Open Access Article
Open Access ArticleOne-pot synthesis of fluorescent cap1 mRNAs using labelled trinucleotide m7G-cap analogs for real-time in vitro and in vivo mRNA tracking
Chou Yang†
a,
Chen Guo†b,
Yiqian Zhou†b,
Suyong Lib,
Mengjie Lia,
Yalin Xieb,
Yanhui Liub,
Xinyu Lua,
Lijun Zhangb,
Guoheng Moa,
Wei Zhua,
Jinlin Hou*a and
Jiancun Zhang *b
*b
aState Key Laboratory of Organ Failure Research, Guangdong Provincial Key Laboratory of Viral Hepatitis Research, Guangdong Institute of Liver Diseases, Department of Infectious Diseases and Hepatology Unit, Nanfang Hospital, Southern Medical University, Guangzhou, 510515, China. E-mail: larry@henovcom.com; Houjlhousmu@163.com
bGuangzhou Henovcom Bioscience Co., Ltd., Guangzhou, Guangdong 510700, China
First published on 14th May 2026
Abstract
Visualizing synthetic labelled mRNAs in cells and animals enables the study of mRNA functions/dynamics and also helps to evaluate the delivery, distribution, intracellular kinetics, and efficacy of mRNA-based therapeutics/vaccines. Current methods for synthesizing labelled mRNAs, suitable for in vitro and in vivo tracking, rely on tedious and strenuous post-transcription modification processes. Herein, we report a direct one-pot co-transcriptional synthesis of labelled cap1 mRNAs by employing a series of fluorescently tagged trinucleotide cap analogs. These synthesized and translatable mRNAs with a single labelled fluorescent tag at the 3′ position of the ribose ring of the m7G cap enabled real-time tracking of mRNA in biological systems without the need for additional chemical modifications. Robust transcriptional yield, capping efficiency and transcript integrity were achieved by the standard enzymatic co-transcription protocol with these fluorophore-labelled m7GpppNN trinucleotide analogues. These synthesized labelled mRNAs retained their translational competence and activity, as confirmed by protein expression assays in mammalian cells. Further, we demonstrated that mRNAs with these labelled caps allowed direct visualization of mRNA uptake, intracellular trafficking and localization, and biodistribution using fluorescence microscopy and whole-animal imaging in mouse models. This strategy streamlines the generation of functional and trackable mRNAs, providing a powerful tool for the study of mRNA function and dynamics, for further evaluating mRNA vaccines and therapeutic delivery systems.
Introduction
Messenger RNA (mRNA) technology has emerged as a transformative modality in the fields of prophylactic vaccines, protein and enzyme replacement therapy, gene editing, immunotherapy and regenerative medicine, as exemplified by the rapid development of COVID-19 mRNA vaccines.1 Its unique ability to transiently produce proteins in vivo without risk of integration into the genome or requiring nuclear delivery has paved the way for innovative applications across a wide spectrum of diseases, including various cancers, genetic disorders and infectious diseases.2 Currently, lipid nanoparticles (LNPs) are the most widely used delivery system for mRNA based-vaccine and therapeutic development.3 Given that the half-life of intracellular mRNAs is short4,5 and their concentrations range from nanomolar to picomolar levels,6 a persistent challenge remains in the real-time monitoring of mRNA dynamics in vitro and in vivo in order to screen appropriate delivery vehicles and to evaluate mRNA localization and endosomal escape efficiency, which are crucial for optimizing delivery systems, enhancing therapeutic precision, and understanding the target organ of delivery and the fate of intracellular mRNAs.Current methods for the preparation of labelled mRNAs often require multiple steps through postmodification of the transcribed mRNAs either by chemical modifications or by combined enzymatic and chemical modifications, making the process time-consuming and complex.7 mRNA conjugation with various labels can also be achieved through either random or site-specific labeling. Since mRNA is a linear polymer, the main challenge lies in attaching a label to a single specific site without compromising its translational activity. Chemical modifications on the mRNA bases are limited as these modifications must not disturb base-pair interactions and should not interfere with ribosome-guided decoding of the open-reading frame. The universal regulatory elements of the 5′ cap and poly (A) tail provide desirable sites for labelling modification.8,9 Other reported approaches to visualize mRNA include in situ hybridization (ISH),10 molecular beacon11 or spinach aptamers;12 however, these methods have limited applications due to the design of probes that only recognize a specific transcript sequence to incorporate a large sequence into the mRNA. Recently, various groups have investigated fluorescent labelled mRNAs for the study of their localization and delivery. Baird has recently reported the DNA template-based site-specific incorporation of a single modified nucleotide at the 3′ terminus of various RNAs, but it requires several enzymatic and chemical steps.13 A dual radionuclide–near-infrared probe was developed allowing for the quantitative, longitudinal and non-invasive monitoring of mRNA vaccines in monkeys.14 A photoaffinity approach was reported using an aptamer ligand functionalized with a photoactivatable diazirine reactive group, resulting in covalent attachment to the RNA of interest upon radiation.15 The method of multicovalent modification at the polyA tail for enhancing the translational activity of mRNAs was also described.16 Site-specific substitution of guanine at the hairpin with modified 7-deazaguananine derivatives for labelling mRNA was also reported with guanine transglycosylase combining chemical attachment of fluorogenic groups.17 In another study, the cytosine base was fully substituted with a fluorescent nucleobase analogue, phenoxazine, to yield translatable and trackable fluorogenic mRNA molecules, but this may alter the secondary structure of mRNA from the one with the natural base.18 Jemielity et al. reported an innovative approach to synthesize either anthraniloyl (Ant) or N-methylanthraniloyl (Mant)-labelled cap0 mRNA directly with dinucleotide cap analogs with moderate capping rates. His group also reported azido-functionalized cap analogs that allow for post-transcriptional fluorescence labeling via bioorthogonal click chemistry, facilitating the study of mRNA dynamics in living cells.19 A similar approach of N-7 of m7G cap modification with an azide moiety was also reported.20
In response to the need for facile syntheses of trackable and translatable cap1 mRNA, we designed and synthesized trinucleotide cap analogs pre-installed with various fluorescent groups, including fluorescein amidite (FAM), methylanthraniloyl (Mant), and cyanine dyes through various linkers (spacers) attached to the 3′-aminomethylgroup of the ribose at the m7G moiety. This novel approach allows direct one-pot syntheses of labelled cap1 mRNAs, eliminating the need for cumbersome post-transcriptional modifications. By leveraging the unique properties of m7G-capped trinucleotide analogs, this method introduces fluorescent labels directly during transcription, without compromising the integrity of the mRNA. Notably, the labelled mRNA retains its translational functionality and can be efficiently encapsulated in lipid nanoparticles, enabling targeted delivery evaluation.
This method not only simplifies the synthesis of labelled cap1 mRNA but also enables the real-time tracking of cap1 mRNA in live cells and animal models, avoiding the immunogenicity issues often associated with cap0 mRNA;21 thus, this approach provides a robust platform for evaluating and optimizing mRNA delivery systems. In summary, this method offers a powerful tool for the facile syntheses of labelled mRNA for studying mRNA behavior and facilitates the development of more effective mRNA-based therapeutics and vaccines.
Materials and methods
Chemical information
The details about all the chemicals and solvents are provided in the SI.Cell lines
HEK293T cells and Huh-7 cells (BeNa Culture Collection, China) were maintained in DMEM supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin. Jurkat T cells (iCell Bioscience, China) were propagated in an RPMI 1640 medium containing 10% heat-inactivated FBS with identical antibiotic concentrations (100 U mL−1 penicillin and 100 µg mL−1 streptomycin). All cell lines were incubated at 37 °C under a 5% CO2 humidified atmosphere.In vitro transcription of mRNA
A 20 µL reaction volume of in vitro transcription (IVT) includes: 1 µg linearized DNA template, 5 mM of ATP, CTP, GTP and N1-Me-pUTP (Henovcom, China), 4 mM of CleanCap®AG (3′-OMe)/LZCap®AG (3′-Acm)/Cy3/Cy5/Cy7/FAM/Mant-labelled LZCap®AG (the wavelengths of each cap are listed in Table 1), 2 µL of T7 RNA polymerase mix (NEB, USA), T7 reaction buffer (NEB, USA) and RNase-free water (Shaghai Titan Scientific, China). The mixture was incubated for 2–3 h at 37 °C. After transcription, 0.5 µL of DNase I (Yeasen, China) was added, followed by incubation at 37 °C for 15 min to remove DNA template. Crude mRNA products were then purified by adding 30 µL of 7.5 M LiCl and 30 µL of RNase-free water and incubated at −20 °C for at least half an hour. The mRNA precipitate was centrifuged at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 8 min and washed three times with icy 75% ethyl alcohol. The precipitate was dissolved in RNase-free water. The concentrations of labelled mRNA products were detected using a Micro spectrophotometer (AllSheng, China), and the fluorescence of CY3/Cy5/Cy7/FAM/Mant was analyzed using a TECAN (INFINITE E PLEX, Switzerland) reader in the fluorescence mode. The mRNA transcripts synthesized by in vitro transcription for the subsequent experiments include luciferase mRNA, eGFP mRNA, and HBs183 mRNA. The transcript lengths and template details are shown in SI S11.
000 rpm for 8 min and washed three times with icy 75% ethyl alcohol. The precipitate was dissolved in RNase-free water. The concentrations of labelled mRNA products were detected using a Micro spectrophotometer (AllSheng, China), and the fluorescence of CY3/Cy5/Cy7/FAM/Mant was analyzed using a TECAN (INFINITE E PLEX, Switzerland) reader in the fluorescence mode. The mRNA transcripts synthesized by in vitro transcription for the subsequent experiments include luciferase mRNA, eGFP mRNA, and HBs183 mRNA. The transcript lengths and template details are shown in SI S11.
| Cap analog | Exciting light (nm) | Emitted light (nm) |
|---|---|---|
| LZCap®AG (3′-Ma-Cy3) | 535 | 570 |
| LZCap®AG (3′-Ma-Cy5) | 640 | 675 |
| LZCap®AG (3′-Ma-Cy7) | 651 | 780 |
| LZCap®AG (3′-Ma-Peg5-FAM) | 480 | 520 |
| LZCap®AG (3′-Ma-C6-MANT) | 356 | 448 |
mRNA capping rate analysis
First, 100 pmol of capped mRNA was annealed with 10 µM biotinylated probes (5′-dCdGdCdTAGCCCAGCUUGGGUCUCCU-biotin-3′) in a 5 × hybrid RNA degeneration buffer in a 100 µL reaction volume. Following annealing, 3 µL of RNase H (180 U) was added, and the mixture was incubated at 37 °C for 3 h. Streptavidin magnetic beads (80 µL, NEB) were washed three times with a TES buffer and then incubated with the reaction product (100 µL) at room temperature for 1 h under shaking. RNA fragments containing the 5′ cap were captured by the probes and magnetic beads, and then separated from the uncaptured material using a magnetic rack. For elution, 80 µL of preheated elution buffer was added, and the sample was incubated at 80 °C for 3 min. The eluted samples were stored at −20 °C or −80 °C until LC-MS analysis (Thermo Fisher, USA). Chromatographic separation was performed on a DNAPac RP column (4 µm, 2 × 50 mm; Waters, USA) maintained at 60 °C. Mobile phase A consisted of 2% hexafluoroisopropanol (HFIP) and 0.1% diisopropylethylamine (DIPEA) in water, while mobile phase B was composed of 0.075% HFIP and 0.0375% DIPEA in methanol. Gradient elution was carried out at a flow rate of 0.3 mL min−1 over a total run time of 20 min. UV detection was set at 260 nm. Mass spectrometry was conducted using an electrospray ionization (ESI) source in the negative ion mode. The following parameters were applied: sheath gas flow rate, 40; auxiliary gas flow rate, 10; spray voltage, 3.3 kV; capillary temperature, 320 °C; lens voltage, 60; and auxiliary gas temperature, 350 °C. Data were acquired over a scan range of 600–3000 m/z at a resolution of 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500.
500.
Preparation of conjugated lipid nanoparticles (LNPs)
Lipid nanoparticles (LNPs) were employed to encapsulate the mRNA using a microfluidic chip (FluidicLab, China) and a pump (DK Infusetek, China). The lipid components, SM102, Cholesterol, DSPC, DMG-PEG2K and DSPE-PEG2K-MAL (all obtained from Sinopeg, China), were dissolved in ethanol at a molar ratio of 48.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 38.7
38.7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 11.1
11.1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.5
1.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.2, while the Cy5/Cy7-labelled mRNA was prepared in a filtered acetate buffer (25 mM, pH 4.0). The two phases were combined at an aqueous-to-organic ratio of 3
0.2, while the Cy5/Cy7-labelled mRNA was prepared in a filtered acetate buffer (25 mM, pH 4.0). The two phases were combined at an aqueous-to-organic ratio of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v). Following encapsulation, the LNP-mRNA mixture was promptly placed into a sealed dialysis cassette to remove residual citrate salt and ethanol. The formulation was then concentrated using a 30 kDa ultrafiltration unit (Millipore, USA) and kept at 4 °C. The particle size and polydispersity index (PDI) were measured via dynamic light scattering (Malvern PANalytical Zetasizer Lab, USA), and both the encapsulation efficiency and the mRNA concentration were determined using a Quant-iT RiboGreen RNA Assay Kit (Invitrogen, USA).
1 (v/v). Following encapsulation, the LNP-mRNA mixture was promptly placed into a sealed dialysis cassette to remove residual citrate salt and ethanol. The formulation was then concentrated using a 30 kDa ultrafiltration unit (Millipore, USA) and kept at 4 °C. The particle size and polydispersity index (PDI) were measured via dynamic light scattering (Malvern PANalytical Zetasizer Lab, USA), and both the encapsulation efficiency and the mRNA concentration were determined using a Quant-iT RiboGreen RNA Assay Kit (Invitrogen, USA).
For antibody conjugation, anti-CD3 antibodies (Bioxcell, USA) or anti-CD7 antibodies (Biointron, China) were first thiolated using Traut's reagent (Thermo Scientific, USA) following the standard protocols. Briefly, antibodies were incubated with a 100-fold molar excess of Traut's reagent at 37 °C for 2 hours, followed by desalting using Zeba spin desalting columns (Sigma Aldrich, Germany), and the thiol-modified antibody solution was collected. Separately, mRNA-LNPs were incubated with modified antibodies at a molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (maleimide
1 (maleimide![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) thiol) at 4 °C for 2 hours. The reaction mixture was purified using a HiTrap column (Cytiva, USA), followed by the collection of the purified sample. The characterization of all LNP samples is shown in SI S12.
thiol) at 4 °C for 2 hours. The reaction mixture was purified using a HiTrap column (Cytiva, USA), followed by the collection of the purified sample. The characterization of all LNP samples is shown in SI S12.
Fluorescence-labelled luciferase mRNA transfection
HEK293T cells were inoculated into a 96-well plate at a density of 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 cells per mL with 125 µL DMEM per well for 18–36 h in a CO2 incubator. The transfection reagents of jetMESSENGER® (Polyplus, France) were mixed according to a system of 10.5 µL jetMESSENGER buffer, 1 µL jetMESSENGER and 0.5 µg fluorescence-labelled luciferase mRNAs and then incubated at RT for 15 min before addition into each well and kept in a CO2 incubator for 24 h. After transfection, half or equal volume of Steady-Glo® Luciferase Assay System (Promega, USA) was added per well, incubated at RT for 5 min before transferring 100–200 µL to a black culture vessel and analyzed using a TECAN (INFINITE E PLEX, Switzerland) reader with luminescence mode.
500 cells per mL with 125 µL DMEM per well for 18–36 h in a CO2 incubator. The transfection reagents of jetMESSENGER® (Polyplus, France) were mixed according to a system of 10.5 µL jetMESSENGER buffer, 1 µL jetMESSENGER and 0.5 µg fluorescence-labelled luciferase mRNAs and then incubated at RT for 15 min before addition into each well and kept in a CO2 incubator for 24 h. After transfection, half or equal volume of Steady-Glo® Luciferase Assay System (Promega, USA) was added per well, incubated at RT for 5 min before transferring 100–200 µL to a black culture vessel and analyzed using a TECAN (INFINITE E PLEX, Switzerland) reader with luminescence mode.
Flow cytometry analysis
HEK293 T cells and Jurkat T cells were transferred to white non-binding 96-well plates. HEK293T cells were transfected with LZCap®AG (3′-Acm) capped/Cy5-labelled luciferase mRNA by jetMESSENGER® (Polyplus, France) for 4 h before the flow cytometry analysis of Cy5 signals. Jurkat T cells were transfected with Cy5-labelled HBs183TCR (ZL 2023 1 1265442.4) mRNA encapsulated in anti-CD3/CD7-conjugated LNP. After incubation at 37 °C for 24 h, the cells were centrifuged at 500×g for 5 minutes at 4 °C and washed twice with cold PBS supplemented with 2% FBS (FACS buffer). Cy5 signals of 293T cells and Jurkat T cells were analyzed in an APC channel. Jurkat T cells were incubated with FITC-conjugated anti-human TCR β chain antibodies (BioLegend, USA) for 30 minutes at 4 °C in the dark. After staining, the cells were washed twice with a FACS buffer and resuspended in 300–500 µL FACS buffer for acquisition. Cy5 fluorescence was excited with a 640 nm laser and detected using a 675/25 nm bandpass filter (APC channel). Data were collected using a BD Accuri C6 Plus flow cytometer (BD Biosciences, USA), and analysis were performed using the FlowJo v10 software.Confocal imaging
Before transfection with Cy5-labelled mRNAs-LNP, 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Huh7 cells were cultured in a 35 mm confocal-specialized disk for 18–24 h. Then, 1 µg of Cy5-luciferase-LNP or Cy5-eGFP-LNP was added into Huh7 cells and incubated for 30 min, 1 h, 3 h, 6 h and 24 h, respectively, under 5% CO2 at 37 °C. After washing three times with PBS, the cells were fixed with 4% paraformaldehyde at room temperature for 15 minutes, followed by DAPI staining for 2–3 minutes. Again, the cells were washed three times with PBS before imaging. For colocalization confocal observation with a lysosome marker, 100 µM of lysosome marker, Green DND-26 (MCE, USA), was added into Huh7 cell media before incubation for 1 h under 5% CO2 at 37 °C and fixed with 4% paraformaldehyde. All confocal imaging studies were performed using an automated spinning disc confocal microscope (Zeiss 2012, Germany). The Z-stack of three image planes, namely, red, green and blue was recorded with one confocal plane.
000 Huh7 cells were cultured in a 35 mm confocal-specialized disk for 18–24 h. Then, 1 µg of Cy5-luciferase-LNP or Cy5-eGFP-LNP was added into Huh7 cells and incubated for 30 min, 1 h, 3 h, 6 h and 24 h, respectively, under 5% CO2 at 37 °C. After washing three times with PBS, the cells were fixed with 4% paraformaldehyde at room temperature for 15 minutes, followed by DAPI staining for 2–3 minutes. Again, the cells were washed three times with PBS before imaging. For colocalization confocal observation with a lysosome marker, 100 µM of lysosome marker, Green DND-26 (MCE, USA), was added into Huh7 cell media before incubation for 1 h under 5% CO2 at 37 °C and fixed with 4% paraformaldehyde. All confocal imaging studies were performed using an automated spinning disc confocal microscope (Zeiss 2012, Germany). The Z-stack of three image planes, namely, red, green and blue was recorded with one confocal plane.
In vivo imaging
BALB/c mice were dosed with 1 mg kg−1 or 0.5 mg kg−1 of Cy7/Cy5-Luciferase-LNP through tail vein injection, while the control mice were not dosed. Cy7/Cy5 fluorescence was measured for the whole body and organs at 30 min, 3 h, 6 h and 24 h using an IVIS Lumina system (PerkinElmer, USA). For each time point, the liver, spleen, heart, lung, kidney, bladder, uterus, intestines and stomach were isolated for Cy7/Cy5 measurement for ex vivo imaging. The wavelengths used for the detection of Cy7 and Cy5 are Ex/Em = 745 nm/800 nm and Ex/Em = 640 nm/680 nm, respectively. The expression of luciferase was tested in the luminescence mode 15 min after P. O. injection of D-luciferin sodium (Bidepharm, China) at 3 hours post-injection of Cy7/Cy5-Luciferase-LNP. The quantitative results of fluorescence intensity were obtained from the images using the Living Image®4.4 software by calculating the average radiance (photon per s1 per cm2 per sr1) in the region of interest and analyzed using GraphPad Prism 8.0.1. All animal studies were conducted under the animal care and use regulations of the Guangzhou Institute of Biomedicine and Health, Chinese Academy of Sciences, with approval from the Institutional Animal Care and Use Committee (IACUC; approval no. 2024096).Results
Synthesis of site-specifically labelled trinucleotide cap analogs
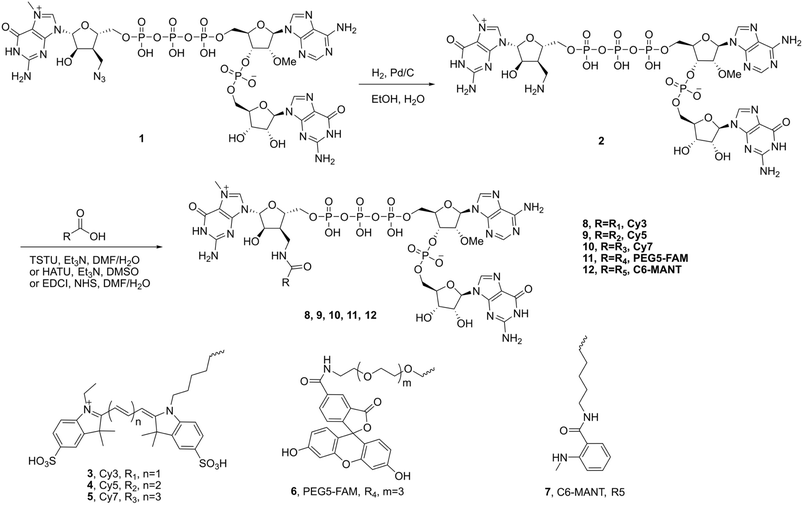
The syntheses of various labelled trinucleotide cap analogs were started with a trinucleotide cap analog 1, which bears an azidomethyl moiety at the 3′ ribose position of the m7G cap replacing the canonical OH group.22 The hydrogenation of analog 1 with Pd/C in EtOH/H2O gave the aminomethyl compound 2 in 93% isolated yield. Subsequent conjugation of compound 2 with various tagging molecules was conducted through different amide formation reactions to afford the labelled trinucleotide cap analogs with TSTU and Et3N in DMF/H2O for 8 (Cy3), 9 (Cy5), and 10 (Cy7) in 21–30% isolated yields, while HATU, Et3N, and DMSO were utilized for 11 (PEG5-FAM) in 9% isolated yield, and EDCI, NHS, and DMF/H2O were used for 12 (C6-MANT) in 31% isolated yield, as illustrated in Scheme 1.One-pot co-transcriptional synthesis of fluorescence-labelled mRNAs
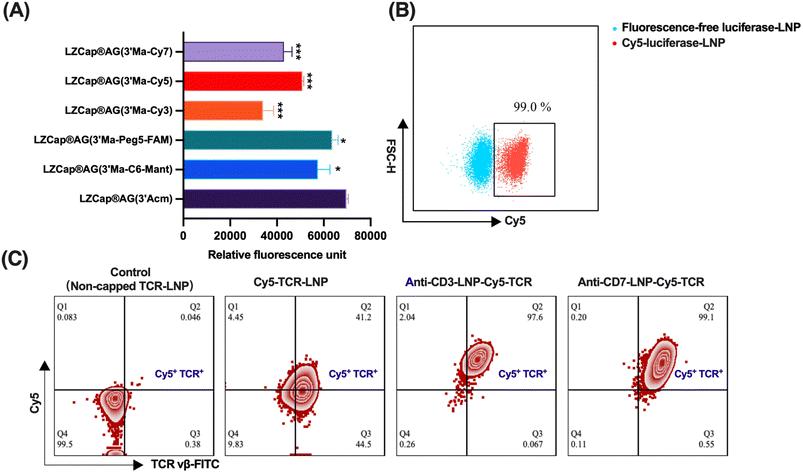
The labelled cap1 mRNA was synthesized using a standard one-pot in vitro transcription procedure. The m7G-capped trinucleotide cap analogs 8, 9, 10, 11, and 12 were co-transcribed with T7 polymerase, NTPs and DNA template encoding firefly luciferase protein in vitro. The average yields and capping efficiencies of capped luciferase mRNAs are summarized in Table 2, showing that both parameters were generally maintained at levels comparable to those obtained with the widely used reference cap structure CleanCap. Fluorescence spectroscopy was employed to measure the fluorescence of labelled luciferase mRNAs (Fig. 1A), and the fluorescence intensity increased with the increasing concentrations of capped mRNA. The integrity of capped mRNAs was analyzed by agarose gel electrophoresis (Fig. 1B), and all mRNAs showed high integrity and accuracy. This streamlined approach is efficient and reproducible by eliminating the need for post-transcriptional labeling.| Cap 1 analogs | CleanCap®AG (3′-OMe) | LZCap®AG (3′-Acm) | LZCap®AG (3′-Ma-fluorescence) | ||||
|---|---|---|---|---|---|---|---|
| Cy3 | Cy5 | Cy7 | Peg5-FAM | C6-MANT | |||
| mRNA yields (mg mL−1) | 5.0 | 5.0 | 4.3 | 5.3 | 7.1 | 5.9 | 5.3 |
| mRNA capping efficiency | 92% | 98% | 79% | 82% | Not detected | 100% | 90% |
Fluorescent cap analogs enable the real-time tracking of mRNA delivery and translation
To evaluate the translational function of mRNAs labelled with fluorescent cap analogs, HEK-293T cells were transfected with firefly luciferase-encoding mRNAs containing these modified caps. Although the fluorescence intensity of the labelled analogs was modestly reduced compared to the unlabelled LZCap®AG (3′-Acm) control, it remained robust, indicating that the labelling moieties did not compromise translational capacity (Fig. 2A). To investigate the intracellular delivery kinetics of LNPs, Cy5-labelled mRNA was introduced to enable the real-time monitoring of cellular transfection. This approach allowed the systematic quantification of delivery efficiency for LNPs and other carriers via flow cytometry. Strikingly, over 95% of HEK-293T cells exhibited efficient internalization of Cy5-capped mRNA within 4 hours post-transfection, validating this strategy for the dynamic tracking of mRNA delivery (Fig. 2B).We further engineered antibody-conjugated LNPs (anti-human CD3/CD7) for targeted mRNA delivery. A Cy5-labelled TCR mRNA was synthesized by IVT using LZCap®AG (3′-Ma-Cy5). Flow cytometry demonstrated that antibody-LNP complexes markedly enhanced both the uptake of Cy5-labelled mRNA and TCR expression in Jurkat T cells. Critically, the overlap of Cy5 fluorescence and TCR expression within the same cell population indicated that Cy5 fluorescence serves as a quantitative proxy for assessing the impact of antibody conjugation on LNP delivery performance (Fig. 2C). Collectively, these findings underscore the utility of fluorescent cap analogs as a dual-functional tool for the real-time monitoring and optimization of precision mRNA delivery platforms.
Translation and intracellular localization of Cy5 cap analog-labelled mRNAs
Cy5-labelled mRNA encoding luciferase or eGFP serves as a versatile dual-purpose tool for simultaneously probing the translational activity and intracellular trafficking dynamics. Upon LNP-mediated delivery into Huh7 cells, confocal microscopy revealed imminent cytoplasmic accumulation of Cy5-labelled mRNA as early as 30 minutes post-delivery. Time-course analysis further demonstrated progressive co-localization of the fluorescent mRNA within the lysosomal compartments (Fig. 3A). Importantly, the parallel detection of eGFP-derived green fluorescence by confocal imaging confirmed that Cy5 capping retains the translational competence of the mRNA, with no observable interference from the fluorophore (Fig. 3B). These findings collectively establish Cy5-labelled mRNA as a reliable platform for the quantitative in vitro evaluation of LNP-based mRNA delivery systems.In vivo tracing of Cy7 and Cy5 cap analog-labelled luciferase mRNAs
To explore the biodistribution and real-time tracking potential of labelled mRNAs in vivo, Cy7 and Cy5-Luciferase-LNPs were administered to mice via intravenous injection, with continuous monitoring of live animals and their isolated organs using fluorescence imaging. The in vivo tracing of Cy7 fluorescence showed that LNP-delivered mRNA was distributed widely in the liver, stomach and intestines initially but mostly accumulated in the liver after 24 hours (Fig. 4A). Notably, luciferase expression was detected in the liver as early as 3 hours post-injection, confirming the translational function of Cy7-labelled mRNA (Fig. 4B). Autopsy at different time points further confirmed the distribution profile of the formulation with a pronounced shift toward hepatic and splenic accumulation 24 hours post-injection (Fig. 4C). Consistently, Cy5-luciferase-LNPs exhibited a comparable biodistribution pattern, predominantly localizing to the liver and spleen at early time points (Fig. 4D and E), with hepatic luciferase activity emerging synchronously at 3 hours (Fig. 4F). These data showed visible fluorescence signals in the Cy7- and Cy5-luciferase mRNA-LNP-treated groups relative to the control group, supporting the feasibility of qualitative tracking under the present experimental conditions. Collectively, the results indicate that Cy7- and Cy5-labelled mRNAs serve as highly sensitive tracers for the spatiotemporal mapping of organ-specific mRNA delivery and expression.Discussion
The 5′ cap of mRNA has been shown to be a viable modification site for mRNA labeling, because it enables site-specific modification while minimizing the perturbation of the coding region and overall transcript architecture. While existing strategies for synthesizing modified cap analogs are well-established,23–25 many of these approaches still depend on post-transcriptional modification or additional bioorthogonal reactions, thereby increasing experimental complexity and limiting their practicality for the routine generation of trackable mRNAs. In this study, we developed fluorescently labelled LZCap®AG through amide linkage chemistry and integrated it into mRNA via a streamlined “one-pot” IVT protocol. This method produced intact mRNA transcripts with good IVT yields and clear detectable fluorescence signals, making it a simple and practical approach for preparing labeled mRNAs.An important finding of this study is that fluorescence modification at the cap was compatible with the biological activity of the mRNA. Although the capping efficiency of labelled LZCap®AG caps was marginally reduced relative to the reference cap, these labelled mRNAs retain activities in transfection and translation, as demonstrated by luciferase expression in HEK-293T cells (Fig. 2A). Notably, the Cy5-labeled mRNA showed a clear fluorescence signal while still retaining translational activity, which makes it useful for tracking delivery and evaluating function at the same time. This could be an advantage compared with the methods based on dense internal nucleotide labeling, because those methods are more likely to affect the RNA structure, transcription, or translation.
The cellular experiments further showed that Cy5-labeled mRNA could be used to evaluate lipid screening and the formulation of mRNA delivery efficiency. Flow cytometry results showed the efficient uptake of Cy5-labeled luciferase mRNA in HEK-293T cells (Fig. 2B). In Jurkat T cells, the Cy5 signal was also detected together with TCR expression after the delivery of Cy5-labeled TCR mRNA by anti-CD3/CD37-conjugated LNPs (Fig. 2C). These results suggest that the fluorescence from the cap label can be used as an indicator of mRNA delivery, while protein expression can be used to evaluate its function at the same time. Consistently, confocal imaging showed rapid intracellular accumulation of Cy5-labeled mRNA and its progressive colocalization with lysosomes. Meanwhile, eGFP expression confirmed that the labeled mRNA still retained translational activity after delivery. Previous studies investigated the dissociation between LNP uptake mechanisms and endosomal escape efficiency, using eGFP/Cy5 signal ratios to quantify the escape efficiency.26 Here, we extended this paradigm by correlating Cy5-labelled LZCap®AG localization with real-time eGFP expression, validating its dual utility for spatial tracking and functional delivery assessment.
The in vivo biodistribution studies further showed the potential of this approach. Both Cy5- and Cy7-labeled mRNAs could be used to visualize biodistribution after LNP administration, with the main signals found in the liver and spleen, and luciferase expression in the liver was also detected 3 h after injection. It should be noted that high-intensity fluorescence signals are essential to overcome tissue autofluorescence. LZCap®AG labelled with Cy5 was disturbed by autofluorescence of alfalfa in mice feed, which also has a λmax value approaching 652 nm. For the usage of Cy5 label, we fed mice with alfalfa-free foods one week before IVIS observation and obtained clear Cy5 signals both in the body and in the organs of animals. Cy7-labelled LZCap®AG, which has a longer conjugated chain and a more red-shifted optical window (λmax/λem = 755/788 nm), showed a clear advantage for whole-body imaging by reducing the background noise from animal feed. To improve mRNA delivery and organ targeting efficiency, the selective organ targeting (SORT) LNP platform delivers mRNA into other target organs,27 like the spleen28,29 or lung.30 Here, ex vivo organ observation showed that the fluorescence of Cy5- andCy7-labelled LZCap®AG can be detected in different organs, including the liver, spleen, lung, heart, kidney, stomach, intestines, bladder and ovary, indicating that Cy5 and Cy7 are also suitable for biodistribution tracing in organs.
Based on the above-mentioned results, fluorescent cap labeling shows clear advantages for both real-time in vitro and in vivo mRNA tracking studies. Previous studies have reported that fluorescent oligomer tags applied for labeling endogenous RNA, such as ISH31 and the MS2 system,32 are useful for RNA detection or tracking, but they are often limited by live-cell imaging applicability, complicated probe design or delivery, or high background noise. In contrast, fluorescent cap analogs circumvent these issues by preserving mRNA integrity while enabling real-time tracking.
This study also has limitations. For the Cy7-labeled analog, the capping efficiency could not be accurately measured by LC-MS under the current analytical conditions. In addition, although fluorescence localization together with protein expression can provide useful information on mRNA delivery, this approach still cannot directly measure the endosomal escape efficiency. Future work should combine this labeling strategy with more quantitative trafficking analysis to further assess its value in delivery optimization.
In summary, we reported a series of labelled LZCap®AG trinucleotide analogs for the one-pot synthesis of labelled and functional cap1 mRNA with high yield, capping rate and mRNA integrity. In particular, with the high IVT yield, rapid transfection kinetics, and uncompromised translational activity, the Cy5- and Cy7-labelled mRNAs obtained from the corresponding Cy5- and Cy7-labelled LZCap®AG trinucleotide cap analogs can serve as useful and valuable tools for in vitro and in vivo mRNA tracking and the evaluation and optimization of LNP-targeted delivery.
Author contributions
J. C. Zhang and J. L. Hou: conceptualization, methodology, supervision, and experimental design; C. Yang, C. Guo, Y. Q. Zhou and S. Y. Li: investigation; C. Yang, C. Guo and J. C. Zhang: writing original draft; C. Yang, C. Guo and M. J. Li: data curation, formal analysis, and visualization; X. Y. Lu, Y. H. Liu and W. Zhu: data curation and formal analysis; and Y. L. Xie, L. J. Zhang and G. H. Mo: review and editing.Conflicts of interest
The authors declare no competing interests.Data availability
The data supporting the findings of this study are available in the article.Supplementary information (SI): general chemical synthesis information and the characterization of the analogs and LNPs. See DOI: https://doi.org/10.1039/d6ra00979d.
Acknowledgements
We acknowledge the support from the National Key Research and Development Program of China (2022YFC2303600) and the China Postdoctoral Science Foundation (2024M75316). The graphical abstract was created with BioRender.com.References
- S. Qin, X. Tang, Y. Chen, K. Chen, N. Fan, W. Xiao, Q. Zheng, G. Li, Y. Teng, M. Wu and X. Song, Signal Transduction Targeted Ther., 2022, 7, 166 CrossRef CAS PubMed.
- N. Pardi, M. J. Hogan, F. W. Porter and D. Weissman, Nat. Rev. Drug Discovery, 2018, 17, 261–279 CrossRef CAS PubMed.
- P. R. Cullisand and M. J. Hope, Mol. Ther., 2017, 25, 467–1475 Search PubMed.
- S. R. Kushner, J. Bacteriol., 2002, 184, 4658–4665 CrossRef CAS PubMed , 4657.
- J. Yu J and J. E. Russell, Mol. Cell. Biol., 2001, 21, 5879–5888 CrossRef PubMed.
- H. Kempe, A. Schwabe, F. Crémazy, P. J. Verschure and F. J. Bruggeman, Mol. Biol. Cell., 2015, 26, 797–804 CrossRef PubMed.
- J. Mattay, M. Dittmar and A. Rentmeister, Curr. Opin. Chem. Biol., 2021, 63, 46–56 CrossRef CAS PubMed.
- M. Warminski, A. Mamot, A. Depaix, J. Kowalska and J. Jemielity, Acc. Chem. Res., 2023, 56, 2814–2826 CrossRef CAS PubMed.
- M. Warminski, P. J. Sikorski, J. Kowalska and J. Jemielity, Top. Curr. Chem., 2017, 375, 16 CrossRef PubMed.
- C. Larsson, I. Grundberg, O. Söderberg and M. Nilsson, Nat. Methods, 2010, 7, 395–397 CrossRef CAS PubMed.
- S. Tyagi and F. R. Kramer, Nat. Biotechnol., 1996, 14, 303–308 CrossRef CAS PubMed.
- J. S. Paige, K. Y. Wu and S. R. Jaffrey, Science, 2011, 333, 642–646 CrossRef CAS PubMed.
- M. N. Mwangi and N. J. Baird, MethodsX, 2024, 13, 102925 CrossRef CAS PubMed.
- K. E. Lindsay, S. M. Bhosle, C. Zurla, J. Beyersdorf, K. A. Rogers, D. Vanover, P. Xiao, M. Araínga, L. M. Shirreff, B. Pitard, P. Baumhof, F. Villinger and P. J. Santangelo, Nat. Biomed. Eng., 2019, 3, 371–380 CrossRef CAS PubMed.
- A. L. Quillin, D. B. Karloff, T. M. Ayele, T. F. Flores, G. Chen, Z. T. McEachin, A. N. Valdez-Sinon and J. M. Heemstra, ACS Chem. Biol., 2025, 20, 707–720 CrossRef CAS PubMed.
- L. Anhäuser, S. Hüwel, T. Zobel and A. Rentmeister, Nucleic Acids Res., 2019, 47, e42 CrossRef PubMed.
- S. C. Alexander, K. N. Busby, C. M. Cole and S. R. Jaffrey, J. Am. Chem. Soc., 2015, 137, 12756–12759 CrossRef CAS PubMed.
- T. Baladi, J. R. Nilsson, A. Gallud, E. Celauro, C. Gasse, F. Levi-Acobas, I. Sarac, M. R. Hollenstein, A. Dahlén, E. K. Esbjörner and L. M. Wilhelmsson, J. Am. Chem. Soc., 2021, 143, 5413–5424 CrossRef CAS PubMed.
- A. Mamot, P. J. Sikorski, M. Warminski, J. Kowalska and J. Jemielity, Angew. Chem., Int. Ed., 2017, 56, 15628–15632 CrossRef CAS PubMed.
- J. M. Holstein, L. Anhäuser and A. Rentmeister, Angew. Chem., Int. Ed., 2016, 55, 10899–10903 CrossRef CAS PubMed.
- S. A. Dilliard, Q. Cheng and D. J. Siegwart, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2109256118 CrossRef CAS PubMed.
- J. Zhang, L. Zhang and Y. Zhou, US Pat., 12060385, 2024.
- Y. Inoue, K. Izawa, S. Kiryu, M. Watanabe, Y. Tojo, H. Ochi and Y. Hiraoka, Mol. Imaging, 2008, 7, 21–27 CrossRef PubMed.
- M. Ziemniak, M. Szabelski, M. Łukaszewicz, P. J. Sikorski, J. Kowalska and J. Jemielity, RSC Adv., 2013, 3, 42769 RSC.
- S. Croce, S. Serdjukow, T. Carell and T. Frischmuth, ChemBioChem, 2020, 21, 1641–1646 CrossRef CAS PubMed.
- N. Aliakbarinodehi, A. Gallud, M. Mapar, E. Wesén, S. Heydari, Y. Jing, G. Emilsson, K. Liu, A. Sabirsh, V. P. Zhdanov, L. Lindfors, E. K. Esbjörner and F. Höök, ACS Nano, 2022, 16, 20163–20173 CrossRef CAS PubMed.
- Q. Cheng, T. Wei, L. Farbiak, L. T. Johnson, S. A. Dilliard and D. J. Siegwart, Nat. Nanotechnol., 2020, 15, 313–320 CrossRef CAS PubMed.
- P. S. Kowalski, U. Capasso Palmiero, Y. Huang, A. Rudra, R. Langer and D. G. Anderson, Adv. Mater., 2018, 30, 1801151 CrossRef PubMed.
- D. Zhang, E. N. Atochina-Vasserman, J. Lu, D. S. Maurya, Q. Xiao, M. Liu, J. Adamson, N. A. Ona, E. K. Reagan, H. Ni, D. Weissman and V. Percec, J. Am. Chem. Soc., 2022, 144, 4746–4753 CrossRef CAS PubMed.
- M. Qiu, Y. Tang, J. Chen, R. Muriph, Z. Ye, C. Huang, J. Evans, E. P. Henske and Q. Xu, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2116271119 CrossRef CAS PubMed.
- J. Ouellet, Front. Chem., 2016, 4, 29 Search PubMed.
- D. S. Peabody, EMBO J., 1993, 12, 595–600 CrossRef CAS PubMed.
Footnote |
| † Chou Yang, Chen Guo, and Yiqian Zhou contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |