Open Access Article
Open Access ArticlecShot: spatial dynamic imaging of cell cycle-dependent miRNA heterogeneity using dynamic DNA patterns
Jie
Zhou
,
Yuwei
Sha
,
Ling'ou
Qin
,
Fengying
Yuan
,
Yu
Ouyang
 *,
Yaqin
Chai
*,
Yaqin
Chai
 *,
Pu
Zhang
*,
Pu
Zhang
 * and
Ruo
Yuan
* and
Ruo
Yuan
 *
*
College of Chemistry and Chemical Engineering, Southwest University, Chongqing 400715, China. E-mail: ouyangyu@swu.edu.cn; yqchai@swu.edu.cn; puzhang@swu.edu.cn; yuanruo@swu.edu.cn
First published on 6th February 2026
Abstract
Profiling cell cycle phase progression and determining how cells respond to external stimuli through miRNA quantification will enable substantial improvements in therapeutic assessment. However, miRNA evaluation is challenging due to its transient nature and the complex cellular environment. Here, we report “cShot” (cell cycle shot), a method for spatial dynamic imaging of miRNA across different cell cycles via dynamic DNA sequences that integrate a pair of competitive probes into a set of static target-binding probes. Compared to traditional irreversible binding probes, cShot enables dynamic binding and dissociation of miRNA through constitutional exchange between two probe sets. Typically, cShot relies on multiple signal outputs, ensuring an accurate signal output through adaptive regulation of the probes. The dynamic behavior of cShot is accompanied by kinetic models, providing a systematic self-checking ability for c-Shot. By tethering DNA tetrahedra to the cShot system and utilizing a hybridization chain reaction, cShot enables real-time monitoring of cell cycle-dependent heterogeneity in synchronized MCF-7 cells under radiomimetic drug stimulation.
Introduction
Cancer remains challenging to treat, requiring clinical drug treatment and an early diagnosis due to its heterogeneity which is manifested in various tumor properties.1–4 A valid method to evaluate the treatment effect of drugs in heterogeneous cancers may greatly promote the recovery of patients. Drug-treated or drug-tolerant subpopulations of cancer cells show typical characteristics in growth, metabolic state and gene regulation pathways.5–8 Traditional methods that monitor cancer cells at the absolute level may lead to inaccuracies in cancer diagnosis and drug assessment.9–12 Therefore, the ability to explore the heterogeneity through intact cellular cycle phases, including growth behaviors, is thought to be a major challenge to achieving shortened therapy processes. It is worth noting that the cell cycle progression is regulated by successive transcription and post-transcriptional pathways.13–15 Among these upstream and downstream pathways, microRNA (miRNA) regulators are essential in almost all biomechanisms.16–19 That is to say that the expression of specific miRNA changes in an orderly way with the cell cycle phases. Therefore, quantification of miRNA with high fidelity in order to map such stages of cell cycle progression is a great challenge.MiRNA molecules in living systems are dynamically distributed, and are present in direct or indirect transient correlation with neighboring biomolecules.20–24 Previous studies have investigated programmable miRNA recognition molecules that quantify the absolute parameters of miRNA expression.25–28 Yet, these methods do not provide information about dynamic changes in miRNA expression due to the irreversible binding between the recognition molecule and the target miRNA.29–32 Recent studies have developed mechanical DNA devices for recognizing and sensing miRNA in response to external stimuli, e.g. light irradiation,33–38 thermal stimulation,39,40 chemical reactions41–43 or interactions with a molecular fuel.44–48 In principle these devices are able to be engineered into a miRNA-quantifying device. However, these DNA devices rely on single signal outputs that are derived from the DNA conformational probe itself. In complex cellular environments, degradation of this probe may compromise dynamic regulation of the DNA structure and hinder real-time detection of the target (Fig. S1A). Another key issue of dynamic miRNA sensing rests, however, on the reverse reconfiguration of the DNA molecule after binding to the miRNA, which greatly limits the promise of application in dosage assessment during disease treatment. Our previous work reports an sequence-structure bispecific RNA aptamers to realize the dynamic distribution of miRNA in living cells.49 However, the relatively dim fluorescence from the labeled molecules and the limited functional integration with the enzyme-free nucleic acid amplification strategy limit the sensitivity of miRNA visualization.
Nucleic acid-based constitutional dynamic networks, CDNs, were recently developed by Willner's laboratory as dynamic supramolecular assemblies that mimic complex living networks in nature.50–55 The simplest [2 × 2] CDN consists of four equilibrated constituents AA′, AB′, BA′ and BB′. The signal-triggered stabilization of one of the constituents, e.g., AA′, leads to the dynamic reconfiguration of the CDN into a new equilibrated CDN where the stabilized constituent, AA′, is up-regulated at the expense of AB′ and BA′ that are down-regulated. The up-regulation of constituent AA′ is accompanied by the concomitant up-regulation of the constituent BB′, that does not share components with AA′, as a result of the triggered separation of AB′ and BA′. By integrating a conformational DNA probe into a dynamic network comprising multiple signal probes, the system can collectively reflect dynamic changes of the probe target. Thus, even if one probe is degraded, the remaining probes can still facilitate structural switching and ensure effective target detection, Fig. S1B. Owing to the anti-degradation capability and multi-signal output performance of the DNA network, this system not only enables dynamic monitoring of the target but also enhances detection accuracy when compared to single recognition probes. However, all stimuli-responsive systems are operated in homogenous aqueous phases and include constituents with flexible two-dimensional structures. Recently, Willner and co-worker Zhou developed a catalytic hairpin assembly (CHA)/entropy-driven evolution of the CDN inside cells where they realized the sensing and imaging of miRNA by means of fluorescence (Förster) resonance energy transfer (FRET) through the labeling of fluorophores at the ends of the constituents strands.56 However, in all these systems, amplification of the signal of the regulated CDN for dynamic and sensitive imaging of the cells is an important challenge to overcome. In particular, the applications of these systems for monitoring of intact cell cycle progressions in synchronized cells are scarce. This calls for the need to design intracellular dynamic sensing molecules that are spatially selectively activated by endogenous miRNA to reveal the heterogeneity of the cell cycle phases under treatment with auxiliary stimuli.
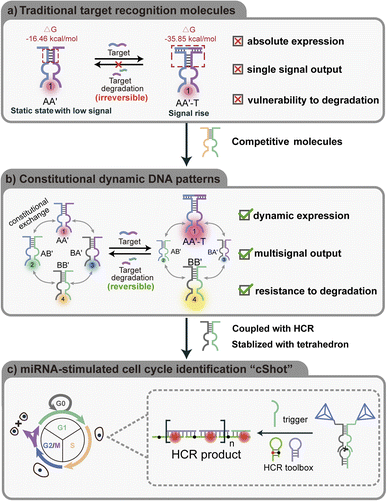
Here, we report a constitutional dynamic DNA pattern as a sensitive and specific tool for the detection of intracellular miRNA to reveal the heterogeneity across cell cycle phases, a process termed “cShot”. Compared to traditional recognition molecules, such as the dimer of DNA molecules A and A′ in Scheme 1a, the fixed binding affinity between the target and the free tether on the molecules enables determination of the absolute, static expression level of miRNA by a single output signal. As shown in Scheme 1b, by ingenious addition of a pair of competitive molecules, B and B′, that share a complementary domain with A and A′, these patterns of molecules achieve thermodynamic equilibrium by generating four constituents AA′, AB′, BA′ and BB′. The engineering design of the CDN reaction system was assessed and verified through NUPACK simulations (Fig. S3). It should be noted that the shared domain (marked with a square) enables dynamic constitutional exchange among these patterns of molecules. These molecules reconstitute into another thermodynamic equilibrium when an intracellular miRNA binds specifically to the exposed tether associated with the recognition molecule dimer, AA′, thus inducing the stabilization of AA′-miRNA by taking the constituents A and A′ from AB′ and BA′. Impressively, when the target content is reduced, the recognition molecules are able to reconfigure through the competition of the complementary shared domains associated with the antergic constituents owing to the high free energies of less bindable target. By embedding Mg2+ ion-dependent DNAzyme units into the constitutional dynamic patterns, four independent output signals are generated from each constituent to quantify the dynamic changes of the target concentration. We utilized Mg2+ ion-dependent DNAzyme units conjugated to the CDN to guide the generation of hybridization chain reaction (HCR) amplifiers, Scheme 1c, and to create a system that acts as a sensitive, dynamic model assembly for spatial selective sensing and imaging of miRNA in synchronized cells. Finally, the dimer DNA tetrahedra that were extended from the constituent strands of the CDN are then integrated as [2 × 2] constitutional dynamic assemblies into different cell lines to improve the cell permeability of the reaction network. Compared to the naked CDN constituents, the tetrahedra-functionalized CDN constituents show improved cell permeation, less degradation, and more sensitive imaging observations. We use miRNA mimics and miRNA inhibitors to show the great capabilities of cShot for imaging the dynamic changes of intracellular targets. Furthermore, cShot can be utilized to assess the efficacy of radiomimetic drug neocarzinostatin in synchronized MCF-7 cell lines, thereby preventing unnecessary cellular damage resulting from overtreatment.
Results and discussion
Mechanisms of nucleic acid conformational transitions
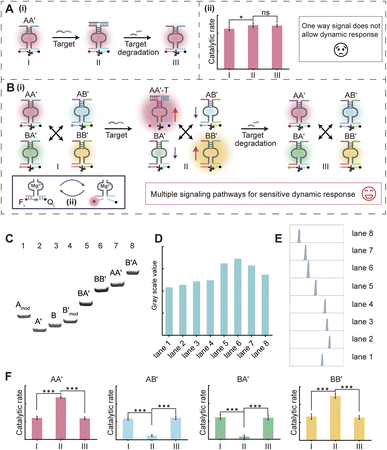
Current DNA sensing analysis methods mostly depend on a single signal output, where the signal output is derived from the binding of the target to a recognition probe. When the biomolecule target, miR-21, is degraded or binds to other biomolecules due to high affinity, the output signal remains in the “on” state due to the absence of a substance capable of degrading the detection system, thereby impeding the dynamic monitoring of the target (Fig. 1A, panel i). Additionally, signals generated by a traditional target recognition molecule are univariate, which not only fail to reflect the dynamic concentration changes of the target but also make it difficult to accurately determine its concentration (Fig. 1A, panel ii). Here, we introduced a CDN system to address these limitations. Based on the pairing of the AA′ constituent (which served as the target recognition probe, ΔG = −21.87 kcal mol−1) with a competitive probe BB′ four constituents AA′, AB′, BA′ and BB′ were generated with shared sequences in the stem domain under thermodynamic equilibrium. The stabilization of AA′ upon target binding to its recognition domain leads to the dehybridization of the A and A′ strands within the AB′ and BA′ constituents, thereby reducing their concentrations, whereas the concentration of the corresponding BB′ constituent increases. Upon subjecting an anti-strand that is fully complementary to the binding domain associated with AA′ (target degradation process), the system dynamically returns to its initial equilibrium state through the hybridization of A (association with AB′) and A′ (association with BA′), thereby realizing the reversible transition of the detection system (Fig. 1B). Firstly, we validated the successful assembly of each constituent, AA′, AB′, BA′ and BB′, through gel electrophoresis. As can be observed in Fig. 1C, lanes 1–4 correspond to the individual DNA single strands Amod, A′, B and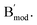 Lanes 5–8 represent the successful hybridization of B and A′, B and B′, A and A′, B′ and A. Fig. 1D and E show the quantitative gray scale values and total gray scale analysis of Fig. 1C. It should be noted that as the DNA molecular weights of the four constituents in the original design were relatively close, we extended the structures of A and B′ with free tethers to differentiate the migration rate of these bands, and this modification did not impact the composition of the original components. The normalized intensity of the catalytic rate of the DNAzyme reporters associated with the corresponding constituents is shown in Fig. 1F. The corresponding dynamic reassembly of these constituents was observed using an integrated Mg2+-dependent DNAzyme (Fig. 1B, panel ii). The concentration of Mg2+ was optimized at 20 mM to ensure robust catalytic signaling of the cShot system (Fig. S4). Column I represents the initial state of the system. After binding of the target to AA′ (column II), the concentration of AA′ and BB′ was up-regulated, and the AB′/BA′ concentrations were down-regulated. Specifically, after the degradation of the target (column III), the content of AB′ and BA′ was up-regulated, and the AA′/BB′ concentrations were down-regulated. Time-dependent fluorescence changes generated by the DNAzyme units associated with columns I, II and III are shown in Fig. S2 in the SI. The transient concentration of the cleaved substrate serves as an indicator of the DNAzyme's catalytic turnover rate, which directly correlates with the dynamics of RNA concentration. Comparison of the catalytic rate of each constituent in an independent AA′ sensing system (Fig. 1A, panel ii) and in an integrated dynamic network (Fig. 1F), the CDN-based sensing probe is able to reflect the dynamic changes that occur in the target. Furthermore, the four signals generated through the dynamic network can synergistically reflect changes in target concentration, providing more accurate detection than a single signal response.
Lanes 5–8 represent the successful hybridization of B and A′, B and B′, A and A′, B′ and A. Fig. 1D and E show the quantitative gray scale values and total gray scale analysis of Fig. 1C. It should be noted that as the DNA molecular weights of the four constituents in the original design were relatively close, we extended the structures of A and B′ with free tethers to differentiate the migration rate of these bands, and this modification did not impact the composition of the original components. The normalized intensity of the catalytic rate of the DNAzyme reporters associated with the corresponding constituents is shown in Fig. 1F. The corresponding dynamic reassembly of these constituents was observed using an integrated Mg2+-dependent DNAzyme (Fig. 1B, panel ii). The concentration of Mg2+ was optimized at 20 mM to ensure robust catalytic signaling of the cShot system (Fig. S4). Column I represents the initial state of the system. After binding of the target to AA′ (column II), the concentration of AA′ and BB′ was up-regulated, and the AB′/BA′ concentrations were down-regulated. Specifically, after the degradation of the target (column III), the content of AB′ and BA′ was up-regulated, and the AA′/BB′ concentrations were down-regulated. Time-dependent fluorescence changes generated by the DNAzyme units associated with columns I, II and III are shown in Fig. S2 in the SI. The transient concentration of the cleaved substrate serves as an indicator of the DNAzyme's catalytic turnover rate, which directly correlates with the dynamics of RNA concentration. Comparison of the catalytic rate of each constituent in an independent AA′ sensing system (Fig. 1A, panel ii) and in an integrated dynamic network (Fig. 1F), the CDN-based sensing probe is able to reflect the dynamic changes that occur in the target. Furthermore, the four signals generated through the dynamic network can synergistically reflect changes in target concentration, providing more accurate detection than a single signal response.
CDN system reconfiguration and kinetic validation
The adaptive triggered transitions of the CDN “S” to form either CDN “X” or “Y” caused by subjecting the CDN to either miR-21, that is complementary to the binding domain of AA′, or a competitive analogue (ca), that is complementary to the binding domain of AB′, are illustrated in Fig. 2A. Next, in order to attain accurate quantification of each constituent of the CDN system, the time-dependent fluorescence changes of the various concentrations of the different DNAzyme reporter units associated with the individual structural constituents of the CDN system were investigated. As shown in Fig. 2B, it was observed that the rates of catalytic activity of the DNAzymes exhibited an upward trend when the concentration of the respective constituents in the CDN system increased. The results were also analysed using the standard curve method to build calibration curves, which can be used to determine the sample concentration. The appropriate calibration curves of each constituent under different concentrations of DNAzyme reporter units are shown in Fig. S5. Fig. 2C, curve i, shows the time-dependent fluorescence variations produced by the DNAzyme reporter units related to the CDN “S” where the four constituents AA′, AB′, BA′, BB′ exist in an equilibrium state. As shown in Fig. 2A, subjecting the CDN to the target miR-21 (T) disturbs the equilibrium among the four constituents. Specifically, there is a significant up-regulation of the concentrations of AA′ and BB′, which increase by 97% and 98%, respectively (Fig. 2C, curve ii), in comparison to their expression levels in CDN “S”. Conversely, the concentrations of AB′ and BA′ experience a down-regulation of 79% and 81% (Fig. 2C, curve ii), respectively. Similarly, curve iii represents the time-dependent fluorescence changes generated by the DNAzyme reporter units associated with the transition of CDN “S” to CDN “Y” triggered by the competitive analogue (ca). The stabilization of AB′ in turn results in an up-regulation of BA′, while AA′ and BB′ experience a down-regulation. Compared to CDN “S”, the constituents AB′ and BA′ are up-regulated by 93% and 91%, respectively (Fig. 2C, curve iii), while the constituents AA′ and BB′ are down-regulated by 89% and 82%, respectively. According to the calibration curves derived from the independent constituent under various concentrations, a summary of the concentrations of the constituents in CDN “S”, “X” and “Y” is represented in Fig. S6. Furthermore, a kinetic model was developed based on the key chemical reactions among the constituents and the external DNA inputs to simulate the time-dependent concentration changes of the constituents in response to DNA triggers. As shown in Fig. 2D and E, the kinetic validation studies demonstrate concentration dependent reconfiguration dynamics of CDN “S” under DNA stimuli of (i) 1 µM T/ca and (ii) 0.5 µM T/ca. Experimental fluorescence trajectories (discrete points) show excellent agreement with the mechanistic kinetic model predictions (solid curves, R2 > 0.95), confirming predictable state transitions in this dynamic DNA system. Full system reconfiguration between the CDN states was achieved within 15 min, as quantified by time-to-threshold analysis. The rate constants (k) for the transitions from CDN “S” to “X” or “Y,” as simulated by the kinetic models, are listed in Tables S1 and S2. The corresponding reaction equations (Fig. S7 and S8) and time-dependent fluorescence signals (Fig. S9 and S10) are provided in the SI. The results demonstrate that the proposed constitutional dynamic networks (CDNs) exhibit exceptional sensitivity toward the target miR-21, facilitated by the amplification effect of the Mg2+-dependent DNAzymes. This process drives the system to re-equilibrate into a new state amenable to subsequent analysis. To further demonstrate the feasibility of the CDN system, the four constituents of CDN “S”, “X” and “Y” were characterized using gel electrophoresis. The gel analysis of the triggered transitions of CDN “S” to either CDN “X” or “Y” are de-scripted in Fig. 2F and G. Lanes 1–4 represent the single constituents, BB′, AA′, AB′, and BA′, respectively, and lane 5 is the annealing solution of the four DNA strands A, A′, B and B′, representing the CDN “S” state. All four constituents exhibited distinctly recognizable bands in lane 5, aligning precisely with the positions of the individual constituent bands. Lane 6 and lane 8 represent the complexes of components AA′/AB′ with T and ca, respectively. Lanes 7 and 9 in Fig. 2F represented the CDN system after the addition of the T and ca, respectively. The quantitative gray scale values of Fig. 2F are shown in Fig. S11. Notably, significant changes in the components of the CDN were observed following the introduction of the targets. The bands corresponding to the components antagonistic to the components that increase in concentration were nearly absent. This observation strongly indicates that the CDN system is capable of undergoing autonomous compositional changes and state switching in response to the addition of specific targets. The gel electrophoresis results provide clear visual evidence of the dynamic behaviour of the CDN system, showcasing its ability to respond to external cues by altering its composition and state.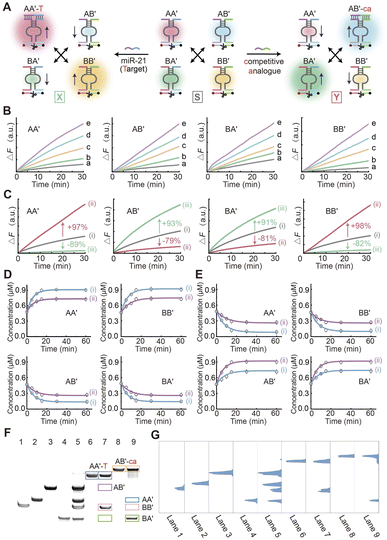 | ||
| Fig. 2 (A) Schematic illustration of the transition of CDN “S” into CDN “X” in the presence of miR-21 and the transition of CDN “S” into CDN “Y” in the presence of the competitive analogue. (B) Time-dependent fluorescence changes generated by the DNAzyme units associated with each constituent at different concentrations (a = 0.2 µM, b = 0.4 µM, c = 0.6 µM, d = 0.8 µM, e = 1.0 µM). (C) Time-dependent fluorescence changes generated by the DNAzyme units associated with the different constituents, gray curve represents CDN “S”, red curve represents CDN “X” after CDN “S” is triggered by miR-21, green curve represents CDN “Y” after CDN “S” is triggered by the competitive analogue. (D) Temporal concentration changes of the constituents during the miR-21-triggered reconfiguration of CDN “S” to CDN “X” at (i) 1 µM and (ii) 0.5 µM. Experimental data are shown as points, and simulated data as solid lines, based on the kinetic model described in Fig. S5. (E) Temporal concentration changes of the constituents during the ca-triggered reconfiguration of CDN “S” to CDN “Y” at (i) 1 µM and (ii) 0.5 µM. Experimental data are shown as points, and simulated data as solid lines, based on the kinetic model described in Fig. S6. (F) PAGE analysis of the CDNs presented in Scheme 1. Lanes 1–4 correspond to the intact individual constituents, 1 µM each, BB′, AA′, AB′, and BA′. Lane 5 is the separate constituents of CDN “S”. Lane 6 is a reference showing the intact constituent AA′ stabilized by miR-21, AA′-T (1 µM). Lane 7 is the separate constituents associated with CDN “X”. Lane 8 is a reference showing the intact constituent AB′ stabilized by competitive analogue, AB′-ca (1 µM). Lane 9 is the separate constituents associated with CDN “Y”. (G) The total gray scale analysis of part (F). Error bars indicate the mean ± standard deviation (n = 3). | ||
CDN-integrated HCR for sensitive miRNA detection
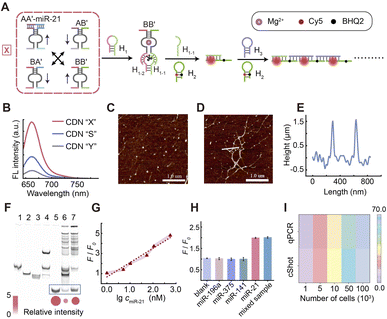
To help advance the delivery of such a CDN system into use in cell sensing research that requires high sensitivity and anti-degradation ability, we note that the engineered Mg2+-ion-dependent DNAzyme units conjugated to the constituents shows several advantageous functionalities. (i) The DNAzyme units act as reporter units for the quantitative evaluation of the compositions of the different CDNs through the DNAzyme-catalyzed cleavage of the different fluorophore/quencher (Fi/Qi)-modified substrates associated with the different reporter units. (ii) The DNAzyme-catalyzed cleavage of the corresponding substrates enables the amplified conversion of the constituent-associated DNAzyme to a cyclic substrate. (iii) Because of the catalytic function of the DNAzyme elements, the structure can translate into the corresponding DNA hairpins that dictate enzyme-free nucleic acid amplification in analytical sensors. (iv) Mg2+ is proven to be abundant inside living cells, which enables the CDN to be adapted for cellular imaging analysis.57–59 In view of this, based on proper delivery of the CDN into living cells, dynamic detection of the changes in intracellular biomarkers associated with the dynamic networks and the sensitive imaging of biomarkers related to the dynamic network-guided enzyme-free amplification, such as the hybridization chain reaction (HCR), could be envisaged.Next, we employed the cShot to detect the target miR-21 in vitro by integrating HCR to increase the sensitivity of the sensing platform. As shown in Fig. 3A, after up-regulation of constituent AA′ by the target miR-21, the subsequently up-regulated constituent BB′ is capable of cleaving hairpin H1 by using the Mg2+-dependent DNAzyme domain, and the resulting product H1–1 can open hairpin H2 by hybridizing to the exposed tether in H2. The opened H2 recognizes the exposed domain of H3, thereby initiating the subsequent HCR reaction through the cross-opening of H2 and H3. The feasibility of a CDN system to initiate HCR responses was evaluated through fluorescence spectra by functionalization of fluorophore/quencher in the stem of the hairpins, atomic force microscopy (AFM) and gel electrophoresis. First, we modified the stem of H2 with a Cy5 fluorophore and a corresponding quencher BHQ2 to track the activation of the HCR microwires by the different states of the CDN. As shown in Fig. 3B, the CDN “S”-induced generation of the HCR wires results in the observation of the fluorescent spectrum of Cy5 because the appropriate amount of BB′ was digested by H1 causing the cross-opening of H2 and H3, leading to the opening of H2 independently of the Cy5 and BHQ2 embedded in the stem of H2. Accordingly, when the miR-21 bound to AA′ (CDN “X”), the up-regulation of BB′ caused increased HCR generation and a the strong fluorescent signal for Cy5. However, when the ca bound to AB′ (CDN “Y”), the down-regulation of both AA′ and BB′ resulted in a decrement in HCR generation, thereby resulting in minimal Cy5 fluorescence intensity. Fig. 3C presents AFM characterization of the H1, H2 and H3 mixture in the absence of the CDN system. No discernible linear HCR products were detected in the sample. Meanwhile, Fig. 3D and E exhibit the specific shape of the HCR product triggered by CDN “X”, where a large number of assembled linear polymers appear with a thickness of 1.8 nm, indicated the successful assembly of H2 with H3. As can be seen in Fig. 3F, complementary quantitative electrophoretic separation of the different states of the CDNs was conducted. The distinct bands observed in lane 1, lane 2, and lane 3 represent H1, H2, and H3, respectively. Lane 4 corresponds to the mixture of H1, H2 and H3. The three separated bands indicate the independence of these hairpins in the absence of the trigger, H1–1. Lane 5 corresponds to a commercial synthesized H1–2 fragment and acts as a reference band for the digested H1. Upon the introduction of the CDN system to the hairpin pool, both lanes 6 and 7 display bands with higher molecular weights, signifying the presence of the HCR product. Additionally, a band appeared at the same position as the reference strand in lane 5 in both lanes 6 and 7, which indicates the successful cleavage of H1 by the Mg2+-dependent DNAzyme associated with BB′. Notably, the bands associated with H1–2 in lane 7 were more prominent, indicating greater production of the HCR derived from CDN “X” than from CDN “S”. Relative intensity analysis of the three bands corresponding to H1–2 in Fig. 3F further illustrates that the elevated content of BB′ in the CDN “X” state resulted in a more substantial production of cleaved products. The above results confirm that the proposed sensing system based on CDN could effectively achieve the detection of target miRNAs. The quantitative performance of target miR-21 detection based on the dynamically triggered adaptive re-equilibration of the CDN-coupled HCR system was evaluated under optimized experimental conditions: reaction time, 2 hours; H2/H3 concentration, 2 µM (Fig. S12). Under these optimized experimental conditions, we applied different concentrations of target miR-21 to investigate the sensitivity of the CDN-based sensing system. Based on Fig. 3A, we recorded the Cy5 fluorescence intensity generated by HCR when using different target concentrations. The results in Fig. 3G demonstrate that the fluorescence intensity increases when the concentration of miR-21 increased in the range of 1 nM to 500 nM. The fluorescence changes of the aqueous solution and the logarithmic concentration of miR-21 exhibited a good linear relationship that obeys the equation I = 1.421 lg cmiR-21 + 0.661, in which I is the relative fluorescence intensity of F/F0, while F and F0 are the fluorescence intensity of CDN “X” and CDN “S”, respectively. The corresponding original fluorescence spectra related to Fig. 3G are shown in Fig. S13. Hence, the proposed cShot exhibits exceptional specificity for the target miR-21 in the presence of foreign miRNAs, including miR-196a, miR-375 and miR-141 (Fig. 3H). The corresponding original fluorescence spectra related to Fig. 3H are shown in Fig. S14. Impressively, the application of our assay system for the detection of miR-21 in real samples (cell lysates of MCF-7 overexpressed with miR-21 and MHCC97L with low expression of miR-21) yielded reasonable results (Fig. S15), and the results were consistent with the gold standard method for miRNA quantification, the quantitative real-time polymerase chain reaction, qPCR (Fig. 3I).
Tetrahedron-stabilized cShot for intracellular miRNA imaging
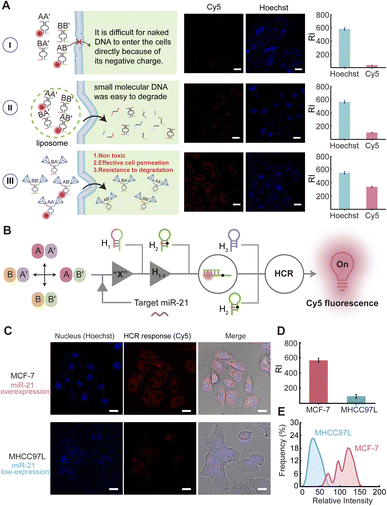
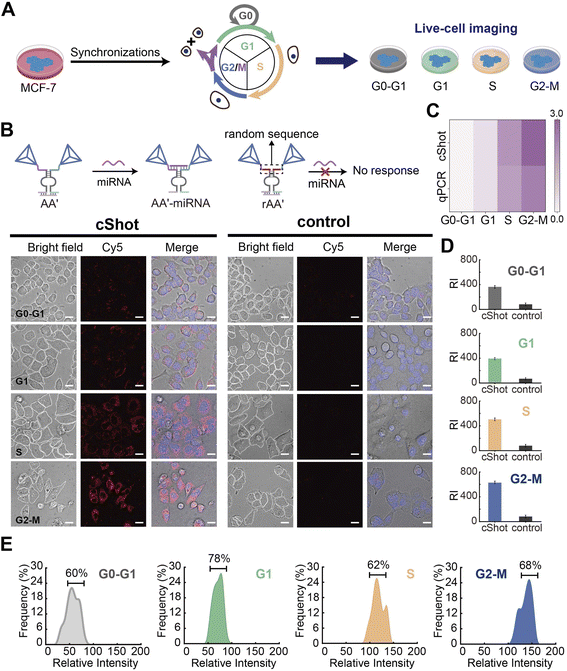
Realizing that CDN “S” can be reconfigured by the target miRNA to form CDN “X” and can be used for the sensitive analysis of miRNA by integrating an amplified HCR strategy, we applied the CDNs-guided HCR to “snapshot” the intracellular miRNA in living cells and across different cell cycles, a process termed as cShot. We utilized breast cancer cells MCF-7 and hepatocellular carcinoma cells MHCC97L as the model systems for our study (the cell lines were purchased from the American Type Culture Collection), with miR-21 showing high expression in the MCF-7 cells, and MHCC97L cells demonstrating relatively low expression levels of miR-21. All constituents of the cShot were transfected into the cells via commercial liposomes. However, despite the high expression of miR-21 in these cancer cells, we did not achieve recognizable fluorescence imaging results by transfecting non-modified CDN constituents (the corresponding confocal microscopy images are shown in Fig. S16). This deficiency can be attributed to the vulnerability of the imported CDN system, which comprises short, single stranded DNA susceptible to degradation within the complex intracellular milieu. Therefore, we extended these strands of these constituents with assembled DNA tetrahedra to stabilize the CDN system (see Fig. S17), enhancing its resistance to degradation when applied to living cells. The high permeability of the tetrahedra into cells was utilized, thereby enhancing the system's cellular permeation efficiency, as can be seen in Fig. 4A. As a comparison, a group of naked DNA constituents (Fig. 4A, panel I) and naked constituents loaded in liposomes (Fig. 4A, panel II) were used to treat living cells to investigate the cellular permeation. As can be observed from Fig. 4A, panel III, the DNA tetrahedra-functionalized CDN system shows better cellular permeation ability than that of the naked constituents or naked constituents loaded in liposomes. The normalized fluorescent intensities of the respective cellular confocal images are shown in Fig. 4A. The successful construction of the four tetrahedra constituents was initially verified through native gel electrophoresis (Fig. S18). The biostability of the cShot system was verified in serum-containing medium, showing minimal degradation over 24 h (Fig. S19), which ensures the fidelity of the dynamic response in living cells. It should be noted that four assistant strands, a1, a2, a3 and a4 were engineered to mask the recognition sites within the four constituents respectively, preventing interactions between hairpin H1 and BB′ (Fig. S20). This effectively suppressed the DNAzyme cleavage activity of BB′ in CDN “S”, reducing the background noise and significantly enhancing cellular miRNA imaging performance. The blockage of the four constituents is required because we observed that when CDN “S” was used for intracellular miRNA imaging, its inherent cleavage activity led to elevated background signals, particularly in cells with low miRNA expression. In contrast, the active CDN “X” state (designed for high miRNA-expressing cells) exhibited minimal increase of the fluorescent imaging signal (Fig. S21). The blocking strands thus improved miRNA imaging sensitivity. Strand a4 was specifically used to block BB′ that recognizes H1 to induce the HCR product, while the other three strands a1, a2 and a3 were engineered to ensure the thermal dynamic balance of the CDN system. Subsequently, the regulatory transitions of the tetrahedra-based CDN across different states in the presence of the target miR-21 was tested. As depicted in Fig. S22, the tetrahedra-based CDN is capable of transitioning from the “S” state to the “X” or “Y” state when the target is present. However, the modulation amplitude is slightly attenuated compared to the non-modified CDN system, which may be because the tetrahedra framework introduces steric resistance to CDN conformational switching and causes shielding of the auxiliary DNA strand. The highly sensitive fluorescence imaging of intracellular miR-21 derived from cShot is shown in Fig. 4B and C. MCF-7 cell lines were employed as miR-21-overexpressed samples and MHCC97L cell lines were employed as a negative control. According to the confocal fluorescence images and quantitative mean fluorescence intensity analysis (Fig. 4D), a strong red fluorescent signal from the Cy5 associated with the HCR product was observed in the MCF-7 cells. In contrast, a much weaker red fluorescent signal was observed in the MHCC97L cells. The expression levels of miR-21 differed significantly between the cancer cells (MCF-7 and MHCC97L), leading to substantial changes in fluorescence intensity. To further characterize the integrated fluorescence intensity, we analyzed five sets of confocal fluorescence images from these two cell lines (MCF-7 and MHCC97L, n = 50 cells per line). The fluorescence intensity distribution frequency was quantified for each cell population (Fig. 4E). Histogram analysis revealed heterogeneous distributions of miR-21 fluorescence intensity between the two cell lines. The integrated fluorescence intensities within the histogram showed positive correlation with the relative integrated fluorescence intensities generated from the different cell lines in Fig. 4C. The observed heterogeneity in miR-21 content in the MCF-7 cells may be attributed to the presence of cells at distinct cell cycle phases.34 Therefore, we decided to further apply the developed cShot method to evaluate miR-21 levels in cells during different cell cycle phases, which will validate its methodological capability in living cell analysis across the cell cycle progression.Cell cycle-resolved miRNA imaging via cShot
The cell cycle progression, regulated through sequential transcriptional and post-transcriptional pathways, is accompanied by dynamic fluctuations in intracellular miRNA levels, which exhibit cell cycle-dependent variations.60,61 Monitoring these cell cycle-specific miRNA signatures offers a promising approach for tracking disease progression, as distinct miRNA expression patterns may serve as molecular indicators of pathological states across various diseases.62 Accordingly, MCF-7 cells were selected as the model system and were synchronized at distinct cell cycle phases (G0–G1, G1, S, and G2–M) using established protocols63 (Fig. 5A). The developed cShot platform was employed for miR-21 imaging in these synchronized cell populations, with control experiments performed by replacing A and A′ with scrambled sequences (rAA′) that lack the specific domain for miR-21 recognition (Fig. 5B). Confocal fluorescence microscopy images demonstrated that cShot generated well-resolved fluorescence signals in MCF-7 cells across different cell cycle phases. In contrast, the control CDN system (lacking the miR-21 recognition sequence) failed to induce detectable signal variations, resulting in negligible background fluorescence. Furthermore, qPCR validation of the miR-21 expression corroborated the cShot results, showing a progressive increase in miR-21 abundance during MCF-7 cell cycle progression (Fig. 5C). Fig. 5D display histograms of the relative Cy5 fluorescence intensity (RI) derived from HCR generation in MCF-7 cells at distinct cell cycle phases. A significant strong fluorescence intensity was observed from the cShot strategy, while the control experiments showed negligible intensity, which demonstrated the sensitive analytical ability of the cShot system. Subsequently, 25 single cells were selected from each cell cycle phase cohort for fluorescence intensity analysis, followed by evaluation of histograms correlating the miR-21-associated fluorescence intensity with the cell cycle phase to assess the collective cellular response across the different stages (Fig. 5E). Notably, distinct phases exhibited characteristic histogram profiles of peak fluorescence intensity. The G0–G1 phase, representing the initial stage of the cell cycle, demonstrated relatively low miR-21 levels, resulting in diminished fluorescence signals. A gradual increase in fluorescence intensity was observed in the G1-phase cells, followed by a marked rise during the S phase, a period characterized by active DNA synthesis. The strongest fluorescence signals were detected in G2–M phase cells, consistent with peak miR-21 expression at mitotic entry.64 Collectively, these results demonstrated cShot to be a robust analytical tool for assessing miRNA expression levels across cell cycle phases under restricted auxiliary conditions.Dynamic miRNA monitoring via cShot under drug stimulation
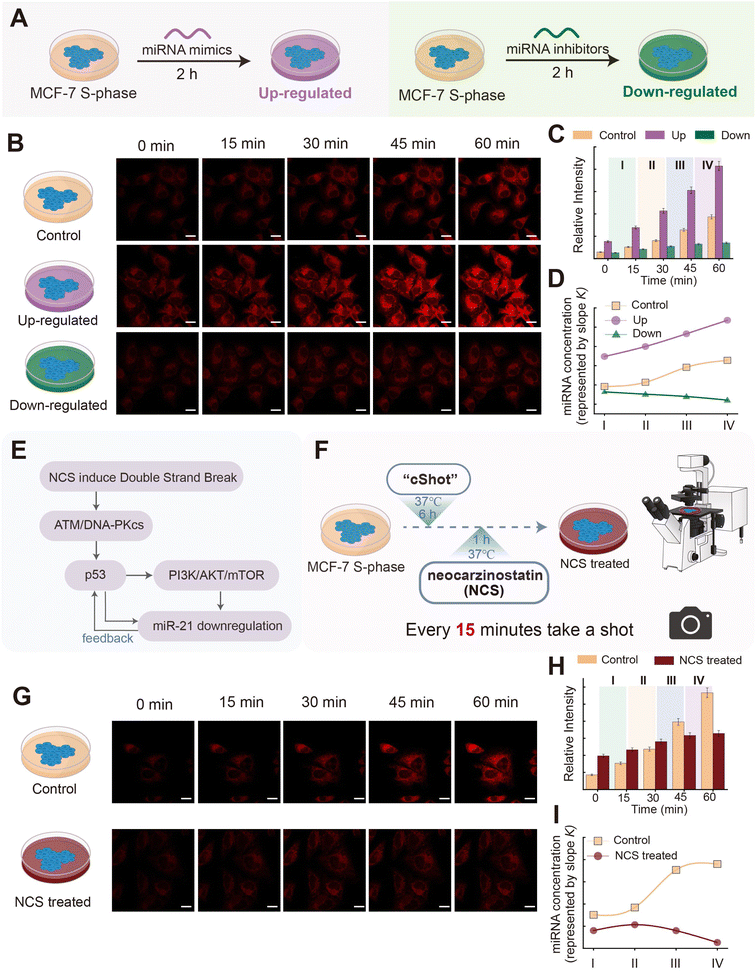
Subsequently, we evaluated the capability of cShot in detecting dynamic changes of target molecule levels by cellular imaging. A prominent advantage of the CDNs lies in their autonomous ability to modulate signal output in response to fluctuating target concentrations. In this study, dynamic regulation of miR-21 levels was achieved in MCF-7 cells through transfection with DNA sequences that were either complementary (miRNA inhibitors) or homologous (miRNA mimics) to miR-21 (Fig. 6A). Following 6 hours of cShot-mediated transfection, cells were subsequently transfected with miRNA mimics and inhibitors using the Lipo8000™ lipid-based transfection system. Cellular imaging was initiated after 2 hours post-transfection, with images captured at 15 minute intervals throughout the observation period. MCF-7 cells synchronized in the S-phase without up-regulation or down-regulation were designated as the control group. As shown in Fig. 6B, the two pretreated cell groups exhibited markedly distinct HCR-generated time-dependent fluorescence trends compared to that of the control group. The relative fluorescence intensities (RI) from each image of these groups were quantified and plotted in Fig. 6C. The time-dependent fluorescence data were divided into four phases: time interval I corresponds to 0–15 min; time interval II corresponds to 15–30 min; time interval III corresponds to 30–45 min; and time interval IV corresponds to 45–60 min. The varying RI revealed increasing fluorescence intensities over time across all three groups, albeit with significant differences in their growth rates. The average rate of fluorescence change (slope k: ΔF/Δt) over consecutive 15 minute imaging intervals is proportional to the change in the target concentration. Therefore, the increase in fluorescence intensity directly reflects the accumulation (or consumption) of the target miR-21 within the corresponding time window. The average rate of fluorescence change (slope k) which represents the miR-21 concentration of the cells was analyzed for each time interval. Fig. 6D demonstrates statistically distinct k-values among the three groups during phase I, indicating different miR-21 levels at this initial stage. Both the control and up-regulated groups exhibited progressively increasing k-values, reflecting continuous accumulation of intracellular miR-21. This observation aligns with the heightened metabolic activity characteristic of S-phase cells.65 The up-regulated group displayed accelerated k-value increase, attributable to the time-dependent transfection process of the miR-21 mimics. Conversely, the down-regulated group showed a sustained decline in k-values, consistent with anti-miR-21 inhibitor-driven depletion of endogenous miR-21. These results robustly validate cShot's capability to monitor dynamic target fluctuations in live cells, highlighting its high potential for real-time therapeutic monitoring during pharmacological interventions.Neocarzinostatin (NCS), a clinically established antineoplastic agent, exhibits broad-spectrum antitumor activity through potent suppression of diverse cancer cell lines.66,67 While both NCS and miRNAs are known to play critical roles in tumorigenesis and cancer progression, their mechanisms of action are mechanistically distinct. NCS acts as a DNA-damaging agent that triggers the p53-mediated stress response and cellular stress responses, which in turn leads to the dynamic regulation of miR-21 expression level (Fig. 6E).68 To probe this interplay, we applied cShot to investigate the dynamic response of miR-21 in MCF-7 cells following NCS treatment. MCF-7 cells were treated with NCS and imaged via confocal microscopy 1 h post-treatment, with images acquired at 15 minute intervals (Fig. 6F). As shown in Fig. 6G, the fluorescence intensities of NCS-treated and control cells exhibited negligible differences during the initial 0–30 minutes period. However, a pronounced divergence emerged between 30–60 min, indicating the occurrence of NCS-induced modulation of the miR-21 dynamics. The transient concentration of the HCR product provides a real-time monitor for the DNAzyme's catalytic rate within CDN constituents, which directly correlates with the dynamics of the miRNA concentrations. Quantitative analysis of the relative fluorescence intensity revealed elevated signals in NCS-treated cells relative to controls during 0–30 min, followed by a significant reduction in the NCS group after 30 min (Fig. 6H and I). This biphasic response suggests an initial elevation of miR-21 levels in NCS-treated cells prior to imaging, with subsequent time-dependent depletion. Kinetic analysis of the fluorescence growth rates (k) further corroborated these findings. Unlike the un-treated S-phase MCF-7 cells, which displayed sustained miR-21 accumulation, NCS-treated cells exhibited continued miR-21 increase from phase I to phase II (0–30 min), potentially reflecting the activation of stress-responsive mechanisms aimed at counteracting NCS-induced DNA damage. Strikingly, phase III–IV (30–60 min) manifested a sharp decline in the k-values, likely attributable to diminished DNA repair capacity, pathway inhibition, and eventual miR-21 depletion, culminating in cellular apoptosis. To further validate the performance of the cShot system, we conducted a comparative imaging study using a conventional molecular beacon (MB) as a reference method (Fig. S23 with accompanying discussion). Notably, under stimulation by NCS, the MB produced substantially weaker fluorescence signals and failed to distinctly resolve the temporal fluctuations of intracellular miR-21 levels. This conclusion underscores the translational potential of temporally resolved monitoring for delineating disease-associated cellular trajectories during therapeutic interventions. The capability to dynamically track molecular responses may enable clinicians to circumvent suboptimal pharmacological outcomes—such as overtreatment or therapeutic inefficacy—by informing personalized dosing regimens aligned with real-time cellular feedback.
Conclusions
The development of “cShot”, a constitutional dynamic DNA network (CDN)-driven sensing platform, addresses the critical challenge of dynamically monitoring intracellular miRNA expression across cell cycle phases. By integrating reversible binding of competitive DNA components with hybridization chain reaction (HCR)-based signal amplification, cShot overcomes the limitations of traditional static detection methods, enabling real-time tracking of miRNA dynamics in synchronized cancer cells. It should be noted, however, that the signal output of the CDN system primarily relies on thermodynamic changes among its constituents, which may partially compromise detection sensitivity; this limitation could be mitigated in future studies by designing initially unbalanced CDN systems to enhance the responsiveness. Beyond the demonstration of miRNA imaging, the architectural flexibility of cShot suggests it has the potential to become a universal platform for sensing diverse molecular targets by simply reconfiguring the corresponding recognition sequences. Moreover, cShot's capacity to map miRNA heterogeneity across cell cycles and drug responses positions it as a transformative tool for precision oncology. By correlating miRNA dynamics with therapeutic outcomes, clinicians could tailor dosages in real time, minimizing toxicity while maximizing efficacy. In addition, the integration of CDNs with functional DNA nanostructures (like the tetrahedron used here) provides a robust framework for spatial–temporal imaging within specific subcellular organelles, offering a powerful tool for studying complex intracellular signaling pathways and evaluating the real-time efficacy of therapeutic drugs in personalized medicine. Notably, by incorporating dissipative network motifs, the system could potentially achieve autonomous regeneration for the continuous, long-term imaging of various biomolecules. In conclusion, cShot bridges the gap between static miRNA quantification and dynamic cellular profiling, offering a robust framework to decode the temporal complexity of biological systems under therapeutic treatment.Author contributions
Jie Zhou, Pu Zhang and Ruo Yuan conceived the project. Jie Zhou designed the sequences, performed the experiments and analyzed the data. Yuwei Sha, Fengying Yuan and Ling'ou Qin performed the experiments and assisted with the revision of the paper. Ruo Yuan, Yu Ouyang and Yaqin Chai supervised the project. Jie Zhou wrote the paper. Jie Zhou, Ruo Yuan, Yaqin Chai, Pu Zhang and Yu Ouyang commented on and revised the paper.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: detailed experimental procedures, oligonucleotide sequences, and apparatus information. See DOI: https://doi.org/10.1039/d5sc09638c.Acknowledgements
This paper was financially supported by the National Natural Science Foundation of China (22374122, 22574132, 22476161, 22174113 and 22176153), Natural Science Foundation of Chongqing, China (CSTB2025NSCQ-JQX0006, CSTB2025NSCQ-GPX0534), Brain-Gain Plan of New Chongqing Young Elite Program (CSTB2025YCJH-KYXM0029) and Fundamental Research Funds for the Central Universities (SWU-KR25035). We are grateful to Dr Yan Li (Analytical and Testing Center, Southwest University) for the assistance with the confocal observations.Notes and references
- Y. Xia, M. Sun, H. Huang and W.-L. Jin, Signal Transduction Targeted Ther., 2024, 9, 1–33 Search PubMed.
- B. A. Helmink, M. A. W. Khan, A. Hermann, V. Gopalakrishnan and J. A. Wargo, Nat. Med., 2019, 25, 377–388 CrossRef CAS PubMed.
- L. A. Korde, M. R. Somerfield, D. L. Hershman and for the Neoadjuvant Chemotherapy, Endocrine Therapy, and Targeted Therapy for Breast Cancer Guideline Expert Panel, J. Clin. Oncol., 2022, 40, 1696–1698 Search PubMed.
- M. Morel, R. Shtrahman, V. Rotter, L. Nissim and R. H. Bar-Ziv, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 8133–8138 CrossRef CAS PubMed.
- L. J. Bugaj, A. J. Sabnis, A. Mitchell, J. E. Garbarino, J. E. Toettcher, T. G. Bivona and W. A. Lim, Science, 2018, 361, eaao3048 CrossRef PubMed.
- H. Chen, J. Yao, R. Bao, Y. Dong, T. Zhang, Y. Du, G. Wang, D. Ni, Z. Xun, X. Niu, Y. Ye and H.-B. Li, Mol. Cancer, 2021, 20, 29 CrossRef CAS PubMed.
- Y. Pu, L. Li, H. Peng, L. Liu, D. Heymann, C. Robert, F. Vallette and S. Shen, Nat. Rev. Clin. Oncol., 2023, 20, 799–813 CrossRef PubMed.
- E. A. Shirshin, M. V. Shirmanova, A. V. Gayer, M. M. Lukina, E. E. Nikonova, B. P. Yakimov, G. S. Budylin, V. V. Dudenkova, N. I. Ignatova, D. V. Komarov, V. V. Yakovlev, W. Becker, E. V. Zagaynova, V. I. Shcheslavskiy and M. O. Scully, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2118241119 CrossRef CAS PubMed.
- W. Wang, L. Liu, J. Zhu, Y. Xing, S. Jiao and Z. Wu, ACS Nano, 2024, 18, 6266–6275 CrossRef CAS PubMed.
- C. Park, S. Chung, H. Kim, N. Kim, H. Y. Son, R. Kim, S. Lee, G. Park, H. W. Rho, M. Park, J. Han, Y. Song, J. Lee, S.-H. Jun, Y.-M. Huh, H. H. Jeong, E.-K. Lim, E. Kim and S. Haam, ACS Nano, 2024, 18, 26297–26314 Search PubMed.
- X. Wang, S. S. Liew, J. Huang, Y. Hu, X. Wei and K. Pu, J. Am. Chem. Soc., 2024, 146, 22689–22698 CrossRef CAS PubMed.
- L. Hao, N. Boehnke, S. K. Elledge, N.-S. Harzallah, R. T. Zhao, E. Cai, Y.-X. Feng, S. Neaher, H. E. Fleming, P. B. Gupta, P. T. Hammond and S. N. Bhatia, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2307802121 CrossRef CAS PubMed.
- H. K. Matthews, C. Bertoli and R. A. M. de Bruin, Nat. Rev. Mol. Cell Biol., 2022, 23, 74–88 CrossRef CAS PubMed.
- L. Liu, W. Michowski, A. Kolodziejczyk and P. Sicinski, Nat. Cell Biol., 2019, 21, 1060–1067 CrossRef CAS PubMed.
- T. Otto, S. V. Candido, M. S. Pilarz, E. Sicinska, R. T. Bronson, M. Bowden, I. A. Lachowicz, K. Mulry, A. Fassl, R. C. Han, E. S. Jecrois and P. Sicinski, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 10660–10665 Search PubMed.
- V. Sekar, E. Mármol-Sánchez, P. Kalogeropoulos, L. Stanicek, E. A. Sagredo, A. Widmark, E. Doukoumopoulos, F. Bonath, I. Biryukova and M. R. Friedländer, Nat. Biotechnol., 2024, 42, 1296–1302 CrossRef CAS PubMed.
- S. Zhao, Q. Zhang, R. Luo, J. Sun, C. Zhu, D. Zhou and X. Gong, Chem. Sci., 2024, 15, 18347–18354 RSC.
- H. Zhang, Y. Tang, Y. Zhou, Y. Wang, H. Si, L. Li and B. Tang, Chem. Sci., 2024, 15, 9345–9352 RSC.
- S. Yue, J. Zhan, X. Xu, J. Xu, S. Bi and J.-J. Zhu, Chem. Sci., 2024, 15, 11528–11539 RSC.
- F. Ma, G.-A. Lu, Q. Chen, Y. Ruan, X. Li, X. Lu and C. Li, Sci. Bull., 2020, 65, 2130–2140 CrossRef CAS PubMed.
- R. Rupaimoole, G. A. Calin, G. Lopez-Berestein and A. K. Sood, Cancer Discov., 2016, 6, 235–246 CrossRef CAS PubMed.
- S. Tian, Y. Asano, T. Das Banerjee, S. Komata, J. L. Q. Wee, A. Lamb, Y. Wang, S. N. Murugesan, H. Fujiwara, K. Ui-Tei, P. J. Wittkopp and A. Monteiro, Science, 2024, 386, 1135–1141 CrossRef CAS PubMed.
- Y. Cui, X. Lyu, L. Ding, L. Ke, D. Yang, M. Pirouz, Y. Qi, J. Ong, G. Gao, P. Du and R. I. Gregory, Nature, 2021, 593, 602–606 CrossRef CAS PubMed.
- Z. Xu, Y. Dong, N. Ma, X. Zhu, X. Zhang, H. Yin, S. Chen, J.-J. Zhu, Y. Tian and Q. Min, J. Am. Chem. Soc., 2023, 145, 26557–26568 CrossRef CAS PubMed.
- H. Yan, Y. Wen, Z. Tian, N. Hart, S. Han, S. J. Hughes and Y. Zeng, Nat. Biomed. Eng., 2023, 7, 1583–1601 CrossRef CAS PubMed.
- M. Wu, L. Liu, Y. Xing, Z. Wu, P. Huang and B. Li, Adv. Funct. Mater., 2025, 2422305 CrossRef CAS.
- H. Lv, N. Xie, M. Li, M. Dong, C. Sun, Q. Zhang, L. Zhao, J. Li, X. Zuo, H. Chen, F. Wang and C. Fan, Nature, 2023, 622, 292–300 CrossRef CAS PubMed.
- M. R. Jones, N. C. Seeman and C. A. Mirkin, Science, 2015, 347, 1260901 CrossRef PubMed.
- Y. He, Q. Wang, C. Hong, R. Li, J. Shang, S. Yu, X. Liu and F. Wang, Angew. Chem., Int. Ed., 2023, 62, e202307418 CrossRef CAS PubMed.
- T. Jet, G. Gines, Y. Rondelez and V. Taly, Chem. Soc. Rev., 2021, 50, 4141–4161 RSC.
- L. Zhou, M. Gao, W. Fu, Y. Wang, D. Luo, K. Chang and M. Chen, Sci. Adv., 2020, 6, eabb0695 CrossRef CAS PubMed.
- Y. Wu, G. A. Wang, Q. Yang and F. Li, J. Am. Chem. Soc., 2024, 146, 18041–18049 CrossRef CAS PubMed.
- C. Wang, M. P. O'Hagan, Z. Li, J. Zhang, X. Ma, H. Tian and I. Willner, Chem. Soc. Rev., 2022, 51, 720–760 RSC.
- H. Liu, J. Dong, Z. Duan, F. Xia, I. Willner and F. Huang, Sci. Adv., 2024, 10, eadp6166 CrossRef CAS PubMed.
- T. Wang and F. C. Simmel, Angew. Chem., Int. Ed., 2023, 62, e202302858 CrossRef CAS PubMed.
- Y. Chen, S.-Y. Yang, X. Ou, H. Wang, F.-C. Kong, P. C. Y. Chow, Y. Wang, Y. Jiang, W. Zhao, J. Sun, R. T. K. Kwok, D.-W. Zheng, W. Yu, F. Wang, J. W. Y. Lam and B. Z. Tang, J. Am. Chem. Soc., 2024, 146, 35462–35477 CrossRef CAS PubMed.
- C. Wang, Y. Xie, X. Song, Z. Chao, K. Wu, Y. Fang, H. Zhao, H. Ju and Y. Liu, Angew. Chem., Int. Ed., 2023, 62, e202312665 CrossRef CAS PubMed.
- S. Chen, Y. Yin, X. Pang, C. Wang, L. Wang, J. Wang, J. Jia, X. Liu, S. Xu and X. Luo, Chem. Sci., 2024, 15, 566–572 RSC.
- Y. Wu, G. A. Wang, Q. Yang and F. Li, J. Am. Chem. Soc., 2024, 146, 18041–18049 CrossRef CAS PubMed.
- C. Zhang, X. Ma, X. Zheng, Y. Ke, K. Chen, D. Liu, Z. Lu, J. Yang and H. Yan, Sci. Adv., 2022, 8, eabl4589 Search PubMed.
- Z. Zhang, W. Wei, S. Chen, J. Yang, D. Song, Y. Chen, Z. Zhao, J. Chen, F. Wang, J. Wang, Z. Li, Y. Liang and H. Yu, J. Am. Chem. Soc., 2024, 146, 7052–7062 CrossRef CAS PubMed.
- D. Mathur, Nat. Nanotechnol., 2024, 19, 143–144 CrossRef CAS PubMed.
- G. A. Wang, J. Xu, S. M. Traynor, H. Chen, F. Eljabu, X. Wu, H. Yan and F. Li, J. Am. Chem. Soc., 2021, 143, 13655–13663 Search PubMed.
- R. Mukherjee, A. Sengar, J. Cabello-García and T. E. Ouldridge, J. Am. Chem. Soc., 2024, 146, 18916–18926 CrossRef CAS PubMed.
- L. Liu, F. Hong, H. Liu, X. Zhou, S. Jiang, P. Šulc, J.-H. Jiang and H. Yan, Sci. Adv., 2022, 8, eabm9530 CrossRef CAS PubMed.
- J. Xu, G. A. Wang, L. Gao, L. Wu, Q. Lei, H. Deng and F. Li, Nat. Commun., 2023, 14, 4248 CrossRef CAS PubMed.
- T. M. Brown, H. H. Fakih, D. Saliba, J. Asohan and H. F. Sleiman, J. Am. Chem. Soc., 2023, 145, 2142–2151 CrossRef CAS PubMed.
- J. Deng and A. Walther, J. Am. Chem. Soc., 2020, 142, 21102–21109 CrossRef CAS PubMed.
- H.-R. Chen, M.-L. Su, Y.-M. Lei, Z.-X. Ye, Z.-P. Chen, P.-Y. Ma, R. Yuan, Y. Zhuo, C.-Y. Yang and W.-B. Liang, J. Am. Chem. Soc., 2023, 145, 12812–12822 CrossRef CAS PubMed.
- S. Wang, L. Yue, Z. Shpilt, A. Cecconello, J. S. Kahn, J.-M. Lehn and I. Willner, J. Am. Chem. Soc., 2017, 139, 9662–9671 Search PubMed.
- Y. Ouyang, J. Dong and I. Willner, J. Am. Chem. Soc., 2023, 145, 22135–22149 CrossRef CAS PubMed.
- L. Yue, S. Wang, S. Lilienthal, V. Wulf, F. Remacle, R. D. Levine and I. Willner, J. Am. Chem. Soc., 2018, 140, 8721–8731 CrossRef CAS PubMed.
- Y. Ouyang, Y. S. Sohn, X. Chen, R. Nechushtai, E. Pikarsky, F. Xia, F. Huang and I. Willner, J. Am. Chem. Soc., 2025, 147, 19282–19295 CrossRef CAS PubMed.
- L. Gao, R. Zhang, Y. Tang and F. Li, Angew. Chem., Int. Ed., 2025, 64, e202425644 Search PubMed.
- X. Wang, S. Wang, L. Yue and W. Tan, Chem, 2025, 11, 102593 CAS.
- Z. Zhou, N. Lin, Y. Ouyang, S. Liu, Y. Zhang and I. Willner, J. Am. Chem. Soc., 2023, 145, 12617–12629 CrossRef CAS PubMed.
- L. Csernoch, J. C. Bernengo, P. Szentesi and V. Jacquemond, Biophys. J., 1998, 75, 957–967 CrossRef CAS PubMed.
- A. R. Nayak, W. Rangubpit, A. H. Will, Y. Hu, P. Castro-Hartmann, J. J. Lobo, K. Dryden, G. D. Lamb, P. Sompornpisut and M. Samsó, Nat. Commun., 2024, 15, 4115 CrossRef CAS PubMed.
- C. C. Daw, K. Ramachandran, B. T. Enslow, S. Maity, B. Bursic, M. J. Novello, C. S. Rubannelsonkumar, A. H. Mashal, J. Ravichandran, T. M. Bakewell, W. Wang, K. Li, T. R. Madaris, C. E. Shannon, L. Norton, S. Kandala, J. Caplan, S. Srikantan, P. B. Stathopulos, W. B. Reeves and M. Madesh, Cell, 2020, 183, 474–489 CrossRef CAS PubMed.
- T. Otto, S. V. Candido, M. S. Pilarz, E. Sicinska, R. T. Bronson, M. Bowden, I. A. Lachowicz, K. Mulry, A. Fassl, R. C. Han, E. S. Jecrois and P. Sicinski, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 10660–10665 CrossRef CAS PubMed.
- R. A. Elbarbary, K. Miyoshi, J. R. Myers, P. Du, J. M. Ashton, B. Tian and L. E. Maquat, Science, 2017, 356, 859–862 CrossRef CAS PubMed.
- H. Xiao, W. Xiao, J. Cao, H. Li, W. Guan, X. Guo, K. Chen, T. Zheng, Z. Ye, J. Wang and H. Xu, Cancer Lett., 2016, 374, 107–116 CrossRef CAS PubMed.
- I. Pellarin, A. Dall'Acqua, A. Favero, I. Segatto, V. Rossi, N. Crestan, J. Karimbayli, B. Belletti and G. Baldassarre, Signal Transduction Targeted Ther., 2025, 10, 1–62 Search PubMed.
- J.-Y. Zhou, W.-L. Ma, S. Liang, Y. Zeng, R. Shi, H.-L. Yu, W.-W. Xiao and W.-L. Zheng, BMB Rep., 2009, 42, 593–598 CrossRef CAS PubMed.
- R. A. Laskey, M. P. Fairman and J. J. Blow, Science, 1989, 246, 609–614 CrossRef CAS PubMed.
- Y. V. Surovtseva, V. Jairam, A. F. Salem, R. K. Sundaram, R. S. Bindra and S. B. Herzon, J. Am. Chem. Soc., 2016, 138, 3844–3855 Search PubMed.
- A. S. Kawale, K. Akopiants, K. Valerie, B. Ruis, E. A. Hendrickson, S. N. Huang, Y. Pommier and L. F. Povirk, Nucleic Acids Res., 2018, 46, 8926–8939 Search PubMed.
- R. L. Hanson and E. Batchelor, Cell Rep., 2020, 33, 108240 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |