Open Access Article
Open Access ArticlePOLARIS: perovskite optimization using LLM-assisted refinement and intelligent screening
Jordan
Marshall†
a,
Sheryl L.
Sanchez†
 a,
Rushik
Desai
b,
Elham
Foadian
a,
Utkarsh
Pratiush
a,
Arun
Mannodi-Kanakkithodi
a,
Rushik
Desai
b,
Elham
Foadian
a,
Utkarsh
Pratiush
a,
Arun
Mannodi-Kanakkithodi
 b,
Sergei V.
Kalinin
a and
Mahshid
Ahmadi
b,
Sergei V.
Kalinin
a and
Mahshid
Ahmadi
 *a
*a
aInstitute for Advanced Materials and Manufacturing, Department of Materials Science and Engineering, The University of Tennessee Knoxville, Knoxville, Tennessee 37996, USA. E-mail: mahmadi3@utk.edu
bSchool of Materials Engineering, Purdue University, West Lafayette, IN 47907, USA
First published on 30th March 2026
Abstract
We present a comprehensive and reproducible pipeline that unites literature mining, molecular graph generation, and uncertainty-aware predictive modeling to accelerate the design of organic spacer cations for two-dimensional (2D) halide perovskites (HPs). Despite the critical influence of spacer chemistry on phase stability, excitonic behavior, transport properties and environmental robustness, the chemical space of HPs remains underexplored due to inconsistent reporting and limited structured datasets. To overcome this, we curated a diverse set of 200 experimental papers from various publishers and research groups into Google's NotebookLM powered by Gemini, utilizing its retrieval-augmented generation (RAG) framework to extract synthesis-relevant metadata with high accuracy and reproducibility. To ensure data quality and consistency, we limited our selection to papers published in peer-reviewed journals with an impact factor above 10, focusing on studies with well-documented experimental protocols. Benchmarking against five other large language models (LLMs) confirmed NotebookLM's superior stability and minimal hallucination rate, making it ideal for hypothesis-driven data curation. From extracted IUPAC names, we constructed SMILES representations and augmented the dataset with over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ammonium-containing molecules from QM9. These were converted into graph-based molecular embeddings and used to train a multitask graph neural network coupled with a Gaussian process (GNN–GP) backend to predict optoelectronic and structural properties with uncertainty quantification. The latent space clustering of the learned embeddings revealed chemically interpretable families of spacer candidates, which we cross-validated against ChatGPT-generated design heuristics. The convergence between unsupervised clustering and transformer-derived guidance highlights the power of combining LLMs with active learning to generate, test, and refine design hypotheses in underexplored chemical domains. This study demonstrates how fragmented literature can be transformed into actionable, structure–property insights through a tightly integrated informatics pipeline available to a broad experimental community, and demonstrates the value of open repositories that can be mined for information. Our approach lays the foundation for closed-loop, autonomous materials discovery and design and provides a scalable strategy for targeted development of next-generation HP optoelectronics.
000 ammonium-containing molecules from QM9. These were converted into graph-based molecular embeddings and used to train a multitask graph neural network coupled with a Gaussian process (GNN–GP) backend to predict optoelectronic and structural properties with uncertainty quantification. The latent space clustering of the learned embeddings revealed chemically interpretable families of spacer candidates, which we cross-validated against ChatGPT-generated design heuristics. The convergence between unsupervised clustering and transformer-derived guidance highlights the power of combining LLMs with active learning to generate, test, and refine design hypotheses in underexplored chemical domains. This study demonstrates how fragmented literature can be transformed into actionable, structure–property insights through a tightly integrated informatics pipeline available to a broad experimental community, and demonstrates the value of open repositories that can be mined for information. Our approach lays the foundation for closed-loop, autonomous materials discovery and design and provides a scalable strategy for targeted development of next-generation HP optoelectronics.
Introduction
Organic spacers fundamentally determine structure–function relationships in two-dimensional (2D) and quasi-2D halide perovskites (HPs), yet molecular mechanisms governing their nucleation pathways, phase formation dynamics, electronic coupling mechanisms, and environmental stability remain inadequately characterized.1,2 These spacer cations introduce critical structural elements through aromatic versus aliphatic backbones, conformational flexibility, hydrogen-bonding capabilities, and permanent dipole orientations that collectively regulate octahedral slab separation distances, preferential crystal orientations, defect passivation efficiencies, and dielectric interfaces.3,4In conventional Ruddlesden–Popper (RP) 2D HP architectures, monovalent organic spacers establish discrete van der Waals gaps between individual octahedral sheets; with increasing octahedral layer indices (n), quasi-2D systems evolve toward multiple-quantum-well configurations exhibiting tailorable exciton confinement properties and directionally-dependent charge transport mechanisms.3,5–7 In contrast, Dion–Jacobson (DJ) phases incorporate divalent cations that form covalent bridges between adjacent slabs. This configuration can enhance mechanical integrity, improve moisture resistance, and, under appropriate processing conditions, promote favorable out-of-plane orientation and charge transport.5,8
While organic spacers play a central role in determining the structure, stability, and optoelectronic behavior of 2D and quasi-2D HPs, systematic exploration of spacer chemistry is hindered by fragmented and inconsistently reported experimental literature. Critical synthesis parameters including precursor stoichiometry, solvent selection, deposition protocols, and thermal processing are often embedded within heterogeneous, narrative Methods sections that vary widely across journals and research groups. Existing literature-mining and information-extraction approaches either rely on rigid document assumptions, require extensive manual configuration, or lack reproducible validation against human-annotated ground truth, limiting their scalability and reliability. More recent prompt-based LLM workflows offer greater flexibility but suffer from session-to-session inconsistency, user-to-user variability, and limited benchmarking of extraction fidelity. These limitations motivated the development of POLARIS: a reproducible, retrieval-augmented literature mining platform designed to systematically extract synthesis-relevant experimental parameters from diverse perovskite publications and to rigorously benchmark extraction performance across models, users, and runs. The primary contribution of this work is not the introduction of a new language model, but the establishment of a defensible, scalable framework for transforming unstructured perovskite literature into reliable, machine-actionable data.
This mechanistic gap, specifically the limited understanding of how spacer cation chemistry governs nucleation pathways, phase formation, electronic coupling, and environmental stability, arises from two interrelated challenges; the vast chemical design space of potential organic spacers, and inconsistent reporting practices across the HP literature.9,10 Substantial variations in precursor stoichiometry, solvent selection criteria, film deposition techniques, and thermal processing protocols significantly complicate cross-study comparisons, thereby impeding development of generalizable design principles.4,9,11 Particularly, odd-even layer number effects12,13 and cooperative hydrogen-bonding networks can profoundly influence phase purity and defect concentrations,7,14–17 yet these phenomena remain inadequately quantified due to heterogeneous experimental methodologies and incomplete metadata reporting.6,10,18
Automated scientific literature analysis has progressed from rule-based systems to modern ML methods, with early approaches relying heavily on topic modeling and named entity recognition (NER).19–24 Tools like Latent Dirichlet Allocation (LDA) enabled coarse clustering of research topics, while domain-specific transformers such as SciBERT and BioBERT significantly improved the extraction of technical terms, chemical names, and entities from biomedical and materials science publications.25–27 BioBERT, trained on over 18 million PubMed abstracts and 750![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 full-text articles, demonstrated substantial improvements over generic language models in biomedical NER tasks, achieving F1 scores exceeding 0.85. The F1 score represents the harmonic mean of precision and recall, balancing how many relevant instances are correctly extracted (recall) against how many extracted instances are actually correct (precision).19,27 Similarly, MatBERT and BatteryBERT established specialized capabilities for materials science domains, with particular strength in identifying synthesis conditions and performance metrics.28,29 In prior literature, XML-based pipelines were commonly used to structure scientific documents for downstream processing.30,31 These formats enabled sentence-level tokenization, allowing rule-based or fine-tuned language models to identify whether specific sentences contained quantitative synthesis data such as precursor ratios or annealing conditions. While effective in constrained environments, such approaches often require strict document formatting and rigid preprocessing protocols. The XML pipelines typically involve aggressive metadata pruning to reduce token count before submission to language models, which is especially important when using API-based services where costs are scaled with input and output size. While this can reduce overhead, it may also discard valuable contextual information.
000 full-text articles, demonstrated substantial improvements over generic language models in biomedical NER tasks, achieving F1 scores exceeding 0.85. The F1 score represents the harmonic mean of precision and recall, balancing how many relevant instances are correctly extracted (recall) against how many extracted instances are actually correct (precision).19,27 Similarly, MatBERT and BatteryBERT established specialized capabilities for materials science domains, with particular strength in identifying synthesis conditions and performance metrics.28,29 In prior literature, XML-based pipelines were commonly used to structure scientific documents for downstream processing.30,31 These formats enabled sentence-level tokenization, allowing rule-based or fine-tuned language models to identify whether specific sentences contained quantitative synthesis data such as precursor ratios or annealing conditions. While effective in constrained environments, such approaches often require strict document formatting and rigid preprocessing protocols. The XML pipelines typically involve aggressive metadata pruning to reduce token count before submission to language models, which is especially important when using API-based services where costs are scaled with input and output size. While this can reduce overhead, it may also discard valuable contextual information.
Despite these advancements, earlier methods have several limitations. They often require extensive manual labeling, retraining on specialized datasets, and complex infrastructure, making them difficult to scale or adapt across diverse literature formats. ChemDataExtractor and similar systems demand significant computational resources and domain-specific ontologies, creating barriers to widespread adoption.32 More critically, these systems lack interactive and modular control and users cannot easily adapt to new research questions or data types without significant reconfiguration. Even minor shifts in task objectives such as transitioning from extracting synthesis methods to identifying degradation pathways often necessitate rewriting prompts, reengineering internal logic, or redeploying entire pipelines.33,34 This rigidity limits their scalability and hinders rapid iteration during exploratory research. Recent work demonstrated that conventional NLP pipelines require approximately 40–60 hours of expert configuration to adapt extraction workflows between closely related materials subdisciplines.35
These constraints make consistent and reproducible information extraction from scientific papers especially challenging in rapidly evolving fields like HP research.31,36 Most recently, LLMs have offered a more flexible and democratized alternative by allowing users to pose natural language questions directly to full-text documents.33,34 When paired with well-structured prompts, these models can extract precise data such as chemical compositions, synthesis conditions, and material names without specialized training.31 However, prompt-based querying introduces its own challenges, for instance, minor changes in phrasing can yield inconsistent or contradictory outputs, a phenomenon known as prompt drift. Previous work shows that variations in prompt temperature, prefix wording, and example ordering can cause output variations exceeding 20% for identical information extraction tasks.37 Iterative refinement and carefully designed prompt templates help reduce this effect. However, ensuring consistent outputs across different sessions and users remains a major challenge. The same prompt can yield different responses when run at various times (session-to-session inconsistency) or by different users (user-to-user variability), making it difficult to reproduce results reliably in collaborative or automated workflows. Earlier research quantified this variability, finding that even advanced prompting strategies such as self-consistency prompting applied to models like ChatGPT produced extraction consistency metrics below 75% when evaluated across multiple sessions and user groups. This underscores the limitations of purely prompt-based approaches and highlights the need for more robust solutions like retrieval augmentation or automated validation mechanisms.38
To address these limitations, we have developed and implemented a scalable literature mining framework leveraging Retrieval-Augmented Generation (RAG) techniques.39 By integrating targeted document retrieval with LLM interrogation protocols, our RAG methodology enables systematic extraction of IUPAC nomenclature, precise precursor formulations, solvent compositions, deposition methodologies, and thermal treatment parameters from extensive publication databases. We have rigorously benchmarked multiple LLM architectures including GPT-4o, NotebookLM, DeepSeek, Elicit, perplexity which uses different LLMs but selects the best model based on the given prompt, and Publication Analyzer against manually curated datasets to quantify extraction accuracy, session consistency, and prompt robustness.
In this study, all references to ChatGPT correspond specifically to the GPT-4o model accessed through the ChatGPT interface, while all benchmarking experiments utilized the GPT-4o to ensure consistent versioning and reproducibility across extraction runs. To maintain uniformity in model attribution, we adopt standardized naming throughout the manuscript for all evaluated LLM architectures, including GPT-4o, DeepSeek-V3, Perplexity, Elicit, NotebookLM, and Publication Analyzer. NotebookLM, used as the primary tool for large-scale literature extraction, refers to the Gemini-based retrieval-augmented platform available during February–May 2025. Its implementation in this study included a 50-document notebook capacity, document-grounded retrieval, and a conservative extraction strategy designed to minimize hallucinations while preserving contextual fidelity, as described in the literature-mining section. Publication Analyzer refers to a custom GPT-based system developed for structured literature mining within the ChatGPT environment. It is designed to process uploaded PDFs, perform section-aware extraction (e.g., abstract, methods, results), and generate grounded summaries and structured outputs while avoiding unsupported information. The system supports configurable analysis depth (summary-only, standard, or exhaustive), phrase- and token-level extraction relevant to perovskite materials, and cross-document comparison to identify recurring themes and patterns. It further incorporates retrieval-augmented generation to keep outputs traceable to the uploaded sources. Importantly, in the present implementation, Publication Analyzer operates within the standard ChatGPT document-upload constraints (up to 10 files per run/session), and thus was used as a controlled, text-grounded comparator rather than a truly unlimited-corpus mining system. For clarity, we further distinguish between LLMs used for source-grounded extraction and benchmarking (GPT-4o, NotebookLM, DeepSeek-V3, perplexity, Elicit, and Publication Analyzer) and the ChatGPT interface, which was used exclusively for hypothesis generation and qualitative reasoning tasks. All figures, tables, and model descriptions have been updated to reflect this consistent nomenclature and usage.
The present literature-mining workflow is restricted to text-based extraction, and all benchmarking analyses were conducted on information contained within the main body of each manuscript, including experimental sections, results descriptions, and chemical nomenclature. Although modern vision-language models permit multimodal extraction from figures, tables, and schematics, we focus here on text-only inputs to ensure reproducibility, controlled model comparison, and consistent ground-truth evaluation. The 200-paper dataset consists of PDFs drawn from a diverse range of publication venues, selected to reflect heterogeneity in document structure, formatting, and reporting conventions rather than the absence of figure-embedded data. The targeted metadata precursor formulations, deposition protocols, solvent systems, temperature-time profiles, and IUPAC nomenclature are typically reported explicitly in text across journals, enabling direct and traceable comparison of extraction performance across large language model architectures. As a result, some quantitative or graphical information available only in figures, tables, or schematics lies outside the scope of the current pipeline. Extending the framework to incorporate multimodal extraction remains an important direction for future development and would enable broader capture of synthesis parameters and structure–property relationships reported visually, as well as phrase- and token-level information currently inaccessible through text alone. Concurrently, we evaluated prompt drift phenomena through systematic variation of query phrasing and implemented iterative prompt refinement workflows, with each prompt validation conducted against representative literature samples (Table S1), ensuring reproducible performance across heterogeneous reporting contexts.38
Our comprehensive platform integrates four functional modules, as schematically illustrated in Fig. 1, to enable a closed-loop system for data-driven spacer discovery in HPs. Fig. 1a presents the overarching workflow, which begins with literature mining facilitated by LLMs to extract structured metadata on synthesis conditions, chemical compositions, performance outcomes, and spacer structure–property relationships. This metadata provides the basis for hypothesis generation and candidate selection. The subsequent digital processing stage, shown in Fig. 1b, converts the extracted synthetic protocols into standardized molecular formats such as SMILES strings, which are then analyzed using RDKit and enriched with quantum-mechanical featurization to yield property-relevant molecular descriptors. Fig. 1c highlights the high-throughput experimental platform, where automated synthesis and optical characterizations are employed to systematically evaluate perovskite–spacer formulations under controlled conditions. Finally, Fig. 1d depicts the automated data analysis and model training pipeline, which incorporates spectral deconvolution, hierarchical clustering, acquisition function optimization, and multidimensional visualization. These processed outputs inform a deep-kernel learning (DKL) algorithm that drives iterative candidate refinement, completing the self-improving discovery loop.40–43 The automated synthesis, characterization, and active learning-based ML methods as illustrated in Fig. 1 are well established in our group; a recent addition to our workflow is the use of LLMs for literature curation and hypothesis generation, leveraging the rich scientific knowledge embedded in published research.44–47 While Fig. 1 illustrates a closed-loop integration of these components in our research, the present work focuses specifically on development and validation of the LLM-driven literature mining and information extraction methodologies. Through demonstration of scalable, reproducible data curation workflows, we establish foundational capabilities for future expansion toward fully autonomous, closed-loop discovery platforms targeting robust, high-efficiency HPs for different functionality.
Results and discussion
Fig. 2 details the end-to-end workflow established in this study to systematically benchmark and validate prompt-based large-language models (LLMs) for extracting critical synthesis metadata from 2D and quasi 2D HPs research publications. The heterogeneity of journal formats, stylistic variations in methodological descriptions, and the proliferation of synonyms for identical reagents and processes pose significant challenges for manual or rule-based text-mining approaches. Our pipeline addresses these challenges by emphasizing reproducibility, scalability, and modularity through a series of interconnected stages including prompt formulation, ground-truth annotation, performance metric computation, and model selection. In contrast to early research, our methodology leverages Retrieval-Augmented Generation (RAG) with large language models (LLMs), eliminating the need for XML parsing and enabling more flexible, prompt-driven interaction with heterogeneous document formats. In this study, our approach instead applies to strategic prompt engineering and targeted section filtering to optimize information density while preserving context. Importantly, OpenAI's currently charges for usage of their API.48 This makes high-throughput extraction workflows significantly more expensive than flat-rate ChatGPT subscriptions. However, API access affords critical advantages, including reproducibility across runs, consistent model versioning, and full control over prompt architecture capabilities that are essential for scalable, automated extraction pipelines. Ultimately, the framework studied here provides a modern, modular, and source-grounded alternative to traditional XML-based approaches. By removing reliance on rigid document structures and enabling controlled LLM interrogation, our scalable strategy balances performance, interpretability, and operational cost supporting rigorous, high-integrity data curation in materials science domains. As shown in Table S2, our approach achieves higher recall and improved extraction stability, particularly when applied across diverse publishing styles and experimental reporting formats. Our framework is designed with modularity and extensibility in mind, because we integrate a retrieval-augmented LLM environment (e.g., NotebookLM) with structured metadata indexing, researchers can flexibly reframe queries, substitute context documents, or reconfigure downstream featurization and modeling components without altering the overall architecture. This enables rapid adaptation to evolving research goals with minimal system-level changes.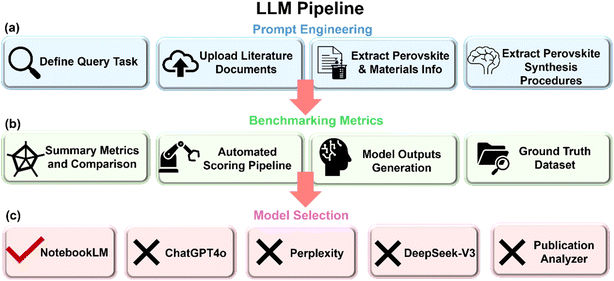 | ||
| Fig. 2 (a) Query-driven LLM workflow: ingest documents, structured prompt interaction, iterative refinement, (b) benchmarking and descriptor output, and (c) model selection. | ||
In the prompt formulation stage, we developed an extensible library of queries targeting key experimental parameters containing precursor stoichiometries, solvent compositions, deposition techniques (e.g., spin-coating vs. drop-casting), annealing temperatures and time, and auxiliary processing steps (e.g., ligand addition, aging protocols, etc.). Each prompt was iteratively refined over multiple cycles using a representative subset of articles to resolve ambiguities arising from chemical nomenclature (e.g., “DMF” vs. “dimethylformamide”), nested procedural narratives, and overlapping domain-specific terminology. Refinements included adding context-specific qualifiers, adjusting prompt structure to focus on paper section headings such as extracting the results and experimental sections, and implementing fallback patterns to capture unexpected phrasing.
Concurrently, we assembled a ground-truth (GT) dataset by manually annotating ten HPs studies selected for their diversity in reporting style (ranging from terse bullet-point protocols to narrative experimental sections). Annotators recorded all relevant materials, operational parameters, and processing conditions, then supplemented each entry with synonyms, abbreviations, and alternate spellings in a structured excel database. For robust evaluation, we defined two critical performance metrics. The first metric is GT phrase recall (%) and the second is GT token recall (%). The GT phrase recall (%) measures the fraction of multi-word ground-truth phrases (complete terminological units) successfully retrieved by each model. This metric assesses the model's ability to capture compound technical terms intact, for example, identifying “methylammonium lead iodide” as a complete entity rather than fragmentary components. GT token recall (%) quantifies the proportion of individual tokens (words, subwords, or the smallest unit of language) from ground-truth phrases that appear in the model outputs. This more granular measure accommodates partial matches, revealing cases where models capture essential information but with minor lexical variations or incomplete terminology. For ground-truth benchmarking, 10 papers were randomly selected from the 200-paper dataset, with the sole requirement being the presence of a clearly identifiable Methods section to enable consistent manual annotation of synthesis-relevant metadata. To ensure diversity in reporting formats, each of the 10 papers originated from a different journal, thereby capturing variation in structure, terminology, and experimental narrative style. This random, cross-journal sampling provides a representative evaluation of extraction performance across heterogeneous literature sources. Although this approach offers broad coverage, a more systematic stratification of papers by extraction difficulty such as contrasting journals with highly structured Methods sections against those with more integrated or narrative-style reporting would enable a more explicit assessment of model robustness. We therefore identify difficulty-aware benchmarking as an important direction for future refinement of the evaluation framework. All documents in this study were processed as directly downloaded PDF files, and no XML or HTML sources were used in any stage of the pipeline. As a result, the workflow does not depend on journal-specific markup conventions or structural schemas, and model performance reflects the LLMs' ability to interpret text extracted from heterogeneous PDF layouts rather than publisher metadata formats. The 200-paper dataset encompasses a wide range of journals with differing formatting styles, section structures, and typographic conventions, enabling evaluation of extraction robustness under realistic variability. Across this corpus, models such as NotebookLM and Publication Analyzer demonstrated stable, source-grounded extraction independent of PDF layout, whereas models like GPT-4o and perplexity exhibited greater sensitivity to formatting differences and narrative structure. This PDF-based design ensures that generalizability assessments center on the LLMs' text-processing capabilities rather than on variations in publisher markup systems.
For performance benchmarking, six LLM architectures including GPT-4o, NotebookLM, DeepSeek-V3, Elicit, perplexity which uses different LLMs and chooses the best model for the given prompt, and Publication Analyzer were evaluated on the ground-truth dataset of literature across three repeated executions of identical queries. The multiple runs were conducted to ensure that prior interactions or user history from different accounts did not influence the outcomes, thereby maintaining consistency and fairness in model comparisons. We captured raw outputs and associated metadata for each run, including character count, word count, line count, and a cleaned version of the response (normalized by lowercasing, punctuation removal, and whitespace standardization). This preprocessing ensured fair comparison across models and eliminated artifacts arising from formatting differences.
We computed six quantitative performance measures for each model-run combination. Beyond GT phrase recall and GT token recall, we measured GT token precision (fraction of output tokens matching the ground-truth tokens), the composite GT token F1 score balancing precision and recall, cosine similarity between vector embeddings of ground truth and model responses, and verbosity ratio (response length relative to ground-truth annotation length). Model-prompt pairings were ranked by F1 score, with statistical significance across runs evaluated using paired t-tests (p < 0.05). Further information on benchmarking metrics calculations can be found in SI, Note S1.
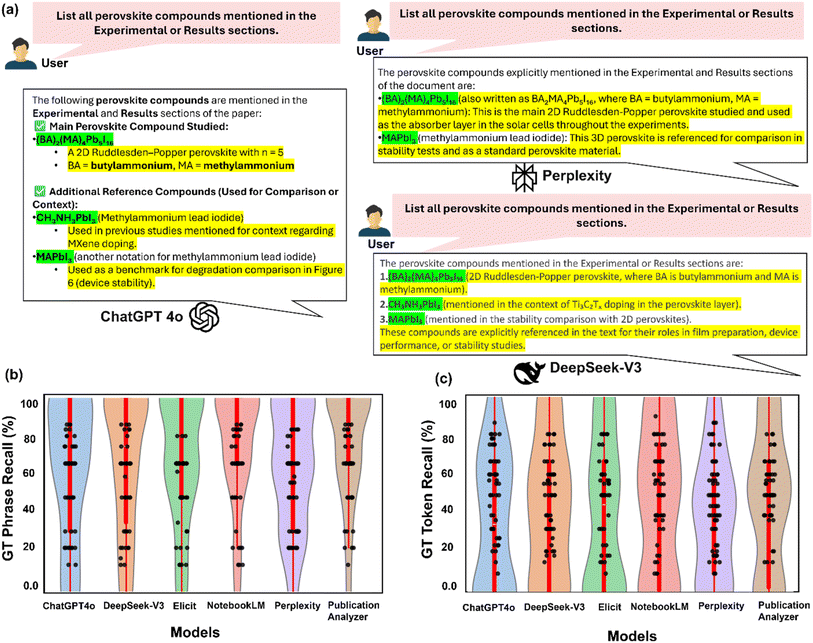
Fig. 3a demonstrates marked differences in response structure and content when identical prompts were submitted to three different LLMs. For example, when tasked with “List all perovskite compounds mentioned in the Experimental or results sections,” models exhibited distinctive response patterns. ChatGPT produced structured outputs with categorical organization, correctly identifying “(BA)2(MA)4Pb5I16” as the main compound while including reference materials like “CH3NH3PbI3” with explanatory notes about their contextual relevance. Perplexity generated verbose explanations, elaborating on chemical notations (e.g., noting that “(BA)2(MA)4Pb5I16” can be written as “BA2MA4Pb5I16”) and providing functional descriptions of each compound's role in experimental workflows. DeepSeek offeres concise, enumerated lists prioritizing factual extraction over contextual elaboration, maintaining a focus on direct identification of perovskite compounds while minimizing ancillary information.
The violin plots in Fig. 3b and c reveal distinctive performance profiles across models. For GT phrase recall (Fig. 3b), Publication Analyzer and NotebookLM achieved the highest median values (85–95%), indicating consistent retrieval of complete technical terms across papers and runs. The narrow distribution width for these models demonstrates remarkable stability across diverse document formats and reporting styles. Elicit exhibited intermediate performance across both GT phrase recall and token recall metrics, with median values around 70% and 50% respectively. While its recall rates surpassed those of perplexity, Elicit showed considerable variance and lacked the consistency observed in top-performing models. In contrast, ChatGPT and DeepSeek displayed lower median values (65–80%) with elongated distribution tails, suggesting inconsistent performance with pronounced paper-to-paper and run-to-run variability. Perplexity exhibited the lowest median phrase recall (∼65%) and widest distribution, highlighting substantial instability in technical term extraction. GT token recall measurements (Fig. 3c) reveal a systematic performance reduction across all models compared to phrase-level metrics, emphasizing the challenges of retrieving complete term-based inventory without omissions or substitutions. NotebookLM and Publication Analyzer maintained relatively better performance (55–75% median token recall) despite this general trend. The decline was most pronounced for perplexity, with median token recall falling to approximately 45%, indicating frequent omission or misrepresentation of individual terms even when broader concepts were identified. In contrast to models that occasionally produced higher raw precision scores through unsupported assumptions or inferred content, NotebookLM delivered stable, grounded, and highly reproducible extractions. Its deliberately conservative extraction strategy limits outputs to content explicitly present in the source text. This restraint virtually eliminated hallucinations and ensured traceability, aligning well with our objective of verifiable, hypothesis-driven discovery. Its lower precision score, relative to some models, reflected cautious omissions rather than factual inaccuracies, which is a favorable trade-off for downstream modeling applications where erroneous inputs can propagate and amplify modeling errors.
Along with the results of Fig. 3, and S1 presents four supporting plots that offer a comprehensive comparison of the five LLMs under evaluation in this study. Fig. S1a shows the total number of extracted phrases per model, categorized by extraction quality with fully correct in (blue), partially correct in (red), and incorrect or hallucinated in (cyan) color bars. While each model extracted a similar total number of phrases, there are subtle but important differences in quality. NotebookLM and DeepSeek-V3 stand out with a larger proportion of fully correct extractions and fewer incorrect responses, suggesting they maintain a better balance between recall and precision. For NotebookLM, this performance likely reflects its document-grounded architecture, such as a retrieval-augmented generation (RAG) system that helps prevent hallucination. In contrast, perplexity shows the lowest number of fully correct extractions and the largest number of incorrect or hallucinated outputs, indicating poor precision. ChatGPT4o and Publication Analyzer fall in between. ChatGPT4o tends to produce more partially correct (but incomplete or paraphrased) extractions, while Publication Analyzer leans toward more incorrect outputs, likely due to overgeneration or loose alignment with the source content. Models that produce longer responses naturally generate larger absolute numbers of both correct and incorrect phrases, Fig. S1a alone cannot be used to assess extraction fidelity. Instead, relative metrics including GT phrase recall, GT token recall, and coefficient of variation across runs provide the appropriate basis for comparison and indicate that NotebookLM and DeepSeek exhibit lower hallucination rates and more stable performance than other models. Although Publication Analyzer shows comparable recall values, its use is constrained by a 10-document upload limit and attempts to exceed this capacity within a single chat session resulted in loss of document tracking, duplication of extracted content, and increased hallucination frequency in multi-document contexts. Representative verbatim extraction outputs from all evaluated models in response to an identical prompt are provided in Note S3.
Fig. S1b presents the coefficient of variation (CV) for each model, a metric reflecting output stability across multiple benchmarking runs. Lower CV values indicate that a model performs consistently across runs, while higher CV values suggest that performance varies significantly depending on prompt phrasing, context, or randomness. Publication Analyzer exhibits the lowest CV, making it the most stable model. Its output is reproducible across sessions, which is especially important for high-integrity extraction workflows. NotebookLM and perplexity follow, with moderate CV values indicating reasonable but imperfect consistency. DeepSeek-V3 and especially ChatGPT4o show the highest CVs, highlighting substantial run-to-run variability despite solid average performance in other metrics. This suggests that even if these models perform well in some runs, they may be unreliable or sensitive to slight prompt variations. Fig. S1c explores the efficiency trade-off between verbosity and GT phrase recall. Ideally, a model should extract correct phrases with as little output as possible. In this plot, Publication Analyzer emerges as the most efficient model, achieving the highest GT phrase recall with the lowest verbosity, meaning it says less but hits the ground truth more often. NotebookLM ranks second in phrase recall but is the most verbose, suggesting that while it successfully identifies many correct phrases, it often produces long or redundant responses to get there. DeepSeek-V3 and perplexity show similar performance profiles, with relatively high verbosity but lower GT phrase recall, indicating a less efficient extraction style. Finally, ChatGPT4o performs the worst on this metric, despite having lower verbosity than some other models, it recovers the fewest ground truth phrases, revealing inefficiency in both content selection and phrasing. Lastly, Fig. S1d displays the total number of GT phrases missed by each model. As expected from earlier plots, Publication Analyzer misses the fewest phrases overall, followed by NotebookLM, then DeepSeek-V3, with Perplexity and ChatGPT4o missing the most. This plot reinforces earlier findings about each model's recall strength and further confirms that Publication Analyzer and NotebookLM are the strongest performers in terms of retrieving relevant, source-grounded information from full-text documents.
These benchmarking results highlight a critical distinction in LLM performance for scientific information extraction. The models optimized for general conversational tasks often prioritize comprehensive responses over factual precision, introducing potential confounds through elaboration and inference. In contrast, models like NotebookLM that prioritize source fidelity and output stability demonstrate superior performance for literature mining in sensitive scientific domains, particularly when integrated within RAG frameworks that emphasize verifiable extraction over generative completeness. Based on these findings, NotebookLM was selected for literature extraction because its RAG-based architecture minimizes hallucinations and preserves source fidelity, as confirmed by benchmarking (Fig. 3 and Table S2). For hypothesis generation, ChatGPT was used because it consistently produced chemically interpretable heuristic rules when prompted in a generative setting. The two LLMs serve distinct roles factual extraction versus design ideation and benchmarking showed no single model performed both tasks optimally.
To support large-scale extraction across the full 200-paper dataset, we therefore employed NotebookLM, which accommodates up to 50 documents per notebook, maintains internal document provenance, and avoids cross-document drift through its retrieval-grounded architecture. All large-language-model interactions in this study were performed through freely accessible or institutionally licensed user interfaces rather than metered APIs, the pipeline incurred no token-based usage costs during benchmarking or large-scale extraction. This distinction is important, as the economic feasibility of LLM-enabled workflows depends strongly on whether models are accessed through subscription-based interfaces or usage-metered APIs. In the context of this work, NotebookLM operates under a flat-rate model with document-number limits, while perplexity, Elicit, and Publication Analyzer can be used at no direct cost for moderate-scale extraction tasks. GPT-4o only introduces substantial expense when accessed via the API for automated high-throughput extraction; however, such API-based workflows were not employed here. As a result, the present pipeline can be replicated without significant financial barriers, though future deployments that incorporate automated or multimodal extraction would require careful cost-performance considerations. The impact factor >10 criterion used to construct the 200-paper corpus reflects the publication landscape of the halide perovskite community, where the most methodologically rigorous studies and the most complete synthesis metadata are concentrated in high-impact journals that enforce standardized reporting practices. In this field, the threshold therefore serves as a practical quality filter rather than a general measure of prestige, ensuring that extracted parameters such as precursor stoichiometry, solvent composition, and thermal processing conditions are documented with sufficient clarity for benchmarking. However, reliance on impact factor alone can introduce selection bias, and influential contributions may also appear in high-quality mid-impact venues. Future extensions of this dataset may incorporate additional metrics, such as citation counts, citation-rate normalization, or composite scoring schemes, to produce a more comprehensive and representative literature set while maintaining methodological rigor. The 200 papers ingested into NotebookLM were divided into four batches of 50 documents each through simple random grouping, reflecting the platform's notebook capacity limit rather than a cross-validation or stratified sampling procedure. This random partitioning ensured that each batch contained a heterogeneous mixture of journals, reporting styles, and experimental formats representative of the broader dataset. NotebookLM processes each notebook independently using its retrieval-grounded mechanism, and extraction stability was evaluated by comparing performance across these four randomly assembled groups. Although this approach was sufficient for assessing batch-to-batch consistency, more formal grouping strategies such as difficulty-aware stratification may be incorporated in future iterations of the workflow. This approach aligns with how typical researchers without premium subscriptions would utilize the platform, providing insight into real-world applicability of our methodology. Through systematic application of our validated prompts across these paper batches, we constructed a comprehensive database of perovskite organic spacers that have been researched, synthesized, or are commercially available. This dataset forms the foundation for subsequent analysis of structure–property relationships and identification of promising candidates for further experimental investigation. For this list of organic spacers, we performed density functional theory (DFT) computations to calculate their HOMO–LUMO gaps and dipole moments. These properties are then utilized in the next exercise on molecular graph-based data visualization.
To address the limited size of our organic spacer dataset (106 molecules), we augmented it with over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ammonium-containing molecules from the QM9 database, processed using identical featurization protocols. We supplemented our organic spacer dataset with 10
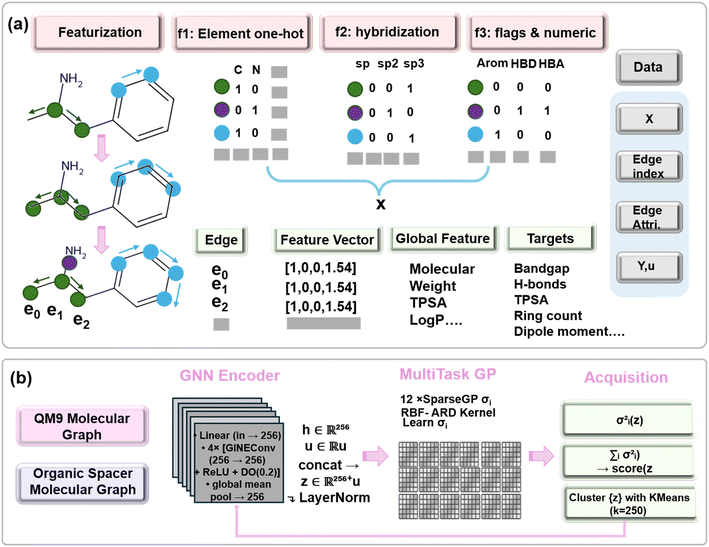
000 ammonium-containing molecules from the QM9 database, processed using identical featurization protocols. We supplemented our organic spacer dataset with 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 QM9 molecules to reduce the risk of severe overfitting and poor generalization when training the GNN model. To ensure that the augmented dataset reflects the chemistry of experimentally reported spacer cations, we filtered QM9 to retain only molecules containing protonated or protonatable amine groups. All 106 literature-extracted spacers contain ammonium motifs (primary, secondary, cyclic, or aromatic), and restricting QM9 accordingly produces a chemically coherent training set. Without this filter, the GNN is exposed to structures (e.g., unfunctionalized hydrocarbons, aldehydes, ketones, carboxylates) that never appear in perovskite spacers and would introduce irrelevant inductive biases. By utilizing the QM9 database we can expose the model to a higher variety of graph topologies and chemical environments. This approach can help us regularize the multi-task process, since related property predictions on the QM9 subset can help constrain the shared network layers and improve the performance on the perovskite targets. Although the QM9-derived molecules numerically dominate the dataset, they primarily serve to inform the GNN's representation learning, whereas the literature-derived perovskite spacers provide the high-fidelity experimental signal that anchors the uncertainty-aware regression and downstream analysis. This pipeline constructs a graph-based molecular dataset suitable for training GNNs using molecular representations derived from SMILES strings shown in Fig. 4a. Each SMILES string is parsed into an RDKit Mol, sanitized, and explicitly hydrogenated before Gasteiger partial charges are computed. We construct an unweighted adjacency matrix and, via NetworkX, compute shortest-path distances from the charged ammonium nitrogen to every atom. Atom-level embeddings (x) concatenate element one-hot vectors (H, C, N, O, F, P, S, Cl, Br, I), atomic degree, formal charge, implicit valence, hybridization one-hots (SP/SP2/SP3/SP3D/SP3D2), aromaticity flags, SMARTS-derived hydrogen-bond donor/acceptor indicators, Gasteiger charge, van der Waals radius, atomic mass, Pauling electronegativity, nitrogen-distance, and ring-membership counts (sizes 3–6). For each directed bond, edge_attr encodes bond order, aromaticity, conjugation, stereochemistry (none/any/Z/E), and estimated bond length (Å). Global descriptors including molecular weight, octanol–water partition coefficient (log
000 QM9 molecules to reduce the risk of severe overfitting and poor generalization when training the GNN model. To ensure that the augmented dataset reflects the chemistry of experimentally reported spacer cations, we filtered QM9 to retain only molecules containing protonated or protonatable amine groups. All 106 literature-extracted spacers contain ammonium motifs (primary, secondary, cyclic, or aromatic), and restricting QM9 accordingly produces a chemically coherent training set. Without this filter, the GNN is exposed to structures (e.g., unfunctionalized hydrocarbons, aldehydes, ketones, carboxylates) that never appear in perovskite spacers and would introduce irrelevant inductive biases. By utilizing the QM9 database we can expose the model to a higher variety of graph topologies and chemical environments. This approach can help us regularize the multi-task process, since related property predictions on the QM9 subset can help constrain the shared network layers and improve the performance on the perovskite targets. Although the QM9-derived molecules numerically dominate the dataset, they primarily serve to inform the GNN's representation learning, whereas the literature-derived perovskite spacers provide the high-fidelity experimental signal that anchors the uncertainty-aware regression and downstream analysis. This pipeline constructs a graph-based molecular dataset suitable for training GNNs using molecular representations derived from SMILES strings shown in Fig. 4a. Each SMILES string is parsed into an RDKit Mol, sanitized, and explicitly hydrogenated before Gasteiger partial charges are computed. We construct an unweighted adjacency matrix and, via NetworkX, compute shortest-path distances from the charged ammonium nitrogen to every atom. Atom-level embeddings (x) concatenate element one-hot vectors (H, C, N, O, F, P, S, Cl, Br, I), atomic degree, formal charge, implicit valence, hybridization one-hots (SP/SP2/SP3/SP3D/SP3D2), aromaticity flags, SMARTS-derived hydrogen-bond donor/acceptor indicators, Gasteiger charge, van der Waals radius, atomic mass, Pauling electronegativity, nitrogen-distance, and ring-membership counts (sizes 3–6). For each directed bond, edge_attr encodes bond order, aromaticity, conjugation, stereochemistry (none/any/Z/E), and estimated bond length (Å). Global descriptors including molecular weight, octanol–water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), and detailed ring statistics (total, aromatic, aliphatic counts; maximum, mean, and per-size counts for rings 3–8) form the graph-level feature vector u. Finally, each data object carries an eleven-dimensional target vector y, in order: band_gap, HOMO energy, LUMO energy, dipole_moment, TPSA, total ring count (n_rings), hydrogen-bond donor count (n_hbd), hydrogen-bond acceptor count (n_hba), conjugated bond count (n_conj_bonds), molecular_weight, and log
P), and detailed ring statistics (total, aromatic, aliphatic counts; maximum, mean, and per-size counts for rings 3–8) form the graph-level feature vector u. Finally, each data object carries an eleven-dimensional target vector y, in order: band_gap, HOMO energy, LUMO energy, dipole_moment, TPSA, total ring count (n_rings), hydrogen-bond donor count (n_hbd), hydrogen-bond acceptor count (n_hba), conjugated bond count (n_conj_bonds), molecular_weight, and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P. By training the shared GNN encoder and sparse-GP heads on both QM9 and perovskite graphs leveraging the broad chemical diversity of QM9 we regularize multi-task learning and significantly improve predictive accuracy on perovskite-specific properties. These properties were chosen to capture key aspects of spacer-inorganic interactions, such as orbital alignment,5,49 dielectric screening,5,49 steric hindrance,50,51 hydrogen bonding capacity52,53 and solubility, which are factors that are known to influence octahedral connectivity,54 defect passivation,55 film morphology56 and charge transport57 in HPs. Each molecule is thus represented as a PyTorch Geometric Data object.
P. By training the shared GNN encoder and sparse-GP heads on both QM9 and perovskite graphs leveraging the broad chemical diversity of QM9 we regularize multi-task learning and significantly improve predictive accuracy on perovskite-specific properties. These properties were chosen to capture key aspects of spacer-inorganic interactions, such as orbital alignment,5,49 dielectric screening,5,49 steric hindrance,50,51 hydrogen bonding capacity52,53 and solubility, which are factors that are known to influence octahedral connectivity,54 defect passivation,55 film morphology56 and charge transport57 in HPs. Each molecule is thus represented as a PyTorch Geometric Data object.
Our multitask Deep Kernel Learning (DKL) framework integrates a graph neural network (GNN) encoder with a sparse Gaussian process (GP) backend to generate expressive molecular representations alongside principled uncertainty estimates. Each molecular graph is encoded with atom-level features (element identity, hybridization, formal charge), bond-level attributes (bond type, estimated bond length, aromaticity, and conjugation), and global molecular descriptors, all assembled into a unified graph data structure. The encoder projects node features into a 256-dimensional hidden space, followed by four Graph Isomorphism Network (GINE) convolutional layers. Nonlinear activation via ReLU and dropout (20%) are applied after each layer to introduce regularization. The final node embeddings are aggregated using global mean pooling to produce a fixed-size graph-level representation. This vector is concatenated with global molecular descriptors and normalized through layer normalization, yielding a composite latent representation that combines local structural detail with holistic molecular information.
A related approach employing DKL for molecular representation learning was recently explored by Ghosh et al.,58 and other similar deep learning framworks59 who demonstrated that active learning with DKL on the QM9 dataset can produce latent spaces that capture molecular functionality more effectively than variational autoencoders (VAEs). Their model used SELFIES-based molecular embeddings as input and showed that DKL's structure–property correlations result in more interpretable and task-aligned latent spaces. Notably, they observed that predictive uncertainty correlated well with error, and that latent spaces contained concentrated maxima associated with specific functionalities. Their findings highlight the utility of DKL in guiding active molecular discovery through meaningful organization of chemical space. While their embeddings were derived from string-based molecular encodings, our framework instead uses graph-based message passing and handcrafted global descriptors to provide a chemically-informed latent representation. Nonetheless, both approaches demonstrate the value of DKL in learning predictive and interpretable molecular embeddings within active learning pipelines.
The sparse GP backend constructs an individual probabilistic regression head for each of the 11 target properties. Each head is instantiated with 128 learnable inducing points in the latent space and utilizes an RBF kernel with automatic relevance determination to capture feature-wise length scales. During training, the GP heads maximize variational evidence lower bound (ELBO) under a Gaussian likelihood, enabling efficient learning of both predictive means and covariances. We jointly optimize the GNN encoder and all GP parameters via stochastic gradient descent using the Adam optimizer with a learning rate of 1 × e−3, applying early stopping based on validation root-mean-squared error and patience of 30 epochs. To accelerate data-efficient learning, we embed this GNN-GP model within an active learning loop. Starting from a small seed set stratified by perovskite versus non-perovskite samples, we iteratively train the model, query posterior variances for unlabeled graphs, and select high-uncertainty candidates via minibatch K-means clustering in the latent space shown in Fig. 4b. During each acquisition iteration, candidate molecules are clustered in the identical latent space that the Gaussian-process regression heads receive as input. The graph neural network encoder produces a 256-dimensional embedding h for each molecule by mean-pooling the final node representations. This learned vector is concatenated with the handcrafted global descriptor vector u which contains properties such as molecular weight, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, and ring statistics yielding a composite representation
P, and ring statistics yielding a composite representation  , where g is the length of u. A feature-wise layer normalization is then applied to standardize scale across dimensions. The set of z vectors for all molecules still in the pool is supplied directly to the K-means algorithm without any intermediate dimensionality-reduction step. Because clustering is performed on this fully normalized concatenation of learned and fixed features, the acquisition strategy probes the same representation that determines both predicted means and posterior variances, ensuring that selected molecules occupy genuinely uncertain regions of the model's feature space. This acquisition strategy balances exploration and exploitation, adding diverse, information-rich samples to the training pool. At each iteration, we record per-task R2, RMSE on a held-out test set, and model training loss, allowing continuous monitoring of performance gains. The resulting pipeline delivers calibrated predictions and uncertainty quantification, facilitating robust property estimation and informed decision-making in high-throughput HP discovery.
, where g is the length of u. A feature-wise layer normalization is then applied to standardize scale across dimensions. The set of z vectors for all molecules still in the pool is supplied directly to the K-means algorithm without any intermediate dimensionality-reduction step. Because clustering is performed on this fully normalized concatenation of learned and fixed features, the acquisition strategy probes the same representation that determines both predicted means and posterior variances, ensuring that selected molecules occupy genuinely uncertain regions of the model's feature space. This acquisition strategy balances exploration and exploitation, adding diverse, information-rich samples to the training pool. At each iteration, we record per-task R2, RMSE on a held-out test set, and model training loss, allowing continuous monitoring of performance gains. The resulting pipeline delivers calibrated predictions and uncertainty quantification, facilitating robust property estimation and informed decision-making in high-throughput HP discovery.
Fig. S2 presents the performance evolution across 30 active learning iterations for eleven molecular targets using a multi-task DKL framework with graph neural network encoding. Performance is assessed through R2 and RMSE heatmaps evaluated against a held-out test set throughout the training process. The results reveal distinct learning trajectories that correlate with the underlying nature of each molecular property. Discrete structural descriptors (n_rings, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) p, and n_conj_bonds) demonstrate rapid convergence to high predictive accuracy (R2 ≥ 0.9), reflecting the model's capacity to effectively encode and quantify graph-topological features through message-passing mechanisms. In contrast, continuous quantum mechanical properties (band_gap, HOMO, and LUMO) exhibit more gradual improvement trajectories, ultimately achieving robust performance (R2 ≈ 0.8–0.9). This delayed convergence is consistent with the computational complexity of electronic structure properties, which require the model to learn subtle orbital interactions and electronic correlations that benefit substantially from iterative data acquisition.
p, and n_conj_bonds) demonstrate rapid convergence to high predictive accuracy (R2 ≥ 0.9), reflecting the model's capacity to effectively encode and quantify graph-topological features through message-passing mechanisms. In contrast, continuous quantum mechanical properties (band_gap, HOMO, and LUMO) exhibit more gradual improvement trajectories, ultimately achieving robust performance (R2 ≈ 0.8–0.9). This delayed convergence is consistent with the computational complexity of electronic structure properties, which require the model to learn subtle orbital interactions and electronic correlations that benefit substantially from iterative data acquisition.
The dipole_moment prediction presents a notable limitation, plateauing at R2 ≈ 0.5 across all learning rounds. This performance ceiling likely stems from the fundamental mismatch between the property's geometric dependence and the model's reliance on 2D molecular representations. Dipole moments are inherently sensitive to three-dimensional molecular conformations and spatial charge distributions information that remains inaccessible to graph-based encoders operating solely on connectivity patterns. The observed learning dynamics for LUMO further support the presence of inter-task dependencies, as its improvement appears contingent upon homo stabilization, suggesting beneficial knowledge transfer through the shared encoder architecture.
A striking asymmetry emerges in hydrogen bonding predictions: while n_hbd achieves near-perfect accuracy (R2 ≈ 1.0), n_hba performance remains consistently poor. This disparity likely reflects the structural unambiguity of hydrogen bond donors typically manifested as explicit –OH or –NH functional groups compared to the more complex and context-dependent nature of acceptor sites, which encompass diverse electron-rich moieties that may not be readily captured by current message-passing protocols.
TPSA and molecular_weight predictions exhibit distinct behavior, showing minimal improvement across active learning rounds despite being deterministic functions of molecular structure. This observation suggests that these properties may require different representational approaches than those naturally captured by the current graph neural network architecture. The apparent difficulty in learning these targets likely reflects the need for more specialized feature extraction mechanisms or alternative inductive biases that better align with the computational pathways required for these specific molecular descriptors.
These findings demonstrate the selective efficacy of uncertainty-based acquisition functions, which effectively prioritize tasks exhibiting both measurable epistemic uncertainty and favorable representational alignment (e.g., band_gap, HOMO). The pronounced improvement in specific tasks validates the efficiency of task-aware uncertainty sampling and highlights opportunities for targeted architectural enhancements. Properties requiring global structural information or geometric features present natural directions for methodological advancement through enhanced input representations such as three-dimensional molecular geometries or physics-informed descriptors complementing the existing framework. Future iterations could benefit from task-conditioned feature augmentation strategies to expand the model's representational capacity for a broader range of molecular targets, potentially through specialized encoder branches or multi-modal input processing.
The t-SNE projections and unsupervised clustering of molecular embeddings reveal that the Graph Neural Network (GNN) has learned chemically meaningful representations across a combined dataset comprising over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ammonium-containing molecules from QM9 and 106 experimentally studied perovskite organic spacers. The two-dimensional embeddings show distinct and compact clusters, indicating that molecules with similar chemical features and functional groups are positioned adjacently in the learned latent space. Notably, the GNN model successfully differentiates between the QM9 and perovskite spacer molecules, while preserving chemically relevant substructure relationships within each group.
000 ammonium-containing molecules from QM9 and 106 experimentally studied perovskite organic spacers. The two-dimensional embeddings show distinct and compact clusters, indicating that molecules with similar chemical features and functional groups are positioned adjacently in the learned latent space. Notably, the GNN model successfully differentiates between the QM9 and perovskite spacer molecules, while preserving chemically relevant substructure relationships within each group.
To visualize and interpret the learned molecular space, we perform t-SNE on the GNN-learned latent embedding h, rather than the full GP input vector z = [h; u] which includes concatenated global descriptors. This choice is motivated by the fact that h is the sole output of the GNN encoder and directly reflects the message-passing over the molecular graph structure. In contrast, the global descriptor vector u consists of features that are fixed and not learned from data. Including u in the t-SNE projection would confound the learned representation with prior feature design, potentially obscuring the model's internal structural understanding.
Moreover, the sparse GP backend operates on the composite input z, and its kernel structure encourages local smoothness in function space. While this is desirable for calibrated prediction, it can also homogenize nearby points in the latent space, potentially diminishing local diversity that reflects fine-grained chemical distinctions. By projecting only h, we isolate the component of the representation that captures learned topological and electronic patterns from graph connectivity, without the smoothing effect imposed by GP regularization. This makes the t-SNE projections better suited for analyzing how the GNN encoder organizes molecular structures based purely on relational and learned attributes. In this sense, using h aligns with the goal of probing the model's internal inductive biases and understanding how chemical space is structured by the encoder prior to probabilistic inference.
Beyond visualization, the learned latent space also underpins the active learning component of our framework, enabling the principled selection of new, unseen molecules. Because the GNN encoder maps each molecular graph to a continuous embedding that captures structural and functional information, the latent space provides a chemically meaningful coordinate system over which model uncertainty can be assessed. By combining the encoder output with global descriptors to form the full representation z = [h; u], and feeding this into task-specific Gaussian processes, we obtain posterior variance estimates that reflect epistemic uncertainty in prediction. This allows us to identify regions of latent space where the model is uncertain either due to lack of training data, structural novelty, or target-specific complexity. During each acquisition round, we cluster the latent representations of unlabeled molecules and select points with high uncertainty from each cluster, ensuring that acquisitions are both diverse and informative. Crucially, this approach allows us to not only refine predictions for known perovskite-relevant molecules, but also to explore chemically distinct candidates that fall outside the original training distribution. In this way, the latent space supports generalization beyond interpolation, guiding the discovery of new molecules whose structural and electronic features may be relevant to perovskite performance, yet were not explicitly represented in the initial dataset. This dual role as both an interpretable molecular map and a sampling surface for targeted acquisition demonstrates the broader utility of the latent representation learned by the DKL model.
Cluster labels assigned in the fourth t-SNE projection show that the model identifies several discrete molecular families. These clusters reflect both chemical similarity and functional group diversity, confirming that the model captures domain-relevant features required for structure–property learning in the context of 2D and quasi-2D HPs. The visual separation of clusters suggests that the GNN's learned representations are sensitive to functional group patterns that influence intermolecular interactions, electronic structure, and potential spacer behavior, as shown in Fig. 5. The latent space visualization colored by target properties reveals substantial information about the clustering structure and validates the chemical coherence of the molecular groupings shown in Fig. S3–5. When examining electronic properties, clusters 0 and 4 form the hottest regions in the band-gap visualization, both lacking extended π paths and giving the widest gaps (7–9 eV and 6–8 eV respectively), with cluster 0 dominated by long saturated alkyl ammoniums and cluster 4 by bifunctional diammonium/dithiol species where two cationic ends lift every frontier level. The coolest patch sits on the brominated fringe (isolated cyan-green arm), reflecting halogen substitution's ability to compress the gap to 2–3 eV by stabilizing the acceptor level through heavy-atom polarization that pulls LUMO down to ≤−3 eV. HOMO visualization shows aromatic clusters appearing in yellow orange with shallower values (−6 to −5 eV) consistent with electron delocalization along rings, while saturated chains remain blue-cyan as σ-bond frameworks keep ionization energy deep (−8 to −7 eV). LUMO coloring reveals diammoniums and long alkyls giving the warmest shades with positive or near-vacuum energies (0 to +2 eV for cluster 0, 0 to +1.5 eV for cluster 4) reflecting strong cationic character, while bromine-bearing members turn blue green through σ–π mixing effects. The physicochemical property visualization demonstrates a continuous blue-to-red sweep from lower-left to upper-right across all panels, with deep blue points at 0–20 Å2 TPSA carrying one donor/acceptor, while red points exceed 80 Å2 with four donors/acceptors, perfectly aligning cluster 0 (long alkyl ammoniums) in the low-polarity corner and cluster 4 (diammonium/dithiol) in the high-polarity corner. Structural descriptors show ring count and conjugated bonds creating identical spatial patterns that isolate the aromatic island (cluster 1 polycyclics and ring-bearing cluster 2), while molecular weight creates two distinct red patches—one for aromatic systems where extra ring carbons add mass, another for long-alkyl zones with additional methylenes. Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P separates hydrophobic species (cluster 0 linear chains and cluster 1 aromatics) in red regions from polar diammoniums (cluster 4) in deep-blue corners.
P separates hydrophobic species (cluster 0 linear chains and cluster 1 aromatics) in red regions from polar diammoniums (cluster 4) in deep-blue corners.
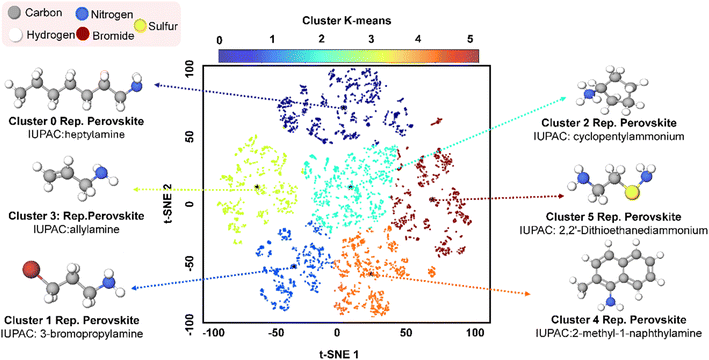 | ||
| Fig. 5 Latent space with t-SNE reduction with clustered k-means and representative perovskite spacer molecules chosen from each cluster with each position marked by a black star. | ||
However, not all property visualizations provide equally interpretable information. Dipole moment follows the same spatial trend as other polar descriptors, though its hottest region is slightly displaced because dipole requires both polar bonds and their vector alignment symmetric placement can mute the overall value even when hetero-atom count is high, making this property less predictable from structural position alone. The reduced accuracy observed for dipole moment (Fig. S2) is expected, because dipole magnitude cannot be inferred reliably from 2D molecular topology alone. It requires 3D conformer geometry, including internal rotation barriers, aromatic plane tilts, and side-chain folding patterns. Our message-passing encoder cannot recover this information from 2D SMILES-derived graphs, which explains the persistent performance ceiling. Incorporating learned 3D geometries or equivariant GNNs (E(n)-GNN, MACE, or SE(3) models) is a natural next step for improving prediction of conformation-dependent descriptors which can be taken from other databases available.60 Molecular weight presents another challenge as it splits along two structural directions rather than tagging any single cluster, creating two separate red patches that don't correspond to a single chemical motif, limiting its utility for systematic design guidance. Additionally, while deep-LUMO traps in the blue zones can be identified, these regions need careful stoichiometry control as they may problematically pull electrons out of the perovskite system. The visualization method itself has inherent limitations since t-SNE preserves local neighborhoods while distorting large-scale distances, meaning that while smooth color transitions across tens of neighbors indicate meaningful chemical relationships, global patterns should be interpreted cautiously. Despite these limitations, the smooth color transitions across most property visualizations, combined with identical geometric neighborhoods sharing similar shades across different target properties, demonstrates that the latent embedding has successfully ranked molecules by their underlying chemical physics, making the clusters chemically coherent and providing reliable visual cues for molecular design in cases where clear spatial–property relationships emerge.
Representative spacers from each cluster (Fig. 5) illustrate the range of structural motifs relevant to 2D and quasi-2D HP design. For each cluster, we isolated the organic-spacer entries identified in our literature survey and computed the Euclidean distance between each spacer's 11-dimensional property vector and the cluster centroid, the property distributions for each cluster can be observed in Fig. S6–11. The spacer with the smallest distance was chosen as that cluster's representative. Although only one spacer per cluster is shown, multiple spacers may occupy the same cluster.61 Across all clusters, molecules contain nitrogen-based functional groups, typically primary amines or cyclic derivatives, which are known to facilitate interaction with the HP lattice.62 Despite overlapping molecular weight ranges, the chemical backbones vary significantly across the six clusters. Cluster 0 features long, linear alkyl ammoniums like heptylamine that provide excellent moisture barriers with limited electronic interaction.61 Cluster 1 encompasses bulky π-systems exemplified by 2-methyl-1-naphthylamine, offering good surface-oriented band offsets and UV filtering capabilities.63 Cluster 2 comprises compact cyclic spacers such as cyclopentylammonium that pack densely and suppress phase transitions.62 Cluster 3 contains very small cations like allylamine with single C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bonds, useful as co-spacers or orientation seeds.64 Cluster 4 includes chelating diammoniums or dithiols such as 2,2′-dithioethanediammonium that bind simultaneously to two surface Pb sites for strong defect passivation.65 Cluster 5 features halogenated propyl chains like 3-bromopropylamine that act as intermediate-polarity spacers with enhanced lattice polarizability.65
C bonds, useful as co-spacers or orientation seeds.64 Cluster 4 includes chelating diammoniums or dithiols such as 2,2′-dithioethanediammonium that bind simultaneously to two surface Pb sites for strong defect passivation.65 Cluster 5 features halogenated propyl chains like 3-bromopropylamine that act as intermediate-polarity spacers with enhanced lattice polarizability.65
These structural variations produce distinct electronic signatures that directly influence optoelectronic and morphological behavior in 2D HPs including both RP and DJ phases.66,67 The calculated band gaps of our representative spacers range from approximately 3.0 to 4.5 eV, reflecting variations in electronic structure that influence optical properties of 2D HPs.68,69 The broad range of dipole moments across clusters (0.2–30 D in the full QM9 envelope, narrowing to 1–5 D in perovskite-relevant subsets) parallels enhanced dielectric screening and suppressed ion migration reported for high-dipole cations.61,70,71 In Cluster 4, the diammonium spacers with their bifunctional anchoring capability reproduce marked improvements in moisture tolerance and reduced Pb leakage.72,73 Aromatic spacers in Cluster 1 foster π–π stacking and lattice rigidity, mirroring thermal and moisture stability enhancements achieved with conjugated cations in both RP and DJ architectures.74–76 Conversely, the flexible aliphatic frameworks in Clusters 0 and 2 facilitate defect passivation, improve film uniformity, and maintain lattice integrity at elevated temperatures.77,78 The halogenated species in Cluster 5 offer tunable polarity while potentially improving film orientation.50,79 Together, these clustering-derived insights validate established structure–property relationships and demonstrate that our approach can effectively identify and prioritize cation chemistries to tune optical, electronic, and morphological properties across layered HPs.
While fundamentally data-driven, our pipeline also functions as a hypothesis-generation engine: by mapping each molecule into an interpretable latent manifold and isolating clusters with distinct electronic, steric, and dipolar signatures, we can formulate precise, experimentally testable hypotheses. For example, high-dipole spacers have been shown to enhance interfacial charge separation, improving energy level alignment and carrier extraction efficiency in HP solar cells.80,81 Aromatic backbones have also been reported to increase lattice rigidity and thermal stability under thermal cycling. This hypothesis-driven selection of mechanistically inspired spacer molecules promises to accelerate design cycles and guide synthetic efforts toward high-performance DJ and RP HPs.82 These structural variations have direct implications for the functionality of the corresponding HPs. To assess whether LLMs can support hypothesis generation in molecular design for 2D HPs, we compared the qualitative suggestions from ChatGPT with the structural groupings identified by our GNN-based clustering of organic spacer cations. Unlike our separate literature-mining model which extracts grounded, source-linked information from primary articles, we engaged ChatGPT specifically for its capacity to generate hypotheses, not verified facts. In response to the question “What type of organic molecule should I use to make stable 2D HPs with high photovoltaic efficiency?” and the follow-up clarification “Ruddlesden–Popper iodide-based lead perovskites,” ChatGPT outlined general design principles: (1) use of large-n phases to reduce exciton binding; (2) vertical layer orientation for better charge transport; and (3) near-linear Pb–I–Pb angles for stronger coupling. It then connected these structural goals to organic spacer properties suggesting rigid, polar, moderately bulky, and hydrogen-bonding-capable molecules and listed several example cations, including phenylethylammonium, butylammonium, and cyclopentylammonium.
Although ChatGPT does not evaluate chemical feasibility or rank candidates, it demonstrates potential as a hypothesis-forming tool: offering plausible mechanistic rationales and proposing candidate classes that can be independently examined.83,84 We then compared its qualitative suggestions to the chemical families identified by our unsupervised GNN clustering, which was trained on a merged dataset of >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 QM9 molecules and literature-reported spacers, using structural and electronic descriptors. Without using phase or performance labels, our model learned six distinct clusters with clear chemical features. Notably, Cluster 1 was enriched in bulky π-systems like 2-methyl-1-naphthylamine, consistent with ChatGPT's emphasis on aromatic structures for surface-oriented band offsets and UV filtering.50 Cluster 4 included chelating diammoniums such as 2,2′-dithioethanediammonium with high dipole moments and bifunctional binding capability, matching ChatGPT's suggestion that polar spacers can aid defect passivation and charge separation. Cluster 2 contained compact cyclic spacers like cyclopentylammonium that suppress phase transitions, aligning with ChatGPT's recommendation for moderately bulky molecules. Meanwhile, Cluster 0 featured long linear alkyl chains like heptylamine that provide excellent moisture barriers, while Cluster 5 included halogenated species like 3-bromopropylamine offering tunable polarity and enhanced lattice polarizability.
000 QM9 molecules and literature-reported spacers, using structural and electronic descriptors. Without using phase or performance labels, our model learned six distinct clusters with clear chemical features. Notably, Cluster 1 was enriched in bulky π-systems like 2-methyl-1-naphthylamine, consistent with ChatGPT's emphasis on aromatic structures for surface-oriented band offsets and UV filtering.50 Cluster 4 included chelating diammoniums such as 2,2′-dithioethanediammonium with high dipole moments and bifunctional binding capability, matching ChatGPT's suggestion that polar spacers can aid defect passivation and charge separation. Cluster 2 contained compact cyclic spacers like cyclopentylammonium that suppress phase transitions, aligning with ChatGPT's recommendation for moderately bulky molecules. Meanwhile, Cluster 0 featured long linear alkyl chains like heptylamine that provide excellent moisture barriers, while Cluster 5 included halogenated species like 3-bromopropylamine offering tunable polarity and enhanced lattice polarizability.
This cross-comparison between ChatGPT-generated hypotheses and our unsupervised GNN clustering suggests that transformer-based language models can contribute meaningfully to hypothesis formation when prompted with well-posed scientific queries. Although ChatGPT does not perform molecular selection based on data or embeddings, it proposed specific spacer molecules alongside general phase-stabilizing strategies and chemical parameter ranges, including spacer rigidity, polarity, hydrogen-bonding capacity, and steric volume. These qualitative suggestions showed partial alignment with structural and electronic trends captured in our GNN-derived clusters, including correlations with dipole moment, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, TPSA, and conjugation extent across the latent space.85,86 The correspondence between ChatGPT's mechanistic intuitions and our data-driven cluster properties such as the hydrophobic alkyl chains in Cluster 0, aromatic π-systems in Cluster 1, and high-dipole bifunctional molecules in Cluster 4 validates both the chemical coherence of our unsupervised clustering and the potential utility of LLMs as hypothesis generators. Rather than serving as a factual source, ChatGPT functioned here as a hypothesis generator offering a scaffold of chemically intuitive heuristics that can be validated or refined through data-driven clustering. As such, integrating LLMs into materials informatics workflows can enhance the interpretability of unsupervised model outputs and help prioritize rational candidates for experimental follow-up, especially in data-sparse design spaces. This demonstrates how combining learned molecular representations with LLM-driven knowledge synthesis can enable a more guided and mechanistically informed exploration of chemical design space.
P, TPSA, and conjugation extent across the latent space.85,86 The correspondence between ChatGPT's mechanistic intuitions and our data-driven cluster properties such as the hydrophobic alkyl chains in Cluster 0, aromatic π-systems in Cluster 1, and high-dipole bifunctional molecules in Cluster 4 validates both the chemical coherence of our unsupervised clustering and the potential utility of LLMs as hypothesis generators. Rather than serving as a factual source, ChatGPT functioned here as a hypothesis generator offering a scaffold of chemically intuitive heuristics that can be validated or refined through data-driven clustering. As such, integrating LLMs into materials informatics workflows can enhance the interpretability of unsupervised model outputs and help prioritize rational candidates for experimental follow-up, especially in data-sparse design spaces. This demonstrates how combining learned molecular representations with LLM-driven knowledge synthesis can enable a more guided and mechanistically informed exploration of chemical design space.
Conclusion
This work establishes an integrated framework combining literature-mined experimental data, high-throughput molecular graph construction, and multitask GNN–DKL modeling to analyze and predict structure–property relationships in 2D HPs. We augmented a curated set of 106 known perovskite organic spacers with over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 chemically diverse, ammonium-containing compounds from the QM9 database, enabling multi-task training across eleven molecular properties selected to probe key aspects of spacer–inorganic interactions—including orbital alignment (band gap, HOMO, LUMO), dielectric response (dipole moment), steric hindrance (molecular weight, log
000 chemically diverse, ammonium-containing compounds from the QM9 database, enabling multi-task training across eleven molecular properties selected to probe key aspects of spacer–inorganic interactions—including orbital alignment (band gap, HOMO, LUMO), dielectric response (dipole moment), steric hindrance (molecular weight, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), hydrogen bonding potential, and detailed ring statistics (aromatic/aliphatic ring counts, size distribution). Each molecule is encoded as a graph with chemically rich atom-, bond-, and global-level features, including formal charge, valence, hybridization state, Gasteiger partial charges, electronegativity, ring membership, and nitrogen proximity, allowing the model to capture molecular factors known to affect octahedral tilt, layer orientation, and phase stability in HPs.
P), hydrogen bonding potential, and detailed ring statistics (aromatic/aliphatic ring counts, size distribution). Each molecule is encoded as a graph with chemically rich atom-, bond-, and global-level features, including formal charge, valence, hybridization state, Gasteiger partial charges, electronegativity, ring membership, and nitrogen proximity, allowing the model to capture molecular factors known to affect octahedral tilt, layer orientation, and phase stability in HPs.
Clustering in the learned latent space reveals chemically meaningful groupings, where each cluster aggregates spacer candidates with shared structural or electronic motifs such as rigid conjugated aromatics, highly polar cyano-substituted molecules, or flexible alkyl chains. These clusters provide a basis for hypothesis formation: by identifying representative molecules from each group, we generate a manageable set of candidates that span the design space and offer targeted functionalities. In parallel, we employed ChatGPT as a generative tool for literature-informed hypothesis framing. Although not grounded in curated data, ChatGPT rapidly articulated structural heuristics (e.g., favoring vertical orientation via rigidity and dipole alignment) and suggested plausible molecular scaffolds. Comparing these LLM-derived suggestions with our GNN clusters revealed partial alignment validating the utility of LLMs for conceptual guidance, while reserving molecular ranking and diversity assessment for model-based evaluation. Together, the ML–LLM integration supports a hybrid hypothesis-generation strategy: ChatGPT proposes mechanism-linked design principles, while GNN clustering filters and expands on them quantitatively across large chemical spaces.
Our future work will deploy this pipeline in an automated high-throughput synthesis and in situ characterization workflow to fabricate and evaluate cluster-representative spacer candidates. By closing the loop between data-driven molecular embeddings, language model hypotheses, and experimental feedback, this framework offers a scalable route for rational discovery and optimization of organic spacers in 2D HPs and other materials systems.
Conflicts of interest
There are no conflicts to declare.Data availability
The code/models/data is available in Zenodo repository: 10.5281/zenodo.18879311.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00378d.
Acknowledgements
J. M, S. S, E. F and M. A. acknowledge support from the National Science Foundation (NSF), Award Number 2043205. S. S. and E. F. acknowledge partial support from the Center for Materials Processing (CMP) at the University of Tennessee, Knoxville. R. D. and A. M. K. acknowledge support from the School of Materials Engineering at Purdue University, and from the NSF FAIROS program award 2226418. U. P. and S. V. K. acknowledge the support by the National Science Foundation Materials Research Science and Engineering Center program through the UT Knoxville Center for Advanced Materials and Manufacturing (DMR-2309083).References
- E.-B. Kim, M. S. Akhtar, H.-S. Shin, S. Ameen and M. K. Nazeeruddin, A review on two-dimensional (2D) and 2D-3D multidimensional perovskite solar cells: Perovskites structures, stability, and photovoltaic performances, J. Photochem. Photobiol., C, 2021, 48, 100405, DOI:10.1016/j.jphotochemrev.2021.100405.
- G. Grancini and M. K. Nazeeruddin, Dimensional tailoring of hybrid perovskites for photovoltaics, Nat. Rev. Mater., 2019, 4(1), 4–22 CrossRef CAS.
- C. T. Triggs, R. D. Ross, W. Mihalyi-Koch, C. F. M. Clewett, K. M. Sanders, I. A. Guzei and S. Jin, Spacer Cation Design Motifs for Enhanced Air Stability in Lead-Free 2D Tin Halide Perovskites, ACS Energy Lett., 2024, 9(4), 1835–1843, DOI:10.1021/acsenergylett.4c00615.
- Y. Li, J. V. Milić, A. Ummadisingu, J.-Y. Seo, J.-H. Im, H.-S. Kim, Y. Liu, M. I. Dar, S. M. Zakeeruddin and P. Wang, et al., Bifunctional Organic Spacers for Formamidinium-Based Hybrid Dion–Jacobson Two-Dimensional Perovskite Solar Cells, Nano Lett., 2019, 19(1), 150–157, DOI:10.1021/acs.nanolett.8b03552.
- Y. Gao, X. Dong and Y. Liu, Recent Progress of Layered Perovskite Solar Cells Incorporating Aromatic Spacers, Nano-Micro Lett., 2023, 15(1), 169, DOI:10.1007/s40820-023-01141-2.
- E. Fransson, J. Wiktor and P. Erhart, Impact of Organic Spacers and Dimensionality on Templating of Halide Perovskites, ACS Energy Lett., 2024, 9(8), 3947–3954, DOI:10.1021/acsenergylett.4c01283.
- T. L. Leung, I. Ahmad, A. A. Syed, A. M. C. Ng, J. Popović and A. B. Djurišić, Stability of 2D and quasi-2D perovskite materials and devices, Commun. Mater., 2022, 3(1), 63, DOI:10.1038/s43246-022-00285-9.
- Z.-J. Bai, J. Xiong, Y. Mao, S. Tian, B.-H. Wang, B. Hu, X. Wang, W. Zhou, C.-T. Au and L. Chen, et al., Lead-free Dion-Jacobson layered double perovskite as a photocatalyst for toluene oxidation, Cell Rep. Phys. Sci., 2023, 4(10), 101591, DOI:10.1016/j.xcrp.2023.101591.
- K. Higgins, S. M. Valleti, M. Ziatdinov, S. V. Kalinin and M. Ahmadi, Chemical Robotics Enabled Exploration of Stability in Multicomponent Lead Halide Perovskites via Machine Learning, ACS Energy Lett., 2020, 5(11), 3426–3436, DOI:10.1021/acsenergylett.0c01749.
- D. Giovanni, S. Ramesh, M. Righetto, J. W. Melvin Lim, Q. Zhang, Y. Wang, S. Ye, Q. Xu, N. Mathews and T. C. Sum, The Physics of Interlayer Exciton Delocalization in Ruddlesden–Popper Lead Halide Perovskites, Nano Lett., 2021, 21(1), 405–413, DOI:10.1021/acs.nanolett.0c03800.
- C. Qin, F. Zhang, L. Qin, X. Liu, H. Ji, L. Li, Y. Hu, Z. Lou, Y. Hou and F. Teng, Charge Transport in 2D Layered Mixed Sn–Pb Perovskite Thin Films for Field-Effect Transistors, Adv. Electron. Mater., 2021, 7(10), 2100384, DOI:10.1002/aelm.202100384.
- S. Wang, M. Mandal, H. Zhang, D. W. Breiby, O. Yildiz, Z. Ling, G. Floudas, M. Bonn, D. Andrienko and H. I. Wang, et al., Odd–Even Alkyl Chain Effects on the Structure and Charge Carrier Transport of Two-Dimensional Sn-Based Perovskite Semiconductors, J. Am. Chem. Soc., 2024, 146(28), 19128–19136, DOI:10.1021/jacs.4c03936.
- M. Choghaei, M. Schiffer, V. Tyagi, M. Righetto, J. Du, M. Buchmüller, K. O. Brinkmann, G. Brocks, P. Görrn and L. M. Herz, et al., Odd-even effects in lead-iodide-based Ruddlesden–Popper 2D perovskites, J. Mater. Chem. A, 2025, 13 10.1039/d5ta01234a.
- M. Moroni, C. Coccia and L. Malavasi, Chiral 2D and quasi-2D hybrid organic inorganic perovskites: from fundamentals to applications, Chem. Commun., 2024, 60(70), 9310–9327, 10.1039/d4cc03314k.
- H. Lee, T. Moon, Y. Lee and J. Kim, Structural Mechanisms of Quasi-2D Perovskites for Next-Generation Photovoltaics, Nano-Micro Lett., 2025, 17(1), 139, DOI:10.1007/s40820-024-01609-9.
- W. Zhang, Z. Liu, L. Zhang, H. Wang, C. Jiang, X. Wu, C. Li, S. Yue, R. Yang and H. Zhang, et al., Ultrastable and efficient slight-interlayer-displacement 2D Dion-Jacobson perovskite solar cells, Nat. Commun., 2024, 15(1), 5709, DOI:10.1038/s41467-024-50018-4.
- X. Shen, K. Kang, Z. Yu, W. H. Jeong, H. Choi, S. H. Park, S. D. Stranks, H. J. Snaith, R. H. Friend and B. R. Lee, Passivation strategies for mitigating defect challenges in halide perovskite light-emitting diodes, Joule, 2023, 7(2), 272–308, DOI:10.1016/j.joule.2023.01.008.
- L. Zhang, C. Sun, T. He, Y. Jiang, J. Wei, Y. Huang and M. Yuan, High-performance quasi-2D perovskite light-emitting diodes: from materials to devices, Light: Sci. Appl., 2021, 10(1), 61, DOI:10.1038/s41377-021-00501-0.
- J. Yang, T. Zhang, C.-Y. Tsai, Y. Lu and L. Yao, Evolution and emerging trends of named entity recognition: Bibliometric analysis from 2000 to 2023, Heliyon, 2024, 10(9), e30053, DOI:10.1016/j.heliyon.2024.e30053.
- X. Fu, C. A. Schuh and E. A. Olivetti, Materials selection considerations for high entropy alloys, Scr. Mater., 2017, 138, 145–150, DOI:10.1016/j.scriptamat.2017.03.014.
- E. Kim, K. Huang, A. Tomala, S. Matthews, E. Strubell, A. Saunders, A. McCallum and E. Olivetti, Machine-learned and codified synthesis parameters of oxide materials, Sci. Data, 2017, 4, 170127, DOI:10.1038/sdata.2017.127.
- E. Kim, K. Huang, A. Saunders, A. McCallum, G. Ceder and E. Olivetti, Materials Synthesis Insights from Scientific Literature via Text Extraction and Machine Learning, Chem. Mater., 2017, 29(21), 9436–9444, DOI:10.1021/acs.chemmater.7b03500.
- R. K. Vasudevan, M. Ziatdinov, C. Chen and S. V. Kalinin, Analysis of citation networks as a new tool for scientific research, MRS Bull., 2016, 41(12), 1009–1015, DOI:10.1557/mrs.2016.270.
- S. R. Young, A. Maksov, M. Ziatdinov, Y. Cao, M. Burch, J. Balachandran, L. Li, S. Somnath, R. M. Patton and S. V. Kalinin, et al., Data mining for better material synthesis: The case of pulsed laser deposition of complex oxides, J. Appl. Phys., 2018, 123(11) DOI:10.1063/1.5009942.
- H. Jelodar, Y. Wang, C. Yuan, X. Feng, X. Jiang, Y. Li and L. Zhao, Latent Dirichlet allocation (LDA) and topic modeling: models, applications, a survey, Multimed. Tool. Appl., 2019, 78(11), 15169–15211, DOI:10.1007/s11042-018-6894-4.
- I. Beltagy; K. Lo; A. CohanSciBERT: A pretrained language model for scientific text. arXiv, 2019, preprint, arXiv:1903.10676, DOI:10.48550/arXiv.1903.10676.
- J. Lee, W. Yoon, S. Kim, D. Kim, S. Kim, C. H. So and J. Kang, BioBERT: a pre-trained biomedical language representation model for biomedical text mining, Bioinformatics, 2020, 36(4), 1234–1240 Search PubMed.
- J. Zhang, L. Zhang, Y. Sun, W. Li and R. Quhe, Named entity recognition in the perovskite field based on convolutional neural networks and MatBERT, Comput. Mater. Sci., 2024, 240, 113014 Search PubMed.
- S. Huang and J. M. Cole, BatteryBERT: A pretrained language model for battery database enhancement, J. Chem. Inf. Model., 2022, 62(24), 6365–6377 Search PubMed.
- M. P. Polak, S. Modi, A. Latosinska, J. Zhang, C.-W. Wang, S. Wang, A. D. Hazra and D. Morgan, Flexible, model-agnostic method for materials data extraction from text using general purpose language models, Digital Discovery, 2024, 3(6), 1221–1235, 10.1039/d4dd00016a.
- M. P. Polak and D. Morgan, Extracting accurate materials data from research papers with conversational language models and prompt engineering, Nat. Commun., 2024, 15(1), 1569, DOI:10.1038/s41467-024-45914-8.
- M. C. Swain and J. M. Cole, ChemDataExtractor: a toolkit for automated extraction of chemical information from the scientific literature, J. Chem. Inf. Model., 2016, 56(10), 1894–1904 Search PubMed.
- A. Matarazzo; R. TorloneA Survey on Large Language Models with some Insights on their Capabilities and Limitations. arXiv, 2025, preprint arXiv:2501.04040, DOI:10.48550/arXiv.2501.04040.
- D. H. Hagos, R. Battle and D. B. Rawat, Recent advances in generative ai and large language models: Current status, challenges, and perspectives, IEEE Trans. Artif. Intell., 2024, 5(12), 5873–5893 Search PubMed.
- O. Fagbohun; S. Yashwanth; A. S. Akintola; I. Wurola; L. Shittu; A. Inyang; O. Odubola; U. Offia; S. Olanrewaju; O. ToluwalekeGreenIQ: A Deep Search Platform for Comprehensive Carbon Market Analysis and Automated Report Generation, arXiv, 2025, preprint, arXiv:2503.16041, DOI:10.48550/arXiv.2503.16041.
- X. Liu; P. Sun; S. Chen; L. Zhang; P. Dong; H. You; Y. Zhang; C. Yan; X. Chu; T.-y. ZhangPerovskite-llm: Knowledge-enhanced large language models for perovskite solar cell research. arXiv, 2025, preprint, arXiv:2502.12669, DOI:10.48550/arXiv.2502.12669.
- S. Schulhoff; M. Ilie; N. Balepur; K. Kahadze; A. Liu; C. Si; Y. Li; A. Gupta; H. Han; S. SchulhoffThe prompt report: A systematic survey of prompting techniques, arXiv, 2024, preprint, arXiv:2406.06608, p. 5, DOI:10.48550/arXiv.2406.06608.
- B. Chen, Z. Zhang, N. Langrené and S. Zhu, Unleashing the potential of prompt engineering for large language models, Patterns, 2025, 101260, DOI:10.1016/j.patter.2025.101260.
- Y. Gao; Y. Xiong; X. Gao; K. Jia; J. Pan; Y. Bi; Y. Dai; J. Sun; H. Wang; H. WangRetrieval-augmented generation for large language models: A survey, arXiv, 2023, preprint, arXiv:2312.10997, 2, 1, DOI:10.48550/arXiv.2312.10997.
- A. P. Bento, A. Hersey, E. Félix, G. Landrum, A. Gaulton, F. Atkinson, L. J. Bellis, M. De Veij and A. R. Leach, An open source chemical structure curation pipeline using RDKit, J. Cheminf., 2020, 12, 1–16 Search PubMed.
- A. G. Wilson, Z. Hu, R. Salakhutdinov, and E. P. XingDeep kernel learning, in Artificial intelligence and statistics, PMLR, 2016, pp. 370–378 Search PubMed.
- Y. Xu, X. Liu, W. Xia, J. Ge, C.-W. Ju, H. Zhang and J. Z. H. Zhang, ChemXTree: A Feature-Enhanced Graph Neural Network-Neural Decision Tree Framework for ADMET Prediction, J. Chem. Inf. Model., 2024, 64(22), 8440–8452, DOI:10.1021/acs.jcim.4c01186.
- M. Ziatdinov Active learning with fully bayesian neural networks for discontinuous and nonstationary data. arXiv, 2024, preprint, arXiv:2405.09817, DOI:10.48550/arXiv.2405.09817.
- E. Foadian, S. Sanchez, S. V. Kalinin and M. Ahmadi, From Sunlight to Solutions: Closing the Loop on Halide Perovskites, ACS Mater. Au, 2025, 5(1), 11–23, DOI:10.1021/acsmaterialsau.4c00096.
- M. Um, S. L. Sanchez, H. Song, B. J. Lawrie, H. Ahn, S. V. Kalinin, Y. Liu, H. Choi, J. Yang and M. Ahmadi, Tailoring Molecular Space to Navigate Phase Complexity in Cs-Based Quasi-2D Perovskites via Gated-Gaussian-Driven High-Throughput Discovery, Adv. Energy Mater., 2024, 15(16), 2404655, DOI:10.1002/aenm.202404655.
- S. L. Sanchez, Y. Tang, B. Hu, J. Yang and M. Ahmadi, Understanding the ligand-assisted reprecipitation of CsPbBr3 nanocrystals via high-throughput robotic synthesis approach, Matter, 2023, 6(9) DOI:10.1016/j.matt.2023.05.023.
- E. Foadian, J. Yang, S. B. Harris, Y. Tang, C. M. Rouleau, S. Joy, K. R. Graham, B. J. Lawrie, B. Hu and M. Ahmadi, Decoding the Broadband Emission of 2D Pb-Sn Halide Perovskites through High-Throughput Exploration, Adv. Funct. Mater., 2024, 34(52), 2411164, DOI:10.1002/adfm.202411164.
- O. AI, Pricing of OpenAI platform, 2025. https://platform.openai.com/docs/pricing accessed.
- Z. Fang, M.-H. Shang, Y. Zheng, Q. Sun, X. Hou and W. Yang, Built-in Electric Field in Quasi-2D CsPbI3 Perovskites Using High-Polarized Zwitterionic Spacer for Enhanced Charge Separation/Transport, J. Phys. Chem. Lett., 2023, 14(32), 7331–7339, DOI:10.1021/acs.jpclett.3c01894.
- P. Liu, X. Li, T. Cai, W. Xing, N. Yang, H. Arandiyan, Z. Shao, S. Wang and S. Liu, Molecular Structure Tailoring of Organic Spacers for High-Performance Ruddlesden–Popper Perovskite Solar Cells, Nano-Micro Lett., 2024, 17(1), 35, DOI:10.1007/s40820-024-01500-7.
- H. Lai, Z. Xu, Z. Shao, B. Cui, B. Tian, H. Wang and Q. Fu, Rational Regulation of Organic Spacer Cations for Quasi-2D Perovskite Solar Cells, Sol. RRL, 2023, 7(10), 2300132, DOI:10.1002/solr.202300132.
- Z. Guan, D. Shen, M. Li, C. Ma, W.-C. Chen, X. Cui, B. Liu, M.-F. Lo, S.-W. Tsang and C.-S. Lee, et al., Effects of Hydrogen Bonds between Polymeric Hole-Transporting Material and Organic Cation Spacer on Morphology of Quasi-Two-Dimensional Perovskite Grains and Their Performance in Light-Emitting Diodes, ACS Appl. Mater. Interfaces, 2020, 12(8), 9440–9447, DOI:10.1021/acsami.9b20750.
- K. Shen, J. Wang, Y. Lu, Y. Shen, Y.-Q. Li, Z. Su, L. Chen, F. Song, X. Gao and J.-X. Tang, Exploration of the Defect Passivation in Perovskite Materials Using Organic Spacer Cations, Adv. Mater. Interfaces, 2022, 9(10), 2102253, DOI:10.1002/admi.202102253.
- H. Di, W. Zeng, B.-H. Li, F. Liao, C. Zhao, C. Liang, H. Li, J.-C. Wang, D.-B. Cheng and Z. Ren, et al., Regulating 3D Phase in Quasi-2D Perovskite Films for High-Performance and Stable Photodetectors, Adv. Sci., 2023, 10(26), 2302917, DOI:10.1002/advs.202302917.
- D. Zhang, Y. Fu, W. Wu, B. Li, H. Zhu, H. Zhan, Y. Cheng, C. Qin and L. Wang, Comprehensive Passivation for High-Performance Quasi-2D Perovskite LEDs, Small, 2023, 19(11), 2206927, DOI:10.1002/smll.202206927.
- W. Yang, X. He, X. Huang, X. Wang, Y. Zhang and C.-H. Gao, Defect Passivation in Quasi-2D Perovskite Light-Emitting Diodes by an Ibuprofen Additive, ACS Appl. Mater. Interfaces, 2024, 16(1), 1628–1637, DOI:10.1021/acsami.3c10337.
- J.-H. Kim, C.-M. Oh, I.-W. Hwang, J. Kim, C. Lee, S. Kwon, T. Ki, S. Lee, H. Kang and H. Kim, et al., Efficient and Stable Quasi-2D Ruddlesden–Popper Perovskite Solar Cells by Tailoring Crystal Orientation and Passivating Surface Defects (Adv. Mater. 31/2023), Adv. Mater., 2023, 35(31), 2370221, DOI:10.1002/adma.202370221.
- A. Ghosh, M. Ziatdinov, S. V. Kalinin, Active Deep Kernel Learning of Molecular Functionalities: Realizing Dynamic Structural Embeddings, arXiv, 2024, preprint, arXiv:2403.01234, DOI:10.48550/arXiv.2403.01234.
- H. Zhou, K. Wang, C. Nie, J. Deng, Z. Chen, K. Zhang, X. Zhao, J. Liang, D. Huang and L. Zhao, et al., Quantitative Analysis of Perovskite Morphologies Employing Deep Learning Framework Enables Accurate Solar Cell Performance Prediction, Small, 2025, 21(18), 2408528, DOI:10.1002/smll.202408528.
- T. Xie, Y. Wan, Y. Zhou, W. Huang, Y. Liu, Q. Linghu, S. Wang, C. Kit, C. Grazian and W. Zhang, et al., Creation of a structured solar cell material dataset and performance prediction using large language models, Patterns, 2024, 5(5), 100955, DOI:10.1016/j.patter.2024.100955.
- X. Liu, H. Yan, Z. Shu, X. Cui and Y. Cai, Theoretical insights into spacer molecule design to tune stability, dielectric, and exciton properties in 2D perovskites, Nanoscale, 2025, 17(5), 2658–2667, 10.1039/d4nr04406a.
- S. Khan, R. Shrestha, M. Jin, D. Kim, G.-L. Chen, R. Li, Y. Gu, Q. Tu, N. Ahn and W. Nie, Designing Robust Quasi-2D Perovskites Thin Films for Stable Light-Emitting Applications, Adv. Mater., 2025, 37(25), 2413412, DOI:10.1002/adma.202413412.
- E. Mahal and B. Pathak, Pressure-Induced Modulation of Band Characteristics in 2D Hybrid Perovskites, ACS Appl. Energy Mater., 2025, 8(2), 1134–1142, DOI:10.1021/acsaem.4c02687.
- S. Choudhary, J. Ghosh, S. Pathak, S. K. Saini, N. K. Tailor, P. Sellin, S. Bhattacharya and S. Satapathi, Organic Spacers Modulated Low Dose X-Ray Detection in Hybrid Halide 2D Perovskites: Unveiling Exciton Dynamics, Small, 2025, 21(7), 2409962, DOI:10.1002/smll.202409962.
- Z. Liu, Z. Zhang, Y. Liu, R. Luo, Y. Cheng, Y. Shen, K. Wang and M. Wang, Boosting Carrier Mobility in 2D Layered Perovskites for High-Performance UV Photodetector, Small Methods, 2025, 9(3), 2400887, DOI:10.1002/smtd.202400887.
- B. Kim, J. Park, D. Kang, N. E. Jung, K. Kim, H. Ryu, J. I. Jang, S. Park and Y. Yi, Tuning electronic structure and carrier transport properties through crystal orientation control in two-dimensional Dion-Jacobson phase perovskites, Nano Converg., 2025, 12(1), 1, DOI:10.1186/s40580-024-00473-y.
- Y. Zhang, J. Xi, Y. Deng, W. Liu, Z. Li, C. Liu and W. Guo, The Crucial Role of Organic Ligands on 2D/3D Perovskite Solar Cells: A Comprehensive Review, Adv. Energy Mater., 2024, 14(48), 2403326, DOI:10.1002/aenm.202403326.
- M. Dyksik, M. Baranowski, J. J. P. Thompson, Z. Yang, M. R. Medina, M. A. Loi, E. Malic and P. Plochocka, Steric Engineering of Exciton Fine Structure in 2D Perovskites, Adv. Energy Mater., 2025, 15(9), 2404769, DOI:10.1002/aenm.202404769.
- J. Jiang, T. van der Heide, S. Thébaud, C. R. Lien-Medrano, A. Fihey, L. Pedesseau, C. Quarti, M. Zacharias, G. Volonakis and M. Kepenekian, Flexible and efficient semiempirical DFTB parameters for electronic structure prediction of 3D, 2D iodide perovskites and heterostructures, Phys. Rev. Mater., 2025, 9(2), 023803 Search PubMed.
- Y. Kim, S. Nussbaum, D. Chen, N. Grandjean, R. Scopelliti, H. Guo, S. G. Lee, H.-H. Cho, J.-H. Yum and K. Sivula, Decoupling Interlayer Spacing and Cation Dipole on Exciton Binding Energy in Layered Halide Perovskites, Chem. Mater., 2024, 36(20), 10133–10141, DOI:10.1021/acs.chemmater.4c01527.
- J. Qian, Y. Li, Y. Shen, X. Zhao, C. Wu, H. Gu, Z. Zhang, Y. Chen, B. Cai and J. Xia, et al., Dion–Jacobson-Phase 2D Sn-Based Perovskite Comprising a High Dipole Moment of π-Conjugated Short-Chain Organic Spacers for High-Performance Solar Cell Applications, ACS Nano, 2024, 18(23), 15055–15066, DOI:10.1021/acsnano.4c02076.
- Z. Luo, J. Wu, R. Lin, W. Zhang, Y. Liu, L. Xiao and Y. Min, Solvent-mediated carboxylic acid diammonium spacer for synthesizing FA-based 2D Dion–Jacobson perovskites toward efficient solar cells, Appl. Phys. Lett., 2024, 125(26) DOI:10.1063/5.0248035.
- Y. Zhou, Y. Zhang, L. Zhang, H. Wu, Y. Zhou, X. Xu, J. Yu, X. Wu, J. Xie and W. Fu, et al., Aromatic Imidazole Diammonium-based 2D Dion–Jacobson Perovskites with Reduced Exciton Binding Energy, Adv. Funct. Mater., 2024, 34(48), 2408774, DOI:10.1002/adfm.202408774.
- J. Gu and Y. Fu, Is There an Optimal Spacer Cation for Two-Dimensional Lead Iodide Perovskites?, ACS Mater. Au, 2025, 5(1), 24–34, DOI:10.1021/acsmaterialsau.4c00101.
- J. Zhang, L. Chu, T. Liu, B. Tian, W. Chu, X. Sun, R. Nie, W. Zhang, Z. Zhang and X. Zhao, et al., Engineering Spacer Conjugation for Efficient and Stable 2D/3D Perovskite Solar Cells and Modules, Angew. Chem., Int. Ed., 2025, 64(1), e202413303, DOI:10.1002/anie.202413303.
- Y. Shen, L. Luo, Y. Zhang, Y. Meng, Y. Yan, P. Xie, D. Li, Y. Ji, S. Hu and S. Yip, et al., High-Performance Nanogap Photodetectors Based on 2D Halide Perovskites with a Novel Spacer Cation, Adv. Funct. Mater., 2024, 34(41), 2403746, DOI:10.1002/adfm.202403746.
- X. Gao, Y. Wu, Y. Zhang, X. Chen, Z. Song, T. Zhang, Q. Fang, Q. Ji, M.-G. Ju and J. Wang, How the Spacer Influences the Stability of 2D Perovskites?, Small Methods, 2025, 9(5), 2401172, DOI:10.1002/smtd.202401172.
- J. Choi, J. Kim, M. Jeong, B. Park, S. Kim, J. Park and K. Cho, Molecularly Engineered Alicyclic Organic Spacers for 2D/3D Hybrid Tin-based Perovskite Solar Cells, Small, 2024, 20(48), 2405598, DOI:10.1002/smll.202405598.
- F. B. Minussi, R. M. Silva, J. C. S. Moraes and E. B. Araújo, Organic cations in halide perovskite solid solutions: exploring beyond size effects, Phys. Chem. Chem. Phys., 2024, 26(31), 20770–20784, 10.1039/d4cp02419b.
- W. Zhang, H. Liu, T. Huang, L. Kang, J. Ge, H. Li, X. Zhou, W. Zhang, T. Shi and H.-L. Wang, Oriented Molecular Dipole-Enabled Modulation of NiOx/Perovskite Interface for Pb-Sn Mixed Inorganic Perovskite Solar Cells, Adv. Mater., 2025, 37(8), 2414125, DOI:10.1002/adma.202414125.
- W. Xu, Y. Zhang, Y. Huang, Y. Tao, J. Wang, W. Liu, D. Wang, Z. Zhang, J. Xiong and J. Zhang, Non-Volatile Multifunctional Dipole Molecules Enable 19.2% Efficiency for Printable Mesoscopic Perovskite Solar Cells, Small, 2025, 21(7), 2407063, DOI:10.1002/smll.202407063.
- Z. Tan, W. liu, R. Chen, S. Liu, Q. Zhou, J. Wang, F. Ren, Y. Cai, C. Shi and X. Liu, et al., Enhancing Interfacial Contact for Efficient and Stable Inverted Perovskite Solar Cells and Modules, Adv. Funct. Mater., 2025, 35(19), 2419133, DOI:10.1002/adfm.202419133.
- K.-H. Cohrs, E. Diaz, V. Sitokonstantinou, G. Varando and G. Camps-Valls, Large language models for causal hypothesis generation in science, Mach. Learn.: Sci. Technol, 2025, 6, 013001, DOI:10.1088/2632-2153/ada47f.
- G. Xiong, E. Xie, A. H. Shariatmadari, S. Guo, S. Bekiranov, A. Zhang, Improving scientific hypothesis generation with knowledge grounded large language models, arXiv, 2024, preprint arXiv:2411.02382, DOI:10.48550/arXiv.2411.02382.
- J. M. Cavanagh, K. Sun, A. Gritsevskiy, D. Bagni, T. D. Bannister, T. S. L. Head-Gordon, Modifying Large Language Models for Directed Chemical Space Exploration, arXiv, 2024, preprint, arXiv:2409.02231, DOI:10.48550/arXiv.2409.02231.
- K. Sakano, K. Furui and M. Ohue, NPGPT: natural product-like compound generation with GPT-based chemical language models, J. Supercomput., 2024, 81(1), 352, DOI:10.1007/s11227-024-06860-w.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |