 Open Access Article
Open Access ArticleAn automated sampling workflow for parallel long-term membrane diffusion cell testing
Claire
Benstead
 ,
Maria
Politi
,
Maria
Politi
 ,
David S.
Bergsman
,
David S.
Bergsman
 * and
Lilo D.
Pozzo
* and
Lilo D.
Pozzo
 *
*
Department of Chemical Engineering, University of Washington, Seattle, WA, USA. E-mail: dbergs@uw.edu; dpozzo@u.washington.edu
First published on 25th February 2026
Abstract
Permeation tests are an important but time-intensive component of membrane characterization that often requires recurring data collection over extended periods of time. Here we present a new workflow for automating long-term membrane permeation testing. Through the use of open-source software and 3D-printed labware, we have automated and parallelized the periodic collection of liquid samples from membrane diffusion cell tests conducted in H-cells. Capable of parallelizing up to 8 H-cell diffusion tests at a time, this workflow allows for round-the-clock data collection without the need for manual sampling. In tandem with UV-vis characterization, we demonstrate the usage of this workflow by measuring the diffusion coefficients of four aqueous dyes of varying composition through poly(ethylene glycol) diacrylate (PEGDA)-based hydrogel membranes. The materials and scripts presented here have been published for easy implementation to other research projects that require repeated sample collection over extended periods of time. Developed with standard labware and tools in mind, this workflow is not only compatible with the Opentrons OT2 robot, but could also be adapted to other commercially available liquid handling platforms.
1 Introduction
Membranes used for chemical separations can be found in a vast array of technologies, ranging from water purification and gas separations, to electrochemistry and bioreactors.1–3 Because of this, the development of new membrane materials and applications has become increasingly of interest to the scientific community. Among methods of characterizing these materials, permeation testing is a critical component of membrane development. However, it is also a time-consuming task, due to the need to set up the permeation experiment, the long time required to measure sufficient flux, and the required regular measurement of permeate concentration. These experiments can also be human resource-intensive and prone to user error.4–6In recent years, self-driving labs (SDLs) and materials acceleration platforms (MAPs) have revolutionized how new materials are discovered, developed, and tested.7,8 While systems with in situ or customized capabilities can accelerate these measurements, these systems are often prohibitively expensive to purchase and build. Instead, democratization of laboratory automation has and continues to allow a more widespread adoption of these practices, lowering the barrier to entry through open-source software and hardware, low-cost automation tools, and customizations of commercially available robotic platforms.9,10 A prime example of this democratization in practice is with the Opentrons OT2 liquid-handling robot, whose open-source nature and widespread availability has made it a popular platform for customization. Accordingly, researchers have developed a variety of modifications and custom protocols to meet their research needs, such as the automation of DNA assemblies, mass spectrometry-enabled high-throughput reaction screening, and Newtonian fluid viscosity testing.11–13 The availability and lowered barrier to entry of these types of workflows have also inspired further developments, such as the TidyTron protocol developed by Bryant Jr et al. for cleaning and reusing single-use plastic waste, the AMPERE-2 protocol inspired by the original AMPERE workflow for electrodeposition and electrochemical testing, and SA-ODG workflow by Detassis, et al. for microRNA detection and quantification in liquid biopsies.14–18
One of the many advantages that SDLs and MAPs can offer is the ability to automate repetitive or time-consuming tasks, which can both increase throughput and improve experimental reproducibility by minimizing human involvement.11,15 When characterizing membrane transport properties, collecting and processing samples throughout the length of a multi-day experiment is one such task that can be improved by SDLs and MAPs. In fact, researchers in the membrane community have been developing new techniques and designs to automate and increase the throughput of membrane permeation testing. Muetzel, et al., and Ouimet, et al., for example, have developed automated data collection workflows for rapid and in situ characterization, which increase the amount of data collected per sample and reduce the demand on the researcher's time.19,20 Kazemi and Latulippe developed a technique known as stirred-well filtration (SWF), which allows for high-throughput, parallelized microscale screening for downstream bioprocessing, and Arias Ponce, et al. have designed 3D-printed millifluidic devices to study the effects of ligand chemistries on hydrogel membrane performance, with the potential for parallelization.21,22
Here, we have developed a workflow to automate the data collection of multi-day membrane diffusion testing. Capable of parallelizing up to 8 H-cell diffusion tests at a time, this workflow has been designed using an Opentrons OT2 platform equipped with custom 3D-printed labware and open source scripts. In this design (Fig. 1), the membrane is assembled at the flange connection of a dual-chamber H-cell. As the test solute diffuses across the membrane from the donor chamber to the receptor chamber, the OT2 collects samples from each chamber at regular intervals and stores each sample in a standard 96-well plate. This design allows for sample collection to occur nonstop throughout the experiment independent of researcher involvement, instead only requiring once- or twice-daily visits to characterize the collected samples with UV-vis spectroscopy – a step which, though not demonstrated here, could itself be automated.
To demonstrate the implementation of this workflow, we have characterized solute diffusion across two poly(ethylene glycol) diacrylate- (PEGDA)-based hydrogel membranes. Hydrogels, due to their high hydrophilicity and tunable structural properties, are being developed for a wide variety of membrane-based applications, including for separations, water purification, anti-fouling coatings, and drug delivery.23–28 They consist of 3D networks of crosslinked hydrophilic polymers, which allows them to retain large amounts of water without sacrificing their structure. Because of the vast array of variables that can be altered during their formulation process – such as polymer type, polymer weight, functionalization chemistry, crosslinking method, chemical additives, and environmental conditions – hydrogels can be tuned and altered extensively to meet performance needs.29 However, with such an expansive design space to work with, optimizing a hydrogel material to meet specific performance criteria can be a lengthy iterative process. For example, while low-MW polymers can be used to create a more restrictive network, the resulting material can also become more brittle.30 Alternatively, high-MW polymers can create flexible membrane coatings, but face issues with high swelling ratios and membrane delamination.31
In cases where permeance is critical, characterizing transport properties within a hydrogel can become a rate-limiting step in the optimization process.32 This is partially because the mobility of solutes within hydrogel networks, such as biological substrates (e.g. sugars), chemical precursors, drugs, proteins, or other polymers, may change due to a number of factors, including structural heterogeneity (i.e., a distribution of mesh sizes and water-filled voids), electrostatics, swelling and deswelling, and adsorption.33 While theories and models regarding solute transport within a hydrogel network have been studied and developed for decades, actual diffusion testing of these hydrogels remains a necessary component to successful materials development.34–38 Through parallelization and automation of the sample collection process, these diffusion and permeability tests can be made far more efficient.
These PEGDA-based membranes, which consist of PEGDA hydrogels crosslinked onto poly(ether sulfone) (PES) support substrates, were fabricated using 25 wt% (PEG25) and 40 wt% (PEG40) pre-polymerization solutions. PEGDA, a popular hydrogel material due to its biocompatibility and ease of cross-linking through photopolymerization, is often used in cell encapsulation and drug delivery, thus making transport dynamics a critical component of characterization.39–41 Using four aqueous dyes (brilliant blue, acid orange 7, rhodamine B, methylene blue) as proxies for small molecules of varying charges, this automation protocol was coupled with UV-vis characterization to determine their effective diffusivities through each membrane.
We present this workflow as a low-cost and easily implementable approach to automating and parallelizing membrane testing. Although we demonstrate the application of this protocol with hydrogel membranes, it could be easily adapted or modified to work with other types of membranes, analytes, and characterization techniques such as fluorescence spectroscopy, conductivity, or liquid chromatography. The scripts and custom hardware could also be implemented with other open-source liquid-handling platforms, such as the Science Jubilee from Machine Agency or the Pipettin' Bot from Open Lab Automata.42
2 Materials and methods
2.1 Chemicals
Poly(ethylene glycol) diacrylate (PEGDA, average Mn 700 g mol−1) and brilliant blue FCF (analytical standard, 793 g mol−1) were purchased from Sigma-Aldrich (St. Louis, Mo). Lithium phenyl (2,4,6-trimethyl benzoyl) phosphinate (LAP, purity ≥98.0% (T)(HPLC)) was purchased from TCI America (Tokyo, Japan). Poly(ether sulfone) membranes (PES, 1.2 µm pore size, 47 mm diameter, 110–150 µm nominal thickness) were purchased from Sterlitech Corporation (Auburn, WA). Rhodamine B (479 g mol−1) and acid orange 7 (orange II, pure, certified, 350.32 g mol−1) were purchased from Fisher Scientific (Waltham, MA). Methylene blue chloride (1% aqueous solution, 319 g mol−1, CAT MM0440) was purchased from ALDON Innovating Science (Avon, NY). All chemicals were used as received without further purification. All water used was deionized.2.2 Labware and robotic platforms
Hydrogel crosslinking was conducted using RealUV LED strip lights (395 nm, Waveform Lighting). Electrochemical H-cells with NW16 flange connections and GL18 threaded top openings (with knuckle clamps and vacuum-seal Teflon-centered Viton O-rings) were purchased from Adams & Chittenden Scientific Glass Coop (Berkeley, CA). Magnetic stir bars (small, almond-shaped) were used in each chamber of each H-cell for mixing. Sample aliquots collected by the automated workflow were prepared and characterized in clear, flat-bottomed 96-well polystyrene plates (Fisherbrand, 12-565-501). Single-use X-Pierce Sealing Films, used to reduce evaporation from the sample plates, were purchased from Excel Scientific (non-sterile, Z722502). An Opentrons OT2 liquid handling robot was equipped with single-channel GEN2 P20 (1–20 µL) and P300 (20–300 µL) pipettes (left and right positions, respectively). The associated 20 µL and 300 µL pipette tips and tip racks were purchased from Opentrons. UV-vis characterization was conducted using a BMG LABTECH VANTAstar plate reader (SN 421-0094).2.3 Hydrogel preparation
All hydrogel solutions were prepared as 3 g samples in single-use 5 mL glass vials (Fisherbrand, cat no: FS60910A-112), which were either wrapped in aluminum foil or kept away from light to avoid premature activation of the photoinitiator. The photoinitiator (LAP) was first added to a small amount of water (∼0.2 g) to ensure it was fully dissolved and would be evenly distributed throughout the gel. The PEGDA was then added, followed by the remaining quantity of water needed to reach the concentrations listed in Table 1. The solutions were mixed by hand until there was no visible separation of aqueous and polymer phases, then allowed to rest for 30 minutes before membrane fabrication.| Membrane | Support substrate | PEGDA wt% | LAP wt% |
|---|---|---|---|
| PES | PES | — | — |
| PEG25 | PES | 25 | 0.125 |
| PEG40 | PES | 40 | 0.125 |
2.4 Membrane fabrication
Commercially available PES membranes (1.2 µm pore size, 110–150 µm nominal thickness) were used as the support substrate for all hydrogel membrane samples. The PES membranes were cut to size to fit the NW16 flange of the H-cells (19 mm diameter, with an exposed area diameter of 16 mm), then allowed to soak in the hydrogel formulation solution for a minimum of 30 minutes. After the soaking step was completed, the membranes were removed from the solution and a Kimwipe was used to wick away any excess solution that could pool unevenly on the membrane during the crosslinking process. This was done by briefly tapping an edge of the substrate to the Kimwipe until drops of excess solution were no longer forming when the membrane was held vertically. The membranes were then crosslinked using 395 nm light for 5 minutes in ambient conditions. For the PES-only control samples, the substrates were pre-wet with water before assembling in the H-cells, to reduce the chances of air becoming trapped at the membrane interface and affecting permeation rates. Membrane thicknesses, which ranged from 110 to 200 µm, were measured with a caliper before and after permeability testing, with the post-experiment thickness being used for diffusivity calculations. Individual membrane sample thicknesses can be found with the data provided in the GitHub and Zenodo repositories.2.5 Test solution preparation
The same preparation method was used for each dye (brilliant blue (BB), acid orange 7 (AO), rhodamine B (RB), and methylene blue (MB)), starting with a 10 mM concentrated stock solution and a calibration curve to determine the working range of concentrations for UV-vis (see: Section 2.7). The working UV-vis ranges were established using the same clear, flat-bottomed 96-well plates and sample volume (200 µL) as the samples prepared during the automation workflow. In these conditions, ∼1 Abs was achieved with 15 µM BB, 80 µM AO, 20 µM RB, and 30 µM MB.With these maximum concentrations for UV-vis characterization in mind, the testing solutions were prepared so that the initial donor chamber concentration (C1,0) was 10× more concentrated (e.g., 150 µM BB, rather than 15 µM). This made it possible to extract small aliquots (20 µL) from each H-cell chamber during each sample collection, rather than the full 200 µL needed for characterization. By working with a higher concentration of dye in the H-cells and diluting the extracted sample to an acceptable concentration for UV-vis (10× dilution) in the sample plate, the impact of successive iterations of sample collection on species concentration in the H-cells was minimized. When the H-cells were being prepared to initiate a diffusion experiment, an additional 1 mL sample of the dye was also prepared at the same testing concentration (150 µM BB, 800 µM AO, 200 µM RB, or 300 µM MB, depending on the experiment) in a 5-mL glass vial. This extra aliquot was subsequently used to create an experiment-specific calibration curve (see: Section 2.7).
2.6 Automation workflow
The hardware layout in the OT2, as seen in Fig. 3: 1: dual H-cell assembly; 2: 12-well scintillation vial plate; 4: dual H-cell assembly; 5: 300 µL tip rack; 6: secondary 96-well sample plate (with X-Pierce sealing film); 7: dual H-cell assembly; 8: primary 20 µL tip rack; 9: primary 96-well sample plate (with X-Pierce sealing film); 10: dual H-cell assembly; 11: secondary 20 µL tip rack. All H-cells were assembled (as seen in Fig. 2d) in the H-cell holders and dual stir plates so that the receptor chambers were located closest to the front of the platform.
 | ||
| Fig. 3 Experimental layout of the OT2. The OT2 footprint can accommodate up to 8 H-cells, with two H-cells within each well-plate footprint. | ||
All custom hardware specs are available in the GitHub repository.43
When preparing an experimental setup, several parameters must be defined in the Jupyter Notebook page to tailor the workflow script accordingly. These parameters include: the volume of the aliquots extracted from each donor and receptor chamber, the volume of water used to dilute the aliquots in the sample well plate (dilution factor), total experimental time, and frequency of sample collection. The parameters described in the following Section (2.6.3) were defined to fit the experimental criteria for testing small aqueous dyes on PEGDA hydrogel membranes, but can easily be modified before each round of experimentation.
For the tests discussed in this paper, all samples were collected as 20 µL aliquots and subsequently diluted to 200 µL in the sample well-plates (a 10× dilution factor), with total experimental times ranging from 44 to 96 hours. Dilution factors may be adjusted by altering the aliquot and dilution volumes (e.g., 10 µL aliquot/190 µL water for a 20× dilution) within the volume limitations of the equipment available (maximum of 20 µL aliquot for the P20 pipette, and maximum sample volume of 300 µL in the well plate). Total experimental time may be increased or decreased to match faster or slower diffusion rates. Additionally, most tests discussed here were conducted with a consistent sampling frequency of every 4 or 6 hours. However, arbitrary frequencies – including variable frequencies – can also be defined. For example, rather than collecting samples every 4 hours for the entirety of the experiment, the researcher may implement 2 or 3 rates, such as sample collection every 2 hours for the first 12 hours of the experiment (when concentrations are changing rapidly), followed by every 6 hours for the remainder of the experimental time. The limitations to sampling frequency and total rounds of sample collection are born from two factors: physical space availability and robot speed. If utilizing more than 4 of the membrane testing spots, a maximum of 12 rounds of sample collection may be executed, due to available space in the sample well-plates. Additionally, the robot cannot collect samples at a frequency faster than the time required to complete a full sampling iteration. For a 4-H-cell experiment (i.e. 4 membranes tested in parallel), one iteration requires approx. 10 minutes.
Upon initiating the automation protocol, the OT2 robot prepared the sample well plate by dispensing 180 µL with the P300 pipette into the assigned wells for the first round of sample collection. Using the P20 pipette, the OT2 then collected a 20 µL aliquot from the donor chamber of the first H-cell, dispensed it to the assigned sample well, and mixed the now-diluted sample by aspirating/dispensing 20 µL 8 times. This process was repeated for the receptor chamber of the first H-cell, and again for each chamber of each subsequent H-cell. A clean 20 µL pipette tip was used for each sample collection and discarded after the mixing step. Once all samples had been collected and prepared, the robot picked up the same P300 tip used for dispensing the water to the sample plate, and used it again to dispense 20 µL water to each side of each H-cell to replenish the volume lost to the aliquot collection. This way, a constant and equal volume was preserved in each side of the H-cells to eliminate effects from potential differences in hydrostatic pressure buildup. The P300 tip was then discarded after the replenish step was complete. This completed one iteration of sampling.
The amount of time required to complete a full iteration was then subtracted from the time left until the next sampling iteration, so that the time between each sample collection was kept consistent. In other words, if the robot was scheduled to sample every 4 hours and each sampling iteration took 10 minutes to complete, then the next iteration would be scheduled to occur 230 minutes after completing the iteration, rather than 240 minutes.
Further discussion on the impacts of successive sampling on solute concentration can be found in the SI.
2.7 Characterization
All solutions and samples that were characterized with UV-vis were prepared as 200 µL samples in 96-well flat-bottom well plates, with a 200 µL DI water sample for background subtraction. As noted in Section 2.5, 1 mL samples of testing solution were prepared on the day of experimentation to establish experiment-specific calibration curves. The relationship between concentration and absorbance is defined by the Beer–Lambert Law:| A = εlc | (1) |
In which absorbance (A) is directly proportional to molar absorptivity (ε), path length (l), and concentration (C).44 Given that sample volume was kept consistent (200 µL), the path length l could be treated as constant; thus, the slope of the (experiment-specific) calibration curve (εl) was used to calculate concentration for each sample.
Dye diffusivities were then determined by assuming a pseudo-steady state (PSS) diffusion approximation based on Fick's Law, as depicted in Fig. 4.45–48
 | (2) |
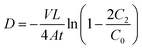 | (3) |
Derived from Colton's theory, the concentration of dye in the receptor chamber (C2) is a function of time t.47 The initial testing dye concentration (C0) (obtained as the donor sample collected at t = 0), membrane thickness (L), exposed membrane area (A), and total volume (V) are treated as constants here, experimentally obtained for each individual membrane permeability test. The diffusion coefficient (D) of each dye-membrane sample was calculated at each discrete C2(t) value and averaged together to obtain the effective diffusion coefficient for one membrane sample. The individual samples for each type of dye-membrane combination were then averaged together to determine the effective diffusivity for that combination. This derivation assumes that the solutions in each chamber are well-mixed (concentrations at the membrane interfaces = bulk solution concentrations), that the diffusion is one-dimensional (no radial diffusion), that species loss due to sampling is negligible, and that the diffusion coefficient (D) is independent of concentration. A full derivation of eqn (3) can be found in the SI.
The diffusion coefficients determined through the use of eqn (3) represent the effective diffusivities of the test solutes through each membrane composition. For the PES substrates, this effective diffusivity, DPES, can be related to the self-diffusivity of the dye in water (Daq) by the following:
| DPES = αDaq | (4) |
In which α is a coefficient accounting for characteristic features of the porous PES substrate, including tortuosity and porosity.47,49,50 For the PEG25 and PEG40 membranes, the diffusivities calculated with eqn (3) are lumped parameters describing effective diffusion across the composite membrane, accounting for the effects of the PES substrate (α), the partitioning coefficient (K), and the true diffusivity across the membrane (Dm).
| Deff = αKDm | (5) |
In this work, Deff is represented by DPES for the PES substrate (sans hydrogel membrane), DP25 for the PEG25 membrane, and DP40 the PEG40 membrane.
3 Results and discussion
Four aqueous dyes (brilliant blue, acid orange 7, rhodamine B, and methylene blue) were tested against 25 wt% PEGDA (PEG25) and 40 wt% PEGDA (PEG40) hydrogel membranes using the ADT automated workflow to assess the transport properties of these membranes. In addition to these PEGDA-based membranes, these four dyes were also tested on the support substrate (PES, pore size 1.2 µm), resulting in a total of 12 different dye-membrane combinations. A minimum of 4 membrane samples were tested for each dye-membrane combination, with more than 80 membrane samples tested across the 12 combinations represented in Fig. 5. The average effective diffusivities for each of the dye-membrane combinations can also be found in Table 2, along with the species charge, dye molecular weight, λmax, and associated free diffusion coefficients in water (Daq) of each dye. | ||
| Fig. 5 Effective diffusivities of four aqueous dyes (BB, AO, RB, MB) through a PES support substrate and two hydrogel membranes: 25 wt% PEGDA (PEG25), and 40 wt% PEGDA (PEG40). | ||
| Dye | Charge | MW | λ max (nm) | D aq | D PES | α | D P25 | D P40 |
|---|---|---|---|---|---|---|---|---|
| a In the absence of a literature value for AO, the Daq of the chemically similar Methyl Orange (MO) at similar concentrations is ∼95.1–98.7 × 10−7 cm2 s−1.56 | ||||||||
| BB | −2 | 793 | 628 | 56.8(ref. 51) | 5.55 ± 1.68 | 0.098 ± 0.030 | 0.06 ± 0.01 | 0.01 ± 0.01 |
| 60 (ref. 52) | ||||||||
| AO | −1 | 350 | 483 | —a | 9.22 ± 3.34 | 0.097 ± 0.035a | 2.41 ± 0.29 | 1.92 ± 0.38 |
| RB | +/− | 479 | 554 | 36 (ref. 53) | 3.71 ± 0.93 | 0.103 ± 0.026 | 4.89 ± 2.05 | 2.51 ± 0.70 |
| 42 ± 3 (ref. 54) | ||||||||
| 42.7 ± 0.4 (ref. 55) | ||||||||
| MB | +1 | 319 | 664 | 19 (ref. 52) | 4.70 ± 0.63 | 0.247 ± 0.035 | 5.28 ± 1.39 | 2.10 ± 0.24 |
3.1 PEGDA hydrogel membrane selectivity
An example of the data collected using the automated workflow can be found in Fig. 6, in which acid orange 7 (AO) was tested on four PEG25 membranes. Over the course of 66 hours, the OT2 collected and prepared 12 samples from each donor and receptor chamber of each H-cell. These samples were then characterized with UV-vis, resulting in the data depicted in the concentration (C) vs. time (t) plot (Fig. 6, left). Using eqn (3), the C2(t) data, and the initial dye solution concentration C0 (C1,0 + C2,0), the diffusivity (D) for each membrane sample was calculated at each collection time t, as depicted in the diffusivity vs. time plot (Fig. 6, right). These discretely calculated values were then averaged to obtain the DP25 for each membrane sample. For the membrane samples represented in Fig. 6 (samples 1, 2, 3, and 4), these DP25 values were determined to be 2.85 ± 0.05, 2.43 ± 0.06, 2.32 ± 0.06, and 2.12 ± 0.04 (×10−7) cm2 s−1, respectively.Given the relatively large pore size of the PES substrate (1.2 µm) in comparison to the organic dyes and the PEGDA hydrogel network (mesh size approx. 3 nm, see SI for mesh size determination), it was expected that the rate of diffusion for each dye would be reduced in comparison to free diffusion in aqueous media, but that the support substrate would not exhibit any selectivity among the four dyes.57,58 This effective diffusivity (DPES) could then be used as a point of comparison for the two hydrogel membrane types (PEG25 and PEG40), to better understand their performance and selectivity, and assess the performance of the automated workflow. As expected, the PES support substrate did not exhibit any noticeable selectivity among the test dyes, with all four DPES values in the same order of magnitude of each other (5.55 ± 1.68 for BB, 9.22 ± 3.34 for AO, 3.71 ± 0.93 for RB, and 4.70 ± 0.63 × 10−7 cm2 s−1 for MB). Compared to their aqueous diffusion coefficients, all four dyes were slowed by an order of magnitude, as seen with the α values (DPES/Daq) provided in Table 2.
For the two hydrogel membranes that were tested using the automated workflow, the effects of the hydrogel network on diffusion was immediately apparent. The PEG25 and PEG40 membranes, which consisted of low-MW PEGDA (700 Mn) embedded and crosslinked into the PES support substrate, had the greatest effect on the diffusion of the divalent anion BB, either screening the dye completely or slowing diffusion to a nearly undetectable rate. The other anionic dye, AO, was able to diffuse across the hydrogel membranes, albeit at a reduced rate compared to the DPES. Alternatively, the zwitterionic RB and cationic MB were relatively unhindered by the presence of a hydrogel network, with DP25 and DP40 values for both dyes on par with their DPES values. These results are in good agreement with diffusion coefficients determined through other techniques, such as fluorescence imaging, microfluidics, and NMR. For a more direct comparison to these techniques, a Dm for each dye-membrane composition has been estimated using eqn (5) (see the SI for the evaluation of partitioning coefficients). These estimates can be found in Table 3. For RB, Dm25 is reasonably similar to the reported values of 3.63 ± 0.44 × 10−7 cm2 s−1 for a 20 vol% hydrogel of 700 Mn PEGDA and 5.891 ± 0.349 × 10−7 cm2 s−1 for 20 wt% 575 Mn PEGDA.59,60 The approximated value for Dm40 of 0.86 × 10−7 cm2 s−1 is somewhat low compared to a reported value of 6.248 ± 1.841 × 10−7 cm2 s−1 for 40 wt% 575 Mn PEGDA.60 For MB, the diffusivities through 30 wt% and 40 wt% 575 Mn PEGDA hydrogels have been reported as 6.77 ± 0.427 and 3.16 ± 0.769 × 10−7 cm2 s−1, respectively, about an order of magnitude faster than the approximated Dm values reported here, though the relatively high α value for MB may contribute to this discrepancy.52
| Dye | K 25 | αK | D m25 | D m40 |
|---|---|---|---|---|
| RB | 33.2 ± 7.9 | 3.42 | 1.43 | 0.73 |
| MB | 22.8 ± 1.8 | 5.63 | 0.94 | 0.37 |
For the anionic dyes, less information is currently available regarding transport properties through PEGDA or similar hydrogels. However, Evans, et al. have reported their work in characterizing the diffusion of acid blue 22 through 575 Mn PEGDA. Acid blue 22 shares several characteristics with BB, including overall charge and similar molecular weights (691.8 vs. 737.7 g mol−1), and thus is likely to exhibit similar behavior within a PEGDA hydrogel. Interestingly, while Evans, et al. were able to report diffusion coefficients of MB with PEGDA using three techniques (optical diffusion, UV-vis, and NMR) with relatively similar results, the authors found that the acid blue 22 was unable to uptake into the PEGDA or, when the hydrogel was prepared with the dye present, unable to release from the polymer network. This resulted in UV-vis-obtained diffusion coefficients on the order of 10−9 cm2 s−1, and NMR-obtained coefficients on the order of 10−7 cm2 s−1, with the authors noting that while the dye was able to move within the gel at a similar rate to the other test solutes, it tended to accumulate at the gel–water interface instead of diffusing out, resulting in the discrepancy between techniques.52 This behavior was similarly observed with BB in this work, in which the PEG25 and PEG40 hydrogel membranes both rejected the divalent anion, resulting in UV-vis-obtained diffusion coefficients on the order of 10−9 cm2 s−1. While BB is larger than the other three dyes tested here and would experience greater counter-ion screening effects in solution, it is unlikely that this rejection is driven by size-based exclusion. Charge-based exclusion likely plays a significant role here, given that the diffusivities of the two anionic dyes (BB and AO) were the most affected by the presence of the PEGDA membranes, while the zwitterionic and cationic dyes (RB and MB) were minimally impacted. This charge preference is further supported by the adsorption of RB and MB into the PEGDA network61 and rejection of the BB during partitioning tests, which can be found in the SI.
3.2 Automation workflow
The workflow presented here was created to automate the most time-intensive component of membrane diffusion testing. As mentioned earlier, the α term reported in Table 2 is a lumped parameter that describes the PES substrate. Given that the same substrate was used for all dye testing, it was expected that all four dyes would produce the same α. Indeed, all four dyes experienced an order of magnitude reduction in diffusivity compared to Daq. In particular, BB, AO, and RB all produced similar α values (0.098 ± 0.030, 0.097 ± 0.035, and 0.103 ± 0.025, respectively), with MB reporting a slightly higher value of 0.247 ± 0.035. This consistency across 4 dyes and 35 samples indicates that the automated workflow provided consistent and reproducible results.The plots in Fig. 7 show a comparison of data collected when conducting these diffusion tests entirely by hand and when the sample collection has been automated. Sample collection by hand was performed every 3–4 hours during a typical work day, whereas the automated sample collection was conducted every six hours for the length of the experiment. Although both experiments presented in the figure produced a similar result (DP25 of 2.12 ± 0.04 × 10−7 cm2 s−1 from the automated sampling and 2.27 ± 0.03 × 10−7 cm2 s−1 from the manual sampling), the automated method facilitated the collection of more data within the same time frame, and could do so at a more consistent, uninterrupted frequency regardless of the time of day. Whereas collecting samples by hand required a researcher to be present in the lab three or more times per day on a strictly timed schedule, the robot could collect and store all samples in a well plate without the need for human oversight. Not only did this reduce the number of times the researcher needed to be physically present down to once or twice a day (without having to adhere to the regimented sampling schedule), but utilizing the sample well plate also made it easy to characterize with UV-vis. As long as the robot was not actively performing a sampling iteration, the sample plate could be removed from the platform, characterized with the plate reader, and returned to the platform without having any impact on the diffusion tests themselves.
 | ||
| Fig. 7 Comparison of data collected using the automated sampling method vs. by hand. Both methods tested the permeation of AO through 25 wt% PEGDA (PEG25) membranes. | ||
The ability to parallelize testing was also greatly beneficial, as it allowed the rate of testing to be increased without increasing time in the lab, which led to an overall increase in efficiency of membrane testing. The robot was able to collect samples overnight and over weekends, so there was a reduced need to intentionally plan when to start experiments in order to optimize data collection. This meant that diffusion tests could be initiated at any time, with no impact on the quality or quantity of the data. As mentioned earlier, more than 80 membrane samples were characterized using this workflow. Assuming each membrane test required a minimum of 44 hours to complete (the shortest experimental time used for the membranes discussed here), a researcher conducting these tests manually would be capable of completing two of these tests within a standard work week. To characterize all 80 membrane samples at that rate would require 40 weeks of nonstop experimentation and more than 600 visits to the lab to collect samples. With the workflow described here, parallelization and automation could drastically reduce the total experimentation time from 10 months to just 5 weeks, with the benefit of nearly double the amount of data collected per sample.
As an added benefit to automating sample collection, this workflow allows for more data and more thorough experimental analysis than might be achievable by manual experimentation. The consistent and repetitive sampling throughout the length of a diffusion experiment makes it possible to monitor and assess experimental design. This could be useful in cases where, for example, results of a test are unexpected, or where the problem may not be observed without sufficient data. An example of this can be found in Fig. 8, which shows the results of a round of experiments testing the diffusivity of BB through PES.
 | ||
| Fig. 8 Data collected (left) and resulting calculated diffusion coefficients (right) of four BB-through-PES diffusion tests conducted in parallel with the automated sampling method. | ||
As seen with all four samples in Fig. 6, the discretely calculated diffusion coefficients for each membrane sample remained relatively consistent throughout the length of the experiment. This consistency would be expected for a system in which diffusion is unidirectional, diffusivity is independent of solute concentration, and conservation of mass applies.45 Alternatively, as seen with sample 8 (Fig. 8), the concentration profile varied slightly from the average behavior of the other three samples tested within the same round of experimentation. Although this could be an indication of a faster permeation rate, the resulting diffusion coefficients calculated at each time t suggest inconsistent diffusion. Thanks to the nonstop data collection made possible with automated workflow, this inconsistency could be tracked throughout the experiment, showing that sample 8 was an outlier from the beginning. When unusual diffusion behavior occurred with one individual sample out of the four tested in parallel, this behavior could generally be linked to an identifiable issue, such as improper assembly of the H-cell allowing for radial diffusion, or the presence of air bubbles trapped at the interface. If the diffusivity had only been calculated using the initial and final concentrations (C0 and C2,t=44), this error might not have been observed at all. As such, the continuous data collection allowed for a more critical analysis of each experiment.
Another type of outlier that was occasionally observed during the course of experimentation was an unusually high or low diffusivity that remained consistent throughout the length of the experiment, such as sample 11 in Fig. 9. In this case, two of three BB-through-PES samples tested in parallel exhibited very similar diffusivities (D = 6.20 ± 0.13 and 6.20 ± 0.59 × 10−7 cm2 s−1), which were consistent with the overall DPES of 5.55 ± 1.68 × 10−7 cm2 s−1 for BB. The third sample, on the other hand, showed an unusually low diffusivity of D = 1.35 ± 0.12 × 10−7 cm2 s−1. Unlike sample 8, which showed inconsistent diffusion over time, sample 11 exhibited this unusually low diffusivity consistently throughout the entirety of the test, suggesting that this was not the result of a mistake in the H-cell assembly. Instead, this low rate of diffusion was linked to a wiring malfunction in the dual stir plate, which caused the magnetic stirrer positioned under sample 11 to stop working. As a result, the data collected from sample 11 represents diffusion in an unmixed H-cell. Since the malfunction occurred early on, this slower rate of diffusion remained consistent throughout the length of the experiment. The individual samples with inconsistent diffusion coefficients that could be linked to an experimental error or mechanical issue have been excluded from the results reported in Table 2, and the discovery of this wiring malfunction prompted a re-design of the dual stir plates to increase robustness and reliability. The updated designs for the dual stir plate are available in the GitHub repository.
3.3 Further applications
As mentioned previously, the python scripts, labware designs, and protocols for this workflow have been made available as a GitHub repository. Additional features have been included, such as dual H-cell holder designs for both 22 mL and 30 mL H-cells, options for testing 4 or 8 H-cells in parallel, and walkthrough instructions on executing the workflow and analyzing the data. There is also an optional feature within the protocol to automate the addition of the concentrated dye stock to each donor chamber, which reduces the time lapse between the addition of the dye at the onset of the experiment (t = 0) and the collection of the first sample aliquot (iteration 0). While the collection of UV-vis data remains a manual component in this workflow, this may be beneficial for research labs with limited resources or shared instrumentation. Whereas other approaches to membrane permeation testing may allow for a fully automated workflow – such as connecting the chamber(s) of an H-cell to a spectrophotometer flow cell with a recirculating system – doing so with multi-day diffusion tests would restrict the availability of the spectrophotometer to a single experiment. To parallelize such a workflow to match the capabilities of the OT2, which can fit up to 8 H-cells in a single round of experimentation, would require purchasing duplicates of all the equipment needed. The cost of purchasing multiple pumps and flow cells, and potentially even multiple spectrometers, can quickly become prohibitive, once again raising the barrier to entry to lab automation. Alternatively, the workflow presented here contains all experimentation within the footprint of an OT2 (or similar liquid-handling robot), and all characterization can be completed using a single well plate and a plate reader. Furthermore, the specialized equipment needed to implement this automation workflow – such as the stir plates, H-cell well plates, and 12-well scintillation vial plate – can all be 3D-printed and built at a low cost, and are easily modifiable to specific parameters, such as H-cells of varying shapes and sizes.While the implementation of this workflow was demonstrated here by measuring effective diffusion across hydrogel membranes, the depth and type analysis can be varied to meet the researcher's needs. Using variable sampling frequencies instead of a single interval, for example, can provide greater resolution into areas of interest. If membranes exhibit non-steady state transport at the onset of the experiment, such as reactive barrier membranes or those with high partitioning coefficients, the frequency of sampling could be increased at the start of the experiment to better identify the lag time/breakthrough time, and at what point the system reaches steady state.62–64 An example of using this variable frequency option has been included in the SI. Sample collection could also be used with other forms of analysis and characterization, such as fluorescence spectroscopy, liquid chromatography, conductivity, or pH, to name a few. With the current design of the workflow, experimental design space is somewhat limited to nonvolatile materials and solvents, as the H-cells must be left open for the OT2 to operate – that said, a robotic platform such as the OT2 can easily be stored in a fume hood for greater safety controls, and add-ons such as a wiper seal insert or pre-slit silicone cap could be used to reduce the size of the chamber openings to reduce evaporation. For a fully-sealed system, the workflow could be adapted to alternative platforms capable of handling liquid transfer with septa and syringe needles, such as the Science Jubilee.
4 Conclusions
In this work, we have presented and demonstrated a workflow for the automation of sample collection in multi-day H-cell diffusion tests. Developed to collect and prepare aliquots from membrane diffusion tests at regular intervals throughout the length of the experiment, this workflow allows for the parallelization of up to 8 membrane tests at a time. We demonstrated the use of this automation workflow in determining the diffusion coefficients of four aqueous dyes (brilliant blue, acid orange 7, rhodamine B, methylene blue) through two compositions of PEGDA hydrogel membranes. Furthermore, the usage of open hardware and software, in tandem with standardized equipment, not only make this workflow compatible with commercially available robots (such as the Opentrons OT2), but can also be translated to other material acceleration platforms, such as the Science Jubilee. The python scripts and custom hardware developed for this workflow have been made available in a GitHub repository, to be used according to the methods described here, or to be modified to fit other experimental parameters and characterization techniques.Author contributions
Claire Benstead: investigation (lead), methodology, software (equal), visualization (lead), writing – original draft preparation (lead), writing – review & editing (equal) Maria Politi: software (equal) David S. Bergsman: conceptualization, project administration, supervision (equal), writing – review & editing (equal) Lilo D. Pozzo: conceptualization, funding acquisition (lead), project administration, resources (lead), supervision (equal), writing – review & editing (equal).Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article are available at Zenodo at [https://doi.org/10.5281/zenodo.17238337]. Data and data analysis scripts are additionally available at GitHub at [https://github.com/pozzo-research-group/OT2-ADT].The code and labware designs for the automated diffusion testing protocol (ADT) can be found at [https://github.com/pozzo-research-group/OT2-ADT] with [https://doi.org/10.5281/zenodo.17676403]. The version of the code used for this study is version 1.0.0.
Supplementary information (SI): for equation derivations, conservation of mass analysis, partitioning tests, and small angle X-ray scattering (SAXS) analysis of hydrogel mesh sizes. See DOI: https://doi.org/10.1039/d5dd00523j.
Acknowledgements
This work was fully supported by the United States National Science Foundation (NSF) Emerging Frontiers in Research and Innovation award EFMA-2029249. The authors acknowledge the use of facilities and instrumentation supported by the U.S. National Science Foundation through the Major Research Instrumentation (MRI) program (DMR-2116265) and the UW Molecular Engineering Materials Center (MEM-C), a Materials Research Science and Engineering Center (DMR-2308979). We thank Benjamin Hornburg for assistance with designing and building the dual stir plates. We also thank David A. C. Beck for assisting with code development and preparation.Notes and references
- J. Zhang, A. N. Shocron, V. Meola, C. Violet, Z. Jiao, P. I. Lenz, Y. Duan, R. Wang, A. Iddya and M. Elimelech, Environ. Sci. Technol. Lett., 2025, 12, 1082–1088 CrossRef CAS.
- J. Kamcev, C. M. Doherty, K. P. Lopez, A. J. Hill, D. R. Paul and B. D. Freeman, J. Membr. Sci., 2018, 566, 307–316 CrossRef CAS.
- X. Tang, K. Guo, H. Li, Z. Du and J. Tian, Biochem. Eng. J., 2010, 52, 194–198 CrossRef CAS.
- M. S. Cha, H. Y. Jeong, H. Y. Shin, S. H. Hong, T.-H. Kim, S.-G. Oh, J. Y. Lee and Y. T. Hong, J. Power Sources, 2017, 363, 78–86 Search PubMed.
- Y. Ye, H. H. Ngo, W. Guo, Y. Liu, S. W. Chang, D. D. Nguyen, J. Ren, Y. Liu and X. Zhang, Chem. Eng. J., 2019, 358, 236–242 Search PubMed.
- J. Kamcev, E.-S. Jang, N. Yan, D. R. Paul and B. D. Freeman, J. Membr. Sci., 2015, 479, 55–66 CrossRef CAS.
- S. P. Stier, C. Kreisbeck, H. Ihssen, M. A. Popp, J. Hauch, K. Malek, M. Reynaud, T. Goumans, J. Carlsson, I. Todorov, L. Gold, A. Räder, W. Wenzel, S. T. Bandesha, P. Jacques, F. Garcia-Moreno, O. Arcelus, P. Friederich, S. Clark, M. Maglione, A. Laukkanen, I. E. Castelli, J. Carrasco, M. C. Cabanas, H. S. Stein, O. Ozcan, D. Elbert, K. Reuter, C. Scheurer, M. Demura, S. S. Han, T. Vegge, S. Nakamae, M. Fabrizio and M. Kozdras, Adv. Mater., 2024, 36, 2407791 CrossRef CAS PubMed.
- M. Politi, F. Baum, K. Vaddi, E. Antonio, J. Vasquez, B. P. Bishop, N. Peek, V. C. Holmberg and L. D. Pozzo, Digital Discovery, 2023, 2, 1042–1057 RSC.
- S. Doloi, M. Das, Y. Li, Z. H. Cho, X. Xiao, J. V. Hanna, M. Osvaldo and L. N. W. Tat, Digital Discovery, 2025, 4, 1685–1721 Search PubMed.
- B. Pelkie, S. Baird, E. Aissi, K. Aspuru-Takata, Y. Cao, J. H. Chang, K. Gambhir, W. S. Hale, L. Hao, C. Hattrick, J. Hein, D. Luo, O. Melville, M. Ngan, L. L. B. Nyeland, N. Peek, M. Politi, E. E. Rajkumar, A. Siemenn, B. Subbaraman, S. Vasquez, J. Watchorn, W. Zhang, R. Ziskason, L. Pozzo, T. Buonassisi and T. Vegge, Democratizing self-driving labs through user-developed automation infrastructure, 2025, https://chemrxiv.org/engage/chemrxiv/article-details/67a4ffb6fa469535b94a3ad9 Search PubMed.
- J. A. Bryant Jr, M. Kellinger, C. Longmire, R. Miller and R. C. Wright, Synth. Biol., 2023, 8, ysac032 CrossRef PubMed.
- H. M. Brown and P. W. Fedick, React. Chem. Eng., 2023, 8, 556–562 RSC.
- B. W. Soh, A. Chitre, S. Z. Tan, Y. Wang, Y. Yi, W. Soh, K. Hippalgaonkar and D. I. Wilson, Digital Discovery, 2025, 4, 711–722 Search PubMed.
- J. A. Bryant, C. Longmire, S. Sridhar, S. Janousek, M. Kellinger and R. C. Wright, SLAS Technol., 2024, 29, 100107 CrossRef PubMed.
- J. Abed, Y. Bai, D. Persaud, J. Kim, J. Witt, J. Hattrick-Simpers and E. H. Sargent, Digital Discovery, 2024, 3, 2265–2274 Search PubMed.
- N. Fisker-Bødker, D. Persaud, Y. Bai, M. Kozdras, T. Vegge, J. Hattrick-Simpers and J. H. Chang, Digital Discovery, 2025, 2491–2501 Search PubMed.
- S. Detassis, M. Grasso, M. Tabraue-Chávez, A. Marín-Romero, B. López-Longarela, H. Ilyine, C. Ress, S. Ceriani, M. Erspan, A. Maglione, J. J. Díaz-Mochón, S. Pernagallo and M. A. Denti, Anal. Chem., 2019, 91, 5874–5880 CrossRef CAS PubMed.
- S. Detassis, F. Precazzini, I. Brentari, R. Ruffilli, C. Ress, A. Maglione, S. Pernagallo and M. A. Denti, Analyst, 2024, 149, 3891–3899 RSC.
- Z. W. Muetzel, J. A. Ouimet and W. A. Phillip, ACS Appl. Polym. Mater., 2022, 4, 3438–3447 CrossRef CAS.
- J. A. Ouimet, F. Al-Badani, X. Liu, L. Lair, Z. W. Muetzel, A. W. Dowling and W. A. Phillip, J. Membr. Sci. Lett., 2024, 4, 100087 CrossRef.
- A. S. Kazemi and D. R. Latulippe, J. Membr. Sci., 2014, 470, 30–39 Search PubMed.
- I. Arias Ponce, R. Sujanani, J. D. Moon, J. M. Urueña, C. J. Hawker and R. A. Segalman, ACS Appl. Polym. Mater., 2024, 6, 14629–14637 CrossRef CAS.
- H. Feil, Y. H. Bae, J. Feijen and S. W. Kim, J. Membr. Sci., 1991, 64, 283–294 Search PubMed.
- H. Chang, H. Zhao, F. Qu, Z. Yan, N. Liu, M. Lu, Y. Liang, B. Lai and H. Liang, Sep. Purif. Technol., 2023, 308, 122948 CrossRef CAS.
- M. K. Yazdi, V. Vatanpour, A. Taghizadeh, M. Taghizadeh, M. R. Ganjali, M. T. Munir, S. Habibzadeh, M. R. Saeb and M. Ghaedi, Mater. Sci. Eng., C, 2020, 114, 111023 Search PubMed.
- A. C. Sagle, H. Ju, B. D. Freeman and M. M. Sharma, Polymer, 2009, 50, 756–766 Search PubMed.
- R. Huang, L. K. Kostanski, C. D. M. Filipe and R. Ghosh, J. Membr. Sci., 2009, 336, 42–49 CrossRef CAS.
- F. M. Andreopoulos, E. J. Beckman and A. J. Russell, Biomaterials, 1998, 19, 1343–1352 Search PubMed.
- Y.-H. P. Zhang, J. Sun and Y. Ma, J. Ind. Microbiol. Biotechnol., 2017, 44, 773–784 CrossRef CAS PubMed.
- N. Rekowska, D. Arbeiter, H. Seitz, R. Mau, A. Riess, T. Eickner, N. Grabow and M. Teske, Curr. Dir. Biomed. Eng., 2022, 8, 181–184 CrossRef.
- A. Augustine, M. Veillerot, N. Gauthier, B. Zhu, C.-Y. Hui, Y. Tran, E. Verneuil and A. Chateauminois, Soft Matter, 2023, 19, 5169–5178 RSC.
- F. V. Lavrentev, V. V. Shilovskikh, V. S. Alabusheva, V. Y. Yurova, A. A. Nikitina, S. A. Ulasevich and E. V. Skorb, Molecules, 2023, 28, 5931 Search PubMed.
- C. T. Reinhart and N. A. Peppas, J. Membr. Sci., 1984, 18, 227–239 CrossRef CAS.
- M. Levin, Y. Tang, C. D. Eisenbach, M. T. Valentine and N. Cohen, Macromolecules, 2024, 57, 7074–7086 CrossRef CAS PubMed.
- B. Amsden, Macromolecules, 1998, 31, 8382–8395 CrossRef CAS.
- B. G. Amsden, Macromolecules, 2022, 55, 8399–8408 Search PubMed.
- N. R. Richbourg and N. A. Peppas, Prog. Polym. Sci., 2020, 105, 101243 Search PubMed.
- G. J. Moridis, Water Resour. Res., 1999, 35, 1729–1740 CrossRef.
- S. J. Rukmani, P. Lin, J. S. Andrew and C. M. Colina, J. Phys. Chem. B, 2019, 123, 4129–4138 CrossRef CAS PubMed.
- K. McAvoy, D. Jones and R. R. S. Thakur, Pharm. Res., 2018, 35, 36 CrossRef PubMed.
- C. A. Durst, M. P. Cuchiara, E. G. Mansfield, J. L. West and K. J. Grande-Allen, Acta Biomater., 2011, 7, 2467–2476 CrossRef CAS PubMed.
- J. Vasquez, H. Twigg-Smith, J. Tran O'Leary and N. Peek, Proceedings of the 2020 CHI Conference on Human Factors in Computing Systems, New York, NY, USA, 2020, pp. 1–13 Search PubMed.
- C. Benstead and M. Politi, OT2-ADT, 2025, https://github.com/pozzo-research-group/OT2-ADT.
- D. C. Harris and M. D. Bertolucci, Symmetry and Spectroscopy: An Introduction to Vibrational and Electronic Spectroscopy, Dover Publications, New York, 1989 Search PubMed.
- K. Engberg and C. W. Frank, Biomed. Mater., 2011, 6, 055006 CrossRef PubMed.
- C. J. Lee, J. A. Vroom, H. A. Fishman and S. F. Bent, Biomaterials, 2006, 27, 1670–1678 CrossRef CAS PubMed.
- C. K. Colton, K. A. Smith, E. W. Merrill and P. C. Farrell, J. Biomed. Mater. Res., 1971, 5, 459–488 CrossRef CAS PubMed.
- A. Cavallo, M. Madaghiele, U. Masullo, M. G. Lionetto and A. Sannino, J. Appl. Polym. Sci., 2017, 134, 44380 CrossRef.
- A. Zech and M. de Winter, Transp. Porous Media, 2023, 146, 475–492 CrossRef.
- H. A. Balogun and R. P. Lively, J. Membr. Sci., 2026, 737, 124731 Search PubMed.
- R. Syms, Biomicrofluidics, 2017, 11, 044116 Search PubMed.
- S. M. Evans, A. L. Litzenberger, A. E. Ellenberger, J. E. Maneval, E. L. Jablonski and B. M. Vogel, Mater. Sci. Eng., C, 2014, 35, 322–334 CrossRef CAS PubMed.
- S. A. Rani, B. Pitts and P. S. Stewart, Antimicrob. Agents Chemother., 2005, 49, 728–732 CrossRef CAS PubMed.
- P.-O. Gendron, F. Avaltroni and K. J. Wilkinson, J. Fluoresc., 2008, 18, 1093–1101 Search PubMed.
- C. T. Culbertson, S. C. Jacobson and J. Michael Ramsey, Talanta, 2002, 56, 365–373 CrossRef CAS PubMed.
- D. G. Leaist, J. Colloid Interface Sci., 1988, 125, 327–332 CrossRef CAS.
- H. Ju, B. D. McCloskey, A. C. Sagle, V. A. Kusuma and B. D. Freeman, J. Membr. Sci., 2009, 330, 180–188 CrossRef CAS.
- P. Malo de Molina, S. Lad and M. E. Helgeson, Macromolecules, 2015, 48, 5402–5411 CrossRef CAS.
- L. T. Hirschwald, S. Brosch, G. Linz, J. Linkhorst and M. Wessling, Adv. Mater. Technol., 2023, 8, 2201857 CrossRef CAS.
- K. O'Donnell, A. Boyd and B. J. Meenan, Materials, 2019, 12, 3381 CrossRef PubMed.
- M. Chen, K. R. Kumrić, C. Thacker, R. Prodanović, G. Bolognesi and G. T. Vladisavljević, Gels, 2023, 9, 849 CrossRef CAS PubMed.
- R. A. Siegel and E. L. Cussler, J. Membr. Sci., 2004, 229, 33–41 CrossRef CAS.
- H. Wu, J. Thibault and B. Kruczek, J. Membr. Sci., 2021, 618, 118715 Search PubMed.
- D. Bai, F. Asempour and B. Kruczek, Chem. Eng. Res. Des., 2020, 162, 228–237 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |





