 Open Access Article
Open Access ArticleDesign, synthesis, bioevaluation, and in silico insights of pyrazole-pyrrolinone and pyridazinone hybrids as potential antimicrobial agents
Sayed K. Ramadan *a,
Selwan Hamedb,
Hoda A. Ahmedc,
Tarfah Al-Warhid,
Nuha S. Alharbie and
Eman A. E. El-Helw
*a,
Selwan Hamedb,
Hoda A. Ahmedc,
Tarfah Al-Warhid,
Nuha S. Alharbie and
Eman A. E. El-Helw *a
*a
aChemistry Department, Faculty of Science, Ain Shams University, Cairo, 11566, Egypt. E-mail: sayed.karam2008@sci.asu.edu.eg; eman.abdelrahman@sci.asu.edu.eg
bDepartment of Microbiology and Immunology, Faculty of Pharmacy, Capital University, Ain Helwan, Helwan, 11795, Egypt
cDepartment of Chemistry, College of Science in Yanbu, Taibah University, Yanbu Governorate, Saudi Arabia
dDepartment of Chemistry, College of Science, Princess Nourah Bint Abdulrahman University, P. O. Box 84428, Riyadh 11671, Saudi Arabia
eDepartment of Chemistry, College of Science, Taibah University, Madinah, Saudi Arabia
First published on 14th May 2026
Abstract
The growing prevalence of antimicrobial resistance demands new molecular scaffolds capable of targeting planktonic pathogens and biofilm-associated infections. In this work, a series of new pyrazole-linked pyrrolinone and pyridazinone derivatives was designed and synthesized, starting from a pyrazolylfuran-2(3H)-one via selective ammonolysis and hydrazinolysis. The acid hydrazide obtained underwent heterocyclization, acylation, benzoylation, and condensation with aldehyde. The resulting compounds were evaluated for antimicrobial, antibiofilm, and mechanistic properties. Biological assessments included antimicrobial activity against five clinically relevant microbial strains, antibiofilm activity against Pseudomonas aeruginosa, cytotoxicity toward human fibroblasts, efflux pump inhibition studies, and molecular docking against Staphylococcus aureus PBP4 and Escherichia coli DNA gyrase B. Several derivatives showed pronounced strain-selective antimicrobial activity. Compounds 4, 6, 7, and particularly 10 showed strong antibacterial potency with MIC ranging from 1 to 8 µg mL−1, while compounds 6, 8, 9, and 10 showed notable activity against Candida albicans. Importantly, compounds 8–10 achieved near-complete eradication of P. aeruginosa biofilms (≥99.999%) and significantly inhibited biofilm formation (≥80%). Efflux inhibition assays displayed a 44–55% decrease in ethidium bromide efflux, supporting a multimodal mechanism of action. Docking studies revealed favorable binding affinities (−7.12 to −8.24 kcal mol−1), and cytotoxicity evaluation confirmed a favorable safety profile (IC50 120–200 µM). Among the evaluated compounds, pyrrolinone derivative 10 appeared as a promising antibiofilm and antimicrobial candidate for further preclinical development.
Introduction
Furan-2(3H)-ones attracted great attention due to their ease transformation into various heterocyclic systems of promising pharmacological efficacy. Our research team has broadly explored and discussed the chemistry and biological activity of these systems, highlighting their potential for developing new therapeutic agents.1–8 Acid hydrazides are versatile developing blocks for synthesizing heterocyclic frameworks of improved pharmacological properties.1,7 Pyrrolinones are a vital class of heterocyclic systems owing to their wide biological activity, synthetic utility, and presence in various natural products and pharmaceuticals. They are known for their roles as enzyme inhibitors, antimicrobial agents, anti-inflammatory drugs, and more recently, as intermediates in the synthesis of complex molecules. They serve as versatile pharmacophores in modern medicinal chemistry.Numerous pyrrolinone derivatives show potent biological activities, including antimicrobial, enzyme inhibition, anti-inflammatory, activities, use as scaffolds in drug design due to their capability to interact with multiple biological targets.9–11 Pyridazinones and pyrrolinones are vital creating blocks for diverse important drugs like naturally occurring antibacterial pyrrolinone (althiomycin), the cardiotonic pyridazinone (pimobendan and levosimendan), and analgesic anti-inflammatory (emorfozan).12–14 Fig. 1 shows some previously reported pyrrolinones I–VI with potent antimicrobial and anticancer properties in addition to some designed compounds showing potent activities.
 | ||
| Fig. 1 Pyrrolinone derivatives with antimicrobial and anticancer activity15–19 and the designed compounds of potent activity. | ||
For instance, pyrrolinone I had significant activity against S. aureus with MIC of 6.5 µg mL−1 and good activity against E. coli with MIC of 15 µg mL−1.15 Pyrrolinone II exhibited equipotent antibacterial activity against S. aureus, E. coli, and P. aeruginosa with ciprofloxacin (MIC: 6.25 µg mL−1).16 Also, pyrrolinone III showed MIC of 6.25 µg mL−1 against E. coli and P. aeruginosa and 12.5 µg mL−1 against S. aureus comparable to ciprofloxacin (MIC = 6.5 µg mL−1).17 Pyrrolinone IV had comparable activity to doxorubicin (reference anticancer drug) against HepG2 cancer cell line.18 Also, pyrrolinones V and VI showed strong cytotoxic activity and tubulin polymerase inhibition comparable to paclitaxel (tubulin inhibitor).19
Earlier investigations implied that pyrrolinone derivatives exert biological activity through multiple mechanisms, including enzyme inhibition (e.g., CDK2 and carbonic anhydrase inhibition) and disruption of cytoskeletal dynamics via tubulin polymerization inhibition, leading to cell cycle arrest and apoptosis.20,21 Additionally, their broad pharmacological profile suggests multi-target interactions, including interference with protein synthesis and other cellular pathways.11 Also, prior studies suggest that pyrrolinone derivatives exert their activity primarily through interaction with key cellular enzymes and redox-sensitive pathways. For example, several reports indicate that substituted pyrrolinones can function as enzyme inhibitors by binding to active or allosteric sites, thereby disrupting essential metabolic or signaling processes.22–24
In parallel, other studies propose that their conjugated structure enables redox cycling, leading to the generation of reactive oxygen species (ROS) and later oxidative stress-mediated damage in target cells.25 There is also evidence that some pyrrolinones can intercalate or associate biomolecular targets such as proteins or nucleic acids, further modulating cellular function.26 Together, these findings suggest a multifaceted mechanism in which pyrrolinones may combine enzyme inhibition with redox activity, ultimately impairing cellular homeostasis.
Pyrrolinones are not only important in medicinal chemistry but also show significant antimicrobial activity. Studies have shown that pyrrolinone derivatives can inhibit bacterial and fungal growth by targeting key enzymes and disrupting cellular processes, including membrane integrity and metabolic pathways. Their versatile reactivity allows further functionalization, enabling the development of diverse heterocyclic libraries with enhanced antimicrobial efficacy and broader spectrum activity. Thus, this work expanded the structural diversity for pyrazole-based furanone, pyridazinone, and pyrrolinones by introducing functional groups, exploring their potential as antimicrobial agents. Computational analyses including molecular docking and pharmacokinetic profiling revealed that these modifications remarkably enhance receptor binding affinity, surpassing conventional drugs in predicted stability and interaction strength.
Results and discussion
Synthesis and characterization
The key synthon, pyrazolyl-2(3H)-furanone 3 was prepared in 81% yield through Perkin condensation of 4-(4-methoxyphenyl)-4-oxobutanoic acid (1) and 3-(4-chlorophenyl)-1-phenylpyrazole-4-carbaldehyde (2)27 using cyclo-dehydrating agent including fused sodium acetate and ethanoic anhydride (Scheme 1). The lactone carbonyl absorption appeared at ν 1746 cm−1 in IR spectrum of furanone 3. Its 1H NMR spectrum (500 MHz, DMSO-d6) offered four singlet signals for methoxy, C4–H furanone, and C5–H pyrazole protons at δ 3.79, 7.43, and 9.14 ppm, in addition to signals corresponding to aromatic protons, referring to its existence as more thermodynamically stable E-isomer. Further, its 13C NMR spectrum (125 MHz, DMSO-d6) displayed signals at δ (ppm): 55.9 (OCH3), 100.1 (C4–furanone), 155.3 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 161.5 (
N), 161.5 (![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) –OCH3), and 169.3 (C
–OCH3), and 169.3 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), in addition to other signals for aromatic carbons. This furanone 3 suffers from the higher electron deficiency at lactonic carbonyl carbon atom, which makes it easier to be attacked by nitrogen nucleophiles.
O), in addition to other signals for aromatic carbons. This furanone 3 suffers from the higher electron deficiency at lactonic carbonyl carbon atom, which makes it easier to be attacked by nitrogen nucleophiles.
Thus, ammonolysis of furan-2(3H)-one 3 with ammonium acetate in refluxing ethyl alcohol afforded pyrrolinone 4 in 79% yield, through lactone ring opening by liberated ammonia followed by 5-exo-trig cyclization (cf. Scheme 1). The IR spectrum of pyrrolinone 4 furnished absorption bands for amide NH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups. A singlet signal for NH proton appeared in the downfield region in its 1H NMR spectrum. Compelling evidence for the assigned structure was acquired from its 13C NMR spectrum, which showed the characteristic signals. In turn, hydrazinolysis of furanone 3 upon stirring with hydrazine hydrate (80%) in ethanolic solution at room temperature (RT) afforded the acid hydrazide 5 in 83% yield, through lactone ring opening by hydrazine molecule (Scheme 1).
O groups. A singlet signal for NH proton appeared in the downfield region in its 1H NMR spectrum. Compelling evidence for the assigned structure was acquired from its 13C NMR spectrum, which showed the characteristic signals. In turn, hydrazinolysis of furanone 3 upon stirring with hydrazine hydrate (80%) in ethanolic solution at room temperature (RT) afforded the acid hydrazide 5 in 83% yield, through lactone ring opening by hydrazine molecule (Scheme 1).
The IR spectrum of hydrazide 5 lacked lactonic carbonyl absorption while ketone and hydrazide carbonyl absorption bands appeared, in addition to bands for NH and NH2 groups. Also, its 1H NMR spectrum showed singlet signals for methylene (CH2), NH2, and NH protons at δ 3.18, 4.32, and 6.54 ppm, respectively, in addition to singlet signals for protons of methoxy, olefinic, and C5–H pyrazole at δ 3.72, 7.13, and 8.67 ppm, respectively. Furthermore, its 13C NMR spectrum provided signals at δ 43.5 (CH2), 55.7 (OCH3), 151.8 (![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) –OCH3), 159.2 (C
–OCH3), 159.2 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O hydrazide), 166.5 (C
O hydrazide), 166.5 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), in addition to other signals for aromatic carbons.
O ketone), in addition to other signals for aromatic carbons.
The acid hydrazide functionality was used to build up two heterocyclic systems including pyridazinone and pyrrolinone. Indeed, refluxing the acid hydrazide 5 with equal mixture of hydrochloric and glacial ethanoic acids achieved 6-exo-trig cyclization through acid-catalyzed pathway to produce pyridazinone 6 in 64% yield (Scheme 2). Its IR spectrum showed NH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption bands at ν 3369 and 1656 cm−1, respectively. Its 1H NMR spectrum (400 MHz, DMSO-d6) provided a singlet signal for methylene protons at δ 3.94 ppm and another singlet signal for NH proton in the downfield region at δ 13.08 ppm, in addition to three singlet signals for protons of methoxy, C5–H pyridazine, and C5–H pyrazole at δ 3.76, 6.82, and 8.42 ppm, respectively. In turn, acylation of acid hydrazide 5 with ethanoic anhydride at RT or heating at 80–90 °C produced N-acetylaminopyrrolinone 7 in 84% yield, referring to the reactivity of hydrazide towards acylation (Scheme 2). Its IR spectrum revealed absorption bands for NH at ν 3279 cm−1 and carbonyl groups at ν 1724 and 1668 cm−1. Its 1H NMR (500 MHz, DMSO-d6) indicated its presence as a mixture of E- & Z-isomers in a ratio of 55
O absorption bands at ν 3369 and 1656 cm−1, respectively. Its 1H NMR spectrum (400 MHz, DMSO-d6) provided a singlet signal for methylene protons at δ 3.94 ppm and another singlet signal for NH proton in the downfield region at δ 13.08 ppm, in addition to three singlet signals for protons of methoxy, C5–H pyridazine, and C5–H pyrazole at δ 3.76, 6.82, and 8.42 ppm, respectively. In turn, acylation of acid hydrazide 5 with ethanoic anhydride at RT or heating at 80–90 °C produced N-acetylaminopyrrolinone 7 in 84% yield, referring to the reactivity of hydrazide towards acylation (Scheme 2). Its IR spectrum revealed absorption bands for NH at ν 3279 cm−1 and carbonyl groups at ν 1724 and 1668 cm−1. Its 1H NMR (500 MHz, DMSO-d6) indicated its presence as a mixture of E- & Z-isomers in a ratio of 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 45%. It offered two singlet signals for protons for each of acetyl-methyl at δ 1.78 & 1.84 ppm, methoxy at δ 3.72 & 3.79 ppm, C4–H pyrrolinone at δ 6.65 & 6.75 ppm, C5–H pyrazole at δ 8.73 & 9.17 ppm, and NH at δ 9.71 & 10.51 ppm. Furthermore, its 13C NMR spectrum (125 MHz, DMSO-d6) offered signals at δ (ppm) 21.0 (CH3), 55.7 (OCH3), 160.8 (
45%. It offered two singlet signals for protons for each of acetyl-methyl at δ 1.78 & 1.84 ppm, methoxy at δ 3.72 & 3.79 ppm, C4–H pyrrolinone at δ 6.65 & 6.75 ppm, C5–H pyrazole at δ 8.73 & 9.17 ppm, and NH at δ 9.71 & 10.51 ppm. Furthermore, its 13C NMR spectrum (125 MHz, DMSO-d6) offered signals at δ (ppm) 21.0 (CH3), 55.7 (OCH3), 160.8 (![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) –OCH3), 165.8 (CO pyrrolinone), and 169.2 (
–OCH3), 165.8 (CO pyrrolinone), and 169.2 (![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) OCH3).
OCH3).
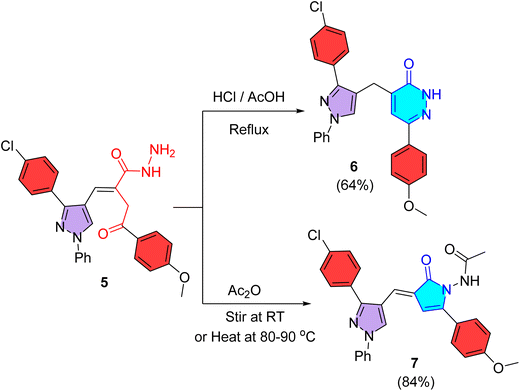 | ||
| Scheme 2 Synthesis of pyridazinone 6 (through refluxing with HCl/AcOH mixture) and pyrrolinone 7 (through stirring or heating with acetic anhydride). | ||
Also, N-benzoylaminopyrrolinone 8 was provided in 78% yield upon treating the acid hydrazide 5 with benzoyl chloride in 1,4-dioxane including triethylamine (Et3N) at an ambient temperature (Scheme 3). About its IR spectrum, an absorption band for NH appeared at ν 3218 cm−1 and carbonyl absorption bands were detected at ν 1696 and 1650 cm−1. Further, its 1H NMR spectrum (400 MHz, DMSO-d6) showed signals at δ (ppm): 3.73 (s, 3H, OCH3), 5.69 (s, 1H, C4–H pyrrolinone), 7.18 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), and 8.57 (s, 1H, C5–H pyrazole), while the expected NH proton signal was not detected, presumably due to rapid proton exchange with residual moisture in the solvent (DMSO). Analogously, treatment of acid hydrazide 5 with chloroacetyl chloride in 1,4-dioxane and Et3N on stirring at RT produced N-chloroacetylaminopyrrolinone 9 in 81% yield (Scheme 3). According to its IR spectrum, an absorption band for NH was seen at ν 3269 cm−1 and carbonyl absorption bands appeared at ν 1731 and 1682 cm−1. Its 1H NMR (400 MHz, DMSO-d6) presented signals at δ (ppm) 3.80 (s, 3H, OCH3), 4.20 (s, 2H, CH2), 6.83 (s, 1H, C4–H pyrrolinone), 7.11 (s, 1H, CH
), and 8.57 (s, 1H, C5–H pyrazole), while the expected NH proton signal was not detected, presumably due to rapid proton exchange with residual moisture in the solvent (DMSO). Analogously, treatment of acid hydrazide 5 with chloroacetyl chloride in 1,4-dioxane and Et3N on stirring at RT produced N-chloroacetylaminopyrrolinone 9 in 81% yield (Scheme 3). According to its IR spectrum, an absorption band for NH was seen at ν 3269 cm−1 and carbonyl absorption bands appeared at ν 1731 and 1682 cm−1. Its 1H NMR (400 MHz, DMSO-d6) presented signals at δ (ppm) 3.80 (s, 3H, OCH3), 4.20 (s, 2H, CH2), 6.83 (s, 1H, C4–H pyrrolinone), 7.11 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 9.21 (s, 1H, C5–H pyrazole), and 11.04 (br.s, 1H, NH), in addition to signals for aromatic protons. Furthermore, its 13C NMR (100 MHz, DMSO-d6) offered signals for carbons of methoxy (δ 55.8 ppm), methylene (δ 66.8 ppm), and carbonyl (δ 166.1 & 168.8 ppm) groups, besides aromatic carbons.
), 9.21 (s, 1H, C5–H pyrazole), and 11.04 (br.s, 1H, NH), in addition to signals for aromatic protons. Furthermore, its 13C NMR (100 MHz, DMSO-d6) offered signals for carbons of methoxy (δ 55.8 ppm), methylene (δ 66.8 ppm), and carbonyl (δ 166.1 & 168.8 ppm) groups, besides aromatic carbons.
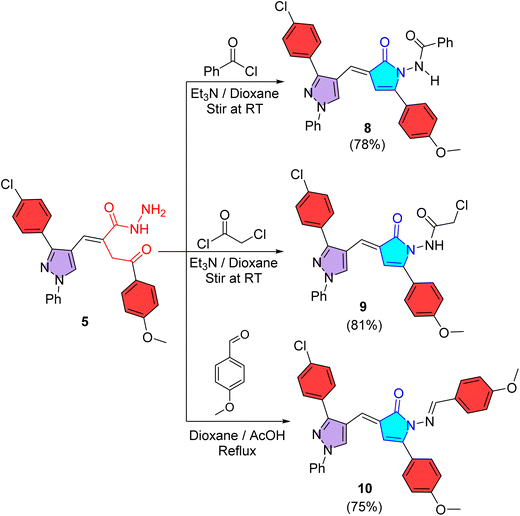 | ||
| Scheme 3 Synthesis of pyrrolinones 8–10, through condensation of acid hydrazide 5 with benzoyl chloride, chloroacetyl chloride, and 4-methoxybenzaldehyde. | ||
Condensation of the acid hydrazide 5 with 4-methoxybenzaldehyde in refluxing 1,4-dioxane and glacial ethanoic acid achieved the construction of pyrrolinone 10 in 75% yield (Scheme 3). Its IR spectrum lacked NH absorption and presented carbonyl and imino absorptions at ν 1683 and 1621 cm−1, respectively. Its 1H NMR spectrum offered two singlet signals for protons of two methoxy groups present at δ 3.80 and 3.82 ppm, in addition to four singlet signals for C4–H pyrrolinone, olefinic (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), C5–H pyrazole, and CH
), C5–H pyrazole, and CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N protons at δ 6.86, 7.13, 9.18, and 9.29, respectively.
N protons at δ 6.86, 7.13, 9.18, and 9.29, respectively.
Antimicrobial activity
Five microbial strains were employed to evaluate the antimicrobial activity of newly prepared compounds 3–10 with starting concentration of 1 mg mL−1 (1000 µg mL−1). Two Gram positive bacteria: Bacillus subtilis (B. subtilis) and Staphylococcus aureus (S. aureus), two Gram negative bacteria: Escherichia coli (E. coli) and Pseudomonas aeruginosa (P. aeruginosa), and one fungal strain: Candida albicans (C. albicans) were used. Antimicrobial potency was expressed as minimum inhibitory concentration (MIC) in µg mL−1 and compared with standard reference drugs, ofloxacin (OFX) for antibacterial activity and ketoconazole (Kn) for antifungal activity (cf. Table 1). Only MICs were determined in this study, and no subculturing assays were performed to show minimum bactericidal (MBC) or fungicidal (MFC) endpoints; this will be addressed in future work. The MICs of evaluated compounds were determined applying the standard broth dilution technique following CLSI guidelines.28| Compds | Microbial strains | ||||
|---|---|---|---|---|---|
| Gram negative | Gram positive | Fungi | |||
| E. coli | P. aeruginosa | S. aureus | B. subtilis | C. albicans | |
| a MIC concentration in µg mL−1. Positive and negative controls were DMSO and microbial growth, respectively. Ketoconazole (Kn) was used as a reference antifungal drug. Ofloxacin (OFX) was used as reference antibacterial drug.b Are the most promising compounds. | |||||
| 3 | 125 | 500 | 28 | 16 | 64 |
| 4b | 64 | 250 | 8 | 4 | 16 |
| 5 | 125 | 500 | 16 | 8 | 32 |
| 6a | 64 | 250 | 8 | 4 | 8 |
| 7b | 64 | 250 | 8 | 4 | 16 |
| 8 | 32 | 125 | 4 | 2 | 8 |
| 9 | 32 | 125 | 4 | 2 | 8 |
| 10b | 32 | 125 | 2 | 1 | 4 |
| OFX | 2 | 4 | 1 | 0.5 | — |
| Kn | — | — | — | — | 2 |
Overall, the tested compounds showed variable antimicrobial activity, with a pronounced selectivity toward Gram-positive bacterial and fungal strains judged to Gram-negative bacteria. This trend may be attributed to the structural differences of bacterial cell envelopes, where the outer membrane of Gram-negative bacteria acts as an added permeability barrier, limiting the intracellular access of antimicrobial agents. All tested compounds displayed moderate to weak activity against E. coli and P. aeruginosa, with MIC values ranging from 32 to 500 µg mL−1. Compounds 8, 9, and 10 showed the highest activity within this category, showing MIC of 32 µg mL−1 against E. coli and 125 µg mL−1 against P. aeruginosa. In contrast, compounds 3 and 5 displayed comparatively weaker activity, with MIC reaching up to 500 µg mL−1 against P. aeruginosa. Despite these findings, all compounds were significantly less potent than ofloxacin, which showed MIC values of 2–4 µg mL−1 against Gram-negative strains.
In contrast, the compounds showed enhanced activity against Gram-positive bacteria. MIC values against S. aureus and B. subtilis ranged from 1 to 28 µg mL−1, showing a substantial improvement compared with their Gram-negative activity. Compounds 4, 6, 7, 8, 9, and 10 exhibited notable antibacterial effects, with MIC values ≤8 µg mL−1 against both Gram-positive strains. Among these, compound 10 appeared as the most active antibacterial candidate, showing MIC values of 2 µg mL−1 against S. aureus and 1 µg mL−1 against B. subtilis, approaching the potency of reference drug, ofloxacin. The enhanced activity of these compounds suggests that specific structural features, particularly those present in the most promising compounds (marked with a in Table 1), may contribute to stronger interactions with Gram-positive bacterial targets, such as cell wall biosynthesis enzymes.
The antifungal evaluation against C. albicans revealed that most compounds have moderate to good antifungal activity. MIC ranged from 4 to 64 µg mL−1, with compounds 6, 8, 9, and 10 showing the most pronounced effects. Notably, pyrrolinone 10 exhibited MIC of 4 µg mL−1, showing strong antifungal potential, though it stayed less potent than ketoconazole (MIC = 2 µg mL−1). The observed antifungal activity suggests that these substrates may interfere with critical fungal cellular processes, potentially including ergosterol biosynthesis or membrane integrity. The improved activity of compounds bearing specific substituents supports SAR favoring antifungal efficacy.
A comparative analysis of MIC values implies that compounds labeled as most promising (4, 6, 7, and 10) consistently showed superior antimicrobial profiles across multiple strains. This observation highlights the importance of structural optimization, as minor chemical modifications markedly influenced biological activity. The increased potency of these compounds may be attributed to enhanced lipophilicity, improved membrane permeability, or stronger binding affinity to microbial targets.
Cytotoxicity testing
The cytotoxic potential of synthesized substrates was evaluated employing a human fibroblast cell line, a widely accepted in vitro model for assessing biocompatibility and preliminary safety of bioactive molecules, using MTT (3-[4,5-dimethylthiazole-2-yl]-2,5-diphenyltetrazolium bromide).29 Cytotoxicity was expressed as IC50 (µM) values, defined as the concentration needed to reduce cell viability by 50%, and the results are summarized in Table 2. All measurements are the mean of three independent experiments, ensuring reproducibility and reliability of the data obtained. Overall, the tested compounds (3–10) showed low cytotoxicity toward fibroblast cells, with IC50 values ranging from 120 to 200 µM. Such relatively high IC50 values show that substantial concentrations are needed to induce cytotoxic effects, suggesting a favorable safety profile for this compound series.| Compds | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|
| a IC50 values are expressed in µM as the concentration needed to inhibit 50% of cell viability. Data represent mean of three independent experiments. | ||||||||
| IC50 (µM) | 200 | 180 | 180 | 160 | 150 | 140 | 130 | 120 |
Compounds 3, 4, and 5 showed the highest IC50 values (200, 180, and 180 µM, respectively), showing minimal cytotoxic impact on normal cells. This observation supports their potential suitability for further biological evaluation, particularly in applications where preservation of healthy cell viability is critical. A gradual increase in cytotoxicity was seen across the series from compounds 6 to 10, as shown by decreasing IC50 values (160–120 µM). Pyrrolinone 10 displayed the lowest IC50 value (120 µM), reflecting comparatively higher cytotoxicity among the tested molecules. However, even this value stays within a range generally considered acceptable for early-stage drug candidates, particularly when balanced against potential therapeutic efficacy. Importantly, none of the compounds showed pronounced toxicity at low micromolar concentrations, which minimizes concerns about nonspecific cellular damage.
The observed trend in cytotoxicity may be attributed to structural variations among the compounds, which could influence cellular uptake, membrane interactions, or intracellular target engagement. Such structure-cytotoxicity relationships warrant further investigation to improve the balance between biological activity and safety. Notably, the absence of sharp cytotoxic effects across the tested concentration range suggests that the molecular scaffold is inherently biocompatible, providing a promising foundation for further chemical modification. From a safety perspective, the high IC50 values against normal fibroblast cells show a wide preliminary therapeutic window, particularly when compared with typical effective concentrations reported for many bioactive compounds. This selectivity toward non-toxic behavior in normal cells is a crucial prerequisite for advancing compounds into more extensive pharmacological and in vivo evaluations.
In conclusion, the cytotoxicity data showed that all tested compounds have acceptable in vitro safety profiles, with low toxicity toward human fibroblast cells. These findings support the continued investigation of this compound series as potential bioactive agents, while future studies should incorporate additional normal cell lines and mechanistic toxicity assays to further substantiate their safety and therapeutic relevance.
Biofilm inhibition/eradication
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log reductions, corresponding to 95.8% to 99.85% reduction after 4 h. They are strong but not sterilizing.
log reductions, corresponding to 95.8% to 99.85% reduction after 4 h. They are strong but not sterilizing.
| Compds | Treatmentc | ||||
|---|---|---|---|---|---|
| 2 h | Log10 reduction | 4 h | Log10 reduction | % Reduction after 4 h | |
| a Values expressed after 2 and 4 h for log10 CFU per well, and represent mean ± SD from triplicate experiments. CFU: colony-forming units (living bacteria).b % Reduction is versus PBS.c 0.5 MIC of each compound was used. “<2.00” shows count below the detection limit; ≥99.999% denotes near-complete eradication. | |||||
| PBSb | 7.45 ± 0.10 | — | 7.42 ± 0.08 | — | — |
| 3 | 6.40 ± 0.12 | 1.05 | 6.00 ± 0.11 | 1.42 | 95.8% |
| 4 | 5.60 ± 0.10 | 1.85 | 4.60 ± 0.08 | 2.82 | 99.85% |
| 5 | 6.20 ± 0.09 | 1.25 | 5.75 ± 0.08 | 1.67 | 97.9% |
| 6 | 5.80 ± 0.10 | 1.65 | 4.95 ± 0.09 | 2.47 | 99.7% |
| 7 | 5.95 ± 0.11 | 1.50 | 5.10 ± 0.07 | 2.32 | 99.5% |
| 8 | 4.90 ± 0.12 | 2.55 | <2.00 | ≥5.42 | ≥99.999% |
| 9 | 4.75 ± 0.10 | 2.70 | <2.00 | ≥5.42 | ≥99.999% |
| 10 | 4.50 ± 0.09 | 2.95 | <2.00 | ≥5.42 | ≥99.999% |
For example, compound 4 showed 4 h CFU = 4.60 and Log reduction = 2.82, that showed reduction from 107.42 to ∼104.6 (≈2500× fewer bacteria) and ≈99.85% killing. These results suggest these compounds significantly damage the biofilm but do not eradicate it completely. On the other hand, compounds 8, 9, and 10 had exceptional, near-total biofilm eradication, and these three compounds were the most potent. Their 4 h results were Log CFU < 2.00 (below detection limit and the biofilm was essentially wiped out), Log reduction ≥ 5.42, and % reduction ≥ 99.999%. That means that 5.42![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log reduction equals ∼260
log reduction equals ∼260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000× fewer bacteria, the remaining bacteria are too few to count, and this is considered near-complete biofilm eradication. Compounds 8, 9, and 10 outperform all other compounds by a wide margin. Thus, these compounds are excellent leads for further development, likely having strong biofilm penetration or strong bactericidal activity at sub-MIC levels.
000× fewer bacteria, the remaining bacteria are too few to count, and this is considered near-complete biofilm eradication. Compounds 8, 9, and 10 outperform all other compounds by a wide margin. Thus, these compounds are excellent leads for further development, likely having strong biofilm penetration or strong bactericidal activity at sub-MIC levels.
At 0.5 MIC, compounds 6, 7, 8, 9, and 10 displayed marked reductions in OD values (0.40–0.25), corresponding to 65–80% inhibition of biofilm biomass (cf. Table 4). Compounds 3 and 5 exhibited moderate inhibition (OD ≈ 0.65–0.55; ∼50% reduction). Markedly, compounds 8–10 were the most effective, achieving >75% inhibition and partial eradication of established biofilms. For eradication assays, treatment of pre-formed biofilms for 4 h with 0.7–0.8 MIC concentrations of 8–10 nearly eliminated biofilm biomass (OD570 ≤ 0.15; >85% eradication).31
| Compds | Biofilm inhibition | % Inhibition vs. PBSb | Biofilm eradication | % Eradication vs. PBS |
|---|---|---|---|---|
| a 0.5 MIC of each compound was used, and the data are expressed as mean ± SD (OD570, mean ± SD).b % Reduction is versus PBS (control). | ||||
| PBS | 1.20 ± 0.05 | — | 1.18 ± 0.06 | — |
| 3 | 0.65 ± 0.04 | 45.8% | 0.55 ± 0.05 | 53.4% |
| 4 | 0.53 ± 0.04 | 55.8% | 0.52 ± 0.04 | 55.9% |
| 5 | 0.55 ± 0.03 | 54.2% | 0.48 ± 0.04 | 59.3% |
| 6 | 0.35 ± 0.03 | 70.8% | 0.28 ± 0.02 | 76.3% |
| 7 | 0.40 ± 0.02 | 66.7% | 0.32 ± 0.03 | 72.9% |
| 8 | 0.28 ± 0.02 | 76.7% | 0.18 ± 0.01 | 84.7% |
| 9 | 0.25 ± 0.02 | 79.2% | 0.15 ± 0.01 | 87.3% |
| 10 | 0.26 ± 0.02 | 78.3% | 0.16 ± 0.01 | 86.4% |
Potential antifungal mechanism
| Compds | MIC without sorbitol | MIC with 0.8 M sorbitol | Fold change |
|---|---|---|---|
| a Concentration was in µg mL−1 and all experiments were performed in triplicate. | |||
| 3 | 16 | 64 | 4× |
| 4 | 8 | 16 | 2× |
| 5 | 32 | 128 | 4× |
| 8 | 16 | 32 | 2× |
Some tested compounds offered moderate to strong efflux pump inhibitory activity. Compounds 3–5 produced relatively modest reductions in k (17–24% inhibition), while compounds 6–10 produced more pronounced effects (cf. Table 6). Compounds 8 (51.7% inhibition), 9 (44.8% inhibition), and 10 (55.2% inhibition) significantly decreased the efflux rate and yielded elevated residual fluorescence levels (43–50%), indicative of substantial intracellular retention of EtBr. Generally, the data support a structure-dependent variation in efflux inhibitory potency, with compounds 8–10 appearing as the most promising EPIs within the tested series.
| Compds | k (mean ± SD) | % Inhibition vs. DMSO | Residual RFU at 15 min |
|---|---|---|---|
| a 0.25 MIC value was used.b CCCP (carbonyl cyanide m-chlorophenylhydrazone, 75 µM) was used as positive control for efflux pump inhibition. | |||
| DMSO | 0.145 ± 0.012 | — | 18% ± 2 |
| bCCCP | 0.040 ± 0.006 | 72.4% | 62% ± 4 |
| 3 | 0.120 ± 0.010 | 17.2% | 26% ± 3 |
| 4 | 0.113 ± 0.009 | 24.3% | 31% ± 2 |
| 5 | 0.110 ± 0.009 | 24.1% | 30% ± 2 |
| 6 | 0.085 ± 0.007 | 41.4% | 41% ± 3 |
| 7 | 0.090 ± 0.008 | 37.9% | 38% ± 3 |
| 8 | 0.070 ± 0.006 | 51.7% | 48% ± 3 |
| 9 | 0.080 ± 0.007 | 44.8% | 43% ± 3 |
| 10 | 0.065 ± 0.005 | 55.2% | 50% ± 4 |
In summary, this work addressed several critical gaps in the field of antimicrobial drug discovery. There is an obvious lack of new heterocyclic scaffolds with dual antimicrobial and antibiofilm activity, as most existing antibiotics show limited efficacy against mature biofilms and are typically active against either bacteria or fungi, but not both. Notably, pyrazole-pyrrolinone scaffolds have not been previously explored for broad-spectrum antimicrobial and antibiofilm applications. Additionally, there is limited research on compounds that combine antimicrobial potency with efflux pump inhibition, despite efflux-mediated resistance being a major challenge, and few small molecules reported to date can simultaneously inhibit efflux while keeping strong antimicrobial activity.
Another key limitation in current literature is the insufficient mechanistic understanding of new heterocycles acting on both bacterial and fungal systems; prior studies rarely investigate multimodal actions, such as targeting bacterial enzymes (e.g., PBP4 and GyrB), disrupting fungal cell-wall integrity, inhibiting efflux pumps, and affecting biofilms within a single chemical scaffold. Furthermore, there is a scarcity of lead compounds that combine high potency with low cytotoxicity, as many candidates fail due to toxicity or poor drug-likeness, emphasizing the need for molecules with balanced efficacy, safety, and physicochemical properties. Finally, there is a notable lack of systematic SAR studies for pyrazole-pyrrolinone analogues, leaving an important knowledge gap about how structural modifications influence antimicrobial performance.
To address these gaps, this work makes several important contributions. It reports the synthesis of new library of pyrazole-pyrrolinone derivatives, introducing previously unreported structures and expanding the chemical diversity of heterocyclic antimicrobial agents. The study also finds a potent lead compound (compound 10) that shows broad-spectrum antibacterial, antifungal, and antibiofilm activity, with MICs of 1–8 µg mL−1, strong antifungal efficacy, and biofilm eradication of at least 99.999%, standing for a rare and noteworthy combination of properties within this scaffold class.
In addition, the research provides evidence for multimodal mechanisms of action, demonstrating efflux pump inhibition (44–55%), biofilm disruption, molecular docking interactions with PBP4 and DNA gyrase B, and interference with fungal cell-wall integrity, thereby highlighting the scaffold's therapeutic potential and reducing the likelihood of resistance development. The study further offers emerging SAR trends for pyrazole-pyrrolinone hybrids by systematically analyzing structural analogues, offering the first insights into how substituent patterns influence antimicrobial and antibiofilm activity. Importantly, the compounds demonstrate low cytotoxicity alongside high antimicrobial potency, with fibroblast IC50 values ranging from 120 to 200 µM, showing a favorable safety profile. Finally, the integration of ADME-relevant properties, including SwissADME analysis, confirms good drug-likeness, acceptable lipophilicity, and suitable physicochemical characteristics, parameters that are essential yet often overlooked in early-stage antimicrobial research.
About the rationale for target choice, both PBP4 and DNA gyrase B are well-established and clinically relevant antibacterial targets. PBP4 plays a critical role in bacterial cell wall biosynthesis and contributes to β-lactam resistance in S. aureus through enhanced peptidoglycan cross-linking, making it a valuable target for overcoming resistance mechanisms. This functional role makes PBP4 an important secondary resistance determinant, especially in strains displaying reduced susceptibility to methicillin and other β-lactams. Moreover, the 5TW8 structure provides a high-resolution model of the PBP4 active site, enabling accurate evaluation of ligand interactions. Its well-defined catalytic domain makes it a reliable template for structure-based drug design and docking studies aimed at discovering new inhibitors capable of overcoming β-lactam resistance mechanisms. Similarly, DNA gyrase B is an essential enzyme involved in DNA replication and supercoiling in E. coli, and its inhibition is a validated strategy in antibacterial drug design. Thus, these targets were selected not only for their general antimicrobial relevance but also because of their essential biological functions, established druggability, and availability of high-resolution crystal structures suitable for reliable molecular docking analysis.
The S-score (docking score, kcal mol−1) represents an estimate of binding free energy between ligand and target protein, where more negative values show stronger predicted binding affinity. This score is not solely dependent on the number of interactions (e.g., hydrogen bonds), but rather reflects a combination of energetic contributions, including van der Waals (VDW) interactions, electrostatic forces, hydrophobic effects, and, to some extent, ligand conformational stability within binding pocket.
The S-score, RMSD values (Å), and amino–acid interaction patterns were benchmarked against the respective co-crystallized ligands (AI8 and E0F) as well as reference antibiotics: Linezolid for 5TW8 and Gentamicin for 6KZV (cf. Table 7). These antibiotic agents are active against most Gram-positive and Gram-negative bacteria, respectively. The protein structure was parted from inhibitor particle and adjusted by molecular minimization with added H-atoms. All prepared complexes were docked in the same place of binding site of native co-crystallized ligands.
| Compds | S-score (kcal mol−1) | RMSD (Å) | aBinding amino acids (bond type, length ‘Å’) | |||
|---|---|---|---|---|---|---|
| 5TW8 | 6KZV | 5TW8 | 6KZV | 5TW8 | 6KZV | |
| a Common amino acid between compounds with co-crystallized ligands and reference agents were italicized.b Reference agent: linezolid for 5TW8; gentamicin for 6KZV. | ||||||
| 4 | −7.2545 | −7.3414 | 1.7053 | 1.8591 | ASN 141 (H-acceptor, 3.19) | ILE 78 (pi–H, 4.55) |
| SER 116 (pi–H, 4.56) | ||||||
| 6 | −7.2781 | −7.7133 | 1.7217 | 1.0368 | SER 116 (H-acceptor, 3.24) | ASN 46 (pi–H, 3.85) |
| ILE 94 (pi–H, 4.29) | ||||||
| 7 | −7.1188 | −8.0359 | 1.5469 | 1.8005 | SER 262 (H-donor, 3.05) | ALA 100 (H-donor, 2.96) |
| SER 116 (H-acceptor, 3.08) | THR 165 (pi–H, 4.50) | |||||
| ASN 141 (H-acceptor, 2.91) | ||||||
| 10 | −7.8841 | −8.2369 | 1.6917 | 1.7899 | SER 116 (H-acceptor, 3.06) | ASN 46 (pi–H, 3.95) |
| TYR 239 (pi–H, 4.32) | ||||||
| Co-crystallized ligand | −8.0713 | −8.3347 | 1.4939 | 1.1922 | SER 139 (H-donor, 3.08) | ASP 73 (H-donor, 2.75) |
| SER 262 (H-donor, 3.02) | ||||||
| SER 262 (H-donor, 3.11) | ||||||
| GLY 181 (H-donor, 2.80) | GLU 50 (H-donor, 2.76) | |||||
| SER 75 (H-acceptor, 2.80) | ||||||
| SER 262 (H-acceptor, 2.85) | ||||||
| SER 139 (H-acceptor, 2.49) | ASP 49 (H-donor, 3.08) | |||||
| LYS 259 (H-acceptor, 2.64) | ||||||
| THR 260 (H-acceptor, 2.52) | ||||||
| LYS 259 (ionic, 2.64) | THR 165 (H-acceptor, 2.81) | |||||
| LYS 259 (ionic, 3.46) | ||||||
| GLU 297 (ionic, 3.27) | ||||||
| ALA 182 (pi–H, 4.11) | ASP 49 (ionic, 3.58) | |||||
| PHE 243 (pi–H, 4.18) | ||||||
| PHE 241 (pi–pi, 3.83) | ||||||
| bReference agent | −7.0120 | −7.3800 | 1.0899 | 1.2303 | SER 262 (H-donor, 3.04) | ASP 49 (H-donor, 3.08) |
| ALA 100 (H-donor, 2.73) | ||||||
| SER 116 (H-acceptor, 3.15) | ASP 45 (H-donor, 2.85) | |||||
| GLU 42 (H-donor, 2.90) | ||||||
Also, Table S1 showed graphical 2D and 3D descriptions of docking simulation. The defined information on the molecular interactions between ligands and CDK2 protein was displayed in a 2D illustration. Also, the binding interactions between ligands and active pockets of protein are shown in red by 3D depiction, which imply H-bond interactions. The studied molecules displayed diverse H-bonding, ionic, and π–H interactions with key amino acids within the enzymatic binding pockets. Several interactions were conserved compared to co-crystallized ligands, supporting binding mode validity.
It was seen that the co-crystallized ligands showed significantly more contacts, particularly H-bonds, compared to the tested compounds, yet yielded comparable S-scores. This might be attributed to the fact that the binding affinity is not determined exclusively by the number of H-bonds. In several cases, the docked compounds achieve favorable binding through hydrophobic interactions, π–π stacking, and VDW contacts within key regions of the active site. These interactions can compensate energetically for fewer hydrogen bonds. Additionally, the co-crystallized ligands are often larger and more complex, enabling multiple polar contacts but also potentially incurring higher desolvation or conformational penalties. In contrast, the studied compounds, although forming fewer hydrogen bonds, may adopt more optimal binding conformations with reduced steric strain and better complementarity to hydrophobic pockets, resulting in comparable overall docking scores.
Thus, the potent compounds showed favorable binding free energies (S-scores ranged from −7.1188 to −8.2369 kcal mol−1) against both protein targets, suggesting thermodynamically favorable complex formation.37,38 Against 5TW8, pyrrolinone 10 displayed the most favorable affinity (−7.8841 kcal mol−1), approaching co-crystallized ligand (−8.0713 kcal mol−1). Against 6KZV, pyrrolinone 10 again showed the highest affinity (−8.2369 kcal mol−1), and compound 7 also performed strongly (−8.0359 kcal mol−1), both comparable to the co-crystallized ligand (−8.3347 kcal mol−1). These results infer that compounds 7 and 10 interact efficiently with the active sites of both Gram-positive and Gram-negative targets.
A recurrent interaction among potent compounds was with SER 116, which functioned as either hydrogen-bond acceptor or donor. Compounds 4, 6, 7, and 10 also formed stabilizing interactions with residues such as ASN 141, SER 262, and TYR 239 depending on the ligand geometry. The co-crystallized ligand displayed an extensive network of hydrogen bonds and ionic interactions involving SER 139, SER 262, GLY 181, LYS 259, and GLU 297, which explains its superior binding energy.
In 6KZV, key residues involved in interactions with the active compounds included ASN 46, ILE 94, ALA 100, and THR 165, frequently through π–H interactions or H-bonds. The co-crystallized ligand interacted strongly through multiple donors and acceptors such as ASP 73, GLU 50, and ASP 49, reflecting the tight binding required for ribosomal inhibition. The reference antibiotics (linezolid for 5TW8 and gentamicin for 6KZV) exhibited characteristic interactions with SER 262, SER 116 (Linezolid) and ASP 49, ALA 100, GLU 42 (Gentamicin). Notably, some compounds shared identical conserved residues (italicized in the dataset), showing that they may exploit similar mechanisms of inhibition.
Overall structure–activity interpretation
Compounds 7 and 10 showed the best performance across both targets, showing high binding affinities comparable to or approaching co-crystallized ligands (AI8 and E0F), low RMSD values showing stable conformations, and interactions with amino acids known to be critical for ligand recognition. Such dual-target inhibitory potential suggests promising broad-spectrum antibacterial activity.All compounds demonstrated RMSD between 1.03 and 1.86 Å, showing stable and reliable docking conformations with acceptable deviation from the optimal pose. The RMSD values for all compounds were comparable to those of reference ligands, confirming consistency in predicted binding modes. The verification of docking program's protocol was performed by contrasting the co-crystallized ligands (AI8 and E0F) with their related protein targets (PDB ID: 5TW8 and 6KZV).37 This visualized superimposition of native co-crystallized ligand (depicted in green) with the redocked co-crystallized ligand (represented in red) through 2D diagrams. The RMSD for these superimpositions, which show the difference between the two structures, were within the favorite ranges (1.4939 and 1.1922 Å, respectively), as shown in Fig. S1.
Modeling pharmacokinetics
The ADME (absorption, distribution, metabolism, and excretion) profiles of new substrates were analyzed featuring physicochemical, lipophilicity, pharmacokinetics, and drug-likeness properties applying SwissADME free web tool39–41 (cf. Fig. S2–S9). Table 8 summarizes key pharmacokinetic descriptors, TPSA (topological polar surface area, Å2), rotatable bonds, lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), hydrogen-bonding features (HBA/HBD), GI absorption, bioavailability score, and efflux/BBB permeability, for the synthesized substrates. These properties collectively determine how well a compound can permeate biological membranes and interact with microbial targets. The violations of the synthesized compounds towards pharmacokinetics rules and filters were shown in Table S2. With the PAINS filter (label compounds that assign false-positive results in biological assays), the acid hydrazide 5 and pyridazinone 6 passed (as no alerts were shown) in contrast to other compounds that showed one alert. This means that these substrates may perform well in further biological or clinical testing. Considering Brenk filters (flag structural features that are correlated with toxicity, instability, or bad pharmacokinetics), pyridazinone 6 passed (no alerts) but compounds 3, 4, 7, and 8 referred to one alert, while compounds 9 and 10 displayed two alerts and compound 5 showed three alerts. Inhibition of the newly synthesized compounds against CYP isoenzymes was described in Table S3, which assumed all compounds as non-inhibitors of CYP1A2 and CYP2D6 isoenzymes.
P), hydrogen-bonding features (HBA/HBD), GI absorption, bioavailability score, and efflux/BBB permeability, for the synthesized substrates. These properties collectively determine how well a compound can permeate biological membranes and interact with microbial targets. The violations of the synthesized compounds towards pharmacokinetics rules and filters were shown in Table S2. With the PAINS filter (label compounds that assign false-positive results in biological assays), the acid hydrazide 5 and pyridazinone 6 passed (as no alerts were shown) in contrast to other compounds that showed one alert. This means that these substrates may perform well in further biological or clinical testing. Considering Brenk filters (flag structural features that are correlated with toxicity, instability, or bad pharmacokinetics), pyridazinone 6 passed (no alerts) but compounds 3, 4, 7, and 8 referred to one alert, while compounds 9 and 10 displayed two alerts and compound 5 showed three alerts. Inhibition of the newly synthesized compounds against CYP isoenzymes was described in Table S3, which assumed all compounds as non-inhibitors of CYP1A2 and CYP2D6 isoenzymes.
| Compds | Rotatable bonds | TPSA (Å2) | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp (cm s−1) Kp (cm s−1) |
HBA | HBD | Consensus log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
GI absorption | Bioavailability score | BBB permeant | Pgp substrate |
|---|---|---|---|---|---|---|---|---|---|---|
a TPSA: topological polar surface area, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp: skin permeation, HBA: hydrogen bond acceptors, HBD: hydrogen bond donors. Kp: skin permeation, HBA: hydrogen bond acceptors, HBD: hydrogen bond donors. |
||||||||||
| 3 | 5 | 53.35 | −4.67 | 4 | 0 | 5.38 | High | 0.55 | No | No |
| 4 | 5 | 56.15 | −5.06 | 3 | 1 | 4.89 | High | 0.55 | Yes | No |
| 5 | 9 | 99.24 | −6.15 | 5 | 2 | 3.93 | High | 0.55 | No | No |
| 6 | 6 | 72.8 | −5.31 | 4 | 1 | 5.02 | High | 0.55 | No | Yes |
| 7 | 7 | 76.46 | −5.63 | 4 | 1 | 4.57 | High | 0.55 | No | No |
| 8 | 8 | 76.46 | −4.84 | 4 | 1 | 5.66 | High | 0.17 | No | No |
| 9 | 8 | 76.46 | −5.40 | 4 | 1 | 4.86 | High | 0.55 | No | No |
| 10 | 8 | 68.95 | −4.61 | 5 | 0 | 6.33 | Low | 0.17 | No | No |
Noteworthy, the most relevant parameters were well described as follows: fewer rotatable bonds often enhance membrane permeability and binding affinity due to improved conformational rigidity. Compounds 4 (5 RBs) and 6 (6 RBs) fall within the optimal range for good oral bioavailability (≤10). Compounds 7 (7 RBs) and 10 (8 RBs) also remain acceptable, supporting good biological activity. TPSA influences passive membrane diffusion. All four active compounds have TPSA values below 90 Å2, which supports good cell penetration: 4 (56.15 Å2), 6 (72.8 Å2), 7 (76.46 Å2), and 10 (68.95 Å2). These values lie well within the range associated with effective antimicrobial penetration.
Lipophilicity is a key determinant of the ability to permeate microbial lipid membranes. Active compounds have moderately high log P values: 4 (4.89), 6 (5.02), 7 (4.57), and 10 (6.33). This suggests stronger membrane interaction, a known mechanism contributing to antimicrobial potency. Balanced hydrogen-bonding capacity improves both permeability and target binding. Active compounds show: HBD: 0–1 and HBA: 3–5. These values align with optimal antimicrobial pharmacophores, sufficient polarity for target binding but not enough to impair permeability.
All active compounds, except pyrrolinone 10, show high GI absorption, supporting good pharmacokinetic behavior. Pyrrolinone 10 shows low absorption, but its strong activity may stem from higher lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 6.33) and favorable structural features. Only pyridazinone 6 is predicted to be a P-gp substrate, but this does not negatively affect antimicrobial action. None of these compounds penetrate the BBB, which is expected and not needed for antimicrobial therapy. The strong antimicrobial activities of compounds 4, 6, 7, and 10 can be rationalized based on the ADME trends in Table 8.
P 6.33) and favorable structural features. Only pyridazinone 6 is predicted to be a P-gp substrate, but this does not negatively affect antimicrobial action. None of these compounds penetrate the BBB, which is expected and not needed for antimicrobial therapy. The strong antimicrobial activities of compounds 4, 6, 7, and 10 can be rationalized based on the ADME trends in Table 8.
All four active compounds have TPSA between 56–76 Å2, enabling efficient cell membrane penetration, which is essential for reaching intracellular microbial targets. Their lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ∼4.5–6.3) enhances interaction with microbial membranes, contributing to disruption of membrane integrity or improved intracellular uptake. This balance explains why these compounds outperform others with either: too high TPSA (e.g., acid hydrazide 5: 99.24 Å2) or less favorable lipophilicity (e.g., compound 5: log
P ∼4.5–6.3) enhances interaction with microbial membranes, contributing to disruption of membrane integrity or improved intracellular uptake. This balance explains why these compounds outperform others with either: too high TPSA (e.g., acid hydrazide 5: 99.24 Å2) or less favorable lipophilicity (e.g., compound 5: log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 3.93).
P 3.93).
The number of rotatable bonds (5–8) in the active compounds allows them to adopt optimal conformations for binding microbial enzymes, while keeping membrane permeability. Excessive flexibility (e.g., compounds 8 and 9 with eight RBs but lower TPSA/lipophilicity synergy) may reduce binding specificity, which may explain their lower activity. Compounds 4, 6, 7, and 10 have moderate HBA (3–5) and low HBD (0–1). This supports both strong interactions with target residues and reduced polarity, which improves membrane permeation. Inactive or weaker compounds show higher polar burden (e.g., compound 5 with two HBD and five HBA), which may hinder permeation. Even though pyrrolinone 10 has low GI absorption, its strong antimicrobial activity can be explained by its high lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 6.33) and favorable TPSA (68.95 Å2). This combination supports: better membrane disruption potential and strong hydrophobic interactions with biological targets.
P 6.33) and favorable TPSA (68.95 Å2). This combination supports: better membrane disruption potential and strong hydrophobic interactions with biological targets.
The plasma protein-binding model predicts the likelihood of a substrate being extensively bound to blood carrier proteins. The newly synthesized compounds showed a high probability of plasma protein binding. These findings were further interpreted using the BOILED-Egg simulation (Fig. 2), which graphically illustrates gastrointestinal absorption and brain penetration. Compounds 3 and 5–9, found in the BOILED-Egg white region, are predicted to undergo passive absorption through the gastrointestinal tract. Those highlighted in red are expected to evade P-glycoprotein-mediated efflux from the central nervous system (CNS). In turn, pyrrolinone 4, which existed inside the yellow chart area, is expected to penetrate the blood–brain barrier. Pyridazinone 6, shown by blue, may be possible candidate for the permeability glycoprotein (PGP).
This matches typical high-activity patterns of thiophene derivative and hydrophobic heterocycles. The excellent antimicrobial activities of compounds 4, 6, 7, and 10 correlate strongly with their favorable ADME properties: TPSA within optimal membrane-penetrating range, high lipophilicity enabling strong cell–membrane interaction, balanced H-bonding profiles, and good molecular flexibility (rotatable bonds). These physicochemical features collectively enhance both target binding and cell permeability, providing a clear explanation for their superior antimicrobial performance.
SAR study
The SAR analysis reveals a clear relationship between physicochemical properties, substituent effects, and biological activity within this scaffold class. The most active compounds fall within a best polarity-lipophilicity window, clustering at c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Log
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values of approximately 4.5–6.3 and TPSA below 90 Å2. This balance appears critical for achieving both efficient membrane penetration and effective target engagement, as demonstrated by compounds 4, 6, 7, and 10, which exhibit strong antibacterial activity (MIC 1–8 µg mL−1), as well as compounds 8–10, which display exceptional antibiofilm efficacy with ≥99.999% eradication. Electronic effects also play a significant role, where moderately electron-donating groups such as para-methoxy enhance antifungal activity, particularly against C. albicans, likely by improving hydrogen bonding and dipole interactions while also helping permeability.
P values of approximately 4.5–6.3 and TPSA below 90 Å2. This balance appears critical for achieving both efficient membrane penetration and effective target engagement, as demonstrated by compounds 4, 6, 7, and 10, which exhibit strong antibacterial activity (MIC 1–8 µg mL−1), as well as compounds 8–10, which display exceptional antibiofilm efficacy with ≥99.999% eradication. Electronic effects also play a significant role, where moderately electron-donating groups such as para-methoxy enhance antifungal activity, particularly against C. albicans, likely by improving hydrogen bonding and dipole interactions while also helping permeability.
In contrast, electron-withdrawing halogens such as para-chloro tend to improve antibacterial potency by increasing lipophilicity and enabling favorable hydrophobic or halogen-bonding interactions within PBP4 and DNA gyrase B binding pockets. Positional effects further highlight the importance of para-substitution, which consistently provides superior activity compared to ortho (due to steric hindrance) and meta positions (due to suboptimal spatial orientation). Additionally, the incorporation of cationic or reactive acyl functionalities, such as chloroacetyl and benzyl/aryl acyl moieties, as seen in compounds 8–10, strongly correlates with enhanced biofilm eradication and efflux pump inhibition (44–55%), suggesting improved penetration of the extracellular polymeric matrix and potential interaction with efflux machinery.
A more detailed SAR analysis by structural regions further clarifies these trends. Substituents on the pyrazole ring significantly influence activity, where para-positioned electron-donating groups enhance antifungal potency, consistent with observed effects on fungal cell-wall pathways, as shown by sorbitol protection assays. Halogen substituents, particularly para-chloro groups, favor antibacterial activity through improved hydrophobic fit and possible halogen bonding interactions within enzyme active sites. Excessively polar substituents, especially when TPSA exceeds ∼90 Å2, tend to reduce antibacterial efficacy due to diminished membrane permeability unless compensated by strong binding interactions.
About the core structure, pyrrolinone derivatives, exemplified by compound 10, demonstrate superior overall activity compared to pyridazinone analogues. This advantage is likely due to their more planar and compact conformation, which helps better accommodation within enzyme binding pockets, while the lactam carbonyl serves as a key hydrogen bond acceptor supporting dual-target interactions. In contrast, pyridazinones contribute chemical diversity and may enhance antifungal activity in certain cases but generally show weaker antibacterial performance.
Substitutions at the acyl, hydrazide, and carbamoyl positions further define activity profiles. Conversion of acid hydrazides to acylated derivatives, particularly benzoyl and chloroacetyl groups, significantly enhances activity. Benzoyl moieties improve hydrophobic and π–π interactions, leading to better binding affinity and improved Gram-positive antibacterial activity, as observed in compounds 4, 6, 7, and 10. Chloroacetyl groups are strongly associated with superior antibiofilm activity and efflux inhibition, likely due to their ability to enhance penetration, disrupt biofilm architecture, and interact with efflux systems. Aryl acyl substituents generally outperform aliphatic ones, reflecting stronger hydrophobic and π-stacking interactions and improved binding thermodynamics. Consistent with earlier observations, para-substitution on aromatic rings optimally orients functional groups toward protein subpockets or facilitates interaction with membranes and biofilm matrices.
These trends collectively define an optimal physicochemical and structural envelope for activity, characterized by moderate lipophilicity (c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Log
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 4.5–6.3), TPSA below 90 Å2, and limited conformational flexibility. Compounds exceeding lipophilicity thresholds (>6.5–7) risk solubility and efflux-related drawbacks, while polar molecules suffer from reduced permeability, explaining the observed mid-range optimum. Overall, antibacterial potency, particularly against Gram-positive strains, is enhanced by para-halogen substitution and benzoyl or chloroacetyl modifications, with pyrrolinone cores showing superior engagement with PBP4 and GyrB targets. Antifungal activity, especially against C. albicans, is favored by para-methoxy substitution and balanced lipophilicity, aligning with mechanisms involving fungal cell-wall disruption. Antibiofilm activity against P. aeruginosa is strongly associated with chloroacetyl and benzoyl functionalities combined with para-substitution, which promotes extracellular matrix penetration and biomass reduction. Finally, efflux inhibition is most pronounced in compounds bearing hydrophobic acyl groups and para-substituted aryl rings, enhancing membrane interaction and helping interference with efflux systems.
P 4.5–6.3), TPSA below 90 Å2, and limited conformational flexibility. Compounds exceeding lipophilicity thresholds (>6.5–7) risk solubility and efflux-related drawbacks, while polar molecules suffer from reduced permeability, explaining the observed mid-range optimum. Overall, antibacterial potency, particularly against Gram-positive strains, is enhanced by para-halogen substitution and benzoyl or chloroacetyl modifications, with pyrrolinone cores showing superior engagement with PBP4 and GyrB targets. Antifungal activity, especially against C. albicans, is favored by para-methoxy substitution and balanced lipophilicity, aligning with mechanisms involving fungal cell-wall disruption. Antibiofilm activity against P. aeruginosa is strongly associated with chloroacetyl and benzoyl functionalities combined with para-substitution, which promotes extracellular matrix penetration and biomass reduction. Finally, efflux inhibition is most pronounced in compounds bearing hydrophobic acyl groups and para-substituted aryl rings, enhancing membrane interaction and helping interference with efflux systems.
In summary, our SAR shows that para-substituted aryl groups on the pyrazole, particularly para-halogens (antibacterial) and para-methoxy (antifungal), and aryl acylation of the hydrazide (notably benzoyl and chloroacetyl) drive activity by improving target binding, membrane/EPS penetration, and efflux inhibition. The pyrrolinone core outperforms pyridazinone for MIC and biofilm endpoints, with promising compound 10 delivering MIC 1–8 µg mL−1, ≥99.999% biofilm eradication, and 44–55% efflux reduction within an optimal physicochemical window (c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Log
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 4.5–6.3; TPSA < 90 Å2). Ortho bulk generally reduces activity (torsional penalties). These trends provide clear design rules that explain the superior performance of 4, 6, 7, and 10, and will guide next-round analog synthesis (Fig. 3).
P 4.5–6.3; TPSA < 90 Å2). Ortho bulk generally reduces activity (torsional penalties). These trends provide clear design rules that explain the superior performance of 4, 6, 7, and 10, and will guide next-round analog synthesis (Fig. 3).
Based on these findings, this work advances scientific knowledge in several important ways. It introduces a new chemotype with significant antimicrobial and antibiofilm potential by reporting a previously unexplored library of pyrazole-pyrrolinone derivatives, thereby expanding the chemical space of heterocyclic antimicrobials and offering new structural templates for future drug discovery efforts. The study also shows a unique and valuable combination of potent antibacterial, antifungal, and antibiofilm activities within a single scaffold class; notably, several compounds, particularly compound 10, exhibit simultaneous efficacy against bacteria, fungi, and mature biofilms, addressing multiple unmet challenges in infectious disease treatment. Furthermore, the work provides compelling evidence for a multimodal mechanism of action, integrating efflux pump inhibition, biofilm disruption, and molecular docking to key bacterial targets such as PBP4 and DNA gyrase B, which collectively suggest a broad mechanistic profile with a reduced likelihood of resistance development.
In addition, by synthesizing systematically varied analogues, the study shows appearing SAR, linking specific chemical modifications to biological outcomes and offering valuable direction for future molecular optimization. Importantly, identification of compound 10 as a promising candidate underscores the translational value of this research, as it combines potent antimicrobial activity with low cytotoxicity and favorable physicochemical properties, positioning it as a promising and credible starting point for the development of new anti-infective therapeutics.
Conclusion
This study introduces a new pyrazole-pyrrolinone chemotype that unites potent planktonic antimicrobial, antifungal, and antibiofilm activities, an uncommon combination for early-stage heterocycles, and shows compound 10 as a credible agent. Quantitatively, several analogues achieved MICs of 1–8 µg mL−1 against clinically relevant bacteria, approaching the activity of the fluoroquinolone benchmark ofloxacin under comparable conditions, while compounds 8–10 eradicated mature P. aeruginosa biofilms by ≥99.999% (≥5-log kill within 4 h), a level that far exceeds the sub-log to ∼2-log reductions typically observed for many conventional antibiotics in biofilm settings. The efflux inhibition magnitude (44–55% reduction in EtBr efflux) provides mechanistic value by potentially boosting intracellular drug exposure and mitigating resistance drivers, and the sorbitol protection phenotype supports fungal cell-wall targeting, explaining the robust activity against C. albicans. Safety and developability signals are encouraging human fibroblast IC50 = 120–200 µM yields a preliminary selectivity index (IC50/MIC) in the 15–200 range, while SwissADME properties (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ∼4.5–6.3; TPSA < 90 Å2) align with drug-like space and suggest tractable paths for permeability and formulation.
P ∼4.5–6.3; TPSA < 90 Å2) align with drug-like space and suggest tractable paths for permeability and formulation.
Scientific value added: (i) a previously unexplored pyrazole-pyrrolinone scaffold is shown to deliver near-benchmark planktonic potency together with rare, rapid, and deep biofilm eradication; (ii) multimodal mechanistic evidence (efflux inhibition + putative PBP4/GyrB engagement + fungal cell-wall effects) advances understanding of how a single scaffold can span antibacterial and antifungal indications; and (iii) emerging SAR links substituent patterns to potency, antibiofilm efficacy, and efflux modulation, guiding rational optimization. These features position compound 10 and close analogues for topical treatment of SSTIs and wound infections, antibiofilm device-related indications (e.g., catheter or dressing coatings), and, with ADME optimization, systemic therapy against Gram-positive pathogens and mixed bacterial-fungal biofilms. The combination of fast ≥5-log biofilm kills, low cytotoxicity, and efflux modulation is particularly relevant where standard-of-care agents underperform in biofilm contexts.
Building on this novelty and value, prioritized next steps include mechanistic validation (target engagement vs. PBP4/GyrB; EPS disruption), SAR-driven tuning to moderate c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Log
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P toward 3–4 while preserving TPSA <90 Å2, and preclinical de-risking (in vitro ADME, hERG/CYP, mouse PK). Success criteria include keeping MIC ≤4–8 µg mL−1, reproducing ≥3–5-log biofilm reductions in device models and showing in vivo efficacy in MRSA skin or catheter-biofilm models. Collectively, the data support pyrrolinone 10 as a first-in-class, antibiofilm-capable antimicrobial agent with clear translational potential. This work could be considered as a starting point for a larger SAR driven optimization program. Ongoing and planned future studies will focus on systematic scaffold modification, substituent diversification, and pharmacokinetic optimization to further refine the SAR and advance these compounds toward preclinical development.
P toward 3–4 while preserving TPSA <90 Å2, and preclinical de-risking (in vitro ADME, hERG/CYP, mouse PK). Success criteria include keeping MIC ≤4–8 µg mL−1, reproducing ≥3–5-log biofilm reductions in device models and showing in vivo efficacy in MRSA skin or catheter-biofilm models. Collectively, the data support pyrrolinone 10 as a first-in-class, antibiofilm-capable antimicrobial agent with clear translational potential. This work could be considered as a starting point for a larger SAR driven optimization program. Ongoing and planned future studies will focus on systematic scaffold modification, substituent diversification, and pharmacokinetic optimization to further refine the SAR and advance these compounds toward preclinical development.
Materials and methods
MEL-TEMP II electrothermal melting point device was employed to measure melting points of the compounds obtained. The elemental analyses were conducted on a PerkinElmer 2400 CHN elemental analyzer (PerkinElmer, Waltham, MA) at Faculty of Science, Ain Shams University. Infrared (IR) spectra (ν, cm−1) were measured utilizing KBr disks on Fourier transform-infrared Thermo Electron Nicolet iS10 Spectrometer (Thermo Fisher Scientific Inc. Waltham, MA) at Faculty of Science, Ain Shams University. The 1H and 13C NMR spectra (δ, ppm) were performed on DELTA2 NMR (500 and 100 MHz) and BRUKER Avance III (400 and 100 MHz) spectrometers at National Research Center and Faculty of Pharmacy, Ain Shams University, using tetramethyl silane (TMS) as an internal standard and deuterated dimethyl sulfoxide (DMSO-d6) as a solvent. The reaction progress and purity of products were checked and confirmed by thin-layer chromatography (TLC) using TLC aluminum sheets silica gel 60F254 (Merck, Darmstadt, Germany) with diverse solvent systems as mobile phases.E/Z-3-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-5-(4-methoxyphenyl)furan-2(3H)-one (3)42
Equi-molar mixture of 4-(4-methoxyphenyl)-4-oxobutanoic acid (1), 3-(4-chlorophenyl)-1-phenyl-1H-pyrazole-4-carbaldehyde (2), and fused sodium acetate (0.01 mol) was heated in the presence of ethanoic anhydride (4 mL) at 190–200 °C for 3 h. After cooling, the reaction mixture was poured onto ice-cold water with stirring. The solid formed was collected and recrystallized by ethyl alcohol to get yellow crystals, mp 210–212 °C, yield 81%. FT-IR (KBr, ν, cm−1): 1746 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (500 MHz, DMSO-d6, δ, ppm): 3.79 (s, 3H, OCH3), 7.04 (d, 2H, Ar–H methoxyphenyl, J = 8.1 Hz), 7.39–7.41 (d, 2H, Ar–H, N-phenyl, J = 7.5 Hz), 7.43 (s, 1H, C4–H furanone), 7.54–7.65 (m, 6H, Ar–H + CH
O); 1H NMR (500 MHz, DMSO-d6, δ, ppm): 3.79 (s, 3H, OCH3), 7.04 (d, 2H, Ar–H methoxyphenyl, J = 8.1 Hz), 7.39–7.41 (d, 2H, Ar–H, N-phenyl, J = 7.5 Hz), 7.43 (s, 1H, C4–H furanone), 7.54–7.65 (m, 6H, Ar–H + CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.76 (d, 2H, Ar–H, N-phenyl, J = 7.6 Hz), 8.00 (d, 2H, Ar–H chlorophenyl, J = 7.6 Hz), 9.14 (s, 1H, C5–H pyrazole); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 55.9 (OCH3), 100.1, 115.0 (2), 117.3, 119.9 (2), 120.9, 123.2, 123.8, 127.5 (2), 127.9, 129.3, 129.5, 130.1 (2), 130.3, 130.9, 131.2 (2), 134.3, 139.3, 153.0, 155.3, 161.5, 169.3 (C
), 7.76 (d, 2H, Ar–H, N-phenyl, J = 7.6 Hz), 8.00 (d, 2H, Ar–H chlorophenyl, J = 7.6 Hz), 9.14 (s, 1H, C5–H pyrazole); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 55.9 (OCH3), 100.1, 115.0 (2), 117.3, 119.9 (2), 120.9, 123.2, 123.8, 127.5 (2), 127.9, 129.3, 129.5, 130.1 (2), 130.3, 130.9, 131.2 (2), 134.3, 139.3, 153.0, 155.3, 161.5, 169.3 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); anal. calcd for C27H19ClN2O3 (454.91): C, 71.29; H, 4.21; N, 6.16; found: C, 71.17; H, 4.14; N, 6.13%.
O); anal. calcd for C27H19ClN2O3 (454.91): C, 71.29; H, 4.21; N, 6.16; found: C, 71.17; H, 4.14; N, 6.13%.
E/Z-3-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-5-(4-methoxyphenyl)-1,3-dihydro-2H-pyrrol-2-one (4)
A solution of furan-2(3H)-one 3 (2 mmol) and ammonium acetate (3 mmol) in ethyl alcohol (20 mL) was refluxed for 1 h. The solid deposited during heating was collected and recrystallized from ethyl alcohol/dioxane mixture (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to offer yellow crystals, mp 230–232 °C, yield 79%. FT-IR (KBr, ν, cm−1): 3350 (NH), 1700 (C
1) to offer yellow crystals, mp 230–232 °C, yield 79%. FT-IR (KBr, ν, cm−1): 3350 (NH), 1700 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (500 MHz, DMSO-d6, δ, ppm): 3.80 (s, 3H, OCH3), 6.82 (s, 1H, C4–H pyrrolinone), 6.93 (s, 1H, CH
O); 1H NMR (500 MHz, DMSO-d6, δ, ppm): 3.80 (s, 3H, OCH3), 6.82 (s, 1H, C4–H pyrrolinone), 6.93 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.02 (d, 2H, Ar–H, methoxyphenyl, J = 8.6 Hz), 7.37–7.48 (m, 3H, Ar–H), 7.55–7.72 (m, 4H, Ar–H), 7.83 (d, 2H, Ar–H, J = 8.6 Hz), 8.04 (d, 2H, Ar–H, J = 7.6 Hz), 9.10 (s, 1H, C5–H pyrazole), 10.43 (br.s, 1H, NH); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 55.5, 99.9, 114.8 (2), 118.1, 119.8, 119.9 (2), 120.5, 123.4, 127.7, 128.0 (2), 128.8, 129.5 (2), 130.1 (2), 130.9 (2), 131.3, 134.0, 139.5, 155.1, 161.0, 168.5; anal. calcd for C27H20ClN3O2 (453.93); C, 71.44; H, 4.44; N, 9.26; found: C, 71.31; H, 4.38; N, 9.28%.
), 7.02 (d, 2H, Ar–H, methoxyphenyl, J = 8.6 Hz), 7.37–7.48 (m, 3H, Ar–H), 7.55–7.72 (m, 4H, Ar–H), 7.83 (d, 2H, Ar–H, J = 8.6 Hz), 8.04 (d, 2H, Ar–H, J = 7.6 Hz), 9.10 (s, 1H, C5–H pyrazole), 10.43 (br.s, 1H, NH); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 55.5, 99.9, 114.8 (2), 118.1, 119.8, 119.9 (2), 120.5, 123.4, 127.7, 128.0 (2), 128.8, 129.5 (2), 130.1 (2), 130.9 (2), 131.3, 134.0, 139.5, 155.1, 161.0, 168.5; anal. calcd for C27H20ClN3O2 (453.93); C, 71.44; H, 4.44; N, 9.26; found: C, 71.31; H, 4.38; N, 9.28%.
E/Z-2-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-4-(4-methoxyphenyl)-4-oxobutanehydrazide (5)42
Hydrazine hydrate (3.2 mmol, 80%) was added dropwise to a stirred solution of furanone 3 (3 mmol) in ethyl alcohol (20 mL) and the reaction mixture was further stirred for 3 h at room temperature. The white precipitate was collected and recrystallized from ethyl alcohol to give white crystals, mp 222–224 °C, yield 83%. FT-IR (KBr, ν, cm−1): 3339, 3206 (br. NH, NH2), 1688 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), 1667 (C
O ketone), 1667 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O hydrazide), 1639 (C
O hydrazide), 1639 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H NMR (500 MHz, DMSO-d6, δ, ppm): 3.18 (s, 2H, CH2), 3.72 (s, 3H, OCH3), 4.32 (br.s, 2H, NH2), 6.54 (br.s, 1H, NH), 6.90 (d, 2H, Ar–H, J = 8.6 Hz), 7.13 (s, 1H, CH
N); 1H NMR (500 MHz, DMSO-d6, δ, ppm): 3.18 (s, 2H, CH2), 3.72 (s, 3H, OCH3), 4.32 (br.s, 2H, NH2), 6.54 (br.s, 1H, NH), 6.90 (d, 2H, Ar–H, J = 8.6 Hz), 7.13 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.29–7.31 (m, 3H, Ar–H), 7.45–7.48 (dd, 2H, Ar–H, J = 8.1 and 7.6 Hz), 7.59–7.64 (m, 4H, Ar–H), 7.95 (d, 2H, Ar–H, J = 8.1 Hz), 8.67 (s, 1H, C5–H pyrazole); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 43.5 (CH2), 55.7 (OCH3), 89.7, 100.0, 114.0 (2), 117.2, 118.4, 119.5 (2), 127.5 (2), 128.5, 128.8, 129.4 (2), 129.9 (2), 130.0, 130.7 (2), 133.9, 136.2, 139.6, 151.8, 159.2, 166.5; anal. calcd for C27H23ClN4O3 (486.96): C, 66.60; H, 4.76; N, 11.51; found: C, 66.51; H, 4.71; N, 11.53%.
), 7.29–7.31 (m, 3H, Ar–H), 7.45–7.48 (dd, 2H, Ar–H, J = 8.1 and 7.6 Hz), 7.59–7.64 (m, 4H, Ar–H), 7.95 (d, 2H, Ar–H, J = 8.1 Hz), 8.67 (s, 1H, C5–H pyrazole); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 43.5 (CH2), 55.7 (OCH3), 89.7, 100.0, 114.0 (2), 117.2, 118.4, 119.5 (2), 127.5 (2), 128.5, 128.8, 129.4 (2), 129.9 (2), 130.0, 130.7 (2), 133.9, 136.2, 139.6, 151.8, 159.2, 166.5; anal. calcd for C27H23ClN4O3 (486.96): C, 66.60; H, 4.76; N, 11.51; found: C, 66.51; H, 4.71; N, 11.53%.
4-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methyl)-6-(4-methoxyphenyl)pyridazin-3(2H)-one (6)
A suspension of hydrazide 5 (2 mmol) in a mixture of hydrochloric and ethanoic acids (10 mL, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) was refluxed for 1 h, then left to stand at RT. The reaction mixture poured onto ice-cold water with stirring. The solid obtained was collected and recrystallized by methyl alcohol to offer white crystals, mp 241–243 °C, yield 64%. FT-IR (KBr, ν, cm−1): 3369 (NH), 1656 (C
1) was refluxed for 1 h, then left to stand at RT. The reaction mixture poured onto ice-cold water with stirring. The solid obtained was collected and recrystallized by methyl alcohol to offer white crystals, mp 241–243 °C, yield 64%. FT-IR (KBr, ν, cm−1): 3369 (NH), 1656 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.76 (s, 3H, OCH3), 3.94 (s, 2H, CH2), 6.82 (s, 1H, C5–H pyridazine), 6.99–7.86 (m, 13H, Ar–H), 8.42 (s, 1H, C5–H pyrazole), 13.08 (br.s, 1H, NH); anal. calcd for C27H21ClN4O2 (468.94): C, 69.16; H, 4.51; N, 11.95; found: C, 69.02; H, 4.44; N, 11.93%.
O); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.76 (s, 3H, OCH3), 3.94 (s, 2H, CH2), 6.82 (s, 1H, C5–H pyridazine), 6.99–7.86 (m, 13H, Ar–H), 8.42 (s, 1H, C5–H pyrazole), 13.08 (br.s, 1H, NH); anal. calcd for C27H21ClN4O2 (468.94): C, 69.16; H, 4.51; N, 11.95; found: C, 69.02; H, 4.44; N, 11.93%.
E/Z-N-(3-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-5-(4-methoxyphenyl)-2-oxo-2,3-dihydro-1H-pyrrol-1-yl)acetamide (7)
A suspension of acid hydrazide 5 (2 mmol) in ethanoic anhydride (5 mL) was stirred at RT for 5 h or heated on water bath at 80–90 °C for 2 h. The solid formed was filtered and recrystallized by ethyl alcohol to obtain yellow crystals, mp 290–292 °C, yield 84%. FT-IR (KBr, ν, cm−1): 3279 (NH), 1724, 1668 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (500 MHz, DMSO-d6, δ, ppm): (E- & Z-isomer, 55
O); 1H NMR (500 MHz, DMSO-d6, δ, ppm): (E- & Z-isomer, 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 45%), 1.78 & 1.84 (s, 3H, CH3), 3.72 & 3.79 (s, 3H, OCH3), 6.65 & 6.75 (s, 1H, C4–H pyrrolinone), 6.88 (d, 2H, Ar–H, J = 8.6 Hz), 7.01–7.05 (m, 3H, Ar–H), 7.20 (s, 1H, CH
45%), 1.78 & 1.84 (s, 3H, CH3), 3.72 & 3.79 (s, 3H, OCH3), 6.65 & 6.75 (s, 1H, C4–H pyrrolinone), 6.88 (d, 2H, Ar–H, J = 8.6 Hz), 7.01–7.05 (m, 3H, Ar–H), 7.20 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.31–7.67 (m, 4H, Ar–H), 7.97 (d, 2H, Ar–H, J = 7.6 Hz), 8.03 (d, 2H, Ar–H, J = 7.6 Hz), 8.73 & 9.17 (s, 1H, C5–H pyrazole), 9.71 & 10.51 (br.s, 1H, NH); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 21.0 (CH3), 55.7 (OCH3), 89.7, 113.7, 114.6, 116.9, 117.3, 119.8, 120.4 (2), 122.3, 126.6, 127.8 (2), 128.8, 129.4 (2), 130.4 (2), 131.4 (2), 135.0, 139.5, 146.9, 153.1, 159.4, 160.8, 165.8, 169.2; anal. calcd for C29H23ClN4O3 (510.98): C, 68.17; H, 4.54; N, 10.96; found: C, 68.08; H, 4.48; N, 10.94%.
), 7.31–7.67 (m, 4H, Ar–H), 7.97 (d, 2H, Ar–H, J = 7.6 Hz), 8.03 (d, 2H, Ar–H, J = 7.6 Hz), 8.73 & 9.17 (s, 1H, C5–H pyrazole), 9.71 & 10.51 (br.s, 1H, NH); 13C NMR (125 MHz, DMSO-d6, δ, ppm): 21.0 (CH3), 55.7 (OCH3), 89.7, 113.7, 114.6, 116.9, 117.3, 119.8, 120.4 (2), 122.3, 126.6, 127.8 (2), 128.8, 129.4 (2), 130.4 (2), 131.4 (2), 135.0, 139.5, 146.9, 153.1, 159.4, 160.8, 165.8, 169.2; anal. calcd for C29H23ClN4O3 (510.98): C, 68.17; H, 4.54; N, 10.96; found: C, 68.08; H, 4.48; N, 10.94%.
E/Z-N-(3-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-5-(4-methoxyphenyl)-2-oxo-2,3-dihydro-1H-pyrrol-1-yl)benzamide (8)
A solution of the hydrazide 5 (2 mmol) and benzoyl chloride (2 mmol) in 1,4-dioxane (10 mL) involving Et3N (0.1 mL) was stirred for 3 h at an ambient temperature. The solid obtained was collected by filtration and recrystallized by methanol to get yellow crystals, mp 245–247 °C, yield 78%. FT-IR (KBr, ν, cm−1): 3218 (NH), 1696, 1650 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.73 (s, 3H, OCH3), 5.69 (s, 1H, C4–H pyrrolinone), 6.95 (d, 2H, Ar–H, J = 8.8 Hz), 7.18 (s, 1H, CH
O); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.73 (s, 3H, OCH3), 5.69 (s, 1H, C4–H pyrrolinone), 6.95 (d, 2H, Ar–H, J = 8.8 Hz), 7.18 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.35 (dd, 2H, Ar–H, J = 7.2 & 7.6 Hz), 7.47–7.65 (m, 12H, Ar–H), 7.93 (d, 2H, Ar–H, J = 8.0 Hz), 8.57 (s, 1H, C5–H pyrazole), NH not observed due to rapid exchange; anal. calcd for C34H25ClN4O3 (573.05): C, 71.26; H, 4.40; N, 9.78; found: C, 71.17; H, 4.32; N, 9.80%.
), 7.35 (dd, 2H, Ar–H, J = 7.2 & 7.6 Hz), 7.47–7.65 (m, 12H, Ar–H), 7.93 (d, 2H, Ar–H, J = 8.0 Hz), 8.57 (s, 1H, C5–H pyrazole), NH not observed due to rapid exchange; anal. calcd for C34H25ClN4O3 (573.05): C, 71.26; H, 4.40; N, 9.78; found: C, 71.17; H, 4.32; N, 9.80%.
E/Z-2-Chloro-N-(3-((3-(4-chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-5-(4-methoxyphenyl)-2-oxo-2,3-dihydro-1H-pyrrol-1-yl)acetamide (9)
To a stirred solution of hydrazide 5 (2 mmol) in 1,4-dioxane (10 mL) involving Et3N (0.1 mL), chloroacetyl chloride (2 mmol) was added dropwise and the reaction mixture was further stirred for 2 h at room temperature. The solid obtained was collected and recrystallized from methanol to give orange crystals, mp 261–263 °C, yield 81%. FT-IR (KBr, ν, cm−1): 3269 (NH), 1731, 1682 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.80 (s, 3H, OCH3), 4.20 (s, 2H, CH2), 6.83 (s, 1H, C4–H pyrrolinone), 7.05 (d, 2H, Ar–H, J = 8.8 Hz), 7.11 (s, 1H, CH
O); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.80 (s, 3H, OCH3), 4.20 (s, 2H, CH2), 6.83 (s, 1H, C4–H pyrrolinone), 7.05 (d, 2H, Ar–H, J = 8.8 Hz), 7.11 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.42–7.74 (m, 9H, Ar–H), 8.07 (d, 2H, Ar–H, J = 8.0 Hz), 9.21 (s, 1H, C5–H pyrazole), 11.04 (br.s, 1H, NH); 13C NMR (100 MHz, DMSO-d6, δ, ppm): 55.8, 66.8, 99.1, 114.6 (2), 117.2, 119.8 (2), 121.1, 121.9 (2), 122.0, 126.2, 127.8 (2), 129.2 (2), 129.4, 130.0 (2), 130.9, 131.1, 134.2, 139.4, 146.3, 153.1, 160.8, 166.1, 168.8; anal. calcd for C29H22Cl2N4O3 (545.42): C, 63.86; H, 4.07; N, 10.27; found: C, 63.72; H, 4.00; N, 10.25%.
), 7.42–7.74 (m, 9H, Ar–H), 8.07 (d, 2H, Ar–H, J = 8.0 Hz), 9.21 (s, 1H, C5–H pyrazole), 11.04 (br.s, 1H, NH); 13C NMR (100 MHz, DMSO-d6, δ, ppm): 55.8, 66.8, 99.1, 114.6 (2), 117.2, 119.8 (2), 121.1, 121.9 (2), 122.0, 126.2, 127.8 (2), 129.2 (2), 129.4, 130.0 (2), 130.9, 131.1, 134.2, 139.4, 146.3, 153.1, 160.8, 166.1, 168.8; anal. calcd for C29H22Cl2N4O3 (545.42): C, 63.86; H, 4.07; N, 10.27; found: C, 63.72; H, 4.00; N, 10.25%.
E/Z-3-((3-(4-Chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-1-((4-methoxybenzylidene)amino)-5-(4-methoxyphenyl)-1,3-dihydro-2H-pyrrol-2-one (10)
A solution of hydrazide 5 (2 mmol) and 4-methoxybenzaldehyde (2 mmol) in 1,4-dioxane (15 mL) involving glacial ethanoic acid (0.2 mL) was refluxed for 3 h. After cooling, the solid was filtered and recrystallized by 1,4-dioxane to obtain red crystals, mp 281–283 °C, yield 75%. FT-IR (KBr, ν, cm−1): 1683 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1621 (C
O), 1621 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.80 (s, 3H, OCH3), 3.82 (s, 3H, OCH3 benzylidene), 6.86 (s, 1H, C4–H pyrrolinone), 7.02–7.08 (m, 4H, Ar–H), 7.13 (s, 1H, CH
N); 1H NMR (400 MHz, DMSO-d6, δ, ppm): 3.80 (s, 3H, OCH3), 3.82 (s, 3H, OCH3 benzylidene), 6.86 (s, 1H, C4–H pyrrolinone), 7.02–7.08 (m, 4H, Ar–H), 7.13 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.41 (t, 1H, Ar–H, J = 7.2 Hz), 7.55–7.69 (m, 8H, Ar–H), 7.78 (d, 2H, Ar–H, J = 8.4 Hz), 8.05 (d, 2H, Ar–H, J = 8.0 Hz), 9.18 (s, 1H, C5–H pyrazole), 9.29 (s, 1H, CH
), 7.41 (t, 1H, Ar–H, J = 7.2 Hz), 7.55–7.69 (m, 8H, Ar–H), 7.78 (d, 2H, Ar–H, J = 8.4 Hz), 8.05 (d, 2H, Ar–H, J = 8.0 Hz), 9.18 (s, 1H, C5–H pyrazole), 9.29 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N); anal. calcd for C35H27ClN4O3 (587.08): C, 71.61; H, 4.64; N, 9.54; found: C, 71.48; H, 4.56; N, 9.51%.
N); anal. calcd for C35H27ClN4O3 (587.08): C, 71.61; H, 4.64; N, 9.54; found: C, 71.48; H, 4.56; N, 9.51%.
Antimicrobial activity
Full details about the five microbial strains used to evaluate antimicrobial activity can be found in SI file.
Conflicts of interest
No potential conflict of interest was reported by the author(s).Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information (SI) files. Supplementary information: docking, ADME, and spectral data. See DOI: https://doi.org/10.1039/d6ra02508k.Acknowledgements
The authors extend their appreciation to the Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R25), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.References
- E. A. E. El-Helw, W. S. I. Abou-Elmagd, E. M. Hosni, M. Kamal, A. I. Hashem and S. K. Ramadan, Synthesis of Benzo[h]quinoline Derivatives and Evaluation of Their Insecticidal Activity Against Culex pipiens L. Larvae, Eur. J. Med. Chem., 2025, 290, 117565 CrossRef CAS PubMed.
- S. K. Ramadan, W. S. I. Abou-Elmagd, E. M. Hosni, M. Kamal, A. I. Hashem and E. A. E. El-Helw, Synthesis, in vivo evaluation, and in silico molecular docking of benzo[h]quinoline derivatives as potential Culex pipiens L. larvicides, Bioorg. Chem., 2025, 154, 108090 CrossRef CAS PubMed.
- A. R. Morsy, S. H. Mahmoud and N. M. Abou Shama, et al., Antiviral activity of pyrazole derivatives bearing a hydroxyquinoline scaffold against SARS-CoV-2, HCoV-229E, MERS-CoV, and IBV propagation, RSC Adv., 2024, 14(38), 27935–27947 RSC.
- S. K. Ramadan, H. S. Abd-Rabboh, N. M. Gad, W. S. I. Abou-Elmagd and D. S. Haneen, Synthesis and characterization of some chitosan-quinoline nanocomposites as potential insecticidal agents, Polycyclic Aromat. Compd., 2023, 43(8), 7013–7026 CrossRef CAS.
- S. K. Ramadan and E. A. E. El-Helw, Synthesis and antimicrobial activity evaluation of some novel heterocycles derived from chromonyl-2(3H)-furanone, J. Chem. Res., 2018, 42(6), 332–336 CrossRef CAS.
- A. K. El-Ziaty, W. S. I. Abou-Elmagd, S. K. Ramadan and A. I. Hashem, Synthesis and biological screening of some chromonyl-substituted heterocycles derived from 2(3H)-furanone derivative, Synth. Commun., 2017, 47(5), 471–480 CrossRef CAS.
- E. A. E. El-Helw, A. R. Morsy and A. I. Hashem, Evaluation of Some New Heterocycles Bearing 2-Oxoquinolyl Moiety as Immunomodulator Against Highly Pathogenic Avian Influenza Virus (H5N8), J. Heterocycl. Chem., 2021, 58(4), 1003–1014 CrossRef CAS.
- Y. M. Youssef, M. E. Azab, G. A. Elsayed, A. A. El-Sayed, A. I. Hassaballah and E. A. E. El-Helw, Synthesis and antioxidant, antimicrobial, and antiviral activity of some pyrazole-based heterocycles using a 2(3H)-furanone derivative, J. Iran. Chem. Soc., 2023, 20(9), 2203–2216 CrossRef CAS.
- B.-C. Ivan, S.-F. Barbuceanu and C. M. Hotnog, et al., New Pyrrole Derivatives as Promising Biological Agents: Design, Synthesis, Characterization, In Silico, and Cytotoxicity Evaluation, Int. J. Mol. Sci., 2022, 23(16), 8854 CrossRef CAS PubMed.
- J. Pohlmann, T. Lampe and M. Shimada, et al., Pyrrolidinedione derivatives as antibacterial agents with a novel mode of action, Bioorg. Med. Chem. Lett., 2005, 15(4), 1189–1192 CrossRef CAS PubMed.
- M. S. Abdelbaset, M. Abdel-Aziz, G. E. A. Abuo-Rahma, M. Ramadan and M. H. Abdelrahman, Pyridazinones and pyrrolinones as promising Scaffolds in Medicinal Chemistry, J. Adv. Biomed. Pharm. Sci., 2019, 2(1), 19–28 Search PubMed.
- P. Zarantonello, C. P. Leslie, R. Ferritto and W. M. Kazmierski, Total synthesis and semi-synthetic approaches to analogues of antibacterial natural product althiomycin, Bioorg. Med. Chem. Lett., 2002, 12(4), 561–565 CrossRef CAS PubMed.
- M. Böhm, I. Morano, B. Pieske, J. Rüegg, M. Wankerl, R. Zimmermann and E. Erdmann, Contribution of camp-phosphodiesterase inhibition and sensitization of the contractile proteins for calcium to the inotropic effect of pimobendan in the failing human myocardium, Circ. Res., 1991, 68(3), 689–701 CrossRef PubMed.
- M. Gökçe, Y. Dündar and E. Küpelġ, Synthesis of (6-substituted-3(2H)-pyridazinon-2-yl)acetic acid and (6-substituted-3(2H)-pyridazinon-2-yl)acetamide derivatives and investigation of their analgesic and anti-inflammatory activities, Fabad J. Pharm. Sci., 2010, 35, 1–11 Search PubMed.
- A. Husain, M. M. Alam and N. Siddiqui, Synthesis, reactions and biological activity of 3-arylidene-5-(4-methylphenyl)-2(3H)-furanones, J. Serb. Chem. Soc., 2009, 74(2), 103–115 CrossRef CAS.
- A. Ahmad, A. Husain, S. A. Khan, M. Mujeeb and A. Bhandari, Design, synthesis, molecular properties and antimicrobial activities of some novel 2(3H)-pyrrolinone derivatives, J. Saudi Chem. Soc., 2015, 19(3), 340–346 CrossRef.
- A. Husain, M. M. Alam, M. Shaharyar and S. Lal, Antimicrobial activities of some synthetic butenolides and their pyrrolinone derivatives, J. Enzyme Inhib. Med. Chem., 2010, 25(1), 54–61 CrossRef CAS PubMed.
- W. S. I. Abou-Elmagd, A. K. El-Ziaty, M. I. Elzahar, S. K. Ramadan and A. I. Hashem, Synthesis and antitumor activity evaluation of some N-heterocycles derived from pyrazolyl-substituted 2(3H)-furanone, Synth. Commun., 2016, 46(14), 1197–1208 CrossRef CAS.
- S. H. Abbas, G. E.-D. Abuo-Rahma, M. Abdel-Aziz, O. M. Aly, E. A. Beshr and A. M. Gamal-Eldeen, Synthesis, cytotoxic activity, and tubulin polymerization inhibitory activity of new pyrrol-2(3H)-ones and pyridazin-3(2H)-ones, Bioorg. Chem., 2016, 66, 46–62 CrossRef CAS.
- R. Singh, V. K. Bhardwaj, J. Sharma, P. Das and R. Purohit, Identification of selective cyclin-dependent kinase 2 inhibitor from the library of pyrrolinone-fused benzosuberene compounds: an in silico exploration, J. Biomol. Struct. Dyn., 2022, 40(17), 7693–7701 CrossRef CAS PubMed.
- F. Ramzan, S. A. Nabi, M. S. Lone, A. Bonardi, A. Hamid, S. Bano, K. Sharma, S. Shafi, M. Samim, K. Javed and C. T. Supuran, Synthesis, biological evaluation and theoretical studies of (E)-1-(4-sulfamoyl-phenylethyl)-3-arylidene-5-aryl-1H-pyrrol-2(3H)-ones as human carbonic anhydrase inhibitors, J. Enzyme Inhib. Med. Chem., 2023, 38(1), 2189126 CrossRef.
- E. A. E. El-Helw and A. I. Hashem, Synthesis and antitumor activity evaluation of some pyrrolinone and pyridazinone heterocycles derived from 3-((2-oxo-5-(p-tolyl)furan-3(2H)-ylidene) methyl)quinolin-2(1H)-one, Synth. Commun., 2020, 50(7), 1046–1055 CrossRef CAS.
- M. S. Abdelbaset, G. A. Abuo-Rahma, M. H. Abdelrahman, M. Ramadan, B. G. M. Youssif, S. N. A. Bukhari, M. F. A. Mohamed and M. Abdel-Aziz, Novel pyrrol-2(3H)-ones and pyridazin-3(2H)-ones carrying quinoline scaffold as anti-proliferative tubulin polymerization inhibitors, Bioorg. Chem., 2018, 80, 151–163 CrossRef CAS.
- R. S. Kumar, A. I. Almansour, N. Arumugam, F. Mohammad and R. R. Kumar, In vitro Mechanistic Exploration of Novel Spiropyrrolidine Heterocyclic Hybrids as Anticancer Agents, Front. Chem., 2020, 8, 465 CrossRef CAS PubMed.
- N. V. Ortiz Zacarías, J. P. D. van Veldhoven, L. Portner, E. van Spronsen, S. Ullo, M. Veenhuizen, W. J. C. van der Velden, A. J. M. Zweemer, R. M. Kreekel, K. Oenema, E. B. Lenselink, L. H. Heitman and A. P. IJzerman, Pyrrolinone Derivatives as Intracellular Allosteric Modulators for Chemokine Receptors: Selective and Dual-Targeting Inhibitors of CC Chemokine Receptors 1 and 2, J. Med. Chem., 2018, 61(20), 9146–9161 CrossRef.
- Y. Lee, K.-Q. Ling, X. Lu, R. B. Silverman, E. M. Shepard, D. M. Dooley and L. M. Sayre, 3-Pyrrolines Are Mechanism-Based Inactivators of the Quinone-Dependent Amine Oxidases but Only Substrates of the Flavin-Dependent Amine Oxidases, J. Am. Chem. Soc., 2002, 124(41), 12135–12143 CrossRef CAS PubMed.
- H.-K. Fun, S. Arshad, S. Malladi, R. Selvam and A. M. Isloor, 3-(4-Chloro-phen-yl)-1-phenyl-1H-pyrazole-4-carbaldehyde, Acta Crystallogr., Sect. E:Struct. Rep. Online, 2011, 67(7), o1783–o1784 CrossRef CAS.
- R. M. Humphries, J. Ambler, S. L. Mitchell, M. Castanheira, T. Dingle, J. A. Hindler, L. Koeth and K. Sei, CLSI methods development and standardization working group best practices for evaluation of antimicrobial susceptibility tests, J. Clin. Microbiol., 2018, 56(4), 01934–01917 CrossRef.
- A. Bahuguna, I. Khan, V. K. Bajpai and S. C. Kang, MTT assay to evaluate the cytotoxic potential of a drug, Bangladesh J. Pharmacol., 2017, 12(2), 115–118 Search PubMed.
- A. Lavrikova, M. Janda, H. Bujdáková and K. Hensel, Eradication of single-and mixed-species biofilms of P. aeruginosa and S. aureus by pulsed streamer corona discharge cold atmospheric plasma, Sci. Total Environ., 2025, 959, 178184 CrossRef CAS PubMed.
- S. Hamed and M. Emara, Antibacterial and antivirulence activities of acetate, zinc oxide nanoparticles, and vitamin C against E. coli O157: H7 and P. aeruginosa, Curr. Microbiol., 2023, 80(2), 57 CrossRef CAS PubMed.
- D. Miron, F. Battisti, F. K. Silva, A. D. Lana, B. Pippi, B. Casanova, S. Gnoatto, A. Fuentefria, P. Mayorga and E. E. Schapoval, Antifungal activity and mechanism of action of monoterpenes against dermatophytes and yeasts, Rev. Bras. Farmacogn., 2014, 24(6), 660–667 CrossRef CAS.
- E. E. Whittle, H. E. McNeil, E. Trampari, M. Webber, T. W. Overton and J. M. Blair, Efflux impacts intracellular accumulation only in actively growing bacterial cells, mBio, 2021, 12(5), 02608–02621 CrossRef.
- A. Arzine, H. Hadni, K. Boujdi and K. Chebbac, et al., Efficient Synthesis, Structural Characterization, Antibacterial Assessment, ADME-Tox Analysis, Molecular Docking and Molecular Dynamics Simulations of New Functionalized Isoxazoles, Pharmaceuticals, 2024, 29(14), 3366 CAS.
- E. A. E. El-Helw, A. Y. Alzahrani and S. K. Ramadan, Synthesis and antimicrobial activity of thiophene-based heterocycles derived from thiophene-2-carbohydrazide, Future Med. Chem., 2024, 16(5), 439–451 CrossRef CAS PubMed.
- S. K. Ramadan, N. A. Ibrahim, S. A. El-Kaed and E. A. E. El-Helw, New potential fungicides pyrazole-based heterocycles derived from 2-cyano-3-(1,3-diphenyl-1H-pyrazol-4-yl)acryloyl isothiocyanate, J. Sulfur Chem., 2021, 42(5), 529–546 CrossRef CAS.
- A. H. Abdelrahman, M. E. Azab, M. A. Hegazy, A. Labena and A. Y. Alzahrani, SK. Ramadan, Synthesis, Computational Analysis, and Exploring Antiproliferative Activity of Triazolo- and Thiazolo-Pyrimidine Derivatives as Potential EGFR Inhibitors, J. Mol. Struct., 2025, 1333, 141789 CrossRef CAS.
- S. K. Ramadan, S. M. Gomha and E. A. E. El-Helw, Straightforward synthesis and in silico evaluation of pyrazolylthiazolidinone derivatives as prospective antiproliferative agents, Bioorg. Chem., 2025, 165, 109036 CrossRef CAS PubMed.
- A. I. Khodair, D. R. Imam, N. A. Kheder, A. M. Fahim and A. A. El-Barbary, Corrigendum to “Synthesis, DFT investigation, molecular docking analysis, ADMET studies, and biological evaluation of a novel series of imidazolidinone derivatives as potential antimicrobial agents, J. Mol. Struct., 2025, 1329, 141415 CrossRef CAS.
- A. El-Sewedy, E. A. El-Bordany, N. F. Mahmoud, K. A. Ali and S. K. Ramadan, One-pot synthesis, computational chemical study, molecular docking, biological study, and in silico prediction ADME/pharmacokinetics properties of 5-substituted 1H-tetrazole derivatives, Sci. Rep., 2023, 13, 17869 CrossRef CAS.
- H. S. El-Hema, S. M. Soliman and W. El-Dougdoug, et al., Design, Characterization, Antimicrobial Activity, and In Silico Studies of Theinothiazoloquinazoline Derivatives Bearing Thiazinone, Tetrazole, and Triazole Moieties, ACS Omega, 2025, 10(9), 9703–9717 CrossRef CAS PubMed.
- A. M. El-Gendy, H. Y. Medrasi, M. A. Al-Sheikh and A. A. Othman, Synthesis, some reactions of 3-((3-(4-chlorophenyl)-1-phenyl-1H-pyrazol-4-yl)methylene)-5-(4- methoxyphenyl)furan-2(3H)-one Containing 1-phenyl-3-p-chlorophenyl pyrazole and investigation of their antibacterial, antifungal and anticancer activities, Am. J. Sci., 2020, 16(6), 8–20 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |



