Open Access Article
Open Access ArticleSynthesis, characterization, and α-glucosidase inhibitory activity of methyl proline derivatives of phenylpropanoid
Weiwei Zhang†
 *a,
Huiwen Yang†a,
Xiangdi Yaoa,
Ruipeng Lia,
Yao Yaoa,
Wei Maa,
Kailiang Maa,
Lufei Shaob and
Pengjuan Zhouc
*a,
Huiwen Yang†a,
Xiangdi Yaoa,
Ruipeng Lia,
Yao Yaoa,
Wei Maa,
Kailiang Maa,
Lufei Shaob and
Pengjuan Zhouc
aSchool of Basic Medical Sciences, Ningxia Medical University, Yinchuan 750004, PR China. E-mail: 20190033@nxmu.edu.cn
bThe Neurology Department, General Hospital of Ningxia Medical University, Yinchuan 750004, PR China
cInstitute of Agricultural Resources and Environment, Ningxia Academy of Agriculture and Forestry Science, Yinchuan 750004, PR China
First published on 5th May 2026
Abstract
To discover novel α-glucosidase inhibitors for diabetes management, a series of six proline methyl ester phenylpropanoid derivatives were synthesized through a seven-step synthetic route involving bromination and nucleophilic substitution. The structures of the target compounds (2f–7f) were confirmed by 1H NMR, 13C NMR, and HRMS. Using the PNPG method, we evaluated their inhibitory activity against α-glucosidase and found that some compounds acted as effective inhibitors. Among them, compound 6f demonstrated the strongest activity, with an IC50 value of 91 µM. Molecular docking elucidated that 6f exploits its specific substitution pattern to form a robust complex, stabilized by a near-parallel π–π stacking with Trp59 and a dense hydrogen-bonding network. The proline ring further enhances stability through deep complementary burial within a hydrophobic cleft, resulting in a highly stable binding pose. Further insights were provided by DFT calculations and wavefunction analyses (including IRI, RDG, and ESP), which revealed that favorable electrostatic complementarity and intramolecular non-covalent interactions contribute to its binding affinity. Preliminary ADME predictions indicated promising drug-like properties for this series. In conclusion, compound 6f shows strong potential as an α-glucosidase inhibitor and merits further investigation for the treatment of diabetes.
1 Introduction
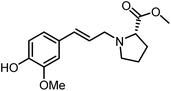
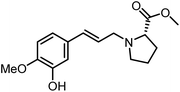
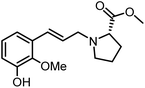
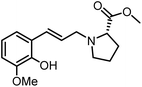
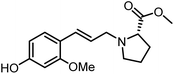
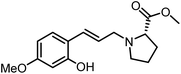
The genus Ficus (Moraceae) represents about 800 species of woody trees, shrubs, vines and climbers which are widely distributed in tropical and subtropical regions.1 Plants of the genus Ficus comprise many varieties, notable generic diversity, and exceptional pharmacological activities which have been used traditionally against an array of human and animal diseases related to digestive, respiratory, endocrine, reproductive systems and also a cure for gastrointestinal and urinary tract infections.2To the best of our knowledge, only two phenylalanine-derived natural products (Phenyliana and N-trans-ferulic-L-proline methyl ester) bearing a proline unit have been reported in the literature (Fig. 1).3,4 Research into their biological activity suggests these compounds may exhibit certain α-glycosidase inhibitory effects.3 Therefore, we endeavour to synthesise the natural molecules and their derivatives, whilst more importantly, to determine their potential biological activity by assessing their α-glycosidase inhibitory effects.
In this study, six phenylpropanoid derivatives featuring proline structures with regionally differentiated substituents on the benzene ring were synthesised and characterised. These compounds underwent bioactivity assessment and antimicrobial experimental studies as potential α-glycosidase inhibitors. Molecular docking and density functional theory calculations were performed on the most optimal inhibitor to elucidate its interaction with α-glycosidase.
2 Materials and methods
2.1 Experimental details
Details of the chemical synthesis are available in the SI.2.2 Antibacterial assay
The culture medium used was LB Broth (BS1302, Basebio). The experimental bacterial strains included Micrococcus luteus, Bacillus sphaericus, Staphylococcus aureus, and Escherichia coli.The minimum inhibitory concentration (MIC) was determined using the broth microdilution (BMD) method to evaluate the in vitro antibacterial efficacy.5 The principle is as follows: the test agent is serially diluted with the bacterial culture, and after incubation, turbidity is observed visually. The lowest concentration showing no bacterial growth is defined as the MIC.
2.3 α-Glucosidase inhibition assay
The α-glucosidase activity was determined using the PNPG (p-nitrophenyl-β-D-galactopyranoside) method as described in the literature.6,7 The enzyme catalyzes the hydrolysis of the substrate PNPG to produce yellow PNP, which exhibits absorbance at 405 nm. The presence of an inhibitor reduces product formation and decreases the absorbance, allowing for the evaluation of inhibitory activity. Each group was tested in triplicate, and the data were analyzed using GraphPad Prism 8.2.4 Molecular docking and molecular dynamics simulations
In this study, the three-dimensional crystal structure of the α-glucosidase was obtained from the Protein Data Bank (PDB), with PDB ID 5KEZ.8 The three-dimensional structure of the ligand molecule, six compounds were obtained following geometric optimisation using the Gaussian 16W software package at the B3LYP/6-31G* (d,p) theoretical level.9 Molecular docking studies were performed using Discovery Studio 2024 software.10 Prior to docking, the receptor protein underwent pre-processing, including dehydration, hydrogenation and residue repair. Prior to molecular docking, the co-crystallized peptide inhibitor piHA was removed from the complex structure (PDB ID: 5KEZ) to expose the active site. Using PyMOL, the piHA chain and crystallographic waters were deleted, retaining only the protein monomer. The structure was then hydrogenated, assigned charges, and energy-minimized to relieve steric clashes. This step is essential as piHA, a competitive substrate analog, obstructs the binding cavity, thereby preventing the docking of novel synthetic ligands.Molecular dynamics simulations were performed using the GROMACS 2025 software package (Linux environment). The optimal complex conformation obtained from molecular docking was used as the initial structure. The protein and ligand were modelled using the GROMACS CHARMM36 all-atom force field and the corresponding force field parameters generated by the CGenFF server, respectively.11 The complex was placed within a cubic periodic box, solvated using the TIP3P water model, and an appropriate number of Na+ or Cl− ions were added to neutralise the net charge of the system. The simulation procedure was as follows: first, energy minimisation was performed using the steepest descent method until the maximum force fell below 1000 kJ (mol−1 nm−1); Subsequently, a 1 ns pre-equilibration was performed under the NVT and NPT ensembles, gradually coupling the system temperature to 300 K and the pressure to 1 bar (using the Berendsen and Parrinello–Rahman methods); finally, a 100 ns production run was carried out under the isothermal-isobaric ensemble. All bond length constraints in the simulation were implemented using the LINCS algorithm; the cutoff for non-bonded interactions was set to 1.2 nm; and long-range electrostatic interactions were handled using the PME method.12 Simulation trajectories were saved every 10 ps for subsequent analysis.
The 100 ns simulation trajectories were analysed using built-in GROMACS tools and custom scripts. The structural stability of the protein backbone atoms, ligand molecules and the complex was assessed by calculating the root-mean-square deviation (RMSD),13,14 the flexibility changes of individual protein residues during the simulation were analysed using root-mean-square fluctuations (RMSF); and the overall structural compactness of the protein was monitored using the radius of gyration (Rg).15,16 Furthermore, the gmx hbond tool was employed to analyse the number of hydrogen bond interactions formed between the ligand and the receptor during the simulation, as well as their evolution over time.
2.5 Computational methods and model development
To investigate in depth the electronic structure and reactivity of the ligand methyl (E)-(3-(4-hydroxy-2-methoxyphenyl)allyl)-L-prolinate (6f), high-precision quantum chemical calculations were performed in the Gaussian 16W software based on the preferred conformation obtained from molecular docking. Using the density functional theory (DFT) method, geometric optimisation and frequency analysis were performed on the ligand molecule at the M062X/6-311G** level (to ensure it represents a stable point on the potential energy surface).17,18 All calculations took into account solvation effects, employing a density-based solvation model (SMD) with water as the solvent.3 Results and discussion
3.1 Chemistry
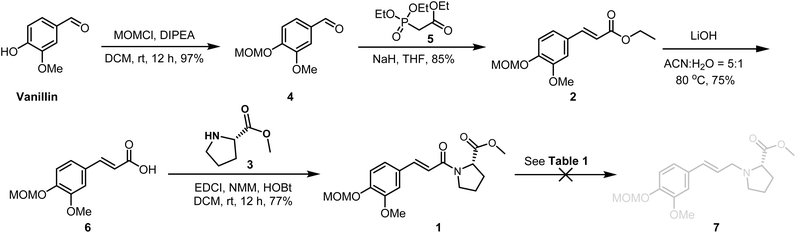
The main text of the article should appear here with headings as appropriate. Our retrosynthetic analysis of Phenyliana and its derivatives is shown in Scheme 1. We envisioned that Phenyliana, N-trans-ferulic-L-proline methyl ester and its derivatives could be generated from the compound 1 through reduction of the amide carbonyl group to a methylene group.19–21 The compound 1 could then be derived from trans methyl phenylacrylate 2 via acids undergoing ester hydrolysis react in amidation reactions.22 Methyl phenylacrylate 2 could be obtained from the readily available commercial raw material vanillin.As depicted in Scheme 2, our synthesis began with commercially available vanillin. Methoxymethoxymethyl protection of the phenolic hydroxyl group afforded aldehyde 4.
Treatment of aldehyde 4 with triethyl phosphonoacetate 5 under Horner–Wadsworth–Emmons reaction olefination conditions afforded trans styrene 2 in 85% yield.23 Subsequently, ethyl acrylate underwent hydrolysis in the presence of lithium hydroxide to yield the corresponding acid 6, which then reacted with L-proline methyl ester via an acylation reaction, successfully producing the amide 1.24 The subsequent amide reduction was not inherently a difficult process, yet in practice proved to be a challenge. As detailed in Table 1, extensive screening of conditions for the selective reduction of the amide group without affecting the ester moiety proved largely unsuccessful.19–21 Consequently, the use of lithium aluminum hydride (LiAlH4) was explored as an alternative to achieve the simultaneous reduction of both functional groups. Following the complete reduction of both amides and esters, the alcohol hydroxyl group could be deoxidized to an acid-a viable alternative.
| Entry | Reaction conditions | Compound 7 |
|---|---|---|
| 1 | Tf2O, HEH, DCM, rt | Not detected |
| 2 | Tetrahydrofuran borane, THF, rt | Not detected |
| 3 | DIBAL-H, THF, 0 °C | Not detected |
| 4 | NaBH4, THF, 0 °C | Not detected |
| 5 | LiHAl4, AlCl3, THF, 0 °C | Complexity |
| 6 | 2,6-Di-tert-butylpyridine, Me3OBF4, DCM, rt | Not detected |
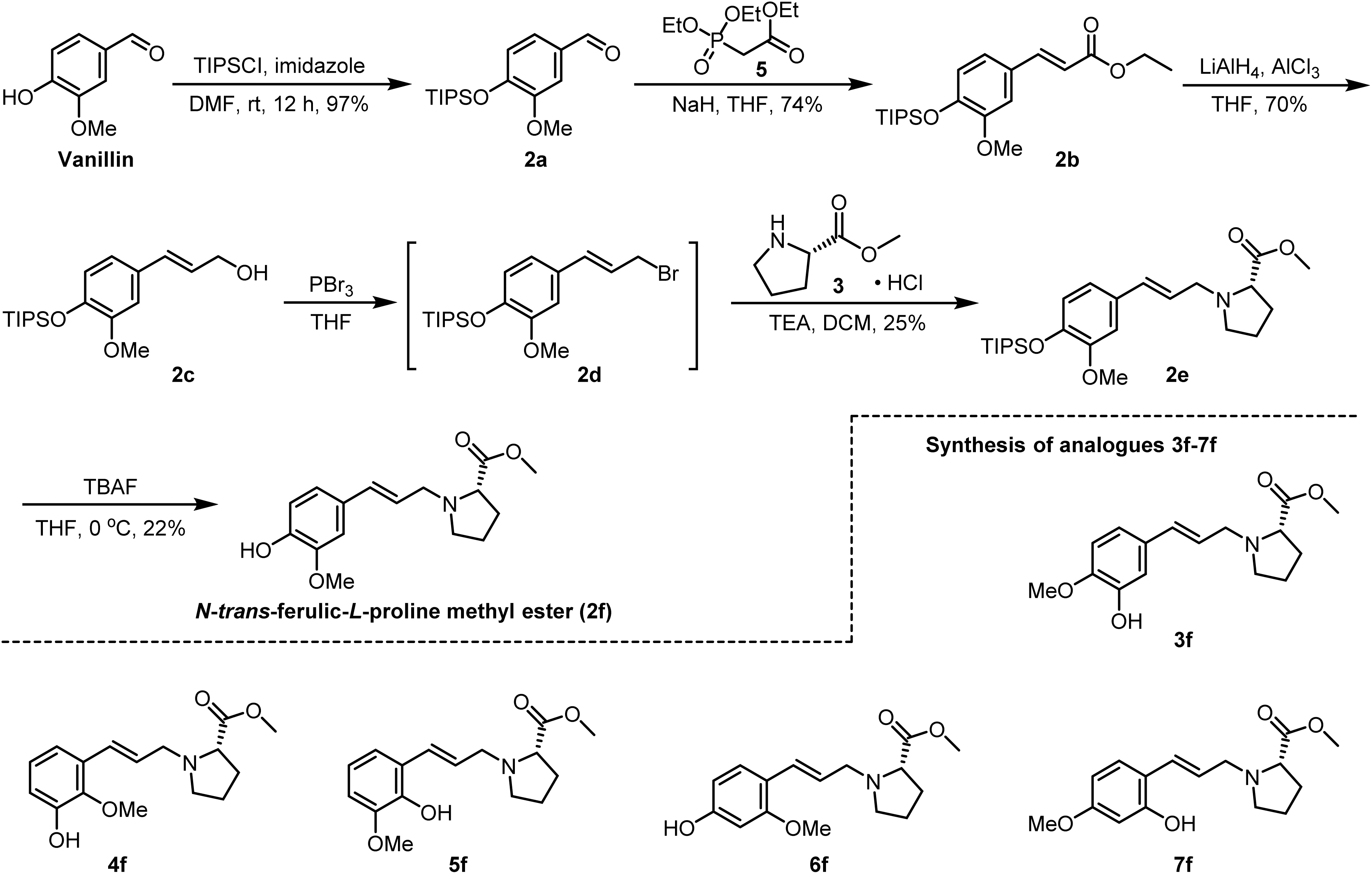
However, the lithium-aluminium hydride system proved complex and failed to yield a clean, single compound. After attempting other conditions, regardless of which reducing agent was altered, the reaction failed to proceed smoothly. Consequently, it became necessary to refine and optimise the aforementioned procedure. Subsequently, we altered the synthetic route, replacing the key amide reduction reaction with a nucleophilic substitution reaction (Scheme 3). Vanillin remained our starting material, with the phenolic hydroxyl group first protected using TIPSCl, treatment of aldehyde 2a with triethyl phosphonoacetate 14 under Horner–Wadsworth–Emmons reaction olefination conditions afforded trans styrene 2b in 85% yield. The reduction of allyl esters to allyl alcohols via lithium–aluminium–hydrogen reactions yielded important intermediate compounds 2c. To convert the alcohol hydroxyl group into a brominated alkyl group (Table 2), phosphorus tribromide, N-iodosaccharin, and N-bromosaccharin were each tested as halogenating reagents.25,26 The use of NISac and NBSac was found problematic, as the desired product co-eluted with the generated triphenylphosphine oxide, complicating purification via conventional chromatography. Therefore, PBr3 proved to be a superior alternative. Following conversion of the allyl alcohol into allyl bromide, the subsequent reaction could proceed directly. The allyl bromide undergoes an intermolecular nucleophilic substitution reaction with L-proline methyl ester hydrochloride, yielding the anticipated compound N-trans-ferulic-L-proline methyl ester (2f).
| Entry | Reaction conditions | Compounds yield |
|---|---|---|
| 1 | PBr3, DIPEA, THF, rt | Trace |
| 2 | PBr3, K2CO3, Et2O, rt | Trace |
| 3 | PBr3, Cs2CO3, Et2O, rt | Trace |
| 4 | PBr3, Cs2CO3, DCM, rt | Trace |
| 5 | PBr3, TEA, CHCl3, rt | Trace |
| 6 | PBr3, TEA, THF, rt | 25% |
| 7 | PBr3, TEA, DCE, rt | Trace |
| 8 | PBr3, TEA, DMF, rt | Trace |
| 9 | PBr3, Cs2CO3, CH3CN, rt | Trace |
| 10 | NBsac, TEA, PPh3, DCM, rt | Trace, contaminated with OPPh3 |
| 11 | NIsac, TEA, PPh3, DCM, rt | Trace, contaminated with OPPh3 |
The immediate next objective is to complete the synthesis of Phenyliana. Proceeding with the synthesis of Phenyliana, analysis of the molecular structure reveals that it can be obtained by demethylating the phenyl ether in N-trans-ferulic-L-proline methyl ester and subsequently hydrolysing the ester group. Indeed, this approach was adopted: first, the methoxy group of the phenolic ether was removed using boron tribromide. Following immediate rotary evaporation, saponification proceeded in a mixed solvent of acetonitrile and water. However, experimental results demonstrated that regardless of the sequence of these two reactions, the resulting product dissolved in water and proved uncollectable alternative methodologies were subsequently employed, yet these too concluded in failure (Scheme 4).
The NMR spectral data of the N-trans-ferulic-L-proline methyl ester synthesized in this study are inconsistent with those previously reported in the literature.3 This discrepancy was unambiguously verified through comprehensive 2D NMR analyses, including DEPT, HSQC, and HMBC spectra. Therefore, it is suggested that the molecular structure reported in prior literature may be inaccurate. Given that this class of compounds is known to exhibit α-glycosidase inhibitory activity, we propose to investigate the structure–activity relationship by systematically modifying the relative positions of the hydroxyl and methoxy substituents on the benzene ring, with the aim of optimizing their biological activity.
By replacing the starting aldehyde and applying an analogous synthetic procedure, a series of six target compounds were successfully obtained (Scheme 3).
3.2 Results of the α-glucosidase inhibition assay
Having obtained these compounds, it was only natural to proceed to investigate their biological activity. Six compounds were initially subjected to antimicrobial testing, employing Micrococcus luteus, Bacillus sphaericus, Staphylococcus aureus and Escherichia coli as test subjects. Results indicated that none of the six compounds exhibited Obvious antimicrobial activity (see Section 4 in SI). Literature review suggested that this class of compounds may possess α-glycosidase inhibitory effects.6,27–30 Consequently, we redirected our experimental focus towards identifying reliable and potential α-glycosidase inhibitors.According to the experimental protocol of the PNPG method, we observed that 3f, 6f and 7f exhibited varying degrees of alpha-glucosidase inhibitory activity (Table 3). Among these, methyl (E)-(3-(4-hydroxy-2-methoxyphenyl)allyl)-L-prolinate (6f) demonstrated a favourable half-maximal inhibitory concentration (IC50) value of 91 µM.
In contrast to the potent activity of compound 6f, the remaining derivatives in the series displayed significantly reduced or no appreciable activity. The structures of these compounds need to be further optimized. The structure–activity relationship of these compounds can provide useful information for the structural optimization of these compounds. Subsequently, we performed comprehensive DFT calculations and molecular docking simulations on compound 6f.
3.3 Analysis and discussion of molecular docking results
Despite their high structural similarity, compounds 2f–7f exhibited markedly divergent binding modes within the active site of pancreatic α-glucosidase (PDB ID: 5KEZ). Docking analyses revealed that while all ligands achieved hydrophobic occlusion within the catalytic pocket, critical variations arose in the π–π stacking geometry between their aromatic rings and the indole side chain of Trp59 (Fig. 2).Inactive isomers (2f–5f, 7f) displayed compromised binding due to suboptimal substitution patterns. Specifically, the positional variation of phenolic hydroxyl and methoxy groups either attenuated the stacking intensity with Trp59 (e.g., compound 5f, which lacked sufficient polar complementarity despite maintaining π–π interactions) or misoriented the proline ester moiety, preventing the formation of stable hydrogen bonds with Thr163 or Asp197. Although some inactive isomers established sporadic hydrogen bonds via phenolic hydroxyl groups with residues such as Asp197 and Gln63, they failed to construct a synergistic anchoring network centered on Trp59, resulting in locally loose conformations.
In stark contrast, the active compound 6f exploited its specific 2-methoxy-4-hydroxy substitution pattern to form a near-parallel π–π stacking interaction with Trp59, serving as a robust hydrophobic anchor. Concurrently, its phenolic hydroxyl and methyl ester carbonyl oxygen atoms engaged in a multipoint hydrogen-bonding network with Asp197, Glu233, and Hie101. Furthermore, the proline ring was deeply embedded within a hydrophobic cleft formed by Ala198, Tyr62, and Leu162, achieving high complementary burial.
The positional isomerism within the 2f–7f series finely tunes the stacking angle relative to Trp59 and the integrity of the polar interaction network. This precise modulation directly determines whether the ligand can achieve a stable, “induced-fit” binding conformation within the active site.
3.4 Results of molecular dynamics simulations
To further evaluate the stability of this complex under dynamic conditions, this study conducted a 100 ns molecular dynamics simulation of the complex formed between α-glucosidase (PDB ID: 5KEZ) and its ligand, methyl (E)-(3-(4-hydroxy-2-methoxyphenyl)allyl)-L-prolinate (6f). The simulation results indicate that the entire system remained highly stable throughout the simulation (Fig. 3). In terms of root-mean-square deviation (RMSD), the RMSD of the protein backbone atoms stabilised at approximately 0.13 nm, with fluctuations not exceeding 0.02 nm, demonstrating good structural stability; the RMSD of the ligand molecule stabilised at around 0.08 nm, with fluctuations within 0.01 nm, indicating that its conformational changes within the binding pocket were extremely limited. The RMSD of the complex as a whole remained at approximately 0.17 nm, with fluctuations not exceeding 0.02 nm, further confirming the overall structural integrity of the system. | ||
| Fig. 3 Molecular dynamics simulations of 6f inhibitors with α-glucosidase: insights into binding stability via RMSD ((A) protein, (B) ligand, (C) complex), (D) RMSF, (E) H-bonds umber, (F) gyrate. | ||
Analysis of the root-mean-square fluctuation (RMSF) revealed that the fluctuations of individual residues in the enzyme were minimal during the simulation, with the overall structure exhibiting good rigidity distribution characteristics. In particular, residues near the active site displayed low flexibility, providing a structural foundation for the stable binding of the ligand. Furthermore, the radius of gyration (Rg) remained stable throughout the simulation, further verifying that the protein did not undergo significant structural relaxation or abnormal folding during the simulation, indicating that its overall conformation was well maintained.
Dynamic analysis of hydrogen bond interactions indicates that, throughout the 100 ns simulation, an average of approximately two hydrogen bonds were maintained between the ligand and the α-glucosidase, with the number of hydrogen bonds remaining relatively stable during the middle and late stages of the simulation, further demonstrating that the key interaction mode is effectively maintained under dynamic conditions. In summary, the molecular dynamics simulation results fully confirm the structural stability of this ligand–enzyme complex under dynamic conditions, providing a reliable theoretical basis for subsequent mechanistic studies and structural optimisation.
3.5 Quantum chemical and wavefunction analysis of ligands
IRI (Interaction Region Indicator) scatter plots were combined with RDG (Reduced Density Gradient) analysis to visualise regions of weak interactions within and between ligand molecules. In the IRI scatter plots, peaks appearing in regions of low density (ρ < 0.05 a.u.) and low RDG values correspond to non-covalent interactions. In conjunction with RDG-coloured isosurface plots, it can be observed that there is a distinct region of van der Waals interactions between the methoxy group on the benzene ring within the ligand and the proline ester moiety, whilst hydrogen-bonding interactions are evident around polar groups such as hydroxyl and methoxy groups. Overall, the ligand molecule stabilises its own conformation through weak interactions at multiple sites, providing a favourable pre-organised structure for target binding.
LOL (Localised Orbital Function) analysis revealed the electronic localisation characteristics of the ligand. The LOL colour-coded plot shows that the benzene ring region exhibits typical delocalised π-electron features (ring-shaped structures formed by high LOL values), whilst the oxygen atoms surrounding the ester and ether bonds display localised lone pair electron features (peaks of high LOL values). These localised electronic regions are precisely the potential donor sites for hydrogen bonding, which is highly consistent with the conclusion observed in molecular docking analysis that the carbonyl oxygen and methoxy oxygen are involved in the interaction.
ESP (electrostatic potential) analysis further clarifies the charge distribution characteristics on the ligand surface. The electrostatic potential contour map of the molecular surface shows that the negatively charged regions (red) are mainly concentrated around the carbonyl, hydroxyl and methoxy oxygen atoms, which are potential electrophilic attack sites; the positively charged regions (blue) are distributed near the methylene of the proline ester and the hydrogen atoms of the benzene ring. This distinct charge separation enables the ligand to form directional electrostatic complementary interactions with the hydrophilic residues of the α-glucosidase (such as Gln and Asp), whilst also generating alkyl interactions with hydrophobic residues (such as Leu and His), thereby providing an electronic explanation for the stable binding mode observed in molecular docking and kinetic simulations.
4 Conclusions
This research successfully designed and synthesized a series of six novel methyl ester derivatives (2f–7f) conjugating phenylpropanoid scaffolds with proline motifs. Among these synthetic analogs, compound 6f, identified as methyl (E)-(3-(4-hydroxy-2-methoxyphenyl)allyl)-L-prolinate, demonstrated the most potent inhibition against α-glucosidase, with a half-maximal inhibitory concentration (IC50) of 91 µM. Docking simulations highlighted that 6f achieves stable inhibition through a multi-pronged mechanism, characterized by a near-parallel π–π stacking interaction with Trp59 and extensive polar contacts with key residues (Asp197, Glu233). This synergistic anchoring network, coupled with the deep hydrophobic burial of the proline moiety, ensures optimal orientation and high binding affinity within the catalytic pocket. The stability of this binding pose and the overall structural integrity of the ligand–enzyme complex were corroborated by 100 ns molecular dynamics simulations, which showed low and stable RMSD values for both the protein backbone and the ligand. Furthermore, density functional theory (DFT) calculations and wavefunction analysis (including IRI, RDG, and ESP) provided electronic-level insights, revealing that favorable electrostatic complementarity and intramolecular weak interactions contribute to its binding affinity. Preliminary ADME predictions indicated promising drug-like properties for these compounds. In summary, compound 6f emerges as a promising lead candidate, characterized by significant inhibitory activity, a clarified binding mechanism, and a stable interaction mode, warranting further investigation for its therapeutic potential in managing diabetes.Author contributions
Weiwei Zhang: writing – review & editing. Huiwen Yang: writing – original draft. Xiangdi Yao: methodology. Ruipeng Li: formal analysis. Yao Yao: validation. Wei Ma: validation. Kailiang Ma: validation. Lufei Shao: validation. Pengjuan Zhou: validation. All authors have given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included the manuscript and supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra02514e.Acknowledgements
This work was supported in part by the National Natural Science Foundation of China (No. 22361039), 2024 Ningxia Hui Autonomous Region Young Scientific and Technological Talent Support and Training Program, Ningxia Hui Autonomous Region Natural Science Foundations (No. 2024AAC03368, No. 2025AAC030827, No. 2026AAC030254).Notes and references
- O. Olaoluwa, O. Taiwo, L. Nahar and S. Sarker, Trends Phytochem. Res, 2022, 6, 46–69 CAS.
- S. B. Badgujar, V. V. Patel, A. H. Bandivdekar and R. T. Mahajan, Pharmaceutical Biology, 2014, 52, 1487–1503 CrossRef CAS PubMed.
- D. Dai, X. Zhang, X. Xu, Z. Li, L. Dong, Q. Chen and R. Zhang, Nat. Prod. Res. Dev., 2024, 36, 1537 CAS.
- D.-C. Dai, X.-F. Xu, H. Yan and Y. Zhang, Biochem. Syst. Ecol., 2022, 105, 104509 CrossRef CAS.
- A. Belley, S. M. Cusick, D. C. Pevear, L. Koeth, J. DiFranco-Fisher, N. Kothari, S. Hawser and G. Moeck, J. Clin. Microbiol., 2025, 63(63), e00661–25 Search PubMed.
- K. Ma, J. Han, L. Bao, T. Wei and W. Liu, J. Nat. Prod., 2014, 77, 942–947 CrossRef CAS PubMed.
- J. Pan, X. Yi, Y. Wang, G. Chen and X. He, J. Agric. Food Chem., 2016, 64, 7475–7480 CrossRef CAS PubMed.
- S. A. K. Jongkees, S. Caner, C. Tysoe, G. D. Brayer, S. G. Withers and H. Suga, Cell Chem. Biol., 2017, 24, 381–390 CrossRef CAS PubMed.
- J. M. Ramos, O. Versiane, J. Felcman and C. A. T. Soto, Spectrochim. Acta, Part A, 2007, 67, 1037–1045 CrossRef PubMed.
- Y. Wu and C. L. Brooks, J. Chem. Inf. Model., 2021, 61, 5535–5549 CrossRef CAS PubMed.
- J. Huang, S. Rauscher, G. Nawrocki, T. Ran, M. Feig, B. L. de Groot, H. Grubmüller and A. D. MacKerell, Nat. Methods, 2017, 14, 71–73 CrossRef CAS PubMed.
- B. Kohnke, C. Kutzner and H. Grubmüller, J. Chem. Theory Comput., 2020, 16, 6938–6949 CrossRef CAS PubMed.
- Y. Kasprzak, J. Rückert, N. Ludolph, C. G. Hübner and H. Paulsen, J. Chem. Phys., 2025, 162, 74107 CrossRef CAS PubMed.
- Y.-K. Kim, L. Cai and Y. Kim, Educ. Psychol. Meas., 2025, 131644251369532 Search PubMed.
- B. A. Bhat, W. R. Mir, B. A. Sheikh, M. Alkanani and M. A. Mir, Sci. Rep., 2022, 12, 7296 CrossRef CAS PubMed.
- F. Beruldsen, M. V. de Freitas and D. A. Antunes, Sci. Rep., 2026, 16, 10035 CrossRef CAS PubMed.
- J. Yang, M. Li, Z. Zhu, F. Han, Y. Ma, J. Hou, Q. Zhao, H. Yuan and X. Li, Front. Bioeng. Biotechnol., 2025, 13, 1618575 CrossRef PubMed.
- Y. Wang, M. Li, P. Duanmu, H. Yang, H. Yuan and X. Li, Sci. Rep., 2025, 16, 263 CrossRef CAS PubMed.
- P.-Q. Huang, Q.-W. Lang and Y.-R. Wang, J. Org. Chem., 2016, 81, 4235–4243 CrossRef CAS.
- P.-Q. Huang and H. Geng, Org. Chem. Front., 2015, 2, 150–158 RSC.
- W. Ou and P.-Q. Huang, Acc. Chem. Res., 2025, 58, 2332–2349 CrossRef CAS PubMed.
- T. Yamada, T. Yoshikawa, N. Tsuji, M. Tamura, M. Gogami, K. Sakata and M. Hatano, Asian J. Org. Chem., 2025, 14, e70171 CrossRef CAS.
- D. Roman, M. Sauer and C. Beemelmanns, Synthesis, 2021, 53, 2713–2739 CrossRef CAS.
- H. Ü. Kaniskan, M. S. Eram, K. Zhao, M. M. Szewczyk, X. Yang, K. Schmidt, X. Luo, S. Xiao, M. Dai, F. He, I. Zang, Y. Lin, F. Li, E. Dobrovetsky, D. Smil, S.-J. Min, J. Lin-Jones, M. Schapira, P. Atadja, E. Li, D. Barsyte-Lovejoy, C. H. Arrowsmith, P. J. Brown, F. Liu, Z. Yu, M. Vedadi and J. Jin, J. Med. Chem., 2018, 61, 1204–1217 CrossRef CAS PubMed.
- H. Firouzabadi, N. Iranpoor and F. Ebrahimzadeh, Tetrahedron Lett., 2006, 47, 1771–1775 CrossRef CAS.
- G.-P. Wu, W.-M. Ren, Y. Luo, B. Li, W.-Z. Zhang and X.-B. Lu, J. Am. Chem. Soc., 2012, 134, 5682–5688 CrossRef CAS PubMed.
- M.-X. Zhao, G.-D. Liang, J.-S. Wu, B.-J. Su, T.-M. Xiao, Y. Lu, S.-Y. Si, L.-J. Yuan and M.-H. Chen, Strepamrisides C and D: two new rhamnosides from Streptomyces amritsarensis13-85, J. Asian Nat. Prod. Res., 2026 DOI:10.1080/10286020.2026.2625948.
- E. Gao, Z.-Q. Zhou, J. Zou, Y. Yu, X.-L. Feng, G.-D. Chen, R.-R. He, X.-S. Yao and H. Gao, Bioactive Asarone-Derived Phenylpropanoids from the Rhizome of Acorus tatarinowii Schott, J. Nat. Prod., 2017, 80, 2923–2929 CrossRef CAS PubMed.
- K. Kocaman, M. Bingul, S. M. Ceyhan, H. Sahin, M. F. Saglam, H. Kandemir and I. F. Sengul, J. Mol. Struct., 2025, 1325, 141057 CrossRef CAS.
- E. Sevimli, G. Seyhan, D. Akkaya, S. Sarı, B. Barut and B. Köksoy, Bioorg. Chem., 2024, 147, 107366 CrossRef CAS PubMed.
- N. R. Sreenatha, T. N. M. Prasad, M. H. Kumar, A. S. J. Chakravarthy, S. N. Chandana, G. K. Puneetha, S. Nagaraju and B. N. Lakshminarayana, J.Mol. Struct., 2024, 1299, 137048 CrossRef CAS.
- F. Shojaie, Comput. Theor. Chem., 2017, 1114, 55–64 CrossRef CAS.
- E. A. Decoteau, A. Raghavan and C. L. Cahill, Inorg. Chem., 2024, 63, 2495–2504 CrossRef CAS.
- M. Ahmed, Sudhanshu, S. S. Malhotra, A. S. Alsubaie, S. M. El-Bahy, R. K. Mohapatra and A. Ansari, J. Mol. Model, 2025, 31, 75 CrossRef CAS PubMed.
Footnote |
| † W. Zhang and H. Yang contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |