Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Subunit fusion unlocks rapid in vitro maturation for slowly activating heterodimeric [FeFe]-hydrogenases
Jan
Jaenecke
 a,
Konstantin
Bikbaev
a,
Konstantin
Bikbaev
 b,
Miriam
Malagnini
b,
Miriam
Malagnini
 c,
Julia
Bronold
a,
Shanika
Yadav
c,
Julia
Bronold
a,
Shanika
Yadav
 d,
Ulf-Peter
Apfel
d,
Ulf-Peter
Apfel
 de,
Christophe
Léger
de,
Christophe
Léger
 c,
James A.
Birrell
c,
James A.
Birrell
 f,
Ingrid
Span
f,
Ingrid
Span
 b,
Nicolas
Plumeré
b,
Nicolas
Plumeré
 *a and
Martin
Winkler
*a and
Martin
Winkler
 *ag
*ag
aProfessorship for Electrobiotechnology, Technical University of Munich, Campus Straubing for Biotechnology and Sustainability, Uferstrasse 53, 94315 Straubing, Germany. E-mail: martin-h.winkler@tum.de; nicolas.plumere@tum.de
bBioinorganic Chemistry, Friedrich-Alexander-Universität Erlangen-Nürnberg, Egerlandstrasse 1, 91058 Erlangen, Germany
cLaboratoire de Bioénergétique et Ingénierie des Protéines (BIP), CNRS, Aix-Marseille Université, 31 chemin Joseph Aiguier, 13009 Marseille, France
dActivation of Small Molecules/Technical Electrochemistry, Faculty of Chemistry and Biochemistry, Ruhr-University Bochum, Universitätsstrasse 150, 44801 Bochum, Germany
eDepartment of Electrosynthesis, Fraunhofer UMSICHT, 46047 Oberhausen, Germany
fSchool of Life Sciences, University of Essex, Wivenhoe Park, Colchester, CO4 3SQ, UK
gInorganic Spectroscopy, Energy Converting Enzymes, Max-Planck-Institute for Chemical Energy Conversion, Stiftstrasse 34-36, 45470 Mühlheim an der Ruhr, Germany
First published on 3rd February 2026
Abstract
Hydrogenases offer a sustainable alternative to noble metals for catalyzing H2-oxidation and H2-production. The heterodimeric [FeFe]-hydrogenase of Desulfovibrio desulfuricans ATCC 7757 (DdHydAB) is most promising due to its exceptional catalytic activity and high-yield heterologous expression of its apo-form. Scalable production of the holo-form relies on in vitro maturation of the apo-enzyme using a chemically synthesized 2FeH cofactor mimic. However, the unusually slow in vitro maturation of DdHydAB raises mechanistic questions and limits its scalability. Through structural and sequence analysis, we identified the cause of this slow maturation and redesigned the enzyme via subunit fusion, inserting short peptide linkers near the active site. This modification facilitates the rearrangement of a critical locking element after cofactor uptake, increasing the maturation rate by up to 41-fold without compromising catalytic performance. Our findings elucidate a key step in the plug-lock-lid mechanism underlying maturation and promote the industrial applicability of DdHydAB.
Introduction
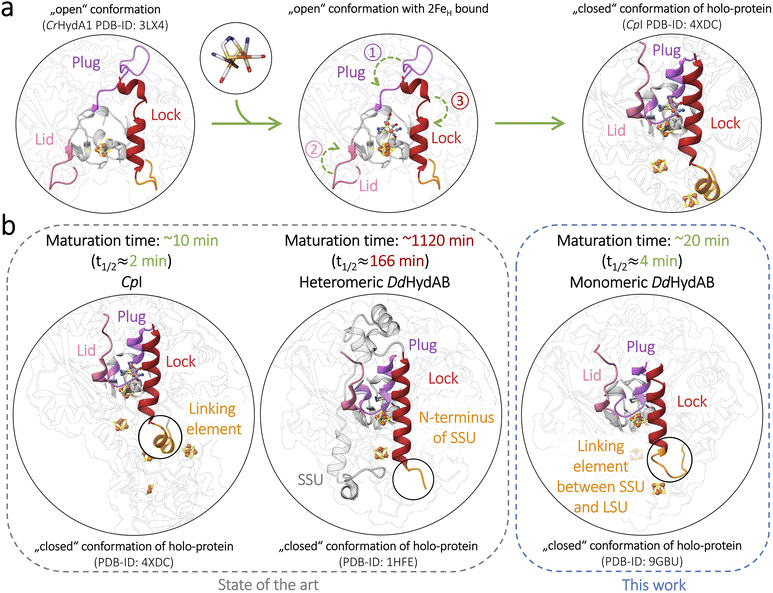
Hydrogenases are increasingly recognized as sustainable catalysts for H2-production and utilization in green energy systems and the (bio)chemical industry.1–7 Among the three existing main classes, called [Fe]-, [NiFe]-, and [FeFe]-hydrogenases, the latter stand out for their bidirectional H+/H2 interconversion at turnover frequencies that rival precious-metal catalysts.8–10 Previous studies highlight the superior features of individual [FeFe]-hydrogenases rendering them as promising candidates for future applications; this includes DdHydAB and CbA5H, both of which can reversibly adopt an inactive, O2-resistant state (Hinact), permitting aerobic purification and facile sample handling.11–15 Recent publications have demonstrated scalable heterologous expression and provide the basis for their implementation in electrochemical systems.6,7,16–18[FeFe]-hydrogenases offer a particularly convenient route to production at scales relevant for industry: their catalytic cofactor (H-cluster) comprises a [4Fe4S]-subcluster (4FeH) covalently linked via one of its Cys ligands to an unusual [2Fe]-subcluster (2FeH). In their apo-state, which lacks the 2FeH-subcluster, [FeFe]-hydrogenases can be effectively overexpressed in Escherichia coli. The purified apo-form can be maturated in vitro to the active holo-enzyme by simply mixing the apo-hydrogenase with an excess of a synthetic 2FeH mimic complex (2FeMIMH), which spontaneously enters the open cofactor-binding site (Fig. 1).19–21
Upon in vitro maturation, the heterodimeric M2-type [FeFe]-hydrogenase DdHydAB of Desulfovibrio desulfuricans (identical to Nitratidesulfovibrio vulgaris and formerly labeled as D. vulgaris; see SI Discussion) reaches exceptionally high activities of 7400 s−1 (H2-evolution) and 56 000 s−1 (H2-oxidation),21–23 combined with the possibility of reversible sulfide-induced O2-protection.14,15 However, the production of the active DdHydAB holo-enzyme still faces a major challenge.21 In contrast to the apo-states of monomeric [FeFe]-hydrogenases that fully activate within 15–30 min upon adding the 2FeMIMH complex,20in vitro maturation of DdHydAB requires more than two days under similar conditions,21 lowering its application potential and motivating a mechanistic re-examination of the maturation process. In vivo, H-cluster maturation is assumed to be conserved among [FeFe]-hydrogenases of eubacteria and eukaryotes.24–27 When heterologously expressed in its apo-form, DdHydAB is already equipped with the 4FeH-subcluster along with its two accessory [FeS]-clusters characteristic of the M2-type architecture, while exhibiting a vacant but accessible 2FeH binding site. The native 2FeH precursor arises via the contribution of three maturases (HydG, HydE, and HydF). HydG, one of the two radical-SAM enzymes involved, generates the mononuclear [FeII(CO)2 (CN)(Cys)]− synthons. The other (HydE) combines two synthons to the 2FeH-pre-complex [FeI2(µ-SH)2(CO)4(CN)2]2− before both of its Fe-sites are cross-bridged by the dithiomethylamine ligand, which is essential for H+-exchange between the proton transfer pathway and the active site (Fig. S1a).27–32 HydF serves as a scaffold protein for the biosynthesis of the 2FeH precursor before it is finally transferred from the mature HydF protein (holo-HydF) to the vacant cofactor-binding site of the apo-hydrogenase during a transient protein–protein interaction (Fig. 1 and S1b).28,33–36 Functional 2FeH insertion entails a rotation of the primary ligand sphere at the distal iron (Fed) of the 2FeH precursor complex, favored by H-bond contacts between the two CN− ligands and individual residues in the secondary ligand sphere,30,37 followed by a local reconfiguration of three structural elements first indicated by Mulder and coauthors in 2010.26 One element turns into a plug-like structure that loosely closes the binding site over the inserted 2FeH precursor, thereby supporting the adoption of its rotated conformation. Next, this plug placement is barred in place from two sides by a restructuring of both remaining loop elements, termed lid and lock (Fig. 1a and S1b).24–26,30,31 The coupling of the 2FeMIMH complex with the 4FeH-subcluster is enabled by one of its Cys ligands, which adopts a bridging configuration (µ-SCys) between both subclusters (Fig. 1 and S1), yielding the transient HredCO state.25,27,38 A single oxidation step leads to the likewise inhibited but rather stable HoxCO state (Fig. S1c). Upon release of the apical CO ligand from Fed, the oxidized active-ready state Hox is formed.25,27,38
We previously hypothesized that DdHydAB's unusually slow maturation results from a higher propensity of the apo-form to adopt the closed conformation.21 However, no attempts have been made to verify this idea or examine other potential reasons for the extremely low maturation rate of DdHydAB, let alone to speed up the maturation process. We propose an alternative, structure-based explanation centered on its heterodimeric state: a superposition of the X-ray structure of DdHydAB (1HFE)39 with those of monomeric enzymes (CrHydA1, PDB ID: 3LX4; CpI, PDB ID: 4XDC)26,30 illustrates that the C-terminus of the small-subunit (SSU) coincides with the end of the lock-element (Fig. S2). In monomeric enzymes, the lock-helix straightens upon 2FeH uptake, bolting the plug in its place over the occupied binding site; in the case of DdHydAB, we hypothesize that the gap between SSU and LSU likely increases the flexibility of the lock-element at the free SSU N-terminus, hindering an efficient lock rearrangement and delaying the final closure of the pocket. Subunit fusion can be a powerful strategy to enhance the features of heterodimeric proteins, such as stability, yield, and catalytic efficiency.40,41 Linker-mediated coupling was also used to create chimeric catalysts with new functions.40–42
Here, we investigated whether a linker-based monomerization approach positively impacts the unusually low in vitro maturation rate of DdHydAB. Employing FTIR spectroscopy and various kinetic measurements, we show that the rates of H-cluster formation and activity development of the monomeric variants increase by nearly two orders of magnitude relative to the heterodimeric wild-type.21,24 Protein-film electrochemistry and X-ray crystallography enable us to ensure the conservation of the advantageous traits of DdHydAB and to rationalize the differences in the variant-specific maturation rates.
Results and discussion
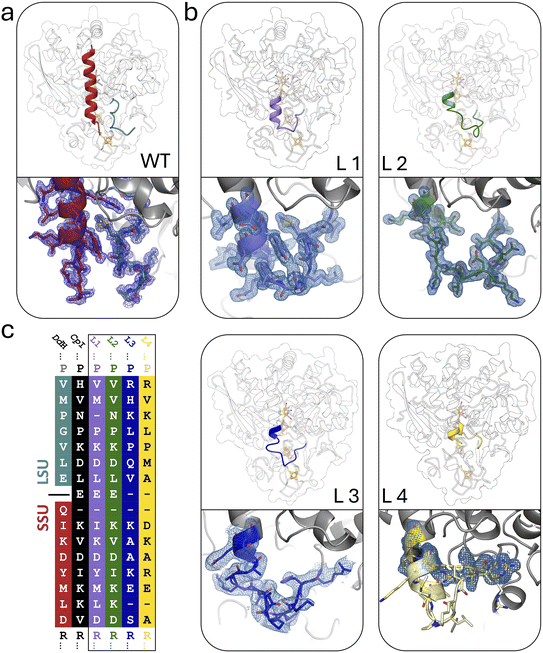
To investigate apo-DdHydAB for possible structural differences from monomeric [FeFe]-hydrogenases, particularly in the terminal regions of the subunits and the structural elements responsible for the closure and locking of the holo-protein, we crystallized DdHydAB in its unmaturated apo-form. The resulting crystal structure exhibited the closed state of the H-cluster binding site but did not show significant differences from the apo-state structure of the fast-maturating CpI enzyme, which was likewise crystallized in its closed state (PDB ID: 4XDD)30 (Fig. S3a–e). As we were unable to support the hypothesis of a significantly higher tendency for premature binding-site closure in the case of DdHydAB, we focused instead on the fact that its heterodimeric state is the most prominent structural difference compared to monomeric enzymes such as CpI and CrHydA1. The gap between the two DdHydAB subunits coincides with the N-terminus of the lock-element, potentially affecting the 2FeH binding-site closure in the H-domain, and thus the rate of in vitro activation. To examine this hypothesis experimentally and with the aim of improving the maturation rate of DdHydAB, we engineered four SSU–LSU fusion constructs (variants L1–L4) by inserting linkers of defined length and sequence (Fig. 2).For the linker design, we followed two strategies: (1) structural mimicry with CpI: based on a structural alignment between DdHydAB (1HFE)39 and the monomeric CpI (PDB ID: 4XDC)30 two linkers were designed (L1–2) with sequence homology and structural features corresponding to the native linker element in CpI (Fig. 2, L1–L2, and S4a, b). (2) Native linker sequences of monomeric homologs: we optimized the structural integration of the linker sequence into the protein environment to avoid destabilization and steric clashes. From a multiple sequence alignment, we selected two monomeric homologs of DdHydAB (M2-type [FeFe]-hydrogenases of Solobacterium moorei and Veillonella atypica; see corresponding sequences in Fig. S5) with a linker sequence coupling the parts that correspond to the SSU and LSU of DdHydAB (Fig. 2, L3–L4, and S4a, b). The resulting fusion proteins were heterologously expressed, purified (Fig. S6), and compared to the dimeric wild-type enzyme for their impact on the rate of in vitro maturation, comparing two independent approaches: (i) we used IR-spectroscopy to monitor the rate of HoxCO formation as a product of 4FeH/2FeH-cluster coupling during maturation in a sealed sample of apo-enzyme upon 2FeMIMH addition. (ii) We followed the activity development of the apo-hydrogenase variants upon addition of a 3-fold excess of 2FeMIMH by periodically testing aliquots for the rate of H2-evolution, which requires CO release from HoxCO.
Electrochemical investigation of CO-binding kinetics and spectroscopic tracking of H-cluster formation rates
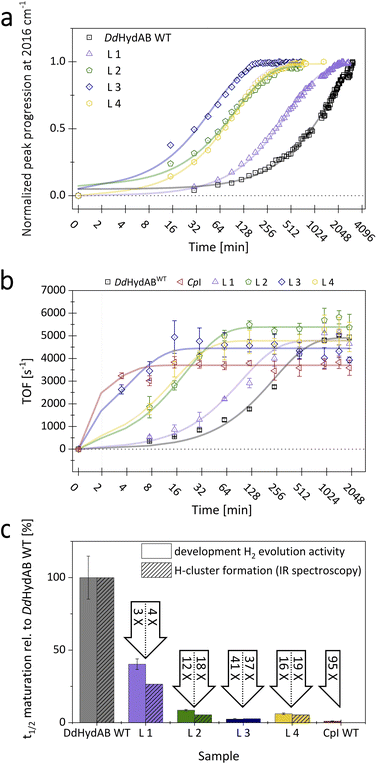
The production of the active enzyme from the HoxCO state requires the final release of the apical CO ligand that blocks the substrate binding site (Fig. S1c). Among the best-characterized [FeFe]-hydrogenases, DdHydAB shows the lowest inhibition constant (Ki) for external CO.39 We exploited this to track the amplitude increase of a characteristic peak at 2016 cm−1 in the IR-spectrum of HoxCO between 0 and 68 h (Fig. 3a; see Fig. S9 for full spectra) as a reporter for plotting the rate of H-cluster formation during the maturation process.To ensure that all monomeric variants retain the high CO affinity of the wild-type enzyme, we employed protein film electrochemistry to monitor the kinetics of CO binding and release. In this classical chronoamperometry experiment, a film of holo-enzyme is prepared on the pyrolytic graphite surface of a rotating disk electrode.43 The H2-oxidation current is measured as a function of time before and after the enzyme is exposed to CO (Fig. S8). For all proteins, we determined and compared the Michaelis–Menten constants for H2 (Fig. S7) and the inhibition constants for CO (KCOi). Both the binding affinity for the substrate (H2) and the affinity for CO (Ki = kout/kin) are similar among all proteins (Table S1) with maximum differences among all proteins remaining well below a factor of two. The final CO-release step is associated with a Ki in the nM range, which confirms that the equilibrium of the reaction HoxCO ⇔ Hox + CO lies on the left side (HoxCO). This especially applies to the conditions of the FTIR-spectroscopy experiment, as throughout the maturation process, the samples are kept in a tightly sealed cell, which maintains a high CO concentration and excludes further competition with the substrate (H2), thus ensuring a homogeneous accumulation of the HoxCO-state (Fig. S1c).21
Although CpI would be an ideal positive control for observing the H-cluster formation rate, its rapid activation and low enzyme-specific CO–ligand binding affinity prevent linear monitoring of the H-cluster formation rate using the state specific IR spectrum of HoxCO.
For each protein, we determined the times required to reach 50% cofactor saturation (half-times; t1/2) (Fig. 3c). All monomeric variants exhibited a significantly higher H-cluster formation rate compared to the heterodimeric wild-type, albeit to varying degrees. Based on the individual t1/2-values for the progression to full H-cluster occupancy, the rates of H-cluster formation can be ranked as follows: L3 > L2 ≈ L4 > L1 >> DdHydAB. With a 3.8-fold enhancement compared to DdHydABWT, the linker of variant L1 showed the lowest impact on the enzyme's maturation rate. Fusion proteins L2 and L4 displayed an 18- and 19-fold increase in the H-cluster formation rate, while variant L3 exhibited the highest impact with a 37-fold increase in the H-cluster formation rate relative to DdHydABWT.
Kinetics of enzyme activation monitored by solution assays
To assess in vitro maturation beyond H-cluster formation, we further monitored the activation to the catalytically competent holo-enzyme by periodically testing aliquots from an in vitro maturation mix for the enzyme's H2-evolution rate. This was achieved through a well-established sodium-dithionite-driven and methyl viologen (MV)-mediated activity assay measured via gas chromatography (Fig. 3b).20,24 This approach enabled the inclusion of CpI as a positive control. The hydrogen evolution assay mirrored the FTIR-based maturation trends, confirming its overall ranking among the compared enzymes. All proteins reached their maximum activity within 34 h, consistent with previous data on the in vitro maturation of DdHydAB.21 While variant L1 exceeded the maturation rate of WT by reaching its full activity after approximately 7.4 h (see Table S2), it was nevertheless the slowest among the four monomeric DdHydAB-variants. In contrast, variant L3 showed an extraordinarily rapid activation with a half-time of merely 4 min, which nearly reached the activation rate of CpI (t1/2: 1.7 min), being one of the fastest-maturating hydrogenases reported to date (Fig. 3c).20 Fusion variants L2 and L4 behaved similarly, peaking at a turnover frequency of around 5000 s−1 after 95 and 68 min, respectively, with variant L2 achieving a higher turnover rate than L4 and the heterodimeric wild-type. To verify that in vitro maturation is discontinued once the aliquot is added to the in vitro assay reaction mix, we prepared a control sample using the fastest-maturating DdHydAB-variant L3. 2FeMIMH and the apo-form of variant L3 were separately diluted and combined in the assay mix right before sample incubation. Under these conditions even L3 showed negligible H2-production (37 s−1), confirming that the in vitro maturation process is effectively stopped when added to the reaction mix of the assay.Although the enzyme to 2FeMIMH ratio (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3) was the same for both approaches compared here, other experimental conditions differed due to the constraints of the respective methods (e.g., enzyme concentration and incubation in open vs. closed vessels). These differences account for the 5-9-fold deviations in absolute t1/2-values for the same enzymes (compare absolute t1/2 values for FTIR and the HER of the same protein in Table S2). We normalized the half-times of in vitro maturation for the different fusion proteins relative to the wild-type (set to 100%). For the same enzyme variants, the relative t1/2 values (rate increase compared to wild-type) obtained on the one hand by monitoring H-cluster formation via FTIR and on the other hand by measuring H2-evolution activity development (HER) aligned closely for all variants (L1: 4-fold/3-fold; L2: 18-fold/12-fold; L3: 37-fold/41-fold; L4: 19-fold/16-fold). Remarkably, L3 shows, in both approaches, by far the highest relative increases compared to the other variants. Its relative HER rate increase is only 2.3-fold lower than the relative rate observed for the monomeric reference enzyme CpI (95-fold; see Table S2). Overall, all linkers enabled a significant increase in the in vitro maturation rate, which was likewise reflected in the rates of H-cluster formation and activity increase. This suggests that the heterodimeric state either slows down subcluster coupling or affects an earlier step in the maturation process. This bottleneck could involve the uptake of the 2FeH precursor, which may be blocked by an unfavorable configuration of the free N-terminus at the lock-element or arise from steps further downstream. It may concern an early state, with the precursor complex 2FeMIMH being embedded in the binding pocket but not yet covalently coupled to the 4FeH-subcluster. Such a state was postulated and experimentally supported to precede subcluster coupling as an inactive intermediate species by Megarity and coauthors.25 Cluster coupling only happens if the applied potential is high enough to oxidize the 4FeH-subcluster, while the uncoupled intermediate state is conserved at potentials below −0.54 V. Megarity et al. described the 2FeMIMH complex to be tightly bound, withstanding several buffer exchanges even at potentials below −0.54 V, where no subcluster coupling occurred. This suggests that after 2FeH uptake, stable binding site closure occurs and is a prerequisite of subcluster coupling. Thus, the unbound 2FeMIMH complex could be easily lost if binding site lockage via the plug, lock, and lid reconfiguration is incomplete.24,30 It can be assumed that subcluster-coupling via the formation of a bridging Cys ligand not only requires a suitable potential range but also a complete and stable H-cluster environment, granted by a sturdy closure of the open 2FeH binding site. The connection between the heterodimeric state and slow maturation rate supports this hypothesis. In DdHydABWT, the flexible N-terminus of the small subunit slows down the locking of the plug-element over the 2FeH binding site after 2FeMIMH uptake, as it lacks the stable resistance and anchoring of the terminal helix segment that is needed for a rapid structural rearrangement at the other end of the helix (Fig. 1a, b, and S1b).
3) was the same for both approaches compared here, other experimental conditions differed due to the constraints of the respective methods (e.g., enzyme concentration and incubation in open vs. closed vessels). These differences account for the 5-9-fold deviations in absolute t1/2-values for the same enzymes (compare absolute t1/2 values for FTIR and the HER of the same protein in Table S2). We normalized the half-times of in vitro maturation for the different fusion proteins relative to the wild-type (set to 100%). For the same enzyme variants, the relative t1/2 values (rate increase compared to wild-type) obtained on the one hand by monitoring H-cluster formation via FTIR and on the other hand by measuring H2-evolution activity development (HER) aligned closely for all variants (L1: 4-fold/3-fold; L2: 18-fold/12-fold; L3: 37-fold/41-fold; L4: 19-fold/16-fold). Remarkably, L3 shows, in both approaches, by far the highest relative increases compared to the other variants. Its relative HER rate increase is only 2.3-fold lower than the relative rate observed for the monomeric reference enzyme CpI (95-fold; see Table S2). Overall, all linkers enabled a significant increase in the in vitro maturation rate, which was likewise reflected in the rates of H-cluster formation and activity increase. This suggests that the heterodimeric state either slows down subcluster coupling or affects an earlier step in the maturation process. This bottleneck could involve the uptake of the 2FeH precursor, which may be blocked by an unfavorable configuration of the free N-terminus at the lock-element or arise from steps further downstream. It may concern an early state, with the precursor complex 2FeMIMH being embedded in the binding pocket but not yet covalently coupled to the 4FeH-subcluster. Such a state was postulated and experimentally supported to precede subcluster coupling as an inactive intermediate species by Megarity and coauthors.25 Cluster coupling only happens if the applied potential is high enough to oxidize the 4FeH-subcluster, while the uncoupled intermediate state is conserved at potentials below −0.54 V. Megarity et al. described the 2FeMIMH complex to be tightly bound, withstanding several buffer exchanges even at potentials below −0.54 V, where no subcluster coupling occurred. This suggests that after 2FeH uptake, stable binding site closure occurs and is a prerequisite of subcluster coupling. Thus, the unbound 2FeMIMH complex could be easily lost if binding site lockage via the plug, lock, and lid reconfiguration is incomplete.24,30 It can be assumed that subcluster-coupling via the formation of a bridging Cys ligand not only requires a suitable potential range but also a complete and stable H-cluster environment, granted by a sturdy closure of the open 2FeH binding site. The connection between the heterodimeric state and slow maturation rate supports this hypothesis. In DdHydABWT, the flexible N-terminus of the small subunit slows down the locking of the plug-element over the 2FeH binding site after 2FeMIMH uptake, as it lacks the stable resistance and anchoring of the terminal helix segment that is needed for a rapid structural rearrangement at the other end of the helix (Fig. 1a, b, and S1b).
Enzyme yields and O2 protection
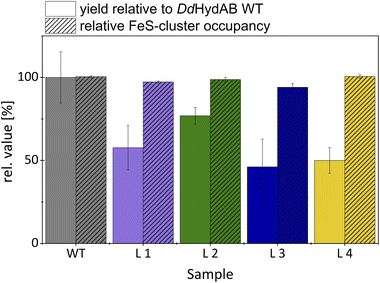
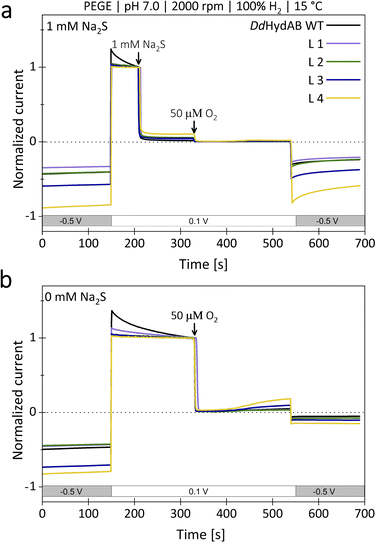
Considering that DdHydAB is a prime candidate for future biotechnological applications that include H2-driven or H2-producing processes, it is crucial to preserve the beneficial characteristics of the original enzyme in its monomeric variants. Therefore, we investigated the monomeric fusion proteins for their heterologous expression yields, [FeS]-cluster occupancy (Fig. 4), and the possibility to provide effective protection against O2 by transiently turning the sample into the sulfide-bound Hinact state (Fig. 5).14We determined differences in the yields from heterologous expression and protein purification between the monomeric variants and the heterodimeric wild-type (Fig. 4). Based on an average of three independent protein purifications, we calculated for L3 the lowest relative yield of 46% compared to DdHydABWT. For fusion variants L4 and L1, the relative yields were slightly higher (50% and 58%). Variant L2, which already enabled an approximately 15-fold maturation rate increase, not only reached and maintained the highest H2-evolution rate (118% compared to wild-type) but also exhibited the highest relative expression yield of 77%. The retention of the protective effect of sulfide treatment against molecular O2 was examined via potential-step voltammetry with (Fig. 5a) and without the addition of 1 mM Na2S prior to an injection of 50 µM O2 (Fig. 5b), as previously described by Rodríguez-Maciá and coworkers.14,15 After determining the original level of catalytic current resulting from H2-evolution at −0.5 V vs. SHE, the potential was stepped up to 0.1 V vs. SHE supporting H2-oxidation. Under such oxidative conditions, as soon as 1 mM Na2S is added to the measuring solution, all four monomeric variants show a rapid and complete loss of catalytic current just like DdHydABWT (Fig. 5a), indicating for all proteins a likewise effective and potential-dependent inactivation via sulfide-binding to the active site. To further verify that the attached sulfide ligand protects the active site against O2-induced damage, O2-saturated buffer was injected to reach a final O2 concentration of 50 µM in the measuring buffer. For the sulfide-treated enzyme sample, the addition of O2-saturated buffer only caused a minimal loss of residual activity. When triggering the release of sulfide from the active site by switching back to −0.5 V vs. SHE, the activities of all investigated proteins recovered accordingly. A comparison between the initial H2-evolution current and the reductive current after O2-exposure suggests for WT and all monomeric variants a comparable decrease in activity, mostly due to film loss from the electrode surface, while in all cases only a very small fraction of the enzyme is irreversibly inactivated by O2. To compare the impact of O2-exposure on enzyme activity between sulfide protected and unprotected samples, a second series of experiments was performed in which O2 exposure was carried out without prior sulfide addition (Fig. 5b). As expected, all enzyme variants show an instantaneous and complete loss of catalytic activity upon O2-addition, which is not recovered upon stepping the potential back to −0.5 V vs. SHE. Thus, all monomeric proteins can be effectively protected against irreversible O2-induced inactivation via pre-exposure to sulfide (Fig. 5a). When kept anaerobically, each variant retains a high catalytic activity comparable to DdHydABWT (Fig. 3b).
Structural analysis
When designing the four linker elements, we aimed at covering a range of lengths and amino acid compositions, considering features such as bulkier side chains that influence the local flexibility based on preceding AlphaFold modelling.44 Accordingly, different characteristics emerged in maturation efficiency, H2-evolution activity, and heterologous expression. To elucidate the structural basis for the strong impact of subunit fusion on enzyme maturation of DdHydAB and to further identify the steric individualities responsible for differences in maturation rate increases between L1 and the other variants, we solved the crystal structures of variants L1–L4. Crystals of variants L1 and L4 were obtained under similar conditions to those previously established for the wild-type,45 whereas variants L2 and L3 required additional optimizations of the crystallization conditions. Diffraction data were collected at beamline P11 at PETRA III (DESY, Hamburg, Germany)46,47 and structures of all monomeric variants were determined at resolutions of 1.05–1.78 Å (Table S3).The linker regions in variants L1, L2, and L3 showed clear electron densities, enabling the modelling of all amino acid residues as well as the tracking of interactions with or within the corresponding loop region (Fig. 2b, S5f, k and p). However, the structures of variant L4 lacked sufficient electron density in the linker segment (Fig. S10), which indicates high mobility of the linker residues. Overall, the protein backbones of the variants are very similar to the wild-type protein, suggesting that the linkers do not affect the overall fold of the protein. We employed ChimeraX48 to superpose a Boltz-2 model49 of L4 with the well-resolved remaining part of the corresponding crystal structure, offering a plausible suggestion for the unresolved linker region in the L4 structure (see Fig. S10).
The 2FeH-subcluster is present in all structures, but with different occupancy levels. In variants L1 and L2, the occupancy of 2FeH has been refined to 0.9 to better match the experimental electron density, while in variants L3 and L4 the 2FeH-subcluster is fully occupied.
L1 and L2 show a well-defined diatomic ligand bound to the open coordination site of Fed with occupancies of 0.5 and 0.9, respectively. We propose that this diatomic ligand corresponds to a CO molecule derived from the synthetic complex used for the maturation of the enzymes. Electron density maps of L3 and L4 show very little positive density at the open coordination site in the Fo–Fc maps, suggesting that most of the molecules in these crystals possess an open coordination site.
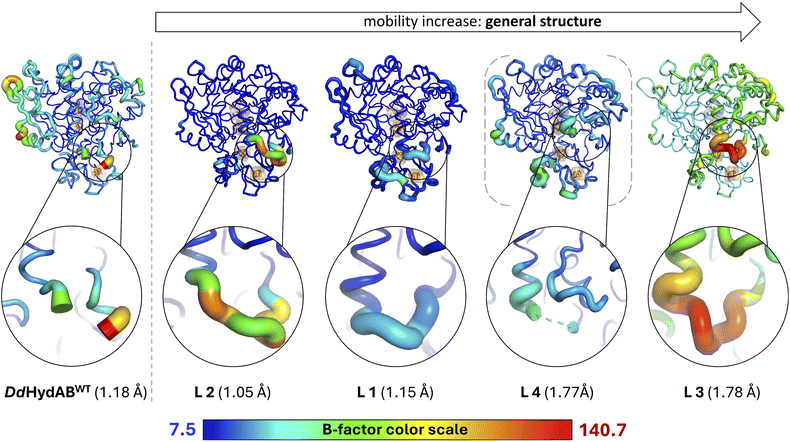
While the structures of all the variants are overall very similar, we observed significant differences in the local mobility between the variants, as highlighted by the visualization of the crystallographic displacement factors (B-factors) in Fig. 6. The displacement of atoms from their mean position in a crystal structure may be the result of temperature-dependent atomic vibrations or static disorder in a crystal lattice. Since all structures were determined from crystals with the same lattice, the different B-factors observed for wild-type and the monomeric variants are more likely to be the result of local mobility. As expected, the heterodimeric DdHydABWT structure shows increased mobility values at the termini of the two subunits, especially at the carboxy terminus of the LSU. The overall mobility of the structures for L1 and L2 is very low; however, the linkers of all variants show individual local flexibilities. The linker of the rather slowly maturating variant L1 is the least mobile one, while the longer linker of L2 is significantly more flexible. Interestingly, the linker of the fastest-maturating variant L3 shows the highest mobility. The L3 structure also shows the highest overall mobility range. However, as the structure of variant L3 was solved at a comparatively lower resolution, the increased B-factors may not be directly comparable with the B-factors determined for the high-resolution structures of L1 and L2. Crystal structures of L3, unlike those of L1 and L2, showed very low occupancies for the surplus CO ligand at the substrate binding site. This may suggest a lower CO-binding affinity or a faster release of the extra ligand, which could result from increased structural mobility compared to the wild-type and the other variants. Even though L3 indeed exhibits the highest kreact value among the five proteins, the difference to the other proteins is not very pronounced (see Table S1). Linker 1 showed by far the least impact on the enzyme maturation rate, being 4.5- to 10-fold lower compared to the other fusion proteins (Fig. 3c). According to a comparison of the side chain packing densities in the linker regions of X-ray structures obtained for L1, L2, and L3, the linker of L1 seems to be more tightly packed, while the side chains of linkers in L2 and L3 are comparatively more loosely embedded (Fig. S11). Thus, they may provide sufficient room for flexible movements (Fig. 2b, S5a, f, k and p). This is likewise reflected in the rather low B-factors assigned to linker 1 (Fig. 6), resulting in high local structural quality, while the higher B-factors for the linkers in structures of L2 and L3 rendered the local quality in the linker regions rather low compared to the remaining structural parts (Fig. 6). These results suggest that stabilization of the N-terminal part of the lock-helix by a linker supports the transition to the stably locked configuration after 2FeH precursor uptake. However, a certain degree of residual flexibility may still be necessary to permit and buffer movements during the structural reconfiguration. In the final section, we examine whether the unusually slow maturation rate of the heterodimeric wild-type enzyme is an intrinsic property of DdHydAB or merely an artifact of the non-native conditions used for in vitro maturation.
Impact of the maturase HydF on maturation
So far, we have only investigated the change in maturation rate upon monomerization with different linkers of DdHydAB by activating the apo-enzyme in vitro with free 2FeMIMH. However, in vivo, the 2FeH precursor is assembled by the maturases HydE and G on the scaffold maturase HydF. Once loaded, HydF is proposed to transfer the 2FeH precursor to the apo-hydrogenase by forming a transient HydF/2FeMIMH/apo-DdHydAB complex. Intermolecular interactions in this ternary complex may stabilize the N-terminus of the small subunit of DdHydAB, thereby enabling fast 2FeH insertion, comparable to the situation in the monomeric DdHydAB versions. To investigate the influence of HydF-supported maturation, we expressed, purified, and loaded DdHydF of D. desulfuricans ATCC 7757 (for sequence of DdHydF, see Fig. S5), as previously done to study the interactions of HydF with different [FeFe]-hydrogenases (Fig. S1).33–35 Loading of DdHydF was verified via electronic absorption spectroscopy (UV-Vis range) (Fig. 7a). HydF-supported activation was tested for the heterodimeric DdHydABWT, the monomeric variant L3, showing the highest activation rate with free 2FeMIMH, and L2, as this variant showed an improved H2-evolution activity even in comparison to DdHydABWT. The activation was tracked using the same assay as before for in vitro maturation, with free 2FeMIMH replaced by equimolar amounts of 2FeMIMH-loaded DdHydF (Fig. 7b). The maturation rates achieved by HydF-mediated activation of the apo-enzyme largely reflect the trends observed when activating the apo-enzyme with free 2FeMIMH. DdHydAB fusion variant L2 shows an even lower t1/2-value of 8.7 ± 0.5 min in the HydF-mediated approach, compared to 14.3 ± 0.9 min when being activated by free 2FeMIMH. L3 reaches a t1/2-value of approximately 4.7 ± 0.9 min in the HydF-supported assay, being only slightly higher than measured for the unmediated assay (2.3 ± 1.1 min). While the monomeric fusion variants were found to show largely comparable half-times for activation in the unmediated and HydF-supported assay, DdHydABWT even showed a two-fold increase in t1/2 (341 ± 154 min compared to 166 ± 25 min). These results suggest that the slow in vitro maturation of DdHydABWT cannot be overcome by the formation of a transient protein–protein complex with DdHydF during the maturation process. Nevertheless, we cannot exclude alternative explanations based on the specific in vivo maturation conditions of periplasmic [FeFe]-hydrogenases such as DdHydABWT (see SI file S2). Our results, however, underscore the importance of subunit fusion, as exemplified by L2 and L3, for achieving high maturation rates, irrespective of the mediator function of HydF.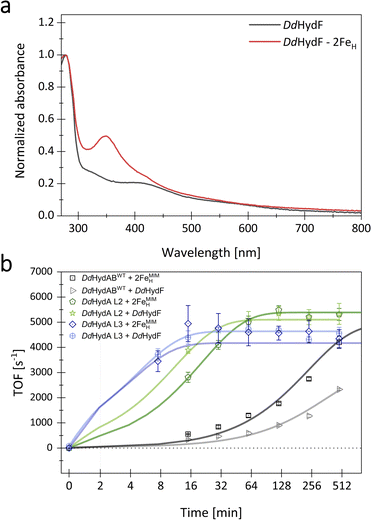 | ||
| Fig. 7 Loading and use of DdHydF as a mediator for in vitro maturation of DdHydABWT and subunit-fusion variant L2. (a) UV-Vis-spectra of as-purified DdHydF and DdHydF loaded with 2FeMIMH. The spectra were normalized by setting the peak at 280 nm to 1.0. Protein samples were diluted with 0.1 M Tris–HCl buffer, pH 8. Both spectra are in line with those earlier reported for the apo- and cofactor-loaded holo-state of HydF from Thermotoga maritima.28 (b) Monitoring the time-dependent increase of H2-evolution activity for DdHydABWT and subunit-fusion variants L2 and L3 during in vitro maturation of 0.16 mM hydrogenase, using either a 3-fold concentration of maturase-free 2FeMIMH or holo-HydF loaded with 2FeMIMH. | ||
Conclusion
We demonstrated that using a linker with specific properties to connect the two subunits of the heterodimeric [FeFe]-hydrogenase DdHydAB significantly enhances the rate of in vitro maturation, while retaining the important catalytic features of this enzyme, such as high activity and affinity for the protecting inhibitor sulfide. Our data provide important insights into the mechanism and rate-limiting factors of cofactor maturation. Our results support the hypothesis of a maturation mechanism where 2FeMIMH must first be tightly bound and stabilized in the binding pocket by reconfigurations in the plug-lock-lid system before subcluster coupling can occur. Consequently, efficient in vitro maturation depends on the right degree of flexibility of the structural elements involved. This flexibility must allow for a fully open conformation to enable efficient and fast 2FeH uptake via the maturation channel to the 4FeH-moiety of the H-cluster. At the same time, the structural elements still need to be rigid enough to sufficiently support the conformational changes required to close and lock up the maturation channel before forcing the 2FeH-subcluster into its rotated, catalytically preferred conformation and enabling sub-cluster coupling. On the path toward establishing DdHydAB as a competitive and fully regenerative biocatalyst in future applications, subunit-fusion allowed us to overcome the hurdle of the extremely slow catalytic activation of the heterodimeric wild-type enzyme.Author contributions
M. W., J. J., N. P., I. S. and C. L. conceived and planned the experiments. J. J., M. W., J. B., M. M. and K. B. carried out the experiments. U. P. A. and S. Y. provided the cofactor for the in vitro maturation experiments. M. W., J. J., N. P., I. S., K. B. and J. A. B. contributed to the interpretation of the results. M. W., J. J. and N. P. took the lead in writing the manuscript. All authors provided critical feedback and helped shape the research, analysis and manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: File S1: materials and methods, text, Fig. S1–S12, and Tables S1–S4. A separate file (deposition reports) comprises all deposition reports associated with the newly published X-ray structure data summarized in Table S3. File S2: genomic data, polypeptide sequences, and further information on the origin and phylogenetic distribution of heterodimeric M2-type [FeFe]-hydrogenases. It also provides a hypothesis for the evolutionary pressures that may have led to the unusual heterodimeric configuration of periplasmic M2-type [FeFe]-hydrogenases and may help explain the low in vitro maturation rate of this M2 subtype. See DOI: https://doi.org/10.1039/d5sc07299a.Acknowledgements
J. J. gratefully acknowledges support by the German Academic Scholarship Foundation. N. P. and M. W. acknowledge financial support by the BMBF funding program “Future Technologies for the Industrial Bioeconomy” (SynHydro3, Project No. 031B1123A and 031B1123C). M. W. was further funded by the Deutsche Forschungsgemeinschaft (DFG) – Project No. 561053011. N. P. was further funded by the FNR project SynergyFuels (16RK34003K) and by the Deutsche Forschungsgemeinschaft (DFG) – Project No. 490864939 (Prosecco). This work was financially supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Priority Program “Iron–Sulfur for Life: Cooperative Function of Iron–Sulfur Centers in Assembly, Biosynthesis, Catalysis and Disease” (SPP 1927) Projects BI 2198/1-1 (J. A. B.) and IS 1476/4-1 (I. S.). I. S. further acknowledges support from COST Action FeSImmChemNet (CA21115) supported by COST (European Cooperation in Science and Technology). U.-P. A. was funded by the DFG under Germany's Excellence Strategy - EXC 2033 – 390677874 – RESOLV. CL acknowledges funding by Agence nationale de la recherche (ANR) (HYDRES, ANR-23-CE50-0016) and by the French government under the France 2030 investment plan, as part of the Initiative d'Excellence d'Aix-Marseille Université – A*MIDEX, AMX-22-RE-AB-097. A*Midex is a foundation of Aix Marseille University. We acknowledge DESY (Hamburg, Germany), a member of the Helmholtz Association HGF, for the provision of experimental facilities. Parts of this research were carried out at PETRA III. Data were collected using beamline P11 operated by DESY Photon Science. We would like to thank Johanna Hakanpää and Guillaume Pompidor for assistance during the experiments. Beamtime was allocated for proposal I-20230763.Notes and references
- M. Qiao, Catalyzing the chemical industry, Nat. Chem. Eng., 2024, 1, 270–272 CrossRef.
- J. M. Le and K. L. Bren, Engineered Enzymes and Bioinspired Catalysts for Energy Conversion, ACS Energy Lett., 2019, 4, 2168–2180 CrossRef CAS.
- A. A. Karyakin, S. V. Morozov, E. E. Karyakina, N. A. Zorin, V. V. Perelygin and S. Cosnier, Hydrogenase electrodes for fuel cells, Biochem. Soc. Trans., 2005, 33, 73–75 CrossRef CAS PubMed.
- K. A. Vincent, J. A. Cracknell, A. Parkin and F. A. Armstrong, Hydrogen cycling by enzymes: electrocatalysis and implications for future energy technology, Dalton Trans., 2005, 3397–3403, 10.1039/B508520A.
- K. A. Vincent, J. A. Cracknell, O. Lenz, I. Zebger, B. Friedrich and F. A. Armstrong, Electrocatalytic hydrogen oxidation by an enzyme at high carbon monoxide or oxygen levels, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 16951–16954 CrossRef CAS PubMed.
- A. A. Oughli, F. Conzuelo, M. Winkler, T. Happe, W. Lubitz, W. Schuhmann, O. Rudiger and N. Plumere, A redox hydrogel protects the O2 -sensitive [FeFe]-hydrogenase from Chlamydomonas reinhardtii from oxidative damage, Angew Chem. Int. Ed. Engl., 2015, 54, 12329–12333 CrossRef CAS PubMed.
- S. Hardt, S. Stapf, D. T. Filmon, J. A. Birrell, O. Rüdiger, V. Fourmond, C. Léger and N. Plumeré, Reversible H2 oxidation and evolution by hydrogenase embedded in a redox polymer film, Nat. Catal., 2021, 4, 251–258 CrossRef CAS PubMed.
- S. V. Hexter, F. Grey, T. Happe, V. Climent and F. A. Armstrong, Electrocatalytic mechanism of reversible hydrogen cycling by enzymes and distinctions between the major classes of hydrogenases, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 11516–11521 CrossRef CAS PubMed.
- J. T. Kleinhaus, F. Wittkamp, S. Yadav, D. Siegmund and U.-P. Apfel, [FeFe]-Hydrogenases: maturation and reactivity of enzymatic systems and overview of biomimetic models, Chem. Soc. Rev., 2021, 50, 1668–1784 RSC.
- K. Pandey, S. T. Islam, T. Happe and F. A. Armstrong, Frequency and potential dependence of reversible electrocatalytic hydrogen interconversion by [FeFe]-hydrogenases, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 3843–3848 CrossRef CAS PubMed.
- M. Heghmanns, A. Rutz, Y. Kutin, V. Engelbrecht, M. Winkler, T. Happe and M. Kasanmascheff, The oxygen-resistant [FeFe]-hydrogenase CbA5H harbors an unknown radical signal, Chem. Sci., 2022, 13, 7289–7294 RSC.
- M. Winkler, J. Duan, A. Rutz, C. Felbek, L. Scholtysek, O. Lampret, J. Jaenecke, U. P. Apfel, G. Gilardi, F. Valetti, V. Fourmond, E. Hofmann, C. Leger and T. Happe, A safety cap protects hydrogenase from oxygen attack, Nat. Commun., 2021, 12, 756 CrossRef CAS PubMed.
- S. Morra, M. Arizzi, F. Valetti and G. Gilardi, Oxygen Stability in the New [FeFe]-Hydrogenase from Clostridium beijerinckii SM10 (CbA5H), Biochemistry, 2016, 55, 5897–5900 CrossRef CAS PubMed.
- P. Rodríguez‐Maciá, L. M. Galle, R. Bjornsson, C. Lorent, I. Zebger, Y. Yoda, S. P. Cramer, S. DeBeer, I. Span and J. A. Birrell, Caught in the Hinact : Crystal Structure and Spectroscopy Reveal a Sulfur Bound to the Active Site of an O2 -stable State of [FeFe] Hydrogenase, Angew Chem. Int. Ed. Engl., 2020, 59, 16786–16794 CrossRef PubMed.
- P. Rodríguez-Maciá, E. J. Reijerse, M. van Gastel, S. DeBeer, W. Lubitz, O. Rüdiger and J. A. Birrell, Sulfide Protects [FeFe] Hydrogenases From O2, J. Am. Chem. Soc., 2018, 140, 9346–9350 CrossRef PubMed.
- A. A. Oughli, S. Hardt, O. Rudiger, J. A. Birrell and N. Plumere, Reactivation of sulfide-protected [FeFe] hydrogenase in a redox-active hydrogel, Chem. Commun., 2020, 56, 9958–9961 RSC.
- D. T. Filmon, J. Jaenecke, M. Winkler, V. Fourmond, C. Leger and N. Plumere, Turning the FeFe hydrogenase from Clostridium beijerinckii into an efficient H(2) oxidation catalyst using a redox-active matrix, Proc. Natl. Acad. Sci. U. S. A., 2025, 122, e2514698122 CrossRef CAS PubMed.
- S. E. Cleary, S. J. Hall, R. Galan-Bataller, T. C. Lurshay, C. Hancox, J. J. Williamson, J. T. Heap, H. A. Reeve and S. Morra, Scalable Bioreactor Production of an O2-Protected [FeFe]-Hydrogenase Enables Simple Aerobic Handling for Clean Chemical Synthesis, ChemCatChem, 2024, 16, e202400193 CrossRef CAS.
- S. Yadav, R. Haas, E. B. Boydas, M. Roemelt, T. Happe, U.-P. Apfel and S. T. Stripp, Oxygen sensitivity of [FeFe]-hydrogenase: a comparative study of active site mimics inside vs. outside the enzyme, Phys. Chem. Chem. Phys., 2024, 26, 19105–19116 RSC.
- J. Esselborn, C. Lambertz, A. Adamska-Venkatesh, T. Simmons, G. Berggren, J. Noth, J. Siebel, A. Hemschemeier, V. Artero, E. Reijerse, M. Fontecave, W. Lubitz and T. Happe, Spontaneous activation of [FeFe]-hydrogenases by an inorganic [2Fe] active site mimic, Nat. Chem. Biol., 2013, 9, 607–609 CrossRef CAS PubMed.
- J. A. Birrell, K. Wrede, K. Pawlak, P. Rodriguez-Maciá, O. Rüdiger, E. J. Reijerse and W. Lubitz, Artificial Maturation of the Highly Active Heterodimeric [FeFe] Hydrogenase from Desulfovibrio desulfuricans ATCC 7757, Isr. J. Chem., 2016, 56, 852–863 CrossRef CAS.
- E. C. Hatchikian, N. Forget, V. M. Fernandez, R. Williams and R. Cammack, Further characterization of the [Fe]-hydrogenase from Desulfovibrio desulfuricans ATCC 7757, Eur. J. Biochem., 1992, 209, 357–365 CrossRef CAS PubMed.
- B. R. Glick, W. G. Martin and S. M. Martin, Purification and properties of the periplasmic hydrogenase from Desulfovibrio desulfuricans, Can. J. Microbiol., 1980, 26, 1214–1223 CrossRef CAS PubMed.
- O. Lampret, J. Esselborn, R. Haas, A. Rutz, R. L. Booth, L. Kertess, F. Wittkamp, C. F. Megarity, F. A. Armstrong, M. Winkler and T. Happe, The final steps of [FeFe]-hydrogenase maturation, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 15802–15810 CrossRef CAS PubMed.
- C. F. Megarity, J. Esselborn, S. V. Hexter, F. Wittkamp, U. P. Apfel, T. Happe and F. A. Armstrong, Electrochemical Investigations of the Mechanism of Assembly of the Active-Site H-Cluster of [FeFe]-Hydrogenases, J. Am. Chem. Soc., 2016, 138, 15227–15233 CrossRef CAS PubMed.
- D. W. Mulder, E. S. Boyd, R. Sarma, R. K. Lange, J. A. Endrizzi, J. B. Broderick and J. W. Peters, Stepwise [FeFe]-hydrogenase H-cluster assembly revealed in the structure of HydA(DeltaEFG), Nature, 2010, 465, 248–251 CrossRef CAS PubMed.
- B. Németh, M. Senger, H. J. Redman, P. Ceccaldi, J. Broderick, A. Magnuson, S. T. Stripp, M. Haumann and G. Berggren, [FeFe]-hydrogenase maturation: H-cluster assembly intermediates tracked by electron paramagnetic resonance, infrared, and X-ray absorption spectroscopy, JBIC, J. Biol. Inorg. Chem., 2020, 25, 777–788 Search PubMed.
- R. Haas, O. Lampret, S. Yadav, U.-P. Apfel and T. Happe, A Conserved Binding Pocket in HydF is Essential for Biological Assembly and Coordination of the Diiron Site of [FeFe]-Hydrogenases, J. Am. Chem. Soc., 2024, 146, 15771–15778 CrossRef CAS PubMed.
- A. Pagnier, B. Balci, E. M. Shepard, W. E. Broderick and J. B. Broderick, [FeFe]-Hydrogenase In Vitro Maturation, Angew. Chem., Int. Ed., 2022, 61, e202212074 Search PubMed.
- J. Esselborn, N. Muraki, K. Klein, V. Engelbrecht, N. Metzler-Nolte, U. P. Apfel, E. Hofmann, G. Kurisu and T. Happe, A structural view of synthetic cofactor integration into [FeFe]-hydrogenases, Chem. Sci., 2016, 7, 959–968 RSC.
- Y. Zhang, L. Tao, T. J. Woods, R. D. Britt and T. B. Rauchfuss, Organometallic Fe2(µ-SH)2(CO)4(CN)2 Cluster Allows the Biosynthesis of the [FeFe]-Hydrogenase with Only the HydF Maturase, J. Am. Chem. Soc., 2022, 144, 1534–1538 Search PubMed.
- R. D. Britt, T. B. Rauchfuss and G. Rao, The H-cluster of [FeFe] Hydrogenases: Its Enzymatic Synthesis and Parallel Inorganic Semisynthesis, Acc. Chem. Res., 2024, 57, 1941–1950 CrossRef CAS PubMed.
- B. Németh, H. Land, A. Magnuson, A. Hofer and G. Berggren, The maturase HydF enables [FeFe] hydrogenase assembly via transient, cofactor-dependent interactions, J. Biol. Chem., 2020, 295, 11891–11901 CrossRef PubMed.
- S. E. McGlynn, E. M. Shepard, M. A. Winslow, A. V. Naumov, K. S. Duschene, M. C. Posewitz, W. E. Broderick, J. B. Broderick and J. W. Peters, HydF as a scaffold protein in [FeFe] hydrogenase H-cluster biosynthesis, FEBS Lett., 2008, 582, 2183–2187 CrossRef CAS PubMed.
- G. Berggren, A. Adamska, C. Lambertz, T. R. Simmons, J. Esselborn, M. Atta, S. Gambarelli, J. M. Mouesca, E. Reijerse, W. Lubitz, T. Happe, V. Artero and M. Fontecave, Biomimetic assembly and activation of [FeFe]-hydrogenases, Nature, 2013, 499, 66–69 CrossRef CAS PubMed.
- G. Caserta, L. Pecqueur, A. Adamska-Venkatesh, C. Papini, S. Roy, V. Artero, M. Atta, E. Reijerse, W. Lubitz and M. Fontecave, Structural and functional characterization of the hydrogenase-maturation HydF protein, Nat. Chem. Biol., 2017, 13, 779–784 CrossRef CAS PubMed.
- O. Lampret, A. Adamska-Venkatesh, H. Konegger, F. Wittkamp, U. P. Apfel, E. J. Reijerse, W. Lubitz, O. Rudiger, T. Happe and M. Winkler, Interplay between CN(-) Ligands and the Secondary Coordination Sphere of the H-Cluster in [FeFe]-Hydrogenases, J. Am. Chem. Soc., 2017, 139, 18222–18230 CrossRef CAS PubMed.
- A. Adamska-Venkatesh, D. Krawietz, J. Siebel, K. Weber, T. Happe, E. Reijerse and W. Lubitz, New redox states observed in [FeFe] hydrogenases reveal redox coupling within the H-cluster, J. Am. Chem. Soc., 2014, 136, 11339–11346 Search PubMed.
- Y. Nicolet, C. Piras, P. Legrand, C. E. Hatchikian and J. C. Fontecilla-Camps, Desulfovibrio desulfuricans iron hydrogenase: the structure shows unusual coordination to an active site Fe binuclear center, Structure, 1999, 7, 13–23 Search PubMed.
- N. Amet, W.-C. Lee Hf Fau-Shen and W. C. Shen, Insertion of the designed helical linker led to increased expression of tf-based fusion proteins, Pharm. Res., 2009, 26, 523–528 Search PubMed.
- J. Guo, Z. Cheng, J. Berdychowska, X. Zhu, L. Wang, L. Peplowski and Z. Zhou, Effect and mechanism analysis of different linkers on efficient catalysis of subunit-fused nitrile hydratase, Int. J. Biol. Macromol., 2021, 181, 444–451 Search PubMed.
- D. K. Patel, D. Vettiatil, D. H. Patel and G. Dave, Linkers: A synergistic way for the synthesis of chimeric proteins, Protein Expression Purif., 2022, 191, 106012 Search PubMed.
- F. Leroux, S. Dementin, B. Burlat, L. Cournac, A. Volbeda, S. Champ, L. Martin, B. Guigliarelli, P. Bertrand, J. Fontecilla-Camps, M. Rousset and C. Léger, Experimental approaches to kinetics of gas diffusion in hydrogenase, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 11188–11193 Search PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. A. A. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler, T. Back, S. Petersen, D. Reiman, E. Clancy, M. Zielinski, M. Steinegger, M. Pacholska, T. Berghammer, S. Bodenstein, D. Silver, O. Vinyals, A. W. Senior, K. Kavukcuoglu, P. Kohli and D. Hassabis, Highly accurate protein structure prediction with AlphaFold, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- M. A. Martini, K. Bikbaev, Y. Pang, C. Lorent, C. Wiemann, N. Breuer, I. Zebger, S. DeBeer, I. Span, R. Bjornsson, J. A. Birrell and P. Rodríguez-Maciá, Binding of exogenous cyanide reveals new active-site states in [FeFe] hydrogenases, Chem. Sci., 2023, 14, 2826–2838 Search PubMed.
- A. Meents, B. Reime, N. Stuebe, P. Fischer, M. Warmer, D. Göries, J. Roever, J. Meyer, J. Fischer, A. Burkhardt, I. Vartiainen, P. Karvinen and C. David, Development of an in-vacuum X-ray microscope with cryogenic sample cooling for beamline P11 at PETRA III, Proc. SPIE, 2013, 8851, 88510K Search PubMed.
- A. Burkhardt, T. Pakendorf, B. Reime, J. Meyer, P. Fischer, N. Stübe, S. Panneerselvam, O. Lorbeer, K. Stachnik, M. Warmer, P. Rödig, D. Göries and A. Meents, Status of the crystallography beamlines at PETRA III, Eur. Phys. J. Plus, 2016, 131, 56 CrossRef.
- E. C. Meng, T. D. Goddard, E. F. Pettersen, G. S. Couch, Z. J. Pearson, J. H. Morris and T. E. Ferrin, UCSF ChimeraX: Tools for structure building and analysis, Protein Sci., 2023, 32, e4792 Search PubMed.
- S. Passaro, G. Corso, J. Wohlwend, M. Reveiz, S. Thaler, V. R. Somnath, N. Getz, T. Portnoi, J. Roy, H. Stark, D. Kwabi-Addo, D. Beaini, T. Jaakkola and R. Barzilay, Boltz-2: Towards Accurate and Efficient Binding Affinity Prediction, bioRxiv, 2025, preprint, DOI:10.1101/2025.06.14.659707.
| This journal is © The Royal Society of Chemistry 2026 |