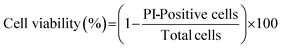Open Access Article
Open Access ArticleA customizable, low-cost 3D-printed device for live cell confinement imaging
Hunter
Richman
a,
Jin
Ou†
a,
Manpreet
Khera
a and
Yan
Yu
 *b
*b
aDepartment of Chemistry, Indiana University, Bloomington, IN 47405, USA
bDepartment of Chemistry, Department of Biomedical Engineering, Washington University in St. Louis, St. Louis, MO 63130, USA. E-mail: yuy1@wustl.edu
First published on 17th March 2026
Abstract
Physical confinement profoundly shapes cell functions from migration to cancer invasiveness. Although multiple in vitro confinement platforms have been designed to replicate physical constraints while enabling simultaneous live-cell imaging, broad adoption across the biology community has been limited because most available platforms require specialized fabrication, offer limited design flexibility, or are prohibitively costly. Here, we introduce an inexpensive, fully customizable confinement platform fabricated using standard 3D printers and readily available materials. The confinement device uses a PDMS pillar to uniformly compress two coverslips against polystyrene spacer beads to define precise confinement heights of 3, 7, or 12 μm. We show that this platform reliably confines both adherent and suspension cells, generates graded morphological changes, and maintains high cell viability for at least 24 hours under live-cell imaging conditions. By combining tunability, reproducibility, and broad accessibility, this device provides a practical alternative to existing confinement methods and enables wider exploration of how mechanical restriction modulates cell function across physiological and pathological contexts.
1. Introduction
In vivo, cells routinely experience physically restricted environments, such as tight interstitial spaces, dense extracellular matrices, and crowded multicellular tissues.1,2 The extent of this confinement varies widely across tissues, from the soft, porous tissue environment of the brain to the densely packed architecture of epithelial layers, and becomes even more pronounced during aging, inflammation, or tumor progression.3–9 These physical constraints profoundly influence cellular behaviors, including migration,10–12 stem cell differentiation,13–15 immune cell activation,16,17 and epithelial repair during wound healing.18–20 In cancer and other pathologies, physical confinement is not simply a byproduct of tissue crowding but an active mechanical stimulus that influences disease evolution.21–24 For example, studies have shown that solid tumors are characterized by extreme cell crowding and elevated interstitial pressure which can suppress healthy cell proliferation, dampen anti-tumor immune responses,25–28 and increase leader-cell phenotypes amongst cancer cells which drive metastasis.23,28,29 Collectively, these findings highlight the need for tools that enable systematic investigation into how the cellular physical microenvironment impacts cell fate and function.Despite broad interest, current in vitro approaches for studying cell functions under physical confinement remain limited as few tools allow systematic and well-controlled confinement during live-cell investigation. In addition, many of these existing tools involve technical barriers that restrict accessibility and hinder widespread adoption (summary in Table S1). Tools such as AFM provide exquisite force control but are restricted to single-cell measurements and require specialized instrumentation.26,27,30 Micropillar arrays, microfluidic chambers, and photolithography-based devices offer controlled confinement at the population level,12,23,31–34 but rely on fabrication facilities and expertise unavailable in many biology laboratories. Microporous scaffolds and crosslinked alginate systems can impose defined pore sizes16,19,22 and even allow tunable compression,7,22 but involve synthetic steps that further limit accessibility. Weighted agarose pads provide a simpler alternative.28 However, their passive loading produces highly non-uniform pressures that depend on gel mass, gel uniformity, and cell morphology, resulting in variable and difficult-to-standardize confinement. Consequently, there remains a critical need for devices that deliver precise, reproducible confinement while being low-cost, easy to assemble, and compatible with standard laboratory workflows.
Here, we present a 3D-printed confinement chamber that directly overcomes these longstanding impediments by providing a simple, customizable, and reproducible platform for applying well-defined mechanical confinement to cells. The device incorporates two coverslips which compress cells, polystyrene bead spacers that set confinement height, and a PDMS (polydimethylsiloxane) pillar to apply uniform, gentle compression. This confinement device can also be modified for brightfield imaging and reagent infusion. Crucially, the device can be fabricated using widely available benchtop 3D printers and inexpensive, commercially sourced materials, thus eliminating the need for microfabrication facilities or specialized expertise.
We demonstrate the versatility and compatibility of the confinement device by confining adherent (RAW 264.7/HEK 293T cells) and suspension cells (Jurkat T cells) to different heights for many hours. By enabling rapid, consistent, and scalable confinement assays at minimal cost, this platform makes mechanobiology experiments accessible to a much wider community. More broadly, it establishes a versatile tool for probing how physical confinement shapes cellular behavior across diverse biological contexts.
2. Materials and methods
2.1 Confinement chamber fabrication
Cell confinement device was designed using FreeCAD 0.21 (design detail in SI), exported into UltiMaker Cura 5.9, then printed on an UltiMaker S Series 3D printer extruding tough polylactic acid (PLA). Cura print settings: resolution, 0.1 mm; shell thickness, 1.2 mm; support type, tree; support placement, everywhere; and adhesion, on.2.2 PDMS pillar fabrication
PDMS elastomer base and curing agent (NC9285739, Fisher Scientific) were mixed at a 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 w/w ratio and stirred for 10 min. The mixture was then degassed under vacuum for 20 min. To mold the pillar, an acrylic tube (20 mm height × 18 mm internal diameter) was placed on top of a coverslip, filled entirely with PDMS, and capped with a 150 g metal weight. Capping both ends of the tube ensured both ends of the PDMS pillar emerged flat. The assembly was cured overnight in an oven at 80 °C. After curing, the PDMS pillar was pushed out of the tube; if stuck, the pillar edges were loosened, and ethanol was added to hasten removal. Cured pillars were sonicated in ultrapure water (18 MΩ cm) for 5 min and sanitized with UV before use.
1 w/w ratio and stirred for 10 min. The mixture was then degassed under vacuum for 20 min. To mold the pillar, an acrylic tube (20 mm height × 18 mm internal diameter) was placed on top of a coverslip, filled entirely with PDMS, and capped with a 150 g metal weight. Capping both ends of the tube ensured both ends of the PDMS pillar emerged flat. The assembly was cured overnight in an oven at 80 °C. After curing, the PDMS pillar was pushed out of the tube; if stuck, the pillar edges were loosened, and ethanol was added to hasten removal. Cured pillars were sonicated in ultrapure water (18 MΩ cm) for 5 min and sanitized with UV before use.
2.3 Spacer bead solution preparation
Polystyrene particles were diluted in ultrapure water to specific concentrations at given particle sizes (Fig. S1): 6.75 × 106 particles per mL for 3 μm polystyrene particles (A37368, ThermoFisher), 2.60 × 105 particles per mL for 7 μm polystyrene particles (ABPS-0700, Abvigen), and 7.95 × 104 particles per mL for 12 μm polystyrene particles (ABPS-1200, Abvigen). The diluted particle suspensions were sonicated for 1 min and immediately added to poly-L-lysine (PLL) coated coverslips.2.4 Confinement coverslip preparation
18 mm coverslips (CS-R18-100, AmScope) were sonicated in ethanol for 3 min and rinsed three times with ultrapure water. Cleaned coverslips were dried in an oven at 80 °C. A 0.1% PLL solution (25988-63-0, MilliporeSigma) was prepared by mixing 100 μL of stock solution with 900 μL of ultrapure water. Next, 330 μL of the diluted PLL solution was applied to each coverslip and incubated at 20 °C for 20 min. Coated coverslips were washed three times with ultrapure water and dried again at 80 °C. Finally, 350 μL of spacer solution was added to each coverslip and the coverslips were incubated at 20 °C for at least 1 h.2.5 Cell culture and seeding
RAW 264.7 GFP-actin macrophages (generous gift from Dr. Sergio Grinstein, The Hospital for Sick Children Research Institute) and GFP-FcγRII 293T cells (generous gift from Prof. Peter Cresswell, Yale University) were cultured in Dulbecco's Modified Eagle Medium (12491015, ThermoFisher) supplemented with 10% fetal bovine serum (MT35010CV, Corning), 2 mM L-glutamine (25030081, ThermoFisher), 100 units ml−1 penicillin–streptomycin (MT30001CI, Corning) at 37 °C in a humidified incubator with 5% CO2. Jurkat E6.1T cells (ATCC) were cultured in RPMI-1640 medium (30-2001, ATCC) supplemented with 10% FBS (fetal bovine serum), 1 units mL−1 penicillin–streptomycin, and 2 mM L-glutamine at 37 °C in a humidified incubator with 5% CO2. For passage, RAW 264.7 GFP-actin cells were washed with PBS (phosphate-buffered saline) then removed with a cell scraper. 293T cells were unadhered from flasks with 0.25% trypsin-EDTA (25200056, ThermoFisher). For experiments, 0.5 million RAW 264.7, 0.2 million GFP-FcγRII HEK 293T cells, or 0.4 million Jurkat E6.1T cells were seeded on piranha solution etched 30 mm coverslips (30-1313-03192, Bioptechs). Lower seeding densities were used for larger cells resulting in approximately 50% confluency at time of confinement.2.6 Confinement device assembly
Cell confinement device assembly is depicted in Fig. 1 and shown in Video S1. First, the top of the confinement device was inverted, and a PDMS pillar was inserted into the central groove of the chamber lid. Next, an 18 mm coverslip containing spacer beads was placed onto the pillar end with the beads facing outward. After that, a 30 mm coverslip with cells was secured in the imaging chamber, which was then pressed into the base of the confiner. Finally, the top and bottom halves of the device were aligned, pressed together gently to apply confinement, then secured with two pins. This assembly process takes approximately 20 s. For cleaning, the device was disassembled and coverslips were discarded. Then, each component of the device was sonicated in soapy water, rinsed thoroughly, and UV sanitized.2.7 Confocal imaging of confined cells
Cells not expressing GFP were stained with cholera toxin subunit B-Alexa Fluor 488 (C22841, ThermoFisher) for visualization of cell morphology. All cells were stained with 12.5 μg mL−1 Hoechst (H21486, ThermoFisher) in cell media for 10 min to label nuclei, followed by a single wash with cell media. Immediately after staining, cells were imaged in FluoroBrite Dulbecco's modified Eagle medium (A1896701, ThermoFisher), then confined and imaged again. Fluorescence channels were acquired as follows: GFP (Ex 488/Em 525), Hoechst (Ex 405/Em 450), and Alexa Fluor 488 (Ex 482/Em 520). Images were acquired using a Leica Stellaris 8 FALCON equipped with a 63× water immersion objective. Confocal images were analyzed using Lecia Microsystem's Las X Life Science microscope software platform.2.8 Absolute confinement measurement
30 mm coverslips were etched in piranha solution then coated with PLL per the protocol used in confinement coverslip preparation. Next, 30 mm coverslips were placed in imaging chambers and 1 mL PBS + 1 μL (0.2 μm) FluoSpheres carboxylate, yellow-green (F8811, ThermoFisher) + 50 μg CF640R (92108, Biotium) were added after brief sonication. 18 mm (confinement coverslips) were prepared as detailed in confinement coverslip preparation except 1 μL of FluoSpheres was added to 1 mL spacer bead solution. After allowing the particles to settle for 1 h, confinement devices were assembled using the FluoSphere coated coverslips and confinement coverslips.2.9 Raw 264.7 membrane and nucleus morphology measurements
RAW 264.7 GFP-actin macrophages were used to visualize cell spreading. Their nuclei were stained with 2 μg mL−1 Hoechst 33342 in culture media 30 min before confinement. Images were captured using an 100× oil-immersion objective lens and filter sets for GFP (Ex 488 nm/Em 525 25 nm) and Hoechst (Ex 402 nm/Em 450 nm) at three time points: before confinement, immediately after, and 1 h post-confinement. Cell and nuclear areas were analyzed using ImageJ via a multistep process. Cells were segmented via thresholding, converted to binary images, then had boundaries refined using the watershed algorithm. Morphological parameters, including area, major diameter, and minor diameter, were quantified using the ‘analyze particles’ function.2.10 Cell viability assay
Cell viability was assessed using propidium iodide (P4170-10MG, Sigma-Aldrich), a membrane-impermeable dye that fluoresces upon DNA intercalation. Propidium iodide cell staining indicates a loss of membrane integrity. For this assay, cells were seeded on 30 mm coverslips. After cells adhered, 5 μg mL−1 propidium iodide was added to cell media. Static images of PI-positive cells were captured using a Nikon Eclipse-Ti inverted fluorescence microscope equipped with a 40× dry objective lens and GFP/PI filter set (ex 488/ex 565). Fluorescence images were captured before confinement and 3, 6, 24, 48, and 72 hours post confinement. For each sample, three replicates were imaged, then random fields of view per replicate were analyzed. Non-viable cells (PI-positive) and total cells (GFP) were counted using ImageJ (v1.54f). Percent cell viability was calculated using the following formula.2.11 Device modification for reagent infusion
To enable reagent infusion, the confiner top was redesigned to include an off-center port accommodating an 18G needle (305196, BD). The needle tip was sanded flat to prevent damage to the chamber surfaces and ensure smooth insertion. Confining coverslips were cut from sterile Petri dish lids to maintain sterility and compatibility. Assembly proceeded as follows: the PDMS pillar was inserted into the modified confiner top, and the 18G needle was advanced through the confiner top and the PDMS pillar. The coverslip was then positioned at the base of the pillar, and the contact point of the needle was marked. A corresponding hole was drilled at this marked location in the coverslip to create a continuous fluidic channel. The hole in the polystyrene confining surface was made using a 1 mm drill bit. Upon final assembly, the needle tip was aligned flush with the end of the PDMS pillar and precisely matched with the hole in the coverslip to ensure direct access to the confined chamber. Reagent infusion was performed using an NE-1000 syringe pump to provide controlled and reproducible flow.2.12 Device modification for brightfield imaging
To enable brightfield compatibility, the confinement device top was fabricated using transparent polyethylene terephthalate glycol-modified (PETG; 32101, Bambu Lab). The component was printed on a Bambu Lab P1S series 3D printer using manufacturer-recommended settings and a 0.4 mm nozzle. Prior to printing, the build plate was coated with adhesive (Bambu Lab glue stick) to promote proper first-layer adhesion and ensure print stability.2.13 Statistical analysis
Nuclear and cellular deformation under confinement data was analyzed using a one-way ANOVA with Tukey's HSD (honestly significant difference) post hoc test. Statistical significance: ns, P > 0.05; *, P < 0.05; **, P < 0.01; ***, P < 0.001, ****, P < 0.0001.3. Results
3.1 Device design and operation
Reproducible cell confinement can be technically challenging because the confining surface must be steadily positioned within 3–12 μm of the substrate without compromising cell viability. Some existing approaches, such as the weighted agar gel method, struggle to meet this requirement. The confinement system developed here addresses these challenges through several key design features (Fig. 1). This device is comprised of a rigid 3D-printed chamber body, a PDMS pillar to apply a uniform compressive load, and two glass coverslips with surface-deposited polystyrene spacer beads to define confinement height. The confinement coverslip was coated with (PLL) to promote the electrostatically driven adsorption of polystyrene beads at sufficiently low density that provided enough space for cell culture while maintaining confinement. These spacer beads set the distance between coverslips (Fig. 1B), while the PDMS pillar ensured even pressure across the cellular plane (Fig. 1D). The entire chamber was laterally stabilized by overlapping chamber walls and dual fastening pins to ensure cell viability was not compromised via sliding coverslips. We chose three specific confinement heights to induce distinct mechanical regimes, primarily based on the expected deformation of RAW 264.7 cells: slight membrane compression (12 μm), intermediate membrane compression (7 μm), and nuclear compression (3 μm). This selection is consistent with previous confinement studies.16,27–303.2 Absolute height and cellular confinement
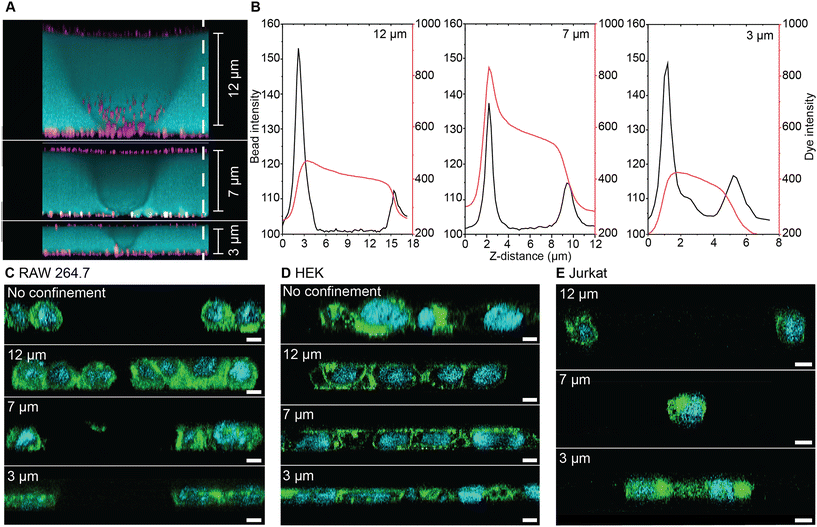
To evaluate device performance, we quantified the absolute height between the confining coverslip and the imaging chamber coverslip. Both coverslips were labeled with adsorbed 0.2 μm fluorescent beads, and CF640R in PBS was introduced to visualize the inter-coverslip gap. Reconstructed confocal Z-stacks and line-scan intensity profiles demonstrated that the spacer beads maintained precise separations of 12, 7, and 3 μm (Fig. 2A and B; more plots in Fig. S3). Next, we confined three representative cell types: adherent RAW 264.7 macrophages and HEK 293T cells, and non-adherent Jurkat T cells, and successfully achieved confinement across all cell types (Fig. 2C–E). Confocal Z-stack reconstructions verified that apical-basal cell heights of those cells precisely matched the spacer-defined confinement levels of 12, 7, and 3 μm. Cell morphology and nuclear shape were visualized by staining cells pre confinement using actin-GFP or cholera toxin subunit B-Alexa Fluor 488, with nuclei labeled by Hoechst. In all cases, Hoechst staining clearly revealed nuclear deformation under increasing compression. At 12 μm, nuclei remained largely spherical with only slight deformation, whereas increased confinement produced pronounced flattening, culminating in oblate, disk-like nuclei at 3 μm.3.3 Quantification of confined cell morphology
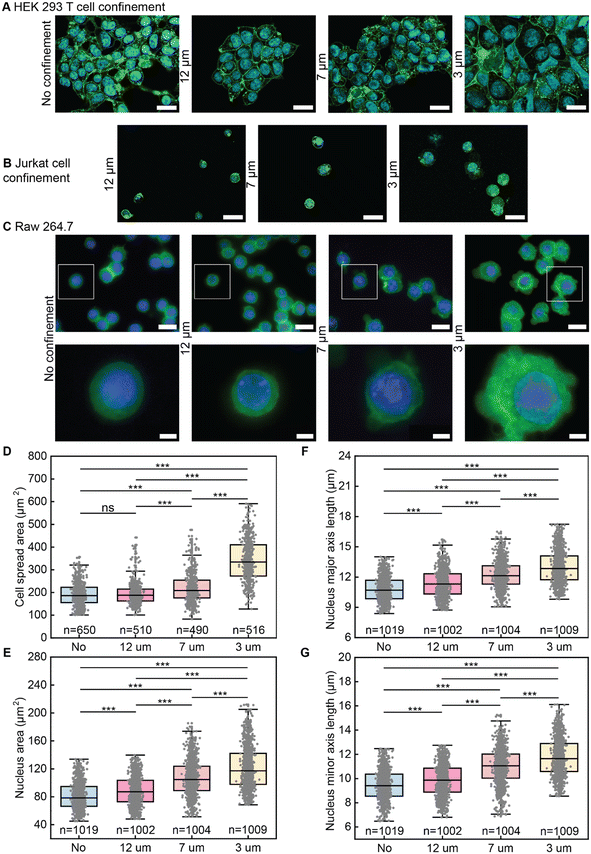
To quantitatively validate the morphological changes we observed in confined cells (Fig. 3A and B), we measured the cell spread area and nuclear dimensions (n ≈ 1000 cells) of GFP-actin RAW 264.7 cells. GFP-actin RAW 264.7 cells with Hoechst-labeled nuclei were imaged one hour after confinement (Fig. 3C). As anticipated, 12 μm confinement produced no significant change in cell spread area, consistent with minimal mechanical restriction (Fig. 3D). In contrast, 7 μm and 3 μm confinement increased spread area by 13.5% and 44.3%, respectively (Fig. 3D). Membrane blebbing emerged under 7 μm confinement and became more pronounced under 3 μm confinement (Fig. 3C). Blebbing was likely a consequence of increased hydrostatic pressure in the cytoplasm which has been previously reported to cause detachment of the plasma membrane from the cortical cytoskeleton.35,36 These blebs can either be resolved if intracellular pressure decreases or a precursor to cell rupture if pressure increases.35,37 Nuclear morphology followed similar trends. Nuclear area increased only modestly from ∼80 μm2 in unconfined cells to a slightly larger value at 12 μm, but expanded markedly at 7 μm and reached ∼120 μm2 at 3 μm confinement (Fig. 3E). The nuclear major axis length increased by 4.7%, 11.6%, and 16.7% at 12, 7, and 3 μm confinement, respectively, while the minor axis length exhibited slightly greater increases of 4.0%, 13.9%, and 19.5% (Fig. 3F and G). To assess the consistency of the confinement height across the sample chamber, in addition to the data in Fig. 3D obtained from RAW 264.7 cells expressing actin-GFP, we measured cell spreading in RAW 264.7 cells expressing RelA-GFP (Fig. S5A), a reporter cell line used to monitor inflammatory NF-κB responses. The results confirmed that both cell types exhibited comparable cell areas under confinement. In addition, when cell areas were averaged from images acquired at multiple locations across the chamber and plotted (Fig. S5B), the values showed negligible variation between locations, indicating that the confinement height was uniform throughout the device. Together, these quantitative measurements confirm that the device reliably produces characteristic increases in cell spreading and nuclear deformation associated with graded mechanical confinement.3.4 Cell compatibility
Lastly, we evaluated cell compatibility, a critical attribute for any confinement device, by quantifying cell viability and proliferation. Viability was assessed using propidium iodide (PI) staining, which selectively labels cells with compromised membranes.38 RAW 264.7 macrophage cells were confined at 12 μm, 7 μm, and 3 μm for durations ranging from 0 to 72 h (Fig. 4). Cell viability was calculated as the percentage of PI-positive cells relative to the total number of GFP-labeled cells. Confinement for up to 24 h maintained high viability (>95%) across all groups, comparable to the unconfined controls, suggesting minimal impact on cellular integrity (Fig. 4B). Extended confinement (48–72 h), however, caused a progressive decline in cell viability inversely related to confinement height (Fig. 4B). After 72 h, unconfined controls showed 78% viability, similar to 74% under 12 μm confinement, which induces minimal membrane compression. In contrast, viability dropped to 67% and 55% under 7 μm and 3 μm confinement, respectively, reflecting increased mechanical stress. These results align with previous reports of reduced viability under extreme confinement.31,39 Collectively, these findings demonstrate that our 3D-printed confinement device is highly compatible with cells for up to 48 h, with longer durations leading to decreased viability due to combined effects of mechanical stress and nutrient limitations. In addition, we observed negligible cell proliferation under all three confinement conditions. This observation is consistent with literature showing that even modest confinement, with confined heights below 20 μm, can inhibit cell proliferation due to restrictions on cell rounding required for mitosis.27,31,404. Discussion
Beyond device fabrication, cell confinement, and assessment of viability and proliferation, we next expanded the platform to enable in situ reagent infusion and exchange (Fig. 5A). To incorporate an infusion port, the top confining surface was fabricated from a sterile Petri dish lid, and a vertical channel was created by punching a hole through the device top, PDMS pillar, and confining coverslip. This channel accommodated an 18G needle (tip sanded flat), allowing controlled introduction of reagents into the confined chamber without disassembly.To validate this modification, 0.2 μm fluorescent beads and 7 μm spacer beads were first adsorbed onto the imaging chamber coverslip. The infusion port was loaded with CF640R in PBS, and the device was assembled under confinement. Two-channel confocal imaging showed that, upon initiation of injection, CF640R rapidly flowed across the bead-labeled surface (Fig. 5C). Quantitative analysis revealed that fluorescence intensity began increasing at approximately 50 s and reached a plateau by ∼90 s (Fig. 5B), confirming efficient reagent exchange within the confined space. Using the same approach, we successfully stained confined cells with Hoechst (Fig. S6). Together, these results establish that reagents can be delivered reliably to confined cells, enabling time-resolved perturbation experiments.
In addition, the device can be adapted for brightfield compatibility when required. By fabricating the confinement top from transparent polyethylene terephthalate glycol-modified (PETG), we enabled brightfield imaging directly through the device (Fig. 5D).
We further systematically compared alternative methods for spacer bead immobilization, particularly in cases where PLL coating is undesirable. The first method, which was used throughout the main study, involved incubating spacer bead suspensions on PLL-coated confinement coverslips for 1 h prior to assembly (Fig. 5E, left). To eliminate PLL, we tested a second method in which spacer beads were introduced into PBS-filled imaging chambers and allowed to adsorb for 10 min before confinement (Fig. 5E, middle). A third approach relied on evaporation: spacer bead suspensions were deposited onto uncoated confinement coverslips, allowed to dry, and then assembled (Fig. 5E, right). We found that 7 and 12 μm spacer beads could be incorporated effectively using all three methods, showing comparable distributions and maintaining the intended confinement heights. In contrast, 3 μm beads aggregated substantially under the evaporation method and therefore could not be reliably incorporated using this approach. Importantly, these findings demonstrate that PLL coating is not required for spacer bead immobilization, which may be advantageous in applications where PLL could influence cellular behavior.
Our PDMS pillar confinement device had two unsuccessful precursors. We previously attempted to confine cells using a weight-based confiner and a screw-based confiner (Fig. S7). These devices were unable to induce consistent, level confinement largely due to resolution limits of affordable 3D-printers and lateral coverslip motion. Thus, we incorporated two key design elements into our final device which ensured proper function. First, uniform compression was achieved by integrating a PDMS pillar. The moderate compressibility of PDMS compensates for minor inaccuracies in 3D printing, ensuring even force distribution across the confining coverslip. Second, lateral stability was enhanced through a nesting shell design fastened with dual pins. In addition, we were able to further improve our confinement device to overcome another limitation of many confinement systems: reliance on diffusion to deliver reagents to cells. Our device was modified to allow rapid delivery of reagents directly to confined cells by incorporating an injection channel through the PDMS pillar. Finally, we were able to achieve brightfield imaging by incorporating a transparent top. Collectively, these design refinements resolve key challenges encountered in earlier prototypes and provide a versatile, accessible, and reproducible platform for studying cellular responses under defined mechanical constraints.
5. Conclusions
Here we present a microscopy-compatible, 3D-printed cell confinement device that provides a robust, tunable, and accessible platform for quantitatively studying cellular responses to mechanical confinement. Our device was able to probe a range of cellular confinement levels from membrane perturbation to nuclear deformation. In addition, cell viability was preserved under short-term confinement (up to 48 h) at all tested heights (3–12 μm) which allows for characterization of both short- and long-term cellular responses to confinement. Prolonged confinement (48–72 h) at 7 and 3 μm, reduced viability and inhibited proliferation, highlighting the impact of sustained mechanical stress on cellular homeostasis. Compared with existing confinement platforms, this device offers advantages in affordability and adaptability, as demonstrated by its tunable confinement range, ease of operation, compatibility with reagent infusion, and support for brightfield imaging.Author contributions
H. R. conceptualized, designed, and fabricated cell confinement devices. H. R. and J. O. conducted experiments. H. R., J. O., and M. K. analyzed data. H. R. drafted the manuscript. H. R. and Y. Y. revised the manuscript.Conflicts of interest
The authors declare no competing interests.Data availability
Data related to the manuscript and confinement device CAD models are provided in the main text and supplementary information (SI) section. Additional raw data is available at Figshare: https://doi.org/10.6084/m9.figshare.30862697.Supplementary information is available. See DOI: https://doi.org/10.1039/d5lc01146a.
Acknowledgements
This work was supported by the National Institutes of Health (NIH) under award R35GM124918. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. HR gratefully acknowledges receipt of a predoctoral fellowship from the Graduate Training Program in Quantitative and Chemical Biology at Indiana University (T32 GM131994).References
- E. Mohagheghian, J. Luo, J. Chen, G. Chaudhary, J. Chen, J. Sun, R. H. Ewoldt and N. Wang, Quantifying compressive forces between living cell layers and within tissues using elastic round microgels, Nat. Commun., 2018, 9(1), 1878, DOI:10.1038/s41467-018-04245-1.
- M. E. Dolega, M. Delarue, F. Ingremeau, J. Prost, A. Delon and G. Cappello, Cell-like pressure sensors reveal increase of mechanical stress towards the core of multicellular spheroids under compression, Nat. Commun., 2017, 8, 14056, DOI:10.1038/ncomms14056.
- C. Schulze, F. Wetzel, T. Kueper, A. Malsen, G. Muhr, S. Jaspers, T. Blatt, K. P. Wittern, H. Wenck and J. A. Kas, Stiffening of human skin fibroblasts with age, Clin. Plast. Surg., 2012, 39(1), 9–20, DOI:10.1016/j.cps.2011.09.008.
- M. Segel, B. Neumann, M. F. E. Hill, I. P. Weber, C. Viscomi, C. Zhao, A. Young, C. C. Agley, A. J. Thompson and G. A. Gonzalez, et al., Niche stiffness underlies the ageing of central nervous system progenitor cells, Nature, 2019, 573(7772), 130–134, DOI:10.1038/s41586-019-1484-9.
- B. González-Bermúdez, H. Kobayashi, A. Abarca-Ortega, M. Córcoles-Lucas, M. González-Sánchez, M. De la Fuente, G. V. Guinea, M. Elices and G. R. Plaza, Aging is accompanied by T-cell stiffening and reduced interstitial migration through dysfunctional nuclear organization, Immunology, 2022, 167(4), 622–639, DOI:10.1111/imm.13559.
- B. Enyedi, M. Jelcic and P. Niethammer, The Cell Nucleus Serves as a Mechanotransducer of Tissue Damage-Induced Inflammation, Cell, 2016, 165(5), 1160–1170, DOI:10.1016/j.cell.2016.04.016 , (acccessed 2025/08/06).
- A. Desmaison, L. Guillaume, S. Triclin, P. Weiss, B. Ducommun and V. Lobjois, Impact of physical confinement on nuclei geometry and cell division dynamics in 3D spheroids, Sci. Rep., 2018, 8(1), 8785, DOI:10.1038/s41598-018-27060-6.
- M. Tan, B. Song, X. Zhao and J. Du, The role and mechanism of compressive stress in tumor, Front. Oncol., 2024, 14, 1459313, DOI:10.3389/fonc.2024.1459313.
- Z. N. Demou, Gene expression profiles in 3D tumor analogs indicate compressive strain differentially enhances metastatic potential, Ann. Biomed. Eng., 2010, 38(11), 3509–3520, DOI:10.1007/s10439-010-0097-0.
- M. Le Berre, Y. J. Liu, J. Hu, P. Maiuri, O. Benichou, R. Voituriez, Y. Chen and M. Piel, Geometric friction directs cell migration, Phys. Rev. Lett., 2013, 111(19), 198101, DOI:10.1103/PhysRevLett.111.198101.
- W. C. Hung, S. H. Chen, C. D. Paul, K. M. Stroka, Y. C. Lo, J. T. Yang and K. Konstantopoulos, Distinct signaling mechanisms regulate migration in unconfined versus confined spaces, J. Cell Biol., 2013, 202(5), 807–824, DOI:10.1083/jcb.201302132.
- E. Desvignes, A. Bouissou, A. Laborde, T. Mangeat, A. Proag, C. Vieu, C. Thibault, I. Maridonneau-Parini and R. Poincloux, Nanoscale Forces during Confined Cell Migration, Nano Lett., 2018, 18(10), 6326–6333, DOI:10.1021/acs.nanolett.8b02611.
- X. Gao, Y. Li, J. W. N. Lee, J. Zhou, V. Rangaraj, J. Marlena and A. W. Holle, Confined Migration Drives Stem Cell Differentiation, Adv. Sci., 2025, 12(21), e2415407, DOI:10.1002/advs.202415407.
- G. Park, J. A. Grey, F. Mourkioti and W. M. Han, 3D Mechanical Confinement Directs Muscle Stem Cell Fate and Function, Adv. Biol., 2025, 9(4), e2400717, DOI:10.1002/adbi.202400717.
- M. Bergert, S. Lembo, S. Sharma, L. Russo, D. Milovanovic, K. H. Gretarsson, M. Bormel, P. A. Neveu, J. A. Hackett and E. Petsalaki, et al., Cell Surface Mechanics Gate Embryonic Stem Cell Differentiation, Cell Stem Cell, 2021, 28(2), 209–216.e204, DOI:10.1016/j.stem.2020.10.017.
- Z. Z. Liu, N. Y. Xu, M. L. Wang, R. Z. Tang and X. Q. Liu, Physical confinement in alginate cryogels determines macrophage polarization to a M2 phenotype by regulating a STAT-related mRNA transcription pathway, Biomater. Sci., 2022, 10(9), 2315–2327, 10.1039/d1bm01719e.
- N. Jain and V. Vogel, Spatial confinement downsizes the inflammatory response of macrophages, Nat. Mater., 2018, 17(12), 1134–1144, DOI:10.1038/s41563-018-0190-6.
- A. Grande-Garcia, A. Echarri, J. de Rooij, N. B. Alderson, C. M. Waterman-Storer, J. M. Valdivielso and M. A. del Pozo, Caveolin-1 regulates cell polarization and directional migration through Src kinase and Rho GTPases, J. Cell Biol., 2007, 177(4), 683–694, DOI:10.1083/jcb.200701006.
- Y. Liu, A. Suarez-Arnedo, E. L. P. Caston, L. Riley, M. Schneider and T. Segura, Exploring the Role of Spatial Confinement in Immune Cell Recruitment and Regeneration of Skin Wounds, Adv. Mater., 2023, 35(49), e2304049, DOI:10.1002/adma.202304049.
- G. Zhang, Z. Zhang, G. Cao, Q. Jin, L. Xu, J. Li, Z. Liu, C. Xu, Y. Le and Y. Fu, et al., Engineered dermis loaded with confining forces promotes full-thickness wound healing by enhancing vascularisation and epithelialisation, Acta Biomater., 2023, 170, 464–478, DOI:10.1016/j.actbio.2023.08.049.
- W. C. Keith, F. Hemmati, R. S. Vaghasiya, F. Amiri and P. Mistriotis, Differential Effects of Confinement-Induced ROS Accumulation on Highly Motile Cancerous and Non-Cancerous Cells, AIChE J., 2024, 70(12), e18598, DOI:10.1002/aic.18598.
- R. Schnellmann, D. Ntekoumes, M. I. Choudhury, S. Sun, Z. Wei and S. Gerecht, Stiffening Matrix Induces Age-Mediated Microvascular Phenotype Through Increased Cell Contractility and Destabilization of Adherens Junctions, Adv. Sci., 2022, 9(22), e2201483, DOI:10.1002/advs.202201483.
- C. Rianna, M. Radmacher and S. Kumar, Direct evidence that tumor cells soften when navigating confined spaces, Mol. Biol. Cell, 2020, 31(16), 1726–1734, DOI:10.1091/mbc.E19-10-0588.
- G. Bastianello, G. R. Kidiyoor, C. Lowndes, Q. Li, R. Bonnal, J. Godwin, F. Iannelli, L. Drufuca, R. Bason and F. Orsenigo, et al., Mechanical stress during confined migration causes aberrant mitoses and c-MYC amplification, Proc. Natl. Acad. Sci. U. S. A., 2024, 121(29), e2404551121, DOI:10.1073/pnas.2404551121.
- G. Bastianello, G. R. Kidiyoor, C. Lowndes, Q. Li, R. Bonnal, J. Godwin, F. Iannelli, L. Drufuca, R. Bason and F. Orsenigo, et al., Mechanical stress during confined migration causes aberrant mitoses and c-MYC amplification, Proc. Natl. Acad. Sci. U. S. A., 2024, 121(29), e2404551121, DOI:10.1073/pnas.2404551121.
- H. K. Matthews, S. Ganguli, K. Plak, A. V. Taubenberger, Z. Win, M. Williamson, M. Piel, J. Guck and B. Baum, Oncogenic Signaling Alters Cell Shape and Mechanics to Facilitate Cell Division under Confinement, Dev. Cell, 2020, 52(5), 563–573 e563, DOI:10.1016/j.devcel.2020.01.004.
- C. J. Cattin, M. Duggelin, D. Martinez-Martin, C. Gerber, D. J. Muller and M. P. Stewart, Mechanical control of mitotic progression in single animal cells, Proc. Natl. Acad. Sci. U. S. A., 2015, 112(36), 11258–11263, DOI:10.1073/pnas.1502029112.
- J. M. Tse, G. Cheng, J. A. Tyrrell, S. A. Wilcox-Adelman, Y. Boucher, R. K. Jain and L. L. Munn, Mechanical compression drives cancer cells toward invasive phenotype, Proc. Natl. Acad. Sci. U. S. A., 2012, 109(3), 911–916, DOI:10.1073/pnas.1118910109.
- M. Pandey, Y. J. Suh, M. Kim, H. J. Davis, J. E. Segall and M. Wu, Mechanical compression regulates tumor spheroid invasion into a 3D collagen matrix, Phys. Biol., 2024, 21(3), 036003, DOI:10.1088/1478-3975/ad3ac5.
- A. J. Lomakin, C. J. Cattin, D. Cuvelier, Z. Alraies, M. Molina, G. P. F. Nader, N. Srivastava, P. J. Sáez, J. M. Garcia-Arcos and I. Y. Zhitnyak, et al., The nucleus acts as a ruler tailoring cell responses to spatial constraints, Science, 2020, 370(6514), eaba2894, DOI:10.1126/science.aba2894.
- A. Prunet, S. Lefort, H. Delanoe-Ayari, B. Laperrousaz, G. Simon, C. Barentin, S. Saci, F. Argoul, B. Guyot and J. P. Rieu, et al., A new agarose-based microsystem to investigate cell response to prolonged confinement, Lab Chip, 2020, 20(21), 4016–4030, 10.1039/d0lc00732c.
- M. T. Doolin, T. S. Ornstein and K. M. Stroka, Nuclear Deformation in Response to Mechanical Confinement is Cell Type Dependent, Cell, 2019, 8(5), 427, DOI:10.3390/cells8050427.
- M. Le Berre, J. Aubertin and M. Piel, Fine control of nuclear confinement identifies a threshold deformation leading to lamina rupture and induction of specific genes, Integr. Biol., 2012, 4(11), 1406–1414, 10.1039/c2ib20056b.
- O. M. Lancaster, M. Le Berre, A. Dimitracopoulos, D. Bonazzi, E. Zlotek-Zlotkiewicz, R. Picone, T. Duke, M. Piel and B. Baum, Mitotic rounding alters cell geometry to ensure efficient bipolar spindle formation, Dev. Cell, 2013, 25(3), 270–283, DOI:10.1016/j.devcel.2013.03.014.
- J. Y. Tinevez, U. Schulze, G. Salbreux, J. Roensch, J. F. Joanny and E. Paluch, Role of cortical tension in bleb growth, Proc. Natl. Acad. Sci. U. S. A., 2009, 106(44), 18581–18586, DOI:10.1073/pnas.0903353106.
- P. Mistriotis, E. O. Wisniewski, K. Bera, J. Keys, Y. Li, S. Tuntithavornwat, R. A. Law, N. A. Perez-Gonzalez, E. Erdogmus and Y. Zhang, et al., Confinement hinders motility by inducing RhoA-mediated nuclear influx, volume expansion, and blebbing, J. Cell Biol., 2019, 218(12), 4093–4111, DOI:10.1083/jcb.201902057.
- C. Jia, J. Shi, Y. Yao, T. Han, A. C. H. Yu and P. Qin, Plasma Membrane Blebbing Dynamics Involved in the Reversibly Perforated Cell by Ultrasound-Driven Microbubbles, Ultrasound Med. Biol., 2021, 47(3), 733–750, DOI:10.1016/j.ultrasmedbio.2020.11.029.
- C. Riccardi and I. Nicoletti, Analysis of apoptosis by propidium iodide staining and flow cytometry, Nat. Protoc., 2006, 1(3), 1458–1461, DOI:10.1038/nprot.2006.238.
- O. Y. Dudaryeva, A. Bucciarelli, G. Bovone, F. Huwyler, S. Jaydev, N. Broguiere, M. Al-Bayati, M. Lütolf and M. W. Tibbitt, 3D Confinement Regulates Cell Life and Death, Adv. Funct. Mater., 2021, 31(52), 2104098, DOI:10.1002/adfm.202104098.
- R. A. Moriarty and K. M. Stroka, Physical confinement alters sarcoma cell cycle progression and division, Cell Cycle, 2018, 17(19–20), 2360–2373, DOI:10.1080/15384101.2018.1533776.
Footnote |
| † Current address: Department of Neuroscience, University of Wisconsin-Madison, Madison, WI 53705, USA. |
| This journal is © The Royal Society of Chemistry 2026 |