Open Access Article
Open Access ArticleMachine learning-driven single-cell phenotyping in size-controlled microenvironments via parallel deterministic droplet microfluidics
Sangmin
Lee†
 abcde,
Steven
O'Donnell†
de,
Zhangli
Peng
e and
Jae-Won
Shin
abcde,
Steven
O'Donnell†
de,
Zhangli
Peng
e and
Jae-Won
Shin
 *abcde
*abcde
aDepartment of Biologic and Materials Sciences & Prosthodontics, University of Michigan, Ann Arbor, MI 48109, USA. E-mail: shinjw@umich.edu
bDepartment of Biomedical Engineering, University of Michigan, Ann Arbor, MI 48109, USA
cBiointerfaces Institute, University of Michigan, Ann Arbor, MI 48109, USA
dDepartment of Pharmacology and Regenerative Medicine, University of Illinois at Chicago, Chicago, IL 60612, USA
eDepartment of Biomedical Engineering, University of Illinois at Chicago, Chicago, IL 60607, USA
First published on 17th March 2026
Abstract
Understanding how individual cells respond to distinct physical microenvironments is critical for mechanobiology, cell therapy, and tissue engineering. Current single-cell encapsulation methods are often limited by Poisson loading and fixed droplet sizes, preventing parallel generation of multiple, size-specific microenvironments and constraining high-resolution phenotypic analyses. Here, we present a droplet microfluidic platform that enables deterministic single-cell encapsulation within microgels of multiple sizes from a single precursor stream, achieved through parallelized flow-focusing combined with cell-selective gelation. This system produces distinct microgel size regimes simultaneously, minimizing empty compartments and enabling direct, side-by-side comparisons of cellular behavior under controlled yet variable confinement. Using machine learning to analyze 3D morphological and cytoskeletal features, we reveal heterogeneous, size-dependent phenotypic responses and demonstrate that cellular phenotypes alone can predict microgel confinement across time. Together, these results establish a data-driven framework for mapping single-cell responses across engineered microenvironments and provide a scalable platform for predictive studies of mechanosensitive behavior in heterogeneous niches.
Introduction
Cells reside within complex microenvironments where variations in physical parameters, such as stiffness, geometry, viscoelasticity, and confinement, critically shape their behavior.1–5 A deeper understanding of how these physical contexts influence single-cell morphology, function, and fate is crucial for advancing both fundamental mechanobiology and therapeutic applications. Droplet-based microfluidic platforms have emerged as a powerful route for high-throughput, single-cell encapsulation within well-defined hydrogel microenvironments, enabling controlled studies of how a cell's physical niche influences its phenotype. These systems provide a more physiologically relevant 3D context for cell culture, offering tunable properties that preserve cell viability and ensure sufficient mass transport.6–9 This ability to precisely control the 3D microenvironment at a single-cell level is a major advantage over traditional bulk methods.Despite recent advances, our ability to examine how microenvironment size, a fundamental physical parameter, shapes single-cell behavior remains limited. Most droplet systems rely on Poisson-limited encapsulation, which yields a large fraction of empty or multi-cell droplets and reduces experimental efficiency.10 In addition, droplet size is typically fixed by device geometry, restricting analyses to a single condition at a time and requiring multiple devices or sequential workflows to compare size variants. Parallel droplet generators11 and droplet-splitting12 strategies have been developed to generate droplets with varying sizes simultaneously, yet these methods have rarely been integrated with deterministic single cell encapsulation. Even when platforms permit size variation, it often demands complex flow adjustments or device changes, limiting straightforward, parallel comparison of cellular responses to different confinement levels. Finally, indiscriminate gelation produces numerous empty microgels, further complicating downstream analyses.
To overcome these shortcomings, deterministic, cell-selective encapsulation strategies that leverage cell adsorption of gel crosslinkers have been developed to enrich single-cell occupancy while preventing gelation of empty compartments.13,14 Droplet-based microgel encapsulation platforms have been implemented to control local microenvironment size, revealing how spatial confinement and 3D biophysical cues influence single-cell morphology, membrane tension, and differentiation potential.15 Mesenchymal stromal cells (MSCs) singly encapsulated within soft, conformal gels have been shown to dynamically remodel their environment in vivo through cytokine-responsive collagenase secretion, resulting in fibrosis resolution.16,17 More recently, asymmetric encapsulation strategies have been developed to impose directional polarity cues within 3D single-cell niches, providing insights into cell polarization and lineage specification.18 While these studies established the importance and feasibility of engineering microenvironmental physical properties, they still lacked a means to simultaneously generate and compare the impact of multiple gel-size conditions on individual cells within a single experiment with deterministic encapsulation.
Together, these limitations define a key gap: current droplet microfluidic systems do not enable deterministic single-cell encapsulation across multiple microgel sizes generated in parallel while minimizing empty compartments. Such a capability is needed for rigorous dissection of how microenvironment size alone regulates single-cell phenotype. To address this gap, we developed a droplet microfluidic platform that combines parallel flow-focusing with cell-selective gelation to produce two distinct microgel size regimes from a single precursor stream. This deterministic design improves encapsulation efficiency, reduces resource waste, and enables direct comparisons under matched biochemical conditions. Moreover, by integrating high-content imaging with machine learning-based morphological profiling, we demonstrate how microgel size shapes cellular morphology and heterogeneity, capturing phenotypic transitions that are difficult to detect in bulk or sequential systems. In doing so, this work extends our previous contributions and establishes a scalable framework for predictive phenotyping in engineered microenvironments.
Results
Device architecture enables parallel generation of multi-sized droplets
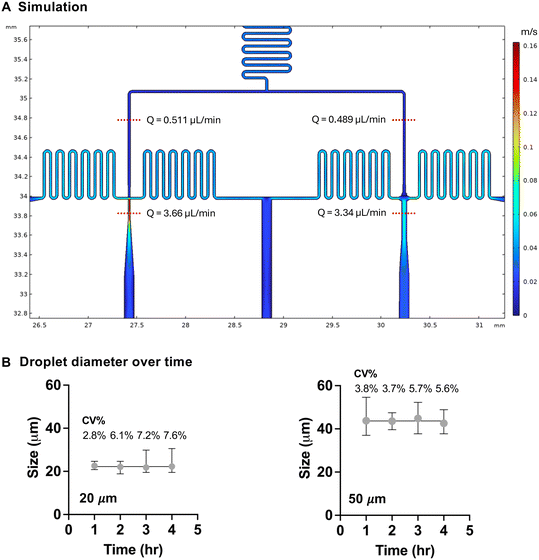
We designed a two-branch droplet microfluidic platform that integrates passive aqueous-phase splitting with selective gelation chemistry (Fig. 1A). A single aqueous precursor stream of 1% w/v alginate (240 kDa, LF200) was evenly split into two downstream channels, each ending in a flow-focusing junction upon contact with the oil phase (HFE 7500 with 1% PFPE surfactant), optimized to produce distinct droplet sizes at 1 μl min−1 aqueous and 6 μl min−1 oil flow rates (Fig. 1B and C). This setup enabled parallel generation of small (21.7 μm in diameter) and large (41.6 μm) droplets from a single input, eliminating the need for multiple devices. Device fidelity was confirmed by profilometry, which verified that SU-8 features matched design specifications (mold height 25 μm; Fig. S1A), with junction widths measured from PDMS averaging 20 μm and 48 μm. No significant channel swelling was observed under operating pressures at 15, 30, or 60 min (Fig. S1B). Finite element simulations predicted that the volumetric flow distribution was comparable between the small- and large-droplet branches, both before and after droplet formation in the microfluidic device (Fig. 2A). Droplet diameters for both size populations remained highly consistent over time, with coefficients of variation (CV) below 8% across all time points (Fig. 2B), demonstrating stable flow focusing and minimal drift in emulsification dynamics. Together, these results validate the predictive accuracy of the simulation model and confirm robust, parallel production of multi-sized droplets suitable for downstream size-controlled single-cell encapsulation applications.Multi-sized droplet and microgel dimensions scale predictably with capillary number and flow rate ratio
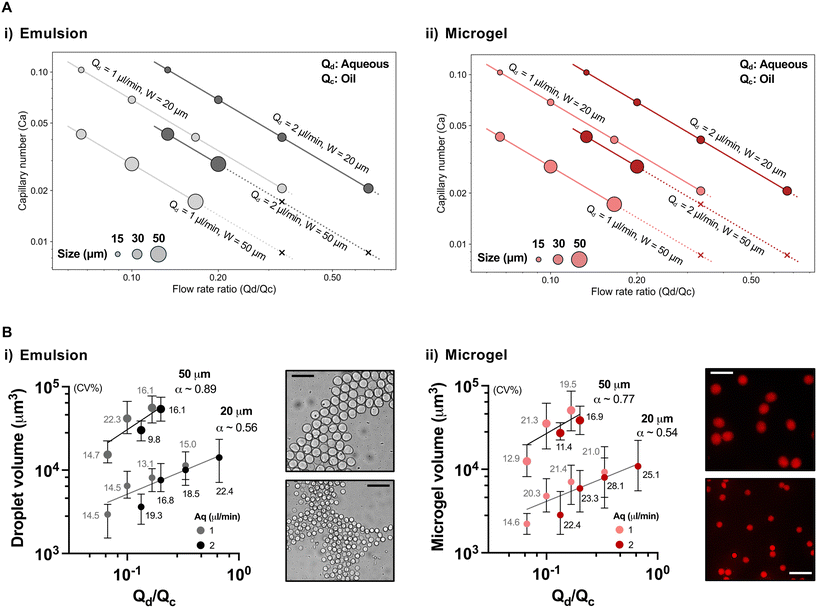
To quantitatively define the physical parameters governing size control in our parallel device, we mapped droplet formation across a range of dispersed-to-continuous phase flow rate ratios (Qd/Qc, where Qd: aqueous phase and Qc: oil phase) and corresponding capillary numbers (Ca = μcvc/γ, where μc is the viscosity of the continuous phase ∼1.24 mPa s, vc is the oil-phase velocity, and γ is the interfacial tension ∼3 mN m−1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 19). The aqueous phase consisted of 1% w/v LF200 alginate with 2 mg mL−1 CaCO3, and the oil phase was HFE 7500 supplemented with 0.03% acetic acid and 1% PFPE surfactant. Across both junctions, droplet formation operated within the squeezing-to-dripping transition regime typical of flow-focusing devices. Increasing Ca via elevated oil-phase velocity reduced droplet size, consistent with enhanced shear-mediated pinch-off. When plotted as Ca versus Qd/Qc, droplet generation remained stable across the tested range in the small channel. Although the large channel showed instability at higher Qd/Qc ratios (>0.2), droplet and microgel generation still remained robust below this threshold, supporting reliable operation under parallelized configurations (Fig. 3A). Droplets were circular under all conditions (Fig. S2), allowing volume estimates to approximate their dependence on Qd/Qc. Droplet volume scaled predictably with Qd/Qc in both small and large junctions, showing the power law exponent α ∼ 0.5 and ∼0.8, respectively, for both droplets and microgels (Fig. 3B). This sublinear scaling is consistent with trends observed in flow-focusing devices, as higher Ca tends to limit droplet growth with increasing Qd/Qc, and the finite viscosity of the aqueous alginate phase can moderate thread thinning, often yielding exponents below unity.20 Together, these results establish a quantitative hydrodynamic framework for multi-size droplet generation in parallel junctions and demonstrate that droplet and microgel dimensions can be predictably tuned through capillary number and flow-rate ratio without compromising stability.
19). The aqueous phase consisted of 1% w/v LF200 alginate with 2 mg mL−1 CaCO3, and the oil phase was HFE 7500 supplemented with 0.03% acetic acid and 1% PFPE surfactant. Across both junctions, droplet formation operated within the squeezing-to-dripping transition regime typical of flow-focusing devices. Increasing Ca via elevated oil-phase velocity reduced droplet size, consistent with enhanced shear-mediated pinch-off. When plotted as Ca versus Qd/Qc, droplet generation remained stable across the tested range in the small channel. Although the large channel showed instability at higher Qd/Qc ratios (>0.2), droplet and microgel generation still remained robust below this threshold, supporting reliable operation under parallelized configurations (Fig. 3A). Droplets were circular under all conditions (Fig. S2), allowing volume estimates to approximate their dependence on Qd/Qc. Droplet volume scaled predictably with Qd/Qc in both small and large junctions, showing the power law exponent α ∼ 0.5 and ∼0.8, respectively, for both droplets and microgels (Fig. 3B). This sublinear scaling is consistent with trends observed in flow-focusing devices, as higher Ca tends to limit droplet growth with increasing Qd/Qc, and the finite viscosity of the aqueous alginate phase can moderate thread thinning, often yielding exponents below unity.20 Together, these results establish a quantitative hydrodynamic framework for multi-size droplet generation in parallel junctions and demonstrate that droplet and microgel dimensions can be predictably tuned through capillary number and flow-rate ratio without compromising stability.
Simultaneous deterministic single-cell encapsulation in multi-sized microgels
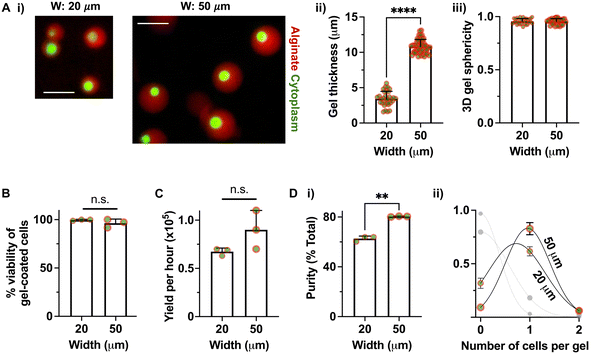
To evaluate biological performance, we assessed whether deterministic single-cell encapsulation was maintained across both droplet size regimes. A suspension of 1% w/v LF200 alginate functionalized with Arg-Gly-Asp (RGD; 60 μM) containing 6 million D1 MSCs per milliliter, pre-incubated with 2 mg mL−1 CaCO3 nanoparticles, was perfused at optimized flow rates of 1 μl min−1 (aqueous) and 6 μl min−1 (oil). The process produced conformal gel-coated cells with two distinct gel thicknesses, ∼3.4 μm (‘small’) and ∼10.7 μm (‘large’), while preserving a spherical 3D morphology (sphericity ∼0.95) (Fig. 4A). The fluorescence intensity profiles of both small and large gel coatings remained uniform across the shell thickness (Fig. S3), suggesting relatively homogeneous alginate distribution and crosslinking throughout the gel shell across both thickness regimes. Cell viability immediately after encapsulation remained high, with Calcein AM staining indicating >96% survival for both sizes (Fig. 4B). The device produced 67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 viable small gels and 90
000 viable small gels and 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 viable large gels per hour (Fig. 4C). Purity analysis showed that 60% of small gels and 80% of large gels contained single cells, substantially exceeding Poisson predictions calculated for the same cell concentration and flow rates (Fig. 4D). Cell-containing droplets averaged ∼5.5% for small and ∼37.7% for large microgels, quantified every 15 min over a 3-hour continuous run, consistent with Poisson predictions (small = 4.2%, large = 27.9%). Surface-localized crosslinking selectively coated cell-containing droplets throughout this period, resulting in higher final purity of cell-containing microgels for both size regimes (small = 60.6%, large = 82.1%) with no detectable decline in throughput or evidence of channel fouling over time (Fig. S4). Together, these results benchmark performance against a Poisson baseline and demonstrate sustained, high-purity single-cell encapsulation across multiple microgel sizes, enabled by cell-surface-localized crosslinking.
000 viable large gels per hour (Fig. 4C). Purity analysis showed that 60% of small gels and 80% of large gels contained single cells, substantially exceeding Poisson predictions calculated for the same cell concentration and flow rates (Fig. 4D). Cell-containing droplets averaged ∼5.5% for small and ∼37.7% for large microgels, quantified every 15 min over a 3-hour continuous run, consistent with Poisson predictions (small = 4.2%, large = 27.9%). Surface-localized crosslinking selectively coated cell-containing droplets throughout this period, resulting in higher final purity of cell-containing microgels for both size regimes (small = 60.6%, large = 82.1%) with no detectable decline in throughput or evidence of channel fouling over time (Fig. S4). Together, these results benchmark performance against a Poisson baseline and demonstrate sustained, high-purity single-cell encapsulation across multiple microgel sizes, enabled by cell-surface-localized crosslinking.
Machine learning-based image analysis reveals time-dependent phenotypic heterogeneity and predicts microgel confinement
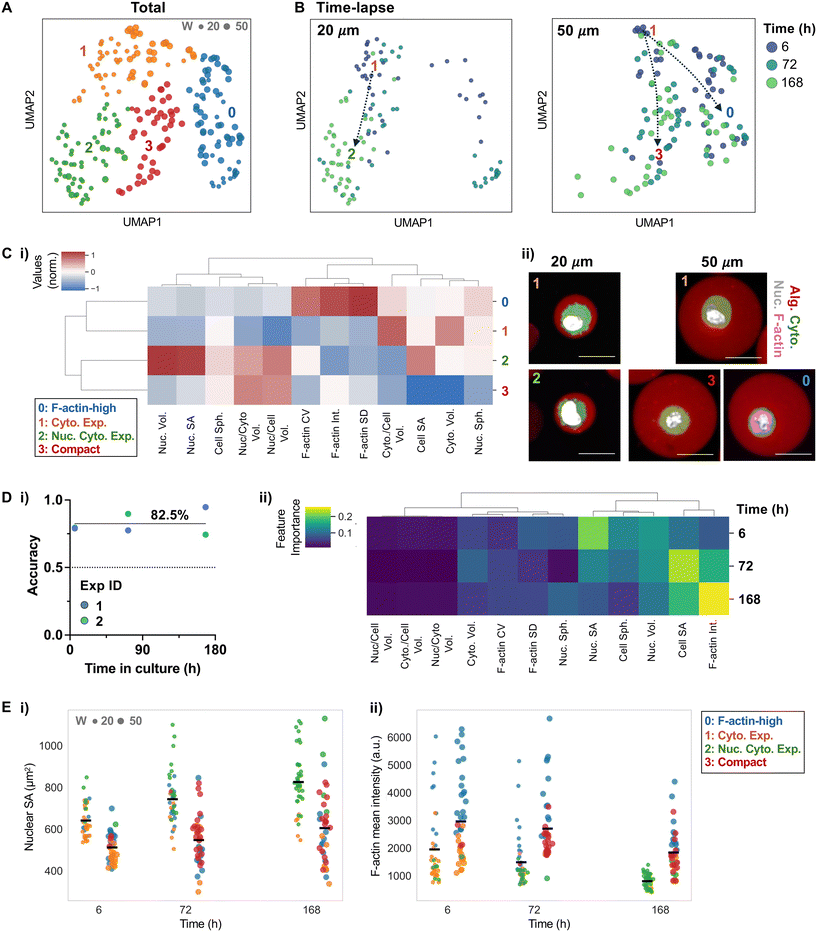
To investigate how physical confinement influences MSC morphology, we performed confocal imaging of cells in small and large microgels over one week. Consistent with our previous studies,13,15–18 most MSCs in high molecular weight LF200 alginate microgels remained viable (Fig. S5A) and in a single-cell state (Fig. S5B), enabling quantitative analysis of cell shape and cytoskeletal organization without confounding effects from cell death, proliferation, or egress. At each time point, cells were stained to visualize cytoplasm, nuclei, and F-actin. Twelve morphological and cytoskeletal features capturing nuclear/cytoplasmic size, shape, and F-actin organization were extracted from segmented 3D images. Microgel-level information was excluded to avoid circularity, ensuring that clustering reflects intrinsic cellular morphology and cytoskeletal organization rather than microgel size.We previously observed that MSCs in small microgels undergo more pronounced isotropic expansion of both cytoplasm and cell volume than in large microgels in culture.15,18 We hypothesized that physical confinement modulates cytoskeletal tension, such that cells in small microgels experience lower resistance, reflected by reduced F-actin, which permits greater volumetric expansion. In contrast, cells in larger microgels likely encounter higher confinement, leading to increased cytoskeletal organization and resistance to shape change. This dynamic may potentially generate heterogeneity in cellular morphological states even within the same microgel size. Unsupervised analysis identified different phenotypical clusters, with cells forming consistent clusters independent of experimental batch (Fig. S6). Four distinct phenotypic clusters were identified (Fig. 5A). Most MSCs in small microgels transitioned from cluster 1 to cluster 2 over one week in culture, whereas MSCs in large microgels transitioned from cluster 1 to either cluster 0 or cluster 3 within the same timeframe (Fig. 5B). Marker phenotype analysis showed that cluster 1 was characterized by increased cytoplasmic volume, while cluster 2 exhibited coordinated nuclear and cytoplasmic expansion. In contrast, cluster 3 displayed a compact morphology, whereas cluster 0 showed prominent F-actin organization accompanied by cytoplasmic enlargement (Fig. 5C; S7). Together, these findings provide new insights into how microgel confinement shapes MSC phenotypic trajectories, indicating that small microgels favor isotropic cell expansion, whereas larger microgels promote either compact states or cytoskeleton-reinforced phenotypes consistent with resistance to confinement.
To complement the unsupervised analysis and test whether these phenotypic differences contain predictive information about confinement, we performed supervised analysis using random forest classifiers trained on morphological and cytoskeletal features to predict microgel size across time points. Across time points, these features predicted microgel size with accuracy significantly above chance (78–85%) (Fig. 5D, i), demonstrating that confinement-dependent phenotypes are sufficiently distinct to enable classification. Feature-importance analysis revealed that nuclear surface area was most informative at early time points, whereas F-actin intensity became increasingly important at later stages (Fig. 5D, ii). Integrated analysis of nuclear surface area and mean F-actin intensity over time further illustrates how cells in different clusters progressively change in both nuclear and cytoskeletal organization (Fig. 5E). Notably, the temporal pattern of feature importance corresponds closely with the cluster transitions observed in small microgels: early nuclear morphology drives cluster 1 to 2 transitions, while later F-actin remodeling underlies progressive cytoskeletal adaptation that distinguishes cell states in large microgels. By enabling side-by-side comparisons under identical biochemical conditions, this platform uncovers physical microenvironment-driven heterogeneity that would be obscured in bulk hydrogels or sequential systems.
Discussion
We present a droplet microfluidic platform that integrates deterministic single-cell encapsulation with tunable 3D microenvironments, enabling parallel generation of multiple droplet sizes from a single precursor stream. Beyond confirming size-controlled droplet production, the system provides a means to systematically probe how physical confinement shapes cellular behavior within a single experiment. By enabling simultaneous comparisons of distinct microenvironments, the platform overcomes limitations of conventional stochastic or sequential workflows and reduces experimental variability.A key feature of this system is the integration of cell-triggered gelation with microfluidic device design, which selectively crosslinks only droplets containing cells while generating multiple droplet sizes simultaneously. This approach minimizes empty microgels and yields reproducible, cell-occupied 3D microniches for single-cell analysis. Unlike conventional “lab-in-droplet” systems, where droplets function as transient assay chambers,21 these crosslinked microgels form persistent, tunable microenvironments that support extended investigation of cell behaviors. Each microniche acts as a self-contained experimental unit with defined physical parameters, enabling direct interrogation of confinement-dependent responses within a single experiment.
Importantly, this platform exposes MSCs to mechanical cues arising from spatial confinement that are directly linked to phenotypic outcomes. Morphological profiling revealed that microgel size drives heterogeneous cellular responses, indicating that spatial constraints are not merely passive boundaries but actively regulate the partitioning of growth between nuclear and cytoplasmic compartments, a process known to be influenced by histone chaperones that regulate chromatin organization.22 Confinement drives sequential, size-dependent adaptation in MSCs: reduced resistance in small microgels permits coordinated nuclear and cytoplasmic expansion, whereas higher confinement in large microgels promotes progressive F-actin reinforcement leading to compact or cytoskeleton-stabilized states. These findings suggest that early nuclear deformation precedes cytoskeletal remodeling, engaging mechanosensitive pathways such as YAP/TAZ to stabilize confinement-specific phenotypes. The emergence of multiple adaptive states indicates that cells interpret confinement as a mechanical signal, activating distinct structural and signaling programs to maintain homeostasis under physical constraints.23 This heterogeneity would likely be obscured in bulk or sequential systems, highlighting the advantage of controlled, side-by-side microniche comparisons.
Although this study examined two discrete microgel sizes, the platform can readily be adapted to generate a broader range of sizes by incorporating additional parallel flow-focusing branches or adjusting channel geometries, enabling simultaneous production of multiple microgel size regimes. Systematic tuning of confinement could reveal thresholds or nonlinear effects in cellular morphology, signaling, and functional outputs. Future studies could leverage live-cell reporters, cytoskeletal perturbations, or pharmacological inhibition to test the mechanistic hypotheses proposed here, linking specific morphological clusters to defined mechanotransduction pathways. For instance, inhibiting actomyosin contractility24,25 or nuclear lamina remodeling26,27 may shift cells from one phenotypic cluster to another, providing causal evidence for the observed confinement-dependent adaptations. Optimizing encapsulation to achieve near-complete single-cell occupancy across multiple sizes within a single experiment would further reduce variability and increase statistical power, enabling robust correlations between early morphological adaptations and long-term functional states. Moreover, the platform could be extended to systematically vary other physical parameters, such as matrix stiffness, viscoelasticity, or ligand presentation, and incorporating downstream readouts, such as live-cell mechanotransduction assays,28 cytoskeletal remodeling,29 or single-cell omics,30 could link immediate confinement responses to persistent phenotypic or molecular programs,31 providing mechanistic insights into how microenvironments shape cell fate.
By integrating precise engineering with data-driven phenotyping, this platform can evolve into a versatile framework for predicting single-cell responses to defined physical and biochemical cues using artificial intelligence.32,33 Its modular architecture supports high-throughput and combinatorial investigations across matrix composition, viscoelasticity, and ligand presentation. This flexibility enables discovery of rare or transient phenotypic subtypes within controlled microenvironments. The system uniquely positions researchers to probe the interplay between confinement and mechanosensitive signaling at single-cell resolution, bridging fundamental mechanobiology with translational applications.6,34
Taken together, this deterministic, modular microfluidic system provides a scalable platform for probing cellular heterogeneity within precisely defined microenvironments. By bridging engineering precision with data-driven analysis, it clarifies how confinement shapes single-cell behavior and establishes a foundation for biomaterial design, predictive screening, and regenerative applications.
Materials and methods
Microfluidic device fabrication
Devices were designed in AutoCAD and fabricated using standard soft lithography based on previous studies.13,15,16,18,35 Designs were transferred to a chromium mask and patterned onto 4-inch silica wafers coated with SU-8 25 photoresist (Kayaku). Photoresist was spin-coated to 25 μm, UV-exposed (Karl SUSS MA-6 mask aligner), and developed per manufacturer instructions. Feature fidelity was confirmed via stereo microscopy, and channel height was measured with a KLA-Tencor P7 profilometer. Masters were silanized with trimethylchlorosilane to facilitate PDMS release. PDMS (Sylgard 184, 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 base
1 base![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) curing agent) was degassed, poured over the master, and cured at 65 °C for 3 h. Inlet/outlet ports were punched, and PDMS slabs bonded to glass slides using oxygen plasma. Channels were rendered hydrophobic with Aquapel (PPG Industries) and primed before use. Devices were interfaced with 27G × ½ needles (BD) and polyethylene tubing (Scientific Commodities, ID 0.38 mm, OD 1.09 mm). The two-branch geometry included a passive splitter dividing the aqueous stream into two flow-focusing junctions optimized to produce droplets with different sizes.
curing agent) was degassed, poured over the master, and cured at 65 °C for 3 h. Inlet/outlet ports were punched, and PDMS slabs bonded to glass slides using oxygen plasma. Channels were rendered hydrophobic with Aquapel (PPG Industries) and primed before use. Devices were interfaced with 27G × ½ needles (BD) and polyethylene tubing (Scientific Commodities, ID 0.38 mm, OD 1.09 mm). The two-branch geometry included a passive splitter dividing the aqueous stream into two flow-focusing junctions optimized to produce droplets with different sizes.
Finite element simulation of microfluidic flow
Finite element simulations were conducted using COMSOL Multiphysics 6.1 to predict the flow velocity field and the flow rate through the channels. The geometry of the channel was imported as CAD file to create the model. Creeping flow interface (spf) without inertial force was used with shallow channel approximation to simulate the microfluidic channel using a 2D geometry. The flow profile in the thickness direction was approximated by the Reynolds lubrication equations. The channel thickness was 25 μm. The viscosities of the dispersed phase (1% LF200 solution) and the oil were 0.3 Pa s and 0.0013 Pa s, respectively, for corresponding water and oil domains in the channels. No-slip boundary conditions were enforced at the walls. The flow rates at the two inlets were given as the experimentally measured values and the outlet was given zero relative pressure boundary condition. Physics-controlled mesh type was used to make sure the mesh convergence was achieved and 928![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 306 degrees of freedom were used. The flow rate through the two junctions were calculated by integrating the velocity across the cross sections.
306 degrees of freedom were used. The flow rate through the two junctions were calculated by integrating the velocity across the cross sections.
Aqueous and oil phase preparation
The aqueous phase was based on a sodium alginate (240 kDa, LF200, FMC Polymer) solution in Dulbecco's modified Eagle's medium (DMEM, Thermo). For crosslinking of cell-free microgels, calcium carbonate (CaCO3; CalEssence® 70 PCC) was suspended in DMEM containing 10% fetal bovine serum, 1% penicillin/streptomycin, and 1% GlutaMAX (Thermo). Aggregates were disrupted by sonication, and the suspension was diluted with HEPES-buffered DMEM (pH 7.8). Equal volumes of the CaCO3 suspension and alginate were mixed to obtain a final working solution of 1% (w/v) alginate with 2 mg mL−1 CaCO3.36 The oil phase consisted of Novec HFE-7500 (3M) supplemented with 1% (w/v) Krytox 157 FSH (PFPE surfactant, Dupont) and 0.03% acetic acid.Preparation of cells in alginate solutions
Mouse D1 mesenchymal stromal cells (MSCs) were cultured in T-175 flasks (Greiner) with complete DMEM. Cells were maintained at 37 °C and 5% CO2 and passaged at 80% confluency using 0.05% trypsin–EDTA (Thermo). Cells from passages 9–15 were used. For cell encapsulation experiments, 6 million MSCs per ml were mixed with 2 mg mL−1 CaCO3 in DMEM and gently rocked for 1 h to enable particle association.36 Cells were washed through a 40 μm strainer to remove aggregates and repeatedly centrifuged in HEPES-buffered DMEM (5× at 550 rpm) to remove excess CaCO3, ensuring that available calcium for crosslinking originated predominantly from cell-associated particles. The final suspension was resuspended in 200 μL HEPES-buffered DMEM and mixed 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 with 2% alginate to generate a working solution containing 1% alginate and CaCO3-coated cells at a final concentration of 15 million cells per mL. A fraction of alginate (1/40) conjugated with Lissamine rhodamine B ethylenediamine (Thermo) was added for fluorescent labeling of the gel coatings.
1 with 2% alginate to generate a working solution containing 1% alginate and CaCO3-coated cells at a final concentration of 15 million cells per mL. A fraction of alginate (1/40) conjugated with Lissamine rhodamine B ethylenediamine (Thermo) was added for fluorescent labeling of the gel coatings.
Droplet generation and characterization
The aqueous and oil phases were loaded into 1 mL and 3 mL syringes, respectively, and infused via a dual syringe pump (Harvard Apparatus 33). Flow rates were varied between 1–2 μl min−1 (aqueous) and 3–15 μl min−1 (oil) to evaluate droplet size distributions. For the main single cell encapsulation experiments, a constant aqueous flow of 1 μl min−1 and oil flow of 6 μl min−1 were used. Droplet formation was monitored at the flow-focusing junctions using brightfield microscopy. Droplet sizes were quantified both in emulsion (brightfield microscopy) and post-breaking (fluorescence microscopy with rhodamine labeling). Microfluidic junction stability was assessed by imaging droplets at 5, 15, 30, and 60 min, and by measuring channel dimensions to confirm minimal swelling immediately after encapsulation. Fabricated microgels were collected at designated time points, and their diameters were measured both in emulsion and after emulsion breaking using Fiji (ImageJ). Size distributions were analyzed, and the coefficient of variation (CV) was calculated to assess size uniformity.Single cell encapsulation and microgel recovery
For encapsulation efficiency studies, cell-containing droplets were collected after 1 h of device operation and incubated with gentle rocking for 30 min to complete crosslinking. Emulsions were destabilized by adding 20% perfluorooctanol (PFO; Alfa Aesar) in HFE 7500, followed by repeated washing with 0.1% bovine serum albumin (BSA; Sigma) in DMEM 5 times. The resulting microgels were resuspended in complete DMEM for analysis. Cell viability was assessed by staining with calcein AM (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution; Biotium) for 30 min at 37 °C, and gels were immobilized on poly-L-lysine coated slides (Sigma-Aldrich) for fluorescence imaging. Encapsulation efficiency was calculated as the ratio of droplets containing single viable cells to the total number of droplets. In continuous-operating encapsulation experiments, cell-containing droplets generated during representative time windows (15–30 min, 75–90 min, and 135–150 min) were collected and analyzed. The fraction of droplets containing cells was quantified both before and after emulsion breaking using brightfield microscopy, following the same imaging and counting criteria described above.
1000 dilution; Biotium) for 30 min at 37 °C, and gels were immobilized on poly-L-lysine coated slides (Sigma-Aldrich) for fluorescence imaging. Encapsulation efficiency was calculated as the ratio of droplets containing single viable cells to the total number of droplets. In continuous-operating encapsulation experiments, cell-containing droplets generated during representative time windows (15–30 min, 75–90 min, and 135–150 min) were collected and analyzed. The fraction of droplets containing cells was quantified both before and after emulsion breaking using brightfield microscopy, following the same imaging and counting criteria described above.
Confocal imaging and 3D image processing for morphological analysis
Cell-containing microgels were immobilized on poly-L-lysine (PLL)-coated glass slides to ensure stable positioning during imaging. Cells were stained membrane-permeable dyes, including calcein AM (cytoplasm; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution; Biotium), Hoechst 33342 (nuclei; 1
1000 dilution; Biotium), Hoechst 33342 (nuclei; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution; Thermo Fisher Scientific), and SPY650-FastAct_X or ‘SiR-XActin’37 (F-actin; 1
1000 dilution; Thermo Fisher Scientific), and SPY650-FastAct_X or ‘SiR-XActin’37 (F-actin; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 dilution; Spirochrome) for 1 h at 37 °C. Full 3D Z-stacks were acquired using a Nikon CSU-W1 SoRa spinning disk confocal microscope. Image stacks were processed in IMARIS (Bitplane) for three-dimensional reconstruction and volumetric analysis using a surface segmentation workflow. Specifically, surface objects were generated with IMARIS's automated surface-detection tools, which use voxel intensity thresholding combined with background subtraction and region-growing algorithms to identify connected fluorescent structures in 3D. Automated segmentation provided object boundaries for nuclei, cytoplasm, and microgels, and these surfaces were then reviewed and refined by adjusting thresholds (within ±10%) as needed to ensure accurate segmentation. Reconstructed surfaces were used to quantify Hoechst-, Calcein-AM-, and rhodamine-positive volumes. Within the Calcein-AM-defined cytoplasmic volume, the mean fluorescence intensity (MFI) and standard deviation of the SPY650-FastAct_X channel were measured, and the coefficient of variation (CV) was calculated to assess intracellular actin heterogeneity. Confocal images of cell-containing microgels were also subjected to Z-stack projection to generate maximum intensity projection images. A line was drawn across the central axis of each microgel, and the fluorescence intensity profile of the rhodamine channel was analyzed using Fiji (ImageJ) software. Rhodamine intensity distribution along the central line was quantified to assess spatial uniformity within the microgels.
1000 dilution; Spirochrome) for 1 h at 37 °C. Full 3D Z-stacks were acquired using a Nikon CSU-W1 SoRa spinning disk confocal microscope. Image stacks were processed in IMARIS (Bitplane) for three-dimensional reconstruction and volumetric analysis using a surface segmentation workflow. Specifically, surface objects were generated with IMARIS's automated surface-detection tools, which use voxel intensity thresholding combined with background subtraction and region-growing algorithms to identify connected fluorescent structures in 3D. Automated segmentation provided object boundaries for nuclei, cytoplasm, and microgels, and these surfaces were then reviewed and refined by adjusting thresholds (within ±10%) as needed to ensure accurate segmentation. Reconstructed surfaces were used to quantify Hoechst-, Calcein-AM-, and rhodamine-positive volumes. Within the Calcein-AM-defined cytoplasmic volume, the mean fluorescence intensity (MFI) and standard deviation of the SPY650-FastAct_X channel were measured, and the coefficient of variation (CV) was calculated to assess intracellular actin heterogeneity. Confocal images of cell-containing microgels were also subjected to Z-stack projection to generate maximum intensity projection images. A line was drawn across the central axis of each microgel, and the fluorescence intensity profile of the rhodamine channel was analyzed using Fiji (ImageJ) software. Rhodamine intensity distribution along the central line was quantified to assess spatial uniformity within the microgels.
Machine learning-based cell phenotype analysis
Quantitative morphological and cytoskeletal features were extracted from segmented nuclei, cytoplasm, and microgels and compiled into a cell-by-feature matrix. Twelve three-dimensional features were measured for each cell, including nuclear volume, cytoplasmic volume, nuclear surface area, cytoplasmic surface area, nuclear sphericity, cell sphericity, mean F-actin intensity, F-actin standard deviation, F-actin coefficient of variation, nucleus-to-cytoplasm volume ratio, nucleus-to-cell volume ratio, and cytoplasm-to-cell volume ratio. These metrics capture key aspects of cell size, shape, and cytoskeletal organization in three dimensions. The resulting feature matrix was standardized using Scanpy's sc.pp.scale function in Python to normalize for scale differences across features.To explore heterogeneity in MSC phenotypes, unsupervised analysis was performed. Twelve morphological and cytoskeletal features per cell were log-transformed and standardized before analysis. Principal component analysis (PCA) was applied to reduce dimensionality and noise. A neighborhood graph was constructed to capture local relationships among cells, followed by Uniform Manifold Approximation and Projection (UMAP) to visualize phenotypic structure in two dimensions. Leiden clustering was applied to identify morphological clusters, with parameters (Leiden resolution, UMAP n_neighbors, min_dist, PCA variance threshold) optimized via grid search to maximize silhouette scores on PCA embeddings. To ensure phenotypic variation reflected intrinsic cell behavior rather than experimental batch, silhouette scores on PCA embeddings (0 = well-mixed, 1 = strong separation) and cluster-wise entropy of replicate labels (higher = better mixing) were calculated. Cluster trajectories were computed separately for each microgel size by tracking cells assigned to Leiden cluster 1 at the earliest time point and inferring transitions at subsequent time points based on proximity in UMAP space. Resulting trajectories were simplified by removing consecutive duplicate cluster assignments and merging identical paths, and branches representing fewer than 5% of the starting population were excluded to retain dominant phenotypic transitions.
To determine whether single-cell morphology and cytoskeletal organization contained predictive information about microgel confinement, random forest (RF) classifiers were trained at each post-encapsulation time point.38 Input features included 12 morphological and cytoskeletal metrics extracted from 3D confocal images, excluding any direct microgel measurements. The target label was the microgel size for each cell. Models were implemented using the RandomForestClassifier in scikit-learn39 with 500 trees (n_estimators = 500) and default Gini impurity as the split criterion. To avoid batch effects, cross-validation was performed using GroupKFold with splits defined by experimental replicate, ensuring that all cells from a single replicate were withheld from training during testing. For each fold, the classifier was trained on n – 1 replicates and evaluated on the held-out replicate. Accuracy was calculated per replicate using accuracy_score and mean ± standard deviation across folds provided the time-resolved prediction accuracy. Feature importance for each time point was computed directly from the trained random forests as the normalized decrease in node impurity (Gini) attributable to each feature, averaged across all trees in the forest. To visualize temporal dynamics, importances were collected across all time points, pivoted to a feature x time matrix, and features were sorted by the variance of their importance over time.
Statistical analyses
All statistical analyses were performed using GraphPad Prism version 10.1.0, as specified in the figure captions. Unless otherwise indicated, comparisons between groups were conducted using one-way ANOVA with Tukey's multiple comparisons test when standard deviations were similar, or one-way Welch ANOVA with Dunnett's T3 multiple comparisons test when standard deviations were unequal. Differences were considered statistically significant at p < 0.05.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article are included in the supplementary information (SI), including microfluidic device characterization, droplet and microgel analysis, cell viability and encapsulation efficiency, and machine-learning-based phenotypic analysis. See DOI: https://doi.org/10.1039/d5lc00839e.Acknowledgements
This work utilized instruments from the Microscope Core Facility at the University of Michigan and the Nanotechnology Core Facility at the University of Illinois at Chicago. It was supported by the National Institutes of Health under Grants R01-HL141255, R01-GM141147, R01-EB034507, and R01-HL171590, as well as by the National Science Foundation CAREER Grant 2143857 (to J.-W. S.) and CAREER Grant 2339054 (to Z. P.).References
- O. Chaudhuri, J. Cooper-White, P. A. Janmey, D. J. Mooney and V. B. Shenoy, Nature, 2020, 584, 535–546 CrossRef CAS PubMed.
- S. Kim, M. Uroz, J. L. Bays and C. S. Chen, Dev. Cell, 2021, 56, 180–191 CrossRef CAS PubMed.
- S. W. Wong, S. Lenzini, R. Giovanni, K. Knowles and J. W. Shin, Acta Biomater., 2021, 133, 126–138 CrossRef CAS PubMed.
- S. Lenzini, D. Devine and J. W. Shin, Front. Bioeng. Biotechnol., 2019, 7, 260 CrossRef PubMed.
- D. E. Discher, L. Smith, S. Cho, M. Colasurdo, A. J. Garcia and S. Safran, Annu. Rev. Biophys., 2017, 46, 295–315 CrossRef CAS PubMed.
- P. Gupta, O. Alheib and J. W. Shin, Adv. Drug Delivery Rev., 2023, 201, 115010 CrossRef CAS PubMed.
- T. Kamperman, M. Karperien, S. Le Gac and J. Leijten, Trends Biotechnol., 2018, 36, 850–865 CrossRef CAS PubMed.
- W. Youn, J. Y. Kim, J. Park, N. Kim, H. Choi, H. Cho and I. S. Choi, Adv. Mater., 2020, 32, e1907001 CrossRef PubMed.
- M. Pires-Santos, S. Nadine and J. F. Mano, Small Sci., 2024, 4, 2300332 CrossRef CAS PubMed.
- L. Mazutis, J. Gilbert, W. L. Ung, D. A. Weitz, A. D. Griffiths and J. A. Heyman, Nat. Protoc., 2013, 8, 870–891 CrossRef CAS PubMed.
- W. Li, E. W. K. Young, M. Seo, Z. Nie, P. Garstecki, C. A. Simmons and E. Kumacheva, Soft Matter, 2008, 4, 258–262 RSC.
- D. R. Link, S. L. Anna, D. A. Weitz and H. A. Stone, Phys. Rev. Lett., 2004, 92, 054503 CrossRef CAS PubMed.
- A. S. Mao, J. W. Shin, S. Utech, H. Wang, O. Uzun, W. Li, M. Cooper, Y. Hu, L. Zhang, D. A. Weitz and D. J. Mooney, Nat. Mater., 2017, 16, 236–243 CrossRef CAS PubMed.
- P. S. Lienemann, T. Rossow, A. S. Mao, Q. Vallmajo-Martin, M. Ehrbar and D. J. Mooney, Lab Chip, 2017, 17, 727–737 RSC.
- S. W. Wong, S. Lenzini, R. Bargi, Z. Feng, C. Macaraniag, J. C. Lee, Z. Peng and J. W. Shin, Adv. Sci., 2020, 7, 2001066 CrossRef CAS PubMed.
- S. W. Wong, C. R. Tamatam, I. S. Cho, P. T. Toth, R. Bargi, P. Belvitch, J. C. Lee, J. Rehman, S. P. Reddy and J. W. Shin, Nat. Biomed. Eng., 2022, 6, 54–66 CrossRef CAS PubMed.
- I. S. Cho, A. Yaghmour, A. Joshi, P. Gupta, M. A. Sanborn, M. P. Zappia, S. W. Wong, G. Cheng, M. V. Frolov, J. Rehman and J. W. Shin, Adv. Mater., 2026, 38, e04590 CrossRef CAS PubMed.
- I. S. Cho, P. Gupta, N. Mostafazadeh, S. W. Wong, S. Saichellappa, S. Lenzini, Z. Peng and J. W. Shin, Adv. Sci., 2023, 10, e2206014 CrossRef PubMed.
- E. Hinderink, B. Bera, C. Schinkel and V. V. Steijn, Sci. Rep., 2025, 15, 5720 CrossRef PubMed.
- P. Zhu and L. Wang, Lab Chip, 2016, 17, 34–75 RSC.
- L. Nan, H. Zhang, D. A. Weitz and H. C. Shum, Lab Chip, 2024, 24, 1135–1153 RSC.
- P. Chen, M. Tomschik, K. M. Nelson, J. Oakey, J. C. Gatlin and D. L. Levy, J. Cell Biol., 2019, 218, 4063–4078 CrossRef CAS PubMed.
- S. Brielle, D. Bavli, A. Motzik, Y. Kan-Tor, X. Sun, C. Kozulin, B. Avni, O. Ram and A. Buxboim, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2016322118 CrossRef CAS PubMed.
- J. W. Shin, J. Swift, K. R. Spinler and D. E. Discher, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 11458–11463 CrossRef CAS PubMed.
- J. W. Shin, A. Buxboim, K. R. Spinler, J. Swift, D. A. Christian, C. A. Hunter, C. Leon, C. Gachet, P. C. Dingal, I. L. Ivanovska, F. Rehfeldt, J. A. Chasis and D. E. Discher, Cell Stem Cell, 2014, 14, 81–93 CrossRef CAS PubMed.
- J. W. Shin, K. R. Spinler, J. Swift, J. A. Chasis, N. Mohandas and D. E. Discher, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 18892–18897 CrossRef CAS PubMed.
- J. Swift, I. L. Ivanovska, A. Buxboim, T. Harada, P. C. Dingal, J. Pinter, J. D. Pajerowski, K. R. Spinler, J. W. Shin, M. Tewari, F. Rehfeldt, D. W. Speicher and D. E. Discher, Science, 2013, 341, 1240104 CrossRef PubMed.
- A. Colom, E. Derivery, S. Soleimanpour, C. Tomba, M. D. Molin, N. Sakai, M. Gonzalez-Gaitan, S. Matile and A. Roux, Nat. Chem., 2018, 10, 1118–1125 CrossRef CAS PubMed.
- G. Lukinavicius, L. Reymond, E. D'Este, A. Masharina, F. Gottfert, H. Ta, A. Guther, M. Fournier, S. Rizzo, H. Waldmann, C. Blaukopf, C. Sommer, D. W. Gerlich, H. D. Arndt, S. W. Hell and K. Johnsson, Nat. Methods, 2014, 11, 731–733 CrossRef CAS PubMed.
- J. de Rutte, R. Dimatteo, M. M. Archang, M. van Zee, D. Koo, S. Lee, A. C. Sharrow, P. J. Krohl, M. Mellody, S. Zhu, J. V. Eichenbaum, M. Kizerwetter, S. Udani, K. Ha, R. C. Willson, A. L. Bertozzi, J. B. Spangler, R. Damoiseaux and D. Di Carlo, ACS Nano, 2022, 16, 7242–7257 CrossRef CAS PubMed.
- M. Meteling, C. Johnbosco, A. Wolfel, F. Conceicao, K. Govindaraj, L. Moreira Teixeira and J. Leijten, Adv. Mater., 2025, 37, e2415981 CrossRef PubMed.
- T. Kulkarni, O. M. Robinson, A. Dutta, D. Mukhopadhyay and S. Bhattacharya, Mater. Today Bio, 2024, 25, 100970 CrossRef CAS PubMed.
- A. M. Bornschlegl and A. B. Dietz, Tissue Cell, 2025, 97, 103041 CrossRef CAS PubMed.
- J. W. Shin and D. J. Mooney, Cell Stem Cell, 2016, 18, 16–19 CrossRef CAS PubMed.
- D. Devine, V. Vijayakumar, S. W. Wong, S. Lenzini, P. Newman and J. W. Shin, Adv. Biosyst., 2020, 4, e2000012 CrossRef PubMed.
- T. Puttrich, S. O'Donnell, S. W. Wong, M. Kotche, A. E. Felder and J. W. Shin, PLoS One, 2023, 18, e0282563 CrossRef CAS PubMed.
- V. Nasufovic, J. Kompa, H. L. Lindamood, M. Blumke, R. Dibsy, B. Koch, V. Levario-Diaz, K. Weber, M. Maager, E. Nomerotskaia, A. Echard, E. A. Cavalcanti-Adam, E. A. Vitriol, H. D. Arndt and K. Johnsson, Angew. Chem., Int. Ed., 2025, 64, e202509285 CrossRef CAS PubMed.
- L. Breiman, Mach. Learn., 2001, 45, 5–32 CrossRef.
- L. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel and B. Thirion, J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed.
Footnote |
| † Equal contribution. |
| This journal is © The Royal Society of Chemistry 2026 |