Open Access Article
Open Access ArticleStable double gyroid network phases in asymmetric linear triblock amphiphiles
Daoyuan
Li
ab,
Mahesh K.
Mahanthappa
 a,
Timothy P.
Lodge
a,
Timothy P.
Lodge
 *ac and
J. Ilja
Siepmann
*ac and
J. Ilja
Siepmann
 *abc
*abc
aDepartment of Chemical Engineering and Materials Science, University of Minnesota, 421 Washington Avenue SE, Minneapolis, Minnesota 55455-0132, USA. E-mail: lodge@umn.edu; siepmann@umn.edu
bChemical Theory Center, University of Minnesota, 207 Pleasant Street SE, Minneapolis, Minnesota 55455-0431, USA
cDepartment of Chemistry, University of Minnesota, 207 Pleasant Street SE, Minneapolis, Minnesota 55455-0431, USA
First published on 30th April 2026
Abstract
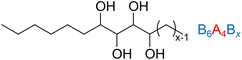
Molecular dynamics simulations are employed to elucidate the phase behavior of a series of asymmetric linear triblock amphiphiles consisting of a sugar-based center segment (A4) containing four hydroxyl groups with the ability to form inter- and intramolecular hydrogen bonds that is flanked by two nonpolar hydrocarbon segments (C6H13 and CxH2x+1, denoted as B6 and Bx with 6 ≤ x ≤ 14). In agreement with prior simulations, the B6A4B6 amphiphile with symmetric hydrocarbon blocks preferentially self-assembles into hexagonally perforated lamellae, whereas those with asymmetric hydrocarbon segments and n ≤ 12 can form stable double gyroid networks. Within these double gyroid morphologies, interconnected channel domains are composed of sugar-based segments embedded in a continuous hydrocarbon matrix. Remarkably small feature sizes ranging from 1.92 to 2.26 nm are observed as the sugar volume fraction (fA) varies from 0.23 to 0.29. Structural analyses further reveal distinct local enhancements in oxygen density—indicative of concentrated sugar-based segments—predominantly at the gyroid nodes, accompanied by reduced densities along the connecting struts. Additionally, a mean curvature analysis confirms that these fairly stiff oligomers form gyroid structures containing a higher fraction of regions with high curvature compared to idealized gyroid structures. Together with investigations of related miktoarm triblock oligomers (AoBpBq), this study highlights the role of asymmetry in the molecular design of amphiphilic block oligomers, which facilitates the stabilization of network morphologies with ultra-small feature sizes and provides insight into the microscopic packing behavior of amphiphiles.
1 Introduction
Block polymers and amphiphilic oligomers have emerged as versatile materials for applications as mesoporous materials and templates for inorganic analogs,1–3 thermoplastic elastomers,4,5 and drug delivery systems.6,7 Microphase separated linear block copolymers typically display domain spacings in the 10–50 nm range; however, high-χ (low-N) block oligomers—characterized by their small molecular size—can generate morphologies with feature dimensions below 5 nm,8–13 opening up new possibilities for nanofiltration, nanopatterning, and other nanoscale technologies.14–18Self-assembly in block polymer and amphiphilic oligomer systems yields a diverse array of ordered structures, including hexagonally perforated lamellae (HPL), hexagonally packed cylinders (CYL), body-centered cubic micelles (BCC), and network (NET) morphologies.19–22 NET structures, characterized by bicontinuous interpenetrating domains,23 are especially attractive for applications such as nanoporous membranes,24–26 energy storage devices,27–30 biophotonic constructs31,32 and photonic crystals.33–35 However, the stabilization of NET morphologies is hampered by packing frustration that restricts their formation to narrow compositional windows,36–38 and by the challenge of designing molecular architectures that can accommodate the varying curvatures inherent in these structures. For instance, the double gyroid (DG) phase exhibits pronounced variations in mean curvature across different local regions of the unit cell,39 necessitating tailored molecular designs. Strategies to broaden the stability window of NET structures include blending components that naturally favor different morphologies11 and developing innovative architectures such as coil–brush or bottlebrush polymers and shape-specific bolapolyphiles.40–46
The drive for miniaturization further underscores the importance of achieving NET structures with ultra-small features. Melt-ordered block polymers are limited by their long chain lengths (N), which restrict the attainable pore sizes and unit cell dimensions in NET morphologies. To overcome these constraints, recent efforts have focused on employing blocks with larger effective segmental interaction parameters (χ), enabling the stabilization of NET phases in low-N systems.47 In this context, amphiphiles—with a large χ between hydrophilic and hydrophobic segments and their low molecular weight—are particularly effective at stabilizing NET structures with extremely small dimensions.48–52
Our previous work, involving molecular simulations, synthesis of high-purity compounds, and experiments, has shown that amphiphilic block oligomers containing sugar-based (in either open or ring forms) and nonpolar hydrocarbon segments can form mesophases with remarkably small domain sizes.8–13 For instance, Shen et al. demonstrated that triblock amphiphiles, such as B6A4B6, can form stable HPL with characteristic length scales approaching 1 nm,10 and Li et al. found that miktoarm triblock oligomers with two tails of different length can stabilize double gyroid NET structures.12 Furthermore, self-consistent field theory (SCFT) studies on asymmetric ABA triblock copolymer melts indicate that introducing molecular asymmetry can stabilize DG structures, suggesting that rigid amphiphilic oligomers with similar triblock architectures may exhibit comparable scale-invariant phase behavior.53
In the present study, molecular dynamics (MD) simulations employing molecular mechanics force fields are used to investigate the self-assembly of neat amphiphilic block oligomers into various morphologies. An asymmetric molecular design converts HPL-forming B6A4B6 amphiphiles into B6A4Bx, which comprise a hydrophilic tetraol segment (A4, consisting of four –CHOH– repeat units) and two hydrophobic alkyl segments (B6 and Bx), each built from –CH2– repeat units (see Fig. 1). Moreover, the microscopic packing of amphiphiles in DG phases is analyzed using methods based on coordinates, density, clustering, and curvature projections, and the order–disorder transition is examined by tracking clustering and level surface distributions along the simulation trajectories.
These simulations are motivated by the possibility of accessing these asymmetric B6A4Bx amphiphiles by an analogous approach to that reported by Barrreda et al. for B13A6 amphiphiles9 and extended by Mattson et al. to related diblock amphiphiles.54 Specifically, tin-mediated allylation of erythrose (or another aldose) followed by olefin cross-metathesis with 1-pentene and tandem hydrogenation of the resulting would yield the B6A4 diblock oligomer. Selective oxidation of the terminal –CH2OH of the A4 segment would yield an aldehyde, which could again undergo tin-mediated allylation, olefin cross metathesis, and hydrogenation to yield the desired B6A4Bx (x ≥ 6) amphiphile. The thermotropic liquid crystalline network morphologies adopted by these amphiphiles could then be identified by small-angle X-ray scattering (SAXS), and they would be expected to exhibit no optical birefringence.9 While these synthesis and characterization experiments are in principle straightforward, optimization of the synthesis and purification to access useful quantities of these amphiphiles for complete molecular and supramolecular assembly characterization could be nontrivial. Thus, this motivates simulation studies of their expected properties prior to embarking on this potentially arduous synthesis task.
2 Methods
2.1 Molecular models and simulation details
The B6A4Bx amphiphiles are modeled using the transferable potentials for phase equilibria united-atom (TraPPE-UA) force field.55–58 Detailed force field parameters are provided in the SI (see Tables S1 and S2). MD simulations are performed in the isobaric–isothermal (NPT) ensemble using GROMACS 2021.3.59,60 System sizes range from 1000 to 2000 molecules. Temperature and pressure are controlled via the Nosé–Hoover thermostat61,62 (with a time constant τT = 0.4 ps) and the Parrinello–Rahman barostat63 (with τp = 2 ps), respectively. Electrostatic interactions are computed using the particle-mesh Ewald method,64 and the p-LINCS algorithm is used to constrain O–H bond lengths,65 which allows for a 2 fs integration time step. Initial disordered configurations are generated by short Monte Carlo (MC) simulations using coupled-decoupled configurational-bias (CD-CB) MC moves56 in the canonical ensemble at a very high temperature of T = 3000 K (but harmonic bonds cannot break). Because the united-atom model utilizes CH pseudo-atoms also for the sugar-based block and the CD-CB regrowth algorithm does not preserve tacticity, each amphiphile should be viewed as a random mixture of stereoisomers. Systems are then equilibrated at T = 445 K and 1 bar using MD trajectories of at least 1 μs, where equilibration is assessed through system energy and volume reaching converged values and analysis of static structure factors performed for 100 ns segments not showing any significant changes. Equilibration is followed by additional trajectories of 200 ns for HPL and 400 ns for DG phases used for detailed structural analyses. Comparisons between experimental and simulated phase boundaries and static structure factors for related A6B13 diblock amphiphiles9 and A2BxA2 (8 ≤ x ≤ 12) triblock amphiphiles10 and for the vapor–liquid equilibria of binary (methanol or ethanol)/n-hexane mixtures57 indicate that the non-polarizable TraPPE-UA force field yields very good predictions for the domain spacing of lamellar phases but overpredicts the effective χ value and, hence, phase transitions occur at higher temperatures than for the corresponding experimental systems. Visualization of simulation snapshots, including those showing dividing surfaces and ball-and-stick representations, is performed using Ovito and VMD.66–682.2 System size tuning and workflow
Initial simulations for all B6A4Bx systems are conducted in orthorhombic cells containing either 1000 or 2000 molecules. This setup permits independent fluctuations in the x-, y-, and z-dimensions, thereby mitigating the effects of incommensurability,69,70 and is well-suited for morphologies with one- or two-dimensional periodicity (e.g., LAM and CYL structures), where domain spacing can be adjusted via rotations and lateral expansions or contractions perpendicular to the lamellar planes or cylinder axes. In contrast, for three-dimensional periodic morphologies (e.g., BCC and NET structures), 1000-molecule systems tend to yield locally segregated but globally disordered micelles or networks due to persistent incommensurability, unless the simulation box dimensions are integer multiples of the unit cell. To achieve ordered NET structures, it is critical to determine both the lattice parameter (a) and the number of molecules per unit cell (NUC) in order to avoid distorted, metastable configurations.71–73 The unit cell dimension is estimated by calculating the structure factor S(q) for disordered NET-like structures and identifying the broad peak position via the relation a = 2πm/q*, where m corresponds to the first observable reflection spacing ratio for the morphology (e.g.,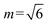 for DG). The value of NUC is then estimated based on the average molecular volume derived from the initial simulations. It should be noted that the broad peak found in the disordered NET-like structures leads to considerable uncertainty in the estimation of a and, in some cases, it is advantageous to explore a range of NUC values. For each B6A4Bx system that initially forms disordered NET structures, simulations are restarted with 8NUC molecules in a cubic box of length Lbox = 2a, which facilitates the formation of ordered NET phases, such as double gyroid (DG), double diamond (DD), single gyroid (SG), and double primitive (DP, also known as plumber's nightmare). To promote ordering, a workflow for 3D NET simulations is employed that includes the application of a guiding field, generated by Gaussian interaction sites with varying strengths for polar and nonpolar beads and distributed within the minority and majority domains, respectively, of the candidate NET structures.11–13 After removal of the guiding field, the stability of each candidate NET morphology is assessed over a 1 μs trajectory in the NPT ensemble but maintaining a cubic box shape. The values of N and additional details regarding the DG unit cells are provided in Table 1.
for DG). The value of NUC is then estimated based on the average molecular volume derived from the initial simulations. It should be noted that the broad peak found in the disordered NET-like structures leads to considerable uncertainty in the estimation of a and, in some cases, it is advantageous to explore a range of NUC values. For each B6A4Bx system that initially forms disordered NET structures, simulations are restarted with 8NUC molecules in a cubic box of length Lbox = 2a, which facilitates the formation of ordered NET phases, such as double gyroid (DG), double diamond (DD), single gyroid (SG), and double primitive (DP, also known as plumber's nightmare). To promote ordering, a workflow for 3D NET simulations is employed that includes the application of a guiding field, generated by Gaussian interaction sites with varying strengths for polar and nonpolar beads and distributed within the minority and majority domains, respectively, of the candidate NET structures.11–13 After removal of the guiding field, the stability of each candidate NET morphology is assessed over a 1 μs trajectory in the NPT ensemble but maintaining a cubic box shape. The values of N and additional details regarding the DG unit cells are provided in Table 1.
| Compound | N | V m [nm3] | f A | Morphology | d [nm] |
|---|---|---|---|---|---|
| a The statistical error for d ranges from 0.020 to 0.034 nm, in terms of the 95% confidence interval estimated by dividing the simulation trajectories into 10 blocks. However, the uncertainty for d introduced due to selecting a specific value of N is estimated to be about 0.05 nm. | |||||
| B6A4B6 | 1504 | 0.53 | 0.30 | HPL | 1.92 |
| B6A4B7 | 1448 | 0.56 | 0.28 | DG | 1.92 |
| B6A4B8 | 1448 | 0.59 | 0.27 | DG | 1.95 |
| B6A4B9 | 1640 | 0.63 | 0.25 | DG | 2.04 |
| B6A4B10 | 1766 | 0.66 | 0.24 | DG | 2.15 |
| B6A4B11 | 1872 | 0.69 | 0.23 | DG | 2.22 |
| B6A4B12 | 1872 | 0.72 | 0.22 | DG | 2.26 |
| B6A4B13 | 2000 | 0.75 | 0.21 | DIS | 2.39 |
| B6A4B14 | 2000 | 0.78 | 0.20 | DIS | 2.44 |
In principle, it would be desirable to estimate the molar free energy for the different NET phases and for different system sizes. Here, we find that only one NET phase remains stable over these long trajectories, so this phase is certainly lower in free energy. However, simulations with slightly different N for the desired NET phase can yield stable trajectories. Since the systems differ in N, one cannot construct a reversible path from one size to another and instead would have to perform thermodynamic integration along a reversible path to the ideal-gas state at a common reference density. Given the complexity of the amphiphiles with inter- and intramolecular hydrogen bonding and a large number of torsional degrees of freedom, such a thermodynamic integration would encounter significant sampling problems and likely not allow for small enough uncertainties to determine which system size corresponds to the free energy minimum. It is reasonable to expect that the optimal value of N should increase with x for the B6A4Bx amphiphiles, and the fact that N = 1448 is used for B6A4B7 and B6A4B8 (and N = 1872 is used for B6A4B11 and B6A4B12) indicates that the robust stability of the equilibrium state allows it to tolerate a certain degree of incommensurability, thereby not requiring the optimal number of molecules in the simulation box.69 Consequently, although the most stable ordered structure is obtained after completion of the workflow, the system size may not be ideal. However, when the incommensurability is large, then the resulting stable NET structure exhibits structural imperfections. For B6A4B9, N = 1448, 1544, and 1640 were explored and found to give d211 values of 1.98, 2.01, and 2.04 nm, respectively. Among these, N = 1640 yields a static structure factor with the minor peaks more clearly resolved (see Fig. S1), and its d211 value falls closer to the midway point between the d211 spacings for B6A4B8 and B6A4B10, thereby indicating that N = 1640 is the better choice among these three candidates.
3 Results and discussion
3.1 Phase behaviors of asymmetric B6A4Bx
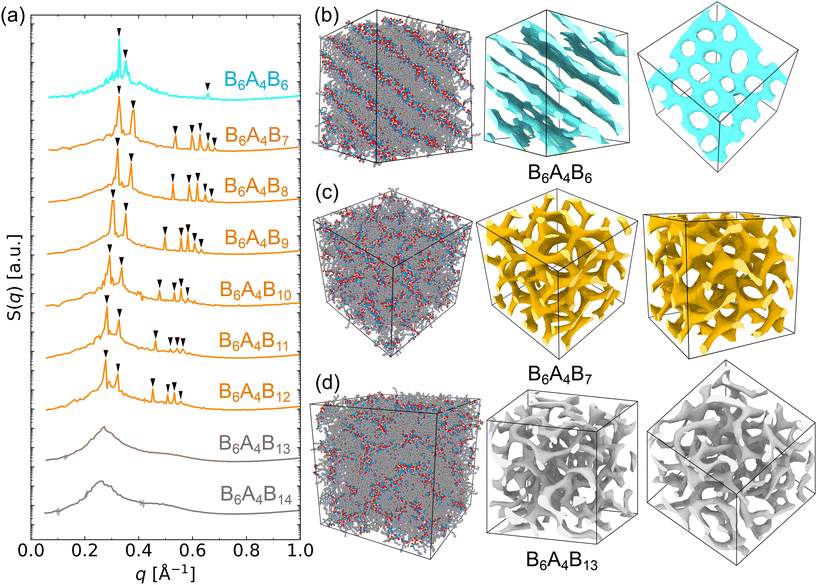
As incrementing x increases the tail length by one CH2 unit at a time, the molecular volume of the B6A4Bx amphiphiles increases linearly (see Table 1 and Fig. S2). A linear regression analysis indicates that the incremental volume of the B6A4 segment is approximately 0.34 nm3 and that of the B6 segment is about 0.19 nm3. Consequently, the volume fraction of the hydrophilic segment (fA) is estimated to vary from 0.30 (for x = 6) to 0.20 (for x = 14) (see Fig. S2).For x = 6 (i.e., B6A4B6), stable hexagonally perforated lamellae (HPL) are observed (see Fig. 2b). The periodicity of these structures is confirmed via the static structure factor (detailed calculation procedures can be found in the SI). Although the temperature is higher by 5 K compared to previous simulations for B6A4B6,10 the structure factor is nearly identical, exhibiting a primary peak along with a closely spaced secondary peak that corresponds to the periodic arrangement of perforations in each layer (see Fig. 2a).
Stable double gyroid (DG) structures are observed for amphiphiles with x ranging from 7 to 12, with d211-spacings increasing from 1.92 nm in B6A4B7 to 2.26 nm in B6A4B12 (see Fig. 2 and Table 1). The characteristic relative peak positions are located at 1, 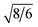 ,
, 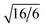 ,
, 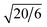 ,
, 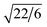 ,
, 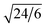 , and
, and 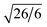 , which are consistent with the Ia
, which are consistent with the Ia![[3 with combining macron]](https://www.rsc.org/images/entities/char_0033_0304.gif) d (Q230) space group symmetry,74 are detected in the structure factors. However, for B6A4Bx with larger x values and closer to the order–disorder boundary anticipated based on linear triblock copolymers, low-intensity peaks at higher q become less discernible. To exclude alternative network structures and to verify that the equilibrium morphology of B6A4Bx amphiphiles is indeed the DG phase, the workflow with guiding fields was also used to generate DD, SG, and DP structures, but these network structures are found to rapidly become disordered within 50 ns after removal of the guiding field (see Fig. S3 for the structural evolution of the B6A4B7 system), thereby eliminating them as candidates for stable network structures and confirming that the DG morphology represents the most stable state formed by self-assembly.
d (Q230) space group symmetry,74 are detected in the structure factors. However, for B6A4Bx with larger x values and closer to the order–disorder boundary anticipated based on linear triblock copolymers, low-intensity peaks at higher q become less discernible. To exclude alternative network structures and to verify that the equilibrium morphology of B6A4Bx amphiphiles is indeed the DG phase, the workflow with guiding fields was also used to generate DD, SG, and DP structures, but these network structures are found to rapidly become disordered within 50 ns after removal of the guiding field (see Fig. S3 for the structural evolution of the B6A4B7 system), thereby eliminating them as candidates for stable network structures and confirming that the DG morphology represents the most stable state formed by self-assembly.
To further validate the quality of the DG structures formed by these amphiphilic oligomers, a level surface analysis is employed.75–79 The DG morphology is approximated by the level surface
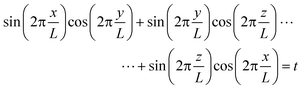 | (1) |
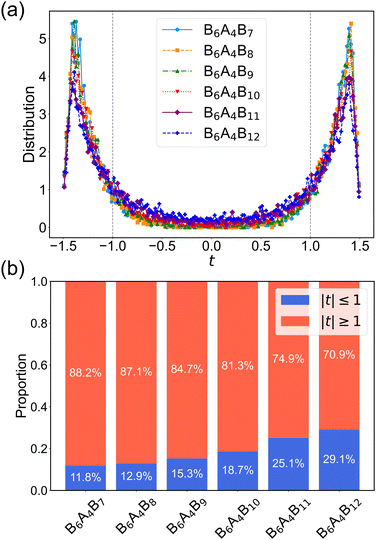
Complementarily, Fig. 3b presents a stacked bar chart that quantifies the proportion of t values inside versus outside the interval [−1, 1]. The histogram shows that less than 30% of the t values fall within this interval, indicating that the majority of oxygen atoms (and thus the A4 segment of the amphiphiles) reside predominantly in the gyroid channels rather than in the interstitial matrix. This detailed analysis reinforces the conclusion that the DG morphology is the most stable equilibrium state. From Fig. 3b, the increasing fraction of t values falling outside of the matrix interval indicates a higher concentration of oxygen atoms within the gyroid matrix. This trend suggests that the interfacial width between the channel and matrix domains is expanding, which constitutes a direct signature of the approach to the order–disorder transition.
For B6A4Bx amphiphiles with larger x values (i.e., x = 13 and 14), the molecules fail to self-assemble into CYL morphology or retain any of the NET morphologies and only a disordered phase (DIS) is observed after a comprehensive exploration of the workflow (see Fig. 2a). This occurs when the volume fraction of the hydrophilic segment (fA) is approximately 0.21, a value that is close to the order–disorder boundary reported in previous SCFT studies on triblock polymers.53 During the workflow exploration, various ordered phases were used as initial configurations for the MD simulations, including relatively stable hexagonally packed cylinders (CYL) and DG phases. When the initial configuration is CYL, B6A4B13 maintains a relatively ordered structure with few defects for around 40 ns, whereas B6A4B14 sustains order for only about 20 ns before transitioning to a DIS phase. Representative snapshots from the simulation trajectories are shown in Fig. S4. In the case of DG being set up as the initial configuration, the DG arrangement in the simulation of B6A4B14 quickly vanishes, while B6A4B13 fully transforms into a DIS phase after approximately 100 ns, but 3-fold connectors persist for a longer time (see Fig. S5). Moreover, static structure factors calculated separately over the 20–50, 50–100, 100–150, 900–1000, and 1900–2000 ns intervals clearly capture the DG-to-DIS transition for the B6A4B13 and B6A4B14 systems (see Fig. S6). For the B6A4B13 system, the structure factor obtained from the 50–100 ns segment exhibits the first two characteristic DG peaks. Extended 2 μs trajectories do not yield re-emergence of the DG morphology for the B6A4B13 and B6A4B14 systems.
3.2 Domain spacing, hydrogen bonding, and amphiphile packing
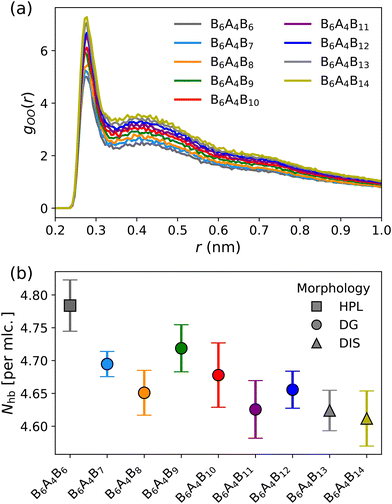
Stable DG structures are observed for amphiphiles with x ranging from 7 to 12, with d211-spacings increasing from 1.92 nm in B6A4B7 to 2.26 nm in B6A4B12. While the increases in d211 values with addition of a methylene group vary from 0.03 to 0.11 nm, an unweighted linear fit yields a correlation coefficient of 0.988 and a slope of 0.075 ± 0.006 nm per CH2 unit. Thus, the variation in step sizes may be caused by the specific system sizes used here not being always optimal.As discussed in our prior work comparing the self-assembly of A2B12A2 and B B6A4B6 triblock amphiphiles, the ability of the B6A4B6 isomer to form intramolecular hydrogen bonds (HB) leads to a preference for U-shaped conformations and a stable HPL phase, whereas the A2B12A2 isomer prefers more stretched conformations that bridge between two polar regions of a LAM structure.10 Here, we focus on the formation of intermolecular HBs in the B6A4Bx amphiphiles as x is increased from 6 to 14 and the morphology changes from HPL to DG to DIS. The oxygen–oxygen radial distribution functions (RDFs) for these systems are shown in Fig. 4(a). Given the low volume fraction of the polar block, fA (see Table 1), the first peak in the OO RDF at 0.28 nm is very pronounced, followed by a broad second peak and a slow decay to a value near unity at 1.0 nm which reflects the cross-sectional size of the polar region. The heights of the first and second peaks decrease monotonically as x decreases. The decrease in peak heights does not indicate a decrease in hydrogen bonding, but reflects the increase in fA which enters into the normalization of the RDFs. The number of HBs per molecules (see Fig. 4(b)) is obtained using an O–O distance less than 0.35 nm and an O–H⋯O angle greater than 150° as the HB criterion. Overall, it is clear that the number of HBs decreases as x increases (i.e., the opposite trend as the peak heights in the RDFs) from a value of 4.78 for B6A4B6 to 4.62 for B6A4B14. The HB data also point to a discontinuous step as the morphology changes from HPL to DG. The statistical uncertainties are too large to conclude that the HB numbers decrease with increasing x for the DG-forming amphiphiles (and the slope would only be ≈−0.01 HB per methylene unit). It should also be noted that, in the case that the optimal NUC could be found for each amphiphile, a more systematic trend may emerge. On the other hand, there does not appear a discontinuous step as the morphology changes from DG to DIS.
A three-dimensional analysis of the DG structures formed by B6A4Bx amphiphiles is needed to elucidate microscopic phenomena in amphiphile packing. Because the hydrophilic A4 segments assemble into gyroid channels, a Gaussian kernel density estimation (KDE) is applied to the oxygen atom coordinates of the A4 segments for DG phases.81,82 The discrete atomic positions are transformed into a continuous probability density function, denoted by ![[f with combining circumflex]](https://www.rsc.org/images/entities/i_char_0066_0302.gif) (r), which represents the estimated probability density of finding an oxygen atom at a given point r in space. A Gaussian kernel is employed, with each atom contributing according to
(r), which represents the estimated probability density of finding an oxygen atom at a given point r in space. A Gaussian kernel is employed, with each atom contributing according to
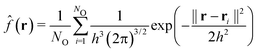 | (2) |
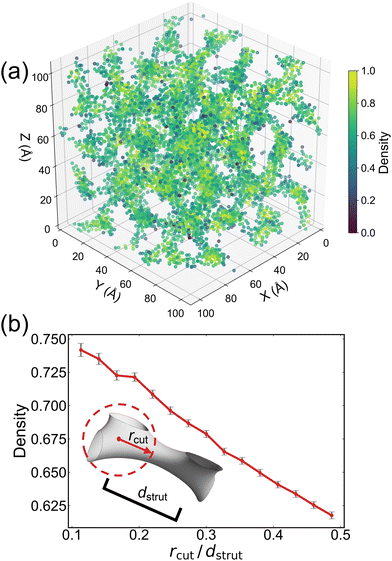
For the DG morphology formed by B6A4B7, the KDE is visualized in three-dimensional space (see Fig. 5a), where each point corresponds to an oxygen atom and the color indicates the normalized density (ranging from 0 to 1). It is observed that isolated points exhibit lower density, whereas points located within the gyroid channels display higher density. To better highlight these high-density regions, Fig. S7 marks all points with density values above the 95th percentile (i.e., 0.87) with red circles. These highlighted points are predominantly situated near the gyroid nodes rather than the struts, indicating that a tighter packing of the hydrophilic A4 segments is required at the nodes to stabilize the DG structure.
To quantitatively analyze the spatial distribution of these points, the gyroid level surface (see eqn (1)) serves as an effective mathematical tool to examine the distribution of the level surface constant t derived from the oxygen atom coordinates. Fig. S8 presents a scatter plot of |t| values versus their KDE density, which clearly shows that high KDE values are only found for |t| > 1. In addition, the frequency histograms along the top and left margins of Fig. S8 reveal that most points are located in the region where |t| ≥ 1.2, with the high-density points concentrated near |t| = 1.3, i.e., close to the center of the nodes.
For DG structures, the right-handed and left-handed gyroid channels each comprise eight nodes, resulting in a total of 16 nodes per unit cell. By calculating the average density of oxygen atoms as a function of the distance from these nodes, the presence of local oxygen density enhancement can be evaluated. Taking the coordinates of the nodes as the centers of spheres with a cutoff radius rcut (with the nearest-neighbor distance between nodes defined as dcut), the average density of oxygen atoms within spherical shells (for increasing rcut bins) is computed to obtain a density versus rcut/dcut curve. The resulting curve is shown in Fig. 5b, which represents an average over DG-forming B6A4Bx amphiphiles with x = 7 to 12; more detailed curves for each individual DG-forming amphiphile are provided in Fig. S9. Because the density is calculated as an average over all points within each spherical shell, the number of points in the shell does not affect the result. Moreover, as most oxygen atoms are located in the gyroid channels (as demonstrated in Fig. 3a), increasing rcut results in the shell comprising progressively more points from the struts, with almost no contribution from the gyroid matrix. Fig. 5b shows a pronounced decrease in density, confirming that the nodes indeed exhibit a higher concentration of oxygen atoms and, consequently, require a greater number of hydrophilic A4 segments for assembly.
3.3 Clustering analysis and order–disorder phase transition
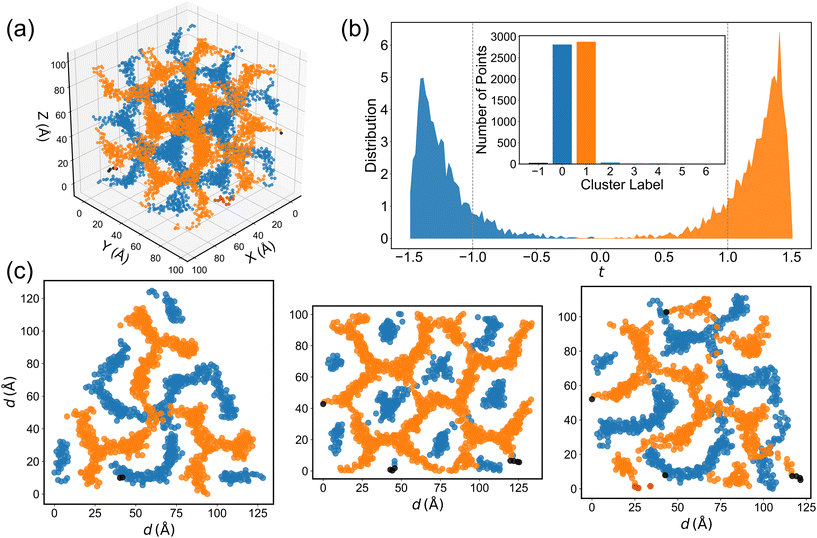
One of the most prominent features of the DG structure is the presence of two interwoven, segregated gyroid channels with opposite chirality.83 Although many visualization methods—such as 2D/3D TEM or software tools like VMD—are available in experiments and simulations, it remains challenging to automatically differentiate the two channels on a microscopic level. To more definitively identify the existence of two segregated clusters within the simulation box, density-based spatial clustering of applications with noise (DBSCAN)84 is applied to the oxygen atom coordinates extracted from the DG phase simulations. In DBSCAN, a point is classified as a core point if the number of neighboring points within a radius (ε) is greater than or equal to a specified minimum (min_samples). To account for periodic boundary conditions, the oxygen atom coordinates are first expanded by replicating the simulation box in all spatial directions. DBSCAN is then executed with ε = 5.0 Å and min_samples = 5, such that regions of high oxygen density are automatically identified while points that do not meet the criteria are classified as noise. We find that box replication is essential to avoid oxygen atoms near the faces, edges, and corners being misclassified as belonging to separate clusters (see Fig. S10). While there is less sensitivity to the choice of ε, using a value of 4.0 Å (i.e., less than the position of the second minimum in the O–O RDF, see Fig. 4) leads to significantly more noise in the DBSCAN analysis and using a very large value would lead to only a single cluster being detected.The DBSCAN analysis for the B6A4B8 system indicates two main clusters that correspond to the two interwoven gyroid channels (see Fig. 6a). To further verify that the two major clusters correspond to gyroid channels with opposite chirality, a level surface analysis is performed. Fig. 6b clearly shows that the oxygen atom coordinates in the orange and blue clusters from Fig. 6a are predominantly located in regions where t > 1 (the right-handed channel) and t < −1 (the left-handed channel), respectively. This indicates that DBSCAN successfully identifies two segregated clusters, with almost no distribution in the region |t| < 1 corresponding to the gyroid matrix, which is mostly occupied by CH2/CH3 groups. Moreover, statistical analysis of the number of points in the DBSCAN clusters reveals that the two major clusters (orange and blue) contain approximately equal numbers of points and significantly more than any other cluster or the noise. Moreover, slices taken along the (111), (110), and (211) directions of the DBSCAN clustering results demonstrate that the internal DG structure is well preserved (Fig. 6c).
In the workflow exploring possible equilibrium morphologies for B6A4Bx amphiphiles, a frequently observed order–disorder phase transition—from the initial structure to a disordered (DIS) state—is of particular interest, especially the transition from the DG structure to a DIS structure. Based on the workflow, the preferred periodicity of the amphiphiles is first determined by MD simulations via the primary peak position q* in the static structure factor. This enables the system size of the different morphologies to be approximately commensurate with the preferred lattice periodicity and, as a result, the primary peak in the structure factor does not shift significantly during the order–disorder transition (see Fig. S6). However, snapshots of the taken after the transition occurred reveal the prevalence of 3-fold connectors in the bicontinuous DIS structures (see Fig. S3–S5).
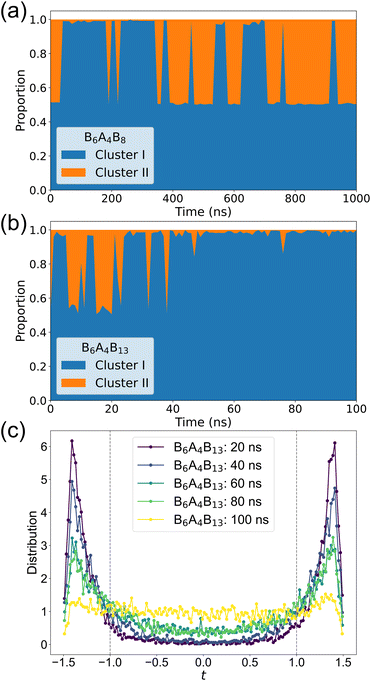
Consequently, DBSCAN is employed to investigate whether multiple locally segregated clusters can form within these DIS structures. Fig. 7a displays the DBSCAN analysis results for the B6A4B8 system, which was initially guided into a DG structure, over a 1 μs MD simulation trajectory with clustering analysis performed every 10 ns. It is observed that, along the simulation trajectory, the initially guided nearly 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 segregation of the two gyroid channels is quickly lost but reappears near 350 ns. This suggests that, at the microscopic scale, a “bridge” forms between the two initially segregated gyroid channels, leading to them merging into one cluster and preventing DBSCAN from clearly differentiating the two channels. As the simulation progresses beyond 350 ns the system re-stabilizes into an arrangement containing two clusters in an almost 1
1 segregation of the two gyroid channels is quickly lost but reappears near 350 ns. This suggests that, at the microscopic scale, a “bridge” forms between the two initially segregated gyroid channels, leading to them merging into one cluster and preventing DBSCAN from clearly differentiating the two channels. As the simulation progresses beyond 350 ns the system re-stabilizes into an arrangement containing two clusters in an almost 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio, but transient occurrences of a dominant cluster caused by bridging defects are still observed. The frequency of these defects appears to decreases over time, with considerably fewer defect-containing snapshots in the 700–1000 ns interval compared to earlier intervals.
1 ratio, but transient occurrences of a dominant cluster caused by bridging defects are still observed. The frequency of these defects appears to decreases over time, with considerably fewer defect-containing snapshots in the 700–1000 ns interval compared to earlier intervals.
For the B6A4B13 amphiphile, which transitions to a DIS structure, the situation is markedly different. Fig. 7b shows the evolution of the proportions of the two largest clusters from the DBSCAN analysis when the initial configuration is DG. Here, only the first 100 ns of the trajectory are considered with DBSCAN clustering performed at 1 ns intervals. During the first 25 ns, the guided DG structure is barely maintained; although some frames exhibit instances of two clusters with intermittent connections, the balance between the two clusters is rapidly lost as the simulation progresses. After 25 ns, a single dominant cluster persists throughout indicating a bicontinuous structure. This observation is consistent with the DIS snapshot shown in Fig. S5. Notably, the static structure factors calculated over the 20–50 ns interval still display distinct DG peaks (see Fig. S6), implying that despite the local absence of two segregated gyroid channels, the spatial arrangement of B6A4B13 amphiphiles retains some periodic characteristics of the DG structure. This observation is corroborated by the level surface analysis (see Fig. 7c), which shows that, at 20 ns, most oxygen atom coordinates remain in regions with |t| > 1 (indicative of the gyroid channels). Although the two peaks corresponding to the regions t > 1 and t < −1 gradually diminish over time, the system still exhibits certain DG features even at 80 ns, with the A4 segments predominantly located within the gyroid channels.
These results suggest that the level surface analysis lags behind both the DBSCAN clustering and the static structure factor analysis. During the DG–DIS transition, defects such as inter-channel “bridges” form first, resulting in the merging of the two gyroid channels at a local scale (as detected by DBSCAN). Subsequently, the DG periodicity is gradually disrupted at larger scales, and finally, the initially guided A4 segments begin to diffuse into other regions of the simulation box. These phenomena are captured sequentially by DBSCAN clustering, static structure factor calculations, and level surface analysis, respectively.
3.4 Mean curvature analysis
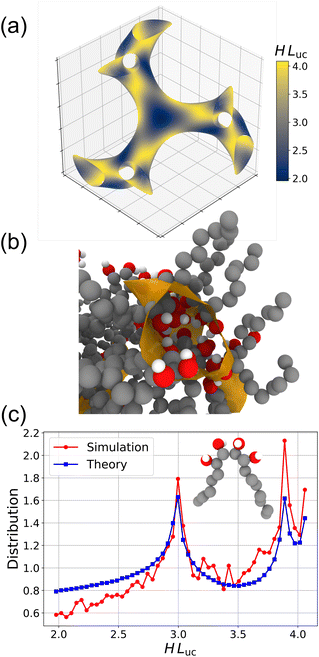
The mean curvature H is an effective approach to explore the dividing surfaces within a DG structure.36,39Fig. 8a presents a three-dimensional visualization of the gyroid level surface, obtained by solving the implicit gyroid level surface equation with the parameter t = 1.1. The iso-surface is extracted via the marching cubes algorithm.85 The average mean curvature at each vertex is computed by evaluating the gyroid level surface equation (see eqn (1)) using analytically derived first and second order derivatives, and the mean curvature values, H, are rendered dimensionless by multiplying them by the unit cell parameter, Luc. As shown in Fig. 8a, the complex topology of the gyroid structure is clearly depicted, highlighting both its geometric features and the local variations in HLuc. Notably, the surface is relatively flat near the gyroid nodes—resulting in smaller values of H Luc—and more curved along the struts, where HLuc is correspondingly larger.At the microscopic level, the orientation of DG-forming B6A4Bx amphiphiles on regions of differing curvature on the gyroid surface can be investigated. Fig. 8b shows a zoomed-in snapshot of a gyroid node from a DG structure formed by B6A4B7. Since the dividing surface separates the hydrophilic A4 segments from the hydrophobic Bx segments, the A4 segments are expected to reside primarily within the dividing surface. As observed in the snapshot, the A4 segments—connected at both ends to hydrophobic Bx segments—do not penetrate deeply into the exterior of the dividing surface. Accordingly, the centers of the second and third hydroxyl carbons are selected as representative markers for the positions of the amphiphiles near the dividing surface.
Fig. 8c presents the distribution of the dimensionless mean curvature HLuc obtained from both simulation and theoretical analyses. For the simulation data, the coordinates of the centers of the second and third hydroxyl carbons are projected onto the implicit gyroid level surface with |t| = 1.1 using a Newton–Raphson scheme. The corresponding curvatures at these projection points are then computed, and the results are averaged over DG-forming B6A4Bx amphiphiles with x = 7 to 12 to reduce the noise resulting from thermal fluctuations prevalent for the short amphiphiles and the relatively small system size. Individual curves for each DG-forming amphiphile are shown in Fig. S11. For the theoretical curve, the marching cubes algorithm is applied to extract the corresponding iso-surfaces, and the mean curvature at the mesh vertices is computed based on local differential geometry. An area-weighted average is subsequently determined for each triangle of the extracted surface, and the resulting data are normalized to yield a distribution. Using different |t| values to define the surface does not qualitatively alter the shape of the curvature distributions but changes the HLuc range and the relative heights for the two peaks (see Fig. S12). To further reduce noise in the simulation data, only the magnitude of the mean curvature is considered, disregarding its sign. It should be noted that, according to the conventional definition of the surface normal, the two channels should exhibit mean curvatures of opposite signs (as shown in Fig. S13). The comparison of these curves provides insight into the extent to which the local curvature statistics obtained from the simulation data generally conform to those predicted by the idealized gyroid level surface. Both curves display two distinct peaks found at the same two HLuc values, and the overall shapes of the two curves match fairly well. However, the distribution obtained from the MD simulations exhibits an increasing trend for the baseline leading to a lower fraction of amphiphile molecules found in regions of lower values of HLuc and a higher peak at HLuc ≈ 3.9 than at 3.0. This discrepancy arises because the stiff B6A4Bx amphiphiles, which preferentially fold into V-shaped conformations,11 respond to packing frustration caused by different regions of the gyroid dividing surface possessing different curvature. Near the gyroid nodes, the idealized surface is relatively flat with low curvature; however, the excluded volume of the hydrophobic tails cannot be geometrically accommodated within the limited space of the matrix. In contrast, in the highly curved strut regions, the shape of the amphiphiles can be more easily accommodated.
4 Conclusion
MD simulations are employed to investigate the phase behavior of B6A4Bx triblock amphiphilic oligomers, which consist of polar, sugar-based (A4) segments and non-polar, hydrocarbon (B6 and Bx) segments. B6A4B6 amphiphiles self-assemble into HPL, while those with x = 7 to 12 form DG structures. In contrast, B6A4B13 and B6A4B14 do not attain stable ordered states, instead forming DIS phases. DG structures are stabilized across a range of A4 volume fractions (fA) from 0.22 to 0.28, with domain spacings varying from 1.92 nm to 2.26 nm. A Gaussian KDE analysis of the oxygen atom coordinates within DG morphologies reveals that the oxygen density is locally enhanced at the gyroid nodes and reduced along the struts. Furthermore, DBSCAN-based clustering accurately distinguishes the two gyroid channels of opposite chirality, which facilitates the detection of a bridging phenomenon during the DG-to-DIS transition. Mean curvature analysis demonstrates that regions with higher curvature are able to accommodate a higher fraction of amphiphile molecules compared to the surface fraction of idealized gyroid structures. This work demonstrates the value of asymmetric molecular design in stabilizing DG structures and provides insight into the microscopic packing behavior of amphiphiles.Author contributions
Daoyuan Li – conceptualization; data curation; investigation; methodology; software; visualization; writing – original draft. Mahesh Mahanthappa – conceptualization; supervision; writing – review & editing. Timothy Lodge – conceptualization; supervision; writing – review & editing. Ilja Siepmann – conceptualization; methodology; supervision; writing – review & editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI).Supplementary information: force field details, calculation of structure factors, tables with force field parameters and additional figures. See DOI: https://doi.org/10.1039/d6lf00030d.
Acknowledgements
This work was supported by the National Science Foundation through the University of Minnesota MRSEC under Award DMR-2011401. Computer resources were provided by this NSF award and by the Minnesota Supercomputing Institute.References
- R. M. Dorin, H. Sai and U. Wiesner, Chem. Mater., 2014, 26, 339–347 CrossRef CAS.
- L. Xiang, Q. Li, C. Li, Q. Yang, F. Xu and Y. Mai, Adv. Mater., 2023, 35, 2207684 CrossRef CAS PubMed.
- D. Zhao, J. Feng, Q. Huo, N. Melosh, G. H. Fredrickson, B. F. Chmelka and G. D. Stucky, Science, 1998, 279, 548–552 CrossRef CAS PubMed.
- R. K. W. Spencer and M. W. Matsen, Macromolecules, 2017, 50, 1681–1687 CrossRef CAS.
- M. Steube, T. Johann, R. D. Barent, A. H. Mueller and H. Frey, Prog. Polym. Sci., 2022, 124, 101488 CrossRef CAS.
- H. He, K. Rahimi, M. Zhong, A. Mourran, D. R. Luebke, H. B. Nulwala, M. Möller and K. Matyjaszewski, Nat. Commun., 2017, 8, 14057 CrossRef PubMed.
- S. Ha, Y. La and K. T. Kim, Acc. Chem. Res., 2020, 53, 620–631 CrossRef CAS PubMed.
- Q. P. Chen, L. Barreda, L. E. Oquendo, M. A. Hillmyer, T. P. Lodge and J. I. Siepmann, ACS Nano, 2018, 12, 4351–4361 CrossRef CAS PubMed.
- L. Barreda, Z. Shen, Q. P. Chen, T. P. Lodge, J. I. Siepmann and M. A. Hillmyer, Nano Lett., 2019, 19, 4458–4462 CrossRef CAS PubMed.
- Z. Shen, J. L. Chen, V. Vernadskaia, S. P. Ertem, M. K. Mahanthappa, M. A. Hillmyer, T. M. Reineke, T. P. Lodge and J. I. Siepmann, J. Am. Chem. Soc., 2020, 142, 9352–9362 CrossRef CAS PubMed.
- Z. Shen, K. Luo, S. J. Park, D. Li, M. K. Mahanthappa, F. S. Bates, K. D. Dorfman, T. P. Lodge and J. I. Siepmann, JACS Au, 2022, 2, 1405–1416 CrossRef CAS PubMed.
- D. Li, Z. Shen, P. Chen, M. K. Mahanthappa, K. D. Dorfman, T. P. Lodge and J. I. Siepmann, JACS Au, 2025, 5, 3387–3398 CrossRef PubMed.
- C. Zheng, K. Luo, S. Das, P. Chen, Z. Shen, J. Jaye, D. Li, M. A. Calabrese, T. M. Reineke and M. K. Mahanthappa, et al. , J. Phys. Chem. B, 2025, 129, 8231–8243 CrossRef CAS.
- J. Zhang, X. Yu, P. Yang, J. Peng, C. Luo, W. Huang and Y. Han, Macromol. Rapid Commun., 2010, 31, 591–608 CrossRef CAS PubMed.
- S. Das, C. Zheng, T. P. Lodge, J. I. Siepmann, M. K. Mahanthappa, M. A. Calabrese and T. M. Reineke, Biomacromolecules, 2024, 25, 1291–1302 CrossRef CAS PubMed.
- S. P. Nunes, Macromolecules, 2016, 49, 2905–2916 CrossRef CAS.
- L. Qiao, D. A. Vega and F. Schmid, Macromolecules, 2024, 57, 4629–4634 CrossRef CAS PubMed.
- S. Maekawa, T. Seshimo, T. Dazai, K. Sato, K. Hatakeyama-Sato, Y. Nabae and T. Hayakawa, Nat. Commun., 2024, 15, 5671 CrossRef CAS PubMed.
- A. C. Finnefrock, R. Ulrich, G. E. S. Toombes, S. M. Gruner and U. Wiesner, J. Am. Chem. Soc., 2003, 125, 13084–13093 CrossRef CAS PubMed.
- S. Cui, E. A. Murphy, S. Santra, F. S. Bates and T. P. Lodge, ACS Nano, 2025, 19, 1211–1221 CrossRef CAS PubMed.
- V. Abetz and P. F. W. Simon, Advances in Polymer Science, in Block Copolymers, ed. V. Abetz, Springer, 2005, vol. 189, pp. 125–212 Search PubMed.
- S. Cui, B. Zhang, L. Shen, F. S. Bates and T. P. Lodge, J. Am. Chem. Soc., 2022, 144, 21719–21727 CrossRef CAS PubMed.
- L. Han and S. Che, Adv. Mater., 2018, 30, 1705708 CrossRef.
- H. Uehara, T. Yoshida, M. Kakiage, T. Yamanobe, T. Komoto, K. Nomura, K. Nakajima and M. Matsuda, Macromolecules, 2006, 39, 3971–3974 CrossRef CAS.
- D. L. Gin, J. E. Bara, R. D. Noble and B. J. Elliott, Macromol. Rapid Commun., 2008, 29, 367–389 CrossRef CAS.
- L. Li, L. Schulte, L. D. Clausen, K. M. Hansen, G. E. Jonsson and S. Ndoni, ACS Nano, 2011, 5, 7754–7766 CrossRef CAS PubMed.
- R. L. Kerr, S. A. Miller, R. K. Shoemaker, B. J. Elliott and D. L. Gin, J. Am. Chem. Soc., 2009, 131, 15972–15973 CrossRef CAS PubMed.
- E. J. W. Crossland, M. Kamperman, M. Nedelcu, C. Ducati, U. Wiesner, D.-M. Smilgies, G. E. S. Toombes, M. A. Hillmyer, S. Ludwigs and U. Steiner, et al. , Nano Lett., 2009, 9, 2807–2812 CrossRef CAS PubMed.
- E. J. W. Crossland, M. Nedelcu, C. Ducati, S. Ludwigs, M. A. Hillmyer, U. Steiner and H. J. Snaith, Nano Lett., 2009, 9, 2813–2819 CrossRef CAS PubMed.
- M. C. Orilall and U. Wiesner, Chem. Soc. Rev., 2011, 40, 520–535 RSC.
- V. Saranathan, C. O. Osuji, S. G. J. Mochrie, H. Noh, S. Narayanan, A. Sandy, E. R. Dufresne and R. O. Prum, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 11676–11681 CrossRef CAS.
- B. D. Wilts, K. Michielsen, H. De Raedt and D. G. Stavenga, J. R. Soc., Interface, 2012, 9, 1609–1614 CrossRef PubMed.
- M. Maldovan, A. M. Urbas, N. Yufa, W. C. Carter and E. L. Thomas, Phys. Rev. B: Condens. Matter Mater. Phys., 2002, 65, 165123 CrossRef.
- M. Stefik, S. Guldin, S. Vignolini, U. Wiesner and U. Steiner, Chem. Soc. Rev., 2015, 44, 5076–5091 RSC.
- J. A. Dolan, B. D. Wilts, S. Vignolini, J. J. Baumberg, U. Steiner and T. D. Wilkinson, Adv. Opt. Mater., 2015, 3, 12–32 CrossRef CAS.
- M. W. Matsen and F. S. Bates, Macromolecules, 1996, 29, 7641–7644 CrossRef CAS.
- S. Hyde, Z. Blum, T. Landh, S. Lidin, B. Ninham, S. Andersson and K. Larsson, The Language of Shape: The Role of Curvature in Condensed Matter: Physics, Chemistry and Biology, Elsevier, 1996 Search PubMed.
- F. S. Bates and G. H. Fredrickson, Phys. Today, 1999, 52, 32–38 CrossRef CAS.
- M. W. Matsen, J. Phys.: Condens. Matter, 2001, 14, R21 CrossRef.
- S. J. Park, G. K. Cheong, F. S. Bates and K. D. Dorfman, Macromolecules, 2021, 54, 9063–9070 CrossRef CAS.
- K. Kawamoto, M. Zhong, K. R. Gadelrab, L.-C. Cheng, C. A. Ross, A. Alexander-Katz and J. A. Johnson, J. Am. Chem. Soc., 2016, 138, 11501–11504 CrossRef CAS PubMed.
- Y. Rokhlenko, K. Kawamoto, J. A. Johnson and C. O. Osuji, Macromolecules, 2018, 51, 3680–3690 CrossRef CAS.
- L. Jiang, D. Nykypanchuk, V. J. Pastore and J. Rzayev, Macromolecules, 2019, 52, 8217–8226 CrossRef CAS.
- S. Poppe, X. Cheng, C. Chen, X. Zeng, R.-b. Zhang, F. Liu, G. Ungar and C. Tschierske, J. Am. Chem. Soc., 2020, 142, 3296–3300 CrossRef CAS PubMed.
- S. Poppe, C. Chen, F. Liu and C. Tschierske, Chem. Commun., 2018, 54, 11196–11199 RSC.
- M. Poppe, C. Chen, H. Ebert, S. Poppe, M. Prehm, C. Kerzig, F. Liu and C. Tschierske, Soft Matter, 2017, 13, 4381–4392 RSC.
- C. Sinturel, F. S. Bates and M. A. Hillmyer, ACS Macro Lett., 2015, 4, 1044–1050 CrossRef CAS.
- S. R. Nowak, K. K. Lachmayr, K. G. Yager and L. R. Sita, Angew. Chem., Int. Ed., 2021, 60, 8710–8716 CrossRef CAS PubMed.
- C. Chen, R. Kieffer, H. Ebert, M. Prehm, R.-b. Zhang, X. Zeng, F. Liu, G. Ungar and C. Tschierske, Angew. Chem., Int. Ed., 2020, 59, 2725–2729 CrossRef CAS PubMed.
- X. Zeng, M. Prehm, G. Ungar, C. Tschierske and F. Liu, Angew. Chem., Int. Ed., 2016, 55, 8329–8332 Search PubMed.
- X. Zeng, S. Poppe, A. Lehmann, M. Prehm, C. Chen, F. Liu, H. Lu, G. Ungar and C. Tschierske, Angew. Chem., Int. Ed., 2019, 58, 7375–7378 CrossRef CAS PubMed.
- F. Liu, M. Prehm, X. Zeng, C. Tschierske and G. Ungar, J. Am. Chem. Soc., 2014, 136, 6846–6849 CrossRef CAS.
- M. W. Matsen, J. Chem. Phys., 2000, 113, 5539–5544 CrossRef CAS.
- I. Mattson, J. Majoinen, M. Lahtinen, T. Sandberg, A. Fogde, T. Saloranta-Simell, O. J. Rojas, O. Ikkala and R. Leino, Soft Matter, 2023, 19, 8360–8377 RSC.
- M. G. Martin and J. I. Siepmann, J. Phys. Chem. B, 1998, 102, 2569–2577 CrossRef CAS.
- M. G. Martin and J. I. Siepmann, J. Phys. Chem. B, 1999, 103, 4508–4517 CrossRef CAS.
- B. Chen, J. J. Potoff and J. I. Siepmann, J. Phys. Chem. B, 2001, 105, 3093–3104 CrossRef CAS.
- J. M. Stubbs, J. J. Potoff and J. I. Siepmann, J. Phys. Chem. B, 2004, 108, 17596–17605 CrossRef CAS.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. Berendsen, J. Comput. Chem., 2005, 26, 1701–1718 CrossRef CAS PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1, 19–25 CrossRef.
- S. Nosé, J. Chem. Phys., 1984, 81, 511–519 CrossRef.
- W. G. Hoover, Phys. Rev. A: At., Mol., Opt. Phys., 1985, 31, 1695–1697 CrossRef PubMed.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, J. Chem. Phys., 1995, 103, 8577–8593 CrossRef CAS.
- B. Hess, H. Bekker, H. J. Berendsen and J. G. Fraaije, J. Comput. Chem., 1997, 18, 1463–1472 CrossRef CAS.
- A. Stukowski, Modell. Simul. Mater. Sci. Eng., 2009, 18, 015012 CrossRef.
- A. Stukowski, JOM, 2014, 66, 399–407 CrossRef CAS.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graphics, 1996, 14, 33–38 CrossRef CAS PubMed.
- A. Arora, D. C. Morse, F. S. Bates and K. D. Dorfman, Soft Matter, 2015, 11, 4862–4867 RSC.
- J. Glaser, J. Qin, P. Medapuram and D. C. Morse, Macromolecules, 2014, 47, 851–869 CrossRef CAS.
- T. Dotera, Phys. Rev. Lett., 2002, 89, 205502 CrossRef PubMed.
- F. J. Martinez-Veracoechea and F. A. Escobedo, J. Chem. Phys., 2006, 125, 104907 CrossRef PubMed.
- P. W. Schönhöfer, L. J. Ellison, M. Marechal, D. J. Cleaver and G. E. Schröder-Turk, Interface Focus, 2017, 7, 20160161 CrossRef PubMed.
- T. H. Epps, E. W. Cochran, T. S. Bailey, R. S. Waletzko, C. M. Hardy and F. S. Bates, Macromolecules, 2004, 37, 8325–8341 CrossRef CAS.
- S. Andersson, S. Hyde, K. Larsson and S. Lidin, Chem. Rev., 1988, 88, 221–242 CrossRef CAS.
- P. J. Gandy, S. Bardhan, A. L. Mackay and J. Klinowski, Chem. Phys. Lett., 2001, 336, 187–195 CrossRef CAS.
- H. Von Schnering and R. Nesper, Z. Phys. B:Condens. Matter, 1991, 83, 407–412 CrossRef.
- M. Wohlgemuth, N. Yufa, J. Hoffman and E. L. Thomas, Macromolecules, 2001, 34, 6083–6089 CrossRef CAS.
- O. Al-Ketan and R. K. Abu Al-Rub, Adv. Eng. Mater., 2019, 21, 1900524 CrossRef.
- J. H. Moon, Y. Xu, Y. Dan, S.-M. Yang, A. T. Johnson and S. Yang, Adv. Mater., 2007, 19, 1510–1514 CrossRef CAS.
- E. Parzen, Ann. Math. Stat., 1962, 33, 1065–1076 CrossRef.
- R. A. Davis, K.-S. Lii and D. N. Politis, Selected Works of Murray Rosenblatt, Springer, 2011, pp. 95–100 Search PubMed.
- P. J. F. Gandy and J. Klinowski, Chem. Phys. Lett., 2000, 321, 363–371 CrossRef CAS.
- M. Ester, H.-P. Kriegel, J. Sander and X. Xuet al., KDD, 1996, pp. 226–231.
- W. E. Lorensen and H. E. Cline, Seminal Graphics: Pioneering Efforts That Shaped the Field, Morgan Kaufmann, 1998, pp. 347–353 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |