroxylation was examined using E. coli strains overexpressing CYP102A2 and CYP102A3, respectively. CYP102A2 exhibited almost no activity, whereas CYP102A3 showed a marked increase in activity upon addition of decoy molecules, and its dependence on decoy molecules closely matched that observed for B. subtilis. These results suggest that CYP102A3 is primarily responsible for the benzene hydroxylation activity in B. subtilis whereas CYP102A2 may also contribute slightly. We confirmed that the B. subtilis strain lacking the CYP102A3 gene (B. subtilis 168 Δcyp102a3) showed a drastic decrease in benzene-hydroxylating activity (Fig. S4 and S5, also see footnote†). In the case of B. licheniformis, the highest activity was observed with PFC9-Ala, which is consistent with the fact that the purified CYP102A7 showed high activity with decoy molecules bearing perfluoroalkyl groups.94 Since P. megaterium and B. subtilis exhibited higher benzene-hydroxylating activity in the presence of decoy molecules than the other bacterial strains tested, we decided to further investigate the potential of these two strains for the biodegradation of environmental pollutants. As part of optimizing the reaction conditions, we examined the effect of reaction temperature on activity. At 30 °C, P. megaterium showed higher activity (3.2%), whereas at 45 °C, B. subtilis exhibited higher activity (5.2%) (Fig. 3b). Based on these results, subsequent reactions were carried out and evaluated at 30 °C for P. megaterium and 45 °C for B. subtilis.
3.2. Evaluation of the reaction system toward aromatic pollutants
To examine whether P. megaterium and B. subtilis were capable of degrading typical environmental pollutants, hydroxylation reactions of toluene and xylenes (the BTX components other than benzene), fluorobenzene and chlorobenzene (halogenated aromatic hydrocarbons), and naphthalene (a polycyclic aromatic hydrocarbon) were conducted under the reaction conditions that showed high activity toward benzene, with substrate and decoy molecule concentrations of 1 mM and 100 µM, respectively. The yields of the corresponding hydroxylated products were quantified by GC-MS or HPLC (Table 2). The hydroxylation of toluene and xylenes proceeded in both P. megaterium and B. subtilis in the presence of decoy molecules with total product yields of 20–40% after 4 hours of reaction, indicating that both strains possess sufficient activity to degrade these compounds. In the hydroxylation of o-xylene by P. megaterium, the product distribution varied differed between the two decoy molecules examined; C7-Pro-Phe promoted benzylic hydroxylation more strongly than (S)-Ibu-Ala. Previous computational studies have shown that the size and shape of the reactive cavity in CYP102A1, as well as the intrinsic reactivity of individual positions on xylene, can influence site selectivity.95 In this context, variation in decoy molecule structure may alter the binding microenvironment experienced by the substrate, which could partly account for the decoy-dependent changes in regioselectivity observed in this study. Although these reactions were carried out in the phosphate buffer, the same reactions are feasible in growth media (Fig. S6). Interestingly, when benzene, toluene, and o-xylene were added simultaneously, hydroxylated products derived from all of them were detected, indicating a promiscuous substrate recognition that leads to broad substrate versatility (Fig. S7).| Substrate | Product | P. megaterium | B. subtilis | ||||
|---|---|---|---|---|---|---|---|
| (S)-Ibu-Ala | C7-Pro-Phe | (S)-Ibu-Leu | |||||
| Sel.a [%] | T. yieldb [%] | Sel. [%] | T. yield [%] | Sel. [%] | T. yield [%] | ||
| a Sel. denotes the regioselectivity of hydroxylation, calculated as the yield of the specified hydroxylated product divided by the total yield of all hydroxylated products, based on GC or HPLC quantification using external calibration curves. b T. yield indicates total GC or HPLC yields calculated as [all hydroxylated products]/[initial substrate] × 100 and represent the mean of three experiments (± standard deviation). Reaction conditions: bacterial cell density (OD600 = 6.3), decoy molecule (100 µM), substrate (1 mM), glucose (40 mM), at 30 °C (for P. megaterium) or 45 °C (for B. subtilis) for 4 h in phosphate buffer (pH 7.4). All products were detected by GC-MS analysis except for naphthalene, for which HPLC analysis was used. | |||||||
| Benzene | Phenol | 9.1 ± 0.9 | 18 ± 3 | 4.1 ± 0.2 | |||
| Toluene | Benzyl alcohol | 1.6 | 28 ± 1 | 4.8 | 28 ± 0 | 6.3 | 30 ± 3 |
| o-Cresol | 97 | 93 | 92 | ||||
| p-Cresol | 1.2 | 2.2 | 1.6 | ||||
| o-Xylene | 2-Methylbenzyl alcohol | 53 | 24 ± 2 | 71 | 22 ± 0 | 88 | 36 ± 3 |
| 2,3-Dimethylphenol | 29 | 18 | 7.2 | ||||
| 3,4-Dimethylphenol | 18 | 11 | 4.6 | ||||
| m-Xylene | 3-Methylbenzyl alcohol | 2.2 | 28 ± 5 | 7.6 | 17 ± 0 | 14 | 27 ± 2 |
| 2,4-Dimethylphenol | 87 | 88 | 79 | ||||
| 2,6-Dimethylphenol | 11 | 4.3 | 7.3 | ||||
| p-Xylene | 4-Methylbenzyl alcohol | 2.1 | 38 ± 2 | 6.4 | 11 ± 0 | 5.5 | 31 ± 0 |
| 2,5-Dimethylphenol | 98 | 94 | 94 | ||||
| Fluorobenzene | 2-Fluorophenol | 57 | 7.7 ± 0.3 | 57 | 7.4 ± 0.4 | 56 | 4.5 ± 0.2 |
| 4-Fluorophenol | 43 | 43 | 44 | ||||
| Chlorobenzene | 2-Chlorophenol | 93 | 15 ± 1 | 93 | 8.5 ± 0.4 | 91 | 8.1 ± 0.2 |
| 4-Chlorophenol | 6.8 | 6.7 | 9.0 | ||||
| Naphthalene | 1-Naphthol | 99 | 26 ± 0 | 99 | 13 ± 0 | 99 | 24 ± 2 |
| 2-Naphthol | 1.0 | 0.9 | 1.1 | ||||
Although halogenated aromatic compounds are generally persistent because their C–H bonds are relatively inert owing to the high electronegativity of halogen substituents, P. megaterium and B. subtilis were still able to produce hydroxylated products from fluorobenzene and chlorobenzene with yields of 5–15%. Naphthalene, which has a relatively larger molecular size, was also efficiently hydroxylated to yield 1-naphthol with an approximate yield of 25%, highlighting the potential of this reaction system for hydroxylating larger aromatic compounds. P. megaterium with C7-Pro-Phe retained hydroxylation activity even when the decoy molecule concentration was reduced to 200 nM, while maintaining >99% regioselectivity of for 1-naphthol (Fig. S8). At this concentration, the amount of hydroxylated products formed per decoy molecule reached 237 ± 63, indicating a highly efficient catalytic contribution of the decoy molecule. In contrast, B. subtilis with (S)-Ibu-Leu produced only background-level amounts of hydroxylation products at decoy concentration below 4 µM, comparable to the no-decoy control. At higher levels, hydroxylation products became clearly quantifiable and increased with the decoy concentration, without compromising its high regioselectivity for 1-naphthol (>97%). The normalized value (product per decoy molecule) reached its maximum of 6.1 ± 0.6 at 20 µM. These results clearly demonstrate that both P. megaterium and B. subtilis exhibit high hydroxylation activity toward representative aromatic pollutants. Since native bacterial cells were used in this study, the reactions were catalyzed by wild-type P450 enzymes (CYP102A1 and CYP102A3), which usually hydroxylate only long-chain fatty acids. The presence of decoy molecules markedly relaxed the substrate specificity, enabling the system to degrade substrates of various molecular sizes and polarities.
It is noteworthy that the two strains exhibited largely overlapping regioselectivity patterns for toluene and xylene hydroxylation. In toluene hydroxylation, ortho-selectivity was remarkably high (>92%) in both strains. Since purified CYP102A1 with decoy molecule also exhibited strong ortho-selectivity,72 the observed regioselectivity further confirms that CYP102A1 is responsible for the hydroxylation reaction in P. megaterium. This finding also suggests that the substrate-binding site of CYP102A3 is very similar to that of CYP102A1, which can be attributed to their high amino acid sequence similarity (Table S2). A similar trend in the regioselectivity of xylene, halobenzene, and naphthalene hydroxylations between P. megaterium and B. subtilis indicates that these substrates bind to the active site in a similar orientation.
3.3. Biodegradation of dioxin model compounds
Building on the finding that relatively large environmental pollutants such as naphthalene were hydroxylated, we finally examined whether the reaction system employing native bacterial cells with decoy molecules could degrade 1- and 2-chlorinated dibenzo-p-dioxins (1- and 2-CDDs), representative model compounds of dioxins. B. subtilis exhibited a marked decrease in 1- and 2-CDD levels upon the addition of (S)-Ibu-Leu as the decoy molecule, resulting in the degradation of most of 1- and 2-CDD within 2 h at 45 °C, whereas B. subtilis alone showed only a slight decrease, probably due to volatilization rather than enzymatic degradation, indicating that oxidation of CDDs was triggered by the decoy molecule (Fig. 4a). It is noteworthy that although B. subtilis JCM 1465T was not isolated from soil contaminated with 1- and 2-CDDs, supplementation with the external additive alone was sufficient to induce the degradation of CDDs. In contrast, P. megaterium did not show any noticeable decrease in 1- and 2-CDD levels, possibly due to a relatively smaller active site compared with that of B. subtilis. Using non-chlorinated dibenzo-p-dioxin (DD) as a surrogate compound to identify degradation products, we observed an additional peak assignable to mono-hydroxylated dibenzo-p-dioxin (samples were derivatized by trimethylsilylation for GC-MS detection), which appeared exclusively in the reaction containing the decoy molecule (Fig. 4b and S9).96 These results suggest that 1- and 2-CDDs, which have even larger molecular sizes than naphthalene, were accommodated in the active site of CYP102A3 and hydroxylated. To evaluate the binding of 1- and 2-CDDs, we constructed protein–ligand complexes based on three-dimensional (3D) structures predicted by AlphaFold2 and refined by molecular dynamics (MD) simulations, followed by docking simulations using AutoDock Vina. Docking simulations revealed that 1- and 2-CDDs can be accommodated within the active-site cavity of CYP102A3 in the presence of (S)-Ibu-Leu as a decoy molecule (Fig. 5, S10, and S11), in a manner similar to the binding of styrene in CYP102A1 complexed with a decoy molecule.97 These results suggest that CYP102A3 can also accommodate these substrates and that their degradation is mediated by the decoy molecule system, leading to efficient degradation of pollutants with relatively large molecular sizes.This study establishes a proof-of-concept for a new bioremediation strategy that harnesses the catalytic potential of native microbial enzymes via small-molecule decoys, eliminating the need for genetic modification. The approach highlights the possibility of reprogramming microbial metabolism for pollutant degradation beyond conventional genetic strategies. Key challenges that remain include expanding the substrate scope, improving catalytic efficiency under variable soil conditions, and ensuring sustained activity and specificity in complex microbial communities. Addressing these points will require optimization of decoy structures, deeper mechanistic insight, and validating this approach under environmentally relevant conditions.
4. Conclusions
In this work, we demonstrated that a variety of naturally occurring soil bacteria can hydroxylate stable and persistent environmental pollutants such as benzene, toluene, and xylenes (BTX) and other chemically stable aromatic compounds upon the addition of decoy molecules that activate their endogenous CYP102 enzymes. Remarkably, B. subtilis degraded the dioxin model compound 2-chlorinated dibenzo-p-dioxin almost completely within 2 h at 45 °C. The use of decoy molecules enables a substantial alteration of the substrate specificity of endogenous CYP102 enzymes without any genetic modification, allowing natural soil bacteria to function as biocatalysts for pollutant degradation while avoiding the legal restrictions associated with genetically engineered microorganisms. As small synthetic organic compounds, decoy molecules are generally evaluated under conventional chemical-substance regulatory frameworks rather than GEM-specific legislation. Importantly, OPERA98-based predictions indicate that the decoy molecules exhibit moderate environmental persistence—moderately slower than benzene but far below that of highly recalcitrant pollutants such as dioxins—and their catalytic efficiency enables effective use at very low concentrations, suggesting that only minimal amounts would be required in practical applications (Table S5 and Fig. S8). These considerations suggest that the decoy strategy provides a practical and environmentally compatible approach for the degradation of persistent organic pollutants in contaminated soils. Overall, this study establishes a groundbreaking concept in which decoy molecules can convert naturally occurring soil bacteria into pollutant-degrading microorganisms without genetic engineering. Although the present work demonstrates this principle in vitro, using CYP102 family enzymes that natively catalyze fatty acid hydroxylation, the same concept could be extended to other enzyme families, offering broad potential for the degradation of diverse environmental pollutants through non-genetic biocatalyst activation. Future studies should evaluate the environmental stability, bioavailability, and practical deployability of decoy molecules in real soil matrices, as well as their environmental fate during field applications, to determine whether this strategy can be safely implemented in contaminated environments.Author contributions
Conceptualization, O. S.; methodology, F. I.; investigation, F. I. and M. K.; visualization, F. I.; writing – original draft, F. I. and O. S.; writing – review and editing, F. I. and O. S.; funding acquisition, F. I., M. K. and O. S.; supervision, O. S.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: experimental details, GC-MS and HPLC analyses of the reaction mixture, chemical structures of the decoy molecules, experiments using E. coli and gene-knockout mutants of B. subtilis 168, and computational details. See DOI: https://doi.org/10.1039/d5ta09218c. The authors have cited additional references within the SI.99–114Acknowledgements
This work was supported by JSPS KAKENHI (JP22H05129 and JP25H00910 to O. S., JP19J23669 to M. K., and JP24KJ1242 to F. I.), Japan. F. I. acknowledges support from the “THERS Interdisciplinary Frontier Next Generation Researcher Project”, supported by JST SPRING (Grant Number JPMJSP2125). F. I. also acknowledges support from the “Graduate Program of Transformative Chem-Bio Research” at Nagoya University, funded by MEXT (WISE Program). The authors thank Dr Yuichiro Aiba and Dr Shinya Ariyasu for their assistance with the experiments and for valuable discussions.Notes and references
- E. L. Rylott and N. C. Bruce, Curr. Opin. Chem. Biol., 2020, 58, 86–95 CrossRef CAS PubMed.
- M. Yoshikawa, M. Zhang and K. Toyota, Microb. Environ., 2017, 32, 188–200 CrossRef PubMed.
- United States Environmental Protection Agency (USEPA), Superfund Remedy Report, 18th edn, 2025 Search PubMed.
- S. Kuppusamy, P. Thavamani, K. Venkateswarlu, Y. B. Lee, R. Naidu and M. Megharaj, Chemosphere, 2017, 168, 944–968 CrossRef CAS PubMed.
- R. A. Rinsky, A. B. Smith, R. Hornung, T. G. Filloon, R. J. Young, A. H. Okun and P. J. Landrigan, N. Engl. J. Med., 1987, 316, 1044–1050 CrossRef CAS PubMed.
- T. Rengarajan, P. Rajendran, N. Nandakumar, B. Lokeshkumar, P. Rajendran and I. Nishigaki, Asian Pac. J. Trop. Biomed., 2015, 5, 182–189 CrossRef CAS.
- M. Honda and N. Suzuki, Int. J. Environ. Res. Public Health, 2020, 17, 1363 CrossRef CAS PubMed.
- M. Van den Berg, L. S. Birnbaum, M. Denison, M. De Vito, W. Farland, M. Feeley, H. Fiedler, H. Hakansson, A. Hanberg, L. Haws, M. Rose, S. Safe, D. Schrenk, C. Tohyama, A. Tritscher, J. Tuomisto, M. Tysklind, N. Walker and R. E. Peterson, Toxicol. Sci., 2006, 93, 223–241 CrossRef CAS PubMed.
- V. F. Doherty and A. A. Otitoloju, Environ. Monit. Assess., 2013, 185, 4159–4170 CrossRef CAS PubMed.
- Agency for Toxic Substances and Disease Registry (ATSDR), Toxicological Profile for Benzene (Draft for Public Comment), U.S. Department of Health and Human Services, Public Health Service, Atlanta, GA, 2024 Search PubMed.
- Agency for Toxic Substances and Disease Registry (ATSDR), Toxicological Profile for Naphthalene, 1-Methylnaphthalene, and 2-Methylnaphthalene, U.S. Department of Health and Human Services, Public Health Service, Atlanta, GA, 2025 Search PubMed.
- O. Kiguchi, T. Kobayashi, Y. Wada, K. Saitoh and N. Ogawa, Chemosphere, 2007, 67, 557–573 CrossRef CAS PubMed.
- A. O. W. Leung, W. J. Luksemburg, A. S. Wong and M. H. Wong, Environ. Sci. Technol., 2007, 41, 2730–2737 CrossRef CAS.
- H. Zhang, Y. Ni, J. Chen, F. Su, X. Lu, L. Zhao, Q. Zhang and X. Zhang, Chemosphere, 2008, 73, 1640–1648 CrossRef CAS PubMed.
- G. J. Zheng, A. O. W. Leung, L. P. Jiao and M. H. Wong, Environ. Int., 2008, 34, 1050–1061 CrossRef CAS PubMed.
- S. Henriksson, J. Hagberg, M. Bäckström, I. Persson and G. Lindström, Environ. Pollut., 2013, 180, 19–26 CrossRef CAS PubMed.
- M. Vidali, Pure Appl. Chem., 2001, 73, 1163–1172 CAS.
- S. Saibu, S. A. Adebusoye and G. O. Oyetibo, Bioresour. Bioprocess., 2020, 7, 7 CrossRef.
- D. T. Gibson, J. R. Koch and R. E. Kallio, Biochemistry, 1968, 7, 2653–2662 CrossRef CAS PubMed.
- J. F. Davey and D. T. Gibson, J. Bacteriol., 1974, 119, 923–929 CrossRef CAS PubMed.
- D. T. Gibson, V. Mahadevan, D. M. Jerina, H. Yagi and H. J. C. Yeh, Science, 1975, 189, 295–297 CrossRef CAS PubMed.
- K. Furukawa, J. R. Simon and A. M. Chakrabarty, J. Bacteriol., 1983, 154, 1356–1362 CrossRef CAS PubMed.
- L. Monna, T. Omori and T. Kodama, Appl. Environ. Microbiol., 1993, 59, 285–289 CrossRef CAS PubMed.
- J. Schneider, R. Grosser, K. Jayasimhulu, W. Xue and D. Warshawsky, Appl. Environ. Microbiol., 1996, 62, 13–19 CrossRef CAS.
- J. N. Rooney-Varga, R. T. Anderson, J. L. Fraga, D. Ringelberg and D. R. Lovley, Appl. Environ. Microbiol., 1999, 65, 3056–3063 CrossRef CAS PubMed.
- J. D. Coates, R. Chakraborty, J. G. Lack, S. M. O'Connor, K. A. Cole, K. S. Bender and L. A. Achenbach, Nature, 2001, 411, 1039–1043 CrossRef CAS PubMed.
- H. Habe, K. Ide, M. Yotsumoto, H. Tsuji, T. Yoshida, H. Nojiri and T. Omori, Chemosphere, 2002, 48, 201–207 CrossRef CAS PubMed.
- I.-H. Nam, Y.-M. Kim, S. Schmidt and Y.-S. Chang, Appl. Environ. Microbiol., 2006, 72, 112–116 CrossRef CAS PubMed.
- J. Dou, A. Ding, X. Liu, Y. Du, D. Deng and J. Wang, J. Environ. Sci., 2010, 22, 709–715 CrossRef CAS PubMed.
- A. Hanano, M. Shaban, D. Almutlk and I. Almousally, Chemosphere, 2019, 216, 258–270 CrossRef CAS PubMed.
- D. Stephenson and T. Stephenson, Biotechnol. Adv., 1992, 10, 549–559 CrossRef CAS PubMed.
- S. El Fantroussi and S. N. Agathos, Curr. Opin. Microbiol., 2005, 8, 268–275 CrossRef CAS PubMed.
- I. P. Thompson, C. J. Van Der Gast, L. Ciric and A. C. Singer, Environ. Microbiol., 2005, 7, 909–915 CrossRef CAS PubMed.
- M. Herrero and D. C. Stuckey, Chemosphere, 2015, 140, 119–128 CrossRef CAS PubMed.
- K. E. Hammel, B. Kalyanaraman and T. K. Kirk, J. Biol. Chem., 1986, 261, 16948–16952 CrossRef CAS.
- S. M. Resnick, K. Lee and D. T. Gibson, J. Ind. Microbiol. Biotechnol., 1996, 17, 438–457 Search PubMed.
- B. Leuthner, C. Leutwein, H. Schulz, P. Hörth, W. Haehnel, E. Schiltz, H. Schägger and J. Heider, Mol. Microbiol., 1998, 28, 615–628 CrossRef CAS PubMed.
- H. Nojiri, J.-W. Nam, M. Kosaka, K.-I. Morii, T. Takemura, K. Furihata, H. Yamane and T. Omori, J. Bacteriol., 1999, 181, 3105–3113 Search PubMed.
- Y. Tao, A. Fishman, W. E. Bentley and T. K. Wood, Appl. Environ. Microbiol., 2004, 70, 3814–3820 Search PubMed.
- K. Koschorreck, S. M. Richter, A. Swierczek, U. Beifuss, R. D. Schmid and V. B. Urlacher, Arch. Biochem. Biophys., 2008, 474, 213–219 CrossRef CAS PubMed.
- C. F. Harford-Cross, A. B. Carmichael, F. K. Allan, P. A. England, D. A. Rouch and L.-L. Wong, Protein Eng., 2000, 13, 121–128 Search PubMed.
- A. B. Carmichael and L.-L. Wong, Eur. J. Biochem., 2001, 268, 3117–3125 CrossRef CAS PubMed.
- Q.-S. Li, J. Ogawa, R. D. Schmid and S. Shimizu, Appl. Environ. Microbiol., 2001, 67, 5735–5739 CrossRef CAS PubMed.
- W. T. Sulistyaningdyah, J. Ogawa, Q.-S. Li, R. Shinkyo, T. Sakaki, K. Inouye, R. D. Schmid and S. Shimizu, Biotechnol. Lett., 2004, 26, 1857–1860 Search PubMed.
- K. Neufeld, J. Marienhagen, U. Schwaneberg and J. Pietruszka, Green Chem., 2013, 15, 2408–2421 RSC.
- A. Sideri, A. Goyal, G. Di Nardo, G. E. Tsotsou and G. Gilardi, J. Inorg. Biochem., 2013, 120, 1–7 CrossRef CAS PubMed.
- H. Suenaga, M. Mitsuoka, Y. Ura, T. Watanabe and K. Furukawa, J. Bacteriol., 2001, 183, 5441–5444 CrossRef CAS PubMed.
- T. Bulter, M. Alcalde, V. Sieber, P. Meinhold, C. Schlachtbauer and F. H. Arnold, Appl. Environ. Microbiol., 2003, 69, 987–995 CrossRef CAS PubMed.
- T. Mori and R. Kondo, FEMS Microbiol. Lett., 2002, 216, 223–227 Search PubMed.
- P. G. de Sousa Júnior, K. M. dos Santos, D. E. P. Silva, D. Nascimento Dari, V. d. J. Andrade Lima, F. I. da Silva Aires, V. de Castro Bizerra, F. Miranda Nunes and J. C. Sousa dos Santos, ACS ES&T Water, 2025, 5, 6217–6245 Search PubMed.
- H. Rafeeq, N. Afsheen, S. Rafique, A. Arshad, M. Intisar, A. Hussain, M. Bilal and H. M. N. Iqbal, Chemosphere, 2023, 310, 136751 CrossRef CAS PubMed.
- R. Boopathy, Bioresour. Technol., 2000, 74, 63–67 CrossRef CAS.
- J. Davison, J. Ind. Microbiol. Biotechnol., 2005, 32, 639–650 CrossRef CAS PubMed.
- J. S. Singh, P. C. Abhilash, H. B. Singh, R. P. Singh and D. P. Singh, Gene, 2011, 480, 1–9 CrossRef CAS PubMed.
- K. M. Tran, H.-M. Lee, T. D. Thai, J. Shen, S.-I. Eyun and D. Na, J. Hazard. Mater., 2021, 419, 126516 CrossRef CAS PubMed.
- A. Saravanan, P. S. Kumar, B. Ramesh and S. Srinivasan, Chemosphere, 2022, 298, 134341 CrossRef CAS PubMed.
- M. B. Amrofell, S. Rengarajan, S. T. Vo, E. S. Ramirez Tovar, L. LoBello, G. Dantas and T. S. Moon, Cells Rep. Methods, 2023, 3, 100669 CrossRef CAS PubMed.
- D. R. George, M. Danciu, P. W. Davenport, M. R. Lakin, J. Chappell and E. K. Frow, Nat. Commun., 2024, 15, 650 CrossRef CAS PubMed.
- T. S. Moon, Curr. Opin. Food Sci., 2024, 56, 101130 CrossRef CAS.
- Y. Ma, A. Manna and T. S. Moon, Curr. Opin. Syst. Biol., 2023, 36, 100483 CrossRef CAS.
- A. M. Hartig, W. Dai, K. Zhang, K. Kapoor, A. G. Rottinghaus, T. S. Moon and K. M. Parker, Environ. Sci. Technol., 2024, 58, 22657–22667 CrossRef CAS PubMed.
- K. E. French, Z. Zhou and N. Terry, Sci. Rep., 2020, 10, 15091 CrossRef CAS PubMed.
- H. Acar Kirit, J. P. Bollback and M. Lagator, Mol. Biol. Evol., 2022, 39, msac220 CrossRef PubMed.
- R. S. Gupta, S. Patel, N. Saini and S. Chen, Int. J. Syst. Evol. Microbiol., 2020, 70, 5753–5798 CrossRef CAS PubMed.
- L. O. Narhi and A. J. Fulco, J. Biol. Chem., 1986, 261, 7160–7169 CrossRef CAS PubMed.
- H. Li and T. L. Poulos, Nat. Struct. Biol., 1997, 4, 140–146 CrossRef CAS PubMed.
- C. J. C. Whitehouse, S. G. Bell and L.-L. Wong, Chem. Soc. Rev., 2012, 41, 1218–1260 RSC.
- N. Kawakami, O. Shoji and Y. Watanabe, Angew. Chem., Int. Ed., 2011, 50, 5315–5318 CrossRef CAS PubMed.
- N. Kawakami, O. Shoji and Y. Watanabe, Chem. Sci., 2013, 4, 2344–2348 Search PubMed.
- O. Shoji, T. Kunimatsu, N. Kawakami and Y. Watanabe, Angew. Chem., Int. Ed., 2013, 52, 6606–6610 CrossRef CAS PubMed.
- Z. Cong, O. Shoji, C. Kasai, N. Kawakami, H. Sugimoto, Y. Shiro and Y. Watanabe, ACS Catal., 2015, 5, 150–156 CrossRef CAS.
- O. Shoji, S. Yanagisawa, J. K. Stanfield, K. Suzuki, Z. Cong, H. Sugimoto, Y. Shiro and Y. Watanabe, Angew. Chem., Int. Ed., 2017, 56, 10324–10329 CrossRef CAS PubMed.
- M. Karasawa, J. K. Stanfield, S. Yanagisawa, O. Shoji and Y. Watanabe, Angew. Chem., Int. Ed., 2018, 57, 12264–12269 CrossRef CAS PubMed.
- K. Yonemura, S. Ariyasu, J. K. Stanfield, K. Suzuki, H. Onoda, C. Kasai, H. Sugimoto, Y. Aiba, Y. Watanabe and O. Shoji, ACS Catal., 2020, 10, 9136–9144 CrossRef CAS.
- Y. Yokoyama, S. Ariyasu, M. Karasawa, C. Kasai, Y. Aiba, H. Sugimoto and O. Shoji, ChemCatChem, 2025, 17, e202401641 CrossRef CAS.
- Y. Sugai, M. Karasawa and O. Shoji, JACS Au, 2025, 5, 4196–4203 CrossRef CAS PubMed.
- P. Le-Huu, T. Heidt, B. Claasen, S. Laschat and V. B. Urlacher, ACS Catal., 2015, 5, 1772–1780 CrossRef CAS.
- J. Akter, T. P. Stockdale, S. A. Child, J. H. Z. Lee, J. J. De Voss and S. G. Bell, J. Inorg. Biochem., 2023, 244, 112209 CrossRef CAS PubMed.
- R. Fasan, M. M. Chen, N. C. Crook and F. H. Arnold, Angew. Chem., Int. Ed., 2007, 46, 8414–8418 CrossRef CAS PubMed.
- K. Zhang, B. M. Shafer, M. D. Demars II, H. A. Stern and R. Fasan, J. Am. Chem. Soc., 2012, 134, 18695–18704 CrossRef CAS PubMed.
- R. Biedendieck, Y. Yang, W.-D. Deckwer, M. Malten and D. Jahn, Biotechnol. Bioeng., 2007, 96, 525–537 CrossRef CAS PubMed.
- R. Biedendieck, R. Beine, M. Gamer, E. Jordan, K. Buchholz, J. Seibel, L. Dijkhuizen, M. Malten and D. Jahn, Appl. Microbiol. Biotechnol., 2007, 74, 1062–1073 CrossRef CAS PubMed.
- M. Ehrhardt, A. Gerber, F. Hannemann and R. Bernhardt, J. Biotechnol., 2016, 218, 34–40 CrossRef CAS PubMed.
- P. S. Vary, R. Biedendieck, T. Fuerch, F. Meinhardt, M. Rohde, W.-D. Deckwer and D. Jahn, Appl. Microbiol. Biotechnol., 2007, 76, 957–967 CrossRef CAS PubMed.
- C. Korneli, F. David, R. Biedendieck, D. Jahn and C. Wittmann, J. Biotechnol., 2013, 163, 87–96 CrossRef CAS PubMed.
- R. Biedendieck, T. Knuuti, S. J. Moore and D. Jahn, Appl. Microbiol. Biotechnol., 2021, 105, 5719–5737 CrossRef CAS PubMed.
- B.-M. Koo, G. Kritikos, J. D. Farelli, H. Todor, K. Tong, H. Kimsey, I. Wapinski, M. Galardini, A. Cabal, J. M. Peters, A.-B. Hachmann, D. Z. Rudner, K. N. Allen, A. Typas and C. A. Gross, Cell Syst., 2017, 4, 291–305 CrossRef CAS PubMed.
- E. Emerson, J. Org. Chem., 1943, 08, 417–428 CrossRef CAS.
- T. S. Wong, N. Wu, D. Roccatano, M. Zacharias and U. Schwaneberg, J. Biomol. Screening, 2005, 10, 246–252 CrossRef CAS PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef.
- O. Trott and A. J. Olson, J. Comput. Chem., 2010, 31, 455–461 CrossRef CAS PubMed.
- S. Salentin, S. Schreiber, V. J. Haupt, M. F. Adasme and M. Schroeder, Nucleic Acids Res., 2015, 43, W443–W447 CrossRef CAS PubMed.
- J. K. Stanfield, H. Onoda, S. Ariyasu, C. Kasai, E. M. Burfoot, H. Sugimoto and O. Shoji, J. Inorg. Biochem., 2023, 245, 112235 CrossRef CAS PubMed.
- X. Wu, Y. Chen, X. Wang, W. Wei and Y. Liang, J. Org. Chem., 2021, 86, 13768–13773 CrossRef CAS PubMed.
- N. Kimura and Y. Urushigawa, J. Biosci. Bioeng., 2001, 92, 138–143 CrossRef CAS PubMed.
- K. Suzuki, Y. Shisaka, J. K. Stanfield, Y. Watanabe and O. Shoji, Chem. Commun., 2020, 56, 11026–11029 RSC.
- K. Mansouri, C. M. Grulke, R. S. Judson and A. J. Williams, J. Cheminf., 2018, 10, 10 Search PubMed.
- J. Kabisch, A. Thürmer, T. Hübel, L. Popper, R. Daniel and T. Schweder, J. Biotechnol., 2013, 163, 97–104 CrossRef CAS PubMed.
- F. Sievers, A. Wilm, D. Dineen, T. J. Gibson, K. Karplus, W. Li, R. Lopez, H. McWilliam, M. Remmert, J. Söding, J. D. Thompson and D. G. Higgins, Mol. Syst. Biol., 2011, 7, 539 CrossRef PubMed.
- D. C. Bas, D. M. Rogers and J. H. Jensen, Proteins: Struct., Funct., Bioinf., 2008, 73, 765–783 CrossRef CAS PubMed.
- K. Shahrokh, A. Orendt, G. S. Yost and T. E. Cheatham III, J. Comput. Chem., 2012, 33, 119–133 CrossRef CAS PubMed.
- W. L. Jorgensen, J. Chandrasekhar, J. D. Madura, R. W. Impey and M. L. Klein, J. Chem. Phys., 1983, 79, 926–935 CrossRef CAS.
- D. A. Case, H. M. Aktulga, K. Belfon, D. S. Cerutti, G. A. Cisneros, V. W. D. Cruzeiro, N. Forouzesh, T. J. Giese, A. W. Götz, H. Gohlke, S. Izadi, K. Kasavajhala, M. C. Kaymak, E. King, T. Kurtzman, T.-S. Lee, P. Li, J. Liu, T. Luchko, R. Luo, M. Manathunga, M. R. Machado, H. M. Nguyen, K. A. O'Hearn, A. V. Onufriev, F. Pan, S. Pantano, R. Qi, A. Rahnamoun, A. Risheh, S. Schott-Verdugo, A. Shajan, J. Swails, J. Wang, H. Wei, X. Wu, Y. Wu, S. Zhang, S. Zhao, Q. Zhu, T. E. Cheatham III, D. R. Roe, A. Roitberg, C. Simmerling, D. M. York, M. C. Nagan and K. M. Merz Jr, J. Chem. Inf. Model., 2023, 63, 6183–6191 CrossRef CAS PubMed.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, J. Chem. Theory Comput., 2015, 11, 3696–3713 CrossRef CAS PubMed.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef CAS PubMed.
- M. R. Shirts, C. Klein, J. M. Swails, J. Yin, M. K. Gilson, D. L. Mobley, D. A. Case and E. D. Zhong, J. Comput.-Aided Mol. Des., 2017, 31, 147–161 CrossRef CAS PubMed.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 CrossRef PubMed.
- M. Bernetti and G. Bussi, J. Chem. Phys., 2020, 153, 114107 CrossRef CAS PubMed.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- B. Hess, J. Chem. Theory Comput., 2008, 4, 116–122 CrossRef CAS PubMed.
- D. R. Nelson, Hum. Genomics, 2009, 4, 59 CrossRef CAS PubMed.
- D. Gront, K. Syed and D. R. Nelson, Protein Sci., 2025, 34, e70057 CrossRef CAS PubMed.
Footnote |
| † It should be noted that the widely used laboratory strain Bacillus subtilis 168, which is the parental strain of the deletion mutants (e.g., B. subtilis 168 Δcyp102a3) used in this study, also catalyzes benzene hydroxylation upon the addition of decoy molecule (Fig. S4). |
| This journal is © The Royal Society of Chemistry 2026 |