Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Synthesis and anti-proliferative activity of new E7010 tethered urea congeners as potential tubulin inhibitors and apoptosis inducers
Shaik Tajab,
Ganga Reddy Velma *cd,
Srinivasa Reddy Telukutla
*cd,
Srinivasa Reddy Telukutla de,
Satyaveni Malasalaf,
Anjali Sharmag,
Irfan Khanac,
Mohd Adil Shareefab,
Suresh K. Bhargava
de,
Satyaveni Malasalaf,
Anjali Sharmag,
Irfan Khanac,
Mohd Adil Shareefab,
Suresh K. Bhargava d,
Magdalena Plebanskie,
Bathini Nagendra Babuab and
Ahmed Kamal
d,
Magdalena Plebanskie,
Bathini Nagendra Babuab and
Ahmed Kamal *ah
*ah
aAcademy of Scientific and Innovative Research (AcSIR), CSIR-Human Resource Development Centre (CSIR-HRDC) Campus, Ghaziabad 201 002, Uttar Pradesh, India
bFluoro-Agrochemicals, CSIR-Indian Institute of Chemical Technology (IICT), Hyderabad 500 007, India
cDepartment of Pharmacology and Toxicology, College of Pharmacy, University of Arizona, Tucson 85721, AZ, USA. E-mail: velmagangareddy47@gmail.com; vgreddy@arizona.edu
dCentre for Advanced Materials & Industrial Chemistry (CAMIC), School of Science, RMIT University, GPO Box 2476, Melbourne 3001, Australia
eSchool of Health and Biomedical Sciences, RMIT University, Melbourne, 3083, Australia
fDepartment of Anatomy & Cell Biology, The Brody School of Medicine, East Carolina University, Greenville, NC, USA
gGuru Gobind Singh College of Pharmacy, Yamunanagar (135001), Haryana, India
hDepartment of Pharmacy, Birla Institute of Technology and Science-Pilani, Hyderabad 500078, India. E-mail: ahmedkamal@hyderabad.bits-pilani.ac.in; ahmedkamal915@gmail.com
First published on 19th January 2026
Abstract
A series of twenty-four 1-phenyl-3-(2-phenylpyridin-3-yl)urea congeners (6a–x) have been designed and synthesized. All these compounds were evaluated for their anti-cancer activity against four human cancer cell lines, including prostate cancer (DU-145), lung cancer (A549), cervical cancer (HeLa) and breast cancer (MDA-MB-231). Compound 6q emerged as the most potent in the series, showing consistently strong cytotoxicity against all tested cell lines, with IC50 values ranging from 2.03 to 8.14 µM. Further investigation into the mechanism of action of compound 6q revealed it can arrest cell cycle progression at the G2/M phase. Immunocytochemistry studies showed a notable disruption of microtubule structure in cells treated with 6q. Molecular docking studies provided strong evidence that compound 6q works by binding to the colchicine binding site of β-tubulin, with sequential hydrophilic and hydrophobic interactions. Additionally, the effects of 6q on cell migration and its ability to induce apoptosis in A549 cells were examined using nuclear and mitochondrial staining techniques.
1 Introduction
Despite extensive efforts in the development of anti-cancer drugs, cancer remains a life-threatening disease.1 Unfortunately, existing anticancer drugs suffer from multiple drawbacks, including serious adverse effects, drug resistance, non-selectivity, and poor bioavailability, which supports the need for the development of new and better drugs.2–4 Microtubules are formed through the polymerization of α–β tubulin heterodimers. They are one of the well-established and validated targets for treating cancers.5 By disrupting the dynamic stability of microtubules, anti-tubulin agents function as spindle poisons, leading to G2/M phase cell cycle arrest and apoptotic cell death.6,7 A wide range of reports in the literature have highlighted that naturally occurring and synthetic hybrids can be potent tubulin agents inhibitors.8–10 Notably, compounds such as Colchicine (I), Combretastatin A-4 (II), Nocodazole (III), and Vinca alkaloids (IVA–IVB) bind to the colchicine binding site on tubulin, inducing depolymerization of microtubule proteins.11 Conversely, Epothilones and Taxanes (V) bind to the taxane binding site, functioning as microtubule polymerization agents or stabilizing agents (Fig. 1). Furthermore, anti-tubulin agents often disrupt tumor vasculature, exhibiting antiangiogenic effects.8On the other hand, E7010 (VI), initially synthesized by Hiroshi Yoshino in 1992, stands as a potent orally active inhibitor of tubulin polymerization. This compound exerts its effect by reversibly binding to the colchicine binding site on tubulin, leading to mitotic arrest.12 Over the past decade, E7010 has emerged as a highly promising entity, characterized by its broad cytotoxicity range, favorable pharmacokinetic profile, and being less susceptible to the target for the development of multidrug resistance.13–16 Furthermore, recent investigations have uncovered the potential of modifying the E7010 compound to yield derivatives with impressive anticancer activity. Our endeavors have been particularly fruitful in harnessing the anti-mitotic capabilities of E7010-based heterocycles for the development of potent antimitotic chemotherapeutics.17 Several of the most promising candidates (VII–XI) are illustrated in Fig. 2.
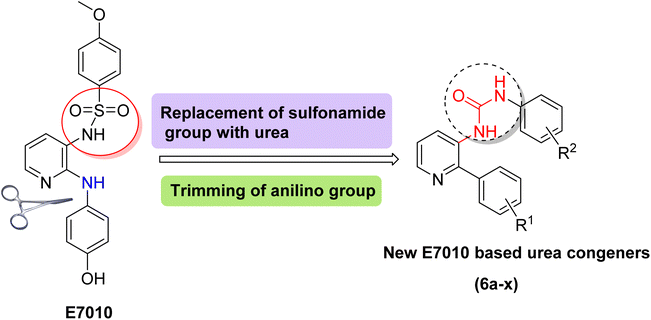
Likewise, the potential of including urea functionality into compounds has been explored, and studies have found that they can exhibit significant anticancer activity, potentially via tubulin inhibition.18–21 Given the remarkable anticancer capabilities observed in E7010 and urea derivatives, and as part of our ongoing efforts to advance the synthesis of novel chemotherapeutic agents,22 our focus was directed toward the synthesis of compounds: E7010-linked urea conjugates. This was achieved by replacing the sulphonamide group in E7010 with an aryl urea motif while introducing phenyl substituents in place of the 2-anilino group (as depicted in Fig. 3). Subsequently, these newly designed compounds were evaluated for their anticancer potential against four human cancer cell lines. Furthermore, comprehensive biological investigations were conducted, with particular emphasis on unraveling the mechanistic properties of the potent compound 6q against A549 cells.
2 Results and discussion
2.1. Chemistry
New phenyl-3-(2-phenylpyridin-3-yl)urea congeners (6a–x) were synthesized as described in Scheme 1. The key intermediates were prepared in two sequential steps. Initially, 2-chloro-3-nitropyridine was subjected to Suzuki coupling conditions with substituted boronic acids at 100 °C for 8 h to furnish substituted 3-nitro-2-phenylpyridines (3a–f), which upon reduction in the presence of H2–Pd/C in EtOAc at room temperature, yield the pre-final amines (4a–f). Lastly, the obtained pre-final amine (4a–f) intermediates were reacted with substituted phenyl isocyanates (5a–d) to get the desired compounds (6a–x) in good to excellent yields. All the designed and synthesized compounds were appropriately characterized by 1H, 13C NMR and mass spectrometry.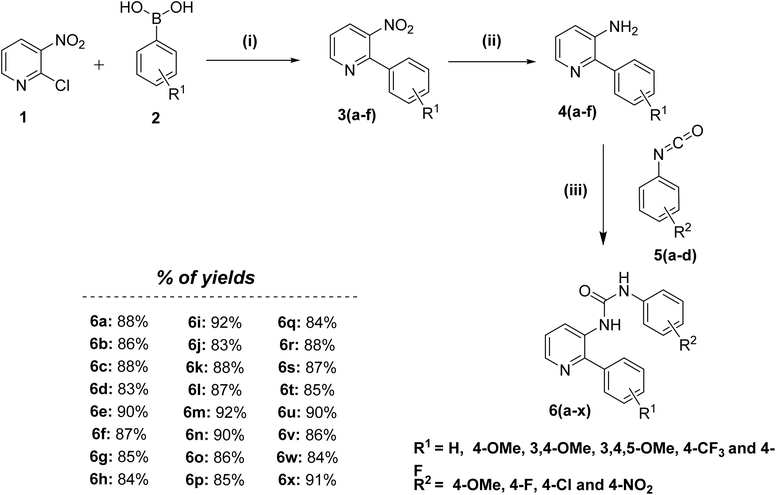 | ||
| Scheme 1 Reagents and conditions: (i) tetrakis(triphenylphosphine)palladium(0), K2CO3, 1,4-dioxane, 100 °C, 8 h, 80–85%; (ii) H2–Pd/C, EtOAc, 0 °C-rt, 8 h, 85–90%; (iii) CHCl3, rt, 12 h, 80–92%. | ||
2.2. In vitro cell growth inhibition properties
The newly synthesized E7010 congeners (6a–x) were examined for their cytotoxic activity against some cancer cell lines, viz., prostate (DU-145), lung (A549), breast (MDA-MB-231), and cervical (HeLa), by employing the MTT assay.23 The IC50 values (µM) are represented in Table 1.| Code | R1 | R2 | DU-145b | A549c | MDA-MB-231d | HeLae |
|---|---|---|---|---|---|---|
| a IC50 values denote the concentrations at which 50% inhibition of the cell growth takes place. Presented data depicts the mean ± standard deviation derived from the three independent studies performed in triplicate.b DU145 – prostate cancer.c A549 – lung cancer.d MDA-MB-231 – breast cancer.e HeLa – vervical cancer cell lines. | ||||||
| 6a | H | 4-OMe | 15.13 ± 1.23 | 9.72 ± 1.21 | >25 | 18.15 ± 2.23 |
| 6b | H | 4-F | 11.5 ± 0.61 | 14.17 ± 1.84 | 16.71 ± 2.11 | 11.37 ± 2.61 |
| 6c | H | 4-Cl | >25 | 20.41 ± 3.14 | >25 | 16.81 ± 1.91 |
| 6d | H | 4-NO2 | >25 | 16.74 ± 1.43 | 12.70 ± 1.83 | >25 |
| 6e | 4-OMe | 4-OMe | 8.14 ± 0.47 | 8.82 ± 0.73 | 9.42 ± 1.12 | 12.13 ± 1.44 |
| 6f | 4-OMe | 4-F | >25 | 11.81 ± 0.97 | 15.62 ± 1.81 | 18.21 ± 2.51 |
| 6g | 4-OMe | 4-Cl | 15.72 ± 1.86 | 8.46 ± 1.13 | >25 | 16.59 ± 0.42 |
| 6h | 4-OMe | 4-NO2 | 7.47 ± 0.62 | 9.76 ± 0.63 | 19.74 ± 2.46 | 6.95 ± 0.81 |
| 6i | 3,4- OMe | 4-OMe | 21.37 ± 3.44 | 14.36 ± 1.29 | 11.26 ± 1.79 | 15.88 ± 2.62 |
| 6j | 3,4- OMe | 4-F | 10.41 ± 1.53 | 12.53 ± 0.89 | 8.72 ± 1.54 | 10.92 ± 1.82 |
| 6k | 3,4- OMe | 4-Cl | 14.45 ± 1.27 | 20.28 ± 3.81 | >25 | 17.91 ± 1.57 |
| 6l | 3,4- OMe | 4-NO2 | 6.78 ± 0.84 | 6.62 ± 0.77 | 9.34 ± 1.18 | 12.01 ± 1.78 |
| 6m | 3,4,5-OMe | 4-OMe | >25 | 19.15 ± 1.67 | 22.74 ± 3.25 | >25 |
| 6n | 3,4,5-OMe | 4-F | 15.24 ± 1.62 | >25 | 16.71 ± 2.97 | 18.32 ± 2.14 |
| 6o | 3,4,5-OMe | 4-Cl | 11.93 ± 1.75 | >25 | >25 | >25 |
| 6p | 3,4,5-OMe | 4-NO2 | 13.31 ± 2.11 | 11.41 ± 0.92 | 13.40 ± 2.12 | 15.95 ± 1.33 |
| 6q | 4-F | 4-OMe | 3.88 ± 0.57 | 2.03 ± 0.15 | 5.79 ± 0.89 | 8.14 ± 1.12 |
| 6r | 4-F | 4-F | 7.32 ± 0.82 | 8.17 ± 1.02 | 14.45 ± 2.27 | 11.45 ± 0.96 |
| 6s | 4-F | 4-Cl | 14.47 ± 2.04 | 15.42 ± 2.11 | 18.64 ± 2.11 | 18.16 ± 2.31 |
| 6t | 4-F | 4-NO2 | 6.53 ± 0.73 | 7.81 ± 1.67 | 8.91 ± 1.31 | 10.17 ± 1.36 |
| 6u | 4-CF3 | 4-OMe | 21.17 ± 2.66 | >25 | 21.14 ± 1.79 | 13.23 ± 1.78 |
| 6v | 4-CF3 | 4-F | 13.67 ± 1.94 | 18.73 ± 2.54 | 12.67 ± 2.03 | 22.06 ± 3.57 |
| 6w | 4-CF3 | 4-Cl | 17.24 ± 1.42 | 11.42 ± 1.85 | 19.14 ± 2.74 | >25 |
| 6x | 4-CF3 | 4-NO2 | 8.21 ± 1.13 | 11.46 ± 2.17 | 10.51 ± 2.15 | 19.21 ± 2.71 |
| Nocodazole | 2.14 ± 0.19 | 2.39 ± 0.14 | 2.13 ± 0.25 | 3.48 ± 0.52 | ||
| E7010 | 2.23 ± 0.17 | 2.67 ± 0.33 | 2.51 ± 0.41 | 2.88 ± 0.24 | ||
| Cisplatin | 1.87 ± 0.21 | 4.54 ± 0.27 | 6.35 ± 0.51 | 3.3 ± 0.1 | ||
These results suggest that most of the newly prepared molecules exhibited moderate to appreciable cytotoxicity against the screened cell lines. Remarkably, compound 6q (R1 = 4-F and R2 = 4-OMe) exhibited substantial cancer cell growth inhibition against the A549 lung cancer cell line with an IC50 of 2.03 µM, equivalent in potency to the positive controls nocodazole and E7010. Noteworthy inhibitory activities were also observed for compounds 6e, 6h, 6l, 6r, and 6t against the DU-145 and A549 cell lines, with IC50 values ranging from 6.5 to 8.2 µM. Furthermore, compounds 6e, 6j, 6l, 6q, and 6t exhibited moderate growth inhibition against the breast cancer cells (MDA-MB-231), having IC50 values below 10 µM. It is observed that when the R1 substitution is 3,4,5-trimethoxy (6m, 6n, 6o) and 4-trifluoromethyl (6u, 6v, 6w) the cytotoxic activity reduces considerably with most of the R2 functional groups. However, they are moderately active, when the R2 group is NO2 in both the cases (6p and 6x). Thus, to assess selectivity towards cancer cells, the most potent compound, 6q, was tested against non-cancerous HEK-293 cells. Compound 6q, along with the standard E7010, exhibited lower inhibitory effects on HEK-293 cell growth, with IC50 values of 10.35 ± 1.12 and 5.18 ± 0.89 µM, respectively, compared to the tested cancer cell lines. These promising preliminary results encouraged us to investigate this compound (6q) for further detailed biological studies (Fig. 4).
 | ||
| Fig. 4 Summarized SAR map showing the influence of R1 and R2 substituents on anti-proliferative activity. Color coding: green = high (IC50 < 5 µM), yellow = moderate (5–10 µM), red = low (>10 µM). | ||
2.3. Cell cycle analysis
Most cytotoxic agents have a characteristic feature of arresting specific phases of the cell cycle, inducing apoptosis, or a combination of both, resulting in the exertion of growth-inhibitory effects.24 In order to elucidate the mechanism through which compound 6q inhibits cell growth, we conducted a cell cycle analysis on A549 cells. For this purpose, the cells were incubated with the compound 6q at different concentrations, 1 and 2 µM, for 48 h, stained with DNA-binding dye propidium iodide, and were then analyzed by flow cytometry. The results obtained clearly showed the compound's ability to arrest the cell cycle in a concentration-dependent manner (Fig. 5). Notably, treatment with compound 6q led to the accumulation of 24.3% and 30.4% of A549 cells in the G2/M phase after exposure to 1 and 2 µM concentrations, respectively, as compared to the untreated control (14.7%).2.4. Inhibition of tubulin polymerization
Tubulin is a central component of the cytoskeletal network and is essential for various cellular functions, including mitosis. Given that compound 6q induced G2/M phase arrest in the cell cycle analysis, it is plausible that it could impede tubulin polymerization, thereby affecting cellular processes.25 To validate this effect, the variations in the microtubule network in A549 cells by immunohistochemistry studies were incubated with this compound for 48 h with individual concentrations of 1 and 2 µM, and the data obtained was compared with the untreated control and nocodazole as positive control. The results confirmed a well-organized microtubular network in untreated cells, whereas cells treated with 1 and 2 µM of 6q showed disorganized microtubules (Fig. 6), confirming the inhibition of tubulin polymerization. Additionally, the positive control, nocodazole, exhibits the expected pattern of disrupted microtubule organization.2.5. Molecular docking studies
Docking studies were conducted to examine the mode of binding and type of interactions between compound 6q and α/β-tubulin employing the Glide docking module within the Schrödinger suite.26 The α/β-tubulin's crystal structure coordinates were obtained from the RCSB protein data bank (PDB ID: 3E22). As shown in Fig. 7, the molecular docking results suggest that the best conformation of 6q is accommodated well within the tubulin's active site. This molecule exhibited two hydrogen bond interactions with Thr179 and Asn101, which are the crucial active site residues. Specifically, the nitrogen atom of the urea functional group established an interaction of hydrogen bond with the backbone carboxylic acid functionality of Thr179, with a separation of 3.18 Å. Furthermore, the urea moieties oxygen atom is predicted to be engaged in a hydrogen bond interaction with the amino functionality of Asn101 with a separation of 3.79 Å. In addition to these hydrogen bond interactions, it is also engaged in many interactions of hydrophobic nature with the α/β-tubulin's key amino acid residues, encompassing Val181, Ala180, Leu248, Leu255, Met259, Val315, Ala250, Ala316 and Tyr224. Collectively, these findings provide compelling evidence that this compound exhibits an intense affinity for the colchicine binding site of the tubulin protein.2.6. Colony forming assay
The long-term cytotoxic and growth-inhibiting capabilities of compound 6q were investigated using a clonogenic cell survival assay. This assay assesses the ability of individual cells to form colonies over an extended period.25 The untreated control displayed colonies, with dense and large, blue-stained clusters, indicating a strong capacity for proliferative growth. Treatment with 1 µM resulted in a relatively similar number of colonies compared to the control, but with visibly smaller colonies, indicating partial suppression of proliferative capacity. A significant decrease in colony density, with primarily small-sized colonies, was observed when the dose was increased to 2 µM. This suggests that higher treatment significantly hinders the ability of surviving cells to divide multiple times. Increasing the dose to 4 µM, A549 cells displayed the fewest colonies, which were very small and faintly stained, indicating a marked cytotoxic or cytostatic effect, Fig. 8. Overall, results indicate that the treatment with this compound reduced clonogenic survival in a dose-dependent manner. This reduction could be attributed to several mechanisms, including induction of apoptosis, cell cycle arrest, or persistent DNA damage that prevents cell proliferation.2.7. Wound healing assay (migration assay)
Cancer cell migration plays a pivotal role in the metastasis process, while endothelial cell migration is a fundamental requirement for angiogenesis, a crucial step in tumor growth.25 Therefore, we evaluated the antimigratory potential of compound 6q using a wound healing assay. In this experiment, wounds were created in confluent monolayers of endothelial HUVEC cells and subsequently treated these cells with various subtoxic concentrations of this molecule. The cells start migrating within this wounded area, which was then monitored employing a phase-contrast microscope. After a 30-hour observation period, the results revealed a concentration-dependent inhibition of HUVEC cell migration induced by this compound. In the untreated control group, the wounds had nearly closed completely (99.5%). In stark contrast, treatment with this molecule at 0.25 µM resulted in a notable slowdown of cell migration, reducing wound closure to 82.15%. Notably, at 1 µM concentration, it significantly inhibited cell migration, with only 37.7% wound closure observed (Fig. 9).2.8. Apoptosis
Apoptosis is the programmed cell death that occurs due to severe, irreversible damage to the cell's prominent organelles.27 Most of the cytotoxic agents act by inducing apoptosis. Therefore, it was considered essential to understand the apoptosis-inducing property of this molecule (6q) on A549 (lung cancer) cells.3 Conclusion
A series of new 1-phenyl-3-(2-phenylpyridin-3-yl)urea congeners were designed, synthesized, and examined for their cytotoxic activities. Among the synthesized compounds, compound 6q showed significant growth inhibition towards some tumor cell lines with IC50 values ranging from 2.03 to 8.14 µM. It has exhibited potent growth inhibition with an IC50 of 2.03 µM towards A549 cells. Mechanism studies indicated that it arrested the A549 cells in the G2/M phase of the cell cycle and inhibited tubulin assembly, thereby disrupting the microtubule dynamics. Furthermore, molecular docking of this compound offered supporting insights into binding modes and protein-ligand interactions at the colchicine binding pocket of tubulin. Additionally, the induction of apoptosis was confirmed through experiments involving Hoechst staining, assessment of mitochondrial membrane potential (ΔΨm), and assays on Annexin V-FITC in A549 cells. Based on these studies, it can be concluded that this molecule exhibits significant cytotoxic activity by inhibiting tubulin polymerization and inducing apoptosis. Therefore, it appears promising for its further detailed exploration and could be considered as a potential lead for the development of newer chemotherapeutic agents.4 Experimental section
4.1. Chemistry
Starting materials and reagents were purchased from Alfa Aesar and Aldrich and used without further purification. Reactions were monitored by TLC analysis using Merck 9385 aluminium plates pre-coated with silica gel containing F254 indicator, which were visualised using UV light. Column chromatography was performed using Merck silica gel of 60–120 mesh with hexane and ethyl acetate as eluents. Melting points were determined on an electrothermal melting point apparatus and are uncorrected. Nuclear Magnetic Resonance spectra (1H and 13C) were recorded on 300 MHz Bruker spectrometer, using TMS as an internal reference. The chemical shifts values are expressed in parts per million (ppm) and coupling constants (J) in Hertz. Splitting patterns of multiplicities are designated as: s, singlet; d, doublet; t, triplet; q, quartet; dd, doublet of doublet; m, multiplet. High-resolution mass spectra were obtained using an ESI-QTOF mass spectrometer (70 eV).![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to afford compounds 4a–f with yields of 85–90%. The obtained product's NMR data matched those reported in the literature.29
1) to afford compounds 4a–f with yields of 85–90%. The obtained product's NMR data matched those reported in the literature.294.1.3.1. 1-(4-Methoxyphenyl)-3-(2-phenylpyridin-3-yl)urea (6a). White solid, yield 88%, Mp: 172–174 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.62 (s, 1H), 8.49 (d, J = 8.4 Hz, 1H), 8.36–8.30 (m, 1H), 7.69–7.63 (m, 3H), 7.55–7.43 (m, 3H), 7.32 (d, J = 9.0 Hz, 2H), 7.26 (dd, J = 8.3, 4.6 Hz, 1H), 6.82 (d, J = 9.0 Hz, 2H), 3.78 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO) δ 154.76, 153.03, 148.30, 142.97, 137.85, 132.92, 131.92, 128.96, 128.74, 128.29, 127.92, 122.12, 120.32, 113.60, 54.97. MS (ESI): m/z 320 [M + H]+. HRMS (ESI) calcd for C19H17N3O2 [M + H]+, 320.1399; found, 320.1392.
4.1.3.2. 1-(4-Fluorophenyl)-3-(2-phenylpyridin-3-yl)urea (6b). White solid, yield 86%, Mp: 181–183 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.85 (s, 1H), 8.47 (dd, J = 8.4, 1.3 Hz, 1H), 8.35 (d, J = 4.0 Hz, 1H), 7.74 (s, 1H), 7.70–7.64 (m, 2H), 7.56–7.44 (m, 3H), 7.40 (dt, J = 6.9, 4.0 Hz, 2H), 7.27 (dd, J = 8.3, 4.6 Hz, 1H), 6.96 (t, J = 8.7 Hz, 2H). 13C NMR (101 MHz, CDCl3) δ 160.06 (d, JCF = 245 Hz), 153.30, 149.16, 144.15, 137.36, 133.22, 132.69, 129.12, 128.84 (d, JCF = 14.0 Hz), 124.26, 123.17, 116.20 (d, JCF = 22.22 Hz). MS (ESI): m/z 308 [M + H]+. HRMS (ESI) calcd for C18H15FN3O [M + H]+, 308.1199; found, 308.1193.
4.1.3.3. 1-(4-Chlorophenyl)-3-(2-phenylpyridin-3-yl)urea (6c). White solid, yield 88%, Mp: 199–201 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.94 (s, 1H), 8.46 (dd, J = 8.3, 1.3 Hz, 1H), 8.35 (dd, J = 4.6, 1.3 Hz, 1H), 7.79 (s, 1H), 7.70–7.64 (m, 2H), 7.52 (dd, J = 14.8, 7.1 Hz, 3H), 7.40 (d, J = 8.8 Hz, 2H), 7.30–7.24 (m, 1H), 7.21 (d, J = 8.8 Hz, 2H). 13C NMR (75 MHz, CDCl3 + DMSO) δ 152.53, 148.51, 143.29, 137.79, 132.50, 129.21, 128.72, 128.25, 128.11, 127.96, 126.12, 122.03, 119.33. MS (ESI): m/z 324 [M + H]+. HRMS (ESI) calcd for C18H15ClN3O [M + H]+, 324.0903; found, 324.0898.
4.1.3.4. 1-(4-Nitrophenyl)-3-(2-phenylpyridin-3-yl)urea (6d). Yellow solid, yield 83%, Mp: 280–282 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.51 (s, 1H), 8.45 (d, J = 8.3 Hz, 1H), 8.39 (dd, J = 4.6, 1.3 Hz, 1H), 8.14 (d, J = 9.2 Hz, 2H), 7.98 (s, 1H), 7.70–7.66 (m, 2H), 7.64–7.60 (m, 2H), 7.58–7.44 (m, 3H), 7.31 (dd, J = 8.3, 4.6 Hz, 1H). MS (ESI): m/z 335 [M + H]+. HRMS (ESI) calcd for C18H15N4O3 [M + H]+, 335.1144; found, 335.1138.
4.1.3.5. 1-(4-Methoxyphenyl)-3-(2-(4-methoxyphenyl)pyridin-3-yl)urea (6e). White solid, yield 90%, Mp: 195–197 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.64 (s, 1H), 8.47 (d, J = 7.0 Hz, 1H), 8.30 (s, 1H), 7.60 (dd, J = 20.2, 13.1 Hz, 4H), 7.32 (d, J = 7.4 Hz, 2H), 6.93 (dd, J = 64.8, 7.1 Hz, 4H), 3.87 (s, 3H), 3.78 (s, 3H). MS (ESI): m/z 350 [M + H]+. HRMS (ESI) calcd for C20H20N3O3 [M + H]+, 350.1505; found, 350.1499.
4.1.3.6. 1-(4-Fluorophenyl)-3-(2-(4-methoxyphenyl)pyridin-3-yl)urea (6f). White solid, yield 87%, Mp: 201–203 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.81 (s, 1H), 8.47 (dd, J = 8.3, 1.4 Hz, 1H), 8.34 (dd, J = 4.6, 1.4 Hz, 1H), 7.70 (s, 1H), 7.66–7.60 (m, 2H), 7.44–7.37 (m, 2H), 7.24 (dd, J = 8.3, 4.6 Hz, 1H), 7.06 (d, J = 8.8 Hz, 2H), 7.02–6.92 (m, 2H), 3.89 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO) δ 157.81 (d, JCF = 235.5 Hz), 156.25, 152.99, 148.33, 143.20, 135.15, 132.71, 130.20, 128.97, 121.87, 120.01 (d, JCF = 7.5 Hz), 114.91 (d, JCF = 22.5 Hz), 113.81, 55.00. MS (ESI): m/z 338 [M + H]+. HRMS (ESI) calcd for C19H17FN3O2 [M + H]+, 338.1305; found, 338.1129.
4.1.3.7. 1-(4-Chlorophenyl)-3-(2-(4-methoxyphenyl)pyridin-3-yl)urea (6g). White solid, yield 85%, Mp: 217–219 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.94 (s, 1H), 8.44 (dd, J = 8.3, 1.3 Hz, 1H), 8.35–8.30 (m, 1H), 7.76 (s, 1H), 7.61 (d, J = 8.7 Hz, 2H), 7.41 (d, J = 8.9 Hz, 2H), 7.26–7.18 (m, 3H), 7.05 (d, J = 8.7 Hz, 2H), 3.87 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 159.24, 152.61, 148.22, 143.15, 137.74, 132.42, 130.06, 128.90, 128.14, 126.32, 121.71, 119.36, 113.67, 54.85. MS (ESI): m/z 354 [M + H]+. HRMS (ESI) calcd for C19H17ClN3O2 [M + H]+, 354.1009; found, 354.1003.
4.1.3.8. 1-(2-(4-Methoxyphenyl)pyridin-3-yl)-3-(4-nitrophenyl)urea (6h). Yellow solid, yield 84%, Mp: 238–240 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.51 (s, 1H), 8.42 (d, J = 8.2 Hz, 1H), 8.37 (d, J = 4.4 Hz, 1H), 8.15 (d, J = 9.1 Hz, 2H), 7.95 (s, 1H), 7.65 (dd, J = 16.1, 9.1 Hz, 5H), 7.26 (dd, J = 8.2, 4.6 Hz, 1H), 7.06 (d, J = 8.6 Hz, 2H), 3.88 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 159.32, 152.03, 148.65, 145.72, 143.69, 141.11, 131.84, 130.08, 129.31, 125.75, 124.46, 121.68, 117.04, 113.69, 112.22, 54.87. MS (ESI): m/z 365 [M + H]+. HRMS (ESI) calcd for C19H17N4O4 [M + H]+, 365.1250; found, 365.1244.
4.1.3.9. 1-(2-(3,4-Dimethoxyphenyl)pyridin-3-yl)-3-(4-methoxyphenyl)urea (6i). White solid, yield 90%, Mp: 198–200 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.57 (s, 1H), 8.52 (d, J = 8.3 Hz, 1H), 8.31 (d, J = 3.4 Hz, 1H), 7.61 (s, 1H), 7.31 (d, J = 8.9 Hz, 2H), 7.26–7.17 (m, 3H), 7.00 (d, J = 8.2 Hz, 1H), 6.82 (d, J = 8.9 Hz, 2H), 3.94 (s, 3H), 3.91 (s, 3H), 3.78 (s, 3H). 13C NMR (101 MHz, CDCl3): δ 156.96, 153.56, 148.89, 148.77, 147.96, 143.09, 133.39, 130.38, 130.19, 127.24, 123.82, 122.92, 121.07, 114.54, 111.63, 110.93, 55.66, 55.61, 55.48. MS (ESI): m/z 380 [M + H]+. HRMS (ESI) calcd for C21H22N3O4 [M + H]+, 380.1610; found, 380.1610.
4.1.3.10. 1-(2-(3,4-Dimethoxyphenyl)pyridin-3-yl)-3-(4-fluorophenyl)urea (6j). White solid, yield 83%, Mp: 192–194 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.90 (s, 1H), 8.48 (d, J = 8.3 Hz, 1H), 8.32 (d, J = 3.4 Hz, 1H), 7.71 (s, 1H), 7.40 (dd, J = 8.9, 4.8 Hz, 2H), 7.27–7.18 (m, 3H), 7.07–7.01 (m, 1H), 6.97 (t, J = 8.7 Hz, 2H), 3.94 (d, J = 5.2 Hz, 3H), 3.92 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 157.72 (d, JCF = 238.5 Hz), 152.83, 148.62, 148.42, 147.99, 142.91, 134.99, 132.65, 130.25, 128.74, 121.84, 121.32, 119.88 (d, JCF = 7.5 Hz), 114.76 (d, JCF = 21.75 Hz), 111.95, 110.95, 55.48, 55.36. MS (ESI): m/z 368 [M + H]+. HRMS (ESI) calcd for C20H19FN3O3 [M + H]+, 368.1410; found, 368.1405.
4.1.3.11. 1-(4-Chlorophenyl)-3-(2-(3,4-dimethoxyphenyl)pyridin-3-yl)urea (6k). White solid, yield 88%, Mp: 208–210 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.92 (s, 1H), 8.50 (d, J = 8.2 Hz, 1H), 8.34 (d, J = 3.6 Hz, 1H), 7.72 (s, 1H), 7.40 (d, J = 8.7 Hz, 2H), 7.29–7.16 (m, 5H), 7.03 (d, J = 8.1 Hz, 1H), 3.94 (s, 3H), 3.92 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 172.55, 152.61, 148.67, 148.43, 148.10, 143.02, 137.78, 132.55, 130.17, 128.90, 128.17, 126.32, 121.84, 121.36, 119.39, 112.00, 110.99, 55.50, 55.38. MS (ESI): m/z 384 [M + H]+. HRMS (ESI) calcd for C20H19ClN3O3 [M + H]+, 384.1115; found, 384.1109.
4.1.3.12. 1-(2-(3,4-Dimethoxyphenyl)pyridin-3-yl)-3-(4-nitrophenyl)urea (6l). Yellow solid, yield 87%, Mp: 259–261 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.54 (s, 1H), 8.52–8.44 (m, 1H), 8.38 (d, J = 3.6 Hz, 1H), 8.15 (d, J = 9.2 Hz, 2H), 7.93 (s, 1H), 7.62 (d, J = 9.2 Hz, 2H), 7.31–7.20 (m, 3H), 7.05 (d, J = 8.2 Hz, 1H), 3.95 (s, 3H), 3.92 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 152.09, 148.75, 148.48, 148.30, 145.56, 143.59, 141.29, 131.97, 130.06, 129.02, 124.52, 121.89, 121.41, 117.03, 111.97, 110.97, 55.52, 55.41. MS (ESI): m/z 395 [M + H]+. HRMS (ESI) calcd for C20H19N4O5 [M + H]+, 395.1355; found, 395.1352.
4.1.3.13. 1-(4-Methoxyphenyl)-3-(2-(3,4,5-trimethoxyphenyl)pyridin-3-yl)urea (6m). Light brown solid, yield 92%, Mp: 209–211 °C; 1H NMR (300 MHz, DMSO): δ 8.79 (s, 1H), 8.54 (d, J = 8.3 Hz, 1H), 8.31 (d, J = 4.5 Hz, 1H), 7.68 (s, 1H), 7.53 (d, J = 2.8 Hz, 1H), 7.31 (t, J = 8.1 Hz, 2H), 7.26 (dd, J = 8.2, 4.7 Hz, 1H), 6.87 (s, 2H), 6.82 (d, J = 8.7 Hz, 2H), 3.89 (d, J = 2.3 Hz, 6H), 3.89 (s, 3H), 3.78 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO) δ 154.89, 153.08, 152.84, 147.82, 142.58, 137.35, 133.01, 131.76, 128.54, 122.21, 120.51, 113.62, 105.69, 60.30, 55.58, 54.96. MS (ESI): m/z 410 [M + H]+. HRMS (ESI) calcd for C22H24N3O5 [M + H]+, 410.1716; found, 410.1710.
4.1.3.14. 1-(4-Fluorophenyl)-3-(2-(3,4,5-trimethoxyphenyl)pyridin-3-yl)urea (6n). Light brown solid, yield 90%, Mp: 222–224 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.00 (s, 1H), 8.54 (d, J = 8.3 Hz, 1H), 8.33 (d, J = 4.6 Hz, 1H), 7.74 (s, 1H), 7.40 (dd, J = 8.9, 4.8 Hz, 2H), 7.27 (dd, J = 8.3, 4.7 Hz, 1H), 6.97 (t, J = 8.7 Hz, 2H), 6.88 (s, 2H), 3.90 (s, 6H), 3.89 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 172.84, 157.86 (d, JCF = 239.25 Hz), 152.91, 147.86, 142.78, 137.44, 134.94, 132.93 (d, JCF = 4.2 Hz), 128.73, 122.29, 120.00 (d, JCF = 7.6 Hz), 114.88 (d, JCF = 22.6 Hz), 105.80, 60.37, 55.64. MS (ESI): m/z 398 [M + H]+. HRMS (ESI) calcd for C21H21FN3O4 [M + H]+, 398.1516; found, 398.1510.
4.1.3.15. 1-(4-Chlorophenyl)-3-(2-(3,4,5-trimethoxyphenyl)pyridin-3-yl)urea (6o). Light brown solid, yield 86%, Mp: 218–220 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.06 (s, 1H), 8.54 (d, J = 8.3 Hz, 1H), 8.33 (d, J = 4.6 Hz, 1H), 7.74 (s, 1H), 7.47 (s, 1H), 7.40 (d, J = 8.8 Hz, 2H), 7.27 (dd, J = 7.8, 4.1 Hz, 1H), 7.22 (d, J = 8.8 Hz, 2H), 6.88 (s, 2H), 3.90 (d, J = 2.3 Hz, 6H), 3.89 (s, 3H). MS (ESI): m/z 414 [M + H]+. HRMS (ESI) calcd for C21H21ClN3O4 [M + H]+, 414.1221; found, 414.1215.
4.1.3.16. 1-(4-Nitrophenyl)-3-(2-(3,4,5-trimethoxyphenyl)pyridin-3-yl)urea (6p). Yellow solid, yield 85%, Mp: 261–263 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.66 (s, 1H), 8.55 (d, J = 8.3 Hz, 1H), 8.40–8.34 (m, 1H), 8.15 (d, J = 9.2 Hz, 2H), 7.94 (s, 1H), 7.62 (t, J = 8.3 Hz, 2H), 7.57 (d, J = 3.7 Hz, 1H), 7.31 (dd, J = 8.3, 4.7 Hz, 1H), 6.88 (s, 2H), 3.90 (s, 6H), 3.89 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 152.85, 152.06, 148.19, 145.58, 143.39, 141.26, 137.50, 132.72, 132.12, 128.87, 124.50, 122.18, 117.05, 105.79, 60.23, 55.60. MS (ESI): m/z 424 [M + H]+. HRMS (ESI) calcd for C21H21N4O6 [M + H]+, 425.1461; found, 425.1455.
4.1.3.17. 1-(2-(4-Fluorophenyl)pyridin-3-yl)-3-(4-methoxyphenyl)urea (6q). White solid, yield 84%, Mp: 191–193 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.52 (s, 1H), 8.48 (d, J = 4.9 Hz, 1H), 8.33 (s, 1H), 7.69–7.61 (m, 2H), 7.58 (s, 1H), 7.41 (s, 1H), 7.31 (d, J = 8.8 Hz, 2H), 7.19 (d, J = 8.6 Hz, 2H), 6.82 (d, J = 8.8 Hz, 2H), 3.78 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 162.24 (d, JCF = 245.25 Hz), 154.88, 153.05, 147.57, 143.04, 133.52 (d, JCF = 3.8 Hz), 131.87, 130.79 (d, JCF = 8.1 Hz), 129.26, 122.40, 120.49, 115.2 (d, JCF = 21.4 Hz), 113.68, 55.03. MS (ESI): m/z 338 [M + H]+. HRMS (ESI) calcd for C19H17FN3O2 [M + H]+, 338.1305; found, 338.1299.
4.1.3.18. 1-(4-Fluorophenyl)-3-(2-(4-fluorophenyl)pyridin-3-yl)urea (6r). White solid, yield 88%, Mp: 202–204 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.75 (s, 1H), 8.48 (d, J = 8.0 Hz, 1H), 8.35 (d, J = 1.7 Hz, 1H), 7.67 (d, J = 9.0 Hz, 3H), 7.43–7.35 (m, 2H), 7.31–7.17 (m, 3H), 7.03–6.92 (m, 2H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 162.22 (d, JCF = 246 Hz), 157.8 (d, JCF = 240 Hz), 152.78, 147.47, 143.25, 134.84, 133.93, 132.71, 130.74 (d, JCF = 7.5 Hz), 129.19, 122.28, 119.92 (d, JCF = 7.5 Hz), 115.13 (d, JCF = 21.75 Hz), 114.67. MS (ESI): m/z 326 [M + H]+. HRMS (ESI) calcd for C18H14F2N3O [M + H]+, 326.1105; found, 326.1099.
4.1.3.19. 1-(4-Chlorophenyl)-3-(2-(4-fluorophenyl)pyridin-3-yl)urea (6s). White solid, yield 87%, Mp: 216–2018 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.83 (s, 1H), 8.46 (d, J = 8.3 Hz, 1H), 8.35 (d, J = 4.0 Hz, 1H), 7.72 (d, J = 6.7 Hz, 1H), 7.67 (dd, J = 8.5, 5.5 Hz, 2H), 7.40 (d, J = 8.8 Hz, 2H), 7.24 (qd, J = 8.4, 5.3 Hz, 5H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 162.14 (d, JCF = 246 Hz), 152.45, 147.62, 143.29, 137.75, 133.87, 132.56, 130.74 (d, JCF = 8.2 Hz), 129.36, 128.13, 126.19, 122.21, 119.33, 115.11 (d, JCF = 21 Hz). MS (ESI): m/z 342 [M + H]+. HRMS (ESI) calcd for C18H14ClFN3O [M + H]+, 342.0809; found, 342.0803.
4.1.3.20. 1-(2-(4-Fluorophenyl)pyridin-3-yl)-3-(4-nitrophenyl)urea (6t). Yellow solid, yield 85%, Mp: 203–205 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.43 (s, 1H), 8.49–8.35 (m, 2H), 8.15 (dd, J = 9.2, 2.8 Hz, 2H), 7.95 (s, 1H), 7.68 (dd, J = 7.2, 4.3 Hz, 2H), 7.62 (dd, J = 9.3, 2.5 Hz, 2H), 7.35–7.18 (m, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 162.27 (d, JCF = 246.3 Hz), 152.04, 147.96, 145.55, 143.91, 141.29, 133.74, 132.04, 130.80 (d, JCF = 8.1 Hz), 129.69, 124.51, 122.31, 117.39, 117.10, 115.22 (d, JCF = 21.3 Hz). MS (ESI): m/z 353 [M + H]+. HRMS (ESI) calcd for C18H14FN4O3 [M + H]+, 353.1050; found, 353.1044.
4.1.3.21. 1-(4-Methoxyphenyl)-3-(2-(4-(trifluoromethyl)phenyl)pyridin-3-yl)urea (6u). White solid, yield 90%, Mp: 211–213 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.59 (s, 1H), 8.50 (d, J = 8.1 Hz, 1H), 8.36 (d, J = 4.4 Hz, 1H), 7.89–7.78 (m, 4H), 7.65–7.57 (m, 1H), 7.32 (d, J = 8.8 Hz, 3H), 6.82 (d, J = 8.9 Hz, 2H), 3.77 (s, 3H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 154.73, 152.79, 146.83, 143.27, 141.71, 133.14, 131.79, 129.48, 129.33, 125.02 (q, JCF3 = 3.3 Hz), 122.76, 121.85, 120.18, 113.55, 54.90. MS (ESI): m/z 388 [M + H]+. HRMS (ESI) calcd for C20H17F3N3O2 [M + H]+, 388.1273; found, 388.1267.
4.1.3.22. 1-(4-Fluorophenyl)-3-(2-(4-(trifluoromethyl)phenyl)pyridin-3-yl)urea (6v). White solid, yield 86%, Mp: 244–246 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.74 (s, 1H), 8.46 (d, J = 12.9 Hz, 1H), 8.38 (s, 1H), 7.83 (d, J = 10.9 Hz, 5H), 7.53 (s, 1H), 7.39 (s, 2H), 7.32 (d, J = 3.4 Hz, 1H), 7.00 (d, J = 8.5 Hz, 2H). 13C NMR (75 MHz, DMSO): δ 159.36, 152.70, 146.96, 143.54, 141.66, 134.82, 132.91, 129.67, 129.35, 125.03 (d, JCF3 = 3.6 Hz), 122.80, 119.88 (d, JCF = 7.7 Hz), 114.95, 114.66. MS (ESI): m/z 376 [M + H]+. HRMS (ESI) calcd for C19H14F4N3O [M + H]+, 376.1073; found, 376.1067.
4.1.3.23. 1-(4-Chlorophenyl)-3-(2-(4-(trifluoromethyl)phenyl)pyridin-3-yl)urea (6w). White solid, yield 84%, Mp: 216–218 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.86 (s, 1H), 8.46 (d, J = 8.4 Hz, 1H), 8.38 (d, J = 4.5 Hz, 1H), 7.86 (d, J = 10.8 Hz, 3H), 7.79 (d, J = 8.4 Hz, 2H), 7.40 (d, J = 8.8 Hz, 2H), 7.33 (dd, J = 8.4, 4.6 Hz, 1H), 7.22 (d, J = 8.8 Hz, 2H). MS (ESI): m/z 392 [M + H]+. HRMS (ESI) calcd for C19H14ClF3N3O [M + H]+, 392.0777; found, 392.0772.
4.1.3.24. 1-(4-Nitrophenyl)-3-(2-(4-(trifluoromethyl)phenyl)pyridin-3-yl)urea (6x). Yellow solid, yield 91%, Mp: 234–236 °C; 1H NMR (300 MHz, CDCl3 + DMSO): δ 9.39 (s, 1H), 8.48–8.40 (m, 2H), 8.15 (d, J = 9.2 Hz, 2H), 8.04 (s, 1H), 7.86–7.80 (m, 4H), 7.61 (d, J = 9.4 Hz, 2H), 7.36 (dd, J = 8.3, 4.7 Hz, 1H). 13C NMR (75 MHz, CDCl3 + DMSO): δ 151.98, 147.76, 145.63, 144.28, 141.61, 141.24, 132.30, 130.37, 129.47, 125.09 (d, JCF3 = 3.2 Hz), 124.53, 122.89, 117.21. MS (ESI): m/z 403 [M + H]+. HRMS (ESI) calcd for C19H14F3N4O4 [M + H]+, 403.1018; found, 403.1012.
4.2. Biology
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells from each sample were analyzed for DNA content (propidium iodide fluorescence) using a BD Accuri C6 flow cytometer. The final flow cytometry data was analyzed by using FlowJo software.24
000 cells from each sample were analyzed for DNA content (propidium iodide fluorescence) using a BD Accuri C6 flow cytometer. The final flow cytometry data was analyzed by using FlowJo software.24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 dilution in blocking solution for 4 h at room temperature. Subsequently, the antibodies were removed, and the cells were washed three times with PBS. Cells were then incubated with the FITC-labeled anti-mouse secondary antibody (1
200 dilution in blocking solution for 4 h at room temperature. Subsequently, the antibodies were removed, and the cells were washed three times with PBS. Cells were then incubated with the FITC-labeled anti-mouse secondary antibody (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500) for 1 h at room temperature. Cells were washed three times with PBS and then mounted in a medium containing DAPI. Images were captured using a confocal microscope.25
500) for 1 h at room temperature. Cells were washed three times with PBS and then mounted in a medium containing DAPI. Images were captured using a confocal microscope.25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells from each sample were used for analysis, where the obtained flow cytometry data was analyzed by using FlowJo software.23
000 cells from each sample were used for analysis, where the obtained flow cytometry data was analyzed by using FlowJo software.23Conflicts of interest
There are no conflicts of interest to declare.Data availability
All data supporting the findings of this study are available within the main article and its supplementary information (SI). Supplementary information: NMR, LC-MS and HRMS spectra. See DOI: https://doi.org/10.1039/d5ra09372d.Acknowledgements
S. T. and V. G. R. acknowledge the UGC, New Delhi, for the award of a research fellowship. This work was performed in part at the RMIT Micro Nano Research Facility (MNRF) in the Victorian Node of the Australian National Fabrication Facility (ANFF).References
- R. L. Siegel, T. B. Kratzer, A. N. Giaquinto, H. Sung and A. Jemal, Cancer statistics, 2025, Ca-Cancer J. Clin., 2025, 75(1), 10–45 Search PubMed.
- S. U. Khan, K. Fatima, S. Aisha and F. Malik, Unveiling the Mechanisms and Challenges of Cancer Drug Resistance, Cell Commun. Signaling, 2024, 22, 109 CrossRef.
- (a) L. Zhong, Y. Li, L. Xiong, W. Wang, M. Wu, T. Yuan, W. Yang, C. Tian, Z. Miao and T. Wang, et al., Small molecules in targeted cancer therapy: advances, challenges, and future perspectives, Signal Transduction Targeted Ther., 2021, 6, 201 CrossRef; (b) B. Kumar, S. Singh, I. Skvortsova and V. Kumar, Promising Targets in Anti-cancer Drug Development: Recent Updates, Curr. Med. Chem., 2017, 24(42), 4729–4752 Search PubMed.
- (a) J. Zugazagoitia, C. Guedes, S. Ponce, I. Ferrer, S. Molina-Pinelo and L. Paz-Ares, Current Challenges in Cancer Treatment, Clin. Ther., 2016, 38(7), 1551–1566 Search PubMed; (b) H. Mellatyar, S. Sattari, A. Nezami Asl and A. Akbarzadeh, Recent Advances and Challenges in Cancer Treatment with Car T Cell Therapy: A Novel Anti-cancer Strategy, BioNanoScience, 2024, 14, 4250–4262 Search PubMed.
- L. Lafanechère, The Microtubule Cytoskeleton: An Old Validated Target for Novel Therapeutic Drugs, Front. Pharmacol, 2022, 13, 969183 Search PubMed.
- K. E. Arnst, S. Banerjee, H. Chen, S. Deng, D.-J. Hwang, W. Li and D. D. Miller, Current Advances of Tubulin Inhibitors as Dual-Acting Small Molecules for Cancer Therapy, Med. Res. Rev., 2019, 39, 1236–1286 CrossRef.
- M. Jordan, Mechanism of action of antitumor drugs that interact with microtubules and tubulin, Curr. Med. Chem.: Anti-Cancer Agents, 2002, 2, 1–17 Search PubMed.
- (a) E. L. Schwartz, Antivascular Actions of Microtubule-Binding Drugs, Clin. Cancer Res., 2009, 15, 2594–2601 Search PubMed; (b) E. Pasquier, N. André and D. Braguer, Targeting Microtubules to Inhibit Angiogenesis and Disrupt Tumour Vasculature:Implications for Cancer Treatment, Curr. Cancer Drug Targets, 2007, 7(6), 566–581 Search PubMed; (c) V. Čermák, V. Dostál, M. Jelínek, L. Libusová, J. Kovář, D. Rösel and J. Brábek, Microtubule-targeting agents and their impact on cancer treatment, Eur. J. Med. Chem., 2020, 99, 151075 Search PubMed.
- M. J. Horgan, L. Zell, B. Siewert, H. Stuppner, D. Schuster and V. Temml, Identification of Novel β-Tubulin Inhibitors Using a Combined In Silico/In Vitro Approach, J. Chem. Inf. Model., 2023, 63(20), 6396–6411 Search PubMed.
- O. Ebenezer, M. Shapi and J. A. Tuszynski, A Review of the Recent Developments of Molecular Hybrids Targeting Tubulin Polymerization, Int. J. Mol. Sci., 2022, 23(7), 4001 CrossRef CAS PubMed.
- (a) Y. Lu, J. Chen, M. Xiao, W. Li and D. D. Miller, An Overview of Tubulin Inhibitors That Interact with the Colchicine Binding Site, Pharm. Res., 2012, 29(11), 2943–2971 CrossRef CAS PubMed; (b) N. Wang, J. Liu, C. Wang, L. Bai and X. Jiang, Asymmetric Total Syntheses of (-)-Jerantinines A, C, and E, (-)-16-Methoxytabersonine, (-)-Vindoline, and (+)-Vinblastine, Org. Lett., 2017, 20(1), 292–295 CrossRef PubMed; (c) N. Wang, S. Du, D. Li and X. Jiang, Divergent Asymmetric Total Synthesis of (+)-Vincadifformine, (-)-Quebrachamine, (+)-Aspidospermidine, (-)-Aspidospermine, (-)-Pyrifolidine, and Related Natural Products, Org. Lett., 2017, 19(12), 3167–3170 Search PubMed.
- K. Yoshimatsu, A. Yamaguchi, H. Yoshino, N. Koyanagi and K. Kitoh, Mechanism of Action of E7010, an Orally Active Sulfonamide Antitumor Agent: Inhibition of Mitosis by Binding to the Colchicine Site of Tubulin, Cancer Res., 1997, 57(15), 3208–3213 CAS.
- Y. Funahashi, N. Koyanagi and K. Kitoh, Antitumor Activity of E7010, an Oral Antimitotic Agent, in Combination with Conventional Chemotherapeutic Agents, Cancer Chemother. Pharmacol., 2001, 47, 179–184 CrossRef CAS PubMed.
- T. M. Mahgoub, E. J. Jordan, A. F. Mahdi, V. Oettl, S. Huefner, N. O'Donovan, J. Crown and D. M. Collins, Evaluation of ABT-751, a Novel Anti-mitotic Agent Able to Overcome Multi-drug Resistance, in Melanoma Cells, Cancer Chemother. Pharmacol., 2024, 93, 427–437 CrossRef CAS PubMed.
- T. M. Mahgoub, E. J. Jordan, A. F. Mahdi, V. Oettl, S. Hufner, N. O'Donovan, J. Crown and D. M. Collins, Evaluation of ABT-751, a Novel Anti-mitotic Agent Able to Overcome Multi-drug Resistance, in Melanoma Cells, Cancer Chemother. Pharmacol., 2024, 93, 427–437 Search PubMed.
- S. Z. Dehghanian, C.-T. Pan, J. M. Lee and Y.-L. Shiue, ABT-751 Induces Multiple Anticancer Effects in Urinary Bladder Urothelial Carcinoma-Derived Cells: Highlighting the Induction of Cytostasis through the Inhibition of SKP2 at Both Transcriptional and Post-Translational Levels, Int. J. Mol. Sci., 2021, 22(2), 945 Search PubMed.
- (a) A. Kamal, M. Ashraf, S. T. Basha, S. M. A. Hussaini, S. Singh, M. V. P. S. Vishnuvardhan, B. Kiran and B. Sridhar, Design, synthesis and antiproliferative activity of the new conjugates of E7010 and resveratrol as tubulin polymerization inhibitors, Org. Biomol. Chem., 2016, 14, 1382–1394 Search PubMed; (b) B. Prasad, V. L. Nayak, P. S. Srikanth, M. F. Baig, N. V. S. Reddy, K. S. Babu and A. Kamal, Synthesis and biological evaluation of 1-benzyl-N-(2-(phenylamino)pyridin-3-yl)-1H-1,2,3-triazole-4-carboxamides as antimitotic agents, Bioorg. Chem., 2019, 83, 535–548 Search PubMed; (c) F. Sultana, S. P. Shaik, V. L. Nayak, S. M. A. Hussaini, K. Marumudi, B. Sridevi, T. B. Shaik, D. Bhattacharjee, A. Alarifi and A. Kamal, Design, Synthesis and Biological Evaluation of 2-Anilinopyridyl-Linked Oxindole Conjugates as Potent Tubulin Polymerisation Inhibitors, ChemistrySelect, 2017, 2, 9901–9910 CrossRef CAS; (d) T. B. Shaik, S. M. A. Hussaini, V. L. Nayak, M. L. Sucharitha, M. S. Malik and A. Kamal, Rational design and synthesis of 2-anilinopyridinyl-benzothiazole Schiff bases as antimitotic agents, Bioorg. Med. Chem. Lett., 2017, 27, 2549–2558 Search PubMed; (e) F. Sultana, M. A. Saifi, R. Syed, G. S. Mani, S. P. Shaik, G. O. Egharevba, C. Godugu, S. Shahjahan and A. Kamal, Synthesis of 2-anilinopyridyl linked benzothiazole hydrazones as apoptosis inducing cytotoxic agents, New J. Chem., 2019, 43, 7150–7161 Search PubMed.
- X. He, Y. Peng, R. Liu, W. Yuan, P. Ye, Z. Wang, C. Fu, Z. Xie, X. Yang, X. Lei, L. Wang, B. Yang, G. Tang and X. Deng, Discovery of Novel Flavonoid Derivatives Targeting VEGFR2 and Tubulin with Potent Anticancer Efficacy, Bioorg. Chem., 2025, 163, 108713 Search PubMed.
- W. Wang, D. Kong, H. Cheng, L. Tan, Z. Zhang, X. Zhuang, H. Long, Y. Zhou, Y. Xu, X. Yang and K. Din, Design, Synthesis, and Biological Evaluation of Novel Sulfonamide-Substituted Chalcone Derivatives as Potent Tubulin Polymerization Inhibitors, Bioorg. Med. Chem. Lett., 2014, 24, 4250–4253 Search PubMed.
- J.-N. Chen, D.-W. Wu, T. Li, K.-J. Yang, L. Cheng, Z.-P. Zhou, S.-M. Pu and W.-H. Lin, Recent Advances in the Development of Colchicine Binding Site Inhibitors Targeting Tubulin Polymerization: An Update of the Structure–Activity Relationships, Curr. Top. Med. Chem., 2017, 17, 3099–3130 Search PubMed.
- M. Gagné-Boulet, S. Fortin, J. Lacroix, C. A. Lefebvre, M. F. Côté and R. C. Gaudreault, Design, Synthesis, and Biological Evaluation of Sulfonamide Derivatives as Tubulin Polymerization Inhibitors with Potent Antiproliferative Activity, Eur. J. Med. Chem., 2015, 100, 34–43 CrossRef.
- (a) D. K. Sigalapalli, V. Pooladanda, P. Singh, M. Kadagathur, S. D. Guggilapu, J. L. Uppu, N. D. Tangellamudi, P. K. Gangireddy, C. Godugu and N. B. Bathini, Discovery of certain benzyl/phenethyl thiazolidinone-indole hybrids as potential anti-proliferative agents: Synthesis, molecular modeling and tubulin polymerization inhibition study, Bioorg. Chem., 2019, 92, 103188 CrossRef CAS; (b) C. Jadala, M. Sathish, P. Anchi, R. Tokala, U. J. Lakshmi, V. G. Reddy, N. Shankaraiah, C. Godugu and A. Kamal, Synthesis of Combretastatin-A4 Carboxamidest that Mimic Sulfonyl Piperazines by a Molecular Hybridization Approach: in vitro Cytotoxicity Evaluation and Inhibition of Tubulin Polymerization, ChemMedChem, 2019, 14, 2052–2064 CrossRef CAS.
- R. Parupalli, R. Akunuri, A. Spandana, R. Phanindranath, S. Pyreddy, M. R. Bazaz, M. Vadakattu, S. V. Joshi, S. Bujji, B. Gorre, V. M. Yaddanapudi, M. P. Dandekar, V. G. Reddy, N. Nagesh and S. Nanduri, Synthesis and Biological Evaluation of 1-Phenyl-4,6-Dihydrobenzo[b]pyrazolo[3,4-d]azepin-5(1H)-one/Thiones as Anticancer Agents, Bioorg. Chem., 2023, 135, 106478 Search PubMed.
- (a) J. A. Choi, J. Y. Kim, J. Y. Lee, C. M. Kang, H. J. Kwon, Y. D. Yoo, T. W. Kim, Y. S. Lee and S. J. Lee, Induction of Cell Cycle Arrest and Apoptosis in Human Breast Cancer Cells by Quercetin, Int. J. Oncol., 2001, 19, 837–844 CAS; (b) N. Mirzadeh, T. S. Reddy, R. Luwor, S. Privér, V. G. Reddy, R. K. Jakku, N. S. Andrew, M. Plebanski, H. Christian and S. Bhargava, Dinuclear orthometallated gold(I)-gold(III) anticancer complexes with potent in vivo activity through an ROS-dependent mechanism, Metallomics, 2021, 13, mfab039 CrossRef.
- S. Sana, V. G. Reddy, T. S. Reddy, R. Tokala, R. Kumar, S. K. Bhargava and N. Shankaraiah, Cinnamide-Derived Pyrimidine-Benzimidazole Hybrids as Tubulin Inhibitors: Synthesis, In Silico, and Cell Growth Inhibition Studies, Bioorg. Chem., 2021, 110, 104765 CrossRef CAS.
- A. Cormier, M. Marchand, R. B. Ravelli, M. Knossow and B. Gigant, Structural Insight into the Inhibition of Tubulin by Antimitotic Agents: Binding of Vinblastine and Maytansine to Tubulin, EMBO Rep., 2008, 9, 1101–1106 CrossRef CAS PubMed.
- V. G. Reddy, T. S. Reddy, C. Jadala, M. S. Reddy, F. Sultana, R. Akunuri, S. K. Bhargava, D. Wlodkowic, P. Srihari and A. Kamal, Pyrazolo-Benzothiazole Hybrids: Synthesis, Anticancer Properties and Evaluation of Antiangiogenic Activity Using In Vitro VEGFR-2 Kinase and In Vivo Transgenic Zebrafish Model, Eur. J. Med. Chem., 2019, 182, 111609 CrossRef.
- S. Sana, V. G. Reddy, S. Bhandari, T. S. Reddy, R. Tokala, A. P. Sakla, S. K. Bhargava and N. Shankaraiah, Exploration of Carbamide-Derived Pyrimidine-Thioindole Conjugates as Potential VEGFR-2 Inhibitors with Anti-Angiogenesis Effect, Eur. J. Med. Chem., 2020, 200, 112457 Search PubMed.
- L.-S. Zheng, F. Wang, X.-Y. Ye, G.-Q. Chen and X. Zhang, Asymmetric Hydrogenation of 2-Aryl-3-phthalimidopyridinium Salts: Synthesis of Piperidine Derivatives with Two Contiguous Stereocenters, Org. Lett., 2020, 22, 8882–8887 Search PubMed.
- S. Malasala, A. Polomoni, M. N. Ahmad, M. Shukla, G. Kaul, A. Dasgupta, S. Chopra and S. Nanduri, Structure-Based Design, Synthesis and Evaluation of New Thienopyrimidine Derivatives as Anti-Bacterial Agents, J. Mol. Struct., 2021, 1234, 130168 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |