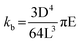Open Access Article
Open Access ArticleStiffness-driven modulation of bactericidal behavior in nanostructured polymer thin films
Ruwen
Tan†
,
Emma
Thomas†
,
Nicolas
Marzolini
and
Yeongseon
Jang
 *
*
Department of Chemical Engineering, University of Florida, Gainesville, FL 32611, USA. E-mail: y.jang@ufl.edu
First published on 17th April 2026
Abstract
The increasing use of medical implants, coupled with rising antimicrobial resistance, has intensified the need for effective antibacterial surface technologies. Protrusive nanostructured surfaces can mechanically disrupt bacterial membranes, leading to cell death, and have inspired extensive biomimetic design strategies. While prior studies have focused on the roles of nanostructure geometry, density, and surface chemistry, the intrinsic stiffness of nanostructured surfaces retaining the same structures and chemistry remains not fully understood. Here, we systematically investigate the relationship between nanostructure stiffness and bactericidal efficacy by fabricating nanostructured polymer surfaces with tunable Young's moduli. Using Escherichia coli and Staphylococcus aureus as model Gram-negative and Gram-positive bacteria, respectively, we demonstrate that bactericidal efficacy against E. coli decreases with reduced nanostructure stiffness, consistent with diminished membrane tension. In contrast, S. aureus exhibits lower sensitivity to stiffness changes at higher moduli, reflecting its thicker and mechanically robust cell envelope. Notably, the softest nanopillars yield the highest bactericidal efficacy against S. aureus, attributed to enhanced nanopillar–cell interactions arising from bacterial cell geometry and increased structural deformability. Furthermore, highly deformable nanostructures promote additional bactericidal effects through lateral squeezing and cell sinking mechanisms. These findings reveal that bacterial cell size and morphology, in conjunction with nanostructure stiffness, critically govern bactericidal performance. This work provides mechanistic insight into stiffness-mediated bacterial membrane disruption and offers design principles for optimizing next-generation antibacterial surfaces.
Introduction
The development of antibacterial surfaces is increasingly important for mitigating microbial contamination in medical and industrial settings.1–3 Traditional approaches rely on bacterial-repelling antifouling coatings4 or bacterial-killing strategies using biocidal agents.5,6 However, these strategies are often limited by reduced long-term efficacy, potential toxicity, and the emergence of antimicrobial resistance.7–11 As an alternative, nanostructured surfaces that physically disrupt bacterial membranes have emerged as a promising non-chemical strategy for bacterial inactivation. Bioinspired nanostructures, such as those found on cicada and dragonfly wings, have demonstrated the ability to mechanically disrupt bacterial membranes.12,13 These studies established that bactericidal activity arises primarily from physical interactions rather than surface chemistry, motivating the development of artificial nanostructured, bactericidal surfaces with controlled geometry.14–16A wide range of nanostructured materials, including silica,17 titanium,18,19 stainless steel,20 aluminum,21 copper,21 gold,22 and polymer,23 have showed bactericidal efficacy, reinforcing the hypothesis that the mechanical interactions between surface nanostructures and adhered bacteria dominate over surface chemistry effects. Previous studies have shown that nanopillar geometry, including height, diameter, and spacing (density), strongly influences bacterial performance.1,10,17,24–27 Reported bactericidal nanopillars typically fall within ranges of 100–900 nm in height, 20–207 nm in diameter, and 9–380 nm in interpillar spacing.25
Bacterial properties, including cell size, membrane rigidity, and surface-associated biomolecules, also influence survival on nanostructured surfaces.24,28 For instance, Gram-negative bacteria such as P. aeruginosa are readily damaged by bioinspired nanopillars, whereas Gram-positive bacteria with thicker and mechanically robust cell envelopes (e.g., B. subtilis, P. maritimus, and S. aureus) often remain intact under similar conditions.13,29 These observations indicate that bacterial membrane rigidity is a key determinant of resistance to mechanical disruption. Given that bactericidal activity arises from physical interactions between the cell and nanostructure, we hypothesize that nanostructure stiffness, analogous to bacterial shape and rigidity, also plays a critical role in governing bactericidal efficacy.
While geometric parameters have been extensively studied for bactericidal performance across various bacterial types, the role of nanostructure stiffness and its coupling with bacterial cell mechanics remains insufficiently understood. In most prior work, nanostructures are fabricated from rigid inorganic materials (e.g., metals,18,20–22 silicon (Si),14,26,30 and silicon oxide17), where deformation during bacterial adhesion is minimal.31,32 In these systems, stiffness has typically been varied by altering nanopillar geometry,17,26,27 making it difficult to decouple mechanical effects from structural parameters and leading to inconsistent observations across studies.33–35 Furthermore, while rigid inorganic nanostructures often exhibit strong bactericidal efficacy, they offer limited tunability and are not well suited for applications requiring flexibility, biocompatibility, or scalable processing. In contrast, in more compliant systems, nanopillars can undergo bending or deformation upon bacterial adhesion,26,36,37 potentially altering cell-surface interactions and bactericidal outcomes.
Poly(ethylene glycol) dimethacrylate (PEGDMA) is a promising material platform due to its antifouling characteristics, biocompatibility, and ability to form well-defined nanostructures.36,38–40 Prior studies have explored PEGDMA-based systems in related contexts: Kim et al. demonstrated that PEGDMA nanopillars possess sufficient stiffness to deform bacteria upon attachment but did not investigate stiffness-dependent bactericidal efficacy.38 Kolewe et al. reported that PEGDMA stiffness influences bacterial adhesion on flat surfaces without incorporating nanostructures.41 Together, these studies highlight the need to systematically investigate stiffness effects in nanostructured polymer systems.
To address this gap, we investigate the effect of nanostructure stiffness on bactericidal efficacy while maintaining consistent surface topography. Specifically, we tune the effective stiffness of nanopillars by varying the Young's modulus of PEGDMA networks with different crosslinking densities. Although this approach enables systematic control of stiffness, we note that changes in polymer modulus may also influence hydration and viscoelasticity, which could contribute to bacterial–surface interactions.42 While these coupled effects cannot be fully decoupled, the use of colloidal lithography and soft molding to maintain consistent nanostructure geometry provides a controlled framework to examine its influence on bactericidal behavior.43 When compared to other nanopatterning methods, such as photolithography,44 electron beam lithography,45 focused ion beam lithography,46 soft lithography,47 and nanoimprinting,48 colloidal lithography offers low costs, high throughput, and accessibility.49
Herein, using nanostructured PEGDMA thin films with controlled nanoscale geometry, we evaluate bactericidal performance against a representative Gram-negative bacterium, E. coli K12, and a Gram-positive bacterium, methicillin-resistant Staphylococcus aureus (MRSA). We further examine the role of externally applied forces by introducing surface-tension-driven loading during drying. This study provides insights into how nanostructure stiffness and bacterial cell properties collectively influence bactericidal behavior and offers design considerations for mechanically tunable antibacterial surfaces.
Experimental section
Materials
Silicon (Si) wafers (4-inch, P-type, <100>), purchased from University Wafer Inc., were used as a substrate to make a master mold. Silica (SiO2) particles with diameters of ∼350 nm (Particle Solutions LLC), serving as masks in colloidal lithography, were prepared by a modified Stöber method.50 Ethanol (Reagent Alcohol) used to clean particles was purchased from Fisher Chemical. Perfluoropolyether (PFPE, Fluorolink MD700, Solvay) was used as a soft mold for pattern transfer. Poly(ethylene glycol) dimethacrylate (PEGDMA, <Mn> ∼350, Polysciences Inc., <Mn> ∼550 and 750, Sigma Aldrich) and Darocur 1173 (photoinitiator, Sigma Aldrich) were used to obtain positive patterns from the Si master mold via soft mold pattern transfer upon ultraviolet (UV) irradiation. Lysogeny broth (LB) and tryptic soy broth (TSB), purchased from Fisher BioReagents and BD Bacto, were used to make LB media and TSB media for bacterial culture.Measurement of mechanical properties by micro-indentation
Micro-indentation was performed by following the protocol described in a previous work.51 Briefly, the flat polymer hydrogel films were placed on the stage and indented with a piezo-driven, quasistatic transducer indentation system (Hysitron BioSoft) with a hemispherical indenter tip. The indentation tip, a borosilicate glass hemisphere with a radius of 1 mm, indented the polymer thin films in the air (or in MilliQ water) at a constant piezo displacement rate of 250 nm s−1 in a displacement-controlled operation mode at 5 random points. Force-response and positions were measured simultaneously by the quasistatic transducer at a rate of 125 Hz. The Young's modulus and other mechanical properties were calculated by analyzing the force-displacement curve with MATLAB.Colloid lithography to create nanopillars on silicon (Si) master mold
Monodispersed colloidal SiO2 microspheres with diameters of 350 nm were cleaned by repeating 5 cycles of centrifugation and redispersion in ethanol. A monolayer of close-packed SiO2 particle array was coated on a 4-inch Si wafer by Langmuir–Blodgett coating.52 Briefly, the wafer was vertically pulled up by a syringe pump at a speed of 5 mm min−1 from water, while the SiO2 particles, dispersed in ethylene glycol (Alfa Aesar, 99%) with a volume fraction of 1%, were added on the water/air interface. The close-packed SiO2 arrays served as masks during reactive ion etching (RIE, Unaxis Shuttlelock Reactive Ion Etcher) using an SF6/O2 gas mixture (5 mTorr, 20 SCCM SF6, 20 SCCM O2, 150 W, for 4 min) to fabricate Si nanopillars beneath.53 The SiO2 particles were finally removed by immersing the etched wafer in a 2 vol% hydrofluoric acid aqueous solution for 2 min.Fabrication of nanostructure on polymer thin film
Si master mold with nanopillars was treated with trichloro-(1H,1H,2H,2H-perfluorooctyl) silane (Sigma Aldrich) to lower the surface energy. A soft negative mold with the inverted pattern of the Si master mold was fabricated by drop-casting PFPE mixed with 4 wt% photoinitiator, followed by UV-induced polymerization at a wavelength of 365 nm for 15 min under N2.54 Using the PFPE soft negative mold, the nanopillar pattern was transferred from the Si master mold to the target polymer thin films. A drop of the PEGDMA oligomer solution containing 0.4 wt% photoinitiator was placed between a PFPE soft mold and a glass slide. Crosslinked PEGDMA thin films with nanopillars were easily peeled off from the PFPE soft molds after 15 min of UV exposure.Bacterial cell culture and bacterial live/dead assay
E. coli K12 KCJ140 (or S. aureus MRSA KCJ3K555) was pre-cultured in 5 mL of LB (or TSB) media overnight. 50 μL of the pre-cultured bacterial solution was transferred to 5 mL fresh LB media (or TSB media) for an additional incubation in a shaking incubator until late log (optical density at 600 nm, OD600 = 1.0) phase. 1 mL of the diluted bacterial solution with OD600 = 0.1 (∼107 cells per mL) in phosphate-buffered saline (PBS) was added to each well of a 24-well plate containing samples. Then, the samples were incubated at 37 °C for 1 h and 4 h. At the end of incubation, each sample was gently rinsed twice with 1 mL PBS to remove non-attached bacteria. To evaluate the viability of bacteria adhered in the wet condition, the samples were stained with a live/dead baclight bacterial viability kit (L7012, Life Technologies) according to the manufacturer's protocol. Briefly, 1.5 μL of 3.34 mM SYTO9 (stain live cells with intact membranes with green fluorescent color) and 1.5 μL of 20 mM propidium iodide (stain dead cells with damaged membranes with red fluorescent color) were mixed in 1 mL of PBS. Each sample was immersed in the staining solution individually and incubated in the dark for 15 min. The samples were transferred to a glass slide, covered with a cover slip to prevent water evaporation. Then, the samples were imaged using a fluorescent microscope (Zeiss, AxioObserver7) equipped with a 20× objective lens. To further evaluate the effect of the nanostructure's mechanical properties on the surface-tension-driven bactericidal efficacy, the samples incubated for 4 h were treated with a sequential short-time drying step, following a protocol from previous work.55 Briefly, the nanostructured samples were incubated in bacterial solution for 4 h and washed with PBS as previously described. Then, the samples were sequentially dried in air for 60 s by allowing the water thin film on the surface to evaporate. The samples were then stained and imaged with the same procedure described above. Images were taken at 5 random points on each sample, and the number of bacteria on each image was analyzed by the image-based tool for counting nuclei (ITCN) plugin in ImageJ. The statistical analysis of bacterial adhesion and viability was performed using a One-way ANOVA in GraphPad.Scanning electron microscopy (SEM) for monitoring morphologies of nanostructured surfaces and adhered bacteria
To visualize the morphology of the nanostructure on the surfaces and adhered bacteria by SEM, samples were prepared and incubated in bacterial solution with the same procedure described above. After 4 h incubation, samples were gently washed with 1 mL PBS twice, fixed in 2.5% glutaraldehyde (TCI America) solution for 1 h at 4 °C, and dehydrated using a series of ethanol concentrations in distilled water (i.e., 50, 70, 90, and 100% for 20 min, respectively). The dehydrated samples were further dried with hexamethyldisilazane (HDMS, Alfa Aesar, 98%) overnight. To visualize the morphology of the nanostructure on the surfaces and adhered bacteria after air-drying for 60 s while minimizing the introduction of additional surface tension, the dried samples were fixed in 2.5% glutaraldehyde and freeze-dried in a lyophilizer (Labconco, FreeZone 2.5 Plus) overnight to ensure complete dehydration. The prepared samples were then sputtered with gold (10 nm). Bacterial morphology was characterized using SEM (FEI Nova NanoSEM 430) at an acceleration potential of 3 kV.Statistical analysis
All quantitative data are reported as mean ± standard error of the mean (SEM). For each sample condition, fluorescence images were collected from five randomly selected locations per sample, and bacterial counts were obtained using the ImageJ ITCN plugin as described above. Statistical analysis of bacterial adhesion and viability between different sample groups was performed using one-way analysis of variance (ANOVA) in GraphPad. Statistical significance between groups was evaluated using the following thresholds: ns (not significant), p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.Results and discussion
Tuning Young's moduli of polymer thin films
Poly(ethylene glycol) dimethacrylate (PEGDMA), a biocompatible polymer with moderate tunability of its properties, has been widely used as the major or added component in polymers for many bio-applications, such as bone generation and tissue scaffolding.56–58 Here, we tailored the Young's moduli of network polymer thin films by changing their crosslinking density through polymerization of the PEGDMA oligomers with different average PEG repeating units (n) of 4.5, 9, and 13.5. Fig. 1a shows the free radical polymerization of PEGDMA oligomers into the networks in the presence of photoinitiator and UV exposure. PEGDMA thin films with 4.5 PEG repeating units showed the highest Young's modulus, 353 MPa in air, while the Young's modulus decreased to 75 MPa and 30 MPa in air when n was increased to 9 and 13.5, respectively (Fig. 1b). The Young's modulus was measured from PEGDMA thin films and used as an estimate of the effective material modulus governing nanopillar behavior, recognizing that geometric confinement and nanoscale effects may lead to deviations from bulk values. It should also be noted that variations in crosslinking density may influence polymer hydration and viscoelastic properties, in addition to stiffness.59The polymerization of PEGDMA oligomers with fewer PEG repeating units leads to higher crosslinking densities in the polymer networks, resulting in higher Young's moduli.60 Because the antibacterial performances of polymer samples are evaluated in aqueous solution, we further assessed the Young's moduli of PEGDMA thin films that were swollen in water for 24 hours (Fig. 1c). The fully swollen PEGDMA thin films also showed a reduction in Young's modulus as their crosslinking densities decreased. The absorption of water molecules on the PEG chains reduces the crosslinking concentration and Young's modulus compared with their moduli in air.60 As a result, we tailored the Young's moduli of PEGDMA thin films made from oligomers containing an average of 4.5, 9, and 13.5 PEG repeating units to 190 MPa, 40 MPa, and 17 MPa in water, respectively. Prior studies on PEGDMA hydrogels typically report moduli in the kPa range and modulate stiffness through polymer weight fraction, with a primary focus on bacterial adhesion.41,61,62 In contrast, the present work investigates PEGDMA thin films with substantially higher moduli in the MPa range and evaluates both bactericidal efficacy and adhesion. Notably, while previous studies have largely examined flat hydrogel substrates, our approach integrates stiffness control within nanostructured architectures.
Fabricating nanopillars on PEGDMA thin films
The nanopillar geometry used in this study (average spacing ∼350 nm, height ∼500 nm, tip diameter ∼110 nm, and base diameter ∼230 nm) was selected based on our prior work, optimizing nanopatterned polymer surfaces for maximal bactericidal efficacy.26 While previous studies from our group compared flat and nanostructured surfaces across varying densities, the present work maintains this optimized geometry and focuses specifically on isolating the effect of stiffness. Fig. 2a shows the schematics of fabricating bactericidal nanopillars on the Si and transferring them to PEGDMA thin films. Colloidal lithography using monodispersed SiO2 nanoparticles as etching masks and SF6/O2 as etching gas enables the creation of ordered, uniform nanopillars across the entire Si master, providing an ideal platform to study the effect of nanopillar stiffness while minimizing the influence of surface topography variation.52,53 A soft molding pattern transfer was then used to replicate the nanopillars from the Si master to target PEGDMA thin films. In the procedures, the perfluoropolyether (PFPE), a fluoro-polymer with low shrinkage upon polymerization and low surface energy, allows the precise replication of inverted structures on the negative mold.Eventually, the nanopillars with different Young's moduli were fabricated on the target PEGDMA thin films by casting and curing their oligomers on the negative mold. Fig. 2b shows scanning electron microscopy (SEM) images of the nanopillars fabricated on the Si master and PEGDMA thin films. The nanopillars fabricated on the PEGDMA thin films shared similar geometric parameters but differed in Young's modulus, enabling investigation of nanopillar stiffness and its impact on bactericidal efficacy. The detailed geometric parameters of nanostructured Si and nanostructured PEGDMA thin films are summarized in Table S1 in the SI.
Understanding the relationship between nanopillar stiffness and the bactericidal efficacy against E. coli
A model Gram-negative bacterium, E. coli K12, which is a completely genome sequenced bacterium and widely used for assessing bactericidal efficacy,63,64 was used to evaluate the effect of nanopillar stiffness on antibacterial performance. We performed bacterial live/dead assays (Fig. 3a) by observing the fluorescent signals of adhered bacteria on nanopillars after 4 h incubation in bacterial solution, allowing for sufficient bacterial adhesion.26 In this assay, SYTO 9 can penetrate into the live cells with intact membranes and stain nucleic acids a fluorescent green, whereas propidium iodide only diffuses into cells with damaged membranes and stains the nucleic acid of non-viable cells a fluorescent red.65 The percent of non-viable cells was calculated by dividing the number of non-viable red cells by the total number of cells adhered to the area of interest (Fig. 3b). We evaluated the bactericidal efficacy by the percentage of non-viable cells among the total bacteria adhered. The bactericidal nanopillars fabricated on Si with a Young's modulus of 169 GPa were used as the control due to their sufficient stiffness and well-known bactericidal efficacy against E. coli.14,31 Importantly, these Si nanopillars represent an upper bound in stiffness, enabling benchmarking of polymer-based nanostructures and isolating the role of mechanical compliance on bactericidal performance.After E. coli incubation with the samples for 4 hours, we confirmed that 66% of adhered E. coli cells were dead on hard Si nanopillars, while the percentage of non-viable cells on the PEGDMA nanopillars decreased from 43% to 28% as the surfaces' elastic moduli decreased from 190 MPa to 17 MPa. We also confirmed that the highest total number of adhered bacteria was observed on Si nanopillars, whereas reduced bacterial adhesion was observed on PEGDMA nanopillars, with variation among surfaces (Fig. 3c). The adhesion density of E. coli on PEGDMA nanopillars ranged from 104 to 105 cells per square centimeter across all Young's moduli. Interestingly, the highest adhesion of E. coli was observed on nanopillars of intermediate stiffness (n = 9). Bacterial adhesion is influenced by multiple factors, including surface topography, wettability, stiffness, and charge.66 The enhanced adhesion at intermediate stiffness may reflect a balance between structural compliance and mechanical support, where sufficient deformation increases bacteria–surface contact area while maintaining stable anchoring points. However, this interpretation remains qualitative.
The dependence of bactericidal efficacy on the surface nanostructure stiffness can be modeled by a biophysical framework describing the different cell-nanopillar interactions under the driving force of the interfacial energy gradient. Three interfaces of bacterium–liquid, bacterium–solid, and solid–liquid exist when a bacterium attaches to a nanostructured surface. The total interfacial energy of the system is determined by the summation of individual interface/surface energies (σ or γ) between the bacterium–liquid (ABLσBL), bacterium–solid (ABSσBS), and solid–liquid (ASLγSL) at given contact areas (A).67 The total interfacial energy decreases as the contact area between a bacterial membrane and the nanostructure increases, driving the bacterium to spontaneously move into the nanopillars.68 The interfacial energy gradient theory is applicable to materials with intrinsic contact angles between 30° and 60°,68 which aligns with the contact angles of Si and PEGDMA used in this study (Fig. S1). While this model does not explicitly account for polymer hydration or viscoelastic effects, it provides a useful framework for interpreting cell–nanopillar interactions.
Following the interfacial energy gradient theory, the driving force to induce bacterial adhesion and membrane expansion into the nanostructures and minimize the interfacial energy gradient13 causes the suspended region of the bacterial membrane between initial contact points with the top of nanostructures to experience a certain deformation, leading to a tensile force equal in magnitude and opposite in direction on the nanopillars. Thus, the stress imposed on the bacterial membrane can only be as large as the nanopillars can tolerate. Nanopillars that lack sufficient stiffness will be bent laterally towards the region of higher bacterial membrane stress, reducing the overall stress on the membrane, as they can no longer sustain the higher stress.
To confirm this hypothesis, the morphology of the bacteria and nanopillars was characterized using SEM. SEM imaging reveals morphological signatures of membrane deformation and rupture, which have been widely associated with mechanically induced bactericidal mechanisms in nanostructured surfaces. Qualitative analysis revealed that E. coli maintained an intact ellipsoidal shape on the flat surface (Fig. 4 and S2), whereas many cells were damaged and flattened due to cytoplasm loss on the Si nanopillars, indicating the physical and mechanical bactericidal efficacy of Si nanopillars. Attributed to the high Young's modulus of Si (169 GPa),31 the Si nanopillars had sufficient stiffness and did not deform underneath the damaged bacterial membrane. However, less drastic deformations of E. coli were observed on the PEGDMA nanopillars, indicating a lower possibility for cell damage and loss of cellular volume compared with cells on Si nanopillars. These observations corroborate the previous results from the bacterial live/dead assay. Moreover, the PEGDMA nanopillars adhered to the bacterial membrane showed bending towards the center of the bacteria, suggesting that the pressure applied to the bacterial membrane induces bending of nanopillars that lack sufficient stiffness along the direction of higher bacterial membrane stress.
 | ||
| Fig. 4 Top-view SEM images of E. coli K12 incubated in a static incubator at 37 °C for 4 h on nanopillars of a) Si, PEGDMA with b) 4.5, c) 9, and d) 13.5 PEG repeating units. All scale bars are 2 μm. | ||
To qualitatively evaluate the effect of nanopillar deformation on the bactericidal efficacy, Euler–Bernoulli beam theory was applied. In this theory, the resistance against bending, the bending modulus kb, is proportional to the Young's modulus of the material.69
The nanopillars were approximated as cylindrical beams using the tip diameter of 110 nm, recognizing that the tapered geometry may lead to a higher effective stiffness due to increased diameter toward the base. Nanopillars with a higher bending modulus showed higher resistance against deformation when similar forces were applied by the stress on the bacterial membrane (Fig. S3), suggesting they can maintain their structure while contending with the stress on the bacterial membrane, while the nanopillars with lower bending moduli will be deformed upon application of this stress. Deformation of the nanopillars may reduce the localized stress exerted on the bacterial membrane while increasing contact area along the nanopillar sidewalls. This combination likely lowers the probability of membrane rupture and subsequent cell death. Collectively, these observations suggest that decreased nanostructure stiffness diminished bactericidal efficacy against E. coli on nanopillar surfaces.
Understanding the relationship between nanopillar stiffness and the bactericidal efficacy against S. aureus
To further understand the relationship between the nanopillar stiffness and bactericidal efficacy against Gram-positive strains, we evaluated the viability of S. aureus, one of the most common bacteria implicated in hospital infections,70 on our surfaces by bacterial live/dead assays. Fig. 5a shows the microscopic fluorescent images of stained S. aureus incubated on surfaces with nanopillars. We confirmed that 55% of adhered S. aureus were stained with red fluorescence and damaged by Si nanopillars, while the percentage of non-viable cells on PEGDMA nanopillars decreased from 17% to 14% as their Young's moduli decreased from 190 MPa to 40 MPa (Fig. 5b). Surprisingly, the percentage of non-viable S. aureus increases to 41% as the Young's modulus of nanopillars further decreases to 17 MPa. Fewer S. aureus bacteria were found on the PEGDMA nanopillars compared to those on Si, while no significant difference in the number of adhered bacteria was found amongst PEGDMA nanopillars with different stiffnesses (Fig. 5c).Similarly, the force to drive S. aureus into the nanopillars and the accompanying pressure that can be applied to the bacterial membrane remained similar from the Si substrate to those made of PEGDMA, with the assumption that all nanopillars maintained their fabricated shape and structural integrity during the cell–nanopillar interaction.68 However, the bactericidal efficacy observed on the nanopillars presented a different trend from this prediction, indicating that other dominant factors exist to govern the bactericidal efficacy in the tested system. To further reveal potential factors affecting the bactericidal efficacy of the nanopillars, we characterized the morphology of S. aureus on the surfaces after 4 h of incubation by SEM (Fig. 6). SEM images revealed that S. aureus cells on the Si nanopillars were embedded into the nanopillars with parts of their membranes lysed. This result corresponds to the bactericidal efficacy of Si nanopillars obtained from the bacterial live/dead assay, where approximately 55% of the cells were found non-viable.
 | ||
| Fig. 6 Top-view SEM images of S. aureus incubated in a static incubator at 37 °C for 4 h on nanopillars of a) Si, PEGDMA with b) 4.5, c) 9, and d) 13.5 PEG repeating units. All scale bars are 2 μm. | ||
We also observed that the bactericidal efficacy of Si nanopillars against S. aureus was lower than that against E. coli after 4 h of cultivation in solution. This is attributed to S. aureus having a stiffer membrane with a thicker layer of peptidoglycan compared to Gram-negative bacteria.71 The thicker peptidoglycan layer offers a higher elastic modulus on the bacterial membrane, reducing the possibility of damage and rupture by nanopillars. Due to the stiffer membrane and bacterial congregation of S. aureus, the Si nanopillars with higher resistance to bending were better able to maintain force on the bacterial membrane sufficient to damage or induce rupture.
Additionally, the SEM images, as well as the live/dead assay, show denser growth of S. aureus compared to E. coli. Staphylococci are known to form an aggressive biofilm that is difficult to eradicate, likely contributing to the increased growth and decreased bactericidal efficacy of S. aureus across all surfaces.72,73 Additionally, the adhesion mechanisms of S. aureus and E. coli to the PEGDMA nanopillars are governed by their unique cell structures and appendages.74 In S. aureus, most adhesins are covalently linked to the cell wall peptidoglycan. Of these, the microbial surface components recognizing adhesive matrix molecules (MSCRAMMs) are primarily responsible for the strong binding of the bacterium to the surface.75 Meanwhile, adhesion of E. coli to abiotic surfaces is aided by extracellular appendages such as type 1 fimbriae and curli amyloid fibers, contributing to their differences in adhesion dynamics.74
Initial bacterial adhesion is thought to be significantly influenced by physicochemical and electrostatic interactions between the surface and the bacterial envelope.76 Surface hydrophobicity, often assessed via water contact angle, is commonly used as an indirect predictor of these interfacial interactions. While this metric does not fully capture the complexity of surface chemistry or hydration-layer effects, it provides a useful comparative framework for interpreting adhesion trends. In our system, the most hydrophilic sample (Si), as indicated by the lowest water contact angle (Fig. S1), exhibited the highest adhesion density. For E. coli, adhesion density generally decreased as the nanostructured surface became more hydrophobic, with the exception of the 40 MPa sample. This trend is consistent with prior reports that the outer membrane of E. coli is predominantly hydrophilic, although it may vary depending on strain and local surface composition.77 In contrast, S. aureus, which possesses a relatively hydrophobic cell wall,78 showed increased adhesion to more hydrophobic surfaces. This behavior aligns with the model proposed by Maikranz et al. in which S. aureus adheres to hydrophobic surfaces through multiple weakly bonded macromolecules, compared to fewer but stronger interactions on hydrophilic surfaces.79 Accordingly, the contributions of surface chemistry and protein adsorption to bacterial adhesion are recognized, but are not explicitly resolved within the scope of this study.
The motility of a bacterium also heavily influences adhesion dynamics, as bacteria must be close to the material's surface to overcome the energy barrier and bind to the surface.42 In a previous study with E. coli K-12, engineered non-flagellated or paralyzed cells exhibited significantly weaker adhesion than the wild-type flagellated strain.80 However, MRSA, generally considered non-motile, exhibited adhesion orders of magnitude higher than that of E. coli, indicating that motility is not a dominant factor in this system.
Our results indicate that the bactericidal efficacy of PEGDMA nanopillars against S. aureus decreased with increasing stiffness, whereas against E. coli it increased with nanopillar stiffness. Visualized in Fig. 6b and c, many S. aureus cells were found to be intact and settled on top of the bent nanopillars of PEGDMA with Young's moduli of 190 MPa (n = 4.5) and 40 MPa (n = 9). However, we found an exception when the Young's modulus of the nanopillars decreases further. We observed the highest number of non-viable S. aureus bacteria on nanopillars with the lowest stiffness (n = 13.5, 17 MPa) compared with substrates containing stiffer nanopillars. Moreover, many S. aureus cells were deformed and sank into the nanopillars, indicating a loss of cytoplasm and damage to the cellular integrity (Fig. 6d). It is thought that the softest nanopillars cannot support S. aureus upon adhesion. This leads to the sinking of the spherical S. aureus (1–2 μm in diameter) into the nanopillars, a phenomenon not observed in the larger, rod-shaped E. coli (1–2 μm length, 0.5 μm width). Due to the 350 nm distance between nanopillars and the higher elastic modulus of the peptidoglycan layer, S. aureus is able to exert enough force to bend the softer nanopillars and create enough space to situate in-between nanopillars. Along with sinking, the interactive radius on the substrate increases due to the spherical shape of S. aureus, leading to greater adhesion to the sidewalls of nanopillars around the bacterium's perimeter.
Based on the Euler–Bernoulli beam theory, we know that the bending modulus, kb, is inversely proportional to the third power of the height of the point of action. Thus, adhesion to multiple sidewalls of the nanopillars significantly increases lateral interaction between the bacterial membrane and the nanopillars, providing greater resistance to bending and enhancing bactericidal efficacy (Fig. 5b and 6d). Choi et al. also reported comparable behavior, showing that S. aureus can be captured and killed by lateral stretching when attached to nanopillar sidewalls spaced similarly to the cell size.81 The lateral compression of S. aureus by the nanopillars results in increased bactericidal efficacy in softer, more deformable nanopillars with lower Young's moduli, where the bacterium is able to sink lower into the nanopillars and increase this interaction. The reverse holds for the effect on E. coli: the rod-shaped bacterium, with a less-rigid outer membrane, adheres on top of the nanopillars. Here, stiffer nanopillars can exert sufficient force on E. coli to puncture the outer membrane without bending. To our best knowledge, our results first confirmed that bactericidal efficacy is affected not only by nanopillar stiffness but also by changes in interactions arising from reductions in nanopillar stiffness and cellular size.
Understanding the relationship between nanopillar stiffness and the surface-tension-driven bactericidal efficacy against Gram-negative and Gram-positive bacteria
As the E. coli and S. aureus are incubated in solution, the interfacial energy gradient plays an important role in driving the bacteria into the nanopillars, leading to bacterial membrane damage and subsequent death. However, as observed in the evaluation of bactericidal efficacy in solution, only up to 65% of E. coli and 55% of S. aureus cells adhering to the nanopillars were inactivated. To further enhance the bactericidal efficacy on the nanopillars, an external force towards the substrate was introduced to the system by the surface tension of the air-liquid interface along the bacterial membrane during surface water evaporation. This creates an additional driving force to pull the bacteria towards the nanopillars, leading to cell rupture on the nanopillars by imposing pressure on the bacterial membrane until it yields to its elastic modulus.55Considering that nanopillar stiffness can also affect bactericidal efficacy under external force, we further evaluated the enhancement of bactericidal efficacy against E. coli and S. aureus on nanopillars with different stiffness upon introduction of an external force. Fig. 7a shows the representative fluorescent images of E. coli and S. aureus upon an external force induced by surface tension during a short exposure of surfaces to air. Based on quantitative image analysis, we found that bactericidal efficacy against E. coli and S. aureus was significantly enhanced on Si nanopillars, reaching 99% and 97%, respectively (Fig. 7b and c). This enhancement can be understood by imposing an additional driving force arising from surface tension to increase the stress on the bacterial membrane, thereby increasing the likelihood of cell rupture and damage.55
To better understand the effect of the surface-tension-induced external force on the bactericidal efficacy against different strains, we calculated the external force and pressure applied to the bacterial membrane as a function of the relative liquid level during water evaporation (Table S2 and Fig. S4). In the calculation, the force is calculated by integrating surface tension along the periphery of the cells, and the pressure is calculated by dividing the force by the projected area in contact with a cylindrical post. The force and pressure reach a maximum when the liquid level is near the half-height of the cells due to surface tension interacting with more bacterial membranes along the cell's perimeter than at other heights. We also noticed that a lower external force was generated with the smaller S. aureus compared to the larger E. coli. However, due to the higher ratio between perimeter and area of S. aureus compared to E. coli, a higher force is applied on the unit area of S. aureus, causing higher pressure applied on the membrane of S. aureus. Thus, similar bactericidal efficacy against S. aureus and E. coli was observed on Si nanopillars, despite the fact that S. aureus has a stiffer bacterial membrane. Moreover, we found that trials with S. aureus exhibited lower bactericidal efficacy on the corresponding soft nanopillars than those with E. coli. Analogous to bactericidal efficacy in solution, surface-tension-driven bactericidal efficacies are reduced as the nanopillar stiffness decreases. We hypothesized that nanopillars with lower stiffness were further deformed upon the introduction of the external force, leading to an inclination of the nanopillars and a change in the cell–nanopillar interaction. As a result, the percentage of non-viable E. coli and S. aureus cells was reduced on the soft nanopillars.
To confirm this hypothesis, we characterized the morphology of E. coli and S. aureus on nanopillars by SEM. E. coli and S. aureus are intact on a flat surface (Fig. S5) but were lysed on Si nanopillars (Fig. 8). These results of drastic cell morphology changes on the Si nanopillars confirm that imposing external force enhances the deformation of adhered bacteria and the bactericidal efficacy. Similarly, bacteria sank into PEGDMA nanopillars upon the addition of the external force, and the surrounding nanopillars inclined directionally towards the bacteria. The stiffer nanopillars, possessing higher resistance to bending, result in larger tension on the bacterial membrane and thus greater bactericidal efficacy. Moreover, the agglomeration of nanopillars was observed on the PEGDMA surface with the lowest stiffness, which is attributed to adhesion-mediated elastocapillary interactions during water evaporation on the surface.82 Due to the agglomeration, the bacterial membrane has a possibility to settle on the side of the nanopillars, which leads to an increase in the contact area and a decrease in the pressure applied to the bacterial membrane. Eventually, the bactericidal efficacy against E. coli and S. aureus decreases on the nanopillars with low stiffness. These observations further confirm the previously mentioned hypothesis.
While this study provides insight into the role of nanopillar stiffness in bactericidal efficacy, several considerations should be noted to contextualize the scope of the work. Tuning the Young's modulus of PEGDMA thin films through crosslinking density may also influence polymer hydration and viscoelastic properties, which could contribute to bacteria–surface interactions alongside stiffness effects. In addition, the proposed bactericidal mechanisms are inferred from qualitative morphological observations and analytical modeling; direct measurements of interaction forces would provide further mechanistic resolution. The present study focuses on short-term behavior to probe primary bacterial adhesion, using two representative bacterial strains and a single nanostructure geometry to enable controlled comparison of stiffness-dependent trends. The selected nanopillar geometry and density were based on our previous work, in which these parameters were optimized to achieve high bactericidal efficacy against E. coli. While surface-tension-driven bactericidal effects demonstrate enhanced efficacy under drying conditions, their influence may vary in fully hydrated environments. Within this context, the results establish a foundation for understanding stiffness-dependent bactericidal behavior and provide guiding principles for the design of mechanically tunable antibacterial surfaces.
Conclusion
In this work, we investigated the effect of nanopillars' stiffness on the bactericidal efficacy against Gram-negative and Gram-positive bacteria. Nanostructured PEGDMA thin films with tunable Young's moduli were fabricated by varying crosslinking density, enabling evaluation of stiffness-dependent trends while maintaining similar geometric features. Using E. coli K12 and methicillin-resistant Staphylococcus aureus (MRSA) as representative models, we observed that bactericidal efficacy against E. coli decreased with decreasing nanopillar stiffness, consistent with reduced mechanical interaction between softer nanopillars and the bacterial membrane. In contrast, nanopillars with both high and low stiffness exhibited strong bactericidal activity against S. aureus, suggesting that bactericidal performance depends not only on nanostructure stiffness but also on bacterial properties such as size and cell envelope structure. Moreover, the introduction of an external force via surface tension was found to enhance bactericidal efficacy, with greater enhancement observed for stiffer nanopillars. These findings suggest that nanopillar stiffness influences bactericidal performance under both passive and externally driven conditions. It should be noted, however, that direct quantification of force and membrane stress will further refine the mechanistic understanding of these interactions. This work provides insight into cell–structure interactions and rational designs for creating synergistic antibacterial nanostructures on materials with different mechanical properties. Longer-term efforts will aim to translate these findings into novel antibacterial surfaces to combat industrial biofouling and medical device infection.Author contributions
The manuscript was written through the contributions of all authors. R. Tan and E. Thomas contributed equally and share first authorship. All authors have given approval to the final version of the manuscript.Conflicts of interest
The authors declare no competing financial interest.Data availability
The data supporting this article (geometric parameters of nanostructures, contact angles, micrographs, force measurements, and symbols and values used in the calculation of the surface-tension-induced force and pressure) have been included as part of the supplementary information (SI).Supplementary information is available. See DOI: https://doi.org/10.1039/d6lf00023a.
Acknowledgements
This research was financially supported by Prof. Jang's startup funds (19030100-211-CRRNT-00128948) provided by the Department of Chemical Engineering and Herbert Wertheim College of Engineering at the University of Florida. The authors acknowledge Prof. Thomas Angelini (UF, Mechanical and Aerospace Engineering) for his assistance in using the micro-indenter for measuring mechanical properties and Prof. Kwangcheol Jeong (UF, Microbiology) for providing the cell strains used in this study.References
- D. P. Linklater, V. A. Baulin, S. Juodkazis, R. J. Crawford, P. Stoodley and E. P. Ivanova, Mechano-bactericidal actions of nanostructured surfaces, Nat. Rev. Microbiol., 2021, 19(1), 8–22 CrossRef CAS PubMed.
- K. Sauer, P. Stoodley, D. M. Goeres, L. Hall-Stoodley, M. Burmølle, P. S. Stewart and T. Bjarnsholt, The biofilm life cycle: expanding the conceptual model of biofilm formation, Nat. Rev. Microbiol., 2022, 20(10), 608–620 CrossRef CAS PubMed.
- Y. Liu and Q. Zhao, Influence of surface energy of modified surfaces on bacterial adhesion, Biophys. Chem., 2005, 117(1), 39–45 CrossRef CAS PubMed.
- Z. Lu, E. A. Q. Mondarte, K. Suthiwanich, T. Hayashi, T. Masuda, N. Isu and M. Takai, Study on Bacterial Antiadhesiveness of Stiffness and Thickness Tunable Cross-Linked Phospholipid Copolymer Thin-Film, ACS Appl. Bio Mater., 2020, 3(2), 1079–1087 CrossRef CAS PubMed.
- X. Yang, J. Hou, Y. Tian, J. Zhao, Q. Sun and S. Zhou, Antibacterial surfaces: Strategies and applications, Sci. China: Technol. Sci., 2022, 65(5), 1000–1010 CrossRef PubMed.
- H. Chouirfa, H. Bouloussa, V. Migonney and C. Falentin-Daudré, Review of titanium surface modification techniques and coatings for antibacterial applications, Acta Biomater., 2019, 83, 37–54 CrossRef CAS PubMed.
- C. L. Ventola, The antibiotic resistance crisis: part 1: causes and threats, Pharm. Ther., 2015, 40(4), 277 Search PubMed.
- T.-F. C. Mah and G. A. O'Toole, Mechanisms of biofilm resistance to antimicrobial agents, Trends Microbiol., 2001, 9(1), 34–39 CrossRef CAS PubMed.
- S. Prabhu and E. K. Poulose, Silver nanoparticles: mechanism of antimicrobial action, synthesis, medical applications, and toxicity effects, Int. Nano Lett., 2012, 2(1), 1–10 Search PubMed.
- R. Tan, J. Yoo and Y. Jang, Engineering Approaches to Create Antibacterial Surfaces on Biomedical Implants and Devices, Racing for the Surface, 2020, pp. 313–340 Search PubMed.
- J. Hasan, R. J. Crawford and E. P. Ivanova, Antibacterial surfaces: the quest for a new generation of biomaterials, Trends Biotechnol., 2013, 31(5), 295–304 CrossRef CAS PubMed.
- E. P. Ivanova, J. Hasan, H. K. Webb, V. K. Truong, G. S. Watson, J. A. Watson, V. A. Baulin, S. Pogodin, J. Y. Wang and M. J. Tobin, Natural bactericidal surfaces: mechanical rupture of Pseudomonas aeruginosa cells by cicada wings, Small, 2012, 8(16), 2489–2494 CrossRef CAS PubMed.
- S. Pogodin, J. Hasan, V. A. Baulin, H. K. Webb, V. K. Truong, T. H. P. Nguyen, V. Boshkovikj, C. J. Fluke, G. S. Watson and J. A. Watson, Biophysical model of bacterial cell interactions with nanopatterned cicada wing surfaces, Biophys. J., 2013, 104(4), 835–840 CrossRef CAS PubMed.
- E. P. Ivanova, J. Hasan, H. K. Webb, G. Gervinskas, S. Juodkazis, V. K. Truong, A. H. Wu, R. N. Lamb, V. A. Baulin and G. S. Watson, Bactericidal activity of black silicon, Nat. Commun., 2013, 4(1), 1–7 Search PubMed.
- V. Truong, N. Geeganagamage, V. Baulin, J. Vongsvivut, M. Tobin, P. Luque, R. Crawford and E. Ivanova, The susceptibility of Staphylococcus aureus CIP 65.8 and Pseudomonas aeruginosa ATCC 9721 cells to the bactericidal action of nanostructured Calopteryx haemorrhoidalis damselfly wing surfaces, Appl. Microbiol. Biotechnol., 2017, 101(11), 4683–4690 CrossRef CAS PubMed.
- X. Li, G. Cheung, G. Watson, J. Watson, S. Lin, L. Schwarzkopf and D. Green, The nanotipped hairs of gecko skin and biotemplated replicas impair and/or kill pathogenic bacteria with high efficiency, Nanoscale, 2016, 8(45), 18860–18869 RSC.
- S. Han, S. Ji, A. Abdullah, D. Kim, H. Lim and D. Lee, Superhydrophilic nanopillar-structured quartz surfaces for the prevention of biofilm formation in optical devices, Appl. Surf. Sci., 2018, 429, 244–252 CrossRef CAS.
- D. P. Linklater, S. Juodkazis, R. J. Crawford and E. P. Ivanova, Mechanical inactivation of Staphylococcus aureus and Pseudomonas aeruginosa by titanium substrata with hierarchical surface structures, Materialia, 2019, 5, 100197 CrossRef CAS.
- C. M. Bhadra, V. K. Truong, V. T. Pham, M. Al Kobaisi, G. Seniutinas, J. Y. Wang, S. Juodkazis, R. J. Crawford and E. P. Ivanova, Antibacterial titanium nano-patterned arrays inspired by dragonfly wings, Sci. Rep., 2015, 5(1), 1–12 Search PubMed.
- Y. Jang, W. T. Choi, C. T. Johnson, A. J. García, P. M. Singh, V. Breedveld, D. W. Hess and J. A. Champion, Inhibition of bacterial adhesion on nanotextured stainless steel 316L by electrochemical etching, ACS Biomater. Sci. Eng., 2018, 4(1), 90–97 CrossRef CAS PubMed.
- J. H. Reed, A. E. Gonsalves, J. K. Román, J. Oh, H. Cha, C. E. Dana, M. Toc, S. Hong, J. B. Hoffman and J. E. Andrade, Ultrascalable multifunctional nanoengineered copper and aluminum for antiadhesion and bactericidal applications, ACS Appl. Bio Mater., 2019, 2(7), 2726–2737 CrossRef CAS PubMed.
- A. Elbourne, V. E. Coyle, V. K. Truong, Y. M. Sabri, A. E. Kandjani, S. K. Bhargava, E. P. Ivanova and R. J. Crawford, Multi-directional electrodeposited gold nanospikes for antibacterial surface applications, Nanoscale Adv., 2019, 1(1), 203–212 RSC.
- M. N. Dickson, E. I. Liang, L. A. Rodriguez, N. Vollereaux and A. F. Yee, Nanopatterned polymer surfaces with bactericidal properties, Biointerphases, 2015, 10(2), 021010 CrossRef PubMed.
- A. Elbourne, J. Chapman, A. Gelmi, D. Cozzolino, R. J. Crawford and V. K. Truong, Bacterial-nanostructure interactions: The role of cell elasticity and adhesion forces, J. Colloid Interface Sci., 2019, 546, 192–210 CrossRef CAS PubMed.
- K. Modaresifar, S. Azizian, M. Ganjian, L. E. Fratila-Apachitei and A. A. Zadpoor, Bactericidal effects of nanopatterns: a systematic review, Acta Biomater., 2019, 83, 29–36 CrossRef CAS PubMed.
- R. Tan, N. Marzolini, P. Jiang and Y. Jang, Bio-Inspired Polymer Thin Films with Non-Close-Packed Nanopillars for Enhanced Bactericidal and Antireflective Properties, ACS Appl. Polym. Mater., 2020, 2(12), 5808–5816 CrossRef CAS.
- Q. Cui, T. Liu, X. Li, K. Song and D. Ge, Nanopillared Polycarbonate Surfaces Having Variable Feature Parameters as Bactericidal Coatings, ACS Appl. Nano Mater., 2020, 3(5), 4599–4609 CrossRef CAS.
- M. E. Davey and G. A. O'toole, Microbial Biofilms: from Ecology to Molecular Genetics, Microbiol. Mol. Biol. Rev., 2000, 64(4), 847–867 CrossRef CAS PubMed.
- Y. Shamis, A. Taube, Y. Shramkov, N. Mitik-Dineva, B. Vu and E. P. Ivanova, Development of a microwave treatment technique for bacterial decontamination of raw meat, Int. J. Food Eng., 2008, 4(3), 8 Search PubMed.
- M. Michalska, F. Gambacorta, R. Divan, I. S. Aranson, A. Sokolov, P. Noirot and P. D. Laible, Tuning antimicrobial properties of biomimetic nanopatterned surfaces, Nanoscale, 2018, 10(14), 6639–6650 RSC.
- M. A. Hopcroft, W. D. Nix and T. W. Kenny, What is the Young's Modulus of Silicon?, J. Microelectromech. Syst., 2010, 19(2), 229–238 CAS.
- E. W. Wong, P. E. Sheehan and C. M. Lieber, Nanobeam mechanics: elasticity, strength, and toughness of nanorods and nanotubes, Science, 1997, 277(5334), 1971–1975 CrossRef CAS.
- E. P. Ivanova, D. P. Linklater, M. Werner, V. A. Baulin, X. Xu, N. Vrancken, S. Rubanov, E. Hanssen, J. Wandiyanto and V. K. Truong, The multi-faceted mechano-bactericidal mechanism of nanostructured surfaces, Proc. Natl. Acad. Sci. U. S. A., 2020, 117(23), 12598–12605 CrossRef CAS PubMed.
- D. P. Linklater, M. De Volder, V. A. Baulin, M. Werner, S. Jessl, M. Golozar, L. Maggini, S. Rubanov, E. Hanssen and S. Juodkazis, High aspect ratio nanostructures kill bacteria via storage and release of mechanical energy, ACS Nano, 2018, 12(7), 6657–6667 CrossRef CAS PubMed.
- S. C. Lohmann, A. Tripathy, A. Milionis, A. Keller and D. Poulikakos, Effect of Flexibility and Size of Nanofabricated Topographies on the Mechanobactericidal Efficacy of Polymeric Surfaces, ACS Appl. Bio Mater., 2022, 5(4), 1564–1575 CrossRef CAS PubMed.
- H.-H. Park, K. Sun, M. Seong, M. Kang, S. Park, S. Hong, H. Jung, J. Jang, J. Kim and H. E. Jeong, Lipid-hydrogel-nanostructure hybrids as robust biofilm-resistant polymeric materials, ACS Macro Lett., 2018, 8(1), 64–69 CrossRef PubMed.
- S. Park, H.-H. Park, K. Sun, Y. Gwon, M. Seong, S. Kim, T.-E. Park, H. Hyun, Y.-H. Choung and J. Kim, Hydrogel nanospike patch as a flexible anti-pathogenic scaffold for regulating stem cell behavior, ACS Nano, 2019, 13(10), 11181–11193 CrossRef CAS PubMed.
- H.-K. Kim, Y.-S. Cho and H.-H. Park, PEGDMA-Based Pillar-Shape Nanostructured Antibacterial Films Having Mechanical Robustness, ACS Appl. Bio Mater., 2022, 5(6), 3006–3012 CrossRef CAS PubMed.
- L. Peng, L. Chang, X. Liu, J. Lin, H. Liu, B. Han and S. Wang, Antibacterial Property of a Polyethylene Glycol-Grafted Dental Material, ACS Appl. Mater. Interfaces, 2017, 9(21), 17688–17692 CrossRef CAS PubMed.
- N.-U. Kang, G. H. Kim, H.-K. Kim, S. H. Kim, Y. Y. Kim, H.-H. Park and Y.-S. Cho, Poly(ethylene glycol) dimethacrylate (PEGDMA) multi-functional pillar-patterned surface for osteogenic differentiation of pre-osteoblast and anti-bacterial effects to Escherichia coli and Staphylococcus aureus, Surf. Interfaces, 2024, 55, 105221 CrossRef CAS.
- K. W. Kolewe, S. R. Peyton and J. D. Schiffman, Fewer Bacteria Adhere to Softer Hydrogels, ACS Appl. Mater. Interfaces, 2015, 7(35), 19562–19569 CrossRef CAS PubMed.
- K. Hori and S. Matsumoto, Bacterial adhesion: From mechanism to control, Biochem. Eng. J., 2010, 48(3), 424–434 CrossRef CAS.
- W. L. Min, B. Jiang and P. Jiang, Bioinspired self-cleaning antireflection coatings, Adv. Mater., 2008, 20(20), 3914–3918 CrossRef CAS.
- M. D. Levenson, N. S. Viswanathan and R. A. Simpson, Improving resolution in photolithography with a phase-shifting mask, IEEE Trans. Electron Devices, 1982, 29(12), 1828–1836 Search PubMed.
- C. Vieu, F. Carcenac, A. Pépin, Y. Chen, M. Mejias, A. Lebib, L. Manin-Ferlazzo, L. Couraud and H. Launois, Electron beam lithography: resolution limits and applications, Appl. Surf. Sci., 2000, 164(1–4), 111–117 CrossRef CAS.
- J. Melngailis, Focused ion beam technology and applications, J. Vac. Sci. Technol., B: Microelectron. Process. Phenom., 1987, 5(2), 469–495 CrossRef CAS.
- Y. Xia, J. A. Rogers, K. E. Paul and G. M. Whitesides, Unconventional Methods for Fabricating and Patterning Nanostructures, Chem. Rev., 1999, 99(7), 1823–1848 CrossRef CAS PubMed.
- S. Y. Chou, P. R. Krauss and P. J. Renstrom, Imprint Lithography with 25-Nanometer Resolution, Science, 1996, 272(5258), 85–87 CrossRef CAS.
- Y. Wang, M. Zhang, Y. Lai and L. Chi, Advanced colloidal lithography: From patterning to applications, Nano Today, 2018, 22, 36–61 CrossRef CAS.
- W. Stöber, A. Fink and E. Bohn, Controlled growth of monodisperse silica spheres in the micron size range, J. Colloid Interface Sci., 1968, 26(1), 62–69 CrossRef.
- M. Garcia, K. D. Schulze, C. S. O'Bryan, T. Bhattacharjee, W. G. Sawyer and T. E. Angelini, Eliminating the surface location from soft matter contact mechanics measurements, Tribol. - Mater. Surf. Interfaces, 2017, 11(4), 187–192 CrossRef.
- P. Kothary, B. M. Phillips, S.-Y. Leo and P. Jiang, Bioinspired broadband midwavelength infrared antireflection coatings on silicon, J. Vac. Sci. Technol., B: Nanotechnol. Microelectron.: Mater., Process., Meas., Phenom., 2016, 34(4), 041807 Search PubMed.
- P. V. Antonov, M. R. Zuiddam and J. W. Frenken, Fabrication of high-aspect ratio silicon nanopillars for tribological experiments, J. Micro/Nanolithogr., MEMS, MOEMS, 2015, 14(4), 044506 CrossRef.
- S. Y. Heo, J. K. Koh, G. Kang, S. H. Ahn, W. S. Chi, K. Kim and J. H. Kim, Bifunctional moth-eye nanopatterned dye-sensitized solar cells: light-harvesting and self-cleaning effects, Adv. Energy Mater., 2014, 4(3), 1300632 CrossRef.
- A. Valiei, N. Lin, J.-F. Bryche, G. McKay, M. Canva, P. G. Charette, D. Nguyen, C. Moraes and N. Tufenkji, Hydrophilic mechano-bactericidal nanopillars require external forces to rapidly kill bacteria, Nano Lett., 2020, 20(8), 5720–5727 CrossRef CAS PubMed.
- J. A. Killion, L. M. Geever, D. M. Devine, J. E. Kennedy and C. L. Higginbotham, Mechanical properties and thermal behaviour of PEGDMA hydrogels for potential bone regeneration application, J. Mech. Behav. Biomed. Mater., 2011, 4(7), 1219–1227 CrossRef CAS PubMed.
- N. R. Patel, A. K. Whitehead, J. J. Newman and M. E. Caldorera-Moore, Poly (ethylene glycol) hydrogels with tailorable surface and mechanical properties for tissue engineering applications, ACS Biomater. Sci. Eng., 2017, 3(8), 1494–1498 CrossRef CAS PubMed.
- J. A. Killion, L. M. Geever, D. M. Devine, L. Grehan, J. E. Kennedy and C. L. Higginbotham, Modulating the mechanical properties of photopolymerised polyethylene glycol–polypropylene glycol hydrogels for bone regeneration, J. Mater. Sci., 2012, 47(18), 6577–6585 CrossRef CAS.
- G. Burke, V. Barron, T. Geever, L. Geever, D. M. Devine and C. L. Higginbotham, Evaluation of the materials properties, stability and cell response of a range of PEGDMA hydrogels for tissue engineering applications, J. Mech. Behav. Biomed. Mater., 2019, 99, 1–10 CrossRef CAS PubMed.
- A. M. Ortega, S. E. Kasprzak, C. M. Yakacki, J. Diani, A. R. Greenberg and K. Gall, Structure–property relationships in photopolymerizable polymer networks: Effect of composition on the crosslinked structure and resulting thermomechanical properties of a (meth) acrylate-based system, J. Appl. Polym. Sci., 2008, 110(3), 1559–1572 CrossRef CAS.
- K. W. Kolewe, J. Zhu, N. R. Mako, S. S. Nonnenmann and J. D. Schiffman, Bacterial Adhesion Is Affected by the Thickness and Stiffness of Poly(ethylene glycol) Hydrogels, ACS Appl. Mater. Interfaces, 2018, 10(3), 2275–2281 CrossRef CAS PubMed.
- M. Techakasikornpanich, K. Jangpatarapongsa, D. Polpanich and A. Elaissari, Impact of polymeric films and hydrogels: Physical characteristics on bacterial growth, Polym. Adv. Technol., 2024, 35(2), e6311 CrossRef CAS.
- L. Teng, S. Lee, A. Ginn, S. M. Markland, R. A. Mir, N. DiLorenzo, C. Boucher, M. Prosperi, J. Johnson and J. G. Morris Jr., Genomic comparison reveals natural occurrence of clinically relevant multidrug-resistant extended-spectrum-β-lactamase-producing Escherichia coli strains, Appl. Environ. Microbiol., 2019, 85(13), e03030-18 CrossRef PubMed.
- F. Song, H. Koo and D. Ren, Effects of material properties on bacterial adhesion and biofilm formation, J. Dent. Res., 2015, 94(8), 1027–1034 CrossRef CAS PubMed.
- L. Boulos, M. Prevost, B. Barbeau, J. Coallier and R. Desjardins, LIVE/DEAD® BacLight™: application of a new rapid staining method for direct enumeration of viable and total bacteria in drinking water, J. Microbiol. Methods, 1999, 37(1), 77–86 CrossRef CAS PubMed.
- S. Zheng, M. Bawazir, A. Dhall, H.-E. Kim, L. He, J. Heo and G. Hwang, Implication of surface properties, bacterial motility, and hydrodynamic conditions on bacterial surface sensing and their initial adhesion, Front. Bioeng. Biotechnol., 2021, 9, 82 Search PubMed.
- P. Müller and A. Saúl, Elastic effects on surface physics, Surf. Sci. Rep., 2004, 54(5–8), 157–258 CrossRef.
- T. Liu, Q. Cui, Q. Wu, X. Li, K. Song, D. Ge and S. Guan, Mechanism study of bacteria killed on nanostructures, J. Phys. Chem. B, 2019, 123(41), 8686–8696 CrossRef CAS PubMed.
- O. A. Bauchau and J. I. Craig, Euler-Bernoulli beam theoryl, in Structural analysis, Springer, 2009, pp. 173–221 Search PubMed.
- M. C. Enright, D. A. Robinson, G. Randle, E. J. Feil, H. Grundmann and B. G. Spratt, The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA), Proc. Natl. Acad. Sci. U. S. A., 2002, 99(11), 7687–7692 CrossRef CAS PubMed.
- C. N. Cornelissen, R. A. Harvey and B. D. Fisher, Microbiology, Lippincott Williams & Wilkins, 2012, vol. 3 Search PubMed.
- M. Otto, Staphylococcal Biofilms, Curr. Top. Microbiol. Immunol., 2008, 322, 207–228 CAS.
- S. Furukawa, Y. Akiyoshi, M. Komoriya, H. Ogihara and Y. Morinaga, Removing Staphylococcus aureus and Escherichia coli biofilms on stainless steel by cleaning-in-place (CIP) cleaning agents, Food Control, 2010, 21(5), 669–672 CrossRef CAS.
- T. Duanis-Assaf and M. Reches, Factors influencing initial bacterial adhesion to antifouling surfaces studied by single-cell force spectroscopy, iScience, 2024, 27(2), 108803 CrossRef CAS PubMed.
- C. Heilmann, Adhesion Mechanisms of Staphylococci, Bacterial Adhesion: Chemistry, Biology and Physics, ed. D. Linke and A. Goldman, Springer Netherlands, Dordrecht, 2011, pp. 105–123 Search PubMed.
- W. M. Dunne, Bacterial adhesion: seen any good biofilms lately?, Clin. Microbiol. Rev., 2002, 15(2), 155–166 CrossRef CAS PubMed.
- I. Rebane, H. Priks, K. J. Levin, İ. Sarigül, U. Mäeorg, U. Johanson, P. Piirimägi, T. Tenson and T. Tamm, Microbial growth and adhesion of Escherichia coli in elastomeric silicone foams with commonly used additives, Sci. Rep., 2023, 13(1), 8541 CrossRef CAS PubMed.
- K. M. Craft, J. M. Nguyen, L. J. Berg and S. D. Townsend, Methicillin-Resistant Staphylococcus aureus (MRSA): Antibiotic Resistance and the Biofilm Phenotype, MedChemComm, 2019, 10(8), 1231–1241 RSC.
- E. Maikranz, C. Spengler, N. Thewes, A. Thewes, F. Nolle, P. Jung, M. Bischoff, L. Santen and K. Jacobs, Different Binding Mechanisms of Staphylococcus aureus to Hydrophobic and Hydrophilic Surfaces, Nanoscale, 2020, 12(37), 19267–19275 RSC.
- L. A. Pratt and R. Kolter, Genetic analysis of Escherichia coli biofilm formation: roles of flagella, motility, chemotaxis and type I pili, Mol. Microbiol., 1998, 30(2), 285–293 CrossRef CAS PubMed.
- G. Choi, Y. Song, H. Lim, S. H. Lee, H. K. Lee, E. Lee, B. G. Choi, J. J. Lee, S. G. Im and K. G. Lee, Antibacterial nanopillar array for an implantable intraocular lens, Adv. Healthcare Mater., 2020, 9(18), 2000447 CrossRef CAS.
- S. H. Kang, B. Pokroy, L. Mahadevan and J. Aizenberg, Control of shape and size of nanopillar assembly by adhesion-mediated elastocapillary interaction, ACS Nano, 2010, 4(11), 6323–6331 CrossRef CAS PubMed.
Footnote |
| † Co-first authors. |
| This journal is © The Royal Society of Chemistry 2026 |