Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
High-throughput and efficient fabrication of engineered skeletal muscle tissue via streamlined 3D multimaterial bioprinting
Tae-Eun
Lim
 a,
Ashfaq
Ahmad
ab,
Yeong-Jin
Choi
a,
Ashfaq
Ahmad
ab,
Yeong-Jin
Choi
 *cd and
Hee-Gyeong
Yi
*cd and
Hee-Gyeong
Yi
 *abe
*abe
aDepartment of Convergence Biosystems Engineering, College of Agriculture and Life Sciences (CALS), Chonnam National University, Gwangju, Republic of Korea. E-mail: hgyi@chonnam.ac.kr
bInterdisciplinary Program in IT-Bio Convergence System, Chonnam National University, Republic of Korea
cAdvanced Bio and Healthcare Materials Research Division, Korea Institute of Materials Science (KIMS), Changwon, Republic of Korea. E-mail: jinchoi@kims.re.kr
dAdvanced Materials Engineering, Korea National University of Science and Technology (UST), Daejeon, Republic of Korea
eInstitute for Biomedical Science, Chonnam National University Hwasun Hospital, Hwasun, Republic of Korea
First published on 2nd March 2026
Abstract
Skeletal muscle diseases have led to an increasing need for a range of physiologically relevant organ-on-a-chip and three-dimensional (3D) culture platforms owing to their clinical significance and ongoing efforts to develop effective therapeutics. However, conventional fabrication approaches still face limitations—such as low throughput, complex fabrication steps, and prolonged production times—making them difficult to apply to efficient drug discovery and development for muscle-related diseases. Here, we present a high-throughput 3D skeletal muscle tissue platform fabricated via a streamlined 3D cell-printing process integrated with a commercially available multi-well plate. Combined with automated 3D printing, the 96-well format enables efficient, rapid, and reproducible fabrication, while significantly reducing processing time and ensuring high tissue uniformity across all wells. To maximize production efficiency, optimized anchor structures and an amphiphilic coating were employed to enhance tissue stability and alignment. In addition, systematic optimization of the tissue geometry and 3D bioprinter-based fabrication parameters enabled consistent production of functional and mature 3D skeletal muscle tissues. Overall, this platform integrates 3D bioprinting, optimized anchor design, and a controlled tissue geometry to enable a rapid, high-throughput, automation-compatible, and error-minimizing fabrication process. These results demonstrate that the proposed platform can serve as a useful tool for diverse biomedical applications, including the development of effective therapeutics for muscle-related diseases.
Introduction
Skeletal muscle is the contractile tissue responsible for generating body movements and accounts for approximately 40% of total body mass.1 It is a multifunctional organ that plays essential roles in locomotion, thermoregulation, joint stabilization, and metabolic homeostasis.2–4 Given these diverse physiological functions, numerous disorders are associated with skeletal muscle. Skeletal muscle can be affected by a wide range of pathological conditions arising from genetic, metabolic, or structural abnormalities, leading to disorders such as myopathies and musculoskeletal diseases.1 Moreover, many severe conditions, including Duchenne muscular dystrophy (DMD) and amyotrophic lateral sclerosis (ALS), currently lack effective therapeutic options.5,6 Although the underlying mechanisms and secondary symptoms vary among muscle disorders, progressive muscle atrophy, weakness, and degeneration commonly occur.7 Consequently, the development of effective therapeutics for muscle diseases remains an urgent and critical challenge in biomedical research.However, the development of new therapeutics requires not only extensive screening of candidate drugs and compounds but also rigorous validation of their efficacy. Preclinical evaluation typically relies on in vitro models and animal models to assess safety and pharmacological activity prior to clinical translation.8,9 Conventional 2D culture systems, however, fail to replicate the three-dimensional complexity and native microarchitecture of skeletal muscle tissue.10 Although animal models provide valuable physiological context, they are often costly, time-consuming, and limited in their ability to accurately recapitulate human-specific drug responses.11–17
To overcome these limitations while enabling more efficient and accurate therapeutic development, 3D tissues and organ-on-a-chip platforms have emerged as powerful tools in biomedical research. These systems can reproduce both the structural and functional characteristics of cellular and tissue microenvironments, and they have been applied across a wide range of biomedical applications.18,19 Among various approaches for developing 3D tissues, 3D cell-printing technology is particularly advantageous, as it allows the construction of complex 3D biological architectures that mimic the native tissue microenvironment with high precision and reproducibility.20
Currently, to fabricate 3D platforms for muscle cell culture, molds are typically designed using computer-aided modeling and fabricated either with polydimethylsiloxane (PDMS) molding or through 3D printing. However, these conventional fabrication methods require long processing times and multiple complex steps to produce a single platform, thereby limiting scalability and accessibility.7,21–23
To address these challenges, we previously developed an in vitro platform that integrates 3D bioprinting technology with commercially available multi-well plates for biomimetic, high-throughput tissue modeling.24 Building upon this concept, the present study employed a universally accessible and commercially available 96-well plate to simplify fabrication and enhance user accessibility. By combining the automation and precision of 3D bioprinting, we successfully established a streamlined fabrication process capable of producing high-throughput skeletal muscle (SKM) tissue platforms within a single day.
Collagen, a hydrogel widely used in tissue engineering, was utilized to encapsulate muscle cells,25–27 and multiple optimization strategies were implemented to improve the accuracy, stability, and efficiency of the fabricated platform. Furthermore, drug testing conducted using the optimized 3D SKM platform demonstrated its feasibility as an efficient high-throughput drug screening system.
Finally, drug testing performed on the optimized skeletal muscle platform validated its applicability for efficient drug screening. This approach not only broadens the range of tissues that can be engineered but also provides a strong foundation for accelerating therapeutic development by improving fabrication efficiency and reducing cost and time requirements.
Materials and methods
Cell culture
C2C12 mouse myoblast cells were cultured in a growth medium (GM) composed of high-glucose Dulbecco's Modified Eagle's Medium (DMEM, Hyclone) containing 10% fetal bovine serum (FBS, Hyclone) and 1% penicillin–streptomycin (P/S, Hyclone) at 37 °C in a humidified 5% CO2 incubator. The culture medium was changed every two days. Extended cultures were conducted until the cells reached 90% confluency. Before hydrogel embedding, the cells were harvested by treatment with 0.25% trypsin in EDTA (Promocell) for 3 minutes, followed by centrifugation at 1200 rpm for 3 minutes. After discarding the supernatant, the cells were either subcultured or used for this study. For this study, cells between passages 3 and 8 were used.Bioink preparation
To prepare the bioink, type I collagen (Atellocollagen, Dalim Tissen) was used as the primary component. Collagen was dissolved in 0.01 N hydrochloric acid (HCl, Biosesang, Korea) at 4 °C for at least 48 hours. During this period, the mixture was vortexed at least twice, resulting in the formation of a pre-gel solution. Once the collagen was completely dissolved, the solution was neutralized using 1 N sodium hydroxide (NaOH, Biosesang), followed by the addition of 10× phosphate-buffered saline (PBS, Biosesang) to maintain osmotic balance. The final volume was adjusted with distilled water. The resulting collagen-based bioink had a final concentration of 0.5 wt% with the pH adjusted to 7.0 and was stored at 4 °C until use.Fabrication of AnchorPlate96
To fabricate the high-throughput platform, a commercially available non-treated 96-well cell culture plate (SPL Life Sciences, Korea) was utilized. Initially, anchors were printed using a multi-head 3D bioprinter (3DX Bioprinter, T&R Biofab, Korea). Printing was performed at room temperature using a double-thread screwed plastic needle (DPN-22G-2, Musashi Engineering, Japan) under pneumatic pressure ranging from 140 to 170 kPa. For the anchor material, Silicone 1700 (Dow Corning) was mixed with a catalyst prior to printing using a Thinky mixer (THINKY, Japan) and centrifuged to remove any bubbles. To prevent the skeletal muscle (SKM) tissue from detaching and floating away from the anchor, a deliberate printing delay was introduced to form mushroom-shaped pin anchors. The total printing time for anchors in the 96-well plate was approximately 25 minutes. Following printing, the anchor-printed plates were cured overnight in a 60 °C oven. This process resulted in the fabrication of the AnchorPlate96. The anchor printing code used in this study is provided in the supplementary source code.3D muscle cell printing
Before bioprinting, the AnchorPlate96 was wiped with 70% ethanol and exposed to UV light for 30 min in a clean bench. After sterilization, 4% Pluronic® F-127 (PF-127, Sigma-Aldrich) was used to coat the surface of the well for 30 minutes and the wells were washed three times with 1× PBS to remove any excess PF-127 solution.22 For bioink preparation, C2C12 myoblasts were trypsinized and resuspended at a density of 1.0 × 107 cells per mL in 0.5 wt% collagen solution at 4 °C using a positive displacement pipette.The bioink was loaded into sterile 3 mL or 10 mL disposable syringes (KOVAX) and printed into the precoated 96-well plates using the 3D bioprinter. Printing was conducted with a plastic needle (PN-27D-B, Musashi Engineering, Japan) at 15 °C under various pneumatic pressures of 8–16 kPa, depending upon desired muscle bundle thickness. In this study, pneumatic pressures in the range of 10–15 kPa were used. Based on our previous studies, bioink printing using a 3D bioprinter did not result in any noticeable difference in cell viability before and after the printing process.28,29 The bioink printing code used in this study is also provided in the supplementary source code. After printing, the constructs were incubated for 1 h to allow gelation of bioink. GM was added on the same day. From day 2, when SKM tissue has fully compacted, the medium was replaced with differentiation medium (DM) consisting of 2% horse serum (HS, Gibco), 1% penicillin–streptomycin (P/S, Hyclone), and 50 ng mL−1 insulin-like growth factor-1 (IGF-1, Sigma-Aldrich). The culture medium was changed every 24 h.
Cell viability assay
After treatment of the fabricated muscle constructs with paclitaxel, cisplatin, dexamethasone, or IGF-1, cell viability was evaluated on days 1, 3, and 5 using a Cell Counting Kit-8 (CCK-8, Dojindo, Japan). Prior to each assay, the constructs were washed with 1× PBS. The culture medium was then mixed with the CCK-8 solution at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 ratio, and 150 μL of the mixture was added to each well. After incubation for 30 min, the medium containing the CCK-8 reagent was collected, and the absorbance was measured at 450 nm using a microplate reader.
10 ratio, and 150 μL of the mixture was added to each well. After incubation for 30 min, the medium containing the CCK-8 reagent was collected, and the absorbance was measured at 450 nm using a microplate reader.
Immunofluorescence
3D SKM tissues were stained with an anti-fast myosin skeletal heavy chain antibody (MHC, Abcam), α-actinin (Sigma-Aldrich), F-actin (phalloidin, Sigma-Aldrich), and 4′,6-diamidino-2-phenylindole (DAPI) at days 7 and 14 after printing on the platform. First, tissues were fixed in 4% paraformaldehyde (Biosesang) for 30 min, followed by permeabilization with 0.2% Triton X-100 (Biosesang) for 10 min at room temperature. The tissues were then blocked with 5% bovine serum albumin (BSA, GenDEPOT) for 15 min. For SKM tissue fluorescence imaging, primary antibodies against MHC and α-actinin (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200) were applied and incubated overnight at 4 °C. Prior to each subsequent step, tissues were washed three times with 1× PBS. After overnight incubation, tissues were washed three times and then incubated for 2 h with secondary antibodies conjugated with fluorophores (Alexa Fluor 594 and Alexa Fluor 488 (Invitrogen)) together with DAPI and phalloidin conjugated to Alexa Fluor 594 (1
200) were applied and incubated overnight at 4 °C. Prior to each subsequent step, tissues were washed three times with 1× PBS. After overnight incubation, tissues were washed three times and then incubated for 2 h with secondary antibodies conjugated with fluorophores (Alexa Fluor 594 and Alexa Fluor 488 (Invitrogen)) together with DAPI and phalloidin conjugated to Alexa Fluor 594 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200) to stain filamentous actin (F-actin). Secondary antibodies and DAPI were diluted at 1
200) to stain filamentous actin (F-actin). Secondary antibodies and DAPI were diluted at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 and 1
500 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200, respectively. After final washing three times, samples were imaged using confocal and fluorescence microscopy (ZEISS) and analyzed with ImageJ (NIH). After additional washing three times, the samples were imaged and analyzed as described above.
200, respectively. After final washing three times, samples were imaged using confocal and fluorescence microscopy (ZEISS) and analyzed with ImageJ (NIH). After additional washing three times, the samples were imaged and analyzed as described above.
Image analysis
Myotubes are defined as multinucleated fibers containing at least three nuclei. Myotubes aligned within ±10° of the major axis of the construct were classified as aligned myotubes.20 Muscle bundle thickness was measured by dividing each tissue into four distinct areas and calculating the average thickness for each area. In addition, myotube length was measured by selecting muscle fibers containing at least three nuclei, as defined above. The fusion index was calculated as the ratio of the number of nuclei within myotubes to the total number of nuclei.20 The CoV was calculated as the standard deviation of muscle bundle thickness divided by the mean thickness.30 To compare the CoV between two groups, samples were obtained from three whole plates per group. The sample size corresponds to the number of wells with successfully formed tissues. All image analysis graphs were generated based on images obtained by selecting multiple regions from each sample. All image analysis graphs were generated based on images obtained from multiple regions within each sample. A minimum of three independent samples were analyzed (n ≥ 3), and analyses were performed using ImageJ software. Detailed methods for the image analyses are described in Fig. S3.Statistical analysis
All quantitative data are presented as the mean ± standard deviation. Statistical significance was assessed using unpaired Student's t-test or one-way ANOVA followed by Dunnett's multiple comparison test, with a minimum of three independent samples per condition (n ≥ 3). Normality was assessed using the Shapiro–Wilk test (α = 0.05). When normality was not satisfied (p < 0.05), statistical comparisons were performed using the Mann–Whitney U test (two groups) or the Kruskal–Wallis test (multiple groups), as appropriate. Statistical analyses and graphs were generated using Microsoft Excel and GraphPad Prism. Statistical significance was indicated as follows: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, and ns: not significant.Results and discussion
1. Fabrication of a 3D skeletal muscle tissue platform using a 3D bioprinter on a commercial 96-well plate
In 2D culture systems, standardized well plates confer advantages for both users and manufacturers. They obviate custom modifications and allow a broad range of laboratory ware to be used alongside various automated instruments such as microplate equipment, ensuring compatibility with fully automated systems like liquid handlers and robotic arms. However, as the generation of three-dimensional tissues within these standardized multi-well plates has not been demonstrated to date, we established a fabrication approach that leverages the automation capabilities of 3D bioprinting in combination with standard multi-well plates to maximize the reproducibility of the developed 3D tissues and minimize the possibility of error. This is important because it enables in situ, non-destructive, high-throughput readouts without moving or detaching the tissues. This platform required three essential components: a commercially available non-treated 96-well plate, a silicone-based anchor ink for anchor printing, and a muscle cell-laden bioink for tissue construction (Fig. 1A).Anchors were printed into each well of the plate using silicone ink with detailed printing procedures, associated videos, and dimensional information provided in Fig. S1 and Video S1. Subsequently, C2C12 myoblasts were encapsulated within neutralized 0.5 wt% collagen and bioprinted along the inner circumference of each well to form circular constructs. The corresponding bioprinting process is shown in Video S2. Following printing, the constructs were cultured until mature muscle bundles were formed (Fig. 1B).
To improve reproducibility and throughput of 3D skeletal muscle (SKM) constructs, three optimization strategies were implemented: (i) optimization of the anchor geometry, (ii) modulation of the SKM construct shape, and (iii) optimization of the 3D SKM fabrication process (Fig. 1C). Collectively, these advances enabled a robust, reproducible high-throughput 3D SKM platform, which was subsequently validated for its applicability in drug screening (Fig. 1D).
2. Comparison of the 3D skeletal muscle tissue morphology and stability according to the anchor geometry
The anchor is an essential component that maintains the overall shape of the skeletal muscle (SKM) tissue and supports cell alignment and rapid contraction during 3D SKM culture. In particular, the anchor is widely recognized as a key element in 3D muscle tissue engineering, as it forms a tendon-like structure that provides uniaxial tension necessary for muscle microtissue formation, compaction, and maturation.21,31 Depending on the research objectives, various types of anchors have been employed in muscle tissue engineering.20,32,33However, the design selection and integration of anchors within different platforms require careful optimization. For instance, previous studies have reported the use of anchors within microfluidic chips, in which a single PDMS-based chip containing anchors was fabricated, and cell-laden hydrogels were injected into enclosed microchambers, leading to the formation of muscle tissue through gradual compaction over time.22,34 Although this enclosed platform provided a stable environment that effectively maintained muscle tissue, it posed a risk of tissue damage during retrieval, limiting its applicability for downstream analyses to evaluate muscle function and physiology. Conversely, in open-top systems, tissues are highly prone to deformation because they are not physically restrained against floating or distortion during culture. One study reported the use of inclined anchors to stabilize the muscle structure during culture; however, this approach required a high level of technical handling, involved multiple molding steps, and was time-consuming to prepare the entire muscle tissue platform.21
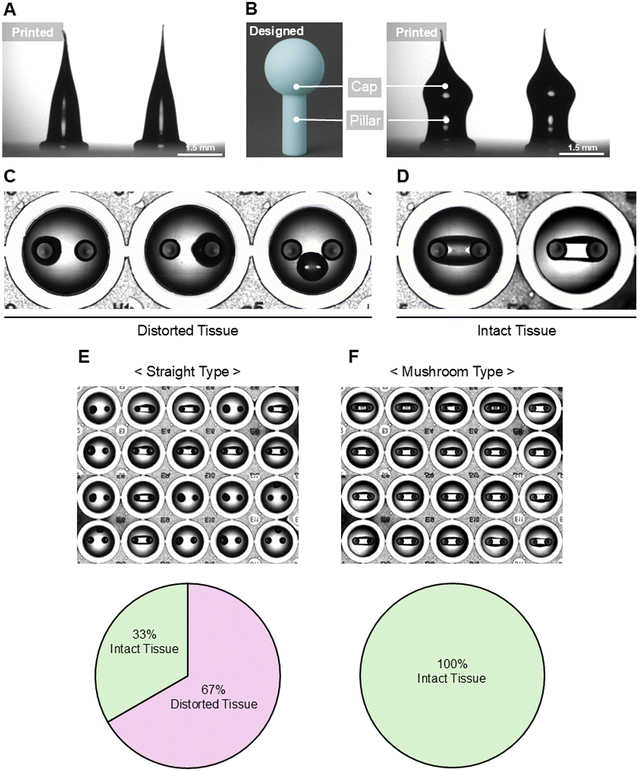
To overcome these limitations in anchor fabrication and platform integration, we utilized the rapid prototyping capability of 3D printing to introduce a mushroom-shaped pin anchor by incorporating a brief printing delay during the layer-stacking process. Although anchors fabricated with printing delays of 1.0 s and 1.5 s appeared to form cap-like structures in representative images, muscle tissue formation experiments using anchors fabricated with a 1.0 s time delay revealed the presence of wells in which failure tissues were generated. In contrast, anchors fabricated with a 1.5 s time delay enabled successful muscle tissue formation in all 12 wells. However, during fabrication across an entire 96-well plate, time-dependent changes in the rheological properties of the silicone ink resulted in a gradual increase in viscosity, leading to reduced extrusion volume even under identical pneumatic pressure. As printing the full plate requires approximately 20 min, this time-dependent viscosity increase caused inconsistent anchor geometries between the first and last wells, particularly due to insufficient material extrusion in later wells, which could compromise reliable formation of the cap-shaped structure.
Therefore, a printing delay of 2 s was selected to ensure stable and uniform fabrication of mushroom-type pin anchors across all wells of the 96-well plate. Under these conditions, the ratio of pillar width to cap width was maintained at approximately 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.35–1
1.35–1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.5. This transient printing delay generated a cap-shaped structure at the tip of the anchor, providing a mechanical interlocking effect that enhanced tissue retention and structural stability (Fig. 2B). The optimization process of the cap-shaped anchor size by adjusting the printing time delay is provided in Fig. S2.
1.5. This transient printing delay generated a cap-shaped structure at the tip of the anchor, providing a mechanical interlocking effect that enhanced tissue retention and structural stability (Fig. 2B). The optimization process of the cap-shaped anchor size by adjusting the printing time delay is provided in Fig. S2.
When SKM tissues were cultured using conventional straight-type pin anchors (Fig. 2A), excessive contraction often led to tissue slippage and detachment from the anchor tips, producing distorted and fragmented constructs (Fig. 2C and E). Such deformation not only disrupted tissue uniformity but also impaired alignment and functional maturation. By contrast, the use of the newly developed mushroom-type pin anchors effectively prevented detachment throughout the culture period. All printed tissues maintained an intact morphology and high structural stability, resulting in reproducible and mechanically robust 3D SKM constructs (Fig. 2D and F).
The fabrication of mushroom-type pin anchors was highly efficient, requiring only ∼20 minutes to fabricate a single AnchorPlate96 (one 96-well plate). This rapid and scalable process enables the production of large numbers of mechanically stable and morphologically uniform SKM tissues, demonstrating its suitability for high-throughput applications in tissue engineering and drug screening.
3. Enhancement of morphological and functional properties of 3D muscle tissues through amphiphilic coating and construct shape optimization
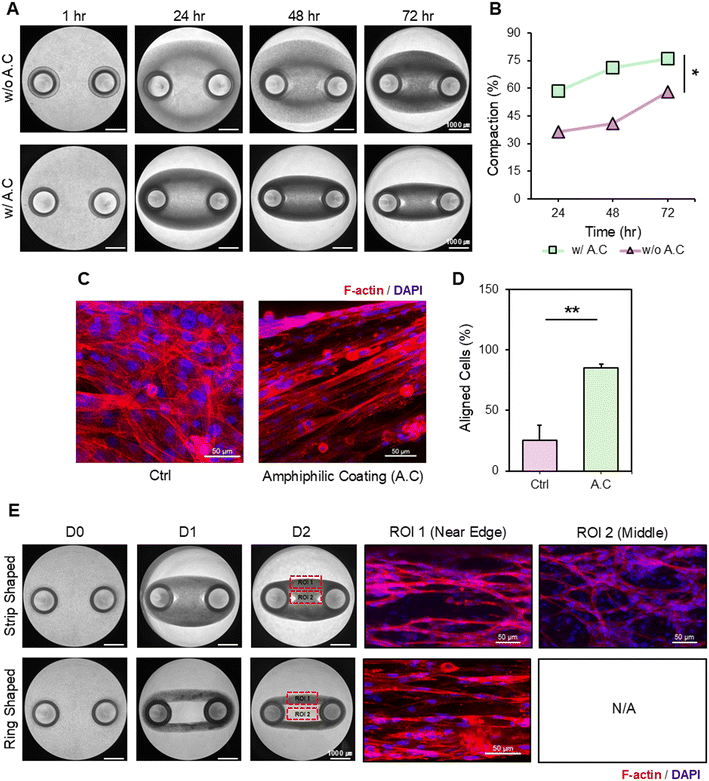
Before muscle tissue culture, an amphiphilic coating (A.C.) using 4% Pluronic F-127 was applied to the wells to prevent cell adhesion to the polystyrene surface. This treatment promoted cell alignment and facilitated rapid contraction and the formation of mature muscle bundles.23,35 It is noteworthy that, despite the use of non-treated 96-well plates to minimize excessive adhesion, both cells and the ECM could still adhere to the polystyrene surface—the primary material of the plate—which can hinder effective cell-mediated compaction. To evaluate the effect of A.C., the contraction rate of tissues was compared with that of the uncoated control. As expected, constructs cultured with amphiphilic coating exhibited an accelerated contraction rate compared to the uncoated control (Fig. 3A and B). In addition to faster tissue compaction, A.C. significantly enhanced cell alignment along the myotube axis induced by uniaxial tension, whereas randomly distributed myoblasts were observed in the control group during culture. The images and quantitative graphs demonstrated that the A.C. group exhibited improved myotube alignment (Fig. 3C and D).To further investigate the morphological determinants of cell alignment, bioink containing C2C12 cells was manually pipetted into 96-well plates equipped with printed anchors. Although the same amount of hydrogel was dispensed into each well, manual pipetting led to operator-dependent variability, producing constructs with heterogeneous geometries such as strip- and ring-shaped structures. The alignment characteristics between these two geometries were then compared. In the region near the edge (ROI 1), both strip- and ring-shaped tissues showed well-aligned myotubes in one direction. However, in the central region (ROI 2), strip-shaped tissues exhibited an irregular, random alignment rather than a consistent unidirectional orientation (Fig. 3E). While both groups appeared similar near the edge, a clear difference was observed when comparing the middle regions, indicating lower structural uniformity in the strip-shaped group.
Based on these findings, the ring-shaped structure was selected for subsequent experiments, as it exhibited higher internal uniformity within a single tissue and more consistent unidirectional myotube alignment (Fig. S5). Therefore, the combination of the amphiphilic coating and optimized construct geometry provides an effective strategy to enhance the morphological integrity of 3D skeletal muscle tissues, thereby improving their reliability for downstream applications.
4. Comparison of embedding methods for muscle cell-laden bioink in 96-well plates
Prior to comparing the pipetting- and bioprinting-based embedding approaches, the optimal bioink volume was determined. Bioink volumes of 20–25 μL, corresponding to pneumatic pressures of 10–15 kPa, were identified as optimal conditions for stable tissue fabrication. When smaller volumes below 25 μL (≤10 kPa) were used, even with repetitive tapping to promote spreading, the bioink was not uniformly distributed across the well bottom, often resulting in incomplete tissue formation or excessively thin tissues that were prone to breakage during culture. In contrast, when larger volumes exceeding 25 μL (≥20 kPa) were applied, excessive bioink deposition frequently resulted in unintended strip-shaped constructs, accompanied by the absence of muscle contraction and impaired cell alignment. Therefore, a bioink volume of 20–25 μL was selected as the optimal condition and applied consistently to both the pipetting-based and bioprinting-based groups.As demonstrated in the previous results, manual bioink injection via pipetting inevitably introduces multiple sources of variability, including pipetting errors, bubble formation, operator-to-operator variability, and differences in technical skill (Fig. S4).
To overcome these limitations, a 3D bioprinter was employed to inject bioink containing muscle cells. In previous studies, manual pipetting was frequently used for bioink dispensing; however, when embedding was performed across the entire 96-well plate using this approach, the tissues within each well exhibited inconsistent geometries due to unintended post-dispensing states of the bioink (Fig. S4). Even though repetitive tapping was applied to distribute the bioink evenly at the bottom of the wells, as observed across three whole plates, bioink that appeared uniform to the naked eye frequently resulted in failed tissues exhibiting slipping-off, floating-off, or strip-shaped morphologies during culture. Consistent with this observation, a comparison of red- and green-colored wells revealed a markedly higher proportion of red-colored wells in this group (Fig. 4A), indicating failed and successful tissues, respectively.
In contrast, as observed across three whole plates, the bioprinting-based approach consistently generated tissues that maintained the intended ring-shaped geometry throughout the culture period, preserving stable tissue structures. Furthermore, comparison of red- and green-colored wells revealed that all wells, except for a single well, were green-colored in this group, indicating successful tissue formation (Fig. 4B).
Taken together, these findings indicate that platforms fabricated via manual pipetting tend to produce undesired and non-uniform tissues, limiting their suitability for true high-throughput applications. Conversely, 3D bioprinting-based embedding enables reproducible fabrication of tissues with well-defined geometrical consistency across most wells, demonstrating its suitability for the reliable establishment of a high-throughput skeletal muscle tissue platform.
5. Uniformity of 3D muscle tissues depending on the embedding method of muscle cell-laden bioink
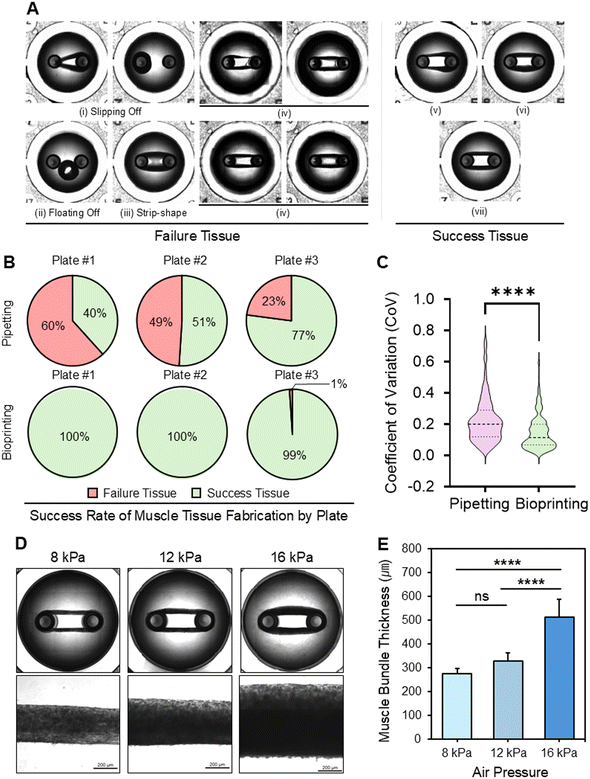
The morphology of tissues generated in each well of the 3D SKM tissue platform varied depending on the embedding method employed. Constructs exhibiting a strip-shaped tissue morphology, tissues that failed to remain positioned on the bottom of the well due to slipping-off or floating-off, and tissues in which a complete ring-shaped structure was not properly formed were collectively defined as failure tissues (Fig. 5A(i–iv)). In contrast, constructs that consistently retained a well-defined ring shape and exhibited a uniform morphology were classified as success tissues (Fig. 5A(v–vii)).Across three pipetting-based plates, the success rate of tissue formation was markedly lower than that observed in the bioprinting-based plates. In contrast, across three bioprinting-based plates, nearly all wells generated successful tissues, demonstrating a notable improvement in reproducibility and structural stability (Fig. 5B).
As shown in the CoV distribution plot, a wider horizontal spread indicates a higher density of values within the corresponding range. Comparison between the two groups demonstrates that embedding via 3D bioprinting markedly improves construct uniformity compared with manual pipetting (Fig. 5C). Furthermore, the 3D bioprinter-based embedding method allowed researchers to fabricate muscle bundles with diverse thicknesses by modulating pneumatic pressure during printing, providing an additional advantage for producing constructs with controlled morphological characteristics tailored to experimental requirements (Fig. 5D and E).
Taken together, these findings demonstrate that 3D bioprinter-assisted embedding not only improves the reproducibility and uniformity of muscle tissue formation but also enables precise control over the construct morphology, thereby enhancing the utility and scalability of the platform for high-throughput applications.
6. Morphological and molecular maturation of 3D muscle tissues
C2C12 cells mixed with hydrogels to prepare the bioink underwent proliferation and compaction, followed by differentiation, during which cells fused to form myotubes (Fig. 6A).36 Using optimized fabrication conditions established through the three strategies described above, the resulting 3D SKM tissues exhibited high uniformity and developed physiologically relevant alignment of myotubes upon completion of differentiation (Fig. 6B).Furthermore, confocal immunofluorescence imaging revealed progressively enhanced alignment and multinucleated myotube formation over time, indicating continuous maturation (Fig. 6C). The emergence of mature muscle bundles was further supported by increases in both myotube length and the fusion index (Fig. 6D and E).
Collectively, these results indicate that the optimized fabrication strategy enables the generation of 3D skeletal muscle tissues exhibiting morphological features and molecular markers associated with myogenic maturation, thereby establishing a robust and reproducible platform for functional muscle studies.
7. Proof-of-concept application of the fabricated high-throughput 3D skeletal muscle tissue plate
Using the stable and mature 3D SKM tissues produced in this study, we performed drug testing to validate the functionality and applicability of the high-throughput platform.21,37,38 Four drugs known to exert negative or positive effects on muscle physiology were selected: paclitaxel,39,40 cisplatin,41,42 and dexamethasone21,43,44 (which induce muscle atrophy and functional decline), and insulin-like growth factor-1 (IGF-1)21,44 (which promotes myotube maturation and myofiber development). It is well established that off-target effects of chemotherapeutic drugs can lead to skeletal muscle loss.45,46 Among various chemotherapeutic agents, paclitaxel and cisplatin were selected in this study. Paclitaxel has been reported to inhibit myotube differentiation and to shorten elongated myotubes.40 To investigate whether these two drugs also induce skeletal muscle loss, they were applied as model chemotherapeutic agents to the 3D skeletal muscle platform developed in this study. The selected drug concentrations were based on previously reported studies and were chosen to remain below the IC50 values.42,47 In addition, the effects of dexamethasone and IGF-1 on skeletal muscle have been well documented in prior studies, and the concentrations used in this study were likewise selected based on previously reported concentrations or values below the IC50.21,44 All four drugs were tested simultaneously at three different concentrations along with a control group. To support the applicability of this platform for drug screening, experiments were conducted to evaluate the cytotoxic effects of each drug on muscle tissues. The cell viability of drug-treated tissues (n = 12 per group) was assessed on day 1 for paclitaxel and cisplatin, on day 3 for dexamethasone, and on day 5 for IGF-1 (Fig. 7B). The drug treatment experiments were conducted according to the timeline shown in Fig. 7A.To assess drug responses using our fabricated high-throughput system plate, tissues were stained for MHC and DAPI, and the myotube morphology was examined by confocal microscopy. For paclitaxel, higher concentrations yielded an increased number of rounded cells rather than elongated, laterally aligned myotubes, accompanied by a decrease in multinucleated fibers (Fig. 7C(i)). For cisplatin, although visually less striking than the paclitaxel and dexamethasone groups, a dose-dependent reduction in the number of multinucleated myotubes was observed, and myotubes appeared as separated structures with a single nucleus rather than aligned multinucleated fibers (Fig. 7C(ii)). In the dexamethasone group, increasing concentration led to thinner myotubes, a decrease in multinucleated myotubes, and a reduction in unidirectionally aligned myotubes, which appeared more separated (Fig. 7C(iii)). Overall, groups treated with negatively acting drugs commonly exhibited detrimental effects on multinucleated myotube formation, thickness, and alignment.
By contrast, samples treated with IGF-1 showed a dose-dependent increase in myotube thickness compared with the control. Alignment between myotubes also improved, and long, laterally oriented, mature multinucleated myotubes were observed (Fig. 7C(iv)).
Collectively, four drugs were tested across four groups per drug (n = 5 per group; total = 80 samples), demonstrating that the high-throughput system developed in this study enables simultaneous, single-run screening of a large number of samples and compounds. Furthermore, these results confirm that the high-throughput 3D SKM tissue platform successfully recapitulates differential drug responses, validating its utility as a proof-of-concept system for drug screening and establishing a robust foundation for future therapeutic development targeting muscle-related diseases.
The fabrication strategy that integrates 3D cell printing using a 3D bioprinter with commercially available multi-well plates effectively overcomes the limitations of conventional PDMS- or mold-based platforms, which typically require considerable time and multiple complex fabrication steps to produce a single chip. Moreover, we developed a collaborative approach that combines 3D bioprinting technology with three optimization strategies, enabling the reproducible fabrication of skeletal muscle tissues with high uniformity, accuracy, and efficiency. Unlike PDMS-based chips, the use of multi-well plates provides direct compatibility with high-throughput imaging systems, allowing evaluation of the physiological quality of all skeletal muscle tissues across an entire plate rather than a single sample. In addition, this platform can be adapted to various multi-well plate formats, enabling the simultaneous acquisition of large-scale experimental datasets within a single run.
Furthermore, the platform can be applied to various multi-well formats besides the 96-well plate, allowing scalable production of skeletal muscle tissues with different dimensions and increased sample throughput.
Looking forward, integration with automated systems such as robotic arms for routine operations—including media exchange and sample washing—could establish a fully automated high-throughput workflow, further enhancing the efficiency, productivity, and simplicity of skeletal muscle tissue fabrication.48 However, the current multi-well plates made of polystyrene may impose certain limitations on achieving high-resolution imaging due to their optical properties. To address this, employing glass-bottom multi-well plates for platform fabrication could yield higher-quality images and enable more precise analysis of the muscle tissue morphology and function.49 In addition, the 3D skeletal muscle (SKM) tissues fabricated in this study exhibited morphological and molecular features associated with myogenic maturation, including increased myotube length and fusion index. However, direct functional assessment of tissue contractility was not performed. In future studies, the proposed platform may be extended to evaluate muscle contractile activity by measuring calcium transients and quantifying ΔF/F0.21,50
Besides these immediate improvements, the high-throughput platform developed in this study—driven by the combination of multiple optimization strategies and 3D bioprinting technology—holds broad potential for diverse biomedical applications. These include accelerating drug discovery, providing physiologically relevant models for muscle disease research, and serving as advanced in vitro testing systems that can complement or even replace conventional preclinical assays.
Conclusions
In this study, we established a streamlined 3D cell-printing strategy using a 3D bioprinter, which reduced fabrication time by minimizing process steps and improved accessibility through the use of a commercially available multi-well plate. While the straight-type pin anchors exhibited low success rates in tissue fabrication, the mushroom-type pin anchors produced with a one-second printing delay achieved ∼100% fabrication success, yielding constructs with an intact morphology. Furthermore, amphiphilic coating combined with ring-shaped constructs provided optimized conditions for generating highly aligned and matured myotubes, thereby enhancing the physiological relevance of engineered skeletal muscle tissues.In addition, 3D bioprinting enhanced uniformity not only within individual wells but also across entire plates, ensuring consistent muscle bundle thickness and morphology. Taken together, the integration of 3D bioprinting with the aforementioned optimization strategies demonstrates that the high-throughput 3D skeletal muscle tissue platform developed here represents a robust and scalable tool with significant potential for drug screening and the development of personalized therapeutics for muscle-related diseases.
Author contributions
Tae-Eun Lim: conceptualization, methodology, investigation, data curation, formal analysis, visualization, writing – original draft, writing – review & editing. Ashfaq Ahmad: conceptualization, methodology, investigation, formal analysis, visualization. Yeong-Jin Choi: writing – review & editing, supervision, funding acquisition, resources. Hee-Gyeong Yi: writing – review & editing, supervision, funding acquisition, resources, project administration.Conflicts of interest
There are no conflicts to declare.Data availability
The data related to this study can be available from the authors upon reasonable request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc01040c.
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grants funded by the Ministry of Science and ICT (MSIT) [No. 2020R1A5A8018367, No. RS-2022-NR067329, and No. RS-2024-00423107], the Korea Evaluation Institute of Industrial Technology (KEIT) funded by the Ministry of Trade, Industry and Energy (MOTIE) [No. 20012378], the Korea Health Industry Development Institute (KHIDI) funded by the Ministry of Health and Welfare [No. RS-2025-19252970], the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through the Ministry of Agriculture, Food and Rural Affairs (MAFRA) [No. RS-2024-00397026], the “Regional Innovation System & Education (RISE)” through the Gwangju RISE Center, funded by the Ministry of Education (MOE) and the Gwangju Metropolitan Government, Republic of Korea [2025-RISE-05-011], and the Fundamental Research Program of the Korea Institute of Material Science [PNKB020].References
- C. McCuller, R. Jessu and A. L. Callahan, in StatPearls, StatPearls Publishing, 2023 Search PubMed.
- S. J. Valberg, Clin. Biochem. Domest. Anim., 2008, 6, 459–484 CrossRef.
- T. C. Gamboa-Martínez, V. Luque-Guillén, C. González-García, J. L. Gómez Ribelles and G. Gallego-Ferrer, J. Biomed. Mater. Res., Part A, 2015, 103, 614–621 CrossRef PubMed.
- J. Simoneau and C. Bouchard, FASEB J., 1995, 9, 1091–1095 CrossRef CAS PubMed.
- T. Osaki, S. G. Uzel and R. D. Kamm, Nat. Protoc., 2020, 15, 421–449 CrossRef CAS PubMed.
- H. Nagy and K. D. Veerapaneni, in StatPearls, StatPearls Publishing, 2023 Search PubMed.
- Myo-MOVES: a custom electrical stimulation system for functional studies of 3D bioengineered muscle - Lab on a Chip (RSC Publishing), https://pubs.rsc.org/en/content/articlelanding/2025/lc/d5lc00614g, (accessed October 9, 2025).
- D. G. Nair and R. Weiskirchen, Curr. Issues Mol. Biol., 2025, 47, 7 CrossRef CAS PubMed.
- Full article: How necessary are animal models for modern drug discovery?, https://www.tandfonline.com/doi/full/10.1080/17460441.2021.1972255, (accessed October 9, 2025).
- E. J. Stevenson, A. Koncarevic, P. G. Giresi, R. W. Jackman and S. C. Kandarian, J. Appl. Physiol., 2005, 98, 1396–1406 CrossRef CAS PubMed.
- J. Von Maltzahn, J.-M. Renaud, G. Parise and M. A. Rudnicki, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 20614–20619 CrossRef CAS PubMed.
- D. B. Thomason and F. W. Booth, J. Appl. Physiol., 1990, 68, 1–12 CrossRef CAS PubMed.
- F. P. Gaschen, E. P. Hoffman, J. R. M. Gorospe, E. W. Uhl, D. F. Senior, G. H. Cardinet III and L. K. Pearce, J. Neurol. Sci., 1992, 110, 149–159 CrossRef CAS PubMed.
- J. W. McGreevy, C. H. Hakim, M. A. McIntosh and D. Duan, Dis. Models Mech., 2015, 8, 195–213 CrossRef CAS PubMed.
- J. A. DiMasi, R. W. Hansen and H. G. Grabowski, J. Health Econ., 2003, 22, 151–185 CrossRef PubMed.
- L. Polonchuk, M. Chabria, L. Badi, J.-C. Hoflack, G. Figtree, M. J. Davies and C. Gentile, Sci. Rep., 2017, 7, 7005 CrossRef PubMed.
- R. J. Wall and M. Shani, Theriogenology, 2008, 69, 2–9 CrossRef CAS PubMed.
- S. Y. Park, H. J. Hong and H. J. Lee, BioChip J., 2023, 17, 24–43 CrossRef CAS.
- R. Driver and S. Mishra, BioChip J., 2023, 17, 1–23 CrossRef CAS.
- Y. Choi, T. G. Kim, J. Jeong, H. Yi, J. W. Park, W. Hwang and D. Cho, Adv. Healthcare Mater., 2016, 5, 2636–2645 CrossRef CAS PubMed.
- M. E. Afshar, H. Y. Abraha, M. A. Bakooshli, S. Davoudi, N. Thavandiran, K. Tung, H. Ahn, H. J. Ginsberg, P. W. Zandstra and P. M. Gilbert, Sci. Rep., 2020, 10, 6918 Search PubMed.
- J. Kim, I. U. Kim, Z. F. Lee, G. Sim and J. S. Jeon, Adv. Funct. Mater., 2024, 34, 2410872 CrossRef CAS.
- T. Osaki, S. G. M. Uzel and R. D. Kamm, Sci. Adv., 2018, 4, eaat5847 Search PubMed.
- A. Ahmad, S.-J. Kim, Y.-J. Jeong, M. S. Khan, J. Park, D.-W. Lee, C. Lee, Y.-J. Choi and H.-G. Yi, J. Mater. Chem. B, 2024, 12, 8633–8646 RSC.
- Y.-J. Choi, Y.-J. Jun, D. Y. Kim, H.-G. Yi, S.-H. Chae, J. Kang, J. Lee, G. Gao, J.-S. Kong and J. Jang, Biomaterials, 2019, 206, 160–169 CrossRef CAS PubMed.
- M. Samandari, J. Quint, A. Rodríguez-delaRosa, I. Sinha, O. Pourquié and A. Tamayol, Adv. Mater., 2022, 34, 2105883 CrossRef CAS PubMed.
- C. Blake, O. Massey, M. Boyd-Moss, K. Firipis, A. Rifai, S. Franks, A. Quigley, R. Kapsa, D. R. Nisbet and R. J. Williams, APL Bioeng., 2021 DOI:10.1063/5.0040764.
- K. Fakhruddin, M. S. A. Hamzah and S. I. A. Razak, IOP Conf. Ser.:Mater. Sci. Eng., 2018, 440, 012042 Search PubMed.
- J. H. Y. Chung, S. Naficy, Z. Yue, R. Kapsa, A. Quigley, S. E. Moulton and G. G. Wallace, Biomater. Sci., 2013, 1, 763–773 RSC.
- J. Son, S. J. Hong, J. W. Lim, W. Jeong, J. H. Jeong and H.-W. Kang, Small Methods, 2021, 5, 2100632 CrossRef CAS PubMed.
- H. H. Vandenburgh, P. Karlisch and L. Farr, In Vitro Cell. Dev. Biol., 1988, 24, 166–174 CrossRef CAS PubMed.
- B. Jo, K. Motoi, Y. Morimoto and S. Takeuchi, Adv. Healthcare Mater., 2024, 13, 2401844 CrossRef CAS PubMed.
- Simulation and Fabrication of Stronger, Larger, and Faster Walking Biohybrid Machines - Pagan-Diaz - 2018 - Advanced Functional Materials - Wiley Online Library, https://advanced.onlinelibrary.wiley.com/doi/full/10.1002/adfm.201801145, (accessed October 19, 2025).
- Z. Roveimiab, F. Lin and J. E. Anderson, Tissue Eng., Part B, 2019, 25, 30–45 CrossRef PubMed.
- Leaf-venation-directed cellular alignment for macroscale cardiac constructs with tissue-like functionalities | Nature Communications, https://www.nature.com/articles/s41467-023-37716-1, (accessed October 8, 2025).
- F. Velasco-Mallorquí, J. M. Fernández-Costa, L. Neves and J. Ramón-Azcón, Nanoscale Adv., 2020, 2, 2885–2896 RSC.
- S. K. A. S. Elfaiomy, S. E. A. Elnasr, A. A. Kassab and T. A. E. Eldeeb, Tanta Med. J., 2025, 53, 129 CrossRef.
- K. Huang, Y. Chiang, M. Ali and S. Hsia, J. Cachexia Sarcopenia Muscle, 2025, 16, e13817 Search PubMed.
- LKB1 Destabilizes Microtubules in Myoblasts and Contributes to Myoblast Differentiation | PLOS One, https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0031583, (accessed October 19, 2025).
- P. B. Antin, S. Forry-Schaudies, T. M. Friedman, S. J. Tapscott and H. Holtzer, J. Cell Biol., 1981, 90, 300–308 CrossRef CAS PubMed.
- Taurine rescues cisplatin-induced muscle atrophy in vitro: a morphological study - PubMed, https://pubmed.ncbi.nlm.nih.gov/24955211/, (accessed October 19, 2025).
- O. Hairuddin, B. H. Yahaya, M. Ibahim, A. Verakumarasivam, C. Choy, M. Mazlan, N. Rahim, S. Dzulkarnain and S. Mansor, Biomed. Res. Ther., 2023, 10, 5810–5830 Search PubMed.
- D.-S. Han, W.-S. Yang and T.-W. Kao, Int. J. Med. Sci., 2017, 14, 434–443 CrossRef CAS PubMed.
- A. Brun, G. Mougeot, P. Denis, M. L. Collin, P. Pouchin, C. Montaurier, S. Walrand, F. Capel and M. Gueugneau, Sci. Rep., 2024, 14, 3108 Search PubMed.
- P. Zhong, X. Li and J. Li, Front. Oncol., 2025, 15, 1551561 CrossRef CAS PubMed.
- Current Opinion in Supportive and Palliative Care, https://journals.lww.com/co-supportiveandpalliativecare/fulltext/2018/12000/preservation_of_muscle_mass_as_a_strategy_to.6.aspx, (accessed January 22, 2026).
- A. Rabzia, M. Khazaei, Z. Rashidi and M. R. Khazaei, Iran. J. Pharm. Res., 2017, 16, 1432–1442 Search PubMed.
- A. Georgescu, J. H. Oved, J. H. Galarraga, T. Cantrell, S. Mehta, B. M. Dulmovits, T. S. Olson, P. Fattahi, A. Wang, P. L. Candarlioglu, A. Muvaffak, M. M. Kim, S. A. Aydin, J. Seo, E. S. Diffenderfer, A. Lynch, G. S. Worthen and D. D. Huh, Cell Stem Cell, 2024, 31, 1847–1864.e6 Search PubMed.
- S. Rajasekar, D. S. Y. Lin, L. Abdul, A. Liu, A. Sotra, F. Zhang and B. Zhang, Adv. Mater., 2020, 32, 2002974 CrossRef CAS PubMed.
- Bioengineered human myobundles mimic clinical responses of skeletal muscle to drugs | eLife, https://elifesciences.org/articles/04885, (accessed January 21, 2026).
| This journal is © The Royal Society of Chemistry 2026 |