 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Dual vision-equipped microfluidic chip for spatiotemporal sequential pick-and-place of oocytes
Shuzhang
Liang
 ,
Hao
Mo
,
Yuguo
Dai
,
Hirotaka
Sugiura
,
Satoshi
Amaya
,
Hao
Mo
,
Yuguo
Dai
,
Hirotaka
Sugiura
,
Satoshi
Amaya
 and
Fumihito
Arai
*
and
Fumihito
Arai
*
Department of Mechanical Engineering, School of Engineering, The University of Tokyo, Tokyo 113-8654, Japan. E-mail: arai-fumihito@g.ecc.u-tokyo.ac.jp
First published on 27th February 2026
Abstract
Oocytes have long been used as a fundamental biological model in bioengineering research such as gene expression analysis, electrophysiological measurements, and drug screening. However, current methods mainly focus on single-microscope setups, which limits the efficient manipulation of multiple oocytes across spatially separated worksites. This study presents a dual vision-equipped microfluidic chip for the manipulation of multiple oocytes between different worksites. The microfluidic chip is equipped with two miniature cameras and then installed on a robotic manipulator. One miniature camera is used to track the position of oocytes in the microfluidic chip. The vision position of the multiple oocytes is utilized to control the flow. The results showed that multiple objects were successfully separated and released in sequence only using the hydrodynamic flow focusing effect. Moreover, a well port is designed to trap single oocytes to deal with the unseparated case of neighboring oocytes in the microchannel based on the vision information. Subsequently, the other camera is installed on the top of the tip part and utilized to detect the single object picking–placing position. Finally, we demonstrate that the dual vision-equipped microfluidic chip on-robot can pick, transport, and place multiple oocytes between different well chip areas. The proposed method has application potential in oocyte biomedical engineering.
1 Introduction
Oocytes have long been used as a significant biological model in bioengineering research and applications such as gene expression analysis, electrophysiological measurements, and drug screening.1,2 Generally, in biology engineering experiments, multiple microscopic worksites are required for different stages of oocyte manipulation due to specific experimental needs.3,4 For example, in the two-electrode voltage clamp (TEVC) process of Xenopus oocytes,3,5 the TEVC plays an important role in AI-driven scientific research and novel compound screening, providing a reliable platform for assessing the efficacy, selectivity, and toxicity of candidate molecules targeting ion channels, transporters, and membrane receptors. Thus, high-quality oocytes are first selected under one microscope; then RNA microinjection is performed at another worksite; subsequently, the oocytes are transported for incubation overnight; finally, electrophysiological measurements are carried out under another microscope. During this multi-step process, repeated operations of picking, transporting, and placing multiple oocytes between different worksites are required. Moreover, since each oocyte has unique experimental performance,6,7 maintaining the order of oocytes during transport is crucial for accurate data recording.Although glass pipettes, combined with fluidic,8 electric,9 or acoustic manipulation techniques,10 have been widely applied for oocyte manipulation,11 such methods suffer from limited efficiency because only one oocyte can be transported at a time. If multiple oocytes are aspirated into glass pipettes, sequence information among them is lost. Recently, microfluidic tools, used as an end effector of robotic manipulators,12,13 have been proposed for manipulation of oocytes.14,15 These microfluidic systems can efficiently control the order of multiple oocytes. Moreover, a capacitance sensor is utilized to detect the oocyte manipulation status inside the channel. With this method, the separation success rate for single oocyte placing is around 87% at one time. Nonetheless, in cases of unsuccessful isolation, the process must be repeated multiple times until successful separation occurs, causing time-consuming and challenging oocyte detection using the capacitance sensor. This limitation arises because capacitance sensors can only capture a single-point electrical signal in the microchannel,14,16 thus failing to provide positional information of multiple oocytes and causing misjudgment in control. Furthermore, it is difficult to monitor oocyte positions in the channel over a long distance between different worksites using the capacitance sensor. To ensure that the compiler continues system operation, such error cases of unsuccessful separation and object positions must be promptly detected and properly handled.
The use of visual sensors has emerged as the most effective method for detecting and manipulating both macro- and micro-scale objects, since they provide accurate morphological and spatial information of multiple targets.17,18 Based on the target image information, system states and error can be analyzed.19 Owing to the benefits of vision-based control, microfluidic systems primarily utilize visual imaging to monitor fluids and particles/cells within microchannels, track object states, and offer real-time feedback for automated or machine learning-driven control.17,20 For instance, W. He et al. developed a neuromorphic-enabled, video-activated framework capable of high-dimensional spatiotemporal characterization for real-time particle sorting in microfluidic chips.21 T. Aoyama et al. utilized real-time vision imaging to extract multi-object moment features for microflow-rate regulation in cellular analysis.22 A. Mudugamuwa et al. designed an active droplet generation platform that employed visual data to determine droplet diameters.23 In general, these visual imaging techniques include optical bright-field microscopy, fluorescence microscopy, confocal laser scanning microscopy, or another microscopy technique.18,24 Specifically, high-resolution CCD (charge coupled device) or CMOS (complementary metal oxide semiconductor) cameras are mounted on bulky microscopes, and functional microfluidic chips are fixed on the limited microscope stage. This configuration has facilitated significant advances in life sciences such as cell motion analysis.
However, this conventional configuration strategy of the vision microscope and microfluidic chip system is not suitable for manipulating multiple oocytes between different worksites due to contradiction between the limited field-of-view under a microscope and the need for large-space microfluidic chip movement. When the microfluidic chip moves from one microscope to another microscope worksite, the position information of oocytes inside the microchannel cannot be observed using the camera on the microscope, causing control mistakes. Therefore, it is necessary to design a system strategy to monitor multi-oocyte information in the microchannel for picking–transporting–placing within a large space.
In this study, we report a new dual vision-equipped microfluidic chip strategy for assisting multi-single oocyte picking–transporting–placing in sequence. The strategy directly installs two miniature cameras on the microfluidic chip, which is an end-effector of a robotic manipulator (called chip-on-robot), as shown in Fig. 1(a). One miniature camera, mounted above the front part of the microchannel, continuously monitors the positions of multiple oocytes within the chip, as shown in Fig. 1(b). The vision position of the multiple oocytes is utilized to control the hydrodynamic flow. The flow effect is analyzed by simulation and applied for isolating oocytes by placing only a single oocyte into a specific well in sequence. Moreover, a well port is designed to trap single oocytes to deal with the unseparated case of neighboring oocytes in the microchannel. Subsequently, the other camera is installed on the top of the tip part and utilized to detect the single oocyte picking–placing position, as shown in Fig. 1(b). Finally, we demonstrate that the dual vision-equipped microfluidic chip on-robot can pick, transport, and place multiple oocytes between different well chip worksites.
Unlike previous vision-based microfluidic systems that rely on stationary microscope platforms, or robotic micromanipulation systems that use external imaging, this work introduces a chip-on-robot architecture integrating dual miniature cameras directly on the microfluidic end-effector, enabling continuous in-channel multi-oocyte monitoring and sequential manipulation between different worksites. The contributions of this work are as follows: (1) dual vision-equipped microfluidic chip extracting multi-oocyte positions for the flow control strategy; (2) well port-assisted fishbone-like microfluidic chip for single oocyte isolation; (3) demonstrating the spatiotemporal sequential pick-transport-and-place of multiple oocytes between different well chips.
2 Materials and methods
In this section, we first present the overall system configuration, in which the microfluidic chip is integrated with two miniature vision cameras. Then, the design and fabrication of the microfluidic chip for manipulation of oocytes is detailed. Next, we prepared the visual detection model and Xenopus oocytes for experiments. Subsequently, the calibration of these two cameras for controlling flow of two pumps and a robotic manipulator is described. Finally, the algorithm control of these two visions is described respectively.2.1 System configuration with two cameras
To obtain more information of multiple oocytes within the microfluidic chip, a vision sensor is more suitable for monitoring than conventional capacitance-based detection.17,18,20,21 In traditional microscope manipulation, visual images are typically captured using a CCD camera mounted on a microscope or a fixed observation platform. However, since the camera and the microfluidic chip are physically independent, cell observation is lost once the chip is moved to another worksite. Here, we directly installed two miniature cameras with a diameter of 5.5 mm and a weight of 59 g (UC-02 Slim USB Camera, Nakabayashi Co., Ltd., Japan) into the microfluidic chip, as shown in Fig. 2(a). Based on the hand-eye visual servoing control system, where a camera fixes at the end of the manipulator to observe objects in real time, the coordination relationship is set as shown in Fig. 2(b) and (c). Then, we set up the miniature camera 1 above the tip region of the chip for obtaining the oocyte picking/placing well information, as shown in Fig. 2(d). On the other hand, to observe the oocyte information in the channel of the microfluidic chip, camera 2 is installed above the front part of the microchannel.The whole system configuration is shown Fig. 2(a). The microfluidic chip is equipped with two miniature cameras and then installed on a robotic manipulator (KWC06020, SURUGA SEIKI Co., Ltd., Japan). Two syringe pumps (KDS 230, KD Scientific Inc., USA) are connected to the microfluidic chip. One computer (i7-12700H central processing unit (CPU), RTX 3060 graphics card, and 32 GB of memory) is used for programming control of systems. The real image of the system is shown in Fig. S1.
2.2 Microfluidic chip design and fabrication
To pick–transport–place multiple oocytes, there are two requirement points. One is a pipette function for direct picking/placing single oocytes from/to the well chip or culture dish. The other is a separation function using a camera for sequential managing of single oocytes in the channel of the microfluidic chip. In this case, we design a well port-assisted fishbone-like microfluidic chip,14 as shown in Fig. 2(e). The microfluidic chip consists of a main channel, symmetrical bilateral branch channels, side channels, a trapping well port, a pipette tip, and two connected ports of pumps. In the microfluidic chip, oocytes are manipulated in the main channel or well port by controlling the flow with two pumps. With the fishbone-like structure, the flow is focused into the main channel from the branch channels. The branch-channel network behaves as a parallel structure. We treat each branch channel flow qi which is governed by branch resistance and coupling side resistance into the main channel, as shown in Fig. 2(e). For i branch nodes (numbered from the entrance), the side flow is lost section by section. Assume that the flow of each section of the main channel is Qi, where i = 1, 2, 3…, Q1 = Q0 + q1, and Qi+1 = Qi + qi+1. We set the i branch node pressure as pb,i, and the outlet tip pressure of the side branch channel as patm. The flow rate in the branch channel is determined by Poiseuille's law:25,26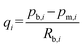 | (1) |
The flow rate between the two sections of the main channel is 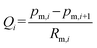 . Thus, we can obtain: pm,i+1 = pm,i − QiRm,i. The liquid resistance of each main channel is equal to Rm, and the fluid resistance of each pair of branch channels is a constant Rb. Since the resistance caused the pressure drop, more fluid enters the front part of the branch channel. Thus, q1 < q2 < q3 < … < qN. We can assume that: qi = q1 + (i − 1)·δqa, 1 < i < k, and qi = q1 + (i − 1)·δqb, k < i < n + 1, where, δqb > δqa > 0. Overall flow into the branch channel satisfies:
. Thus, we can obtain: pm,i+1 = pm,i − QiRm,i. The liquid resistance of each main channel is equal to Rm, and the fluid resistance of each pair of branch channels is a constant Rb. Since the resistance caused the pressure drop, more fluid enters the front part of the branch channel. Thus, q1 < q2 < q3 < … < qN. We can assume that: qi = q1 + (i − 1)·δqa, 1 < i < k, and qi = q1 + (i − 1)·δqb, k < i < n + 1, where, δqb > δqa > 0. Overall flow into the branch channel satisfies:
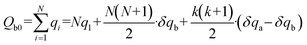 | (2) |
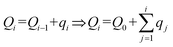 | (3) |
Therefore, the flow increases from the rear to the front of the channel. Because the main channel velocity 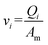 , the velocity increases toward the front. The flow velocity profile in the primary channel serves as the basis for oocyte separation. The oocyte suffers from the hydrodynamic force Fh (drag force, Fd) which is expressed as:27
, the velocity increases toward the front. The flow velocity profile in the primary channel serves as the basis for oocyte separation. The oocyte suffers from the hydrodynamic force Fh (drag force, Fd) which is expressed as:27
| Fh = −Fd = 6πμrV | (4) |
In addition, based on previous work,14 the separation success rate at one time does not reach 100% at real practical experiments and cases of unseparated oocytes frequently occur. Although unseparated oocytes can be finally separated by continuous repeated operation steps, the number of repetitions of flow control is uncertain, making the process time-consuming. Furthermore, since capacitance sensors only provide single-point electrical signals without more information, this can lead to misjudgments during multi-oocyte manipulation. Therefore, to efficiently solve the unseparated case, we design a trapping well port-assisted structure to trap single oocytes in the microfluidic chip, as shown in Fig. 2(e). The size of the well port depends on the diameter of single oocytes, as shown in Fig. S2. During the microfluidic chip releasing process, there are three cases of the oocyte position relationship, as shown in Fig. S3. In the separation case, oocytes are fully isolated within the channel. In unseparated cases, two situations arise. When only the first oocyte reaches the trapping well port position, pump 2 operates in withdrawal mode and the first oocyte is trapped within the well port, achieving single-oocyte isolation. When multiple oocytes move beyond the well port position, pump 2 withdraws and the oocyte nearest to the port is captured. In this case, although the order of the oocyte changes, visual images can still record the whole process and match information.
Subsequently, the fabrication requirements were determined based on the geometric dimensions of the microfluidic chip, and a 3D printing process was employed to achieve the desired precision. The overall workflow includes three-dimensional modeling, print parameter configuration, additive manufacturing, post-processing, and final device assembly. First, a three-dimensional structure was designed via CAD software and exported in STL format. For printing preparation, the digital model underwent slicing at a layer thickness of 25 μm. A high-temperature resistant resin was chosen owing to its physicochemical characteristics, which aid in clearing uncured materials from microstructures and reduce channel blockage risks. The sliced file was subsequently transferred to a Form 3 3D printer (Formlabs Inc., USA) for layer-by-layer fabrication. After printing, the residual resin was removed through 100 kHz ultrasonic cleaning for 10 minutes, followed by 15 minute UV curing to relieve internal stress and improve structural integrity. Finally, the microfluidic chip was sealed with a transparent adhesive film, as shown in Fig. S4, and two access ports were connected to flexible silicone tubing to complete the device assembly.
2.3 Oocyte preparation and visual detection model
Xenopus oocytes are utilized in the experiments. A representative image of the oocytes is presented in Fig. S5. These oocytes were in the growth phase,14,28 with an average diameter of approximately 1.2 mm. Healthy oocytes typically exhibit a distinct coloration pattern, appearing half dark brown and half white.29 The oocytes were maintained in Barth's buffer within 96 well plates and stored at 4 °C. Here, Barth's buffer is prepared and sterilized by high temperature as described in our previous work.14Since the oocyte has two colors on the surface and is managed between different scenes, the traditional detection method by image processing is difficult. For automated oocyte detection and manipulation, an image-based deep learning model was employed. YOLOv5 was selected for this task due to its wide adoption in object recognition,30 high accuracy, and ease of integration into external applications. Its compact structure also facilitates straightforward deployment. To train the YOLOv5 model for oocyte identification, an annotated dataset was constructed, consisting of 200 oocyte images allocated for both training and validation. The dataset was randomly split in an 8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio, resulting in 160 images for training and 40 images for validation. Moreover, the dataset included images captured under different spatial configurations corresponding to various manipulation stages. The training process was conducted for 300 epochs using the pretrained weights “yolov5s.pt” as the initialization model.
2 ratio, resulting in 160 images for training and 40 images for validation. Moreover, the dataset included images captured under different spatial configurations corresponding to various manipulation stages. The training process was conducted for 300 epochs using the pretrained weights “yolov5s.pt” as the initialization model.
For integration into the automatic control system, the trained YOLOv5 model was converted into a “.dll” library, enabling calls from the C# WinForm application. The model was further converted into an “.engine” format to enable TensorRT-based acceleration during inference. Image acquisition within the WinForm environment was achieved using OpenCVSharp. Additionally, to allow user-assisted calibration and control, a mouse-click function was implemented to select oocyte positions directly on the interface when necessary.
2.4 Coordination calibration with vision
In this system, the microfluidic chip is used as an end-effector of the robotic manipulator (chip-on-robot). Therefore, the microfluidic chip moves synchronously with the motion of the manipulator. Since the two cameras are fixed on the microfluidic chip, the two cameras move with the robotic manipulator. Here, camera 1 is applied to detect the target object out of the microfluidic chip for controlling the robotic manipulator. This configuration corresponds to an eye-in-hand system, in which the camera is mounted directly on the robot end-effector.31 To achieve accurate manipulation, an eye-in-hand calibration procedure is performed to determine the coordinate transformation between the camera's coordinate system and the robotic manipulator's coordinate system. Through this calibration, the camera coordinates of a target can be precisely mapped into the robot's spatial coordinate frame. The conversion process includes the camera part and the robot part. The camera part includes the GUI coordinate {G} and camera 1 coordinate {C1}. The robot part includes the manipulator coordinate system {M} and well chip coordinate system {W}, as shown in Fig. 2(b). In this study, a three-axis robotic micromanipulator was employed. For calibration simplification, the camera, manipulator, and well chip coordinate systems were manually adjusted to be parallel. Therefore, the relationship between the target well chip position and the manipulator position can be expressed as: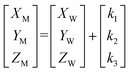 | (5) |
The relationship between the target and camera 1 is:
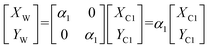 | (6) |
The conversion relationship between the camera and the GUI is:
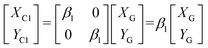 | (7) |
Due to camera 1 moving with the manipulator, it is necessary to record the home position between camera 1 and the manipulator. Thus, the position relationship between camera 1 and the manipulator is:
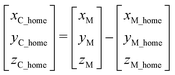 | (8) |
 | (9) |
 | (10) |
 | (11) |
On the other hand, camera 2, which is fixed above the microchannel, also requires calibration. When the camera is mounted on the microfluidic chip to monitor oocyte movement, the calibration process is analogous to that in micromanipulation systems. The essential objective is to convert the camera 2 coordinates {C2} in the captured image into physical microfluidic chip coordinates {F} within the microscope's field of view, thereby enabling accurate analysis of the oocyte's motion trajectory. Since the camera is fixed on the microfluidic chip, the coordinate transformation process mainly involves establishing a relationship between the camera coordinates of the image and the corresponding physical coordinates in the microchannel, as shown in Fig. 2(c). After calibration, the scaling factor between the pixel image distance and the real physical distance can be determined. This factor allows the camera coordinates obtained from the captured images to be accurately transformed into physical coordinates within the observation field.
 | (12) |
The conversion relationship between the camera and the interface is:
 | (13) |
Through two known calibration points (XC2_1, YC2_1) and (XC2_2, YC2_2) with a distance d between the two calibration points, the corresponding ratio can be calculated.
 | (14) |
 | (15) |
In this way, the image coordinates can be converted into physical coordinates under the camera for detecting the oocyte position inside the microchannel of the microfluidic chip.
2.5 Vision algorithm for flow control
The microfluidic chip is equipped with two miniature cameras. To manipulate the oocytes efficiently, camera 2 is employed to detect and track the positions of oocytes within the microfluidic channel, providing real-time feedback for flow control via the syringe pumps. The objective of this control process is to achieve the release of a single oocyte at a time. To realize this, the system must identify the positions of at least the first two oocytes in the channel and determine their relative spacing for accurate decision-making.The vision-based control algorithm is summarized in Algorithm 1 and Fig. S3. Typically, oocytes begin to move sequentially from the rear section toward the front of the channel. As they advance, the distance between adjacent oocytes gradually increases. Once the leading oocyte reaches a predefined release threshold point, it is expelled through the pipette tip of the microfluidic chip. However, as previously noted, incomplete separation may occasionally occur. Therefore, detecting and confirming whether oocytes are fully separated based on visual feedback are essential to ensure that only one oocyte is placed into each well. The vision feedback system dynamically adjusts the flow direction and pump operation to maintain precise separation control. When the vision system detects that the first oocyte has reached the threshold point, pump 1 immediately stops infusion. The position of the second oocyte is then analyzed to determine whether the oocytes are separated. If they are not, pump 2 is activated in the withdrawal mode, trapping the nearest oocyte into the designed well port. Subsequently, the remaining oocytes are aspirated back by pump 1 withdrawal. After successful separation, the first isolated oocyte in the queue is released into the target well. By repeating these vision-guided feedback steps, all oocytes within the microchannel can be sequentially and individually released into well chips.
2.6 Vision algorithm for manipulator control
For the manipulator control, we combined image recognition to calculate the target picking/placing position to achieve automatic control of the chip-on-robot. The control algorithm is shown in Algorithm S1. This process receives images of well chips and oocytes for feedback to manipulator control. In the well-chip part, oocyte images were collected using a camera. Then, the images were detected through the trained YOLOv5 model. The program used detection data to calculate the target picking/placing position. Next, the target position was sent to the robot manipulator. Based on imaging control of the detected oocytes and the movement rule, the chip-on-robot automatically moved to the target location. Finally, after picking/placing one oocyte, the program continued to identify the next target loading position.3 Experiments and results
In this section, the vision-based observation and flow field simulation within the microfluidic chip are performed. Then, we train the vision detection model and use the visual detection for feedback control of the oocyte position in the microfluidic chip and the manipulator position. Finally, we validated the sequential manipulation of the Xenopus oocytes in sequence using the vision-equipped microfluidic chip on robot.3.1 Vision-based observation field
In the vision-equipped microfluidic chip for oocyte observation within a microchannel, the installation distance of the cameras is determined to ensure high-resolution, distortion-free imaging of the observation region. In this case, a miniature camera (300![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 pixels, VGA 640 × 480 resolution, viewing angle approximately 60°) was mounted directly above the microfluidic chip to enable top-view imaging of the oocytes. The installation distance between the camera and the chip surface was determined according to the relationship between the camera's viewing angle and the desired field of view (FOV).32,33 The FOV increases linearly with distance according to the geometric relation:
000 pixels, VGA 640 × 480 resolution, viewing angle approximately 60°) was mounted directly above the microfluidic chip to enable top-view imaging of the oocytes. The installation distance between the camera and the chip surface was determined according to the relationship between the camera's viewing angle and the desired field of view (FOV).32,33 The FOV increases linearly with distance according to the geometric relation:  , where, L is the distance from the camera to the chip surface and θ is the viewing angle.34,35
, where, L is the distance from the camera to the chip surface and θ is the viewing angle.34,35
As shown in Fig. 3(a), the actual width and height of the overlay scene are:  , and
, and  , where W is width of the observation scene, H is the height of the observation scene, and θh and θv are the horizontal viewing angle and vertical viewing angle, respectively. For a camera with a 60° horizontal viewing angle, as the camera distance increases, a larger area of the microfluidic chip can be captured, as shown in Fig. 3(b), but the spatial resolution decreases accordingly. When setting the distance at 35 mm, the calculated view area is shown in Fig. 3(c) and is around 1200 mm2. In the real experiment, the field of view is approximately 33.6 mm × 25.3 mm, resulting in a spatial resolution of approximately 52.5 μm per pixel. The resolution corresponds to approximately 22 pixels across the oocyte diameter. This distance is optimized to ensure that the oocyte region within the chip is clearly focused and fully captured within the field of view. In addition, the FOV also changes with the camera rotating along the axis, as shown in SI Fig. S6 to S9 and section S1.
, where W is width of the observation scene, H is the height of the observation scene, and θh and θv are the horizontal viewing angle and vertical viewing angle, respectively. For a camera with a 60° horizontal viewing angle, as the camera distance increases, a larger area of the microfluidic chip can be captured, as shown in Fig. 3(b), but the spatial resolution decreases accordingly. When setting the distance at 35 mm, the calculated view area is shown in Fig. 3(c) and is around 1200 mm2. In the real experiment, the field of view is approximately 33.6 mm × 25.3 mm, resulting in a spatial resolution of approximately 52.5 μm per pixel. The resolution corresponds to approximately 22 pixels across the oocyte diameter. This distance is optimized to ensure that the oocyte region within the chip is clearly focused and fully captured within the field of view. In addition, the FOV also changes with the camera rotating along the axis, as shown in SI Fig. S6 to S9 and section S1.
3.2 Flow simulation in the microfluidic chip
To evaluate the flow velocity within the microfluidic chip, finite element analysis was conducted using COMSOL Multiphysics. The three-dimensional chip model was imported into COMSOL, where both the channel height and the main channel width were set to 1.6 mm, as shown in Fig. S2. The two side channels also had a width of 1.6 mm and each branch channel was positioned at an angle of relative to the main channel. The branch channel had an entrance width of 0.4 mm, and the spacing between adjacent branches was 3.2 mm. Inlet 1, located at the rear end of the chip, was assigned an initial flow rate of 10 mm s−1, while the outlet was positioned at the chip tip. Following geometric meshing, the velocity field distribution throughout the channel was computed.
relative to the main channel. The branch channel had an entrance width of 0.4 mm, and the spacing between adjacent branches was 3.2 mm. Inlet 1, located at the rear end of the chip, was assigned an initial flow rate of 10 mm s−1, while the outlet was positioned at the chip tip. Following geometric meshing, the velocity field distribution throughout the channel was computed.
We obtained the velocity distribution of different regions, as shown in Fig. 4 and S10. The results indicate that the flow velocity within the main channel increases progressively with infusion from pump 1, as shown in Fig. 4(a). When pump 2 was activated to trap a single oocyte, the flow distribution is shown in Fig. 4(b). The velocity profile along the centerline of the main channel (Fig. 4(c)) confirmed that the designed structure exhibits a stepwise increase in velocity. The reason is that the flow from the side and branch channels converge into the main channel. The magnitude of this stepwise increment depends on the input flow rate. Finally, designing the branch channel into two sections was conducive to the gradual increase of the flow velocity in the main channel. To confirm the release performance for placing single oocytes into the well chip, the flow field in the outlet tip region was simulated, as shown in Fig. 4(d). The velocity gradually decreases from the chip tip toward the bottom of the well chip, as shown in Fig. 4(e) and (f), demonstrating that a lower inlet flow rate yields a gentler release speed, effectively minimizing potential mechanical impact on the oocytes. Finally, the separation gap effect was confirmed through simulations with the particle tracking module, as shown in Fig. 4(g) and (h). The distance between the neighboring oocytes indicated that multiple oocytes were successfully separated.
3.3 Object visual position in the microchannel
To deploy vision detection, the oocyte recognition model was first trained, and the resulting performance metrics are shown in Fig. S11. Evaluation of precision and recall demonstrated that YOLOv5 effectively identified the oocytes. The loss curve indicated a gradual convergence of the model, and the mAP@0.5 metric steadily increased with training. When the training reached 300 epochs, the mAP@0.5 exceeded 0.98, confirming the model's strong performance. The single-image recognition time was under 20 ms, confirming that the model can perform real-time visual detection during microfluidic operations.Next, we evaluated the performance of the microfluidic chip in effectively separating and manipulating multiple hydrogel bead objects (diameter 1.2 mm) through the camera 2 vision information, as shown in Fig. 5. The beads were aspirated into the microfluidic chip for manipulation. The timing control sequence is detailed in SI section S2 and Fig. S12. When beads moved from the rear side to the front side in the main channel, they were separated due to the hydrodynamic focusing effect on the microfluidic chip, as shown in Fig. 5(a) and Video S1. The separation gap between beads was then utilized to release individual beads by controlling the operation of the two pumps. Generally, the first few beads started to move from the rear side to the front side. As objects move toward the front of the channel, the gap distance between adjacent beads increases, as shown in Fig. 5(b) and S13. Once the first bead reaches the releasing position, the maximal separation distance reaches approximately 34 mm, which is equivalent to 28 times the oocyte's diameter (1.2 mm). The separation success rate with the gap effect is around 87% in a single attempt in seven experiments. The speed of oocyte movement increases accordingly along the channel, as shown in Fig. S14, consistent with the simulation results induced by the additional inflow from branch and side channels. Subsequently, by switching the pump, the first bead that reaches the release position is pushed out from the pipette tip of the microfluidic chip.
During the release process, three distinct cases were observed, as shown in Fig. S3. In the separation case, beads were fully isolated. If beads are not separated, pump 2 runs withdrawing, and one bead is trapped into the well port as determined by the vision-based detection of these error cases, as shown in Fig. 5(c), Video S2, and Fig. S15. The position of beads is shown in Fig. 5(d) and (e). Next, the remaining beads were then aspirated backward by withdrawing pump 1. Finally, after separating the beads, the first bead in sequence was placed to the designated well chip. This dealing process takes less than 3.5 s. This difference significantly reduces cumulative operation time compared to previous work,14 as shown in Fig. S16. In addition, with feedback of image information of bead positions, we can determine the flow control strategy between pump 1 and pump 2. Moreover, there is no misjudgment signal in control. To study the flow effect from cooperation of the two pumps, we simulated the flow distribution from the two pumps, as shown in SI Fig. S17 and S18 and section S3. The flow control can be used to optimize the picking/placing process.
3.4 Manipulator motion control with vision
Subsequently, the movement control of the pipette tip of the chip-on-robot by vision camera 1 was confirmed. We moved the chip-on-robot to the target position through multiple single-axis movements,14 as shown in Fig. 6(a). The driving speed of the robotic manipulator was set to 2 mm s−1. The actual trajectory of the tip position was analyzed using software – Tracker and MATLAB. The motion is shown in Fig. S19. Furthermore, the point to point control was assessed through multiple repeated movements. From Fig. 6(b) and (c), and S20, we obtained that the accuracy is less than 0.15 mm, which is significantly smaller than the diameter of a single oocyte (1.2 mm). Based on imaging control of the detected oocytes and the movement rule, the chip-on-robot automatically moved to the target location. | ||
| Fig. 6 Point to point motion of the microfluidic chip on the robot manipulator. (a) Movement path (speed 2 mm s−1). Repeatable of 3 points (b) and point 2 (c). | ||
3.5 Manipulation of multiple single oocytes in sequence
In oocyte experiments, pick-and-place are among the most fundamental tasks, such as placing oocytes at specific worksites for subsequent manipulation or picking target oocytes for further analysis. To demonstrate the capability of our dual vision-equipped chip-on-robot system, we performed sequential pick-and-place manipulation of multiple single oocytes. Patterning and repatterning oocytes in the same well chip are first conducted, as shown in Fig. 7(a). Under the two-vision control, the oocytes are placed as letter “H” and then one oocyte is picked and replaced as letter “n”. The flow rate is set to 2 ml min−1. We repeated these processes 3 times and the patterning all succeeded. After oocytes are loaded into the well chip from the microfluidic chip, we observed the shape and surface of the oocytes under a microscope to assess any potential damage. The results indicate that there are no obvious deformities or breakages observed in the oocytes.In addition, we confirmed the isolation of different oocytes from the well chip, as shown in Fig. 7(b). The different oocyte (blue-treated) is detected by camera 1, and the signal is sent to control the manipulator. Subsequently, the oocytes are picked by the chip-on-robot. When an oocyte is aspirated into the microfluidic chip, camera 2 can detect the signal. After aspiration, oocytes sequentially remained within the main channel of the microfluidic chip. The chip-on-robot then transported them to another area, where the blue oocytes were replaced individually. During the pick-up process, a 10 ml syringe was employed, providing a maximum flow rate of 24 ml min−1. To analyze the pick-up performance, different flow rates were tested (as shown in Fig. 7(c)). The results indicate that oocytes can be reliably captured when the flow rate exceeds 10 ml min−1, while flow rates between 6 ml min−1 and 8 ml min−1 occasionally result in unsuccessful pick-up.
Finally, we demonstrated the transportation of multiple oocytes between these different working well chips. Specifically, oocytes were first picked from one worksite and then transported for rearrangement in another well chip, as shown in Fig. 8(a). The selected oocytes in well chip 1 are unloaded in sequence, as shown in Fig. 8(b) and Video S3. Subsequently, the chip-on-robot with multiple oocytes inside of the microchannel transported to the other well chip. The speed of movement is 2 mm s−1. After reaching the other well chip 2, the chip-on-robot started placing oocytes one by one. The rearrangement process in the target chip is shown in Fig. 8(c). The positions of picking and placing multiple oocytes in different well chips are shown in Fig. 8(d). It shows that the transported distance is beyond 30 mm. In the process, the time is consumed by two parts: the movement of the chip-on-robot and picking/placing a single oocyte, as shown in Fig. 8(e). When the chip-on-robot moves from one well chip to another, it needs to traverse a long distance. Therefore, the movement time is longer at the same speed. During the chip movement, the oocyte remains stationary within the channel. After the chip stop moving, single oocyte can be released at a specific time. Based on these results, the system can reliably perform spatiotemporal sequential manipulation of oocyte groups and transport them into designated worksites.
4 Discussion
This study introduces a dual vision-equipped microfluidic chip capable of achieving spatiotemporally controlled, sequential pick-and-place manipulation of oocytes. Unlike conventional point-sensing methods such as capacitance sensors,14,16,36 which provide only localized single-point data and are susceptible to misjudgment in multi-object environments, the proposed vision-based strategy offers comprehensive spatiotemporal information for dynamic manipulation. This information provides unseparated error case analysis and keeps the compiler continuing system operation. The integration of two miniature cameras directly on the microfluidic chip effectively overcomes the observation constraints of traditional microscope-based systems, which are limited by a fixed field of view. Compared with conventional microscope setups,17,18,20,21 this configuration eliminates the need for mechanical re-focusing or stage adjustments when the chip position changes. Through on-chip visual control, the manipulation process can be continuously monitored even during large-space robotic movement. This strategy bridges the gap between micro-scale fluidic control and macro-scale robotic operation, enabling flexible oocyte transfer between distinct experimental worksites. Moreover, the two cameras ensure a stable imaging foundation for control in microfluidic oocyte manipulation.The well-port assisted fishbone-like microchannel structure,14 in combination with vision-guided flow control, was demonstrated to be highly effective for sequential oocyte isolation and dealing with unseparated error cases. Both experimental observations and flow field simulations confirmed that the fishbone-like channel facilitates the hydrodynamic focusing and progressive separation of multiple oocytes through gradient velocity distributions along the main channel. The addition of well-port structures further improved operation robustness by trapping single oocytes and reducing repetitive release cycles. During this process, the vision-based platform enables spatially resolved, real-time observation of oocyte dynamics within the flow field. This capability ensures precise single-cell-level control during both release and capture operations while improving reproducibility and system stability.
Furthermore, the dual-vision microfluidic chip on robot system enables precise loading of oocytes into designated wells. The overall cycle time for one complete pick-transport-and-place operation, including picking/placing and transporting time of oocytes. The total time for inter-chip transport is dominated by the robot's movement speed (2 mm s−1) and the camera framerate. For larger worksites, increasing the robot's speed or optimizing the movement path could reduce the total operation time. This performance represents a substantial improvement compared to manual handling. This visual–mechanical configuration has rarely been reported in microfluidic oocyte handling and represents a significant advancement toward intelligent micromanipulation platforms.
Nevertheless, certain limitations remain. The resolution of the embedded cameras, although sufficient for oocyte-scale imaging (∼1.2 mm diameter), may not resolve finer subcellular structures or smaller cell types. For a smaller size, tracking stability and placement accuracy would progressively degrade due to reduced pixel coverage and increased sensitivity to noise. Therefore, it is important to choose a suitable lens of the system for different scale objects. On the other hand, since the smaller cell (such as 120 μm) in the current channel (width 1.6 mm, height 1.6 mm) is easy to overlap, it makes separation more difficult. It is also important to set a suitable size of the microchannel for different scale objects. The camera framerate limits the detection speed, which is a challenge posed by the inability of small cameras to achieve high framerates. Additionally, flow control accuracy is still influenced by intrinsic delays in the syringe pump response and subtle surface irregularities in channel fabrication. Furthermore, extending the design toward parallel multi-channel architectures could enhance throughput and experimental efficiency. Our proposed system structure with suitable accessory selection can be used to manipulate different scale cells in large worksites and different worksites. Future work will focus on integrating higher-resolution micro-optical sensors, incorporating stereoscopic depth perception, and developing adaptive flow control algorithms that respond dynamically to visual feedback.
Conclusions
In this study, we demonstrated that a dual vision-based microfluidic chip integrated with a robotic manipulator can achieve sequential manipulation of oocytes. The microfluidic chip was equipped with an embedded camera for real-time tracking of oocyte positions and flow control based on visual feedback. When the camera is set at (0, 0, 35), the view area reaches around 1200 mm2. The results confirmed that multiple oocytes are effectively separated and released in sequence only through hydrodynamic flow focusing within the vision-guided microfluidic chip. Additionally, the incorporation of well-port structures enables selective trapping of individual oocytes, effectively solving cases of incomplete separation among adjacent cells in the microchannel. The other camera, mounted on the tip of the robotic manipulator, was employed to detect and guide the pick-and-place operations of single oocytes. Using dual vision configuration, the chip-on-robot system successfully performed automated pick-up, transport, and placement of multiple oocytes across different well-chip worksites. The proposed method has the advantage of flexible and automatic single-oocyte engineering.Author contributions
S. Liang and F. Arai: conceptualization and methodology. S. Liang, H. Mo, and Y. Dai: resources, data curation, and investigation. S. Liang, H. Sugiura, and S. Amaya: software, writing – review & editing. H. Sugiura and F. Arai: funding acquisition and project administration. S. Liang and F. Arai: supervision, writing – original draft, and writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information (SI), including videos, is provided to illustrate additional details of the system, simulation, and results. The SI videos demonstrate the functionality and effectiveness of the dual vision-equipped microfluidic chip, offering a clearer understanding of oocyte management at the microchannel and the pick-and-place process.Supplementary information: supporting data and videos are available. See DOI: https://doi.org/10.1039/d5lc01118c.
Acknowledgements
This work was supported by JST Moonshot R&D—MILLENNIA Program grant number JPMJMS2033-08.References
- I. Ivorra, A. Alberola-Die, R. Cobo, J. M. González-Ros and A. Morales, Membranes, 2022, 12, 986 CrossRef CAS PubMed.
- M. X. Rodriguez, A. M. Van Keuren and M.-F. Tsai, STAR Protoc., 2021, 2, 100979 CrossRef CAS PubMed.
- S. L. Zeng, L. C. Sudlow and M. Y. Berezin, Expert Opin. Drug Discovery, 2020, 15, 39–52 CrossRef CAS PubMed.
- M. Bhatt, A. Di Iacovo, T. Romanazzi, C. Roseti, R. Cinquetti and E. Bossi, Membranes, 2022, 12, 927 CrossRef CAS PubMed.
- T. Kalstrup and R. Blunck, J. Visualized Exp., 2017, 55598 Search PubMed.
- K. Otani, H. Sugiura, S. Watanabe, T. Bilal, S. Amaya and F. Arai, 2024 IEEE International Conference on Robotics and Automation (ICRA), 2024, pp. 17723–17728 Search PubMed.
- R. Cinquetti, F. G. Imperiali, S. Bozzaro, D. Zanella, F. Vacca, C. Roseti, B. Peracino, M. Castagna and E. Bossi, SLAS Discovery, 2021, 26, 798–810 CrossRef CAS PubMed.
- P. Palay, D. Fathi and R. Fathi, Biol. Reprod., 2022, 108, 393–407 CrossRef PubMed.
- Y. Yamanishi, H. Kuriki, S. Sakuma, M. Hagiwara, T. Kawahara and F. Arai, 2011 International Symposium on Micro-NanoMechatronics and Human Science, 2011, pp. 113–115 Search PubMed.
- X. Liu, Y. Li, F. Liu, Q. Shi, L. Dong, Q. Huang, T. Arai and T. Fukuda, Sci. Adv., 2025, 11, eads8167 CrossRef CAS PubMed.
- Y. Ma, M. Gu, L. Chen, H. Shen, Y. Pan, Y. Pang, S. Miao, R. Tong, H. Huang, Y. Zhu and L. Sun, Theranostics, 2021, 11, 7391–7424 CrossRef CAS PubMed.
- P. Saha, T. Duanis-Assaf and M. Reches, Adv. Mater. Interfaces, 2020, 7, 2001115 CrossRef CAS.
- J. Cheng, R. Anne and Y.-C. Chen, Lab Chip, 2025, 25, 6100–6125 RSC.
- S. Liang, S. Amaya, H. Sugiura, H. Mo, Y. Dai and F. Arai, Adv. Intell. Syst., 2024, 6, 2400185 CrossRef.
- S. Liang, S. Amaya, H. Sugiura, H. Mo, Y. Dai and F. Arai, 2024 IEEE International Conference on Robotics and Automation (ICRA), 2024, pp. 8–13 Search PubMed.
- S. Amaya, H. Sugiura, B. Turan, S. Kaneko and F. Arai, 2023 IEEE 36th International Conference on Micro Electro Mechanical Systems (MEMS), 2023, pp. 57–60 Search PubMed.
- S. Zhou, B. Chen, E. S. Fu and H. Yan, Microsyst. Nanoeng., 2023, 9, 1–15 Search PubMed.
- J. Howell, T. C. Hammarton, Y. Altmann and M. Jimenez, Lab Chip, 2020, 20, 3024–3035 RSC.
- O. Gerhard, S. Schneider, M. Dehne, J. Bahnemann, K. Palme, R. Welsch, O. Dovzhenko, Q. Yu, M. Köhler, J. Cao and A. Groß, Lab Chip, 2025 10.1039/D5LC00550G.
- M. Sesen and G. Whyte, Sci. Rep., 2020, 10, 8736 CrossRef CAS PubMed.
- W. He, J. Zhu, Y. Feng, F. Liang, K. You, H. Chai, Z. Sui, H. Hao, G. Li, J. Zhao, L. Deng, R. Zhao and W. Wang, Nat. Commun., 2024, 15, 10792 CrossRef PubMed.
- T. Aoyama, A. D. Zoysa, Q. Gu, T. Takaki and I. Ishii, J. Rob. Mechatronics, 2016, 28, 854–861 CrossRef.
- A. Mudugamuwa, S. Hettiarachchi, G. Melroy, S. Dodampegama, M. Konara, U. Roshan, R. Amarasinghe, D. Jayathilaka and P. Wang, Sensors, 2022, 22, 6900 CrossRef CAS PubMed.
- E. Dotan, D. Yagoda-Aharoni, E. Shapira and N. T. Shaked, Lab Chip, 2025, 25, 5856–5862 RSC.
- S. Sakuma, K. Nakahara and F. Arai, IEEE Robot. Autom. Lett., 2019, 4, 2973–2980 Search PubMed.
- K. W. Oh, K. Lee, B. Ahn and E. P. Furlani, Lab Chip, 2012, 12, 515–545 RSC.
- J.-L. Bretonnet, J.-F. Wax, J.-L. Bretonnet and J.-F. Wax, AIMS Mater. Sci., 2021, 8, 809–822 CAS.
- F. Meneau, A. Dupré, C. Jessus and E. M. Daldello, Cell, 2020, 9, 1502 CrossRef CAS PubMed.
- K. L. Mowry, Cold Spring Harb. Protoc., 2020, 2020(4), 095844 Search PubMed.
- Z. Yücel, F. Akal and P. Oltulu, Signal Image Video Process., 2023, 17, 4107–4114 CrossRef.
- Y.-R. Li, W.-Y. Lien, Z.-H. Huang and C.-T. Chen, Actuators, 2023, 12, 253 CrossRef.
- P. Fasogbon and L. Fan, 2018 24th International Conference on Pattern Recognition (ICPR), Beijing, 2018, pp. 1875–1881 Search PubMed.
- C. Steger, Int. J. Comput. Vis., 2017, 123, 121–159 CrossRef.
- J. Park, J. Ryu and H. Choi, Appl. Sci., 2024, 14, 9097 CrossRef CAS.
- Z. Ji, Y. Liu, C. Zhao, Z. L. Wang and W. Mai, Adv. Mater., 2022, 34, 2206957 CrossRef CAS PubMed.
- A. I. Egunov, Z. Dou, D. D. Karnaushenko, F. Hebenstreit, N. Kretschmann, K. Akgün, T. Ziemssen, D. Karnaushenko, M. Medina-Sánchez and O. G. Schmidt, Small, 2021, 17, 2002549 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |








