Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Control of mucoid Pseudomonas aeruginosa biofilms in a highly porous skin model (ex vivo/in vitro model) with carbosilane dendrimer-endolysin complexes
Karolina
Lach
a,
Samuel
Takvor-Mena
b,
Oscar
Barrios-Gumiel
b,
Javier
Sanchez-Nieves
 bc,
Jacek
Kuchinka
a,
Piotr
Furmańczyk
d,
Małgorzata
Łysek-Gładysińska
a and
Karol
Ciepluch
bc,
Jacek
Kuchinka
a,
Piotr
Furmańczyk
d,
Małgorzata
Łysek-Gładysińska
a and
Karol
Ciepluch
 *e
*e
aDivision of Medical Biology, Jan Kochanowski University, 25-406 Kielce, Poland
bDepartment of Organic and Inorganic Chemistry, Research Institute in Chemistry “Andres M. del Río” (IQAR), University of Alcaláá (UAH), 28805 Alcala de Henares, Madrid, Spain
cNetworking Research Center for Bioengineering, Biomaterials and Nanomedicine (CIBERBBN), Av. Monforte de Lemos, 3-5, 28029 Madrid, Spain
dDepartment of Machine Design, Faculty of Mechatronics and Mechanical Engineering, Kielce University of Technology, 25-314 Kielce, Poland
eDepartment of Basic Medical Sciences, Faculty of Medical Sciences and Health Sciences, Casimir Pulaski University of Radom, Radom, Poland. E-mail: k.ciepluch@urad.edu.pl
First published on 2nd March 2026
Abstract
The alarming rise in antibiotic resistance in Pseudomonas aeruginosa, resulting from its ability to undergo enzymatic modifications, mutate, or form complex structures such as biofilms, has rendered many conventional therapies ineffective, creating an urgent need to search for new antimicrobial agents. Ligand-modified nanoparticles are a promising alternative to conventional agents. For example, silver-containing dendrimers (metallodendrimers), which permeabilize the bacterial outer membrane, enhance the production of reactive oxygen species, and, in combination with peptidoglycan-degrading proteins—CHAP-domain endolysins—can cross the barrier of Gram-negative bacteria. However, their activity against mature biofilms and potential cytotoxicity to mammalian tissues remain poorly understood. In this study, we used an ex vivo pig ear skin model to evaluate the ability of three carbosilane dendrimers and their combinations with endolysin to inhibit biofilm formation with P. aeruginosa PAO1. The irregular skin texture and richer nutrient resources in the ex vivo model may limit the effectiveness of nanomaterials compared to the biophilic P. aeruginosa in the in vitro model on a smooth plastic surface. Early application of dendrimers with endolysin significantly reduces bacterial survival, while late application causes excessive damage to the epidermis, making it difficult for dendrimers to reach the bacteria and hindering the assessment of the effectiveness of antibacterial drugs. These results clearly suggest that studies on the antibacterial activity of various nanomaterials on smooth surfaces differ significantly from the highly porous structure of the skin model, which has an important impact on their limited application in the treatment of skin wounds.
1. Introduction
In 2028, we will celebrate the 100th anniversary of the discovery of the first antibiotic, penicillin, by Alexander Fleming, which changed the outlook for fighting infections. However, it was not expected that within 100 years, bacteria would acquire such resistance to these types of compounds that it would be almost impossible to fight them. Current medicines face a serious challenge due to the resistance of bacteria, which causes the deaths of millions each year.1,2 The problem is amplified by a decrease in the production of new and effective antibiotics, which limits the ability to successfully treat infections. Current therapies usually rely on modifications of existing drugs, but these strategies provide only a temporary solution.3,4 With conventional antibiotics losing their effectiveness, it has become necessary to search for alternative treatments that are capable of combating MDR (multi-drug resistant) pathogens.One of the most common and “critical” pathogens responsible for hospital-acquired infections is Pseudomonas aeruginosa.5,6 It is a rod-shaped, opportunistic, enveloped and highly adaptive Gram-negative bacterium that is widely distributed in aquatic and soil environments and is capable of growth under both aerobic and oxygen-limited conditions (facultatively anaerobic). Its remarkable metabolic flexibility allows it to survive in a variety of conditions.7,8P. aeruginosa is particularly dangerous for immunosuppressed patients, in whom it can lead to chronic and nosocomial infections, hard-to-treat wounds, and urinary tract infections (UTI); it is also associated with pneumonia and bloodstream and surgical site infections.9,10
Mechanisms of bacterial resistance include limited cell membrane permeability, active antibiotic removal systems (efflux pumps), various mutations that reduce drug sensitivity, and horizontal gene transfer that allows rapid transmission of resistance traits.10–13 One of the most effective ways in which Pseudomonas aeruginosa avoids antibiotics is its ability to form a biofilm.
A bacterial biofilm is a highly organized three-dimensional community of microorganisms embedded in the extracellular polymeric substances (EPS), consisting mainly of exopolysaccharides, such as alginate, proteins and nucleic acids.14,15 Bacteria adhere tightly to each other and/or to non-biological solid surfaces, such as catheters or endoprostheses, biological surfaces, such as wounds, burn injuries, ulcers, or cystic fibrosis.11,16 Bacteria possess many features that help them attach to a surface, such as hydrophobicity of the cell wall, surface potential, size and shape, the presence of quorum sensing, pili and curli, and the ability to produce extracellular adhesins. However, it is important to note that bacterial adhesion to a surface has different levels, depending on several surface factors, such as hydrophobicity, heterogeneity, patterns, roughness, stiffness, surface potential and charge. In addition, environmental conditions are crucial for bacterial attachment, e.g., pH, temperature and ionic strength.17–19 Free-living/planktonic bacteria, once in contact with the surface of a medical device or injured tissue, due to the action of van der Waals forces, the presence of adhesins, type IV pili, and lipopolysaccharide (LPS), start producing extracellular polymers that organize into a spatial matrix, enabling firm adhesion and biofilm development.20,21 The process of biofilm formation consists of several steps: (1) the adhesion of individual bacterial cells to the surface, which can take place through various surface structures, such as fimbriae, pili, or adhesion substances; (2) bacteria enter the microcolony phase, and their cell divisions lead to an increase in the number of cells within the structure; (3) biofilm maturation, during which the bacteria produce an extracellular matrix (EPS) that provides stability and protection against external agents.22,23
In the mature biofilm, intensive cell–cell interactions take place, and the bacteria diversify in terms of metabolic functions, which promotes their survival under changing conditions.24 In time, a dispersal phase occurs, in which some cells detach from the biofilm and can colonize new environments. The structure of the biofilm makes bacteria embedded in it up to hundreds or thousands of times more resistant to antibiotics than their planktonic analogs.25,26 This is due to several factors, including the limited penetration of drugs deep into the biofilm, the presence of spore forms, the ability of bacteria to enter a metabolic dormant state, the secretion of protective substances, and many others, as mentioned above, which reduce their vulnerability to drugs. The biofilm also provides protection from the immune system's response as a result of dynamic changes in gene expression, resulting in the differentiation of surface antigens and leading to the chronic nature of the infection.21 Because of these properties, biofilms are a major cause of difficult-to-treat infections. The increase in antibiotic resistance and the ability of bacteria to form biofilms make classic treatments less and less effective.27,28 Consequently, the search is on for new therapeutic strategies that could inhibit biofilm formation or effectively eliminate it. Research into new methods of biofilm control is crucial for the future of treating bacterial infections and reducing the spread of multidrug-resistant strains.
Nanomaterials, such as dendrimers, also known as ligand-modified nanoparticles, are widely known for their antimicrobial properties and are a promising alternative to the currently studied non-functioning antibiotics.29 The improvement of their structure and therapeutic potential is the subject of ongoing research. It is important to note that dendrimers can differ significantly from one another, and these differences in structure and surface modifications result in different mechanisms of antimicrobial action. Additional modification with metal ions, such as silver, zinc, copper or gold, as well as functional groups, including imidazolium or pyrimidine derivatives, has been shown to significantly increase their antimicrobial and biological efficacy.30,31
There is increasing number of evidence that demonstrates the potential of combining dendrimers with bacteriophages or antimicrobial proteins (AMPs), such as lysins, to enhance antibacterial activity.32,33 Endolysins, for instance, are capable of specifically targeting and degrading bacterial peptidoglycan by cleaving bonds within peptide cross-links, such as tetra-peptides and penta-glycine bridges (endolysin CHAP domain). In Gram-positive bacteria, endolysins can directly access and degrade the exposed peptidoglycan.34–36 In contrast, the presence of an outer membrane (OM) in Gram-negative bacteria provides a significant barrier, which requires the use of membrane-permeabilizing agents, such as dendrimers, to enable the effective action of endolysins.37 Based on numerous scientific reports, the use of dendrimers containing silver ions, particularly in combination with antimicrobial proteins, such as endolysin, represents a promising strategy for the effective elimination of planktonic bacterial cells. These compounds exhibit antibacterial effects through multiple mechanisms, including permeabilization of the outer membrane, degradation of the peptidoglycan layer, and increased reactive oxygen species (ROS) production.38–40
Dendrimers are widely used as drug and gene carriers in nanomedicine and targeted pharmacology.41 Appropriately designed and modified, they can efficiently transport active substances without adversely affecting target cells or tissues.42 Nevertheless, in addition to their antimicrobial properties, dendrimers can also exhibit harmful effects on eukaryotic cells, manifested as cytotoxicity or genotoxicity. According to scientific reports, the intensity of these effects can be strongly dependent on the structure of the dendrimer, the type of surface ligands and the concentration used.43 Since it is known that dendrimers containing silver ions can enhance the production of reactive oxygen species (ROS) and cationic dendrimers permeabilize the outer membrane in bacteria, there is a high probability that a similar effect will also occur in eukaryotic cells. This can lead to increased cytotoxicity, resulting in membrane destabilization and oxidative damage to lipids, proteins, and genetic material (DNA and RNA). As a consequence, dysfunction of cellular organelles, including mitochondria, can occur, and long-term accumulation of free radicals can trigger pathways leading to cell death (caspase-3 enzyme or p53 gene after genotoxic stress initiates apoptosis).44 The use of antimicrobial proteins such as endolysins is considered safe in terms of cytotoxicity because eukaryotic cells do not contain peptidoglycan, which is the main substrate of these enzymes. Consequently, endolysins have no activity against mammalian cells, making them promising candidates for therapeutic applications.
While dendrimers and antimicrobial proteins hold significant therapeutic potential, especially as targeted nanocarriers, their broader biomedical application requires a thorough understanding of their toxicity and biological interactions. Although the initial results are promising, there is a persistent lack of data regarding their effects on complex bacterial structures, such as biofilms, and eukaryotic cells or tissues. Therefore, further in vitro and ex vivo/in vivo studies are essential to optimize therapeutic efficacy while minimizing adverse effects.
This study aimed to evaluate the inhibition of biofilm formation by planktonic Pseudomonas aeruginosa cells using both an in vitro laboratory model and an ex vivo approach utilizing pork ear skin. It is crucial to understand not only how bacterial biofilms respond to treatment under controlled laboratory conditions, but also how they might behave during therapeutic interventions targeting human or animal infections. Additionally, the influence of a complex of dendrimers and endolysin38,39 on normal, uninfected skin was examined to allow a comparative analysis between healthy and P. aeruginosa-infected tissue. Prior to testing the dendrimers and endolysin on ex vivo skin models, it was necessary to assess the cytotoxic effects of the compounds by evaluating the viability of VH10 fibroblast cells exposed to various concentrations of each tested agent.
2. Materials and methods
2.1. Carbosilane dendrimers
The silver and imidazolium metal carbosilane dendrimers Ag-D1,45Ag-D2 and Im-D238 (Scheme 1) were prepared according to the procedures described previously.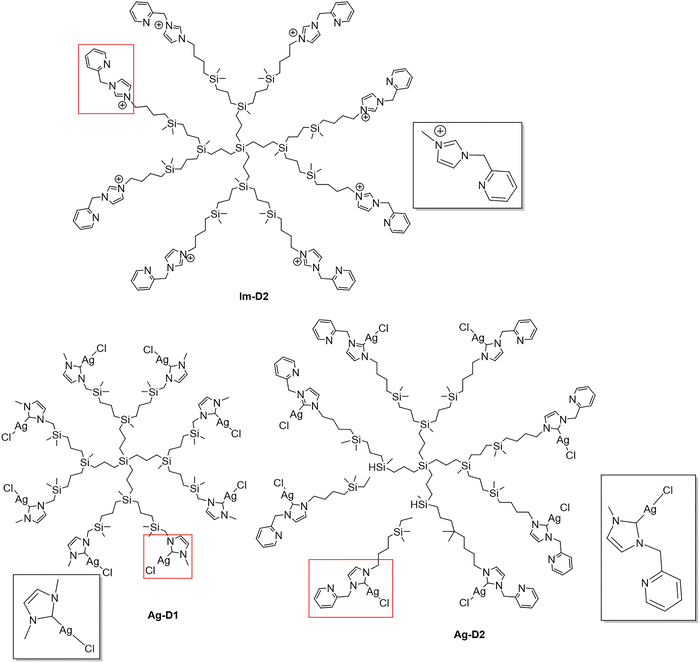 | ||
| Scheme 1 Structures of CBS dendrimers Im-D2 (chloride anions omitted for clarity), Ag-D1 and Ag-D2. The functional groups are highlighted in red squares. | ||
2.2. Recombinant endolysin
The recombinant CHAP endolysin fragment of the phage Φ812 catalytic domain was purchased from Protean, Czech Republic.2.3. Bacterial biofilm formation assay
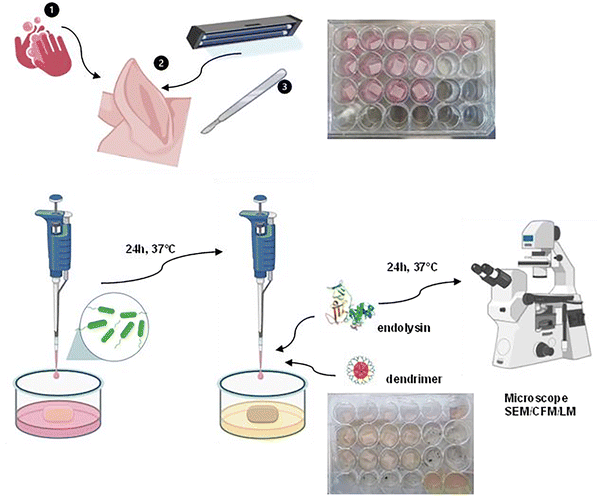
To analyze the formation of the P. aeruginosa PAO1 bacterial biofilm (Scheme 2), the culture medium was prepared using DMEM (Dulbecco's Modified Eagle Medium) containing L-glutamine and phenol red and further enriched with 10% fetal bovine serum (FBS). For the experiment, 24-well culture plates were used, in which previously prepared and UV-sterilized filter paper discs, matched in size to the diameter of the wells, were placed. Here, 250 µL of DMEM medium was added to each well, followed by the placement of sterile skin sections (1 × 1 cm). The bacteria were prepared as described above. Next, 10 µL of diluted bacteria were spotted in the center of a piece of skin and allowed to form a biofilm under the conditions of 37 °C & 5% CO2 for 24 hours. Appropriate volumes of dendrimers (8 and 32 µg mL−1) and endolysin (2 µg mL−1) and fresh DMEM medium (appropriate volume, in total 400 µL) were added to the formed biofilm on skin and incubated at 37 °C with 5% CO2 for 24 hours. After this time, the old medium was gently removed, followed by washing twice with dH2O. The biofilm formed on the skin was stained by spotting 30 µL of dye and left for 20 min in the dark. The dye was then rinsed three times with distilled water and the skin was gently transferred with tweezers to a coverslip. The prepared samples were observed under a confocal microscope, no longer than 30 min after staining.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, and 1
1, and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, and then they were suspended in pure propylene oxide. The samples were placed on a glass slide and left to dry completely, then placed on carbon discs, covered with gold using a JEOL JFC 110E Fine Coat Ion Sputter, and analyzed by SEM (JEOL JSM-7100F, JEOL Ltd, Japan) at an accelerating voltage of 15 kV.
3, and then they were suspended in pure propylene oxide. The samples were placed on a glass slide and left to dry completely, then placed on carbon discs, covered with gold using a JEOL JFC 110E Fine Coat Ion Sputter, and analyzed by SEM (JEOL JSM-7100F, JEOL Ltd, Japan) at an accelerating voltage of 15 kV.
2.4. Cytotoxic effect on fibroblasts
The VH10 human fibroblast cell line (ATCC, Manassas, VA, USA; catalogue no. PCS-201-012) was maintained and cultured in Fibroblast Growth medium (Cat no. 116–500, Cell Application, Inc.) at 37 °C in a humidified atmosphere with 5% CO2. The culture medium was changed every 2 days. The viability of fibroblast cells treated with dendrimers (concentrations of 1, 2, 4, 8, 16, 32, 64, 128 µg mL−1) and their complexes with endolysin (at concentrations of 1 and 2 µg mL−1) was determined using an MTS assay. The fibroblasts were seeded in 96-well plates. After reaching the appropriate confluence, the cells were treated with dendrimers, endolysin, and dendrimer + endolysin for 24 h. The MTS Cell Proliferation Assay Kit (Colorimetric) (Abcam) was used according to the manufacturer's recommendations, and absorbance at 490 nm was measured using a TECAN SPARK Magellan V2.2 STD spectrometer (Tecan Group Ltd, Switzerland).3. Results and discussion
3.1. Bacterial biofilm formation in the presence of dendrimers and complexes with endolysin – in vitro studies
Fig. 1 shows the effect of the tested dendrimers on mature P. aeruginosa PAO1 bacterial biofilm after 24 hours of incubation at various concentrations. In each of the three systems analyzed, a relationship was observed between the concentration of the dendrimer and the degree of inhibition of bacterial growth. For the metallodendrimer Ag-D1 alone (Fig. 1A), none of the concentrations used caused a reduction in biofilm, and an increase in its amount was even observed. Only the highest concentration, 128 µg mL−1, completely degraded the biofilm and bacteria (∼14%). Similarly, the addition of endolysin to selected concentrations, 8, 32, and 64 µg mL−1 did not cause significant degradation of the biofilm. When the imidazolium Im-D2 dendrimer alone was used (Fig. 1B), as described above, no reduction in biofilm quantity was observed, while an increase in its mass was noted at all concentrations tested, reaching an average of ∼161% of the control value. However, the addition of endolysin to the Im-D2 dendrimer at concentrations of 8, 32, and 64 µg mL−1 resulted in a significant decrease in biofilm quantity, to an average value of ∼40% compared to the control. A similar trend was observed for the metallodendrimer Ag-D2 dendrimer (Fig. 1C). The use of the dendrimer alone did not inhibit biofilm formation; an upward trend was observed for all concentrations tested, reaching an average of ∼157%. In contrast, in the presence of endolysin, a significant reduction in biofilm quantity was observed, particularly at a concentration of 8 µg mL−1, where the value dropped to approximately 48%, and when using the highest concentration of the dendrimer alone (128 µg mL−1), which resulted in a further reduction in biofilm quantity to approximately 21% of the control value.To confirm the reduced viability of bacteria in formed biofilm due to CBS dendrimers and endolysin, a FilmTracerTM LIVE/DEADTM Biofilm Viability was performed. Fig. 2A represents the untreated bacterial biofilm used as a control; among a large number of living normal bacteria and stable biofilm, a few red spots appeared. A similar effect was observed for biofilm in the presence of endolysin (Fig. 2B). Higher numbers of red spots were observed near the biofilm exhibiting green fluorescence, but with reduced density after 8 µg mL−1Ag-D1 treatment, compared to the control (Fig. 2C). Applying 8 µg mL−1Im-D2 (Fig. 2D) caused an increase in the number of dead cells and a decrease in living cells was observed among the dense biofilm; orange and red signals dominated, with a small amount of green. A similar effect was observed for 8 µg mL−1Ag-D2 (Fig. 2E), but with a lower intensity of dead cells. Treatment of bacterial biofilm with the additional presence of endolysin did not significantly reduce the biofilm structure (Fig. 2F − 8 µg mL−1Ag-D1 + 2 µg mL−1 endolysin), but several red signals were observed. Fig. 2G presents bacterial biofilm after 8 µg mL−1Im-D2 + 2 µg mL−1 endolysin, where a reduction in biomass was observed, while small amounts of red spots were present. In the case of 8 µg mL−1Ag-D2 + 2 µg mL−1 endolysin (Fig. 2H), the red signal dominated the green one, and the biofilm biomass was reduced.
The use of a higher dose of dendrimers (32 µg mL−1) Ag-D1 (Fig. 2I), Im-D2 (Fig. 2J) and Ag-D2 (Fig. 2K) did not significantly increase the number of dead bacterial cells or decrease the biofilm biomass compared to the lower dose used. Moreover, the additional presence of endolysin with higher doses of dendrimers resulted in an increase in the number of red spots and a reduction in biomass after Ag-D1 + 2 µg mL−1 endolysin (Fig. 2L), while after Im-D2 + 2 µg mL−1 endolysin (Fig. 2M), the biofilm was more condensed, and in the case of Ag-D2 + 2 µg mL−1 endolysin (Fig. 2N), it was diluted and thin.
To explain the mechanism of biofilm inhibition by the tested compounds, both the activity of silver ions and the specific properties of dendrimers must be taken into account. Ag+ ions, present in dendrimers Ag-D1 and Ag-D2, are characterized by high affinity for thiol groups, especially cysteine, which is present in numerous enzymes responsible for the metabolism of exopolysaccharides. The breaking of S–H bonds leads to a weakening of bacterial cell adhesion to the substrate and disruption of the quorum-sensing system. Besides, dendrimers with Ag+ ions increase the permeability of the bacterial OM and cause an increase in ROS production, leading to planktonic cell disintegration and death,39,46 which can occur in biofilm. Additionally, a small dose of the tested compound may increase EPS and protein matrix production, which would change the biofilm structure, and the metabolic activity of bacterial cells would decrease due to the disruption of intracellular processes.47 At the same time, the electrical charge of the EPS matrix, which is mainly negative, must be taken into account. Positively charged dendrimers can be strongly attracted to the biofilm by electrostatic forces, which may facilitate the delivery of silver ions and increase matrix penetration. Moreover, it may promote troublesome aggregation of dendrimers or bacterial cells, leading to increased biofilm formation or changing physical properties, such as density and adhesion capacity to the surface.48
3.2. Bacterial biofilm formation on skin tissue in the presence of dendrimers and complexes with endolysin – ex vivo studies
The skin tissue consists of three main layers: the epidermis with the stratum corneum (the outermost part of the epidermis), the dermis, and the subcutaneous tissue. It acts as a basic protective barrier against pathogens, thanks to several key mechanisms: acidic pH (around 5.5), production of antimicrobial proteins (lysozyme, cathelicidin, β-defensins, lactoferrin, and others), and the presence of salts. The stratum corneum, composed of keratinized cells, prevents the permanent adhesion of microorganisms and promotes their removal through continuous exfoliation. Although it contains few nutrients, it is rich in keratin, which some bacteria can use as a food source.49The surface of the skin, unlike the smooth surface of a 24- or 96-well laboratory plate, is characterized by numerous depressions, protrusions, and pores that drain sebaceous and sweat gland secretions, as well as hair deposits (Fig. 3). These microscopic structures provide excellent anchorage points for bacteria, which can colonize them and develop biofilms under natural conditions or infection. The resulting complex bacterial matrix is then highly resistant to mechanical and chemical removal methods, which significantly hinders the effective treatment of skin infections. Furthermore, due to the presence of cholesterol and other lipids in the stratum corneum, the surface charge of the skin is mostly negative, which is important for bacterial adhesion.50,51
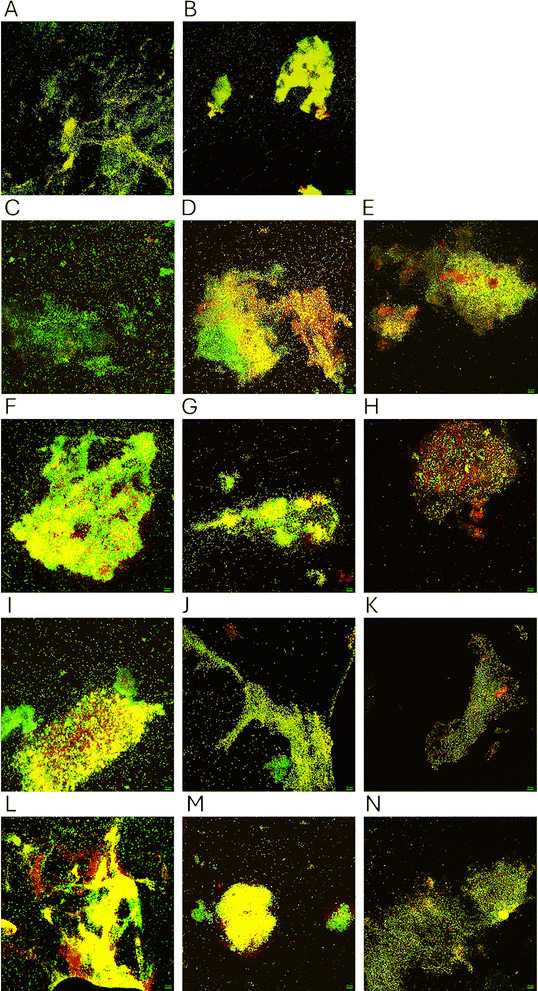
To assess the effects of the CBS imidazolium dendrimer and metallodendrimers and endolysin on the formed biofilm of Pseudomonas aeruginosa PAO1, an ex vivo model was used on pork skin pieces, which, after infection and treatment with dendrimers, were subjected to staining to differentiate between live and dead bacterial cells in the biofilm to be analyzed by confocal fluorescence microscopy. The results are shown in Fig. 4. Fig. 4A shows a dense, unchanged biofilm of the P. aeruginosa PAO1 (control group). After treatment with endolysin alone (2 µg mL−1, Fig. 4B), no significant changes were observed – the biofilm structure was characterized by high bacterial viability, comparable to the control. Treatment of the biofilm with Ag-D1 at a concentration of 8 µg mL−1 (Fig. 4C) led to partial thinning of the matrix, with gaps appearing in the lower layers and red signals indicating dead cells. Similar changes, with a predominance of intense green fluorescence of surface bacteria and periodic red foci, were observed after the application of Im-D2 (8 µg mL−1, Fig. 4D) and the Ag-D2 complex (8 µg mL−1, Fig. 4E). The addition of endolysin increased the number of dead cells in the biofilm in combination with Ag-D1 (Fig. 4F) and Ag-D2 (Fig. 4H), and a clear increase in red signal was observed, while in combination with Im-D2 (Fig. 4G), this effect was weaker. Increasing the concentration of dendrimers to 32 µg mL−1 enhanced the bactericidal effect. AgD1 (Fig. 4I) and Im-D2 (Fig. 4J) caused a significant increase in the red signal, while Ag-D2 (Fig. 4K) slightly increased the proportion of dead cells. The combination of 32 µg mL−1Ag-D1 with endolysin (Fig. 4L) showed the highest efficacy, as the biofilm then contained mostly dead bacteria with a minimal proportion of live ones. Under similar conditions, the Im-D2 + endolysin complex (Fig. 4M) showed a slightly weaker effect, while Ag-D2 + endolysin (Fig. 4N) showed a moderate but noticeable improvement in efficacy compared to the dendrimer alone.
Scanning electron microscopy (SEM) was used to evaluate bacterial biofilm morphology after treatment with dendrimers and their endolysin complexes (Fig. 5), and the results are summarized in Table 1. The negative control group (Fig. 5A) shows a sterile, uninfected skin surface. The positive control group (Fig. 5B) shows a properly formed biofilm and intact bacterial cell structure. The administration of endolysin at a concentration of 2 µg mL−1 (Fig. 5C) did not significantly reduce EPS continuity; the matrix remained dense and wavy and was associated with moderate visibility of bacteria with an intact appearance. In contrast, treatment of the biofilm with 8 µg mL−1Ag-D1 (Fig. 5D) resulted in a reduction in the compact EPS structure, which became looser, and the bacteria, despite their very high numbers, exhibited shortened, irregular shapes. The use of 8 µg mL−1Im-D2 (Fig. 5E) caused a weakened and flattened EPS matrix, restoring the average level of bacteria and their normal appearance. A lower dose of Ag-D2 (8 µg mL−1, Fig. 5F) resulted in dense, wavy EPS and poorly visible bacteria. The combination of Ag-D1 with endolysin (8 µg mL−1Ag-D1 + 2 µg mL−1 endolysin, Fig. 5G) resulted in dilution and cracking of the EPS. Im-D2 at a concentration of 8 µg mL−1 and 2 µg mL−1 endolysin (Fig. 5H) caused a compact and dense EPS layer in which the visibility of bacteria was low, and their shapes were irregular. The addition of endolysin to Ag-D2 (Fig. 5I) diluted the EPS but increased the visibility of the bacteria, which retained their irregular shape. A high dose of Ag-D1 (Fig. 5J) resulted in a dense, layered EPS and, at the same time, low visibility of irregularly shaped cells. A concentration of 32 µg mL−1 of Im-D2 (Fig. 5K) led to severe matrix fibrosis, low bacterial visibility, and cell wall disruption. The use of Ag-D2 at a concentration of 32 µg mL−1 (Fig. 5L) resulted in diluted EPS and high visibility of cells. For Ag-D1 in combination with endolysin (Fig. 5M), the matrix became compact and wrinkled, and the visibility level dropped to average values. For Im-D2 (Fig. 5N) in combination with endolysin, EPS remained dense, and bacteria were hardly visible due to abundant coverage. Finally, the Ag-D2 complex with endolysin (Fig. 5O) led to a dilution of the matrix and an increase in the number of damaged cells, which exhibited numerous cell wall defects.
| Group | EPS | Bacterial visibility | Appearance of bacteria | |
|---|---|---|---|---|
| A. | Control | None | None | — |
| B. | PAO1 | Present, dense | High | Undamaged, normal |
| C. | 2 µg mL−1 E | Present, dense, undulating | Medium | Normal |
| D. | 8 µg mL−1Ag-D1 | Present, less compact | Very high | Shortened, irregular shape |
| E. | 8 µg mL−1Im-D2 | Weakened, flat | Medium | Normal |
| F. | 8 µg mL−1Ag-D2 | Present, dense, undulating | Low (EPS coated) | |
| G. | 8 µg mL−1Ag-D1 + 2 µg mL−1 E | Diluted, cracked, delaminated | Low (EPS coated) | |
| H. | 8 µg mL−1Im-D2 + 2 µg mL−1 E | Present, dense, compact | Low (EPS coated) | Irregular shapes |
| I. | 8 µg mL−1Ag-D2 + 2 µg mL−1 E | Diluted | High | Irregular shapes |
| J. | 32 µg mL−1Ag-D1 | Dense, stratified | Low (EPS coated) | |
| K. | 32 µg mL−1Im-D2 | Severely fibrosis | Low (EPS coated) | |
| L. | 32 µg mL−1Ag-D2 | Diluted | High | Irregular shapes |
| M. | 32 µg mL−1Ag-D1 + 2 µg mL−1 E | Undulating, cracked, dense | Medium | Damaged, irregular shapes |
| N. | 32 µg mL−1Im-D2 + 2 µg mL−1 E | Present, dense | Low (EPS coated) | Numerous cell wall damages |
| O. | 32 µg mL−1 2Ag-D2 + 2 µg mL−1 E | Diluted | High | |
3.3. Histomorphology of uninfected and Pseudomonas aeruginosa-infected pork skin after treatment with dendrimers and their complexes with endolysin
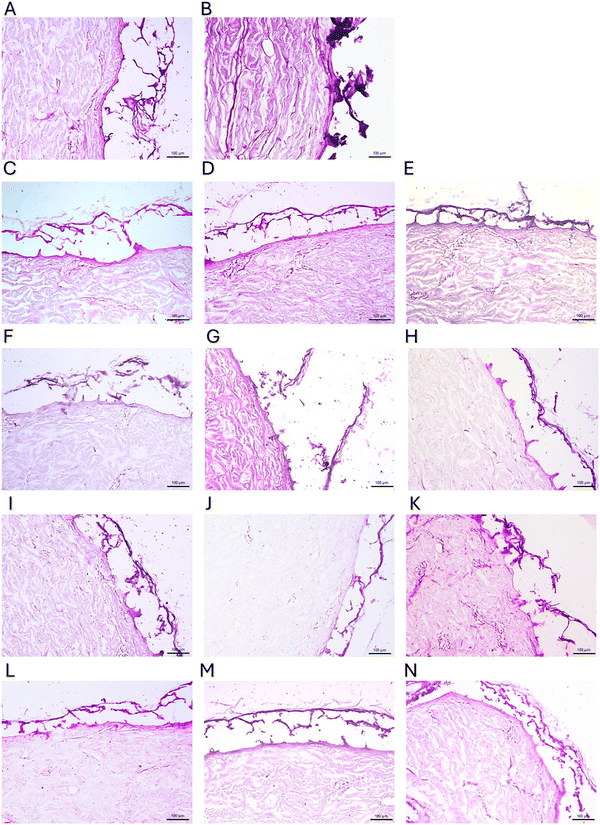
The effects of individual therapies on ex vivo skin structures, indicating changes in the stratum corneum (SC), stratum lucidum (SL), stratum granulosum (SG), stratum spinosum (SS), stratum basale (SB) and dermis (D), are shown in Fig. 6. In the control group (Fig. 6A) and after the application of endolysin (2 µg mL−1) (Fig. 6B), the entire skin structure remained intact. Treatment with the Ag-D1 dendrimer (8 µg mL−1) (Fig. 6C) caused local flattening (↕) and abrasion of keratinocytes in the stratum corneum, without significant changes in the deeper layers of the epidermis; the dermis remained intact. The addition of endolysin Ag-D1 (8 µg mL−1 + 2 µg mL−1) (Fig. 6F) also caused local flattening of the stratum corneum, while hypertrophy (★) of some cells was observed in the spinous layer. A higher concentration of Ag-D1 (32 µg mL−1) (Fig. 6I) caused dilatation of the stratum corneum, disruption of the continuity (↑) of the cells of the stratum spinosum with the stratum basale of the epidermis, atrophy of the cells of the stratum spinosum, without changes in the dermis. The complex of 32 µg mL−1Ag-D1 + 2 µg mL−1 endolysin (Fig. 6L) caused thickening of the stratum corneum, hyperplasia and hypertrophy of keratinocytes in the stratum spinosum, while maintaining the integrity of the dermis. Significant hyperplasia of the stratum corneum and stratum spinosum was observed, without significant changes in the dermis after using the pyrimidine imidazolium Im-D2 - 8 µg mL−1 (Fig. 6D) and its complex with endolysin (8 µg mL−1Im-D2 + 2 µg mL−1 endolysin) (Fig. 6G). At a higher concentration of 32 µg mL−1Im-D2 (Fig. 6J), a strong flattening of the stratum corneum was observed, with a break in continuity between the SC and SL, while the dermis remained unchanged. The complex with endolysin (Fig. 6M) caused incomplete atrophy of the stratum corneum, as well as hypertrophy of the cells of the stratum spinosum, with the dermis remaining intact. The introduction of pyrimidine Ag-D2 (8 µg mL−1) (Fig. 6E) led to significant SC atrophy and severe hyperplasia of the spinous layer, while the dermis remained unchanged. The complex of 8 µg mL−1Ag-D2 + 2 µg mL−1 endolysin (Fig. 6H) caused the same changes as in the case of the dendrimer itself. The use of a higher concentration of Ag-D2 (32 µg mL−1) (Fig. 6K) caused local reduction and disruption between the stratum corneum and stratum lucidum, without significant changes in the stratum spinosum and dermis, while the addition of endolysin (32 µg mL−1 + 2 µg mL−1) (Fig. 6N) caused sporadic disruption of the continuity between the stratum corneum and the stratum lucidum, while maintaining the normal thickness of the stratum corneum and leaving the stratum spinosum and dermis unchanged. The reduction in the toxicity of dendrimers in the presence endolysin could be due to their complexation. Due to its charge, the protein is able to bind to the dendrimer, thus obscuring the active surface groups. Furthermore, local aggregation of dendrimers in the presence of endolysin can quite often occur, thus reducing their properties. All measurements of the thickness of the stratum corneum and the surface area of individual inner layers of the epidermis, i.e., the granular layer, spinous layer, and basal layer, are summarized in Table 2. Overall, the tested compounds have concentration-dependent effects on the structure of the epidermis, while maintaining the integrity of the dermis, and the presence of endolysin tended to mitigate the extent of epidermal disruption induced by the dendrimers, although the magnitude of this effect varied between dendrimer types and concentrations.| Mean thickness of the stratum corneum (SC) [µm] | Mean surface area of the stratum granulosum, stratum spinosum and stratum basale (SG + SS + SB) [µm2] | |||
|---|---|---|---|---|
| Control | 26.68 ± 3.68 | p < 0.001 | 89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 008.5 ± 11 008.5 ± 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 367,57 367,57 |
p < 0.001 |
| Endolysin 2 µg mL−1 | 16.01 ± 6.6 | 52![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 648.05 ± 12 648.05 ± 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 989.59 989.59 |
||
| 8 µg mL−1Ag-D1 | 14.04 ± 4.65 | 42![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 352.25 ± 10 352.25 ± 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 198.6 198.6 |
||
| 8 µg mL−1Im-D2 | 22.69 ± 6.59 | 34![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 443.78 ± 7608.09 443.78 ± 7608.09 |
||
| 8 µg mL−1Ag-D2 | 13.59 ± 3.49 | 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 013.19 ± 5580.38 013.19 ± 5580.38 |
||
| 8 µg mL−1Ag-D1 + 2 µg mL−1 E | 10.02 ± 3.34 | 58![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 781.8 ± 7462.56 781.8 ± 7462.56 |
||
| 8 µg mL−1Im-D2 + 2 µg mL−1 E | 14.51 ± 5.44 | 64![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 693.54 ± 10 693.54 ± 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 175.75 175.75 |
||
| 8 µg mL−1Ag-D2 + 2 µg mL−1 E | 14.79 ± 4.94 | 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 489.31 ± 6394.41 489.31 ± 6394.41 |
||
| 32 µg mL−1Ag-D1 | 16.1 ± 4.47 | 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 132.42 ± 13 132.42 ± 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 635.32 635.32 |
p = 0.008 | |
| 32 µg mL−1Im-D2 | 12.63 ± 3.97 | 52![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 961.31 ± 8276.02 961.31 ± 8276.02 |
p < 0.001 | |
| 32 µg mL−1Ag-D2 | 12.69 ± 3.19 | 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 602.58 ± 4384.04 602.58 ± 4384.04 |
||
| 32 µg mL−1Ag-D1 + 2 µg mL−1 E | 16.26 ± 5.95 | 51![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 156.31 ± 7975.58 156.31 ± 7975.58 |
||
| 32 µg mL−1Im-D2 + 2 µg mL−1 E | 13.66 ± 3.89 | 52![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 731.35 ± 6431.55 731.35 ± 6431.55 |
||
| 32 µg mL−1Ag-D2 + 2 µg mL−1 E | 21.28 ± 5.98 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 038.99 ± 7576.00 038.99 ± 7576.00 |
Structural changes in the biofilm of Pseudomonas aeruginosa PAO1 and layers of skin after 24 hours of treatment with dendrimers at concentrations of 8 and 32 µg mL−1, their complexes with endolysin (2 µg mL−1) are presented at Fig. 7. In the control group (Fig. 7A) with the stratum corneum (SC) detached in places and surrounded by dense biofilm, the epidermis (E) was degraded, while the dermis (D) was partially degraded by bacteria, which manifested itself in places with a flat surface. The administration of endolysin (Fig. 7B) did not change this condition. Treatment with 8 µg mL−1Ag-D1 (Fig. 7C) and 8 µg mL−1Ag-D1 + 2 endolysin (Fig. 7F) did not reduce biofilm formation or SC or E reconstruction, and affected the D as described above. The use of a higher concentration of 32 µg mL−1Ag-D1 (Fig. 7I) or its complex with endolysin (Fig. 7L) did not prevent damage to the epidermis and dermis. Similarly, 8 µg mL−1Im-D2 (Fig. 7D) and Im-D2 + 2 endolysin (Fig. 7H) did not restore the integrity of SC or E, while 32 µg mL−1Im-D2 (Fig. 7J) and Im-D2 + 2 endolysin (Fig. 7M) caused their complete detachment, leaving SC intact. The introduction of chelated dendrimer Ag-D2 at 8 µg mL−1 (Fig. 7E) and 8 µg mL−1 + 2 endolysin (Fig. 7G) also did not prevent SC detachment or E loss, similar to 32 µg mL−1Ag-D2 (Fig. 7K) and Ag-D2 + 2 endolysin (Fig. 7N). In each of these cases, the biofilm adhered to the completely detached stratum corneum, no deeper layers of the epidermis were observed, and the dermis remained degraded, manifesting itself as a flattened surface. This means that none of the tested combinations prevented the mechanical destruction of the skin layers by the PAO1 biofilm (Fig. 8).
Pseudomonas aeruginosa forms clusters of cells that facilitate its attachment to the wound and secretes various enzymes and toxins, such as type A exotoxins, the endotoxin LPS, and elastase, which damage host tissues and immune system cells, leading to necrosis and intensifying the inflammatory process.52 Additional tests must be performed to check whether, after such an advanced bacterial infection, tissue regeneration would be possible in connection with the preservation of the structure of the dermis, despite the penetration of bacteria deep into the remaining tissue.
3.4. Cytotoxic effects on eukaryotic cells
Dendrimers used as wound dressings in skin infections exhibit antibacterial properties; however, their impact on the viability and function of skin cells cannot be overlooked. Therefore, an experiment was conducted to assess the viability and proliferative capacity of VH10 cells in the presence of increasing concentrations of dendrimers. As shown in Fig. 9A, low concentrations of the tested metallodendrimer dendrimer (Ag-D1) – (Fig. 9A) (1–4 µg mL−1) did not significantly affect cell viability, which remained at an average of ∼96%. A slightly higher concentration (8 µg mL−1) led to a marked decrease in survival (∼50%), and the highest concentrations (16, 32, 64, and 128 µg mL−1) further reduced viability to approximately 28%. Based on the results, concentrations that exhibited low or no cytotoxicity were chosen, and endolysin was added at doses of 1 and 2 µg mL−1. As shown in graphs, endolysin alone, at both tested concentrations, did not exhibit cytotoxic effects. Furthermore, the addition of endolysin to Ag-D1 at 4 and 8 µg mL−1 did not induce cytotoxicity (cell viability ∼91%). However, at 16 µg mL−1, survival decreased to ∼52% and ∼42%, respectively, and at 32 and 64 µg mL−1, viability dropped to ∼30% in the presence of endolysin, indicating that high dendrimer concentrations remain cytotoxic despite the addition of the protein.Imidazolium dendrimer Im-D2 (Fig. 9B) exhibited lower cytotoxicity than Ag-D1; at concentrations of 8 and 16 µg mL−1, no significant reduction in cell survival was observed, with viability maintained at around 95%. A moderate decrease to ∼50% was observed at 32 µg mL−1, while the highest concentrations (64 and 128 µg mL−1) reduced survival to ∼30%. In contrast, the addition of endolysin to Im-D2 at lower concentrations (4–8 µg mL−1) did not result in cytotoxicity, with average cell viability exceeding ∼100%. At an Im-D2 concentration of 16 and 32 µg mL−1 combined with a low dose of endolysin, the cell survival rate was approximately 60%. The use of endolysin with dendirmer concentrations of 64 µg mL−1 and 128 µg mL−1 resulted in a fibroblast survival rate of ∼30%. The metallodendrimer Ag-D2 (Fig. 9C) showed an intermediate cytotoxic profile.
Concentrations ranging from 1 to 8 µg mL−1 maintained the cell viability at ∼102%, while 16 and 32 µg mL−1 resulted in a survival rate of ∼77%. At dendrimer concentrations of 64 and 128 µg mL−1, the cell viability decreased to ∼30%. In contrast, Ag-D2 at 16 µg mL−1 in combination with endolysin showed no significant effect on cell viability. Moreover, the addition of 1 or 2 µg mL−1 of endolysin to 32 and 64 µg mL−1 of dendrimer did not reduce cytotoxicity, resulting in a comparable cell survival of ∼65% and ∼31%, respectively.
4. Conclusions
In this study, we have demonstrated that carbosilane CBS dendrimers, imidazolium (Im-D2) and carbosilane metallodendrimers-based silver nanocomposites (Ag-D1 and Ag-D2), when combined with a catalytic CHAP-domain endolysin, act as a modern combat tool against fully formed and highly resistant wild-type Pseudomonas aeruginosa PAO1 bacterial biofilm in in vitro and ex vivo imitations of animal skin infection. On comparing the effectiveness of dendrimer therapy combined with endolysin against P. aeruginosa biofilm in an in vitro model on a smooth plastic surface and ex vivo on a rough, irregular pork skin, a more pronounced reduction in bacterial viability and cellular damage was observed in vitro. This is due to both the higher toxicity of the tested compounds in a simple laboratory environment and the fact that the irregular texture of the skin and richer nutrient resources in the ex vivo model favored the survival of microorganisms. The absence of significant skin damage after the application of dendrimers and their complexes with endolysin indicates that early implementation of therapy could effectively eliminate bacteria without compromising tissue integrity. At the same time, histomorphological analysis revealed that even a 24-hour infection leads to excessive damage to the epidermis, and the bacteria became highly resistant to dendrimers, making it difficult to assess the effectiveness of late interventions. It is worth noting that a significant amount of time often passes between the onset of infection and the appearance of clear pathological changes, diagnosis, and obtaining medical assistance (including the implementation of targeted antibiotic therapy). For example, in an analysis of severe soft tissue infections, the average time from symptom onset to hospital admission was approximately 4.5 days. During this period, the infection may already reach an advanced stage, accompanied by tissue damage and biofilm maturation, which hinders effective pathogen elimination. Therefore, the 24-hour infection variant used in our study has significant translational and therapeutic value; it reflects the stage at which intervention is undertaken after a certain delay, and the effectiveness of treatment may be limited by developed biofilm and tissue changes.Author contributions
Karolina Lach: writing – original draft, methodology, investigation, formal analysis, data curation. Samuel Takvor-Mena: methodology, investigation, writing – review & editing. Oscar Barrios-Gumiel: methodology, investigation. Javier Sanchez-Nieves: writing – review & editing, supervision, funding acquisition, conceptualization. Jacek Kuchinka: methodology, investigation. Piotr Furmańczyk: methodology, investigation. Małgorzata Łysek-Gładysińska: methodology, investigation. Karol Ciepluch: writing – review & editing, supervision, project administration, methodology, funding acquisition, conceptualization.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The data will be made available from the corresponding author upon reasonable request.Acknowledgements
This study was supported by a Polish National Science Centre Research Grant No. 2021/43/D/NZ6/00560. This work was supported by grants from UAH (2024/00300/001-GP2024-02)PID2020112924RB-I00 (MINECO, Spain), EPUINV2020014 (UAH, CAM), and PIUAH21/CC-040 (UAH). This research was also supported by CIBER (Bioengineering, Biomaterials and Nanomedicine CIBER-BBN, CB06/01/1021), Instituto de Salud Carlos III, MICIU European Regional Development Fund. CIBER-BBN is an initiative funded by the VI National R-D-I Plan 2008-2011, Iniciativa Ingenio 2010, Consolider Program, CIBER Actions, and financed by the Instituto de Salud Carlos III (Spain) with assistance from the European Regional Development Fund.References
- B. Van Nieuwenhuyse, D. Van der Linden, O. Chatzis, C. Lood, J. Wagemans, R. Lavigne, K. Schroven, J. Paeshuyse, C. de Magnée, E. Sokal, X. Stéphenne, I. Scheers, H. Rodriguez-Villalobos, S. Djebara, M. Merabishvili, P. Soentjens and J. P. Pirnay, Bacteriophage-Antibiotic Combination Therapy against Extensively DrugResistant Pseudomonas Aeruginosa Infection to Allow Liver Transplantation in a Toddler, Nat. Commun., 2022, 13(1), 1–12, DOI:10.1038/s41467-022-33294-w.
- R. G. Maset, L. Pasquina-Lemonche, A. Hapeshi, L. A. Clifton, J. K. Hobbs, F. Harrison, S. Perrier and S. C. L. Hall, Assessing the Mechanism of Action of Synthetic Nanoengineered Antimicrobial Polymers against the Bacterial Membrane of Pseudomonas Aeruginosa, Biomacromolecules, 2025, 26, 6854–6868, DOI:10.1021/acs.biomac.5c01175.
- R. Urban-Chmiel, A. Marek, D. Stępień-Pyśniak, K. Wieczorek, M. Dec, A. Nowaczek and J. Osek, Antibiotic Resistance in Bacteria—A Review, Antibiotics, 2022, 11(8), 1079, DOI:10.3390/antibiotics11081079/s1.
- J. Laanoja, M. Sihtmäe, H. Vija, I. Kurvet, M. Otsus, K. Šmits, A. Kahru and K. Kasemets, Particle-Driven Synergistic Antibacterial Effect of Silver–Chitosan Nanocomposites Against Escherichia Coli, Pseudomonas Aeruginosa, and Staphylococcus Aureus, ACS Omega, 2025, 10(26), 27904–27919, DOI:10.1021/ACSOMEGA.5C01067.
- A. R. Varadarajan, R. N. Allan, J. D. P. Valentin, O. E. Castañeda Ocampo, V. Somerville, F. Pietsch, M. T. Buhmann, J. West, P. J. Skipp, H. C. van der Mei, Q. Ren, F. Schreiber, J. S. Webb and C. H. Ahrens, An Integrated Model System to Gain Mechanistic Insights into Biofilm-Associated Antimicrobial Resistance in Pseudomonas Aeruginosa MPAO1, npj Biofilms Microbiomes, 2020, 6(1), 1–17, DOI:10.1038/s41522-020-00154-8.
- G. Otis, S. Bhattacharya, O. Malka, S. Kolusheva, P. Bolel, A. Porgador and R. Jelinek, Selective Labeling and Growth Inhibition of Pseudomonas Aeruginosa by Aminoguanidine Carbon Dots, ACS Infect. Dis., 2018, 5(2), 292–302, DOI:10.1021/ACSINFECDIS.8B00270.
- S. Wu, Q. Jiang, D. Lu, X. Zhai, J. Duan and B. Hou, The Effect of Antibacterial Peptide ε-Polylysine against Pseudomonas Aeruginosa Biofilm in Marine Environment, npj Mater. Degrad., 2024, 8(1), 1–11, DOI:10.1038/s41529-02400539-6.
- G. Mancuso, A. Midiri, E. Gerace and C. Biondo, Bacterial Antibiotic Resistance: The Most Critical Pathogens, Pathogens, 2021, 10(10), 1310, DOI:10.3390/PATHOGENS10101310/S1.
- Y. Y. Chen, P. F. Wu, C. S. Chen, I. H. Chen, W. T. Huang and F. D. Wang, Trends in Microbial Profile of Burn Patients Following an Event of Dust Explosion at a Tertiary Medical Center, BMC Infect. Dis., 2020, 20(1), 1–11, DOI:10.1186/s12879-020-4920-4.
- L. L. Burrows, The Therapeutic Pipeline for Pseudomonas Aeruginosa Infections, ACS Infect. Dis., 2018, 4(7), 1041–1047, DOI:10.1021/ACSINFECDIS.8B00112.
- F. Kunisch, C. Campobasso, J. Wagemans, S. Yildirim, B. K. Chan, C. Schaudinn, R. Lavigne, P. E. Turner, M. J. Raschke, A. Trampuz and M. Gonzalez Moreno, Targeting Pseudomonas Aeruginosa Biofilm with an Evolutionary Trained Bacteriophage Cocktail Exploiting Phage Resistance Trade-Offs, Nat. Commun., 2024, 15(1), 1–18, DOI:10.1038/s41467-024-52595-w.
- F. Zhang and W. Cheng, The Mechanism of Bacterial Resistance and Potential Bacteriostatic Strategies, Antibiotics, 2022, 11(9), 1215, DOI:10.3390/ANTIBIOTICS11091215.
- S. Hernando-Amado, M. A. Gomis-Font, J. R. Valverde, A. Oliver and J. L. Martínez, Ceftazidime-Avibactam Use Selects Multidrug-Resistance and Prevents Designing Collateral Sensitivity-Based Therapies against Pseudomonas Aeruginosa, Nat. Commun., 2025, 16(1), 1–12, DOI:10.1038/s41467-02558597-6.
- Y. M. Cai, F. Hong, A. De Craemer, J. G. Malone, A. Crabbé and T. Coenye, Echinacoside Reduces Intracellular C-Di-GMP Levels and Potentiates Tobramycin Activity against Pseudomonas Aeruginosa Biofilm Aggregates, npj Biofilms Microbiomes, 2025, 11(1), 1–13, DOI:10.1038/s41522-025-006732.
- X. Tan, Y. Huang, A. Rana, N. Singh, T. C. Abbey, H. Chen, P. T. Toth and Z. P. Bulman, Optimization of an in Vitro Pseudomonas Aeruginosa Biofilm Model to Examine Antibiotic Pharmacodynamics at the Air-Liquid Interface, npj Biofilms Microbiomes, 2024, 10(1), 1–12, DOI:10.1038/s41522-024-00483-y.
- S. Wang, L. Cai, A. N. Edwards, S. J. Melton, S. T. Retterer, M. J. Doktycz and J. L. Morrell-Falvey, Pseudomonas Aeruginosa Adhesion and Biofilm Formation on Poly(lLysine)-Tethered Hydrogels: Synergistic Effect of Substrate Stiffness and Positive Charge Density, Langmuir, 2025, 41(39), 26628–26638, DOI:10.1021/acs.langmuir.5c02709.
- M. Mu, S. Liu, W. DeFlorio, L. Hao, X. Wang, K. S. Salazar, M. Taylor, A. Castillo, L. Cisneros-Zevallos, J. K. Oh, Y. Min and M. Akbulut, Influence of Surface Roughness, Nanostructure, and Wetting on Bacterial Adhesion, Langmuir, 2023, 39(15), 5426–5439, DOI:10.1021/acs.langmuir.3c00091/asset/images/large/la3c00091_0006.jpeg.
- R. W. Huigens, J. J. Richards, G. Parise, T. E. Ballard, W. Zeng, R. Deora and C. Melander, Inhibition of Pseudomonas Aeruginosa Biofilm Formation with Bromoageliferin Analogues, J. Am. Chem. Soc., 2007, 129(22), 6966–6967, DOI:10.1021/JA069017T.
- R. Rosenzweig, K. Perinbam, V. K. Ly, S. Ahrar, A. Siryaporn and A. F. Yee, Nanopillared Surfaces Disrupt Pseudomonas Aeruginosa Mechanoresponsive Upstream Motility, ACS Appl. Mater. Interfaces, 2019, 11(11), 10532–10539, DOI:10.1021/ACSAMI.8B22262.
- A. Zhao, J. Sun and Y. Liu, Understanding Bacterial Biofilms: From Definition to Treatment Strategies, Front. Cell. Infect. Microbiol., 2023, 13, 1137947, DOI:10.3389/FCIMB.2023.1137947/FULL.
- S. Alfei and D. Caviglia, Prevention and Eradication of Biofilm by Dendrimers: A Possibility Still Little Explored, Pharmaceutics, 2022, 14(10), 2016, DOI:10.3390/PHARMACEUTICS14102016.
- P. Shree, C. K. Singh, K. K. Sodhi, J. N. Surya and D. K. Singh, Biofilms: Understanding the Structure and Contribution towards Bacterial Resistance in Antibiotics, Med. Microecol., 2023, 16, 100084, DOI:10.1016/j.medmic.2023.100084.
- P. P. Mahamuni-Badiger, P. M. Patil, M. V. Badiger, P. R. Patel, B. S. Thorat-Gadgil, A. Pandit and R. A. Bohara, Biofilm Formation to Inhibition: Role of Zinc Oxide-Based Nanoparticles, Mater. Sci. Eng., C, 2020, 108, DOI:10.1016/j.msec.2019.110319.
- Y. Zhang, Y. Cai, B. Zhang and Y. H. P. J. Zhang, Spatially Structured Exchange of Metabolites Enhances Bacterial Survival and Resilience in Biofilms, Nat. Commun., 2024, 15(1), 1–15, DOI:10.1038/s41467-02451940-3.
- K. R. Rouillard, M. R. Markovetz, L. G. Bacudio, D. B. Hill and M. H. Schoenfisch, Pseudomonas Aeruginosa Biofilm Eradication via Nitric Oxide-Releasing Cyclodextrins, ACS Infect. Dis., 2020, 6(7), 1940–1950, DOI:10.1021/ACSINFECDIS.0C00246.
- E. Zahorska, L. M. Denig, S. Lienenklaus, S. Kuhaudomlarp, T. Tschernig, P. Lipp, A. Munder, E. Gillon, S. Minervini, V. Verkhova, A. Imberty, S. Wagner and A. Titz, HighAffinity Lectin Ligands Enable the Detection of Pathogenic Pseudomonas Aeruginosa Biofilms: Implications for Diagnostics and Therapy, JACS Au, 2024, 4(12), 4715–4728, DOI:10.1021/JACSAU.4C00670.
- S. Ellairaja, N. Krithiga, S. Ponmariappan and V. S. Vasantha, Novel Pyrimidine Tagged Silver Nanoparticle Based Fluorescent Immunoassay for the Detection of Pseudomonas Aeruginosa, J. Agric. Food Chem., 2017, 65(8), 1802–1812, DOI:10.1021/acs.jafc.6b04790.
- R. Sean Norman, J. W. Stone, A. Gole, C. J. Murphy and T. L. Sabo-Attwood, Targeted Photothermal Lysis of the Pathogenic Bacteria, Pseudomonas Aeruginosa, with Gold Nanorods, Nano Lett., 2007, 8(1), 302–306, DOI:10.1021/NL0727056.
- Y. You, X. Yu, J. Jiang, Z. Chen, Y. X. Zhu, Y. Chen, H. Lin and J. Shi, Bacterial Cell Wall-Specific Nanomedicine for the Elimination of Staphylococcus Aureus and Pseudomonas Aeruginosa through Electron-Mechanical Intervention, Nat. Commun., 2025, 16(1), 1–15, DOI:10.1038/s41467-02558061-5.
- S. J. Strydom, W. E. Rose, D. P. Otto, W. Liebenberg and M. M. De Villiers, Poly(Amidoamine) Dendrimer-Mediated Synthesis and Stabilization of Silver Sulfonamide Nanoparticles with Increased Antibacterial Activity, Nanomedicine, 2013, 9(1), 85–93, DOI:10.1016/j.nano.2012.03.006.
- N. Gómez-Casanova, Á. Martín-Serrano Ortiz, I. Heredero-Bermejo, J. Sánchez-Nieves, J. Luis Copa-Patiño and F. Javier de la Mata, Potential Anti-Adhesion Activity of Novel Carbosilane Zwitterionic Dendrimers against Eukaryotic and Prokaryotic Pathogenic Microorganisms, Eur. J. Pharm. Biopharm., 2023, 191, 158–165, DOI:10.1016/j.ejpb.2023.07.021.
- V. Patrulea, B. H. Gan, K. Perron, X. Cai, P. Abdel-Sayed, E. Sublet, V. Ducret, N. P. Nerhot, L. A. Applegate, G. Borchard, J. L. Reymond and O. Jordan, Synergistic Effects of Antimicrobial Peptide Dendrimer-Chitosan Polymer Conjugates against Pseudomonas Aeruginosa, Carbohydr. Polym., 2022, 280, 119025, DOI:10.1016/j.carbpol.2021.119025.
- P. Padnya, O. Mostovaya, D. Ovchinnikov, I. Shiabiev, D. Pysin, A. Akhmedov, T. Mukhametzyanov, A. Lyubina, A. Voloshina, K. Petrov and I. Stoikov, Combined Antimicrobial Agents Based on Self-Assembled PAMAM-Calix-Dendrimers/Lysozyme Nanoparticles: Design, Antibacterial Properties and Cytotoxicity, J. Mol. Liq., 2023, 389, 122838, DOI:10.1016/j.molliq.2023.122838.
- X. Shang and D. C. Nelson, Contributions of Net Charge on the Plyc Endolysin Chap Domain, Antibiotics, 2019, 8(2), 1–11, DOI:10.3390/antibiotics8020070.
- M. Sanz-Gaitero, R. Keary, C. Garcia-Doval, A. Coffey and M. J. Van Raaij, Crystal Structure of the Lytic CHAPKdomain of the Endolysin LysK from Staphylococcus Aureus Bacteriophage K, Virol. J., 2014, 11, 133, DOI:10.1186/1743-422X-11133.
- P. K. Thorén Edvardsen, A. N. Englund, Å. Kjendseth Rohr, S. Mesnage and G. VaajeKolstad, Pseudomonas Aeruginosa Cryptic Prophage Endolysin Is a Highly Active Muramidase, Biochemistry, 2025, 10, 53, DOI:10.1021/acs.biochem.5c00142.
- K. Ciepluch, B. Maciejewska, K. Gałczyńska, D. Kuc-Ciepluch, M. Bryszewska, D. Appelhans, Z. Drulis-Kawa and M. Arabski, The Influence of Cationic Dendrimers on Antibacterial Activity of Phage Endolysin against P. Aeruginosa Cells, Bioorg. Chem., 2019, 91, 103121, DOI:10.1016/j.bioorg.2019.103121.
- K. Skrzyniarz, S. Takvor-mena, K. Lach, M. Łysek-, Ó. Barrios-gumiel, J. Cano, K. Ciepluch, K. Skrzyniarz, S. Takvor-mena, K. Lach and M. Łysek, Molecular Mechanism of Action of Imidazolium Carbosilane Dendrimers on the Outer Bacterial Membrane – from Membrane Damage to Permeability to Antimicrobial Endolysin, J. Colloid Interface Sci., 2024, 665, 814–824, DOI:10.1016/j.jcis.2024.03.130.
- K. Lach, K. Skrzyniarz, S. Takvor-Mena, M. Łysek-Gładysińska, P. Furmańczyk, O. Barrios-Gumiel, J. Sanchez-Nieves and K. Ciepluch, Endolysin CHAP DomainCarbosilane Metallodendrimer Complexes with Triple Action on Gram-negative Bacteria: Membrane Destabilization, Reactive Oxygen Species Production and Peptidoglycan Degradation, Int. J. Biol. Macromol., 2024, 278, 134634, DOI:10.1016/j.ijbiomac.2024.134634.
- M. Lasak, M. Łysek-Gładysińska, K. Lach, V. P. Nirwan, D. Kuc-Ciepluch, J. SanchezNieves, F. J. de la Mata, A. Fahmi and K. Ciepluch, Electrospun Nanofibers for the Delivery of Endolysin/Dendronized Ag-NPs Complex Against Pseudomonas Aeruginosa, Nanotechnol., Sci. Appl., 2025, 18, 57–70, DOI:10.2147/nsa.s498942.
- I. Bargathulla, B. Aadhil Ashwaq, S. Sathiyaraj, A. Sultan Nasar and V. Elangovan, Pegylated Bis-Indolyl Polyurethane Dendrimer: Empty Drug Carrier with Prominent Anticancer Activity, Eur. Polym. J., 2021, 153, 110491, DOI:10.1016/j.eurpolymj.2021.110491.
- P. Mittal, A. Saharan, R. Verma, F. M. A. Altalbawy, M. A. Alfaidi, G. E. S. Batiha, W. Akter, R. K. Gautam, M. S. Uddin and M. S. Rahman, Dendrimers: A New Race of Pharmaceutical Nanocarriers, BioMed Res. Int., 2021, 2021(1), 8844030, DOI:10.1155/2021/8844030.
- A. Sripunya, C. Chittasupho, S. Mangmool, A. Angerhofer and W. Imaram, Gallic AcidEncapsulated PAMAM Dendrimers as an Antioxidant Delivery System for Controlled Release and Reduced Cytotoxicity against ARPE-19 Cells, Bioconjugate Chem., 2024, 35(12), 1959–1969, DOI:10.1021/acs.bioconjchem.4c00475.
- A. Abdul Hussein Jameel, N. Hussan Hayder, A. Rami Abdullah and L. Ahmed Yaaqoob, Cytotoxic Effect of Silver Nanoparticles Prepared by Biosurfactant Produced from Pathogenic Bacteria, Indian J. Forensic Med. Toxicol., 2021, 15(4), 63–70, DOI:10.37506/ijfmt.v15i4.16663.
- T. Rodríguez-Prieto, P. F. Popp, J. L. Copa-Patiño, F. Javier de la Mata, J. Cano, T. Mascher and R. Gómez, Silver (I) N-Heterocyclic Carbenes Carbosilane Dendritic Systems and Their Imidazolium- Terminated Analogues as Antibacterial Agents: Study of Their Mode of Action, Pharmaceutics, 2020, 12(10), 1–28, DOI:10.3390/pharmaceutics12100968.
- D. Dhumal, B. Maron, E. Malach, Z. Lyu, L. Ding, D. Marson, E. Laurini, A. Tintaru, B. Ralahy, S. Giorgio, S. Pricl, Z. Hayouka and L. Peng, Dynamic Self-Assembling Supramolecular Dendrimer Nanosystems as Potent Antibacterial Candidates against Drug-Resistant Bacteria and Biofilms, Nanoscale, 2022, 14(26), 9286–9296, 10.1039/d2nr02305a.
- B. Hosnedlova, D. Kabanov, M. Kepinska, V. H. B. Narayanan, A. A. Parikesi, C. Fernandez, G. Bjørklund, H. V. Nguyen, A. Farid, J. Sochor, A. Pholosi, M. Baron, M. Jakubek and R. Kizek, Effect of Biosynthesized Silver Nanoparticles on Bacterial Biofilm Changes in S. Aureus and E. Coli, Nanomaterials, 2022, 12(13), 2183, DOI:10.3390/NANO12132183.
- X. Fu, U. Rehman, L. Wei, Z. S. Chen, M. A. S. Abourehab, P. Kesharwani and Z. H. Cheng, Silver-Dendrimer Nanocomposite as Emerging Therapeutics in Anti-Bacteria and Beyond, Drug Resist. Updates, 2023, 68, 100935, DOI:10.1016/j.drup.2023.100935.
- S. L. Percival, C. Emanuel, K. F. Cutting and D. W. Williams, Microbiology of the Skin and the Role of Biofilms in Infection, Int. Wound J., 2011, 9(1), 14, DOI:10.1111/J.1742-481x.2011.00836.x.
- E. A. Gantwerker and D. B. Hom, Skin: Histology and Physiology of Wound Healing, Clin. Plast. Surg., 2012, 39(1), 85–97, DOI:10.1016/j.cps.2011.09.005.
- K. Chiller, B. A. Selkin and G. J. Murakawa, Skin Microflora and Bacterial Infections of the Skin, J. Invest. Dermatol. Symp. Proc., 2001, 6(3), 170–174, DOI:10.1046/J.0022-202x.2001.00043.x.
- J. Hou, Q. Wu, R. Xiong, P. K. Malakar, Y. Zhu, Y. Zhao and Z. Zhang, A Standardized Mouse Model for Wound Infection with Pseudomonas Aeruginosa, Int. J. Mol. Sci., 2024, 25(21), 11773, DOI:10.3390/IJMS252111773.
| This journal is © The Royal Society of Chemistry 2026 |