Open Access Article
Open Access ArticleA feature-aligned diffusion model for controllable generation of 3D drug-like molecules
Hao
Lu
 ab,
Zhiqiang
Wei
ae,
Xiancong
Hou
a,
Wenzheng
Han
a,
Yang
Zhang
*bcd and
Hao
Liu
*a
ab,
Zhiqiang
Wei
ae,
Xiancong
Hou
a,
Wenzheng
Han
a,
Yang
Zhang
*bcd and
Hao
Liu
*a
aCollege of Computer Science and Technology, Ocean University of China, Qingdao, 266100, Shandong Province, China. E-mail: liu.hao@ouc.edu.cn; luhao@stu.ouc.edu.cn; weizhiqiang@ouc.edu.cn; houxiancong@stu.ouc.edu.cn; hanwenzheng@stu.ouc.edu.cn
bDepartment of Computer Science, School of Computing, National University of Singapore, 117417, Singapore. E-mail: zhang@nus.edu.sg
cCancer Science Institute of Singapore, National University of Singapore, 117599, Singapore
dDepartment of Biochemistry, Yong Loo Lin School of Medicine, National University of Singapore, 117596, Singapore
eCollege of Computer Science and Technology, Qingdao University, Qingdao, 266071, Shandong Province, China
First published on 25th February 2026
Abstract
Structure-based drug design has increasingly leveraged deep learning approaches, particularly diffusion models. Nevertheless, their practical application is hindered by noise-induced deterioration of biological relevance and binding affinity. We propose a dynamic, feature-aligned diffusion framework guided by expert knowledge. The core innovation lies in a feature fusion mechanism that extracts molecular features in real time through a pre-trained expert network and aligns them cross-modally with the latent space of the diffusion trajectory. Additionally, we introduce a dynamic weight adjustment module, which adaptively adjusts the strength of the feature alignment constraint based on the noise intensity, enabling a progressive optimization from coarse- to fine-grained structures. Experimental results demonstrate that our model achieves an average Vina docking score of −10.06, together with favorable binding free energies. Moreover, it reduces structural clashes and improves drug-likeness with respect to bonding and geometry. This work presents a new paradigm that synergistically integrates diffusion models with autoregressive mechanisms and holds strong potential for advancing AI-driven drug discovery.
1 Introduction
Structure-based drug design (SBDD) has advanced significantly in recent years, substantially improving the potency and rationality in finding novel molecular structures.1–3 Given a defined protein pocket, the primary objective is to produce drug-like compounds that not only exhibit strong binding affinity but also conform to the physicochemical and pharmacokinetic properties typical of therapeutic agents. Traditional methods rely heavily on physics-based simulations and experimental trial-and-error, which are inherently limited by the size of available molecular libraries and the accuracy of the force fields. In contrast, deep learning offers the potential to transcend these limitations by leveraging data-driven, implicit rule learning.4–7 Deep learning-based generative models, such as Generative Adversarial Networks (GANs),8 Variational Autoencoders (VAEs),9 and diffusion models,10 have been widely applied to molecular generation tasks, capable of learning effective molecular representations in large chemical spaces and generating molecular structures that comply with chemical rules.11–16 Compared to GANs and VAEs, diffusion models have the advantage of generating higher quality and more diverse molecular samples, while maintaining chemical plausibility and providing stronger modeling capabilities for optimizing target binding affinity.Despite the promising capabilities of diffusion models, existing strategies can be broadly categorized into two paradigms in SBDD: fragment-based optimization relying on known pharmacophores, and de novo design exploring novel chemical space. Recent successes like Delete17 demonstrate high-affinity linker design using fragment-based strategies. However, these methods rely on predefined pharmacological fragments, limiting their ability to explore novel and diverse molecular structures. In contrast, de novo design aims to generate molecules from scratch, but often struggles to achieve strong bioactivity.18,19 Furthermore, effectively controlling the impact of noise and improving the rationality and feasibility of the generated molecules during the complex training process remain open issues that current methods have not fully resolved.
In diffusion models, the molecular generation process reconstructs structures progressively through a Markov chain.10,20 However, the injection of Gaussian noise during the forward process makes it difficult for the model to accurately recover stable molecular structures. The intermediate feature representations become increasingly unstable, and relying solely on stepwise denoising is often insufficient for preserving critical chemical information. While some methods integrate chemical knowledge or impose bonding constraints,21–25 their effectiveness in significantly improving bioactivity remains modest. Moreover, since noise levels vary across diffusion steps, there are notable differences in the feature distribution at each stage. This variation often causes structural clashes between the generated molecules and the residues of the protein pocket, undermining binding compatibility and overall molecular feasibility.
To address these challenges in de novo molecular generation, we introduce a diffusion-based framework that integrates expert-guided feature alignment to improve molecular generation under protein pocket constraints. The core idea is to stabilize and steer the generative process by incorporating a pre-trained expert network that can provide semantic feedback during denoising. This expert-guided mechanism enables the model to better retain binding-relevant information across the noisy diffusion trajectory and enhances compatibility with pharmacologically meaningful features. Furthermore, our framework incorporates a noise-aware alignment strategy that dynamically adapts to the varying uncertainty at different diffusion steps, which is crucial for preserving structural validity and target specificity. By embedding this expert knowledge into the generative process, our method advances beyond existing approaches in producing molecules with lower docking scores, stronger binding free affinity, improved biological relevance, and greater structural coherence. This study aims to bridge the gap between generative modeling and real-world applicability in de novo structure-based drug design, offering a scalable and generalizable solution across diverse protein targets.
2 Results
We propose an expert network-aligned diffusion model, termed ExpDiff, for the generation of high-affinity, drug-like molecules from scratch conditioned on protein binding pockets. This task poses a significant challenge due to the need for simultaneous modeling of intricate intermolecular interactions and compliance with pharmacological constraints. The overall framework of ExpDiff can be seen in Fig. 1a. Both protein pockets and ligands are represented as point clouds, with atomic identities and coordinates subjected to noise perturbation. An equivariant graph neural network (EGNN) is employed as the denoising backbone, iteratively predicting and removing noise to reconstruct valid molecular structures.Our method introduces a dynamic feature alignment mechanism guided by an expert network (see Section 4.2) to maintain the stability of intermediate features. The expert network provides feedback to the diffusion model, enhancing its ability to generate molecules that better match the target pharmacological properties and binding requirements. In addition, we suggest diffusion step-aware alignment (see Section 4.3), which allows our approach to maximize feature alignment at various noise levels, lessen noise interference, and enhance the quality of molecules produced, in contrast to conventional fine-tuning techniques. To guide the generation process toward high-affinity candidates, we train an affinity predictor jointly with the expert network and use its gradients to steer inference-time sampling (see Fig. 1b). In addition, the final point cloud is post-processed into chemically valid molecules by reconstructing bonds based on interatomic distances within a cutoff threshold (see Fig. 1c).
Together, these mechanisms empower ExpDiff to more effectively explore the conditional chemical space and yield ligands with enhanced protein–ligand interactions and physical realism. Quantitative and qualitative evaluations confirm that ExpDiff achieves enhanced docking scores and binding free energy while maintaining drug-likeness and geometric plausibility, representing a substantial advancement in SBDD (see Sections 2.1–2.6) for detailed analysis.
2.1 Molecular generation based on protein pockets
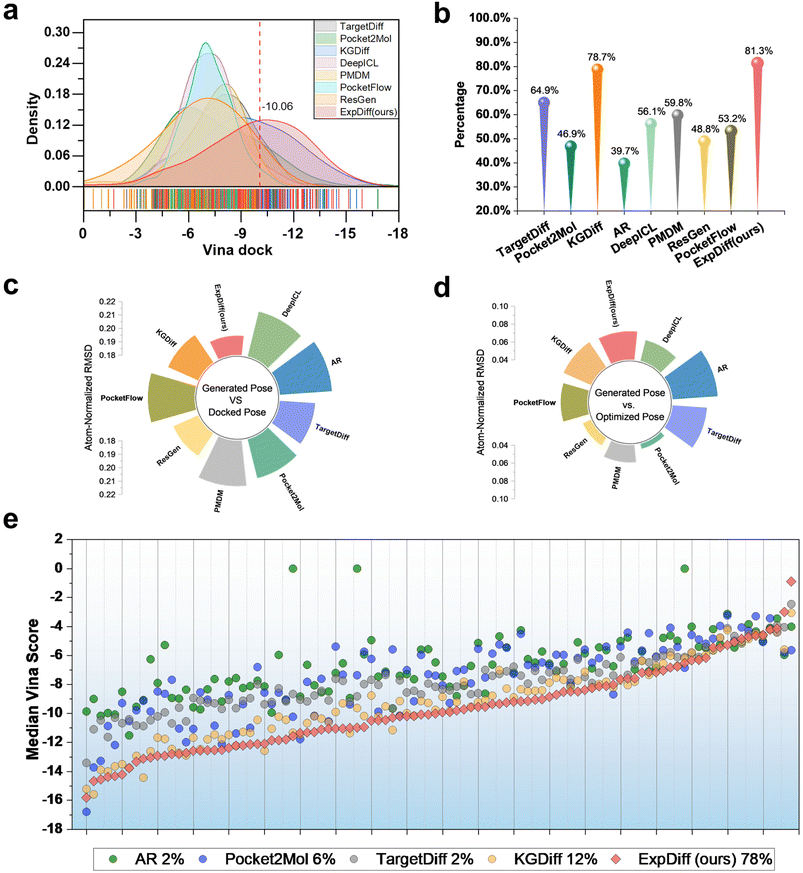
SBDD requires generating molecules with robust predicted binding affinity. To evaluate the binding potential of molecules generated by ExpDiff, we employed molecular docking against 100 test targets from the CrossDocked2020 dataset, generating 100 molecules per target and comparing their docking results with those of baseline models.As can be seen from Table 1, the diffusion models consistently outperform autoregressive approaches across all evaluation metrics, highlighting their clear advantage in molecular generation tasks. Among the diffusion-based methods, ExpDiff achieves superior performance on nearly all metrics, particularly excelling Vina-related evaluations. Notably, ExpDiff achieves the best results on three key metrics, namely, Vina score, Vina minimize, and Vina dock(see Fig. 2a). Specifically, ExpDiff surpasses the second-best model by 1.1 in Vina score and 0.63 in Vina dock score. It is also worth noting that some baseline models produce positive Vina scores, suggesting that the 3D molecules they generate often exhibit severe steric clashes within the docking pocket residues. In contrast, ExpDiff generates structurally coherent molecules with fewer geometric violations, resulting in more favorable docking scores.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 generated molecules sampled using ExpDiff on 100 test targets trained on the CrossDocked 2020 dataset with those using the baseline models. The bold values indicate the best performance results in each category. The SA values here are normalized. The bold values indicate the best performance results. The SA values here are normalized. ↑ means the higher the better and ↓ means the lower the better. Ave. indicates average. Med. indicates median. - indicates that the Vina result is greater than 0, indicating substantial clashes
000 generated molecules sampled using ExpDiff on 100 test targets trained on the CrossDocked 2020 dataset with those using the baseline models. The bold values indicate the best performance results in each category. The SA values here are normalized. The bold values indicate the best performance results. The SA values here are normalized. ↑ means the higher the better and ↓ means the lower the better. Ave. indicates average. Med. indicates median. - indicates that the Vina result is greater than 0, indicating substantial clashes
| Method | Vina score (↓) | Vina minimize (↓) | Vina dock (↓) | High affinity (↑) | QED (↑) | SA (↑) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ave | Med | Ave | Med | Ave | Med | Ave | Med | Ave | Med | Ave | Med | ||
| Reference | −6.36 | −6.46 | −6.71 | −6.49 | −7.45 | −7.26 | — | — | 0.48 | 0.47 | 0.73 | 0.74 | |
| Auto-regressive model | ResGen | — | — | — | −3.42 | −6.14 | −6.92 | 50.3% | 55.9% | 0.50 | 0.51 | 0.73 | 0.73 |
| AR | −5.75 | −5.64 | −6.18 | −5.88 | −6.75 | −6.62 | 37.9% | 31.0% | 0.51 | 0.50 | 0.63 | 0.63 | |
| Pocket2Mol | −5.14 | −4.70 | −6.42 | −5.82 | −7.15 | −6.79 | 48.4% | 51.0% | 0.56 | 0.57 | 0.74 | 0.75 | |
| PocketFlow | −3.02 | −4.10 | −5.49 | −5.55 | −7.18 | −7.19 | 51.3% | 53.1% | 0.53 | 0.53 | 0.81 | 0.82 | |
| Diffusion model | AUTODIFF | −5.25 | −5.33 | −6.91 | −7.06 | −8.84 | −8.94 | 73.0% | 77.0% | 0.57 | 0.58 | 0.76 | 0.77 |
| TargetDiff | −5.50 | −6.32 | −6.69 | −6.86 | −7.83 | −7.92 | 59.2% | 60.4% | 0.49 | 0.49 | 0.59 | 0.58 | |
| TAGMol | −7.02 | −7.77 | −7.95 | −8.07 | −8.59 | −8.69 | 69.8% | 76.4% | 0.55 | 0.56 | 0.56 | 0.56 | |
| DeepICL | −3.79 | −4.12 | −5.88 | −5.78 | −7.13 | −7.29 | 53.7% | 59.0% | 0.61 | 0.62 | 0.33 | 0.30 | |
| PMDM | — | −3.48 | −2.68 | −5.86 | −7.06 | −7.66 | 56.6% | 60.6% | 0.56 | 0.57 | 0.62 | 0.62 | |
| KGDiff | −8.04 | −8.61 | −8.78 | −8.85 | −9.43 | −9.43 | 79.2% | 87.0% | 0.51 | 0.51 | 0.54 | 0.54 | |
| BINDDM | −5.92 | −6.81 | −7.29 | −7.34 | −8.41 | −8.37 | 64.8% | 71.6% | 0.51 | 0.52 | 0.58 | 0.58 | |
| DecompOpt | −5.87 | −6.81 | −7.35 | −7.72 | −8.95 | −9.01 | 73.5% | 93.3% | 0.48 | 0.45 | 0.65 | 0.65 | |
| DecompDiff | −5.67 | −6.04 | −7.04 | −7.09 | −8.39 | −8.43 | 64.4% | 71.0% | 0.45 | 0.43 | 0.61 | 0.60 | |
| IRDiff | −6.03 | −6.89 | −7.27 | −7.37 | −8.42 | −8.42 | 67.4% | 72.7% | 0.53 | 0.54 | 0.59 | 0.58 | |
| IPDiff | −6.42 | −7.01 | −7.45 | −7.48 | −8.57 | −8.51 | 69.5% | 75.5% | 0.52 | 0.53 | 0.61 | 0.69 | |
| FlexSBDD | −6.64 | −7.25 | −8.27 | −8.46 | −9.12 | −9.25 | 78.5% | 84.2% | 0.58 | 0.59 | 0.69 | 0.73 | |
| BokDiff | −5.92 | −5.89 | −7.50 | −7.17 | −8.58 | −8.41 | — | — | 0.48 | 0.49 | 0.60 | 0.60 | |
| ALIDiff | −7.07 | −7.95 | −8.09 | −8.17 | −8.90 | −8.81 | 73.4% | 81.4% | 0.50 | 0.50 | 0.57 | 0.56 | |
| ExpDiff(ours) | −9.14 | −9.69 | −9.65 | −9.92 | −10.06 | −10.25 | 82.0% | 93.2% | 0.54 | 0.55 | 0.56 | 0.55 | |
As shown in Fig. 2b, ExpDiff achieves the highest proportion of molecules with Vina docking scores below −7 when compared with baseline models, indicating a strong capacity to generate candidates with favorable predicted binding potential. Structural analysis further supports this finding: Fig. 2c and d report the atom-normalized RMSD between the generated poses and the docked poses, and between the generated poses and the optimized poses, respectively. The RMSD relative to docked poses is the lowest, whereas the RMSD relative to optimized poses lies at an intermediate level. This observation suggests that the conformations generated by ExpDiff more closely approximate the docked conformations, highlighting its ability to capture spatial interactions within protein binding pockets. These results collectively demonstrate that ExpDiff not only improves molecular quality but also enhances target binding potential, making it a highly effective model for SBDD.
To further evaluate the docking-based binding potential of ExpDiff to generate molecules at a single target, we compared the median docking scores of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 sampled molecules generated for each individual target against those of baseline models. Targets were ranked based on the lowest docking scores achieved by ExpDiff. As shown in Fig. 2e, ExpDiff achieved the best docking-based binding potential on 78% of the 100 targets, outperforming other models such as KGDiff (12%) and Pocket2Mol (6%). These results underscore ExpDiff's robust generalization and superior capability in generating molecules with strong target binding across a wide range of protein pockets. Collectively, these findings demonstrate that ExpDiff consistently outperforms both diffusion-based and autoregressive baselines in producing molecules with enhanced predicted binding affinity and structural validity, making it a top-performing model for SBDD.
000 sampled molecules generated for each individual target against those of baseline models. Targets were ranked based on the lowest docking scores achieved by ExpDiff. As shown in Fig. 2e, ExpDiff achieved the best docking-based binding potential on 78% of the 100 targets, outperforming other models such as KGDiff (12%) and Pocket2Mol (6%). These results underscore ExpDiff's robust generalization and superior capability in generating molecules with strong target binding across a wide range of protein pockets. Collectively, these findings demonstrate that ExpDiff consistently outperforms both diffusion-based and autoregressive baselines in producing molecules with enhanced predicted binding affinity and structural validity, making it a top-performing model for SBDD.
2.2 Molecular drug-like properties
To analyze the drug-like properties of the generated molecules by ExpDiff, we tested several key physicochemical and pharmacological criteria, including quantitative estimate of drug-likeness (QED), synthetic accessibility score (SAS), logarithm of the partition coefficient (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), topological polar surface area (TPSA), molecular weight, and the number of rotatable bonds, which have been widely used in drug discovery to assess the drug-like potential of generated molecules.
P), topological polar surface area (TPSA), molecular weight, and the number of rotatable bonds, which have been widely used in drug discovery to assess the drug-like potential of generated molecules.
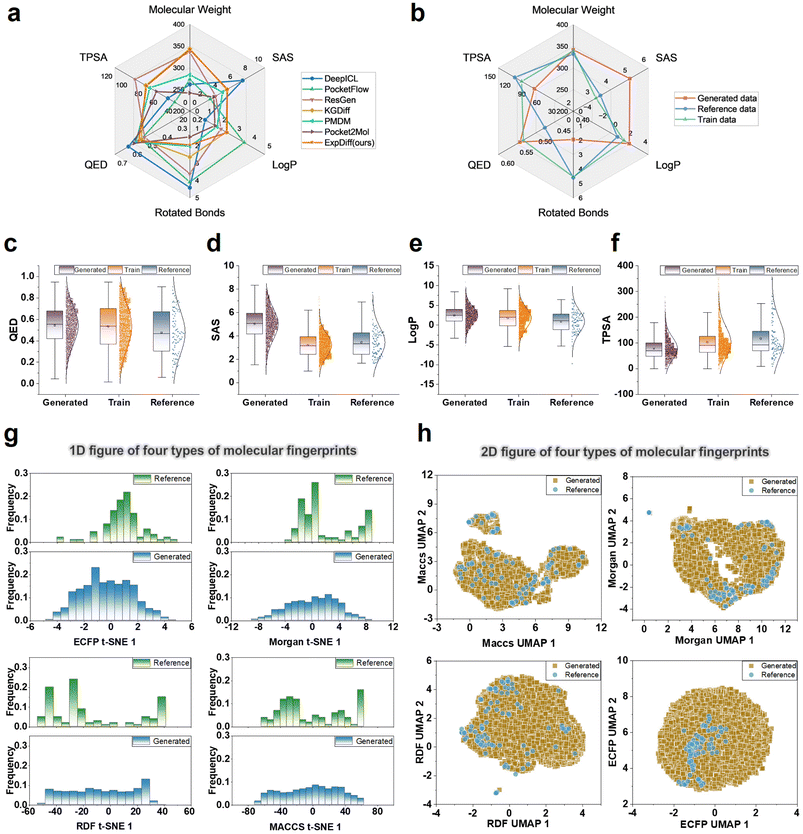
As shown in Fig. 3a, DeepICL improves QED. However, this gain is largely offset by compromises in other molecular properties, such as SAS. In contrast, molecules generated by ExpDiff exhibit all six properties within the intermediate range of the baseline models, while simultaneously achieving enhanced predicted docking potential. Compared with the training and reference data, the generated molecules show a moderate increase in synthetic difficulty, reflecting a balance between drug-likeness and chemical feasibility (see Fig. 3b). Fig. 3c presents the QED values used to measure the drug-like properties of the molecules, which range from 0 to 1. Most of the QED values of the data generated by ExpDiff are clustered in the interval of 0.4 to 0.6, closely matching the training set. In contrast, the reference set exhibits lower QED scores, indicating that the generated data tend to be at a higher level of drug-like properties. The wider QED distribution of the generated data may imply the complexity and diversity of the generated molecules. Fig. 3d exhibits the SAS values reflecting the synthesizability of the molecules, with lower values indicating lower synthetic difficulty. In this figure, the SAS values of the generated molecules are indeed higher than those of the training and reference sets, suggesting that the generated compounds tend to adopt more structurally complex chemotypes and therefore lower synthesizability. However, when compared with existing baseline generative models, ExpDiff does not exhibit a substantial loss in synthetic accessibility. Across the eighteen baseline methods evaluated, the average SA score is 0.62, whereas ExpDiff achieves an average score of 0.56. Although our method focuses on generating molecules with significantly improved predicted binding affinity, the resulting SA values remain competitive and in some cases superior to those of prior models. These results indicate that ExpDiff enhances docking performance without imposing an undue increase in synthetic difficulty. Fig. 3e depicts the hydrophobicity of the molecule. Considering that the desirable log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values are usually between 1 and 3, such compounds are neither too hydrophobic nor too hydrophilic, contributing to an efficient distribution in vivo and in vitro. Relative to the training set and the reference molecules, ExpDiff generates a more concentrated range of molecules and greatly concentrates between 0 and 5, showing the ability to strike a balance between hydrophilicity and cell membrane penetration. Fig. 3f depicts the polar surface area of the molecule, which is essential for the absorption and penetration of biological membranes. The overall low TPSA values of the generated data may affect the bioavailability of the molecules. In contrast, the TPSA values of the training and reference sets are clustered in a higher range, showing their advantages in terms of biocompatibility.
P values are usually between 1 and 3, such compounds are neither too hydrophobic nor too hydrophilic, contributing to an efficient distribution in vivo and in vitro. Relative to the training set and the reference molecules, ExpDiff generates a more concentrated range of molecules and greatly concentrates between 0 and 5, showing the ability to strike a balance between hydrophilicity and cell membrane penetration. Fig. 3f depicts the polar surface area of the molecule, which is essential for the absorption and penetration of biological membranes. The overall low TPSA values of the generated data may affect the bioavailability of the molecules. In contrast, the TPSA values of the training and reference sets are clustered in a higher range, showing their advantages in terms of biocompatibility.
To further evaluate the chemical properties of the molecules generated by ExpDiff, we obtained the molecular fingerprints in four different ways, reduced the dimensionality using the t-SNE and UMAP, and then examined the distribution of the chemical properties. The one-dimensional plots of the generated molecular four molecular fingerprints are displayed in Fig. 3g. The figure indicates that the features of the generated molecules are distributed fairly uniformly, whereas the reference molecules exhibit some variation. This can be because the reference molecule only has 100 molecules, which is a minimal quantity of data. Fig. 3h shows the two-dimensional plots of the four molecular fingerprints of the generated molecules, and it can be seen that the properties of the generated molecules and the reference molecules have the same space. Overall, the ExpDiff-generated molecules differ very little from the training set and the molecular properties in the reference.
Our method demonstrates advantages in generating larger ring systems compared to existing methods. Prior studies, such as those by Diao et al.,26 who focused on macrocyclic molecule design, and Patrick et al.,27 who developed macrocyclic peptides, have highlighted the therapeutic potential of macrocycles in drug discovery. As shown in Table 2, ExpDiff markedly outperforms previous methods in generating macrocyclic structures (ring size bigger than 6), with higher proportions of 7-membered rings (17.2%), 8-membered rings (4.7%) and 9-membered rings (1.5%). These results suggest that our model can capture the structural features needed to generate larger and more diverse ring systems. Our intention is not to claim any inherent superiority of macrocycles, but rather to highlight the model′s ability to explore broader chemical space. Although macrocyclic structures may be synthetically challenging, they represent an important class of molecules with distinctive conformational properties. In this sense, ExpDiff's enhanced capability to produce varied macrocyclic scaffolds underscores its usefulness for exploratory molecular design.
| Ring size | Ref. | liGAN | AR | Pocket2Mol | TargetDiff | ExpDiff (ours) |
|---|---|---|---|---|---|---|
| 3 | 1.7% | 28.1% | 29.9% | 0.1% | 0.0% | 0.0% |
| 4 | 0.0% | 15.7% | 0.0% | 0.0% | 2.6% | 1.3% |
| 5 | 30.2% | 29.8% | 16.0% | 16.4% | 30.8% | 21.7% |
| 6 | 67.4% | 22.7% | 51.2% | 80.4% | 50.7% | 53.7% |
| 7 | 0.7% | 2.6% | 1.7% | 2.6% | 12.1% | 17.2% |
| 8 | 0.0% | 0.8% | 0.7% | 0.3% | 2.7% | 4.7% |
| 9 | 0.0% | 0.3% | 0.5% | 0.1% | 0.9% | 1.5% |
2.3 Molecular geometry properties
Molecules are composed of various types of chemical bonds, such as single, double, and aromatic bonds, and a model's ability to accurately reproduce the distribution of these bond types serves as a direct indicator of its capacity to learn underlying chemical principles and ensure molecular stability. To assess the structural plausibility of generated molecules, we compute the Jensen Shannon Divergence (JSD) between the bond type distributions of generated molecules and those in the real dataset. A lower JSD value indicates a higher similarity, suggesting that the model generates molecules whose bond structures are more consistent with those found in real-world chemical compounds.
Table 3 shows that ExpDiff achieves the lowest JSD for all key bond types, demonstrating its excellent ability to generate molecular structures that closely match real-world distributions. In particular, ExpDiff outperforms all baseline models in reproducing aromatic (C@C, JSD = 0.140) and conjugate bonds (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, JSD = 0.198; C@N, JSD = 0.224), which are critical for molecular stability and biological activity. Notably, ExpDiff improved the generation of heteroatomic bonds such as C–N (JSD = 0.258) and C
C, JSD = 0.198; C@N, JSD = 0.224), which are critical for molecular stability and biological activity. Notably, ExpDiff improved the generation of heteroatomic bonds such as C–N (JSD = 0.258) and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (JSD = 0.340), reflecting a better balance between functional group diversity and chemical effectiveness. Compared to state-of-the-art methods such as TargetDiff and TAGMol, our model has a more refined understanding of chemical patterns and can generate structurally realistic molecules with improved fidelity. We further demonstrate the advantages of ExpDiff in bond lengths, bond angles, and atomic clashes of the generated molecules. More experimental procedures can be found in Text S1 (SI).
O (JSD = 0.340), reflecting a better balance between functional group diversity and chemical effectiveness. Compared to state-of-the-art methods such as TargetDiff and TAGMol, our model has a more refined understanding of chemical patterns and can generate structurally realistic molecules with improved fidelity. We further demonstrate the advantages of ExpDiff in bond lengths, bond angles, and atomic clashes of the generated molecules. More experimental procedures can be found in Text S1 (SI).
| Bond | liGAN | Pocket2Mol | TargetDiff | KGDiff | TAGMol | ExpDiff (ours) |
|---|---|---|---|---|---|---|
| C–C | 0.601 | 0.496 | 0.374 | 0.379 | 0.384 | 0.344 |
C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
0.665 | 0.561 | 0.500 | 0.559 | 0.501 | 0.198 |
| C@C | 0.497 | 0.416 | 0.265 | 0.205 | 0.269 | 0.140 |
| C–N | 0.634 | 0.416 | 0.365 | 0.380 | 0.365 | 0.258 |
C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
0.749 | 0.629 | 0.553 | 0.583 | 0.559 | 0.225 |
| C@N | 0.638 | 0.487 | 0.224 | 0.327 | 0.252 | 0.224 |
| C–O | 0.656 | 0.454 | 0.418 | 0.443 | 0.422 | 0.320 |
C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O O |
0.661 | 0.516 | 0.468 | 0.435 | 0.430 | 0.340 |
2.4 Binding free energy
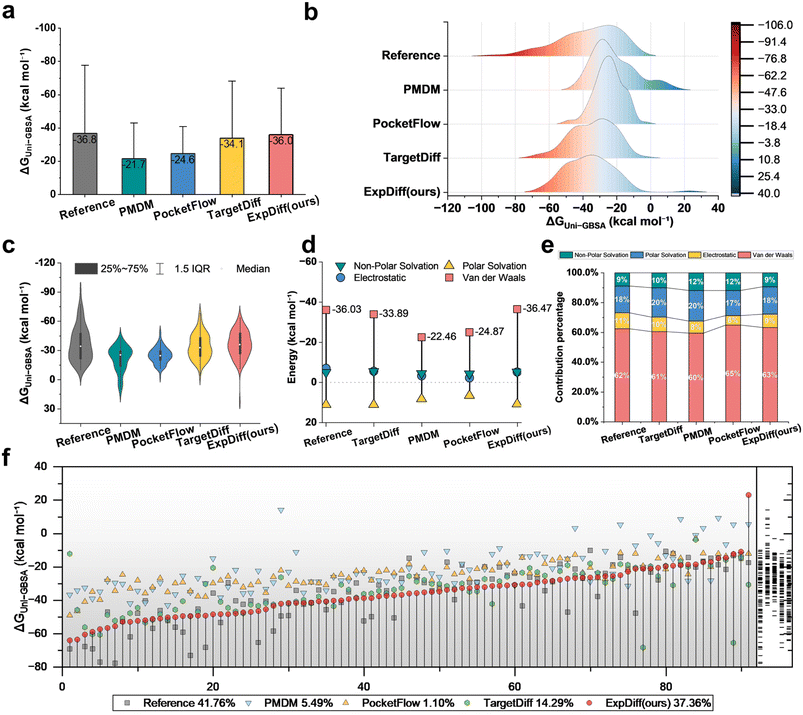
We first compare the overall binding free energy ΔG predictions of Reference, PMDM, PocketFlow, TargetDiff, and our proposed ExpDiff. As shown in Fig. 4a, the Reference method yields the most negative average ΔG (−36.8 kcal mol−1), whereas PMDM, PocketFlow, and TargetDiff systematically underestimate binding free affinity, producing values closer to zero. In contrast, ExpDiff achieves an average ΔG of approximately −36.0 kcal mol−1, closely matching Reference, indicating its ability to recover Reference-level binding free affinities. Density plots and violin plots (Fig. 4b and c) further demonstrate that ExpDiff reproduces the overall distribution shape of Reference while maintaining reduced tail deviations. The median and interquartile range are more concentrated, suggesting consistent and reliable predictions across diverse ligand–receptor pairs without generating numerous extreme outliers.Next, we examine the energy decomposition profiles to assess whether ExpDiff captures detailed physical contributions. Fig. 4d presents representative values for van der Waals, electrostatic, polar solvation, and nonpolar solvation terms. ExpDiff closely reproduces Reference values, particularly for van der Waals and electrostatic contributions, with total energies at roughly −36 kcal mol−1. Percentage decomposition (Fig. 4e) confirms that ExpDiff accurately estimates the relative contributions of different energy components, reducing systematic biases in van der Waals and solvation partitioning. Individual-case scatter plots (Fig. 4f) further highlight that ExpDiff (red dots) tracks Reference trends (gray squares) while minimizing large deviations across many systems. The bottom percentage labels indicate that ExpDiff achieves approximately 37.36% of cases where the generated molecules exhibit superior binding free energy, which is substantially higher than that of PMDM and PocketFlow and approaches the reference level, reflecting robust generalization across diverse receptor spaces.
Collectively, these results demonstrate that ExpDiff produces ligand structures that closely reproduce Reference binding free energies and energy component distributions. This ability to generate energetically favorable and physically consistent molecules is critical for large-scale virtual screening and rational molecular design. Owing to its superior performance in recovering mean ΔG, distribution consistency, and energy decomposition patterns, ExpDiff can serve as a high-confidence generator or be integrated as an auxiliary module in molecular design pipelines, enhancing the likelihood of identifying potent candidates while reducing the need for extensive experimental or computational validation.
2.5 Application scope
The size (area and volume) of the protein pocket binding site has a significant impact on the molecular binding capacity. To study the generalizability of ExpDiff, we analyzed different sizes of binding sites with docking results. Fig. 5a shows the area as well as the volume of different binding sites. Fig. 5b and c show the linear relationship between Vina score and Vina dock with the volume and area of the protein pocket. It can be seen that the area of the protein pocket has a strong correlation (−0.629) with Vina dock. Next, to exemplify the ability to generate molecules across various protein target areas and volumes, we categorized the size of the protein pockets into small, median, and large. The docking scores of ExpDiff and three baseline models were compared across protein pockets of different sizes. As can be seen from Fig. 5d and e, the molecules generated by ExpDiff all reflect the best docking results.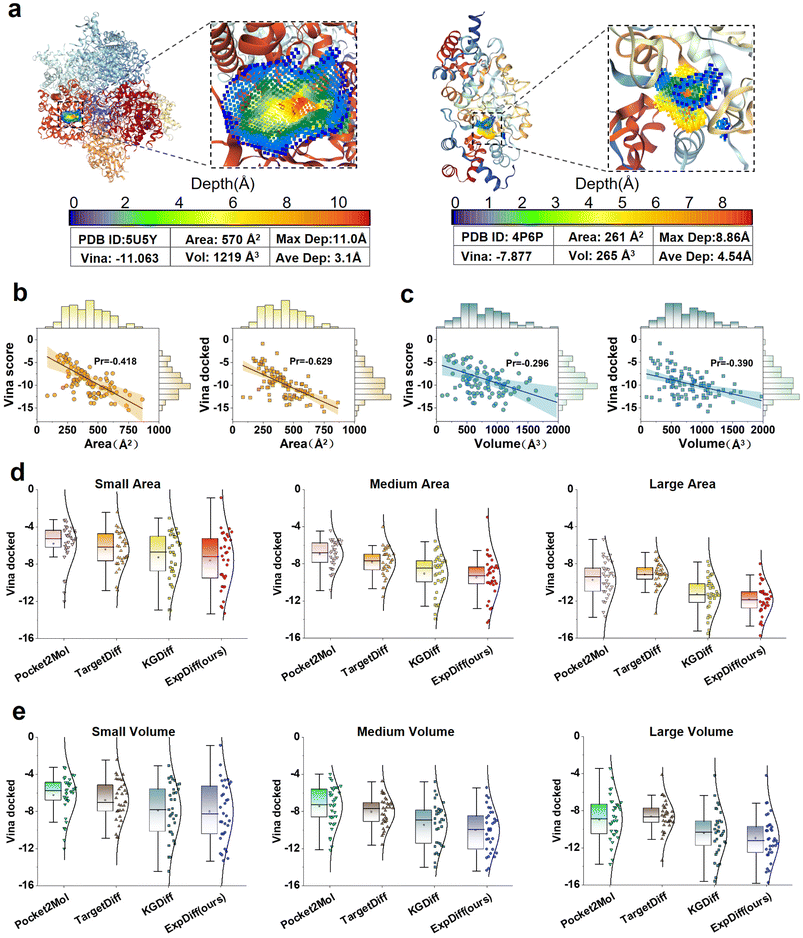 | ||
| Fig. 5 ExpDiff compared to the baseline model for varying docking pocket areas and volumes. (a) Larger and smaller protein pockets. (b) The relationship between the Vina score and Vina dock results and protein pocket area. (c) The relationship between the Vina score and Vina dock results and protein pocket volume. (d and e) Generated molecular binding affinity using Expdiff and baseline models at small, medium, and large protein pocket areas and volumes. The volumes and areas of the docking pockets were obtained from KVFinder.57 | ||
As an additional analysis, we tested four types of protein pockets to test the generalizability of ExpDiff across different target types, including nuclear receptor, protease, epigenetic regulation, and transporter. Except for 1AQ7 and 1B0U, it was found that for most targets, ExpDiff can generate molecules with the best docking results (see SI Table S4).
2.6 Case study
To evaluate ligand–pocket interactions, we conducted case studies by selecting representative generated molecules together with the corresponding test data. In Fig. 6, we first show the structure of the protein where the docking target is located, followed by visualization of the residues of the protein pockets and their interaction with the generated molecules. All three examples show good docking results with some of the residues of the binding site, surpassing those observed in the reference data. Non-covalent interactions such as hydrogen bonds and salt bridges were formed between the generated molecules and the binding site, thus enhancing the predicted binding affinity. This demonstrates that ExpDiff effectively learns the non-covalent relationships between protein pockets and molecules, a capability that empowers the model to design high-affinity drug candidates. In contrast to the binding interactions depicted in Fig. 6a and b, the 5W2G target in Fig. 6c exhibits the presence of a Mg2+ ion (represented by the red orb), which plays a critical role in mediating the molecular interactions. Remarkably, the molecules generated by ExpDiff maintain high binding affinity, underscoring the robustness of the model and its capability to design high-affinity ligands even in the presence of metal ion coordination. | ||
| Fig. 6 Visualization of generated molecules and reference docking results. (a–c) The docking results of target PDB ID: 3DAF, 1A2G, and 5W2G with molecules generated by ExpDiff and the interaction relationship with the target, respectively. 2D interaction diagrams of protein pockets with molecules were obtained using LigPlot+.58 | ||
3 Discussion
Diffusion models have shown promise in molecular generation; however, they still suffer from limitations such as inaccurate noise prediction, instability in feature distribution, and reduced reliability in molecular structure formation. To address these issues, the core objective of this study is to develop a dynamically adaptive alignment strategy that responds to changes in latent features across different diffusion steps, thereby improving the quality and affinity of the generated molecules. Molecular docking and binding free energy analyses demonstrate that our method substantially improves both binding affinity and biological relevance, achieving an average docking score of −10.06 and a binding free energy of −36.0 kcal mol−1, substantially outperforming existing molecular design models. By introducing the feature alignment mechanism of the expert network in multiple denoising steps, our method successfully enhances the stability of the model under high noise conditions and optimizes the pharmacological properties of the final generated molecules. The strategy enables fine-grained control over feature distribution during the generation process, resulting in generated molecules that exhibit superior performance in terms of structural rationality, chemical feasibility, and functional optimization.Although ExpDiff facilitates the design of molecules with stronger predicted interactions, the model still shares certain limitations inherent to fully automated drug discovery. While ExpDiff achieves competitive SA performance compared with baseline generative models, some generated structures still exhibit higher synthetic complexity than molecules in the training and reference sets. This highlights the need for future integration of synthesis-aware constraints or retrosynthetic guidance into the generative process. We have noticed that ExpDiff produces lower affinity molecules under a few protein pockets, which is related to the complexity of the target. Additionally, the molecules need to meet multiple pharmacological objectives. Future research can further extend the method in two directions. First, the generalization ability of the method can be verified in more complex drug design tasks, such as optimizing ADMET properties or improving the multi-target optimization ability of molecule generation, especially with respect to synthetic accessibility. Next, deeper generative theoretical modeling can be explored, such as incorporating physically guided diffusion processes or adaptive alignment strategies with strengthened learning control, to further improve the search efficiency and reliability of diffusion models in high-dimensional spaces.
Beyond small molecule design, the core principles of our alignment strategy offer transformative potential for biomolecular engineering. Building on protein language models (e.g., ESM28) and protein diffusion models (e.g., AlphaFold3 (ref. 29) and RFdiffusion30), future work will aim to extend the method proposed in this study to the protein domain by exploring how to utilize diffusion models and expert network alignment strategies for protein design and optimization. Collectively, this study lays an important foundation for future research on the controllability of generative modeling in the field of scientific computing and opens new avenues for further expansion to other biomolecular domains.
In summary, the ExpDiff model facilitates SBDD and demonstrates clear advantages in generating therapeutic candidates with enhanced binding potential. Conceptually, the expert alignment mechanism imposes external knowledge-driven guidance on the diffusion process, improving the controllability of molecular generation in high-dimensional latent spaces, which constitutes a long-term challenge inherent to traditional diffusion models. This represents a meaningful advancement toward controllable and interpretable molecular generation. This work explores a novel paradigm for SBDD through knowledge-guided diffusion modeling, offering a promising direction to accelerate the drug discovery pipeline, particularly in the generation of high-affinity therapeutic compounds.
4 Method
We begin by formally defining the structure-based molecular design task. An atom in three-dimensional space is represented as A = (x, z), where denotes the Euclidean coordinates and
denotes the Euclidean coordinates and  denotes the k-dimensional atomic features (e.g., atom type, charge, and hybridization). A protein pocket can subsequently be defined as
denotes the k-dimensional atomic features (e.g., atom type, charge, and hybridization). A protein pocket can subsequently be defined as  for i = 1,…,Np, and a molecule can be denoted as
for i = 1,…,Np, and a molecule can be denoted as  for i = 1,…,Nm, where Np and Nm denote the number of atoms in the protein pocket and molecule, respectively. The objective is to identify the molecule Bind = argmaxf(M, P) that maximizes the binding score with the given protein P, where M represents the set of all valid molecules in chemical space and f is a scoring function indicating the binding score between the molecule and the protein. In essence, the goal is to find the compound that optimizes the binding score and crucial drug-like characteristics for a particular protein.
for i = 1,…,Nm, where Np and Nm denote the number of atoms in the protein pocket and molecule, respectively. The objective is to identify the molecule Bind = argmaxf(M, P) that maximizes the binding score with the given protein P, where M represents the set of all valid molecules in chemical space and f is a scoring function indicating the binding score between the molecule and the protein. In essence, the goal is to find the compound that optimizes the binding score and crucial drug-like characteristics for a particular protein.
4.1 Diffusion model
 | (1) |
The denoising process, or reverse process, aims to reconstruct the original data from the noisy sample Mt by iteratively removing noise. This process is parameterized by a neural network and is described by
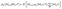 | (2) |
 are the denoising networks parameterized by the neural network for the purpose of predicting the denoising parameters for each diffusion step.
are the denoising networks parameterized by the neural network for the purpose of predicting the denoising parameters for each diffusion step.
Additionally, the model predicts the distribution of the added noise, as specified in eqn (3), by comparing the predicted noise  with the actual noise ϵθ introduced.
with the actual noise ϵθ introduced.
 | (3) |
By minimizing the difference between the predicted and actual noise, the model learns to effectively reverse the diffusion process and generate valid molecular structures.
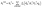 | (4) |
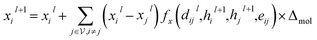 | (5) |
4.2 Alignment of the expert network
We introduced an expert network called DrugClip31 to enhance the quality of the molecular latent representation in the diffusion model. The choice of DrugClip is motivated by the suitability of its molecular encoder for our architecture, rather than an attempt to conduct a comparative evaluation among different encoders. Our goal is to guide molecular generation by aligning the intermediate molecular features generated by the diffusion model with the molecular output features of the expert model. It is worth noting that the molecular latent representation from the diffusion model is mixed with noise.As shown in Fig. 7, the first step in the alignment module involves applying a linear transformation to the latent tensor of the diffusion model ligand, mapping the input features to a higher-dimensional space. Subsequently, a Sigmoid activation function is applied to each element for non-linear transformation. This process transforms the input feature set into a new space, making it more suitable for computing attention scores. The computation is described by eqn (6), where zt represents the atomic latent features of the ligand after removing the noise at the tth step.
| attention_score = Sigmoid(Linear(zt)) × Wattention | (6) |
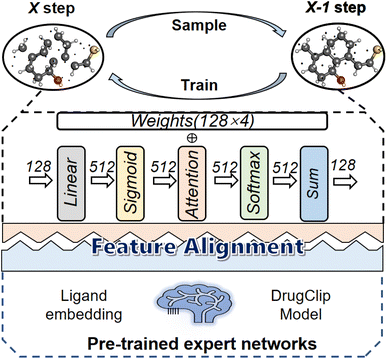 | ||
| Fig. 7 Molecular latent feature alignment module between the expert network and the diffusion model. | ||
To ensure that the attention scores form a valid probability distribution, the Softmax function is employed to normalize the attention scores. This normalization step transforms the scores into attention weights  which are positive and sum to one, thereby defining a probability distribution over the atomic features, as illustrated by
which are positive and sum to one, thereby defining a probability distribution over the atomic features, as illustrated by
| attention_weight = Softmax(attention_socre) | (7) |
The final step involves computing the weighted sum of the atomic features, where the input features are weighted using the attention weights. The attention weights are element-wise multiplied with the atomic features, followed by summation over all atoms (eqn (8)). The result is a global feature representation, which serves as the final output of the module.
 | (8) |
We align the molecular representations obtained from the expert network with those processed by the alignment module. Given the denoised molecular latent features at step t from the diffusion model and the output features, we aim to minimize the discrepancy between the outputs of the diffusion model and the expert model using cosine similarity, as shown in eqn (9).
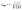 | (9) |
4.3 Feature alignment based on diffusion step awareness
As the diffusion model has different noise levels at each diffusion step, we need to use different alignment strategies at each step to adjust the impact of the expert network. In the early step (close to t = 0), the alignment with the expert network should be more stringent to ensure that the generated molecules are moving towards the target features due to the weaker noise and the molecules are more clearly characterized. While in the later step (close to t = 1000), approaching Gaussian noise, the alignment strategy of the expert model can be more relaxed at this point.We add a dynamic adjustment coefficient s(t), which functions similarly to a Sigmoid function and regulates the alignment strength at each step, to allow for flexible degree modification. In particular, s(t) has the following definition:
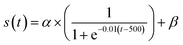 | (10) |
 | (11) |
During the training process, the adjusted alignment loss function is shown in eqn (11). This loss function allows the alignment strategy of the expert network to be dynamically adjusted to changes in the noise level at different stages of training, thus improving the quality of the generated molecules.
4.4 Optimization objectives
We introduced a module for predicting affinity and used the predicted affinity values for guided molecule generation during the inference process, a computational approach that is consistent with the KGDiff32 model. In eqn (12), ν represents the affinity value and the fully connected layer is used to predict the affinity value based on the atomic latent layer representation of the ligand in the diffusion model. The Shifted Softplus activation function is denoted as S. The total loss function is shown in eqn (13), and the loss scaling factors γ and δ are set to 100 and 1, respectively. ε denoted as the alignment loss weight between the expert network and the molecular vector of the diffusion model latent features mixed with noise. By optimizing this objective function, we can train the diffusion model so that the generated molecules not only conform to the distribution of the target data, but also align with the expert network in terms of atomic features, thus improving the quality of the generated molecules.| ν = Sigmod(W2 × S(W1x + b1) + b2) | (12) |
 | (13) |
4.5 Experimental set-up
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 990 complexes for training and 100 additional complexes as tests, and we used PyMol35 to draw protein as well as molecular diagrams.
990 complexes for training and 100 additional complexes as tests, and we used PyMol35 to draw protein as well as molecular diagrams.
The Affinity-related metrics include:
1. Vina score refers to the binding score between the generated molecular conformation and the protein pockets.
2. Vina minimize means the binding score between the optimized generated molecular conformation and the protein pockets.
3. Vina dock means the binding score between the generated molecules and the protein pockets after docking.
4. High affinity means the percentage of generated molecules with a vina score higher than or equal to that of the reference compound.
The Molecular property metrics include:
1.QED (Quantitative Estimate of Drug-likeness)52 evaluates molecular drug-likeness by reflecting the typical distribution of molecular properties in successful drug candidates.
2. SA (Synthetic Accessibility)53,54 evaluates the ease of molecular synthesis.
3. Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P55 evaluates the lipophilicity of the generated molecules.
P55 evaluates the lipophilicity of the generated molecules.
4. TPSA (Topological Polar Surface Area)56 evaluates the topological polar surface area of generated molecules.
5. JSD (Jensen–Shannon divergences) assesses the molecular structures in the empirical distributions of atomic and bond distances by comparing those of the generated molecules to those of the reference molecules.
Author contributions
H. L. was responsible for the design and development of the software, conducted the experiments, and contributed to manuscript editing. Z. W. and Y. Z. proposed the method for model design and made revisions to the manuscript. X. H. and W. H. assisted with the implementation and drawing of several figures. H. L. designed the overall architecture of the research.Conflicts of interest
No competing interest is declared.Data availability
All data and code supporting this research are publicly available. The CrossDocked 2020 dataset used for model development is accessible at https://doi.org/10.1021/acs.jcim.0c00411. Our implementation code, pre-trained models, and processed datasets are permanently archived on Zenodo at https://doi.org/10.5281/zenodo.18232614.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00440c.
Acknowledgements
This work was supported by the Natural Science Foundation of Shandong Province (ZR2023MF104) and the overseas joint training of PhD students from Ocean University of China.References
- O. A. Ntintas, T. Daglis and V. G. Gorgoulis, Harnessing deep learning to build optimized ligands, Nat. Comput. Sci., 2024, 4(11), 809–810, DOI:10.1038/s43588-024-00725-1.
- F. Soleymani, E. Paquet, H. L. Viktor and W. Michalowski, Structure-based protein and small molecule generation using EGNN and diffusion models: A comprehensive review, Computat. Struct. Biotec., 2024, 23, 2779–2797, DOI:10.1016/j.csbj.2024.06.021.
- S. Allenspach, J. A. Hiss and G. Schneider, Neural multi-task learning in drug design, Nat. Mach. Intell., 2024, 6(2), 124–137, DOI:10.1038/s42256-023-00785-4.
- F. Grisoni, Chemical language models for de novo drug design: Challenges and opportunities, Curr. Opin. Struct. Biol., 2023, 79, 102527, DOI:10.1016/j.sbi.2023.102527.
- A. Lavecchia, Navigating the frontier of drug-like chemical space with cutting-edge generative AI models, Drug Discov. Today, 2024, 29(9), 104133, DOI:10.1016/j.drudis.2024.104133.
- A. V. Sadybekov and V. Katritch, Computational approaches streamlining drug discovery, Nature, 2023, 616(7958), 673–685, DOI:10.1038/s41586-023-05905-z.
- M. McGibbon, S. Shave, J. Dong, Y. Gao, D. R. Houston, J. Xie, Y. Yang, P. Schwaller and V. Blay, From intuition to AI: evolution of small molecule representations in drug discovery, Brief. Bioinform., 2023, 25(1), 422, DOI:10.1093/bib/bbad422.
- I. J. Goodfellow, J. Pouget-Abadie, M. Mirza, B. Xu, D. Warde-Farley, S. Ozair, A. Courville and Y. Bengio, Generative adversarial nets, in Proceedings of the 28th International Conference on Neural Information Processing Systems - Volume 2. NIPS’14, MIT Press, Cambridge, MA, USA, 2014, vol. 2, pp. 2672–2680 Search PubMed.
- D. P. Kingma and M. Welling, Auto-Encoding Variational Bayes. arXiv, 2022, DOI:10.48550/arXiv.1312.6114.
- J. Ho, A. Jain and P. Abbeel, Denoising diffusion probabilistic models, in Proceedings of the 34th International Conference on Neural Information Processing Systems. NIPS ’20, Curran Associates Inc., Red Hook, NY, USA, 2020, pp. 6840–6851 Search PubMed.
- T. Ochiai, T. Inukai, M. Akiyama, K. Furui, M. Ohue, N. Matsumori, S. Inuki, M. Uesugi, T. Sunazuka, K. Kikuchi, H. Kakeya and Y. Sakakibara, Variational autoencoder-based chemical latent space for large molecular structures with 3D complexity, Commun. Chem., 2023, 6(1), 249, DOI:10.1038/s42004-023-01054-6.
- H. Y. I. Lam, R. Pincket, H. Han, X. E. Ong, Z. Wang, J. Hinks, Y. Wei, W. Li, L. Zheng and Y. Mu, Application of variational graph encoders as an effective generalist algorithm in computer-aided drug design, Nat. Mach. Intell., 2023, 5(7), 754–764, DOI:10.1038/s42256-023-00683-9.
- C. Hu, S. Li, C. Yang, J. Chen, Y. Xiong, G. Fan, H. Liu and L. Hong, ScaffoldGVAE: scaffold generation and hopping of drug molecules via a variational autoencoder based on multi-view graph neural networks, J. Cheminform., 2023, 15(1), 91, DOI:10.1186/s13321-023-00766-0.
- S. R. Atance, J. V. Diez, O. Engkvist, S. Olsson and R. Mercado, De Novo Drug Design Using Reinforcement Learning with Graph-Based Deep Generative Models, J. Chem. Inf. Model., 2022, 62(20), 4863–4872, DOI:10.1021/acs.jcim.2c00838.
- W. Zhung, H. Kim and W. Y. Kim, 3D molecular generative framework for interaction-guided drug design, Nat. Commun., 2024, 15(1), 2688, DOI:10.1038/s41467-024-47011-2.
- L. Wang, S. Wang, H. Yang, S. Li, X. Wang, Y. Zhou, S. Tian, L. Liu and F. Bai, Conformational Space Profiling Enhances Generic Molecular Representation for AI-Powered Ligand-Based Drug Discovery, Adv. Sci, 2024, 11(40), 2403998, DOI:10.1002/advs.202403998.
- S. Chen, O. Zhang, C. Jiang, H. Zhao, X. Zhang, M. Chen, Y. Liu, Q. Su, Z. Wu, X. Wang, W. Qu, Y. Ye, X. Chai, N. Wang, T. Wang, Y. An, G. Wu, Q. Yang, J. Chen, W. Xie, H. Lin, D. Li, C.-Y. Hsieh, Y. Huang, Y. Kang, T. Hou and P. Pan, Deep lead optimization enveloped in protein pocket and its application in designing potent and selective ligands targeting LTK protein, Nat Mach Intell, 2025, 7(3), 448–458, DOI:10.1038/s42256-025-00997-w.
- P. Wu, H. Du, Y. Yan, T.-Y. Lee, C. Bai and S. Wu, Guided diffusion for molecular generation with interaction prompt, Briefings Bioinf., 2024, 25(3), 174, DOI:10.1093/bib/bbae174 https://academic.oup.com/bib/article-pdf/25/3/bbae174/57295476/bbae174.pdf .
- M. Sako, N. Yasuo and M. Sekijima, DiffInt: A Diffusion Model for Structure-Based Drug Design with Explicit Hydrogen Bond Interaction Guidance, J. Chem. Inf. Model., 2025, 65(1), 71–82, DOI:10.1021/acs.jcim.4c01385.
- A. Q. Nichol and P. Dhariwal, Improved Denoising Diffusion Probabilistic Models, in Proceedings of the 38th International Conference on Machine Learning. PMLR ’21, 2021, pp. 8162–8171, ISSN: 2640-3498. https://proceedings.mlr.press/v139/nichol21a.html Search PubMed.
- Z. Huang, L. Yang, X. Zhou, C. Qin, Y. Yu, X. Zheng, Z. Zhou, W. Zhang, Y. Wang and W. Yang, Interaction-based Retrieval-augmented Diffusion Models for Protein-specific 3D Molecule Generation, 2024, https://openreview.net/forum?id=eejhD9FCP3.
- Z. Huang, L. Yang, Z. Zhang, X. Zhou, Y. Bao, X. Zheng, Y. Yang, Y. Wang and W. Yang, Binding-Adaptive Diffusion Models for Structure-Based Drug Design. arXiv. Version Number: 1, 2024, DOI:10.48550/ARXIV.2402.18583.
- X. Zhou, X. Cheng, Y. Yang, Y. Bao, L. Wang and Q. Gu, DecompOpt: Controllable and Decomposed Diffusion Models for Structure-based Molecular Optimization, arXiv. Version Number: 1, 2024, DOI:10.48550/ARXIV.2403.13829.
- W. Zhung, H. Kim and W. Y. Kim, 3D molecular generative framework for interaction-guided drug design, Nat. Commun., 2024, 15, 2688, DOI:10.1038/s41467-024-47011-2.
- L. Huang, T. Xu, Y. Yu, P. Zhao, X. Chen, J. Han, Z. Xie, H. Li, W. Zhong, K.-C. Wong and H. Zhang, A dual diffusion model enables 3D molecule generation and lead optimization based on target pockets, Nat. Commun., 2024, 15(1), 2657, DOI:10.1038/s41467-024-46569-1.
- Y. Diao, D. Liu, H. Ge, R. Zhang, K. Jiang, R. Bao, X. Zhu, H. Bi, W. Liao, Z. Chen, K. Zhang, R. Wang, L. Zhu, Z. Zhao, Q. Hu and H. Li, Macrocyclization of linear molecules by deep learning to facilitate macrocyclic drug candidates discovery, Nat. Commun., 2023, 14(1), 4552, DOI:10.1038/s41467-023-40219-8.
- P. J. Salveson, A. P. Moyer, M. Y. Said, G. Gokçe, X. Li, A. Kang, H. Nguyen, A. K. Bera, P. M. Levine, G. Bhardwaj and D. Baker, Expansive discovery of chemically diverse structured macrocyclic oligoamides, Science, 2024, 384(6694), 420–428, DOI:10.1126/science.adk1687.
- T. Hayes, R. Rao, H. Akin, N. J. Sofroniew, D. Oktay, Z. Lin, R. Verkuil, V. Q. Tran, J. Deaton, M. Wiggert, R. Badkundri, I. Shafkat, J. Gong, A. Derry, R. S. Molina, N. Thomas, Y. A. Khan, C. Mishra, C. Kim, L. J. Bartie, M. Nemeth, P. D. Hsu, T. Sercu, S. Candido and A. Rives, Simulating 500 million years of evolution with a language model, Science, 2025, 387(6736), 850–858, DOI:10.1126/science.ads0018.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O'Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulyte, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Accurate structure prediction of biomolecular interactions with AlphaFold 3, Nature, 2024, 630(8016), 493–500, DOI:10.1038/s41586-024-07487-w.
- K. E. Wu, K. K. Yang, R. Van Den Berg, S. Alamdari, J. Y. Zou, A. X. Lu and A. P. Amini, Protein structure generation via folding diffusion, Nat. Commun., 2024, 15(1), 1059, DOI:10.1038/s41467-024-45051-2.
- B. Gao, B. Qiang, H. Tan, Y. Jia, M. Ren, M. Lu, J. Liu, W.-Y. Ma and Y. Lan, Drugclip: Contrasive protein-molecule representation learning for virtual screening. in, NeurIPS 2023, 2023, https://openreview.net/forum?id=lAbCgNcxm7 Search PubMed.
- H. Qian, W. Huang, S. Tu and L. Xu, KGDiff: towards explainable target-aware molecule generation with knowledge guidance, Brief. Bioinform., 2023, 25(1), 435, DOI:10.1093/bib/bbad435.
- P. G. Francoeur, T. Masuda, J. Sunseri, A. Jia, R. B. Iovanisci, I. Snyder and D. R. Koes, Three-Dimensional Convolutional Neural Networks and a Cross-Docked Data Set for Structure-Based Drug Design, J. Chem. Inf. Model., 2020, 60(9), 4200–4215, DOI:10.1021/acs.jcim.0c00411.
- R. Wang, X. Fang, Y. Lu and S. Wang, The PDBbind Database: Collection of Binding Affinities for Protein Ligand Complexes with Known Three Dimensional Structures, J. Med. Chem., 2004, 47(12), 2977–2980, DOI:10.1021/jm030580l.
- PyMOL | pymol.org. https://www.pymol.org/.
- O. Zhang, J. Zhang, J. Jin, X. Zhang, R. Hu, C. Shen, H. Cao, H. Du, Y. Kang, Y. Deng, F. Liu, G. Chen, C.-Y. Hsieh and T. Hou, ResGen is a pocket-aware 3D molecular generation model based on parallel multiscale modelling, Nat. Mach. Intell., 2023, 5(9), 1020–1030, DOI:10.1038/s42256-023-00712-7.
- S. Luo, J. Guan, J. Ma and J. Peng, A 3D Generative Model for Structure-Based Drug Design, in Advances in Neural Information Processing Systems, 2021, vol. 34, pp. 6229–6239, https://papers.nips.cc/paper/2021/hash/314450613369e0ee72d0da7f6fee773c-Abstract.html Search PubMed.
- Y. Jiang, G. Zhang, J. You, H. Zhang, R. Yao, H. Xie, L. Zhang, Z. Xia, M. Dai, Y. Wu, L. Li and S. Yang, PocketFlow is a data-and-knowledge-driven structure-based molecular generative model, Nat. Mach. Intell., 2024, 6, 326–337, DOI:10.1038/s42256-024-00808-8.
- X. Peng, S. Luo, J. Guan, Q. Xie, J. Peng and J. Ma, Pocket2Mol: Efficient Molecular Sampling Based on 3D Protein Pockets, in Proceedings of the 39th International Conference on Machine Learning. PMLR ’22, 2022, pp. 17644–17655 Search PubMed.
- V. Dorna, D. Subhalingam, K. Kolluru, S. Tuli, M. Singh, S. Singal, N. M. A. Krishnan and S. Ranu, TAGMol: Target-Aware Gradient-guided Molecule Generation, arXiv, 2024, DOI:10.48550/ARXIV.2406.01650.
- X. Li, P. Wang, T. Fu, W. Gao, C. Li, L. Shi and J. Liu, AUTODIFF: Autoregressive Diffusion Modeling for Structure-based Drug Design, arXiv. Version Number: 2, 2024, DOI:10.48550/ARXIV.2404.02003.
- J. Guan, W. W. Qian, X. Peng, Y. Su, J. Peng and J. Ma, 3D Equivariant Diffusion for Target-Aware Molecule Generation and Affinity Prediction, arXiv. arXiv:2303.03543, 2023, http://arxiv.org/abs/2303.03543 Search PubMed.
- J. Guan, X. Zhou, Y. Yang, Y. Bao, J. Peng, J. Ma, Q. Liu, L. Wang and Q. Gu, DecompDiff: Diffusion Models with Decomposed Priors for Structure-Based Drug Design, arXiv. Version Number: 1, 2024, DOI:10.48550/ARXIV.2403.07902.
- Z. Huang, L. Yang, X. Zhou, Z. Zhang, W. Zhang, X. Zheng, J. Chen, Y. Wang, B. Cui and W. Yang, Protein-Ligand Interaction Prior for Binding-aware 3D Molecule Diffusion Models, 2023, https://openreview.net/forum?id=qH9nrMNTIW.
- Z. Zhang, M. Wang and Q. Liu, FlexSBDD: Structure-Based Drug Design with Flexible Protein Modeling, arXiv, 2024, DOI:10.48550/ARXIV.2409.19645.
- A. K. Yalabadi, M. Yazdani-Jahromi and O. O. Garibay, BoKDiff: Best-of-K Diffusion Alignment for Target-Specific 3D Molecule Generation, arXiv, 2025, DOI:10.48550/ARXIV.2501.15631.
- S. Gu, M. Xu, A. Powers, W. Nie, T. Geffner, K. Kreis, J. Leskovec, A. Vahdat and S. Ermon, Aligning Target-Aware Molecule Diffusion Models with Exact Energy Optimization, arXiv, 2024, DOI:10.48550/ARXIV.2407.01648.
- W. Zhung, H. Kim and W. Y. Kim, 3D molecular generative framework for interaction-guided drug design, Nat. Commun., 2024, 15(1), 2688, DOI:10.1038/s41467-024-47011-2.
- J. Eberhardt, D. Santos-Martins, A. F. Tillack and S. Forli, AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings, J. Chem. Inf. Model., 2021, 61(8), 3891–3898, DOI:10.1021/acs.jcim.1c00203.
- M. Yang, Z. Bo, T. Xu, B. Xu, D. Wang and H. Zheng, Uni-GBSA: an open-source and web-based automatic workflow to perform MM/GB(PB)SA calculations for virtual screening, Brief. Bioinform., 2023, 24(4), 218, DOI:10.1093/bib/bbad218.
- G. Landrum, P. Tosco, B. Kelley, D. Ric, D. CosgroveG. Sriniker, R. Vianello, E. NadineSchneider, E. Kawashima, G. Jones, N. D., A. Dalke, B. Cole, M. Swain, S. Turk, A. AlexanderSavelyev, A. Vaucher, M. Wójcikowski, I. Take, V. F. Scalfani, D. Probst, K. Ujihara, g. godin, A. Pahl, R. Walker, J. Lehtivarjo, F. Berenger, jasondbiggs and strets123, rdkit/rdkit: 2023_09_4 (Q3 2023) Release, Zenodo, 2024, DOI:10.5281/zenodo.10460537.
- G. R. Bickerton, G. V. Paolini, J. Besnard, S. Muresan and A. L. Hopkins, Quantifying the chemical beauty of drugs, Nature Chem, 2012, 4(2), 90–98, DOI:10.1038/nchem.1243.
- P. Ertl and A. Schuffenhauer, Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions, J. Cheminf., 2009, 1(1), 8, DOI:10.1186/1758-2946-1-8.
- J. You, B. Liu, R. Ying, V. Pande and J. Leskovec, Graph Convolutional Policy Network for Goal-Directed Molecular Graph Generation, arXiv, 2019, DOI:10.48550/arXiv.1806.02473.
- M. H. Abraham, H. S. Chadha, R. A. E. Leitao, R. C. Mitchell, W. J. Lambert, R. Kaliszan, A. Nasal and P. Haber, Determination of solute lipophilicity, as log P(octanol) and log P(alkane) using poly(styrene–divinylbenzene) and immobilised artificial membrane stationary phases in reversed-phase high-performance liquid chromatography, J. Chromatogr. A, 1997, 766(1–2), 35–47, DOI:10.1016/S0021-9673(96)00977-6.
- H. Zhong, V. Mashinson, T. Woolman and M. Zha, Understanding the Molecular Properties and Metabolism of Top Prescribed Drugs, Curr. Top. Med. Chem., 2013, 13(11), 1290–1307, DOI:10.2174/15680266113139990034.
- J. V. S. Guerra, H. V. Ribeiro-Filho, J. G. C. Pereira and P. S. Lopes-de-Oliveira, KVFinder-web: a web-based application for detecting and characterizing biomolecular cavities, Nucleic Acids Res., 2023, 51(W1), 289–297, DOI:10.1093/nar/gkad324.
- R. A. Laskowski and M. B. Swindells, LigPlot+: Multiple Ligand–Protein Interaction Diagrams for Drug Discovery, J. Chem. Inf. Model., 2011, 51(10), 2778–2786, DOI:10.1021/ci200227u.
| This journal is © The Royal Society of Chemistry 2026 |