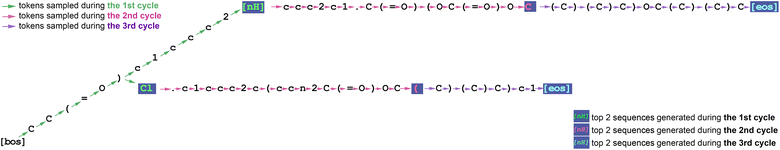Open Access Article
Open Access ArticleFast and scalable retrosynthetic planning with a transformer neural network and speculative beam search
Natalia
Andronova†
 e,
Mikhail
Andronov†
e,
Mikhail
Andronov†
 *a,
Jürgen
Schmidhuber
ab,
Michael
Wand
*a,
Jürgen
Schmidhuber
ab,
Michael
Wand
 ad and
Djork-Arné
Clevert
ad and
Djork-Arné
Clevert
 c
c
aSUPSI, IDSIA, Lugano, Switzerland. E-mail: mikhail.andronov@idsia.ch
bCenter of Excellence for Generative AI, KAUST, Thuwal, Saudi Arabia
cMachine Learning Research, Pfizer Research and Development, Berlin, Germany
dInstitute for Digital Technologies for Personalized Healthcare, SUPSI, Lugano, Switzerland
eIndependent Researcher, Lugano, Switzerland
First published on 30th March 2026
Abstract
AI-based computer-aided synthesis planning (CASP) systems are in demand as components of AI-driven drug discovery workflows. However, the high latency of such CASP systems limits their utility for high-throughput synthesizability screening in de novo drug design. We propose a transformer-based single-step retrosynthesis model with reduced inference latency based on speculative beam search combined with a scalable drafting strategy called Medusa. Replacing the standard transformer and beam search with our approach accelerates the expansion stage of the planning algorithm, leading to higher solvability in CASP when planning under stringent time limits, and saves hours of computation when synthesis is constrained by the number of iterations. Our method brings AI-based CASP systems closer to meeting the stringent latency requirements of high-throughput synthesizability screening and improving the overall user experience.
1 Introduction
The modern pharmaceutical industry is heavily investing in Artificial Intelligence (AI) technologies to reduce the substantial time and financial costs associated with drug development.1 Thus far, AI tools have been most impactful in the preclinical stage of drug discovery, becoming an integral part of the traditional Design-Make-Test-Analyze (DMTA) cycle. Numerous de novo drug design tools can generate virtually unlimited numbers of AI-designed molecular structures for the first stage of the DMTA cycle, already helping to identify potential drug candidates.2 Naturally, the initially generated molecules must undergo extensive filtering, including the filtering for synthesizability.Synthesizability, i.e., the existence of a valid synthesis route from a given target molecule to available building blocks, depends on factors such as route length, yield, cost, the available stock of building blocks, and permitted reaction types.3 Constructing a complete retrosynthetic tree with a Computer-Aided Synthesis Planning (CASP) system provides the most rigorous and flexible assessment of synthesizability. Often, in practical workflows,4,5 full synthesis planning may be replaced with filtering of generated molecules by molecular complexity scores such as SA score6 or SYBA,7 or by machine-learned retrosynthesis prediction scores such as RA score,8 which estimate the likelihood that a CASP system will identify a synthetic route. However, these surrogate metrics provide only approximate assessments and do not replace explicit retrosynthetic tree construction; they may miss feasible routes or overestimate synthetic accessibility. Therefore, accelerating full retrosynthesis planning remains necessary for reliable synthesizability estimation of large amounts of novel molecules.
Like other areas of drug discovery, synthesis planning is also being transformed by AI, and AI-powered CASP systems are now in demand. Open-source AI-based CASP systems such as AiZynthFinder,9,10 ASKCOS,11 SynPlanner,12 and Syntheseus13 combine a single-step retrosynthesis model with a planning algorithm (e.g., MCTS14 or A*15), implementing the design proposed by Segler et al.14
A major challenge limiting the integration of AI-based CASP systems into the DMTA cycle is the stringent latency requirements that a CASP tool must meet to keep up with the flood of structures produced by de novo generators. Current AI CASP systems are not fast enough for applications in the high-throughput setting, taking up to several hours to generate a synthesis plan for a molecule.16,17 Therefore, AI CASP systems will greatly benefit from accelerated inference.
The single-step retrosynthesis models that enable state-of-the-art accuracy are template-free models based on a general sequence-modeling neural network architecture called the transformer.18–20 Typically, single-step retrosynthesis is formulated as a conditional SMILES generation task in transformer models: the model “translates” a query product SMILES into a set of candidate precursor SMILES using beam search in inference.21–23 Since transformers also serve as the backbone for most Large Language Models (LLMs),24 SMILES-to-SMILES transformers as single-step models provide unique opportunities for latency optimization inspired by advances in LLM inference acceleration.
In this work, we demonstrate the acceleration of multi-step retrosynthesis that relies on a SMILES-to-SMILES transformer as a single-step model using two complementary techniques. We accelerate the parallel decoding of single-step expansions using the recently proposed speculative beam search (SBS),25 which extends speculative decoding26 to beam search, in combination with a state-of-the-art drafting approach called Medusa. By integrating both techniques into AiZynthFinder,27 we achieve substantial speed gains in multi-step retrosynthesis. Our method brings AI-based synthesis planning closer to the real-time performance required for integration into modern drug discovery pipelines.
2 Methods
2.1 Speculative decoding
Speculative decoding is a method of reducing the generation latency of autoregressive transformer models. It was originally introduced in the field of Large Language Models (LLMs), where inference speed is a critical issue. Autoregressive transformers are the foundation of both LLMs and template-free SMILES-to-SMILES synthesis prediction; they generate one text token per run, requiring multiple sequential runs to complete a text or a SMILES string. Each model run (forward pass) involves substantial computational overhead and processing time. Speculative decoding accelerates generation by reducing the number of model calls without sacrificing accuracy. It does so by accepting or rejecting entire subsequences that serve as guesses for potential sequence continuations, where these guesses, called “drafts”, may come from arbitrary sources. For example, they can be generated by a dedicated draft model or assembled based on heuristics.The transformer decoder accepts a sequence of N tokens as input and predicts the next token for each position. In standard autoregressive generation, we discard all predictions except the last one, append it to the input sequence, and re-run the transformer. However, in speculative decoding, we first concatenate the input sequence with a draft sequence of M tokens to leverage predictions for multiple positions simultaneously. If the prediction for the last input token matches the first token in the draft sequence, we accept the first draft token and check the prediction for the next position. We repeat this process until either a predicted token differs from the corresponding draft token or we reach the end of the draft sequence. This approach generates between 1 token in the worst case and M + 1 tokens in the best case per transformer forward pass. The probability of accepting a token from the draft is called acceptance rate.26 The empirical mean acceptance rate on the test set is the proportion of accepted speculative tokens to the total number of speculative tokens.
SMILES-to-SMILES generation is a task that is remarkably well-suited for speculative decoding. In chemical reactions, only some of the reactant atoms typically change their connectivity, while large fragments of the reactants remain unchanged and appear the same in the products. Therefore, instead of constructing the target SMILES token by token, the transformer can quickly assemble it from fragments of the query SMILES if they are presented as draft sequences. Extracting multiple fragments of a fixed length from a query sequence, trying them all as drafts at every generation step, and choosing the draft with the most accepted tokens is the essence of the heuristic drafting scheme for speculative decoding applied to SMILES-to-SMILES generation.25
2.2 Speculative beam search
The limitation of basic speculative decoding is that it supports generating only one output sequence per input sequence. While sufficient for text generation in most cases, it poses a major obstacle to the potential of speculative decoding to accelerate template-free reaction prediction and single-step retrosynthesis models. When working as a component of a CASP system, a SMILES-to-SMILES transformer must produce multiple candidate predictions for every query, typically through beam search. We recently developed a method called “speculative beam search (SBS)”25 in an attempt to introduce speculative decoding into CASP. SBS achieves up to 3× acceleration of Molecular Transformer21 inference for reaction prediction and single-step retrosynthesis. The core idea of SBS is an extra step before appending the accepted draft tokens to the growing sequence. After selecting the accepted tokens, we determine the top-K most probable next tokens for each accepted token using the model's forward pass. With that, we obtain a set of candidate subsequences of different lengths, which we then sort by probabilities and extract the top-K most probable continuations to use as beams. Both shorter and longer sequences may be the most probable. The drafts in SBS are generated by a heuristic drafting scheme that uses multiple fragments of the query SMILES as drafts.2.3 Medusa
While SBS achieves good inference acceleration in reaction prediction and single-step retrosynthesis, the heuristic drafting strategy presents a scalability problem.25 To achieve a high acceptance rate, multiple drafts should be used in parallel. It increases the effective batch size as O(BKN), where B is the basic batch size, K is the number of beams, and N is the number of drafts. The latency of a forward pass of the model increases with batch size, and may quickly outweigh the benefits of the high throughput given by batching.A recently proposed method, “Medusa”,28 offers a simple solution for generating single drafts with a high acceptance rate. The fundamental idea of the method is to add extra subnetworks (decoding heads) to the transformer neural network that predict multiple tokens ahead of the next token in parallel. Instead of the usual transformer logits output of shape (B, L, V), a Medusa model gives (B, L, M, V), where B is the input batch size, L is the decoder input length, V is the vocabulary size, and M is the number of Medusa model heads. While the main prediction head generates the next token as usual, the additional Medusa heads predict the second next token, the third next token, and so on up to the M-th next token.
The tokens predicted by the additional heads are the draft sequences for the main head to verify. The first Medusa call is used to generate a draft. In our experiments, the model has 20 heads, so the draft length is 20. We use greedy decoding to create only one draft per given input sequence to avoid inflating the effective batch size. The second Medusa call uses only the main head's output data to verify draft tokens. At least one draft token will always be approved (since it was generated by the main model head in greedy mode), so the worst case is 2 tokens generated across 2 model calls. Of course, the worst-case scenario is still undesirable, as additional heads necessitate additional weights in the model architecture, which may result in a slightly slower forward pass. In our architecture, adding extra heads increased the number of weights by 7.5%. Thus, a high acceptance rate for draft tokens is important. In the best case, the Medusa model with 20 heads (1 main head and 19 extra heads) produces 21 tokens in 2 model calls.
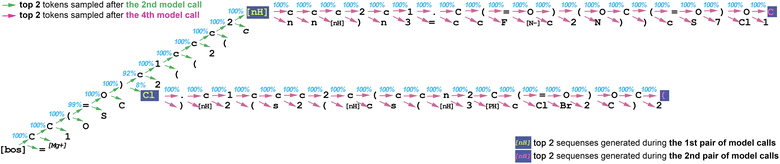
As a verification procedure, we use one similar to top-p sampling. We sort the predicted probabilities for every token in the vocabulary in descending order and calculate the cumulative probabilities for every token. If the cumulative probability for a given draft token is less than the nucleus parameter (99.75%), then that token is considered probable enough and approved. Additionally, the highest probability token among all vocabulary tokens is always approved. Fig. 1 and 2 provide an example.
To train the model, we select the “joint training, combined loss” recipe from the original Medusa paper.28 To give priority to the accuracy of the main head, we divide each head's contribution to the loss function by the number of that head.
When we replace the heuristic drafting in the Molecular Transformer with the Medusa approach, we observe a significant improvement in SBS scalability at larger batch sizes, expanding the potential of speculative decoding and transformer models for fast synthesis planning.
2.4 Multi-step synthesis planning
Since the ultimate goal of building accelerated template-free SMILES-to-SMILES prediction models is to enable fast AI-powered CASP, we evaluate our single-step retrosynthesis model as a component of a multi-step synthesis planning system. We choose AiZynthFinder 4.0 (ref. 9 and 27) because of its straightforward support for arbitrary template-free models and to maintain continuity with prior work that benchmarked various single-step retrosynthesis models as the components of AiZynthFinder.16 We mostly focus on Retro* (ref. 15) as the search algorithm for building the synthesis tree. We use only the reactant probability from the single-step model as the guiding probability in tree search, as done by Torren-Peraire et al.16 AiZynthFinder allows constraining the planning by both total time and the maximum number of iterations, and these must be set simultaneously. Therefore, those constraints should be chosen carefully, because ultimately, the strictest of the two is the limiting constraint. In our experiments, we chose either a time limit (assigning a very large maximum number of iterations) or an iteration limit (assigning a very large maximum time). This made it easier to analyze the experiments. AiZynthFinder also features two search modes: either the search stops when at least one route is found, or it continues until the iteration or time limit is reached. When planning is set to build multiple routes until the time limit is reached, the acceleration of the single-step model does not save time; instead, it enables a more thorough exploration of the route space.It is important to note that AiZynthFinder runs all single-step expansions with a batch size of 1 by design. Therefore, in our multi-step synthesis experiments, the comparison between single-step models is limited to performance at a batch size of 1. Currently, all ML-based open source CASP systems (AiZynthFinder,27 SynPlanner,12 Syntheseus,13 and ASKCOS11) support single-step expansions with only a batch size of 1.
2.5 Model
We train a custom variant of the Molecular Transformer,21 an encoder-decoder transformer for SMILES-to-SMILES translation. Our model consists of six encoder and decoder layers, eight heads in the multi-head attention mechanism, an embedding dimensionality of 256, a feedforward dimensionality of 2048, and 20 additional Medusa heads to predict tokens 1–20 positions ahead of the next token. All Medusa heads are implemented as a single MLP with a hidden layer of dimension 20 × 50 = 1000, followed by a residual connection and layer normalization. The base transformer has 17.4 million parameters, and the Medusa heads have 1.3 million, for a total of 18.7 million. We train a transformer for product prediction to evaluate the round-trip accuracy of the retrosynthesis models. We use the same hyperparameters as in Schwaller et al., with four encoder and decoder layers, eight heads, an embedding dimensionality of 256, and a feedforward dimensionality of 2048, resulting in 11.4 million parameters. We use the Adam29 optimizer for training. We train and test our models on one NVIDIA Tesla V100 GPU with 32 GB of memory.2.6 Data
For training the single-step retrosynthesis model and its isolated evaluation, we employ the standard USPTO 50K dataset with the same train/validation/test splits as in GLN (ref. 30) (40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 008/5001/5007 reactions). We apply the 20-fold R-SMILES augmentation31 to the USPTO 50K training subset. The test set comprises 5007 reactions; we do not augment it. We follow the standard atomwise tokenization procedure21 to tokenize SMILES.
008/5001/5007 reactions). We apply the 20-fold R-SMILES augmentation31 to the USPTO 50K training subset. The test set comprises 5007 reactions; we do not augment it. We follow the standard atomwise tokenization procedure21 to tokenize SMILES.
For multi-step synthesis planning experiments, we used two sets of building blocks: PaRoutes and ZINC. PaRoutes stock contains 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 414 molecules designated as purchasable in the PaRoutes-n1 dataset.10,16 ZINC stock contains 17
414 molecules designated as purchasable in the PaRoutes-n1 dataset.10,16 ZINC stock contains 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422
422![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 831 molecules and is available for download through the AiZynthFinder codebase.32
831 molecules and is available for download through the AiZynthFinder codebase.32
3 Results and discussion
3.1 Single-step retrosynthesis
We first test our models in a single-step retrosynthesis setting using the USPTO 50K dataset. We compare the base transformer with beam search decoding, the base transformer with speculative beam search decoding in a “smart” heuristic drafting variant,25 and the Medusa variant of the transformer with speculative beam search that relies on draft tokens predicted by the Medusa heads. We denote the first model as T-BS (Transformer + Beam Search), the second as T-HSBS (Transformer + Heuristic Speculative Beam Search), and the latter as M-SBS (Medusa + Speculative Beam Search). We also include a separate “optimized” beam search that does not call the model to predict the pad token after the EOS token. This optimization of the beam search does not affect the accuracy or the number of model calls. However, it reduces the effective batch size, thereby reducing calculation time at larger batch sizes. Since T-HSBS and M-SBS do not call their models to produce the pad token after the EOS token, we outline T-BSO (Transformer + Beam Search (Optimized)) as a separate position to ensure a more accurate comparison. Table 1A provides a schematic that describes the difference between the four decoding variants.| Beam size 10 | ||||
|---|---|---|---|---|
| (A) Model description | Architecture | Inference | Drafts | [PAD] generation optimized |
| T-BS | Transformer | Beam search | — | No |
| T-BSO | Transformer | Beam search | — | Yes |
| T-HSBS | Transformer | Speculative beam search | Src. SMILES substrings | Yes |
| M-SBS | Medusa | Speculative beam search | One learnable draft | Yes |
| (B) Decoding wall time, min | B = 1 | B = 4 | B = 8 | B = 16 | B = 32 |
|---|---|---|---|---|---|
| T-BS | 50.0 ± 3.8 | 26.9 ± 3.5 | 18.7 ± 1.2 | 14.9 ± 0.1 | 16.2 ± 0.1 |
| T-BSO | 50.0 ± 2.2 | 16.2 ± 0.3 | 9.4 ± 0.2 | 7.3 ± 0.1 | 5.5 ± 0.1 |
| T-HSBS | 22.7 ± 1.3 | 10.1 ± 0.2 | 7.4 ± 0.2 | 6.1 ± 0.1 | 5.2 ± 0.0 |
| M-SBS | 11.4 ± 0.4 | 4.0 ± 0.2 | 2.4 ± 0.2 | 2.1 ± 0.1 | 1.5 ± 0.1 |
| (C) Model calls | B = 1 | B = 4 | B = 8 | B = 16 | B = 32 |
|---|---|---|---|---|---|
| T-BS | 295![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 947 947 |
99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 030 030 |
54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 934 934 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 941 941 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 170 170 |
| T-BSO | 295![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 947 947 |
99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 030 030 |
54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 934 934 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 941 941 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 170 170 |
| T-HSBS | 92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 538 538 |
36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 960 960 |
28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 056 056 |
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 807 807 |
8817 |
| M-SBS | 59![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 502 502 |
19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 240 240 |
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 730 730 |
5906 | 3224 |
| (D) Average effective batch size | B = 1 | B = 4 | B = 8 | B = 16 | B = 32 |
|---|---|---|---|---|---|
| T-BS | 10 | 40 | 80 | 160 | 320 |
| T-BSO | 8 | 25 | 45 | 82 | 151 |
| T-HSBS | 23 | 40 | 29 | 52 | 93 |
| M-SBS | 6 | 18 | 32 | 58 | 105 |
| (E) Acceptance rate, % | B = 1 | B = 4 | B = 8 | B = 16 | B = 32 |
|---|---|---|---|---|---|
| T-HSBS | 74 | 70 | 64 | 64 | 64 |
| M-SBS | 91 | 91 | 91 | 91 | 91 |
As Table 1B shows, M-SBS significantly outperforms T-BS and T-HSBS at various batch sizes in terms of inference speed. T-HSBS outperforms T-BS and T-BSO at smaller batch sizes but suffers from scalability limitations. Due to the throughput-latency tradeoff inherent to processing multiple draft sequences simultaneously, the heuristic drafting scheme requires careful tuning of draft number and length to achieve optimal performance. At larger batch sizes, the computational overhead of processing multiple drafts negates the acceleration benefits, and the optimal number of drafts becomes 1, making T-HSBS similar to M-SBS, as it also uses only one draft. At the same time, M-SBS achieves a higher acceptance rate (Table 1E) through its integrated architecture, maintaining consistent acceleration even at batch size 32, which establishes M-SBS as the superior acceleration approach for single-step retrosynthesis with transformers. M-SBS requires fewer forward passes of the model to complete generation (Table 1C) and achieves an acceptance rate of 91%, leaving T-HSBS far behind.
In terms of accuracy and prediction validity, all three methods demonstrate nearly identical performance (Table 2). While our speculative beam search approach does not guarantee output distributions identical to those of standard beam search, the practical differences are negligible. A slightly larger difference in accuracy and SMILES validity between M-SBS and T-HSBS stems from marginal performance differences across model checkpoints rather than algorithmic effects. M-SBS implies a custom transformer architecture and requires training a separate model, whereas T-HSBS is a drop-in replacement for beam search.
| Single-step retrosynthesis | Top-1 | Top-3 | Top-5 | Top-10 | |
|---|---|---|---|---|---|
| Accuracy, % | T-BS/T-BSO | 52.08 | 75.16 | 82.97 | 89.08 |
| T-HSBS | 52.08 | 75.16 | 82.07 | 89.12 | |
| M-SBS | 54.06 | 75.95 | 82.90 | 89.20 | |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
|||||
| Pred. 1 | Pred. 3 | Pred. 5 | Pred. 10 | ||
| Invalid SMILES, % | T-BS/T-BSO | 0.8 | 1.8 | 3.5 | 8.1 |
| T-HSBS | 0.8 | 1.8 | 3.5 | 8.2 | |
| M-SBS | 0.5 | 1.7 | 3.1 | 9.3 | |
3.2 Multi-step retrosynthesis
We incorporated our single-step retrosynthesis models into AiZynthFinder to test their performance in multi-step synthesis planning. Since M-SBS consistently outperforms T-HSBS in terms of the speed of single-step prediction generation, we decided to compare M-SBS only with the standard Transformer + Optimized Beam Search (T-BSO) in multi-step synthesis planning. We will further refer to M-SBS as “Medusa” and to T-BSO as “Transformer”. We selected Caspyrus10k as the benchmark dataset for multi-step synthesis planning, drawing inspiration from the experiments of Torren-Peraire et al.16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 414 molecules) or the ZINC stock (17
414 molecules) or the ZINC stock (17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422
422![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 831 molecules). We set the models to generate 10 candidate precursor sets with every call of a single-step model, constrained the maximum route length to 5 or 7, and the maximum number of algorithm iterations to 35
831 molecules). We set the models to generate 10 candidate precursor sets with every call of a single-step model, constrained the maximum route length to 5 or 7, and the maximum number of algorithm iterations to 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 (enough to ensure the priority of the time constraint). When the algorithm identifies the first route from a query molecule to the building blocks, it stops, and the molecule is considered solved. Table 3 summarizes the results of our multi-step retrosynthesis experiments under the time constraints of 5 and 15 seconds for solving a molecule. The results reveal that Medusa consistently outperforms Transformer across all experimental conditions, with improvements in both the number of solved molecules and computational efficiency.
000 (enough to ensure the priority of the time constraint). When the algorithm identifies the first route from a query molecule to the building blocks, it stops, and the molecule is considered solved. Table 3 summarizes the results of our multi-step retrosynthesis experiments under the time constraints of 5 and 15 seconds for solving a molecule. The results reveal that Medusa consistently outperforms Transformer across all experimental conditions, with improvements in both the number of solved molecules and computational efficiency.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 414 molecules) (A–C) or ZINC stock (D and E) is used as a building block set. The maximum synthesis route length is 5 (A–C and D) or 7 (E). The search is stopped when at least one route for a given molecule is found
414 molecules) (A–C) or ZINC stock (D and E) is used as a building block set. The maximum synthesis route length is 5 (A–C and D) or 7 (E). The search is stopped when at least one route for a given molecule is found
| (A) DFPN, PaRoutes stock, depth 5; time limit 5 seconds | Transformer | Medusa | |
|---|---|---|---|
| Solved molecules | 1117 | 2080 | |
| Common solved molecules | 1017 | ||
| Avg. time per solved molecule, sec | 2.01 | 1.85 | |
| Statistics on common solved molecules: | |||
| Avg. time per molecule, sec | 1.88 | 0.86 (×2.) | |
| Avg. alg. iterations per molecule | 6.52 | 9.51 | |
| Avg. alg. iteration time, sec per iteration | 0.34 | 0.10 (×3.) |
| (B) Retro*, PaRoutes stock, depth 5; time limit 5 seconds | Transformer | Medusa | |
|---|---|---|---|
| Solved molecules | 3890 | 5287 | |
| Common solved molecules | 3628 | ||
| Avg. time per solved molecule, sec | 2.14 | 1.41 | |
| Statistics on common solved molecules: | |||
| Avg. time per molecule, sec | 2.06 | 0.99 (×2.) | |
| Avg. alg. iterations per molecule | 5.51 | 7.38 | |
| Avg. alg. iteration time, sec per iteration | 0.43 | 0.16 (×3.) |
| (C) Retro*, PaRoutes stock, depth 5; time limit 15 seconds | Transformer | Medusa | |
|---|---|---|---|
| Solved molecules | 5341 | 6715 | |
| Common solved molecules | 5050 | ||
| Avg. time per solved molecule, sec | 4.25 | 2.86 | |
| Statistics on common solved molecules: | |||
| Avg. time per molecule, sec | 4.00 | 1.84 (×2.) | |
| Avg. alg. iterations per molecule | 12.44 | 18.99 | |
| Avg. alg. iteration time, sec per iteration | 0.44 | 0.14 (×3.) |
| (D) Retro*, ZINC stock, depth 5; time limit 15 seconds | Transformer | Medusa | |
|---|---|---|---|
| Solved molecules | 7708 | 8343 | |
| Common solved molecules | 7439 | ||
| Avg. time per solved molecule, sec | 3.21 | 1.73 | |
| Statistics on common solved molecules: | |||
| Avg. time per molecule, sec | 3.06 | 1.26 (×2.) | |
| Avg. alg. iterations per molecule | 6.15 | 10.13 | |
| Avg. alg. iteration time, sec/iteration | 0.60 | 0.16 (×4.) |
| (E) Retro*, ZINC stock, depth 7; time limit 15 seconds | Transformer | Medusa | |
|---|---|---|---|
| Solved molecules | 7888 | 8608 | |
| Common solved molecules | 7666 | ||
| Avg. time per solved molecule, sec | 3.37 | 1.89 | |
| Statistics on common solved molecules: | |||
| Avg. time per molecule, sec | 3.23 | 1.41 (×2.) | |
| Avg. alg. iterations per molecule | 6.5 | 9.5 | |
| Avg. alg. iteration time, sec/iteration | 0.61 | 0.19 (×3.) |
The iteration of the planning algorithm (Retro* or DFPN search) is accelerated with Medusa by approximately 3 times, which results from the acceleration of the single-step model by approximately 4 times (Table 1B). Due to the different number of the planning algorithm iterations, the first route is identified by Medusa approximately 2 times faster. It leads to the increase of success rate under the stringent time limit. When using DFPN search with a 5 second limit, Medusa solves 2080 molecules out of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, which is 86% more compared to the 1117 solved by Transformer. For the 1017 molecules that both methods successfully solve, Medusa requires on average less than half the time (0.86 seconds vs. 1.88 seconds) (Table 3A). With the Retro* algorithm, Medusa maintains its advantage, solving 36% more molecules than Transformer within 5 seconds (5287 vs. 3890, Table 3B) and 26% more within 15 seconds (6715 vs. 5341, Table 3C). When using the ZINC stock with a depth limit of 5 and a 15 second time limit (Table 3D), Medusa solves 8343 molecules compared to 7708 for the Transformer, while also reducing the average solution time (1.73 s vs. 3.21 s). Finally, with ZINC and the depth limit increased to 7 (Table 3E), Medusa solves 8608 molecules versus 7888 for the Transformer and maintains a lower average solution time (1.89 s vs. 3.37 s). Across all conditions, Medusa consistently achieves faster average solution times while solving substantially more molecules.
000, which is 86% more compared to the 1117 solved by Transformer. For the 1017 molecules that both methods successfully solve, Medusa requires on average less than half the time (0.86 seconds vs. 1.88 seconds) (Table 3A). With the Retro* algorithm, Medusa maintains its advantage, solving 36% more molecules than Transformer within 5 seconds (5287 vs. 3890, Table 3B) and 26% more within 15 seconds (6715 vs. 5341, Table 3C). When using the ZINC stock with a depth limit of 5 and a 15 second time limit (Table 3D), Medusa solves 8343 molecules compared to 7708 for the Transformer, while also reducing the average solution time (1.73 s vs. 3.21 s). Finally, with ZINC and the depth limit increased to 7 (Table 3E), Medusa solves 8608 molecules versus 7888 for the Transformer and maintains a lower average solution time (1.89 s vs. 3.37 s). Across all conditions, Medusa consistently achieves faster average solution times while solving substantially more molecules.
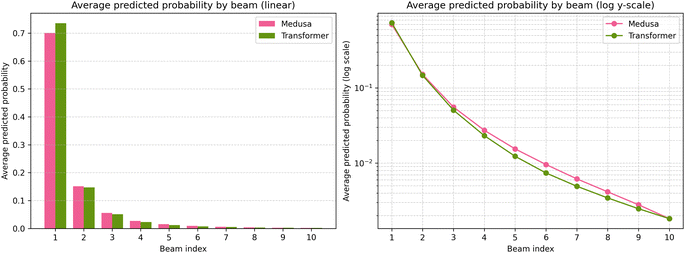
Interestingly, Medusa required more algorithm iterations per commonly solved molecule than Transformer. This likely reflects differences in probability distributions: Transformer tends to concentrate probability mass on the top candidate, while Medusa produces more smooth distributions across candidates, leading to more exploratory search behavior. Fig. 3 shows that the top-1 probability of Medusa is lower than that of the Transformer. The effect is small, but it is noticeable. Similarly, the number of commonly solved molecules under different parameter settings (Table 3) never coincides with the Transformer's number of solved molecules, indicating that Medusa produces a slightly different probability distribution.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422
422![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 831 molecules) of building blocks, Retro* as a search algorithm, and the maximum number of algorithm iterations per molecule of 200. Since we conducted our experiments on a GPU (Tesla V100 32 GB), we set the time limit to 3 minutes instead of the original 480 minutes, which is enough for Transformer and Medusa to be limited only by the iteration limit, and not the time limit, in the case of a complex input molecule. We also use the setting in which the search is stopped as soon as at least one route for a given molecule is found. It does not affect the success rate (the number of molecules for which a full synthesis route is found), but it speeds up experiments by avoiding the calculation of all 200 Retro* iterations. According to Table 6, Transformer, a model of 17 million parameters, is sufficient to achieve a high percentage of successfully solved molecules, approaching 100%. With the limit of 200 iterations, Transformer reaches 97.13% of solved molecules, spending 0.67 seconds per Retro* iteration, while Medusa achieves 95.90% of solved molecules, spending 2 times less, only 0.31 seconds per iteration. This means that if Transformer ran for all 200 iterations, as in the experiments of Torren-Peraire et al.,16 it would spend about 134 seconds per molecule and about 372 hours on the entire dataset. Meanwhile, for Medusa, these values would be 62 seconds and 172 hours, respectively. Thus, Medusa would save approximately 200 hours of GPU calculations while maintaining a 95.9% solved-molecule rate.
831 molecules) of building blocks, Retro* as a search algorithm, and the maximum number of algorithm iterations per molecule of 200. Since we conducted our experiments on a GPU (Tesla V100 32 GB), we set the time limit to 3 minutes instead of the original 480 minutes, which is enough for Transformer and Medusa to be limited only by the iteration limit, and not the time limit, in the case of a complex input molecule. We also use the setting in which the search is stopped as soon as at least one route for a given molecule is found. It does not affect the success rate (the number of molecules for which a full synthesis route is found), but it speeds up experiments by avoiding the calculation of all 200 Retro* iterations. According to Table 6, Transformer, a model of 17 million parameters, is sufficient to achieve a high percentage of successfully solved molecules, approaching 100%. With the limit of 200 iterations, Transformer reaches 97.13% of solved molecules, spending 0.67 seconds per Retro* iteration, while Medusa achieves 95.90% of solved molecules, spending 2 times less, only 0.31 seconds per iteration. This means that if Transformer ran for all 200 iterations, as in the experiments of Torren-Peraire et al.,16 it would spend about 134 seconds per molecule and about 372 hours on the entire dataset. Meanwhile, for Medusa, these values would be 62 seconds and 172 hours, respectively. Thus, Medusa would save approximately 200 hours of GPU calculations while maintaining a 95.9% solved-molecule rate.
We compared the multistep success rate of the models with their round-trip accuracy, diversity, and percentage of invalid SMILES (Table 4). Instead of the relative round-trip accuracy, we calculated the number of round-trip feasible reactions, i.e., predicted reactions that lead to the initial target molecule when evaluated with the Molecular Transformer forward model in top-10 mode. Diversity is measured as the number of unique reaction names, assigned by Rxn-INSIGHT.34 For comparison, we took the default template-based single-step model from AiZynthFinder 4.0. Despite the template-based model (42![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 554 templates in total) offering 50 templates at a time, some templates cannot be applied, some predictions are filtered out by the default neural filter, and duplicate predictions also occur. As a result, the model, on average, offers only 15 reactions per target molecule during testing. The model runs on a CPU. We assigned it a longer time limit of 5 hours per molecule to ensure the programs' running times would be comparable. It increased the success rate from 72.6% to 82.6%.
554 templates in total) offering 50 templates at a time, some templates cannot be applied, some predictions are filtered out by the default neural filter, and duplicate predictions also occur. As a result, the model, on average, offers only 15 reactions per target molecule during testing. The model runs on a CPU. We assigned it a longer time limit of 5 hours per molecule to ensure the programs' running times would be comparable. It increased the success rate from 72.6% to 82.6%.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422
422![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 831 molecules) is used as building blocks; the search is stopped when at least one route for a given molecule is found. Maximum synthesis route length is 7. Transformer stands for transformer with beam search, which produces pad-token after EOS-token automatically without extra calls. Medusa is a Transformer-like model that uses speculative beam search with Medusa heads as draft source. These models run on a GPU. AZF stands for the AiZynthFinder 4.0 default single-step model with a cumulative probability of 99.5% in combination with top-50; it runs on 55 CPU processes in parallel. The multi-step experiments are conducted with a limit of 200 iterations of the planning algorithm and a time limit of 3 minutes (in the case of AZF model, 5 hours per molecule, due to its fast operation). The single-step metrics are measured on the USPTO 50k test set. Eff. N stands for the number of valid reaction SMILES without repetitions. R.-t. (round-trip) feasible reactions stands for the number of reactions that lead to the initial target molecule when evaluated with the Molecular Transformer forward model in top-10 mode. The diversity counts the number of unique reaction names, classified by Rxn-INSIGHT.34
831 molecules) is used as building blocks; the search is stopped when at least one route for a given molecule is found. Maximum synthesis route length is 7. Transformer stands for transformer with beam search, which produces pad-token after EOS-token automatically without extra calls. Medusa is a Transformer-like model that uses speculative beam search with Medusa heads as draft source. These models run on a GPU. AZF stands for the AiZynthFinder 4.0 default single-step model with a cumulative probability of 99.5% in combination with top-50; it runs on 55 CPU processes in parallel. The multi-step experiments are conducted with a limit of 200 iterations of the planning algorithm and a time limit of 3 minutes (in the case of AZF model, 5 hours per molecule, due to its fast operation). The single-step metrics are measured on the USPTO 50k test set. Eff. N stands for the number of valid reaction SMILES without repetitions. R.-t. (round-trip) feasible reactions stands for the number of reactions that lead to the initial target molecule when evaluated with the Molecular Transformer forward model in top-10 mode. The diversity counts the number of unique reaction names, classified by Rxn-INSIGHT.34
| Multistep | Single-step, average per target molecule | |||||||
|---|---|---|---|---|---|---|---|---|
| Model | Top N | Limit | Success rate, % | Total time, h | Invalid SMILES, % | Eff. N | R.-t. feasible reactions | Diversity, classes |
| Transformer | 10 | 200 it | 89.49 | 29.57 | 4.03 | 9.2 | 5.8 | 5.77 |
| 180 s | 94.08 | 58.27 | ||||||
| 50 | 200 it | 97.13 | 26.42 | 13.98 | 39.6 | 18.5 | 13.57 | |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||||
| Medusa | 10 | 200 it | 87.90 | 10.32 | 4.07 | 9.4 | 5.9 | 5.80 |
| 180 s | 94.90 | 41.32 | ||||||
| 50 | 200 it | 95.90 | 15.42 | 15.22 | 39.9 | 18.5 | 13.43 | |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||||||
| AZF (template-based) | 50 | 200 it | 72.63 | 2.73 | 0 | 15.1 | 9.4 | 7.87 |
| 5 h | 82.60 | 24.01 | ||||||
According to all the single-step metrics, Transformer and Medusa are very similar. As we trained our models on data without reagents, they achieved relatively low round-trip accuracies of 62.8% and 63.0%, respectively (considering beam size 10 mode). These numbers are close to the template-based model's round-trip accuracy of 62.3%. One can see that in the 3 minutes per molecule time constraint, Medusa speeds up the program by 1.4×. In the iteration constraint, the 4.4× acceleration of the single-step model and, as a result, faster multi-step iteration, leads to 1.7× and 2.9× acceleration of the entire program.
The PaRoutes-n1 dataset contains 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 target molecules, each associated with a single reference synthesis route and a predefined stock of building blocks. From this dataset, we randomly selected 1000 target molecules for evaluation. The search configuration provided with the PaRoutes benchmark was used without modification. Both models successfully completed the full 500-iteration search procedure for all evaluated targets.
000 target molecules, each associated with a single reference synthesis route and a predefined stock of building blocks. From this dataset, we randomly selected 1000 target molecules for evaluation. The search configuration provided with the PaRoutes benchmark was used without modification. Both models successfully completed the full 500-iteration search procedure for all evaluated targets.
The results of this experiment are summarized in Table 5. During the search, we collected up to 50 solved synthesis routes per target molecule. Both models demonstrated a high overall solvability, defined as the fraction of targets for which at least one valid synthesis route was found. Medusa solved 937 molecules (93.7%), while the Transformer solved 941 molecules (94.1%). To assess the models' ability to reproduce the reference routes provided in PaRoutes, we evaluated route accuracy, defined as the fraction of targets for which the exact reference route appears among the top-50 predicted synthesis routes. Medusa successfully recovered the reference route for 276 molecules (27.6%) and completed the evaluation in 10 hours. The Transformer recovered the reference route for 286 molecules (28.6%) and required 24 hours of computation. Again, Medusa maintains the Transformer's performance while being significantly faster.
| Model | Success rate, % | Route accuracy | Route accuracy overlap | Building block accuracy on unsolved molecules | Runtime, h |
|---|---|---|---|---|---|
| Transformer | 94.1 | 286 | 248 | 8/28 | 24 |
| Medusa | 93.7 | 276 | 22/38 | 10 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 422
422![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 831 molecules) is used as building blocks. Beam size is 50. Maximum synthesis route length is 7. Transformer stands for transformer with beam search, which produces pad-token after EOS-token automatically without extra calls. Medusa is a Transformer-like model that uses speculative beam search with Medusa heads as draft source. AZF stands for AiZynthFinder single-step model. Av. Retro* iter. time represents the average time of a single Retro* iteration. GPU stands for Tesla V100 32 GB. CPU experiments data is taken from ref. 16
831 molecules) is used as building blocks. Beam size is 50. Maximum synthesis route length is 7. Transformer stands for transformer with beam search, which produces pad-token after EOS-token automatically without extra calls. Medusa is a Transformer-like model that uses speculative beam search with Medusa heads as draft source. AZF stands for AiZynthFinder single-step model. Av. Retro* iter. time represents the average time of a single Retro* iteration. GPU stands for Tesla V100 32 GB. CPU experiments data is taken from ref. 16
The sets of molecules solved by both Transformer and Medusa do not fully overlap. Among the molecules for which the reference route was recovered, 248 targets were common to both models. Therefore, Transformer solved 38 molecules that Medusa did not solve, while Medusa solved 28 molecules that Transformer did not solve. This observation is consistent with our earlier results (e.g., Table 3) and indicates that the two models explore different regions of the retrosynthetic solution space, despite achieving similar overall performance. For the 38 molecules solved only by the Transformer, Medusa generated routes with exactly matching reference building block sets for 22 molecules, indicating that it was often close to recovering the correct route topology. Conversely, among the 28 molecules solved only by Medusa, the Transformer produced routes with matching building block sets for 8 molecules.
These observations suggest that even when the exact reference route is not recovered, the models frequently identify chemically consistent precursor sets, reflecting partial convergence toward the reference synthesis strategy.
3.3 Limitations
Similar to the paper by Torren-Peraire et al.,16 we focus here solely on the speed of building the retrosynthetic graph. However, Aizynthfinder also spends a significant amount of time splitting the tree into separate routes. The more model calls are made within the time limit, the more nodes are added to the retrosynthesis tree, and the more complex the tree becomes for splitting. We alleviated this problem by allowing AiZynthFinder to extract only the successful routes in which all leaves are building blocks. This approach requires only a small fraction of the time required for the exhaustive process of extracting all retrosynthetic routes in AiZynthFinder. In cases where unsolved routes are required, the tree-splitting algorithm can be easily optimized by first extracting the most probable reactions rather than making a random choice of unsolved reactions, and by decreasing the maximum route number limit.
Table 4 shows that the round-trip accuracy of Transformer rapidly decreases from 63%  to 47%
to 47%  when going from top-10 to top-50, although the success rate is noticeably improved from 90% to 97% (with Medusa showing similar behavior). We attribute this to the fact that the quality of the beams decreases with an increase in their number, and the template-free models sometimes return short answers, i.e., SMILES that are valid but meaningless, for instance, “Cl” as a single precursor for “ClCc1csc(-c2ccccc2)n1”. At the same time, the metrics show that after the tenth beam, valuable predictions are generated (the absolute number of round-trip feasible reactions is larger in the top-50 than in the top-10), but they are mixed with low-quality predictions that need to be filtered out. Although it is easy to filter out invalid reactions or those that contain the product itself among the reactants, short answers are more difficult to filter out. The default neural filter of AizynthFinder 4.0 defines “Cl ≫ClCc1csc(-c2ccccc2)n1” as a feasible reaction with 99.95% probability. The round-trip filter successfully recognizes this case, but it is heavy and also may erroneously filter out reactions written with missing reagents. Generating reactions with reagents would, in turn, reduce the diversity of autoregressively generated reactions and, consequently, the diversity of routes. Thus, it is necessary to find an elusive balance between all these factors. It may be useful to combine heuristic filters of nonsensical reactions with transformer models in order to avoid formally solved, but in fact incomplete routes.
when going from top-10 to top-50, although the success rate is noticeably improved from 90% to 97% (with Medusa showing similar behavior). We attribute this to the fact that the quality of the beams decreases with an increase in their number, and the template-free models sometimes return short answers, i.e., SMILES that are valid but meaningless, for instance, “Cl” as a single precursor for “ClCc1csc(-c2ccccc2)n1”. At the same time, the metrics show that after the tenth beam, valuable predictions are generated (the absolute number of round-trip feasible reactions is larger in the top-50 than in the top-10), but they are mixed with low-quality predictions that need to be filtered out. Although it is easy to filter out invalid reactions or those that contain the product itself among the reactants, short answers are more difficult to filter out. The default neural filter of AizynthFinder 4.0 defines “Cl ≫ClCc1csc(-c2ccccc2)n1” as a feasible reaction with 99.95% probability. The round-trip filter successfully recognizes this case, but it is heavy and also may erroneously filter out reactions written with missing reagents. Generating reactions with reagents would, in turn, reduce the diversity of autoregressively generated reactions and, consequently, the diversity of routes. Thus, it is necessary to find an elusive balance between all these factors. It may be useful to combine heuristic filters of nonsensical reactions with transformer models in order to avoid formally solved, but in fact incomplete routes.
Currently, AiZynthFinder and its alternatives do not readily support batch sizes other than one for single-step retrosynthesis models in multi-step synthesis planning. Our future work will focus on generalizing multi-step synthesis planning algorithms to support larger batch sizes, which should further reduce the latency in CASP systems, considering the signigicant speed benefits of Medusa at larger batch sizes.
4 Conclusions
We demonstrate that a Medusa variant of the Transformer combined with speculative beam search offers significant speed advantage in multi-step synthesis planning in AiZynthFinder compared to the usual combination of SMILES-to-SMILES Transformer and beam search while retaining the same level of accuracy. In isolated single-step retrosynthesis experiments, Medusa and SBS accelerate the generation of 10 expansions per query by approximately 4 times across batch sizes from 1 to 32 without compromising the accuracy of the baseline Transformer. When Transformer is compared against Medusa in multi-step synthesis planning in AiZynthFinder under stringent time constraints, Medusa allows a single iteration of the planning algorithm to run approximately 3 times faster across varying time limits, search algorithms, and building blocks, enabling AiZynthFinder to solve significantly more molecules under the same time constraints compared to the Transformer. When the multi-step search is constrained by a maximum number of iterations instead of maximum time, the search with Medusa completes much faster than with Transformer, saving dozens of hours of compute while demonstrating negligible discrepancy in solvability and route accuracy. Our approach presents a promising alternative for SMILES-to-SMILES transformers in synthesis planning when planning speed is critical and brings the speed of AI-based CASP closer to the level required for high-throughput synthesizability screening in pharmaceutical research.Author contributions
MA and NA conceptualized the paper idea. NA and MA wrote the code and conducted the experiments. MA and NA wrote the manuscript with inputs from all co-authors. JS, DC, and MW acquired the research funding, administered the project, and provided supervision.Conflicts of interest
There are no conflicts to declare.Data availability
The instructions for downloading the data and running the code to replicate the results are at https://github.com/Academich/faster-ml-casp and https://doi.org/10.5281/zenodo.18002214.Acknowledgements
This study was partially funded by the European Union's Horizon 2020 research and innovation program under the Marie Skłodowska-Curie Innovative Training Network European Industrial Doctorate grant agreement no. 956832 “Advanced machine learning for Innovative Drug Discovery”.References
- R. Vijayan, J. Kihlberg, J. B. Cross and V. Poongavanam, Drug discovery today, 2022, 27, 967–984 CrossRef CAS PubMed.
- Y. Ivanenkov, B. Zagribelnyy, A. Malyshev, S. Evteev, V. Terentiev, P. Kamya, D. Bezrukov, A. Aliper, F. Ren and A. Zhavoronkov, ACS Med. Chem. Lett., 2023, 14, 901–915 CrossRef CAS PubMed.
- M. Stanley and M. Segler, Curr. Opin. Struct. Biol., 2023, 82, 102658 CrossRef CAS PubMed.
- C.-H. Liu, M. Korablyov, S. Jastrzebski, P. Włodarczyk-Pruszynski, Y. Bengio and M. Segler, J. Chem. Inf. Model., 2022, 62, 2293–2300 CrossRef CAS PubMed.
- A. K. Hassen, M. Šícho, Y. J. van Aalst, M. C. Huizenga, D. N. Reynolds, S. Luukkonen, A. Bernatavicius, D.-A. Clevert, A. P. Janssen and G. J. van Westen, et al. , J. Cheminf., 2025, 17, 41 Search PubMed.
- P. Ertl and A. Schuffenhauer, J. Cheminf., 2009, 1, 1–11 Search PubMed.
- M. Voršilák, M. Kolář, I. Čmelo and D. Svozil, J. Cheminf., 2020, 12, 35 Search PubMed.
- A. Thakkar, V. Chadimová, E. J. Bjerrum, O. Engkvist and J.-L. Reymond, Chem. Sci., 2021, 12, 3339–3349 RSC.
- L. Saigiridharan, A. K. Hassen, H. Lai, P. Torren-Peraire, O. Engkvist and S. Genheden, J. Cheminf., 2024, 16, 57 Search PubMed.
- S. Genheden and E. Bjerrum, Digital Discovery, 2022, 1, 527–539 RSC.
- Z. Tu, S. J. Choure, M. H. Fong, J. Roh, I. Levin, K. Yu, J. F. Joung, N. Morgan, S.-C. Li, X. Sun and et al., arXiv, preprint, arXiv:2501.01835, 2025, DOI:10.48550/arXiv.2501.01835.
- T. Akhmetshin, D. Zankov, P. Gantzer, D. Babadeev, A. Pinigina, T. Madzhidov and A. Varnek, J. Chem. Inf. Model., 2025, 65, 15–21 CrossRef CAS PubMed.
- K. Maziarz, A. Tripp, G. Liu, M. Stanley, S. Xie, P. Gaiński, P. Seidl and M. H. Segler, Faraday Discuss., 2025, 256, 568–586 RSC.
- M. H. Segler, M. Preuss and M. P. Waller, Nature, 2018, 555, 604–610 CrossRef CAS PubMed.
- B. Chen, C. Li, H. Dai and L. Song, International Conference On Machine Learning, 2020, pp. 1608–1616 Search PubMed.
- P. Torren-Peraire, A. K. Hassen, S. Genheden, J. Verhoeven, D.-A. Clevert, M. Preuss and I. V. Tetko, Digital Discovery, 2024, 3, 558–572 RSC.
- A. K. Hassen, P. Torren-Peraire, S. Genheden, J. Verhoeven, M. Preuss and I. Tetko, arXiv, preprint, arXiv:2212.11809, 2022, DOI:10.48550/arXiv.2212.11809.
- A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, Ł. Kaiser and I. Polosukhin, Adv. Neural Inf. Process. Syst., 2017, 30, 6000–6010 Search PubMed.
- J. Schmidhuber, Neural Comput., 1992, 4, 131–139 CrossRef.
- I. Schlag, K. Irie and J. Schmidhuber, Proceedings of the 38th International Conference on Machine Learning, 2021, pp. 9355–9366 Search PubMed.
- P. Schwaller, T. Laino, T. Gaudin, P. Bolgar, C. A. Hunter, C. Bekas and A. A. Lee, ACS Cent. Sci., 2019, 5, 1572–1583 CrossRef CAS PubMed.
- I. V. Tetko, P. Karpov, R. Van Deursen and G. Godin, Nat. Commun., 2020, 11, 5575 CrossRef CAS PubMed.
- R. Irwin, S. Dimitriadis, J. He and E. J. Bjerrum, Machine Learning: Science and Technology, 2022, 3, 015022 Search PubMed.
- T. Brown, B. Mann, N. Ryder, M. Subbiah, J. D. Kaplan, P. Dhariwal, A. Neelakantan, P. Shyam, G. Sastry and A. Askell, et al. , Adv. Neural Inf. Process. Syst., 2020, 33, 1877–1901 Search PubMed.
- M. Andronov, N. Andronova, M. Wand, J. Schmidhuber and D.-A. Clevert, J. Cheminf., 2025, 17, 31 CAS.
- Y. Leviathan, M. Kalman and Y. Matias, International Conference on Machine Learning, 2023, pp. 19274–19286 Search PubMed.
- S. Genheden, A. Thakkar, V. Chadimová, J.-L. Reymond, O. Engkvist and E. Bjerrum, J. Cheminf., 2020, 12, 70 Search PubMed.
- T. Cai, Y. Li, Z. Geng, H. Peng, J. D. Lee, D. Chen and T. Dao, arXiv, preprint, arXiv:2401.10774, 2024, DOI:10.48550/arXiv.2401.10774.
- D. P. Kingma and J. Ba, arXiv, 2014, preprint, arXiv:1412.6980, DOI:10.48550/arXiv.1412.6980.
- H. Dai, C. Li, C. Coley, B. Dai and L. Song, Adv. Neural Inf. Process. Syst., 2019, 32, 8872–8882 Search PubMed.
- Z. Zhong, J. Song, Z. Feng, T. Liu, L. Jia, S. Yao, M. Wu, T. Hou and M. Song, Chem. Sci., 2022, 13, 9023–9034 RSC.
- S. Genheden, A. Thakkar, V. Chadimová, J.-L. Reymond, O. Engkvist and E. Bjerrum, J. Cheminf., 2020, 12, 70 Search PubMed.
- A. Kishimoto, B. Buesser, B. Chen and A. Botea, Adv. Neural Inf. Process. Syst., 2019, 32, 7226–7236 Search PubMed.
- M. R. Dobbelaere, I. Lengyel, C. V. Stevens and K. M. Van Geem, J. Cheminf., 2024, 16, 37 Search PubMed.
- S. Chen and Y. Jung, JACS Au, 2021, 1, 1612–1620 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |

![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O)c1ccc2ccccc2.C′ as the draft of 20 tokens for ′CC(
O)c1ccc2ccccc2.C′ as the draft of 20 tokens for ′CC(