Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Nanoplasmonic core–shell nanoraspberry chip for ultrasensitive surface-enhanced Raman scattering detection of SARS-CoV-2: a modular nanobiosensor for respiratory virus diagnostics
Nnamdi
Nwahara
 a,
Saba
Niaz
a,
Muthumuni
Managa
a,
Saba
Niaz
a,
Muthumuni
Managa
 b,
Christian I.
Nkanga
c,
Oluwasesan
Adegoke
b,
Christian I.
Nkanga
c,
Oluwasesan
Adegoke
 d and
Ojodomo J.
Achadu
d and
Ojodomo J.
Achadu
 *a
*a
aSchool of Health and Life Sciences, and National Horizon Centre, Teesside University, Middlesbrough TS1 3BA, UK. E-mail: O.Achadu@tees.ac.uk
bInstitute for Nanotechnology and Water Sustainability (iNanoWS), Florida Campus, College of Science, Engineering and Technology, University of South Africa, Johannesburg 1710, South Africa
cCentre de Recherche en Nanotechnologies Appliquées aux Produits Naturels (CReNAPN), Department of Medicinal Chemistry and Pharmacognosy, Faculty of Pharmaceutical Sciences, University of Kinshasa, B.P. 212, Kinshasa XI, Democratic Republic of the Congo
dLeverhulme Research Centre for Forensic Science, School of Science & Engineering, University of Dundee, Dundee DD1 4GH, UK
First published on 3rd February 2026
Abstract
The development of robust and ultrasensitive point-of-care diagnostics for viral pathogens, particularly for seasonal respiratory viruses, remains a critical challenge, as traditional biosensing platforms often suffer from signal instability and poor performance in complex biological matrices. Here, we report a dual-component surface-enhanced Raman scattering (SERS) immunosensor designed to mitigate some of these limitations through a new hotspot engineering strategy. Our platform is centred on a core–shell Au@Ag nanotag fabricated from branched gold nanoraspberries (AuNRBs), which provides a high density of plasmonic hotspots. A conformal Ag shell confines the 4-aminothiophenol (4-ATP) reporter within these nanogaps, establishing a unique “gap-confined” Raman reporter architecture that enhances signal stability and reproducibility. This nanotag is complemented by a second SERS-active component: a capture sensor chip of fluorine-doped tin oxide (FTO) modified with a dense layer of Au nanoparticles (AuNPs). When implemented in a sandwich immunoassay for the SARS-CoV-2 S2 spike protein, this dual-enhancement platform achieved a sensitive detection range of 1–500 ng mL−1 with an ultralow limit of detection of 1.07 pg mL−1 (15.3 fM) in PBS and 1.30 pg mL−1 (18.6 fM) in human serum albumin (HSA). Although these results demonstrate strong analytical sensitivity and rapid assay performance within 30 min, further studies are required to assess clinical robustness, scalability and long-term stability. Overall, this work demonstrates the potential of the proposed SERS system as a versatile platform for rapid, on-site pathogen diagnostics.
Introduction
Acute respiratory infection (ARI) remains a leading global health challenge,1 with SARS-CoV-2 as a predominant cause of severe respiratory illnesses.2,3 Given the increasing frequency and severity of pathogenic viruses, there is a growing need for the development of next-generation diagnostic nanosensors for efficient and rapid virus detection. For frontline applications, such diagnostic tools should be affordable, sensitive, specific, and rapid, in line with the ASSURED criteria.4 Currently, reverse transcription-polymerase chain reaction (RT-PCR) is considered the gold standard for SARS-CoV-2 detection.5 However, it requires specialised facilities and personnel, is plagued by cumbersome operational procedures,6 and can be complicated by viral mutations.7,8 Viral cultures, while highly specific, are limited by protracted turnaround times.9 Alternatively, protein-based detection methods offer simpler workflows and target viral antigens, with comparatively greater stability.10,11A particularly relevant biosensing target is the S2 subunit of the SARS-CoV-2 spike protein. Unlike the S1 subunit, which is prone to frequent mutations, the S2 subunit is highly conserved across SARS-CoV-2 strains.12 This conserved nature is advantageous for maintaining diagnostic performance as new variants with different S1/RBD sequences emerge. The S2 domain mediates viral and host membrane fusion, a critical step in viral entry; thus, detecting the S2 subunit not only confirms the presence of the virus, but may also support therapeutic monitoring.13,14 Targeting S2 also complements existing methods that primarily focus on the nucleocapsid (N) protein or the S1/RBD region, thereby potentially improving test robustness and reducing false negatives associated with mutations in other domains.15
Advances in nanofabrication have enabled the development of robust nanomaterial-based SARS-CoV-2 sensor systems, including colorimetric,16 fluorescence-based,17 and surface-enhanced Raman scattering (SERS)-based systems.18 Among these, SERS has attracted considerable interest due to its high reported sensitivities and ease of multiplex integration.19 SERS offers versatile ultrasensitive biosensing platforms, combining molecular fingerprinting with trace-level detection.18 The technique benefits from tailored plasmonic nanostructures, such as Au and Ag nanoparticles, nanorods, nanostars,20 and hybrid nanostructures,21 which generate intense electromagnetic “hotspots” for signal amplification.22 SERS nanotags, which are metal nanoparticles functionalized with Raman reporter molecules, serve as the primary signal-generating units and have been widely applied for detecting viral pathogens,18 cancer biomarkers and other clinically relevant targets.23
Despite these advances, conventional SERS nanotags often suffer from signal instability arising from reporter desorption or nanoparticle aggregation, particularly in complex biological matrices. Protective coatings, such as silica shells or hydrophilic polymers (e.g., PEG), have been employed to enhance stability, and improve biocompatibility.24,25 However, these strategies have limitations: PEG provides only moderate protection against desorption and photodegradation,26 while silica is susceptible to hydrolytic degradation and aggregation under physiological conditions.27 These vulnerabilities limit their reliability and broader deployment in real world settings.21,26 Therefore, there remains an urgent need for the development of rapid ultrasensitive SERS biosensors that combine high sensitivity with improved signal stability and practical applicability for the accurate detection of SARS-CoV-2 and other infectious viruses.
To address these challenges, this work reports on the fabrication of a SARS-CoV-2 S2 biosensor employing gold-nanoraspberries (AuNRBs) functionalised with 4-aminothiophenol (4-ATP) and encapsulated within the conformal Ag shell to form robust SERS nanotags. Here, the term “nanoraspberry” is used as a conceptual descriptor of a hierarchical core–shell plasmonic architecture, rather than implying a strict morphological resemblance to a macroscopic raspberry. This core–shell architecture, inspired by gap-enhanced Raman tags (GERTs),18,28 spatially confines the Raman reporter within nanogaps between the AuNRBs and the Ag shell, contributing to enhanced signal stability and reproducibility. The multiscale roughness of the AuNRBs is poised to promote dense hotspot formation, while the anchoring of 4-ATP by strong Au–S bonds facilitates stable reporter integration. The Ag shell provides plasmonic amplification and protection. The capture sensor chip substrate consists of a fluorine-doped tin oxide (FTO) chip modified with Au nanoparticles and antibodies to create a plasmonically active capture surface. Detection proceeds via a sandwich immunoassay in which the SARS-CoV-2 S2 spike protein bridges the sensor chip and the antibody-functionalized nanotags, generating strong and quantifiable SERS signals. The resulting platform demonstrated a large detection range from 1 ng mL−1 to 500 ng mL−1 range with a limit of detection of 18.6 fM in human serum albumin (HSA). When coupled with a portable Raman spectrometer, rapid point-of-care testing within 30 min may be enabled. While further validation is required to assess clinical performance, the proposed platform shows potential for adaptation to multiplexed detection of multiple pathogens, providing a versatile, high-performance platform for rapid viral pathogen monitoring.
Experimental
Materials
Gold(III) chloride trihydrate, silver nitrate, trisodium citrate, ascorbic acid, sodium hydroxide, 4-aminothiophenol (4-ATP), 11-mercaptoundecanoic acid (11-MUA), N-(3-dimethylaminopropyl)-N′-ethylcarbodiimide hydrochloride (EDC), N-succinimide (NHS), (3-mercaptopropyl)trimethoxysilane (MPTMS), tetraethyl orthosilicate (TEOS), 2-(4-morpholino)ethanesulfonic acid (MES), acetone, and ammonium hydroxide solution were obtained from Sigma-Aldrich. Phosphate-buffered solution (PBS, 0.1 M, pH 7.4) without calcium and magnesium was also obtained from Sigma-Aldrich. N-2-Hydroxyethylpiperazine-N′-2-ethane sulfonic acid (HEPES) buffer (1 M, pH 6.0), the Pierce™ BCA protein assay kit, SARS-CoV-2 spike protein S2 monoclonal and polyclonal antibodies and SARS-CoV spike S1 and S2 proteins were obtained from Thermo Fisher Scientific. Human serum albumin (HSA), a soluble, globular, and monomeric protein, was obtained from a commercial supplier (Sigma-Aldrich) and used as received. Deionized water (18.2 MΩ cm) from a Millipore purification system was used to prepare all buffer solutions. Human respiratory syncytial (RSV) (A2) fusion glycoprotein was obtained from Sino Biological. The FTO glass was purchased from Ossila. All other reagents and solvents were purchased from commercial suppliers and were of analytical grade and used as received.Equipment
Raman measurements were performed on an inVia™ confocal Raman microscope (Renishaw, Gloucestershire, U.K.) under ambient conditions with a 100× objective lens. Calibration was performed by using an internal silicon reference before each set of measurements. The FTO surface was examined using a 785 nm laser, with three spectra captured per time point. Spectral pre-processing, including cosmic ray removal, spline baseline subtraction, and smoothing, was performed using the Raman Environment WiRE software. All SERS spectral data were processed using Microsoft Excel (Microsoft, Redmond, WA) and OriginPro, Version 2025 (OriginLab Corporation, Northampton, MA, USA). The laser power was 0.1% (0.3 mW), and the time of acquisition was set to 1 s at each pixel. Transmission electron microscopy (TEM) images were acquired using a JEOL JEM-1200EX operated at 80 kV. UV-vis measurements were performed on a Spark® multimode microplate reader system from Tecan (Männedorf, Switzerland). Scanning electron microscope images were analysed with a 20 kV Hitachi S-3400N SEM. EDX analysis was performed with a Hitachi TM4000 Plus SEM. Dynamic Light Scattering (DLS) and zeta potential (ZP) analysis were performed using a Zetasizer Nano ZS series (ZEN3600, Malvern). A Siemens D500 powder diffractometer equipped with Cu radiation was used for capturing X-ray diffraction data within a 10° to 70° 2θ range, at a 40 kV and 20 mA setting, with a 0.01° phase scale and 0.2 s step time. FTIR spectra were obtained using a Thermo Nicolet iS5 FT-IR spectrometer in the 450–4000 cm−1 range over 64 scans.Methods
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1), followed by extensive washing with ultra-pure water. Into a 250 mL round bottom flask, 100 mL of 0.25 mM HAuCl4 solution was added. The solution was brought quickly to boil under vigorous stirring. Following this, 2 mL of 1% (w/v) trisodium citrate was quickly added and the solution changed to a deep red colour. The mixture was stirred for an additional 14 min before cooling to room temperature.
1), followed by extensive washing with ultra-pure water. Into a 250 mL round bottom flask, 100 mL of 0.25 mM HAuCl4 solution was added. The solution was brought quickly to boil under vigorous stirring. Following this, 2 mL of 1% (w/v) trisodium citrate was quickly added and the solution changed to a deep red colour. The mixture was stirred for an additional 14 min before cooling to room temperature.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 mixture of 2 mM EDC and 8 mM NHS. The reaction was carried out under gentle agitation in a shaker at room temperature for 15 min to activate the carboxylic (COOH) groups on the surface of the AuNRBs@ATP@Ag NPs. To the activated AuNRBs@ATP@Ag, SARS-CoV-2 polyclonal antibody (SARS-CoV-2 Ab) (9 μg mL−1 in PBS 7.2) was added and incubated at 4 °C overnight. Following this, the SARS-CoV-2 Ab-labelled AuNRBs@ATP@Ag NPs were washed with PBS (pH 7.2). After washing steps through low-speed centrifugation, the amount of antibody conjugated onto the surface of the NPs was determined by micro bicinchoninic acid (MicroBCA) assay. To block unreacted active esters and prevent non-specific binding, the antibody-conjugated AuNRBs@ATP@Ag was resuspended in a buffer containing 1% (w/v) BSA and 0.1 mM Tris–HCl and incubated at room temperature for 1 h. The conjugate was isolated by centrifugation (1000 rpm, 3 min), and the pellet was resuspended in 1
4 mixture of 2 mM EDC and 8 mM NHS. The reaction was carried out under gentle agitation in a shaker at room temperature for 15 min to activate the carboxylic (COOH) groups on the surface of the AuNRBs@ATP@Ag NPs. To the activated AuNRBs@ATP@Ag, SARS-CoV-2 polyclonal antibody (SARS-CoV-2 Ab) (9 μg mL−1 in PBS 7.2) was added and incubated at 4 °C overnight. Following this, the SARS-CoV-2 Ab-labelled AuNRBs@ATP@Ag NPs were washed with PBS (pH 7.2). After washing steps through low-speed centrifugation, the amount of antibody conjugated onto the surface of the NPs was determined by micro bicinchoninic acid (MicroBCA) assay. To block unreacted active esters and prevent non-specific binding, the antibody-conjugated AuNRBs@ATP@Ag was resuspended in a buffer containing 1% (w/v) BSA and 0.1 mM Tris–HCl and incubated at room temperature for 1 h. The conjugate was isolated by centrifugation (1000 rpm, 3 min), and the pellet was resuspended in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 diluted PBS buffer to stabilize the colloid and minimize aggregation.
10 diluted PBS buffer to stabilize the colloid and minimize aggregation.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH4OH
NH4OH![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O (1
H2O (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5) at 60 °C for 30 min in a water bath, rinsed with water and then dried using nitrogen gas. Separately synthesized 500 nm SiO2 from TEOS was then used to modify the surface by submerging the FTO glasses in a 2.5 mg mL−1 ethanol solution of SiO2. Following overnight submersion, the FTO–SiO2 NP surface was washed with ethanol and deionized water to remove the loosely bound SiO2. Then, the surface was further submerged overnight in an ethanol solution of 3-mercaptopropyltrimethoxysilane (MPTMS). The modification with MPTMS (40 mM) introduced surface SH groups, which could be exploited for AuNP adsorption via self-assembly. This was achieved by incubating the modified FTO surface in a saturated solution of citrate capped AuNPs as synthesised above. The citrate capping would readily allow Au adsorption on the surface since it readily exchanged. Then the surface was modified with 11-mercaptoundecanoic acid (MUA) by overnight submersion in an ethanoic solution of MUA (10 mM). This effectively introduced surface COOH, which would allow immuno-labelling via EDC/NHS coupling, depicted in Scheme 1.
5) at 60 °C for 30 min in a water bath, rinsed with water and then dried using nitrogen gas. Separately synthesized 500 nm SiO2 from TEOS was then used to modify the surface by submerging the FTO glasses in a 2.5 mg mL−1 ethanol solution of SiO2. Following overnight submersion, the FTO–SiO2 NP surface was washed with ethanol and deionized water to remove the loosely bound SiO2. Then, the surface was further submerged overnight in an ethanol solution of 3-mercaptopropyltrimethoxysilane (MPTMS). The modification with MPTMS (40 mM) introduced surface SH groups, which could be exploited for AuNP adsorption via self-assembly. This was achieved by incubating the modified FTO surface in a saturated solution of citrate capped AuNPs as synthesised above. The citrate capping would readily allow Au adsorption on the surface since it readily exchanged. Then the surface was modified with 11-mercaptoundecanoic acid (MUA) by overnight submersion in an ethanoic solution of MUA (10 mM). This effectively introduced surface COOH, which would allow immuno-labelling via EDC/NHS coupling, depicted in Scheme 1.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 ratio of CuSO4 to BCA) in a 96-well plate. Following incubation at 37 °C for 30 min, the absorbance at 562 nm for each sample was recorded. SARS-CoV-2 S2 polyclonal Ab concentration in the supernatant sample was determined by comparing its absorbance at 562 nm to the absorbance of the standard BSA sample standard curve and multiplying with the dilution factors. The protein concentration of the initial antibody stock solution and the supernatant following conjugation were measured to determine the mass of antibody successfully bound.
50 ratio of CuSO4 to BCA) in a 96-well plate. Following incubation at 37 °C for 30 min, the absorbance at 562 nm for each sample was recorded. SARS-CoV-2 S2 polyclonal Ab concentration in the supernatant sample was determined by comparing its absorbance at 562 nm to the absorbance of the standard BSA sample standard curve and multiplying with the dilution factors. The protein concentration of the initial antibody stock solution and the supernatant following conjugation were measured to determine the mass of antibody successfully bound.
The characteristic Raman peaks of SARS-CoV-2 viral proteins were relatively weak and indistinguishable from background spectra in our experiments, necessitating the use of a Raman label in the sandwich assay. All SERS spectral data were subsequently post-acquisition processed and analysed using Microsoft Excel (Microsoft, Redmond, WA) and Origin software.
Results and discussion
Design of the SERS sensing platform and detection mechanism
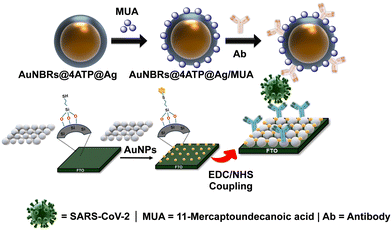
This work reports on the development of a SERS sensing chip for the accurate detection of the SARS-CoV 2 S2 spike protein. The design of the as-reported chip is shown in Scheme 1. The SERS sensing chip can recognise the target protein via antigen–antibody interactions, while the antibody on the SERS nanotag can specifically bind to the captured target SARS-CoV-2 S2, forming a “sandwich” nanostructure. This optical biosensor platform consists of two principal components: (I) SERS nanotag design and (II) SERS sensor chip design.SERS nanotag design
The SERS nanotags employed in this work were constructed from AuNRBs with a conformal Ag shell. The Au core provides chemical stability and tunable plasmonic properties, while the Ag shell significantly enhances localized surface plasmon resonance (LSPR), generating strong electromagnetic fields for SERS signal amplification. The AuNRB cores were synthesized using HEPES as a mild reducing agent, which promotes the slow, anisotropic growth of the branched, raspberry-like nanostructure.30 These branched NPs possess exceptional SERS properties compared to traditional spheres, rods or sphere dimers due to their high-density protrusions and multiscale roughness, which create numerous plasmonic “hotspots” and increase the surface area for reporter adsorption.31 Following synthesis, the 4-ATP Raman reporter was adsorbed onto the AuNRB core surface via a strong Au–SH bond, providing a robust and reproducible spectral signature for quantitative detection. The terminal amine group of 4-ATP also serves to stabilize the interface with the Ag shell. The addition of a conformal Ag shell further amplifies plasmonic activity due to the superior SERS enhancement of Ag, while leveraging the Au core's chemical stability.32To facilitate specific Ab conjugation, the AuNRBs@ATP@Ag surface was further functionalized with 11-MUA. The SH-group of 11-MUA forms a robust self-assembled monolayer via Ag–SH interactions, providing strong surface anchoring. The terminal COOH groups serve as reactive handles for EDC/NHS-mediated covalent coupling with the lysine amine groups of the SARS-CoV-2 S2 polyclonal Ab. 11-MUA was selected over shorter chain carboxylic acids to ensure better spacing and reduce steric hindrance, which is critical for maintaining SERS activity and improving Ab accessibility.
SERS sensor chip design
The sensor chip was constructed on FTO glass, chosen for its transparency, electrical conductivity, and chemical robustness.33 FTO allows for uniform nanostructure deposition, while maintaining optical accessibility for SERS excitation and collection.34 FTO was first coated with a layer of SiO2 to provide abundant hydroxyl groups for subsequent chemical modification and to enhance surface uniformity and stability. Subsequent modification with MPTMS introduced surface SH groups, which served as robust anchors for the deposition of Au nanospheres (AuNPs). The modification with MPTMS introduced surface SH groups, which were exploited for Au adsorption via self-assembly, creating a densely packed, SERS-active surface with multiple plasmonic “hot spots”. Similar to the nanotags, the sensor chip was then functionalized with 11-MUA to introduce COOH groups for the covalent attachment of SARS-CoV-2 S2 monoclonal antibodies, completing the capture surface design. Such design ensures adaptation for multiplex detection.Dual-component Sandwich immunoassay for SERS detection
The developed platform operates as a sandwich immunoassay, providing a highly specific and sensitive detection mechanism. The detection proceeds in a two-step process (Scheme 1). First, the target SARS-CoV-2 S2 spike protein is captured by the Ab immobilized on the AuNP-coated FTO sensor chip. Subsequently, the Ab-functionalized AuNRBs@ATP@Ag SERS nanotags bind to the captured protein, forming a sandwich complex. This design brings the 4-ATP Raman reporter into close proximity with the plasmonically active surfaces of both the nanotags and the sensor chip. The resulting high density “hot spots” at the sandwich interface generate a strong and quantifiable SERS signal. This dual-component design ensures high specificity through dual-Ab recognition and sensitivity via plasmonic amplification from both the SERS nanotags and the AuNP decorated FTO sensor chip. The platform's modular nature can be adapted for multiplex detection by incorporating different Raman reporters and antibodies, thereby offering a versatile tool for rapid point-of-care viral diagnostics.SERS nanotag characterisation
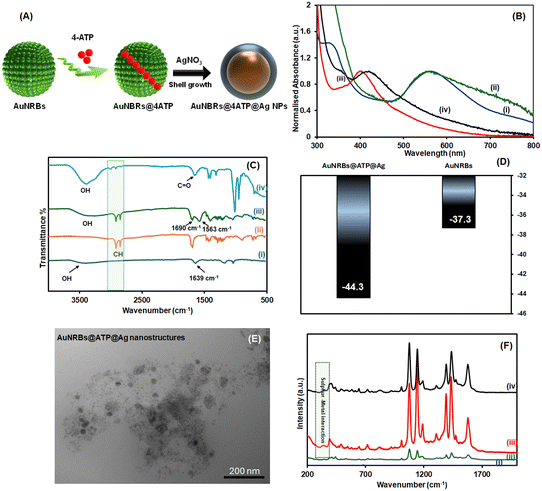
Our SARS-CoV-2 SERS biosensor utilises a core–shell architecture, where a Raman reporter, ATP, is embedded at the Au/Ag interface. UV-vis analysis of the as-synthesised AuNRBs in an aqueous medium exhibited a primary SPR at 562 nm with a shoulder peak at 800 nm (Fig. 1B(i)). This is characteristic of the multi-branched morphology and associated with chain plasmon modes.20 Following 4-ATP assembly, the SPR remained largely unchanged, suggesting a robust Au–SH bond (Fig. 1B(ii)). However, a slight broadening of the shoulder at 800 nm was observed, suggesting minor molecule-induced inter-particle clustering rather than a change in individual particle dimensions (Fig. 1B(ii)). In contrast, the subsequent transition to AuNRBs@ATP@Ag NPs was marked by a significant dampening of the 562 nm Au resonance and the emergence of a dominant Ag peak at ∼400 nm (Fig. 1B(iii)). The disappearance of the core-specific resonance is indicative of a conformal silver shell of sufficient thickness to induce an optical screening effect, wherein the dielectric properties of the Ag shell effectively mask the plasmonic contribution of the underlying Au core.35This structural evolution was further validated by DLS analysis. Ag shell formation was accompanied by a change in DLS sizes, from 51 nm for AuNRBs to 83 nm for AuNRBs@ATP@Ag, consistent with the addition of a shell.19
The stepwise functionalization of the HEPES derived AUNRBs was monitored using FTIR spectroscopy. As can be observed in Fig. 1C, the FTIR spectrum of the AuNRBs is dominated by peaks stemming from HEPES with a broad OH peak from 3000–3600 cm−1, and NH bending vibrations at around 1639 cm−1. Following ATP adsorption, aromatic CH stretching vibrations were subsequently observed, which are attributed to the presence of 4-ATP molecules on the surface of the AuNRBs. Also, no peaks attributable to SH functionality were observed, which implied successful nanostructure decoration with ATP. The introduction of the Ag shell was characterised by the presence of asymmetric and symmetric carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) stretching vibrations around 1690 cm−1 and 1563 cm−1, stemming from the citrate capping agent over the Ag shell. The successful functionalization of MUA was accompanied by the retained C
O) stretching vibrations around 1690 cm−1 and 1563 cm−1, stemming from the citrate capping agent over the Ag shell. The successful functionalization of MUA was accompanied by the retained C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching vibration observed at 1650 cm−1.
O stretching vibration observed at 1650 cm−1.
Zeta potential is a measure of the charges carried by particles suspended in a liquid. It is an important parameter in the fabrication of nanostructures as a high zeta potential implies colloidal stability. which is an important property in biosensor development. The zeta potential for the as-synthesised AuNRBs was found to be −37.3 mV (Fig. 1D). Following ATP adsorption and subsequent growth of the Ag shell, a slight increase in the magnitude of the zeta potential to −44.3 mV was observed (Fig. 1D).
HEPES as a zwitterion buffer and at pH 7.4 carries a negative charge, hence the negative zeta potential observed for the AuNRBs. The negative zeta potential following Ag shell growth may be attributed to the presence of citrate ions, which act as capping agents for Ag. The change in the zeta potential following modification may be attributed to differences in the dielectric constants of the surface agents and hence confirms the successful formation of AuNRBs@ATP@Ag. The high magnitude zeta potential values imply that the as-synthesized nanostructures possess high aqueous dispersibility and high colloidal stability, favourable properties for biological applications.
The transmission electron microscopy (TEM) images show quasi-spherical particles attributed to the synthesized nanostructures (Fig. 1E). In addition to the larger particles, smaller contrast features are also visible in the surrounding region. At the resolution of this image, it is impossible to unambiguously determine whether these smaller features are directly associated with the larger particles or represent separate secondary nanoparticles or surface-deposited species. The obtained DLS average sizes appear slightly larger than what was observed using TEM. This observation is commonly observed in literature,36 and typically explained by the fact that DLS accounts for water layers resulting from solvation as well as adsorbed molecules from the dispersing medium.37 One might consider the possible contribution of aggregates; however, TEM analysis reveals no such evidence. This is likely due to the presence of highly dense surface charges, as indicated by zeta potential measurements, which cause the particles to repel each other and prevent aggregation.
Raman scattering activity and enhancement mechanisms
Fig. 1F shows the Raman spectrum of the AuNRBs, 4-ATP adsorbed AuNRBs (AuNRBs@ATP) and AuNRBs@ATP@Ag. AuNRBs@ATP@Ag was surface functionalised with 11-MUA, which introduced surface COOH groups to allow SARS-COV-2 polyclonal Ab modification via EDC/NHS coupling. The Raman signal of 4-ATP within the AuNRBs@ATP@Ag NP structure is significantly enhanced compared to AuNRBs@ATP alone without the Ag shell. To assess the effect of the respective contribution of the Au core and the Ag shell, the SERS enhancement factor of the nanostructured system was calculated (calculation details in the SI). The calculated enhancement factor (EF) in each case was determined to be 2.43 × 105 for the AuNRBs@ATP and 1.22 × 106 for the AuNRBs@ATP@Ag NPs, lending credence to the superior structure of the core–shell conformation of the as-synthesised AuNRBs@ATP@Ag NPs. The superior EF of the core–shell system over other anisotropic Au NP systems27 may be attributable to minimal aggregation and enhanced hotspot generation between the anisotropic AuNRBs and Ag shell. This enhanced Raman signal behaviour was sustained even following MUA functionalization, ensuring the sustained biosensor performance and is comparable to other core–shell systems.38The strong enhancement can be attributed to several synergistic mechanisms. The primary contribution is a plasmon resonance effect, not unlike the gap-enhanced Raman tag (GERT) mechanism facilitated by the core–shell architecture. This design spatially confines the Raman reporter (4-ATP) within the nanogap between the AuNRB core and the conformal Ag shell, creating highly intense “hotspots” that dramatically increase the scattering cross-section. This is complemented by the multiscale roughness of the AuNRB surface, which promotes the formation of a dense population of these hotspots, further amplifying the effect.19 The enhancement is also reinforced by a charge-transfer mechanism induced by the interaction between the adsorbed 4-ATP molecules and the noble metal surface, which has been observed in other such heterostructures.24 The combination of these physical and chemical enhancement mechanisms provides the SERS system with outstanding amplification capabilities, making it a highly sensitive platform for biosensing.
Moreover, the presence of the characteristic Raman peak of 4-ATP at 1076 cm−1, and the sulphur–metal peak at ∼310 cm−1 (Fig. 1F) is indicative of the successful incorporation of 4-ATP and fabrication of the AuNBs@Ag/MUA NPs.39 The presence of the characteristic peak at 1076 cm−1, which is attributable to the C–S stretching mode and peak at 1580 cm−1, attributable to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching of the aromatic ring (benzene skeletal vibration), also serve as important diagnostic peaks due to their strong SERS intensities. These characteristic peaks remained stable across all composites as stable and efficient Raman reporter peaks and confirm the successful fabrication of AuNRBs@ATP@Ag/MUA NPs.
C stretching of the aromatic ring (benzene skeletal vibration), also serve as important diagnostic peaks due to their strong SERS intensities. These characteristic peaks remained stable across all composites as stable and efficient Raman reporter peaks and confirm the successful fabrication of AuNRBs@ATP@Ag/MUA NPs.
Characterization of the FTO–SiO2–AuNPs sensor chip
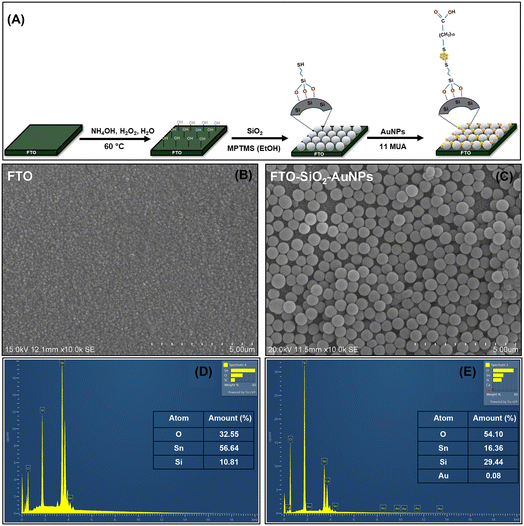
As described in the biosensing principle section, the SERS platform support was prepared by self-assembly of Au onto an FTO support. Fig. 2A shows the fabrication process of FTO–SiO2 arrays by dip coating. SEM (Fig. S2A) and UV-vis (Fig. S2B) of the as-synthesised SiO2 showed the successful synthesis of monodispersed SiO2 with a narrow size distribution, with an average size of 500 nm (Fig. S2C). Similarly, the FTIR results supported the successful preparation of the SiO2 NPs (Fig. S2D). Fig. 2B shows the SEM micrographs of plain FTO and the modified FTO–SiO2–AuNPs. The FTO–SiO2–AuNPs show a large monolayer distribution of SiO2 on the surface of FTO, showing successful SiO2 modification (Fig. 2C). Following SiO2 modification, the surface was further functionalised with MPTMS, which allowed surface SH-groups for AuNP decoration via self-assembly. Although the Au decoration was not as clearly observable at this magnification, the EDX spectra of the bare and modified FTO confirmed the successful decoration of the surface with Au nanoparticles (Fig. 2D and E). This was accompanied by significant reductions in the % Sn compositions between the bare FTO (at 56%) and the modified FTO–SiO2–AuNP surface (at 16%), Fig. 2D and E (inset).X-ray diffraction was used to characterize the crystalline structures of the bare FTO and AuNP-modified FTO surface (Fig. S3). The diffractogram of the bare FTO exhibits patterns largely stemming from the SnO peak. However, following fabrication of the FTO–SiO2–AuNPs, the successful deposition of AuNPs was confirmed by the emergence of well-defined Bragg peaks at 2Θ = 38.1 and 64.5°, corresponding to the (111) and (220) reflections, respectively, characteristic of the face-centred cubic (fcc) structure of Au with reference to PCPDFWIN: Au (#04-0784).40 No pronounced diffraction halo associated with amorphous SiO2 was observed. This is attributed to the ultrathin, near-monolayer nature of the SiO2 coating, whose weak and broad scattering signal is expected to be masked by the strong diffraction peaks of the crystalline FTO substrate and Au nanoparticles. In addition, a noticeable attenuation of the FTO diffraction intensities was observed following surface modification, providing indirect evidence for the successful formation of SiO2 and AuNP overlayers on the FTO surface.
Analytical performance of the SERS biosensor
Following fabrication, the amount of Ab immobilized on the surface of the AuNRBs@ATP@Ag nanotags was quantified using the MicroBCA assay as described above. The initial stock solution had a concentration of 8.22 μg mL−1. After conjugation, the protein concentration remaining in the supernatant was measured to be 3.3 μg mL−1. Based on these results, a total of 4.92 μg of antibody was immobilized per mL of reaction volume, corresponding to a coupling efficiency of 59.85%.The blocked nanoparticles were washed once by centrifugation and resuspended in PBS for storage at 4 °C. The analytical performance of the SERS biosensing platform was evaluated to assess its potential for sensitive and specific detection of the SARS-CoV-2 spike protein. Following the optimised fabrication steps described above, the Ab-functionalised FTO–SiO2–AuNP sensor chip and the complementary AuNRBs@ATP@Ag nanotags were used in sandwich immunoassay detection. The entire detection protocol was optimised, with a 30 min incubation period used for both the antigen capture on the sensor chip and the subsequent binding of the nanotags.
Quantitative SERS analysis and sensitivity
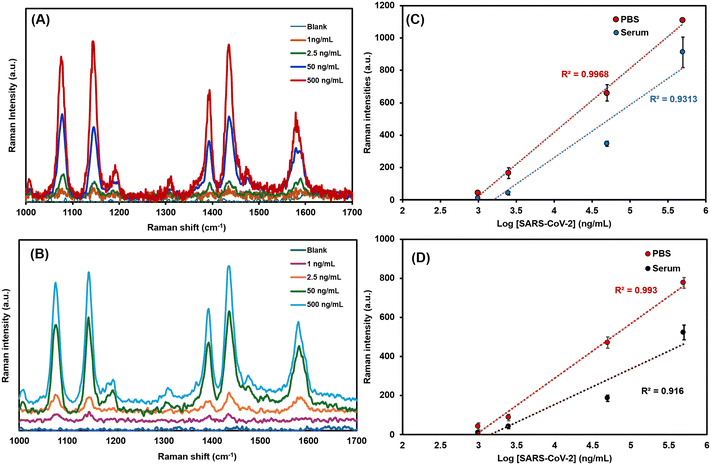
To assess the developed biosensor's sensitivity and quantitative accuracy, we measured the SERS signal of the 4-ATP Raman reporter in response to varying concentrations of the SARS-CoV-2 spike protein. The SERS spectra as shown in Fig. 3A and B exhibit a clear concentration-dependent increase in the 4-ATP Raman signal. The two most prominent and diagnostically important peaks of 4-ATP at 1076 cm−1 (attributed to the C–S stretching mode) and 1580 cm−1 (attributed to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching of the aromatic ring) were used for quantitative analysis.
C stretching of the aromatic ring) were used for quantitative analysis.
A linear relationship was observed between the SERS intensity and the log-function of the SARS-CoV-2 concentration. This was evaluated in both PBS medium and a more complex HSA matrix to simulate real samples. The linear calibration curves are presented in Fig. 3C and D. For the peak at 1076 cm−1, the sensor exhibited a strong correlation coefficient (R2 = 0.99) in PBS across a dynamic range of 1 ng mL−1 to 500 ng mL−1. In HSA, this performance was largely maintained, with an R2 = 0.93 over a similar concentration range. The data from the 1580 cm−1 peak yielded similar strong correlations, with R2 = 0.99 in PBS and R2 = 0.91 in HSA. The use of these two distinct Raman peaks for quantification enhances the robustness and reliability of our assay. By confirming the concentration-dependent signal through two distinct vibrational modes of the same Raman reporter molecule, this approach provides an internal cross-validation of assay performance. This dual-peak strategy helps to mitigate potential signal fluctuations or matrix-related interference that may affect single-peak analyses, particularly in complex biological matrices. The comparable correlation coefficients obtained from both peaks support the reliability of the quantitative analysis. The analytical sensitivity of the SERS biosensor was quantified by calculating the limit of detection (LOD) and limit of quantification (LOQ). A semi-log fitting model was employed to determine these values, which provides a more accurate representation of the binding kinetics observed in such immunoassay systems. The full description is outlined in section S2 of the SI. Using the 1076 cm−1 peak, the LOD was calculated to be 1.07 pg mL−1 in PBS and 1.30 pg mL−1 in HSA. The corresponding LOQ values were 3.57 pg mL−1 in PBS and 4.33 pg mL−1 in HSA. Using the 1580 cm−1 peak, the LOD was 1.04 pg mL−1 in PBS and 1.42 pg mL−1 in HSA. The corresponding LOQ values were 3.47 pg mL−1 in PBS and 4.73 pg mL−1 in HSA.
Mechanism of SERS signal amplification
The high sensitivity of the SERS biosensor is attributed to a dual-component SERS amplification mechanism that generates numerous plasmonic “hotspots”. This is a result of the sandwich immunoassay design, which brings the AuNRBs@ATP@Ag nanotags into close proximity with the AuNPs on the FTO–SiO2–AuNP sensor chip. Upon binding of the target protein, the SERS nanotags are localized onto the capture sensor chip. This controlled self-assembly creates plasmonic nanogaps between the Au and Ag nanostructures, leading to a substantial electromagnetic field enhancement. The 4-ATP Raman reporter molecules are located within these nanogaps, where they experience considerable amplification of their Raman signal. This synergized SERS enhancement from both the nanotags and the sensor chip, facilitated by the target protein, enables the ultrasensitive detection demonstrated by the low LOD values.The observed low LOD values, which fall in the low femtomolar range of the SARS-CoV-2 spike protein, confirm the high sensitivity of our SERS biosensor. The ability of the biosensor to maintain a low LOD in the presence of a complex matrix like HSA highlights its potential for clinical application. The analytical performance of the developed biosensor was also compared with other SARS-CoV-2 biosensor systems in literature, as summarized in Table 1. The LoD of the developed biosensor (18.6 fM) is comparable to that of other Ab-based biosensor systems (14 fM).41 SERS systems based on Au and Ag NPs coupled with magnetic capture have allowed SARS-CoV-2 detection in the femtomolar range.42,43 Morphologically complex nanostructures including MoO3@Au substrates, Nb2C and Ta2C MXenes have been explored as effective SERS substrates, capable of detecting SARS-CoV-2 at the nanomolar and femtogram level, respectively.18,44 However, the developed AuNRBs@ATP@Ag biosensor system exhibits a wide linear dynamic range, which may be attributed to the high surface area and surface-to-volume ratio of the employed nanotags. In addition, the analysis of two Raman label peak signals provides a built-in internal reference, which helps to improve the accuracy of the biosensor.
| Substrate | Target | Limit of detection | Ref. |
|---|---|---|---|
| AuE/CoFeBDCCNH2-CoFe2O4-MOF/SARS-CoV-2 Ab | SARS-CoV-2 | 14 fM | 41 |
| Au-coated MNPs | SARS-CoV-2 | 18 fM | 42 |
| Ag NP-deposited plate | Japanese encephalitis virus (JEV) | 7.6 ng mL−1 | 43 |
| Nb2C and Ta2C MXenes | SARS-CoV-2 | 5 nM | 44 |
| AuNRBs@ATP@Ag on an FTO sensor chip | SARS-CoV-2 | 18.6 fM | This work |
Specificity and selectivity
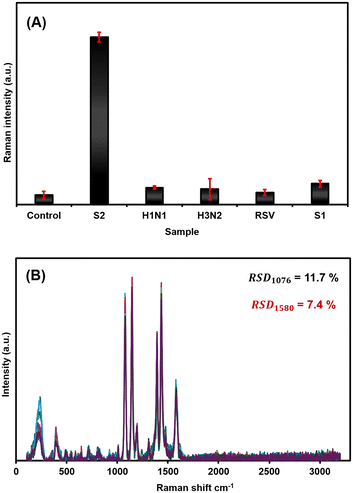
The specificity and selectivity of the developed SERS biosensor were evaluated to assess its ability to uniquely discriminate the target SARS-CoV-2 spike protein within a complex biological matrix. To assess selectivity, control experiments were performed using non-target viral antigens, including influenza viruses A/H1N1 and A/H3N2, and the human respiratory syncytial virus (RSV) fusion glycoprotein. Immunoassays were conducted with these viral pathogens at a concentration of 100 ng mL−1, which lies within the detectable range of the target SARS-CoV-2 S2. As shown in Fig. 4A and S5, the biosensor exhibited a strong and distinct SERS signal for the target S2 protein, while showing negligible SERS signals for any of the other tested antigens. This indicates the high specificity of the immobilized antibodies on both the nanotags and the sensor chip, suggesting preferential binding toward the target SARS-CoV-2 S2 protein and minimising non-specific binding. Furthermore, the nanochip can, in principle, be functionalised with multiple target-specific bioreceptor antibodies, enabling simultaneous detection of viruses. Ongoing work is focused on developing strategies for the detection of multiple pathogens within a single assay, with particular focus on influenza viruses, human respiratory syncytial virus (RSV), and SARS-CoV-2 – key pathogens involved in seasonal “tripledemic” respiratory co-infections. This nanochip system offers key advances in this direction with SARS-CoV-2 serving as a test pathogen in this work to demonstrate the feasibility of this approach.Reproducibility and stability
The reproducibility of the SERS biosensor was evaluated to assess its robustness and consistency. While a dedicated long-term shelf-life study was not conducted, the AuNRBs@4-ATP@Ag nanotags utilize a gap-enhanced Raman tag (GERT)-inspired core–shell architecture. This design is expected to contribute to enhanced signal stability by confining the Raman reporter molecules within the metallic nanogap between the Au core and Ag shell, protecting them from environmental desorption and photodegradation. The conformal Ag shell further acts as a physical barrier, providing additional chemical protection.For reproducibility assessment, SERS mapping on at least 10 randomly selected spots on a single sensor chip was performed to assess intra-batch reproducibility (Fig. 4B and S6). The SERS intensity of the 1076 cm−1 peak was measured, and the relative standard deviation (RSD) was calculated to be 7.5%, indicating good spot-to-spot reproducibility. These results, with an RSD well below the 10% benchmark, support the reproducibility and reliability of the developed biosensing system and detection protocol. A representative back-calculation analysis performed at a concentration of 2.5 ng mL−1 yielded a recovery of approximately 96% (calculation details in the SI, Table S1), indicating good agreement between the nominal and calculated concentrations and supporting the semi-quantitative performance of the sensor in spiked HSA.
Conclusions
This work reports on a sensitive and reliable SERS biosensor for the rapid and specific detection of the SARS-CoV-2 S2 spike protein with potential for accurate diagnostic turnaround times. The platform utilizes a novel core–shell AuNRBs@ATP@Ag nanotag with a gap-enhanced Raman tag (GERT) architecture to achieve signal enhancement and improved signal stability. We herein demonstrate a dual-component hotspot engineering strategy by complementing this novel nanotag with a plasmonically active capture sensor chip, which is a fluorine-doped tin oxide (FTO) surface modified with AuNPs. This combined SERS amplification strategy enabled low limits of detection (LOD) of 1.07 pg mL−1 (15.3 fM) in PBS and 1.30 pg mL−1 (18.6 fM) in HSA, with a rapid assay time of 30 min. The biosensor demonstrated high selectivity against other common respiratory viruses and maintained performance even in complex matrices. This hybrid material-driven approach provides a versatile SERS platform that shows potential for rapid, point-of-care pathogen diagnostics and future adaptability for multiplex (e.g., tripledemic) virus detection. The performance of this biosensor highlights the potential of nanomaterials engineering to address critical challenges in clinical diagnostics.Author contributions
Nnamdi Nwahara: writing – original draft, methodology, investigation, formal analysis, data curation, conceptualization. Saba Niaz: investigation, formal analysis. Muthumuni Managa: writing – review & editing, funding acquisition. Christian I. Nkanga: writing – review & editing, funding acquisition. Oluwasesan Adegoke: writing – review & editing, investigation, formal analysis. Ojodomo J. Achadu: writing – original draft, conceptualization, methodology, investigation, formal analysis, data curation, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting the findings of this study will be made available through the Teesside University Research Repository (PURE) at https://research.tees.ac.uk. All relevant datasets generated and analysed during the current study will be deposited and accessible following publication. Additional data supporting this article have been included as part of the supplementary information (SI) and research materials can also be obtained from the corresponding author upon reasonable request.Supplementary information is available. See DOI: https://doi.org/10.1039/d5sd00185d.
Acknowledgements
The authors gratefully acknowledge financial support from the Royal Society through grant number ICAO/R1/231012 – International Science Partnership Fund (ISPF) – International Collaboration Awards. O. J. A. gratefully acknowledges financial support from the Royal Society through grant numbers RG\R2\232090 – Research Grant. OA acknowledges financial support from the Royal Society of Chemistry (RSC) through E22-7965264821, the Engineering and Physical Sciences Research Council (EPSRC) through EP/X029956/1 and the Royal Society through RG\R2\232243.References
- A. Hill-Ricciuti, E. E. Walsh, W. G. Greendyke, Y. Choi, A. Barrett, L. Alba, A. R. Branche, A. R. Falsey, M. Phillips, L. Finelli, L. Saiman and L. Saiman, Infect. Control Hosp. Epidemiol., 2023, 44, 433–439 Search PubMed.
- S. Iravani, Mater. Adv., 2020, 1, 3092–3103 RSC.
- B. Al Waili, L. Al Yazidi, H. Al Hashami, M. Al Gheilani, F. Al Muharrami, R. Al Jadder, G. Al Hashami, F. B. Alawi, K. Al Maamari, A. Al Mamari, H. Al Khatri and Z. Alhinai, Int. J. Infect. Dis., 2024, 139, 195–201 Search PubMed.
- K. J. Land, D. I. Boeras, X. S. Chen, A. R. Ramsay and R. W. Peeling, Nat. Microbiol., 2019, 4, 46–54 CrossRef CAS.
- S. Li, B. Han, D. Zhou, Y. Gu, B. Li, J. Ma, R. Fu, X. Qi and P. Liu, Anal. Chem., 2023, 95, 2339–2347 CrossRef CAS PubMed.
- K. Leirs, P. Tewari Kumar, D. Decrop, E. Pérez-Ruiz, P. Leblebici, B. Van Kelst, G. Compernolle, H. Meeuws, L. Van Wesenbeeck, O. Lagatie, L. Stuyver, A. Gils, J. Lammertyn and D. Spasic, Anal. Chem., 2016, 88, 8450–8458 CrossRef CAS PubMed.
- H. Jindal, S. Jain, T. K. Suvvari, Lvs. Kutikuppala, S. Rackimuthu, I. C. N. Rocha, S. Goyal and P. R. Radha, SN Compr. Clin. Med., 2021, 3, 2383–2388 CrossRef CAS.
- R. T. Hayden, X. Yan, M. T. Wick, A. B. Rodriguez, X. Xiong, C. C. Ginocchio, M. J. Mitchell and A. M. Caliendo, J. Clin. Microbiol., 2012, 50, 337–345 CrossRef CAS.
- H. Wu, M. Zou, X. Fan, F. Su, F. Xiao, M. Zhou, Y. Sun, F. Zhao and G. Wu, ACS Omega, 2022, 7, 15074–15081 Search PubMed.
- R. Rodriguez-Moncayo, D. F. Cedillo-Alcantar, P. E. Guevara-Pantoja, O. G. Chavez-Pineda, J. A. Hernandez-Ortiz, J. U. Amador-Hernandez, G. Rojas-Velasco, F. Sanchez-Muñoz, D. Manzur-Sandoval, L. D. Patino-Lopez, D. A. May-Arrioja, R. Posadas-Sanchez, G. Vargas-Alarcon and J. L. Garcia-Cordero, Lab Chip, 2021, 21, 93–104 RSC.
- Z. Li, Y. Yi, X. Luo, N. Xiong, Y. Liu, S. Li, R. Sun, Y. Wang, B. Hu, W. Chen, Y. Zhang, J. Wang, B. Huang, Y. Lin, J. Yang, W. Cai, X. Wang, J. Cheng, Z. Chen, K. Sun, W. Pan, Z. Zhan, L. Chen and F. Ye, J. Med. Virol., 2020, 92, 1518–1524 Search PubMed.
- R. N. Kirchdoerfer, N. Wang, J. Pallesen, D. Wrapp, H. L. Turner, C. A. Cottrell, K. S. Corbett, B. S. Graham, J. S. McLellan and A. B. Ward, Sci. Rep., 2018, 8, 15701 Search PubMed.
- L. Guo, S. Lin, Z. Chen, Y. Cao, B. He and G. Lu, Signal Transduction Targeted Ther., 2023, 8, 197 CrossRef CAS.
- A. Z. Mykytyn, R. A. Fouchier and B. L. Haagmans, Curr. Opin. Virol., 2023, 62, 101349 Search PubMed.
- B. C. Dhar, Anal. Bioanal. Chem., 2022, 414, 2903–2934 CrossRef CAS PubMed.
- Z. Li, X. Ding, K. Yin, L. Avery, E. Ballesteros and C. Liu, Biosens. Bioelectron., 2022, 199, 113865 Search PubMed.
- O. Adegoke, K. Oyinlola, O. J. Achadu and Z. Yang, Anal. Chim. Acta, 2023, 1281, 41926 Search PubMed.
- O. J. Achadu, N. Nwaji, D. Lee, J. Lee, E. M. Akinoglu, M. Giersig and E. Y. Park, Nanoscale Adv., 2022, 4, 871–883 Search PubMed.
- Y. Liu, H. Weng, Z. Chen, M. Zong, S. Fang, Z. Wang, S. He, Y. Wu, J. Lin, S. Feng and D. Lin, Biosens. Bioelectron., 2025, 271, 117015 Search PubMed.
- M. Mehta, W. Skinner, B. Gardner, S. Mosca, F. Palombo, P. Matousek and N. Stone, ACS Omega, 2025, 10, 4588–4598 Search PubMed.
- N. G. Khlebtsov, L. Lin, B. N. Khlebtsov and J. Ye, Theranostics, 2020, 10, 2067–2094 Search PubMed.
- J. Langer, D. J. de Aberasturi, J. Aizpurua, R. A. Alvarez-Puebla, B. Auguié, J. J. Baumberg, G. C. Bazan, S. E. J. Bell, A. Boisen, A. G. Brolo, J. Choo, D. Cialla-May, V. Deckert, L. Fabris, K. Faulds, F. Javier García de Abajo, R. Goodacre, D. Graham, A. J. Haes, C. L. Haynes, C. Huck, T. Itoh, M. Käll, J. Kneipp, N. A. Kotov, H. Kuang, E. C. Le Ru, H. K. Lee, J. F. Li, X. Y. Ling, S. A. Maier, T. Mayerhöfer, M. Moskovits, K. Murakoshi, J. M. Nam, S. Nie, Y. Ozaki, I. Pastoriza-Santos, J. Perez-Juste, J. Popp, A. Pucci, S. Reich, B. Ren, G. C. Schatz, T. Shegai, S. Schlücker, L. L. Tay, K. George Thomas, Z. Q. Tian, R. P. van Duyne, T. Vo-Dinh, Y. Wang, K. A. Willets, C. Xu, H. Xu, Y. Xu, Y. S. Yamamoto, B. Zhao and L. M. Liz-Marzán, ACS Nano, 2020, 14, 28–117 CrossRef CAS PubMed.
- T. Nagy-Simon, A. S. Tatar, A. M. Craciun, A. Vulpoi, M. A. Jurj, A. Florea, C. Tomuleasa, I. Berindan-Neagoe, S. Astilean and S. Boca, ACS Appl. Mater. Interfaces, 2017, 9, 21155–21168 Search PubMed.
- Y. Zhang, Y. Qiu, L. Lin, H. Gu, Z. Xiao and J. Ye, ACS Appl. Mater. Interfaces, 2017, 9, 3995–4005 Search PubMed.
- S. Harmsen, M. A. Wall, R. Huang and M. F. Kircher, Nat. Protoc., 2017, 12, 1400–1414 Search PubMed.
- Y. Wang and S. Schlücker, Analyst, 2013, 138, 2224–2238 RSC.
- L. A. Lane, J. Zhang and Y. Wang, J. Colloid Interface Sci., 2024, 663, 295–308 Search PubMed.
- Y. Gu, Y. Zhang, Y. Li, X. Jin, C. Huang, S. A. Maier and J. Ye, RSC Adv., 2018, 8, 14434–14444 RSC.
- J. J. Storhoff, R. Elghanian, R. C. Mucic, C. A. Mirkin and R. L. Letsinger, J. Am. Chem. Soc., 1998, 120, 1959–1964 CrossRef CAS.
- J. E. Ortiz-Castillo, R. C. Gallo-Villanueva, M. J. Madou and V. H. Perez-Gonzalez, Coord. Chem. Rev., 2020, 425, 213489 Search PubMed.
- R. Alvarez-Puebla, L. M. Liz-Marzán and F. J. García De Abajo, J. Phys. Chem. Lett., 2010, 1, 2428–2434 Search PubMed.
- A. I. Pérez-Jiménez, D. Lyu, Z. Lu, G. Liu and B. Ren, Chem. Sci., 2020, 11, 4563–4577 RSC.
- D. Burnat, P. Sezemsky, K. Lechowicz, M. Koba, M. Janczuk-Richter, M. Janik, V. Stranak, J. Niedziółka-Jönsson, R. Bogdanowicz and M. Śmietana, Sens. Actuators, B, 2022, 367, 132145 Search PubMed.
- V. Eskandari, H. Sahbafar, L. Zeinalizad, F. Sabzian-Molaei, M. H. Abbas and A. Hadi, J. Mol. Struct., 2023, 1274, 134497 Search PubMed.
- G. Sirgedaite, M. Talaikis, A. Drabavicius, G. Niaura and M. Lina, Spectrochim. Acta, Part A, 2025, 338, 126160 Search PubMed.
- J. K. Kabeya, N. K. Ngombe, P. K. Mutwale, J. B. Safari, G. G. Matlou, R. W. M. Krause and C. I. Nkanga, Artif. Cells, Nanomed., Biotechnol., 2025, 53, 29–42 Search PubMed.
- C. M. Hoo, N. Starostin, P. West and M. L. Mecartney, J. Nanopart. Res., 2008, 10, 89–96 CrossRef CAS.
- M. G. Ellis, U. Pant, J. Lou-Franco, N. Logan and C. Cao, ACS Appl. Nano Mater., 2023, 6, 10431–10440 Search PubMed.
- N. Nwahara, O. J. Achadu and T. Nyokong, J. Photochem. Photobiol., A, 2018, 359, 131–144 Search PubMed.
- I. Uddin, P. Poddar and N. Phogat, Mater. Focus, 2013, 2, 80–85 Search PubMed.
- S. Palanisamy, L. Y. Lee, C. F. Kao, W. L. Chen, H. C. Wang, S. T. Shen, J. W. Jian, S. S. F. Yuan, Y. A. Kung and Y. M. Wang, Sens. Actuators, B, 2023, 390, 133960 Search PubMed.
- P. C. Guan, H. Zhang, Z. Y. Li, S. S. Xu, M. Sun, X. M. Tian, Z. Ma, J. S. Lin, M. M. Gu, H. Wen, F. L. Zhang, Y. J. Zhang, G. J. Yu, C. Yang, Z. X. Wang, Y. Song and J. F. Li, Anal. Chem., 2022, 94, 17795–17802 Search PubMed.
- M. N. Tripathi, K. Singh, U. Yadav, R. R. Srivastava, M. Gangwar, G. Nath, P. S. Saxena and A. Srivastava, Antiviral Res., 2022, 205, 105382 Search PubMed.
- Y. Peng, C. Lin, L. Long, T. Masaki, M. Tang, L. Yang, J. Liu, Z. Huang, Z. Li, X. Luo, J. R. Lombardi and Y. Yang, Nano-Micro Lett., 2021, 13, 52 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |