Open Access Article
Open Access ArticleA pre-synthetic conjugation methodology of DNA and RNA: combining on-column conjugation and tandem oligonucleotide synthesis
Jagandeep S.
Saraya
 and
Derek K.
O'Flaherty
and
Derek K.
O'Flaherty
 *
*
Department of Chemistry, University of Guelph, 50 Stone Rd. E, Guelph, ON N1G 2 W1, Canada. E-mail: doflaher@uoguelph.ca
First published on 27th February 2026
Abstract
Amino modifiers are valuable chemical tools for nucleic acid bioconjugation applications and for probing potential mechanisms relevant to the origins of life. Herein, we describe a streamlined and cost-effective strategy for the incorporating amine functionalities at the 3′-end of DNA/RNA constructs through a commercially available diethyl 2,2-bis(hydroxymethyl)malonate-derived controlled pore glass (a so-called version of chemical phosphorylation reagent II CPG, or CPR II CPG). This platform mitigates certain limitations associated with previously explored sulfonyl-based supports, thereby improving the robustness of amine handle conjugation. CPR II CPG is first detritylated, converted to a mixed N-hydroxysuccinimide carbonate followed by efficient derivatization with the amine of interest. Oligonucleotides are then assembled using standard solid-phase synthesis and isolated by standard purification workflows. Using 1,3-diamino-2-propanol as a multivalent conjugation handle allows for the installation of ligands prior to strand elongation, which may be further derivatized post-synthetically. Finally, the integration this advancement, and others, with tandem oligonucleotide synthesis (TOS) enables a versatile conjugation TOS methodology, thereby advancing nucleic acid engineering and broader biotechnology applications.
Introduction
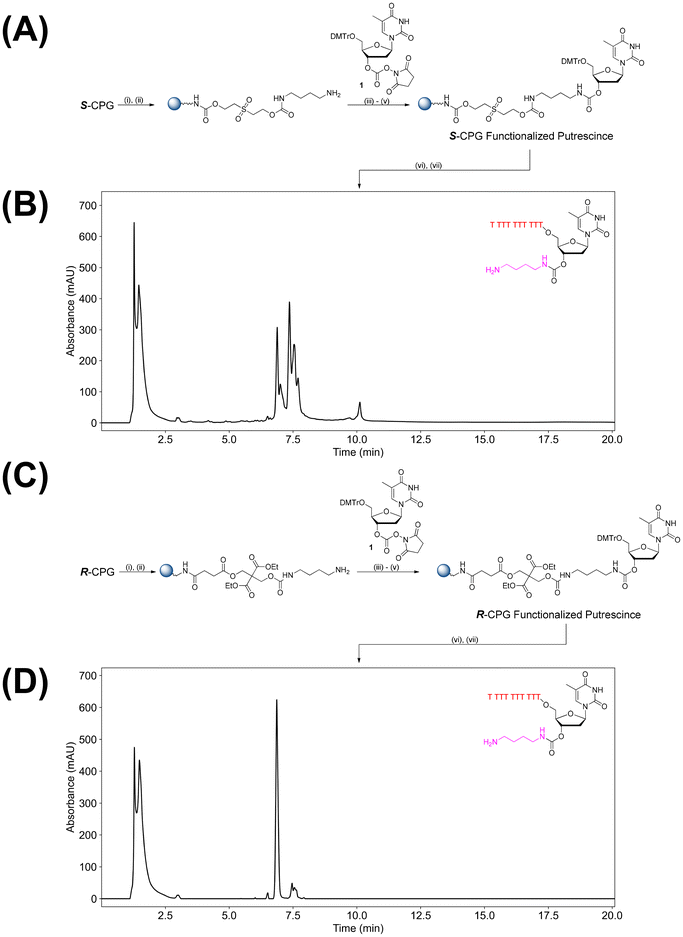
Amino-modifiers are a convenient class of chemical modifications introduced in nucleic acid sequences for (bio)conjugation and origins-of-life research.1–4 Their nucleophilicity, relative to other nucleophiles present in nucleic acids, allows for an orthogonal means of linking ligands (e.g. via NHS esters or by template-directed synthesis).5,6 We and others have developed technologies for the facile introduction of amino-modifiers in DNA and RNA.7–12 The conjugation of the 3′-position is particularly important, as it allows the 5′-position to be utilized for other purposes like enzymatic activity, and may have future applications in real time fluorescence monitoring of PCR.13 This position is typically accessible by using solid supports already pre-derivatized by the ligand of interest, or by using a “reverse” synthesis strategy.14 This makes these pathways typically costly, sometimes laborious, and not necessarily amenable to ligand screening.Maximizing versatility in oligonucleotide conjugation is desirable for many applications including screening ligands and/or sequence contexts.15–17 As a result, we sought to develop a methodology that optimizes on-column functionalization of nucleic acid scaffolds. Using 3′-Phosphate-ON controlled pore glass (CPG), we previously reported a means of derivatizing nucleic acids with 3′-amino groups in DNA and RNA via a carbamate linkage (Fig. 1A).11 Albeit versatile, a notable deficiency was identified as certain amino-modifiers caused significant immolation of the sulfone-containing linker, inevitably lowering overall yields. Furthermore, during our efforts to derivatize nucleic acids with specific amino-modifiers (e.g. putrescine), we observed a substantial accumulation of impurities during and/or after the cleavage and deprotection processing of the modified oligonucleotides.
Based on these limitations, we developed the use of a commercially available dimethyl 2,2-bis(hydroxymethyl)malonate derivatized CPG (a so-called version of chemical phosphorylating reagent II CPG (CPR II CPG), Fig. 1B) to mitigate the issues observed with S-CPG. We find that both the yield and purity of nucleic acid sequences improve significantly for previously problematic syntheses. Building on these findings, we develop a versatile pre-synthetic conjugation platform that allows for a versatile conjugate synthetic pathway. Coupled with previous work on tandem oligonucleotide synthesis (TOS), we introduce a flexible on-column conjugation methodology that allows for selective conjugation of the upstream and/or downstream oligonucleotides.11,12,18
Results and discussion
Somewhat to our surprise, conjugating putrescine to model DNA sequences proved problematic viaS-CPG (Fig. 1A), as significant by-products were observed by SAX-HPLC (Fig. 2B) in conditions tested. We were originally interested in this conjugate as putrescine, and other polyamines, have shown to stabilize nucleic acid structures.19–21 The observed by-products, presumably from the reaction with reactive species generated during deprotection (e.g. divinyl sulfone, acrylonitrile, and/or transamination of nucleobase protecting group, incomplete immolation), was not as prominent when using 3′-amino-2′,3′-dideoxynucleosides (e.g. 3′-NH2-dT, see SI Fig. S3–S6 and Table S2 for head-to-head comparison between S-CPG and R-CPG) or other linear diamines like cystamine.11 Using the commercially available alternative to 3′-Phosphate-ON CPG22 (i.e. CPR II CPG, Fig. 1B![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 23) significantly minimized by-product formation with satisfactory product yields. Here, the CPR II CPG was detritylated and then treated with a saturated solution (1 mL) of N,N′-disuccinimidyl carbonate (DSC) in 1
23) significantly minimized by-product formation with satisfactory product yields. Here, the CPR II CPG was detritylated and then treated with a saturated solution (1 mL) of N,N′-disuccinimidyl carbonate (DSC) in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) pyridine
1 (v/v) pyridine![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) acetonitrile to produce R-CPG (Fig. 1B). The activated solid support was then washed (see SI for details) and treated with putrescine (50 mM) in 1
acetonitrile to produce R-CPG (Fig. 1B). The activated solid support was then washed (see SI for details) and treated with putrescine (50 mM) in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) DMSO
1 (v/v) DMSO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Py for 10 min at room temperature. The resulting amino-derivatized CPR II CPG was subject to a solution containing 5′-O-(4,4′-dimethoxytrityl)-3′-O-(2,5-dioxo-1-pyrrolidinyloxycarbonyl)-thymidine (compound 1, SI Scheme S1) (50 mM) in 1
Py for 10 min at room temperature. The resulting amino-derivatized CPR II CPG was subject to a solution containing 5′-O-(4,4′-dimethoxytrityl)-3′-O-(2,5-dioxo-1-pyrrolidinyloxycarbonyl)-thymidine (compound 1, SI Scheme S1) (50 mM) in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) DMF
1 (v/v) DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN.11 We were thrilled to observe almost complete mitigation of the issue when using CPR II CPG, after SPS and cleavage/deprotection of the synthetic DNA (Fig. 2D, see S8 for MS). These results suggest that the immolated vinyl sulfone linker (or degraded products thereof) is likely causing by-product formation. Although the immolation of the malonyl linker (or malonamide version) leads to a proposed Michael acceptor, it appears that the free amino-modifier functionality does not get captured in our conditions (see SI for details).23
MeCN.11 We were thrilled to observe almost complete mitigation of the issue when using CPR II CPG, after SPS and cleavage/deprotection of the synthetic DNA (Fig. 2D, see S8 for MS). These results suggest that the immolated vinyl sulfone linker (or degraded products thereof) is likely causing by-product formation. Although the immolation of the malonyl linker (or malonamide version) leads to a proposed Michael acceptor, it appears that the free amino-modifier functionality does not get captured in our conditions (see SI for details).23
We next explored whether a branched trifunctional linker, specifically 1,3-diamino-2-propanol, could be compatible with CPR II CPG derivatization. This linker was selected due to its low cost, commercial availability, the presence of potentially orthogonal hydroxyl and amino groups, and the ability to functionalize an oligonucleotide prior to synthesis, directly on solid support. Initial attempts to conjugate this linker to S-CPG resulted in significant immolation of the sulfone linkage, which limited its utility in that context (data not shown). Alternatively, we hoped that treating R-CPG with unprotected 1,3-diamino-2-propanol would generate a modified support bearing a free hydroxyl and amino group. We could then functionalize the remaining amino group using various small molecule handles through our recently developed aqueous-compatible on-column 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC)-mediated conjugation strategy,8 as well as other established methods with slight modifications (Fig. 3).24 Standard SPS could then carried out to generate ligand-modified oligonucleotides at the 3′-terminus.
 | ||
| Fig. 3 Proposed conjugation pathway for the R-CPG functionalized with 1,3-diamino-2-propanol prior to oligonucleotide synthesis various ligand handles can be installed at the free amino group using streamlined chemistries (N-hydroxysuccinimide (NHS) ester or carbonate chemistries to furnish amide and carbamate linkages, respectively, as well as via EDC-mediated amide bond formation).5 “R”, “R′”, “R″” denotes chemical variations in the ligand handles evaluated. | ||
However, LC analysis of our initial attempts revealed incomplete ligand conjugation, which we hypothesized could be attributed to undesired cross-linking of the bis-amino linker during R-CPG treatment (data not included). That is, the second amino group could potentially react with neighboring activated mixed-carbonate groups, thereby reducing its overall availability for subsequent derivatization.
To address this issue, we employed a protection strategy using the 4-monomethoxytrityl (MMTr) group (compound 2, Scheme 1) to temporarily mask one of the amino groups on the linker, thereby inhibiting unwanted cross-linking. After coupling, the support-bound MMTr-protected 1,3-diamino-2-propyl linker was subject to on-instrument detritylation to unmask the desired reactive amino-group for ligand conjugation (Fig. 4). After detritylation, the solid support was washed sequentially with H2O, 50% (v/v) MeCN/H2O, and MeCN to eliminate residual impurities. Given the abundance of commercially available NHS ester ligand handles, and their efficient reactivity with primary/secondary amines, we opted to investigate their on-column derivatization compatibility first.25–27 We chose Boc-Val-OSu and propargyl NHS ester as useful examples to represent amino acid and alkyne-containing conjugates. Ligand conjugation was carried out using a 100 mM solution of the NHS esters in 100 mM HEPES buffer (pH 8.0) containing 50% (v/v) DMF (600 µL final volume) for 2 h. After thorough washing, a mixed-sequence DNA strand (d(TGAC)3) was assembled, with the initial phosphoramidite coupling performed using an extended coupling time (10 min) to mitigate potential steric effects arising from the appended ligand. The oligonucleotide was cleaved and deprotected in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 (v/v/v) EtOH
2 (v/v/v) EtOH![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeNH2(aq)
MeNH2(aq)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NH4OH(aq) (AMA, see SI for additional details) at 65 °C for 2 h and analyzed by SAX-HPLC (Fig. 4). In both cases, a single major product peak was observed. Isolation and characterization by mass spectrometry confirmed successful incorporation of the desired ligands (Fig. S12 and S14).
NH4OH(aq) (AMA, see SI for additional details) at 65 °C for 2 h and analyzed by SAX-HPLC (Fig. 4). In both cases, a single major product peak was observed. Isolation and characterization by mass spectrometry confirmed successful incorporation of the desired ligands (Fig. S12 and S14).
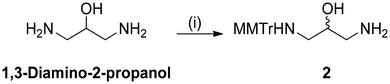 | ||
| Scheme 1 Synthesis of tritylated diamino conjugator 2. Reagents and conditions: (i) MMTr-Cl (1.0 eq.), DBU (3.0 eq.), MeCN, 22 °C, 1 h, 52%. | ||
To evaluate the generality of this approach, the same protocol was applied to the synthesis of a mixed RNA sequence (r(UGAC)3), bearing either a valyl or propargyl functionality (Fig. 4). Conjugation efficiency and overall performance were satisfactory. Importantly, all coupling steps were conducted under aqueous-compatible conditions, thereby avoiding elaborate reaction setups and enabling operational simplicity. Organic conditions may also be employed, when necessary, to facilitate ligand solubilization (see SI for further details). Subsequent functionalization of the free amine post-deprotection/cleavage of the oligonucleotide was also possible using previously reported procedures with slight modifications (Fig. S21–S24).28
Encouraged by the successful incorporation of NHS esters to furnish amide linkages on solid support, we next examined whether the diaminopropyl scaffold could support conjugation via EDC-mediated amide bond formation (Fig. 5). This strategy is particularly attractive given the broad commercial availability and lower cost of carboxylic acids relative to their pre-activated NHS ester counterparts. Following reaction of compound 2 with R-CPG, detritylation, and thorough washing, the functionalized support was treated with a reaction mixture containing the desired carboxylic acid ligand (100 mM), N-methylimidazole (N-MeIm, 100 mM), and EDC·HCl (400 mM) in 100 mM HEPES buffer (pH 8.0) with 50% (v/v) DMF.5 As in the previous scenario, two representative examples were tested: a lysine-derived carboxylic acid employed to generate an amino acid–oligonucleotide conjugate, and 4-bromobenzoic acid selected as a representative ligand based on its relevance to DNA-encoded library (DEL) synthesis platforms and its applicability to biochemical and biotechnological contexts.29–32 In both separate reactions, suspensions were allowed to react for 8 h, followed by washing of the solid supports with H2O, 50% (v/v) MeCN/H2O, and MeCN (10 mL each) to remove residual reagents. Each reaction was repeated for an additional 8 h to drive the coupling step, followed by extensive washing both manually (as described above) and on the automated DNA synthesizer with dry MeCN. A mixed-sequence DNA strand (d(TGAC)3) was elongated from the modified supports, cleaved and deprotected under standard AMA conditions. SAX-HPLC analysis revealed a single predominant product peak for each conjugate (Fig. 5). Isolation and characterization by mass spectrometry supported formation of the desired products (Fig. S26 and S28). Finally, the conjugation reactions were successfully reproduced using mixed-sequence RNA constructs (Fig. 5).
As a final pathway, we investigated whether this strategy could be extended to the formation of carbamate-linked conjugates on solid support using commercially available succinimidyl carbonate (OSu) handles (Fig. 6). As described above, the MMTr was removed and the solid support extensively washed. Subsequently, conjugation was performed using Alloc-OSu or Boc-OSu as two additional representative coupling reagents (one bearing an olefinic moiety and the other a bulky tert-butyl group) each distinct from those evaluated above. Coupling was carried out using the desired OSu carbonate ligand (100 mM) in either 100 mM HEPES buffer (pH 8.0) containing 50% (v/v) DMF or in DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Py (1
Py (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) for 2 h. Following reaction completion, the support was thoroughly washed and subsequently extended using a mixed DNA sequence (Fig. 6). Notably, the use of fully organic conditions, for the OSu carbonate ligand examples tested, provided a modest improvement (∼5–10%) in crude strand purity, which we attributed to enhanced ligand solubility relative to aqueous-compatible conditions. RNA synthesis under analogous conditions was also successful. As we previously reported, the requisite OSu carbonate derivatives are readily accessible via DSC-mediated carbonate formation from corresponding alcohol-containing ligand handles, and may be employed without extensive purification for on-solid support functionalization.11,12 Collectively, these findings (Fig. 4–6) suggest that aqueous-compatible conditions are particularly well-suited for hydrophilic ligands (e.g. PEGylated substrates), whereas organic media may be advantageous for more hydrophobic ligands.
1, v/v) for 2 h. Following reaction completion, the support was thoroughly washed and subsequently extended using a mixed DNA sequence (Fig. 6). Notably, the use of fully organic conditions, for the OSu carbonate ligand examples tested, provided a modest improvement (∼5–10%) in crude strand purity, which we attributed to enhanced ligand solubility relative to aqueous-compatible conditions. RNA synthesis under analogous conditions was also successful. As we previously reported, the requisite OSu carbonate derivatives are readily accessible via DSC-mediated carbonate formation from corresponding alcohol-containing ligand handles, and may be employed without extensive purification for on-solid support functionalization.11,12 Collectively, these findings (Fig. 4–6) suggest that aqueous-compatible conditions are particularly well-suited for hydrophilic ligands (e.g. PEGylated substrates), whereas organic media may be advantageous for more hydrophobic ligands.
Building on previous work from our group and others, we sought to evaluate how our newly developed conjugation strategy could be applied to tandem oligonucleotide synthesis (TOS).33–41 TOS offers distinct advantages for applications involving multiple nucleic acid strands, such as PCR and siRNA-based studies.42,43 In addition to increasing oligomer production throughput, TOS reduces overall cost by lowering the required amount of solid support and minimizing pre- and post-synthesis handling steps, including column packing and oligomer mixing. To the best of our knowledge, reports integrating on-column conjugation strategies with TOS remain sparse. As an initial proof-of-concept (Fig. 7A), we generated a lauric acid-functionalized DNA conjugate (as a hydrophobic ligand example, see SI for synthetic details), followed by the incorporation of a base labile cleavable linker (via the X-amidite, Fig. 7A), and a second strand bearing a terminal amino modifier (5′-amino-5′-deoxythymidinyl unit, NH2T, Fig. 7A) capable of further derivatization. The 5′-end of this downstream strand was first deblocked, then conjugated with an propargyl NHS ester in organic conditions (Fig. 7A).24 To be noted, sequence identities were selected arbitrarily, with the sole criterion that the two resulting strands were capable of hybridization to one another (Fig. 7B). Standard deprotection yielded two major species, as observed by SAX-HPLC (Fig. 7C), with mass spectrometric analysis of both isolated species showing m/z values consistent with those expected (Fig. S54 and S56). To broaden the applicability of this approach, the analogous 5′-OH-strand, instead of the 5′-NH2 was synthesized (Fig. S57). In this case, rather than employing NHS ester-mediated amide bond formation, the 5′-hydroxyl was conjugated to amine-PEG3-azide via DSC-mediated carbamate formation.24 The two predominant species were detected by SAX-HPLC and characterized by mass spectrometry, with the measured m/z values aligning with the calculated m/z values (Fig. S59 and S61).
As a final expansion on our technology, we sought to establish a method that would facilitate a modular post-synthetic TOS conjugation, via an azide-containing handle (compound 3, see Scheme S2). We have recently reported a robust methodology for the incorporation of pyrimidine nucleosides modified at the 3′-position that have traditionally posed synthetic challenges (such as azide-containing substrates).28 This was achieved through the installation of a disulfide linker covalently attached to the C4-position of the nucleobase, enabling efficient and versatile means of conjugation post-elongation of the nucleic acid sequence. Given its post-elongation utility in (bio)conjugation strategies, we asked whether the amino-containing mononucleoside 3 could be coupled to R-CPG (Fig. 8A). A second DNA strand, separated by the base-labile linker (via the X-amidite, Fig. 8A), was subsequently assembled. The 5′-terminus of the downstream strand was then functionalized with a hydrophilic amino-PEG7-alcohol ligand using a previously established protocol, with minor modifications.24 The upstream strand was functionalized via the azide functionality using strain-promoted azide–alkyne cycloaddition (SPAAC) with a model cyclooctyne ligand (BCN-OH, Fig. 8A). Upon standard deprotection, two predominant species were observed by SAX-HPLC (Fig. 8C), with m/z values matching those calculated for the desired products (Fig. S63 and S65). Together, results described in Fig. 7 and 8 (and Fig. S57) demonstrate a robust platform for pre- and post-elongation (bio)conjugations, compatible with TOS applications.
Conclusion
Motivated by challenges in preparing certain nucleic acid strands bearing a 3′-amino-functionality, we aimed to streamline solid support functionalization using solid supports alternative to S-CPG. Leveraging the improved stability of CPR II CPG in mild basic conditions, we showcase the successful incorporation of previously problematic amino-containing handles such as putrescine. Our methodology enabled the incorporation of a trifunctional linker to introduce a variety of conjugation handles via a modular on-column process prior to strand elongation. Although branched 3′-amino modifiers are commercially available (e.g. 3′-amino-modifier serinol CPG), our approach uses standard acid-mediated unmasking of the reactive amino group, and provides an orthogonal amino handle that can be exploited for post-synthetic conjugation. We anticipate that this technology will be broadly adopted for on-column synthesis of 3′-conjugates, particularly because many pre-installed solid supports are either cost-prohibitive for low-resource laboratories, or not commercially available. Furthermore, we established tandem oligonucleotide synthesis of two DNA strands incorporating ligand handles, enabling both pre-synthetic and post-synthetic conjugation at the respective 3′-termini of the upstream strand. Employing our novel methodologies, we aim to streamline the synthesis of highly modified oligonucleotides within duplex contexts.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: experimental methods, supporting figures, small molecule and oligonucleotide synthesis/characterization. See DOI: https://doi.org/10.1039/d5ob01870f.Acknowledgements
The authors are grateful to the O'Flaherty group for helpful discussions. This work was supported by a NSERC Discovery Grant (RGPIN 2020-05043), a NSERC Alliance Catalyst Grant (ALLRP 57555822), Canadian Foundation for Innovation – John R. Evans Leaders Fund (projects 43757 and 45118), and the HFSP Research Grant-Early Career (RGY0062/2022: https://doi.org/10.52044/HFSP.RGY00622022.pc.gr.153594).References
- K. Klabenkova, A. Fokina and D. Stetsenko, Chemistry of Peptide-Oligonucleotide Conjugates: A Review, Molecules, 2021, 26(17), 5420, DOI:10.3390/molecules26175420.
- M. Egli and M. Manoharan, Chemistry, Structure and Function of Approved Oligonucleotide Therapeutics, Nucleic Acids Res., 2023, 51(6), 2529–2573, DOI:10.1093/nar/gkad067.
- K. Renault, J. W. Fredy, P.-Y. Renard and C. Sabot, Covalent Modification of Biomolecules through Maleimide-Based Labeling Strategies, Bioconjugate Chem., 2018, 29(8), 2497–2513, DOI:10.1021/acs.bioconjchem.8b00252.
- H. Lönnberg, Solid-Phase Synthesis of Oligonucleotide Conjugates Useful for Delivery and Targeting of Potential Nucleic Acid Therapeutics, Bioconjugate Chem., 2009, 20(6), 1065–1094, DOI:10.1021/bc800406a.
- J. S. Saraya, S. R. Sammons and D. K. O'Flaherty, Aqueous Compatible Post–Synthetic On–Column Conjugation of Nucleic Acids Using Amino–Modifiers, ChemBioChem, 2025, 26(1), e202400643, DOI:10.1002/cbic.202400643.
- L. Zhou, D. K. O'Flaherty and J. W. Szostak, Template–Directed Copying of RNA by Non–enzymatic Ligation, Angew. Chem., Int. Ed., 2020, 59(36), 15682–15687, DOI:10.1002/anie.202004934.
- U. Asseline and N. T. Thuong, New Solid-Phase for Automated Synthesis of Oligonucleotides Containing an Amino-Alkyl Linker at Their 3′-End, Tetrahedron Lett., 1990, 31(1), 81–84, DOI:10.1016/S0040-4039(00)94339-9.
- D. L. McMinn and M. M. Greenberg, Novel Solid Phase Synthesis Supports for the Preparation of Oligonucleotides Containing 3′-Alkyl Amines, Tetrahedron, 1996, 52(11), 3827–3840, DOI:10.1016/S0040-4020(96)00053-1.
- A. Aviñó, R. G. Garcia, F. Albericio, M. Mann, M. Wilm, G. Neubauer and R. Eritja, New Carbamate Supports for the Preparation of 3′-Amino-Modified Oligonucleotides, Bioorg. Med. Chem., 1996, 4(10), 1649–1658, DOI:10.1016/0968-0896(96)00156-3.
- R. Eisenhuth and C. Richert, Convenient Syntheses of 3′-Amino-2′,3′-Dideoxynucleosides, Their 5′-Monophosphates, and 3′-Aminoterminal Oligodeoxynucleotide Primers, J. Org. Chem., 2009, 74(1), 26–37, DOI:10.1021/jo8018889.
- J. S. Saraya, N. G. Horton, S. R. Sammons and D. K. O′Flaherty, A Robust Strategy for Introducing Amino–Modifiers in Nucleic Acids: Enabling Novel Amino Tandem Oligonucleotide Synthesis in DNA and RNA, Chem. – Eur. J., 2025, 31(26), e202500448, DOI:10.1002/chem.202500448.
- J. S. Saraya, N. G. Horton, M. J. Capperauld, E. Zakaria and D. K. O’Flaherty, A Versatile Disulfide–Containing Solid–Support Strategy for 3′–Modifiers in Oligonucleotides: Introducing Modular Tandem Oligonucleotide Synthesis, Chem. – Asian J., 2025, 20(16), e00537, DOI:10.1002/asia.202500537.
- I. M. Mackay, K. E. Arden and A. Nitsche, Real-Time PCR in Virology, Nucleic Acids Res., 2002, 30(6), 1292–1305, DOI:10.1093/nar/30.6.1292.
- A. P. Guzaev and R. T. Pon, Attachment of Nucleosides and Other Linkers to Solid–Phase Supports for Oligonucleotide Synthesis, Curr. Protoc. Nucleic Acid Chem., 2013, 52(1), 3.2.1–3.2.23, DOI:10.1002/0471142700.nc0302s52.
- A. Gironda-Martínez, E. J. Donckele, F. Samain and D. Neri, DNA-Encoded Chemical Libraries: A Comprehensive Review with Succesful Stories and Future Challenges, ACS Pharmacol. Transl. Sci., 2021, 4(4), 1265–1279, DOI:10.1021/acsptsci.1c00118.
- M. A. Clark, R. A. Acharya, C. C. Arico-Muendel, S. L. Belyanskaya, D. R. Benjamin, N. R. Carlson, P. A. Centrella, C. H. Chiu, S. P. Creaser, J. W. Cuozzo, C. P. Davie, Y. Ding, G. J. Franklin, K. D. Franzen, M. L. Gefter, S. P. Hale, N. J. V. Hansen, D. I. Israel, J. Jiang, M. J. Kavarana, M. S. Kelley, C. S. Kollmann, F. Li, K. Lind, S. Mataruse, P. F. Medeiros, J. A. Messer, P. Myers, H. O'Keefe, M. C. Oliff, C. E. Rise, A. L. Satz, S. R. Skinner, J. L. Svendsen, L. Tang, K. Van Vloten, R. W. Wagner, G. Yao, B. Zhao and B. A. Morgan, Design, Synthesis and Selection of DNA-Encoded Small-Molecule Libraries, Nat. Chem. Biol., 2009, 5(9), 647–654, DOI:10.1038/nchembio.211.
- R. A. Goodnow, C. E. Dumelin and A. D. Keefe, DNA-Encoded Chemistry: Enabling the Deeper Sampling of Chemical Space, Nat. Rev. Drug Discovery, 2017, 16(2), 131–147, DOI:10.1038/nrd.2016.213.
- J. S. Saraya and D. K. O'Flaherty, A Facile and General Tandem Oligonucleotide Synthesis Methodology for DNA and RNA, ChemBioChem, 2024, 25(6), e202300870, DOI:10.1002/cbic.202300870.
- M.-H. Hou, S.-B. Lin, J.-M. P. Yuann, W.-C. Lin, A. H.-J. Wang and L.-S. Kan, Effects of Polyamines on the Thermal Stability and Formation Kinetics of DNA Duplexes with Abnormal Structure, Nucleic Acids Res., 2001, 29(24), 5121–5128, DOI:10.1093/nar/29.24.5121.
- T. Thomas and T. J. Thomas, Selectivity of Polyamines in Triplex DNA Stabilization, Biochemistry, 1993, 32(50), 14068–14074, DOI:10.1021/bi00213a041.
- T. Antony, T. Thomas, A. Shirahata and T. J. Thomas, Selectivity of Polyamines on the Stability of RNA−DNA Hybrids Containing Phosphodiester and Phosphorothioate Oligodeoxyribonucleotides, Biochemistry, 1999, 38(33), 10775–10784, DOI:10.1021/bi990180t.
- T. Horn and M. S. Urdea, A Chemical 5′-Phosphorylation of Oligodeoxyribonucleotides That Can Be Monitored by Trityl Cation Release, Tetrahedron Lett., 1986, 27(39), 4705–4708, DOI:10.1016/S0040-4039(00)85043-1.
- A. Guzaev, H. Salo, A. Azhayev and H. Lönnberg, A New Approach for Chemical Phosphorylation of Oligonucleotides at the 5′-Terminus, Tetrahedron, 1995, 51(34), 9375–9384, DOI:10.1016/0040-4020(95)00544-I.
- M. I. Meschaninova, D. S. Novopashina, O. A. Semikolenova, V. N. Silnikov and A. G. Venyaminova, Novel Convenient Approach to the Solid-Phase Synthesis of Oligonucleotide Conjugates, Molecules, 2019, 24(23), 4266, DOI:10.3390/molecules24234266.
- D. R. Dempsey, H. Jiang, J. H. Kalin, Z. Chen and P. A. Cole, Site-Specific Protein Labeling with N -Hydroxysuccinimide-Esters and the Analysis of Ubiquitin Ligase Mechanisms, J. Am. Chem. Soc., 2018, 140(30), 9374–9378, DOI:10.1021/jacs.8b05098.
- O. Koniev and A. Wagner, Developments and Recent Advancements in the Field of Endogenous Amino Acid Selective Bond Forming Reactions for Bioconjugation, Chem. Soc. Rev., 2015, 44(15), 5495–5551, 10.1039/C5CS00048C.
- G. W. Anderson, J. E. Zimmerman and F. M. Callahan, The Use of Esters of N-Hydroxysuccinimide in Peptide Synthesis, J. Am. Chem. Soc., 1964, 86(9), 1839–1842, DOI:10.1021/ja01063a037.
- N. G. Horton, J. S. Saraya and D. K. O'Flaherty, Novel Disulfide-Containing Linkages of Pyrimidine Nucleobases to Solid Supports: A Versatile Platform for Oligonucleotide Conjugation, Bioconjugate Chem., 2025, 36(10), 2237–2246, DOI:10.1021/acs.bioconjchem.5c00326.
- J. P. Phelan, S. B. Lang, J. Sim, S. Berritt, A. J. Peat, K. Billings, L. Fan and G. A. Molander, Open-Air Alkylation Reactions in Photoredox-Catalyzed DNA-Encoded Library Synthesis, J. Am. Chem. Soc., 2019, 141(8), 3723–3732, DOI:10.1021/jacs.9b00669.
- J.-Y. Li and H. Huang, Development of DNA-Compatible Suzuki-Miyaura Reaction in Aqueous Media, Bioconjugate Chem., 2018, 29(11), 3841–3846, DOI:10.1021/acs.bioconjchem.8b00676.
- J. H. Hunter, L. Prendergast, L. F. Valente, A. Madin, G. Pairaudeau and M. J. Waring, High Fidelity Suzuki–Miyaura Coupling for the Synthesis of DNA Encoded Libraries Enabled by Micelle Forming Surfactants, Bioconjugate Chem., 2020, 31(1), 149–155, DOI:10.1021/acs.bioconjchem.9b00838.
- R. J. Fair, R. T. Walsh and C. D. Hupp, The Expanding Reaction Toolkit for DNA-Encoded Libraries, Bioorg. Med. Chem. Lett., 2021, 51, 128339, DOI:10.1016/j.bmcl.2021.128339.
- K. Yamamoto, Y. Fuchi, M. Okabe, T. Osawa, Y. Ito and Y. Hari, New Cleavable Spacers for Tandem Synthesis of Multiple Oligonucleotides, Synthesis, 2021, 53(23), 4440–4448, DOI:10.1055/a-1538-9883.
- K. Yamamoto, Y. Fuchi, Y. Ito and Y. Hari, Expansion of Phosphoramidite Chemistry in Solid-Phase Oligonucleotide Synthesis: Rapid 3′-Dephosphorylation and Strand Cleavage, J. Org. Chem., 2023, 88(5), 2726–2734, DOI:10.1021/acs.joc.2c02195.
- R. T. Pon and S. Yu, Tandem Oligonucleotide Synthesis Using Linker Phosphoramidites, Nucleic Acids Res., 2005, 33(6), 1940–1948, DOI:10.1093/nar/gki333.
- R. T. Pon, S. Yu and Y. S. Sanghvi, Multiple Oligonucleotide Synthesis in Tandem on Solid-Phase Supports for Small and Large Scale Synthesis, Nucleosides, Nucleotides Nucleic Acids, 2001, 20(4–7), 985–989, DOI:10.1081/NCN-100002474.
- R. T. Pon, S. Yu and Y. S. Sanghvi, Tandem Oligonucleotide Synthesis on Solid-Phase Supports for the Production of Multiple Oligonucleotides, J. Org. Chem., 2002, 67(3), 856–864, DOI:10.1021/jo0160773.
- S. Slott and K. Astakhova, MicroRNA Pools Synthesized Using Tandem Solid-Phase Oligonucleotide Synthesis, J. Org. Chem., 2023, 88(14), 9760–9768, DOI:10.1021/acs.joc.3c00376.
- F. Ferreira, A. Meyer, J.-J. Vasseur and F. Morvan, Universal Solid Supports for the Synthesis of Oligonucleotides via a Transesterification of H -Phosphonate Diester Linkage, J. Org. Chem., 2005, 70(23), 9198–9206, DOI:10.1021/jo051172n.
- M. Madaoui, A. Meyer, J. Vasseur and F. Morvan, Thermolytic Reagents to Synthesize 5′– or 3′–Mono(Thio)Phosphate Oligodeoxynucleotides or 3′–modified Oligodeoxynucleotides, Eur. J. Org. Chem., 2019,(17), 2832–2842, DOI:10.1002/ejoc.201900302.
- P. M. Hardy, D. Holland, S. Scott, A. J. Garman, C. R. Newton and M. J. McLean, Reagents for the Preparation of Two Oligonucleotides per Synthesis (TOPSTM), Nucleic Acids Res., 1994, 22(15), 2998–3004, DOI:10.1093/nar/22.15.2998.
- B. Xiao, S. Wang, Y. Pan, W. Zhi, C. Gu, T. Guo, J. Zhai, C. Li, Y. Q. Chen and R. Wang, Development, Opportunities, and Challenges of siRNA Nucleic Acid Drugs, Mol. Ther. – Nucleic Acids, 2025, 36(1), 102437, DOI:10.1016/j.omtn.2024.102437.
- H. Zhu, H. Zhang, Y. Xu, S. Laššáková, M. Korabečná and P. Neužil, PCR Past, Present and Future, BioTechniques, 2020, 69(4), 317–325, DOI:10.2144/btn-2020-0057.
| This journal is © The Royal Society of Chemistry 2026 |