Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Deep set model for the automated NMR fingerprinting of unknown mixtures
Jens
Wagner
 ,
Kerstin
Münnemann
,
Kerstin
Münnemann
 ,
Thomas
Specht
,
Thomas
Specht
 ,
Hans
Hasse
,
Hans
Hasse
 and
Fabian
Jirasek
and
Fabian
Jirasek
 *
*
Laboratory of Engineering Thermodynamics (LTD), RPTU University Kaiserslautern-Landau, Germany. E-mail: fabian.jirasek@rptu.de; Tel: +49 (0)631 205 4685
First published on 20th February 2026
Abstract
Elucidating unknown mixtures is a critical challenge in chemistry and chemical engineering. Nuclear magnetic resonance (NMR) spectroscopy is a powerful analytical technique generally suited for this purpose. However, component-wise elucidation with NMR is tedious for complex mixtures, requires expert knowledge, and often yields ambiguous results. In contrast, identifying and quantifying structural groups in a mixture from NMR spectra is much more straightforward. In prior work, we have introduced ‘NMR fingerprinting’ for the automated elucidation of carbon-, hydrogen-, and oxygen-containing structural groups in unknown mixtures based on standard NMR experiments and a support vector classification (SVC) from machine learning (ML). In the present work, we present a substantially advanced NMR fingerprinting method that employs a deep set model (DSM), addressing major shortcomings of the SVC, and integrates additional information from 2D NMR experiments. The DSM was trained on experimental NMR spectra of pure components from open-source databases, augmented with synthetic spectral data, and comprises invariant and equivariant network structures to ensure predictions independent of the input order of the NMR signals. Tested on experimental pure-component test data, the DSM performs excellently, significantly outperforming our previous approaches. Furthermore, we demonstrate the applicability of the DSM to unknown mixtures by predicting the structural groups from NMR spectra of test mixtures measured using a benchtop NMR spectrometer. The predictions agree very well with the true mixture compositions, highlighting the method's potential for efficient automated mixture analysis and providing a reliable basis for downstream tasks, such as thermodynamic modeling using group-contribution methods.
Introduction
Complex mixtures of unknown compositions containing unknown components are ubiquitous in chemistry and chemical engineering, constituting a stiff challenge to process design and optimization. Nuclear magnetic resonance (NMR) spectroscopy is a powerful analytical technique well-suited for component elucidation, particularly if used with computer-assisted structure elucidation (CASE) programs,1 which follow a set of predefined rules and incorporate predicted spectra to propose possible molecular structures. However, CASE programs often require some prior knowledge of the molecular formula of the component to be identified, typically obtained by high-resolution mass spectrometry,2–5 adding experimental complexity to their application. Recently, machine-learning (ML) approaches that rely solely on NMR information6–8 or incorporate additional spectroscopic input9,10 have emerged as promising alternatives. Nonetheless, all these methods are restricted to identifying pure components, severely limiting their applicability in chemical engineering practice, where mixtures are usually present.NMR spectroscopy has also successfully been applied for the qualitative and quantitative analysis of mixtures.11–16 If the mixture components are known, a variety of automated quantification methods are available, even if signals in the NMR spectra overlap.17–23 However, elucidating unknown components in mixtures remains a significant challenge, whose solution often depends on expert knowledge, which becomes infeasible if complex mixtures are studied. In cases where mixtures contain unknown components with signal overlap, already the first step of separating the relevant signals, the so-called deconvolution of the NMR spectrum, becomes inherently ambiguous, though some ML approaches for automated deconvolution of NMR spectra have been introduced.24–26
An alternative approach to elucidating components in complex mixtures that avoids the ambiguities of assigning the signals to the unknown components is dereplication,27–32 which identifies individual components by comparing the NMR spectrum of the mixtures to those of pure compounds retrieved from reference databases. However, the limited coverage of these databases confines dereplication to those molecules already represented within them.33 Moreover, methods relying solely on spectral comparisons remain sensitive to experimental conditions due to inherent biases in the reference data.4 Consequently, no broadly applicable solution currently exists for the automated elucidation of unknown components in mixtures by NMR spectroscopy.
While, for the reasons discussed above, elucidating components in unknown mixtures by NMR spectroscopy still poses a significant challenge, identifying the structural groups that constitute these components is considerably more straightforward. This group-based task, which we call ‘NMR fingerprinting’, is based on the fact that in an NMR spectrum, the chemical shift of an analyzed nucleus reflects its electronic environment, thereby revealing the structural group containing it. Traditionally, chemical shift tables that outline characteristic ranges in NMR spectra have been used to assign structural groups to NMR signals.34 However, overlapping characteristic ranges in chemical shift tables lead to ambiguity in assigning structural groups based solely on them. Also, the “static” nature of these tables leads to problems in practice.
From an ML perspective, assigning the correct structural group to signals in an NMR spectrum represents a classification problem. Therefore, we have recently developed a support vector classification (SVC) for the automated NMR fingerprinting of carbon-, hydrogen-, and oxygen-containing structural groups in unknown mixtures based on standard NMR experiments.35,36 Trained on thousands of pure-component spectra from the open-source databases Biological Magnetic Resonance Data Bank (BMRB)37 and NMRShiftDB,38 the SVC automatically assigns structural groups to signals in 13C NMR spectra, leveraging additional information from 1H and 13C DEPT (distortionless enhancement by polarization transfer) NMR spectroscopy. Utilizing SMARTS39 strings as a machine-readable representation of the respective structural groups during model training enables straightforward modification and extension of the considered structural group list. Applied to test mixtures, the predictions by the SVC achieved good agreement with the true mixture compositions, making it a reliable method for the structural group elucidation of unknown mixtures. The results of NMR fingerprinting can subsequently be used for the rational definition of pseudo-components40 and thermodynamic modeling using group-contribution methods,41–45 enabling the conceptual design of fluid separation processes.46,47
However, due to the characteristics of NMR data, SVC-based NMR fingerprinting has significant limitations in its application. Specifically, the signals in the NMR spectrum classified into structural groups can vary substantially in number, depending on the complexity and number of different components in the mixture of interest. This poses a challenge for developing SVCs for NMR fingerprinting, as an SVC requires inputs of constant length, which we have solved by binning the NMR spectra in multiple regions of defined chemical shift width. However, binning leads to the problem that signals with very similar chemical shifts, which in consequence are assigned to the same bin, cannot be distinguished, leading to classification errors. Furthermore, while there are natural choices for ordering the NMR signals in the input of the ML models, particularly with increasing chemical shift, it is not guaranteed that all data sets consistently comply with this ordering. Similarly, there is no inherent physical order of the different NMR spectra, e.g., 1H, 13C, 13C DEPT. Since SVCs are not permutation-invariant, i.e., their results depend on the input order, this poses another source of error for the NMR fingerprinting.
Within the realm of ML, these properties suggest that NMR signals and their corresponding nuclei information are best modeled as elements of sets rather than as fixed-length data instances.48
In this work, we overcome these limitations by developing a classification model based on a deep-set architecture.49 Deep set models (DSM) are a specialized neural network (NN) class within the field of geometric deep learning,50 specifically designed to preserve the symmetries inherent in set-structured data while introducing only minimal additional model complexity.48 Our DSM incorporates both invariant and equivariant network structures, ensuring that predictions are independent of input size and permutation, allowing the model to efficiently handle the unordered and variable-sized nature of NMR signals and nuclei. To fully capture the set-based characteristics of the NMR data, we extend our approach by incorporating information on the carbon–hydrogen correlations from 1H –13C HSQC (heteronuclear single quantum coherence) NMR spectroscopy as the first 2D NMR experiment in our mixture analysis. In doing so, the HSQC information is not directly used as additional input to the DSM but instead serves to construct the set structure of the model input by linking the information gathered from the 1H and 13C NMR experiments.
Additionally, we address the challenge of limited and incomplete training data in the used open-source NMR databases by augmenting incomplete NMR spectra with information derived from magnetically identical nuclei and predicted spectra using the open-source tools RDkit51 and NMRium.52 In this way, we have obtained complete spectral information for 2767 pure components, which we have used to train the model and rigorously test its predictive performance exclusively on unseen experimental NMR spectra. Finally, we have applied the model to test mixtures whose spectra were measured using a 60 MHz benchtop NMR device, demonstrating the approach in practical low-field NMR applications.
Methods
Overview
Fig. 1 provides an overview of the NMR fingerprinting method developed in this work to predict the structural groups and assign them to signals in the 13C NMR spectra of unknown samples using additional information from 1H, 13C DEPT, and 1H –13C HSQC NMR experiments. Central to our method is the DSM, which integrates invariant and equivariant network architectures to ensure predictions independent of the input order of the NMR signals and their associated nuclei information. The DSM was trained on the NMR spectra of 2767 pure components taken from the open-source databases BMRB37 and NMRShiftDB.38 To address the issue of incomplete spectral data within these databases, we employed augmentation techniques that utilize information from magnetically equivalent nuclei identified via RDKit51 and synthetic spectra predicted using NMRium.52 The details on the individual steps of our NMR fingerprinting are explained in the following subsections. | ||
| Fig. 1 Overview on the NMR fingerprinting method based on a deep set model (DSM)49 architecture for predicting the structural groups in unknown mixtures from NMR spectra and assigning them to signals in the 13C NMR spectrum. The DSM was trained on pure-component NMR spectra from the open-source databases BMRB37 and NMRShiftDB,38 with missing information augmented from magnetically identical nuclei and predicted spectra using the open-source tools RDkit51 and NMRium.52 The architecture of the DSM and its input, which is obtained from NMR experiments, are described in Section Deep-set architecture. | ||
Currently, the NMR fingerprinting method distinguishes 13 structural main groups. The method can also distinguish between different substitution degrees, so that, in total, 30 different subgroups can be identified, which are the same as in our previous works36,44 and summarized in Table 1. The quantification of the identified structural groups is finally achieved through signal integration in the 13C NMR spectra.36,44
| Label | Structural group | SMARTS representation |
|---|---|---|
| CH3 | Methyl | [CX4;D1;!$(C[!#6])] |
| CHx | Alkyl; x ∈ {0, 1, 2} | [CX4;D2,D3,D4;!$(C[!#6]);!R] |
| CHcyx | Cyclic alkyl; x ∈ {0, 1, 2} | [CX4;!$(C[!#6]);R] |
| CHxOH | Alcohol; x ∈ {0, 1, 2, 3} | [CX4;!$(C[OX2H0][CX3H1,CX3](![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O))][OX2H] O))][OX2H] |
| CHxO | Ether; x ∈ {0, 1, 2, 3} | [CX4;$(C[OD2]);!$(C[OX2H0][CX3H1,CX3](![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O));!$(C[OX2H])] O));!$(C[OX2H])] |
CHx![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) |
Aliphatic double bond; x ∈ {0, 1, 2} | [CX3;!$(C∼[!#6])] |
| CHxar | Aromatic carbon; x ∈ {0, 1} | [cX3;!$(c∼[!#6])] |
| RO-CHxar | Aromatic carbon with oxygen substituent; x ∈ {0, 1} | [cX3;!$(c![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O);$(c∼[#8X2])] O);$(c∼[#8X2])] |
| COOR | Ester/lactone/anhydride carbonyl | [CX3H1,#6X3](![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O)[#8X2H0] O)[#8X2H0] |
| ROOCHx | Alkyl next to ester/lactone oxygen; x ∈ {0, 1, 2, 3} | [CX4;$(C[OX2H0;$(O(C(![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O)))])] O)))])] |
| COOH | Carboxylic acid | [CX3](![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O)[OX2H1] O)[OX2H1] |
| COald | Aldehyde | [CX3H1;!$(C[!#6])](=O) |
| COket | Ketone | [#6X3H0;!$([#6][!#6])](![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) O) |
Deep-set architecture
The input of the DSM consists of a set of xi, where i denotes one of the N signals in the 13C NMR spectrum of the studied sample, cf.Fig. 1. Each xi contains NMR-spectroscopic information on the respective 13C nucleus associated with that signal (x13Ci) and on the 1H nuclei directly bonded to it (x1Hi). Specifically, x13Ci contains the respective chemical shift δ13Ci determined from the 13C NMR spectrum and the substitution degree Si derived from the intensities in the 13C DEPT 90/135 NMR spectra.36,53x1Hi comprises the chemical shifts δ1Hi of the j … Si1H nuclei directly bonded to the respective 13C nucleus, obtained from the 1H NMR spectrum of the studied sample. The 1H nuclei are thereby assigned to their corresponding 13C nuclei by the cross-signals observed in the 1H –13C HSQC spectrum. Additionally, the DSM uses the boolean input L indicating the presence of labile protons in the sample, which is also obtained from the 1H –13C HSQC spectrum.36The DSM developed in this work combines invariant and equivariant network structures. In the first step, the input information x13Ci for the 13C nuclei and x1Hi for the 1H nuclei is independently processed by dedicated embedding networks ϕ13C and ϕ1H, respectively. Unlike classical neural networks, these embeddings are computed in parallel rather than jointly,49 ensuring that the set-based nature of the nuclei data is respected. Subsequently, the nuclei embeddings are aggregated using the summation as a permutation-invariant function α, leading to the intermediate prediction α(xi) for each structural group based only on NMR-spectroscopic information on the respective nuclei. By employing parallel embeddings and the permutation-invariant function α, the DSM ensures an invariant prediction independent of the input order of the nuclei information.
In the second step, the intermediate predictions α(xi) are refined within the context of all structural groups in the studied sample to account for mutual influences on their respective NMR signals. Therefore, the intermediate predictions α(xi) for each signal in the 13C NMR spectrum are processed in parallel by the main embedding network ϕ, directing them to the equivariant layer σ(α). The equivariant layer σ(α) is a specialized NN layer that combines a standard per-element feed-forward layer σ with summation-based aggregation α,54 allowing the interaction of the embedded structural group predictions α(xi) while maintaining the relation between input and output.48 This summation-based aggregation captures inter-signal relationships in the context of all signals, which is the simplest form of contextualization and does not explicitly encode pairwise interactions between individual signals, as employed, for example, in self-attention-based architectures.48 Finally, through parallel processing by the prediction network ρ, which uses the additional input regarding the presence of labile protons L, the prediction ρ(xi) for each 13C NMR signal is obtained, independent of the input order of the signals.
The DSM does not provide absolute predictions for structural groups; instead, it assigns a probability to each group in Table 1 for every 13C NMR signal, with many groups receiving a probability of zero. This probability is interpreted as the model's confidence in the corresponding group assignment. The structural group with the highest probability (i.e., highest model confidence) is selected as the absolute prediction.
Augmentation of pure-component NMR data
Collecting and processing pure-component NMR data from BMRB and NMRShiftDB was conducted analogous to our previous work.36 Only pure components for which the following conditions are fulfilled were considered: composed exclusively of carbon, hydrogen, and oxygen; can be unambiguously segmented into the structural groups presented in Table 1; and for which both an experimental 13C and 1H NMR spectrum are available.However, some of the NMR spectra from these databases are incomplete, lacking assignments of chemical shifts to the respective nuclei. Upon closer examination, these omissions generally fall into two categories. Sometimes, only one of multiple magnetically equivalent nuclei has an assigned chemical shift. This partial assignment is likely attributable to non-standardized data structures within the databases, which manage redundant information inconsistently. In other cases, none of the magnetically equivalent nuclei have assigned chemical shifts, suggesting that the missing assignment is probably the result of human error during NMR spectra recording or evaluation. Since the DSM classifies 13C signals in the context of all structural groups in the sample rather than individually, it is essential to provide complete spectral information of the components as input. To address the issue of missing spectral data, we implemented a two-step augmentation process:
(1) Missing chemical shifts were supplemented by automatically identifying magnetically equivalent nuclei within each component using RDKit and adopting their corresponding spectral information from magnetically equivalent nuclei for which information was available.
(2) Any remaining gaps in the spectra were filled using data from synthetic spectra predicted for each pure component with NMRium. In this step, no completely synthetic spectra were used; only existing but incomplete experimental spectra were augmented.
Through these augmentation steps, the number of pure components with complete spectral information increased from 839 to 2767, substantially extending the data set available for model training. Additional details on the data augmentation from synthetic spectra are provided in the SI.
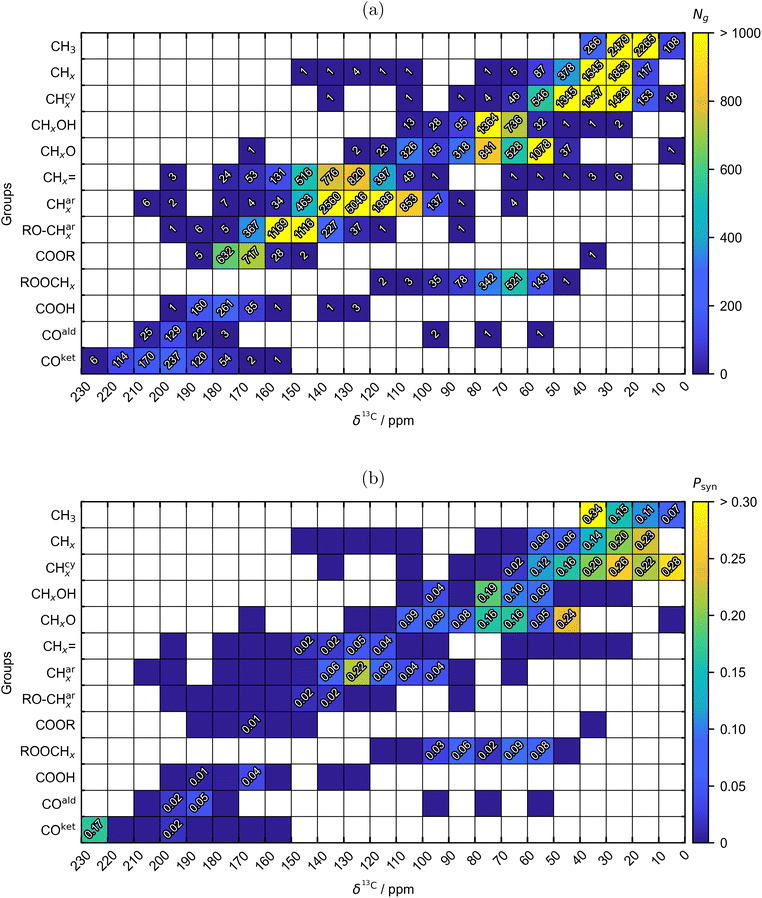
In Fig. 2, the final augmented data set covering 2767 pure components and consisting of a total of 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 838 structural groups is visualized considering the information from the 13C NMR spectrum. The analogous presentation of the data set for the respective 1H NMR-spectroscopic information is provided in Fig. S.1 in the SI.
838 structural groups is visualized considering the information from the 13C NMR spectrum. The analogous presentation of the data set for the respective 1H NMR-spectroscopic information is provided in Fig. S.1 in the SI.
Fig. 2a denotes the number of each of the 13 distinguished structural groups Ng (cf.Table 1) in the augmented pure-component data set, broken down to segments in the 13C NMR spectrum where their respective signals occur. It is important to note that the segmentation of the spectrum used in Fig. 2 is solely for visualization purposes but not used in the DSM, which is in contrast to our previous SVC-based approach, where spectral segmentation was required.35,36 The structural groups exhibit significant overlap in their chemical shift distributions, i.e., they are not confined to specific regions but span a wide range of the 13C NMR spectrum.
Fig. 2b gives an overview of the proportion Psyn = Nsyng/Ng of the number of structural groups Nsyng that incorporate synthetic data for either 13C, 1H, or both. Separate visualizations showing the distribution of Psyn for structural groups containing synthetic data exclusively for 13C or 1H are provided in Fig. S.2 in the SI. Overall, structural groups containing synthetic spectral data account for 11.80% of the entire data set, with 0.84% containing synthetic data for 13C and 11.14% for 1H. Most structural groups with synthetic data are concentrated in the regions below 80 ppm and between 110 and 140 ppm in the 13C NMR spectrum. This distribution likely results from the high density of various structural groups, i.e., aliphatic, cyclic, and double bound carbon groups, in these regions for organic molecules, which complicates signal differentiation and accurate assignment of chemical shifts δ1H in the crowded 1H NMR spectrum (cf. SI for details). Furthermore, augmentations with synthetic data are necessary for carbonyl ketones with signals exceeding 220 ppm, as experimental spectra do not extend to these elevated chemical shifts δ13C by default.
In the SI, we provide a detailed analysis of the influence of synthetic NMR data on the training and predictive performance of the DSM, thereby demonstrating the robustness of the model with respect to the composition of the training data.
Generation of input and output data
The chemical shifts for the 13C and 1H nuclei, the substitution degree of the 13C nuclei, the boolean variable L denoting the presence or absence of labile protons, and the correct structural groups for each pure component in our data set were automatically obtained from the pure-component NMR data using RDKit, as described in our previous work.36 The input data for each pure component and its structural groups were organized into a set-based structure, as illustrated in Fig. 1 and detailed in Section Deep-set architecture. This set-structured input data was automatically generated by determining the carbon-hydrogen connections based on the pure-component structures using RDKit. The associated output data was generated by one-hot encoding the structural groups contained in the respective pure component, as defined in Table 1.Training and evaluation of the deep set model
We trained and evaluated the DSM using the generated pure-component data set to assess predictive performance and robustness. The measured test mixture data are used solely to demonstrate the practical applicability of the method.The generated data set was randomly split into a training, a validation, and a test set, comprising 80%, 10%, and 10% of the pure components from our data set, respectively. The test set was constrained to include only pure components with entirely experimental spectral data, i.e., not including synthetic data, to demonstrate the model's performance in the most realistic scenario. Furthermore, the training set was augmented by synthetic binary and ternary mixture data obtained by simply “mixing” the spectra of the respective pure components in the training set. As a result, each pure component present in the training set appeared three times in the final training set: once with its pure component spectra, once with the spectra of a binary mixture with a randomly chosen other component from the training set, and once with the spectra of a ternary mixture with two randomly chosen other components. In Table S.1 in the SI, we provide an analysis demonstrating the robustness of the DSM to different random splits of the data set.
All models and scripts for training and evaluation were implemented in Python 3.6.8 using PyTorch 2.2.1.55 Training was performed on an A40 GPU using the CrossEntropyLoss function with default PyTorch settings. The Adam optimizer was employed for weight optimization, and a learning rate scheduler with a decay factor of 0.1 and a patience of 20 epochs based on validation loss was utilized. Training was terminated early if the validation loss did not improve for 30 consecutive epochs, and the model achieving the lowest validation loss was selected. Typical training times ranged between one and two hours, while typical inference times were between four and six milliseconds.
Hyperparameter optimization, including the weight decay λ of the Adam optimizer, the initial learning rate, the batch size, and the number of layers and nodes in each network, was performed using a grid search based on validation loss. In the SI, we discuss the sensitivity of the model to the varied hyperparameters and present the validation loss results. The following hyperparameters were selected as final settings: a weight decay of λ = 5 × 10−4, an initial learning rate of 1 × 10−4, and a batch size of one. The network architectures were defined with three layers containing eight nodes each for ϕ13C and ϕ1H and two layers containing 256 nodes each for ϕ and ρ. In all networks, the Sigmoid Linear Unit (SiLU) activation function with default PyTorch settings was applied. In all cases, the number of nodes for the equivariant layer σ(α) was chosen to match those of the networks ϕ and ρ. The input dimensions of ϕ13C and ϕ1H were set to five according to the input data dimension, while the network ρ included an additional node, to account for the boolean variable L indicating the presence of labile protons, and had an output dimension of 13, corresponding to the number of distinct structural groups.
The predictive performance of the DSM on unseen test data was evaluated using the F1 score F1,g for each structural group g:
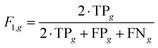 | (1) |
For comparison, we have also retrained and evaluated the SVC from our previous work36 using this work's data set and the same partitioning of the data into training, validation, and test sets, as employed for the DSM. Further information on the training and evaluation of the SVC is provided in the SI.
Furthermore, a final version of the DSM was trained by randomly using the data for 90% of the pure components from our data set for training and the remaining 10% for validation. Unlike the primary evaluation approach described above, this model was not evaluated on a separate pure-component test set. Instead, it was directly applied to experimentally studied test mixtures to demonstrate its practical applicability in predicting the structural groups in real mixtures, as detailed below.
Experimental methods
The compositions of the test mixtures studied in this work, as determined gravimetrically and used a ground truth here, are given in Table 2. Details on the chemicals and protocols used for mixture preparation are given in the SI. The test mixtures were selected so that each structural group considered in the developed NMR fingerprinting method, cf.Table 1 is represented in at least one of the mixtures.| Mixture | Components i | x i mol mol−1 |
|---|---|---|
| I | Water | 0.9266 |
| Acetone | 0.0244 | |
| Tartaric acid | 0.0246 | |
| 1,4-Butanediol | 0.0244 | |
| II | Diethyl ether | 0.7005 |
| Butanal | 0.1494 | |
| Butyl acetate | 0.1501 | |
| III | Cyclohexane | 0.5001 |
| Hexene | 0.3000 | |
| Diglyme | 0.1999 | |
| IV | Anisole | 0.7998 |
| 1-Octanol | 0.1198 | |
| 3-Methylbutan-2-one | 0.0804 |
1H NMR, 1H –13C HSQC NMR, 13C NMR, and 13C DEPT NMR spectra with pulse angles of 90° and 135° were recorded for each test mixture using a 60 MHz benchtop NMR spectrometer (Spinsolve 60 Ultra, Magritek). The settings of the NMR experiments, spectral processing procedures, and extraction of spectral information are reported in the SI.
In 1H –13C HSQC NMR spectra of mixtures, significant signal overlap is a common challenge, obscuring the cross-signals between 1H and 13C nuclei at low concentrations, especially when using benchtop NMR spectrometers with limited sensitivity and resolution. In our experiments, we encountered this exact problem: despite clear evidence of 1H bonded to 13C nuclei as determined by the substitution degree via13C DEPT NMR, in some cases, the absence of observable cross-signals in the 1H –13C HSQC spectra prevented the determination of the chemical shifts δ1Hi necessary for applying the DSM. To address this challenge, we have developed a model for the relationship between the chemical shifts δ13Ci of 13C nuclei and the chemical shifts δ1Hi of their connected 1H nuclei using linear regression, which we have fitted to our comprehensive data set for pure components. This regression model enables the determination of the most likely value of δ1Hi of the connected 1H nuclei for a given δ13Ci. In cases where no cross signals for a13C signal in the 1H –13C HSQC were identified in the studied mixture but the 13C DEPT results indicated that there should be a cross-signal, we have supplemented the missing experimental spectral information by estimating the δ1Hi based on the respective δ13Ci using the regression model. Further details on the regression model are provided in the SI.
The processed spectral information was then fed as input to the DSM to identify the structural groups in the test mixtures. The task here is to assign a group from Table 1 to each signal in the 13C NMR spectrum. The resolution, even of the benchtop NMR spectrometer, is generally high enough to avoid that two different groups produce signals that cannot be distinguished. Even though this case cannot be strictly excluded, we do not consider it here. While the identification and assignment of structural groups is fully automated, the subsequent quantification step is currently performed manually. Quantitative information on the identified structural groups was finally obtained by manually integrating their signals in the 13C NMR spectrum and calculating the group mole fractions xg from the signal areas Ag, see eqn (2):
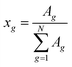 | (2) |
Results and discussion
Prediction of structural groups from pure-component spectra
Fig. 3 presents the F1,g scores of the DSM in predicting the structural groups of the pure components from the test set. The model generally achieves high F1,g scores, indicating high prediction accuracy, across all structural groups. Decreases in the F1,g score are observed only at the boundaries of the characteristic ranges in the 13C NMR spectrum for each structural group. Specifically, F1,g scores below 0.5 are found exclusively for CHarx, RO-CHarx, and ROOCHx groups, with chemical shifts δ13Ci outside their characteristic ranges (cf.Fig. 2). Overall, the developed DSM achieves an average F1 score of 0.92 across all structural groups, demonstrating excellent predictive performance, while significantly outperforming our previous SVC model, attaining a F1 score of 0.85 (cf. Fig. S.3 in the SI for details).Prediction of structural groups from mixture spectra
In the following, the results from the analysis of the studied test mixtures (cf.Table 2) with the DSM are presented and discussed.For the test mixtures studied in this work, no overlapping 13C NMR signals were observed that would affect peak integration and, consequently, the predicted mole fractions xg. In the event of signal overlap in the analysis of a mixture, integration could be performed by integrating the observable peak envelopes according to their shapes. In cases of more pronounced overlap, recently proposed ML-based deconvolution methods24–26 could be employed to facilitate signal separation prior to integration. If individual peaks cannot be reliably distinguished even after such treatment, the overlapping signals could be treated as a single contribution and assigned to all corresponding predicted structural groups.
Conclusions
In this work, we have introduced a deep set model (DSM) to automatically elucidate the structural groups in unknown pure samples and mixtures using the spectral information from standard NMR experiments of the sample, a task we call NMR fingerprinting. The DSM was specifically engineered to process the characteristics of NMR data, e.g., to allow inputs of varying length as generally different numbers of NMR signals must be expected for different samples, and trained on experimental pure-component NMR spectra from open-source databases to predict the structural groups and assign them to the corresponding signals in the 13C NMR spectrum of any given sample. To overcome problems with limited experimental training data, we have augmented the experimental data set with synthetic spectral data generated from predicted NMR spectra.Furthermore, we have incorporated 1H –13C HSQC NMR data as the first 2D NMR information into the NMR fingerprinting approach. This integration opens the possibility of mapping the structural groups identified in the 13C spectrum to the corresponding signals in the 1H spectrum, improving the deconvolution and interpretability of complex 1H spectra of mixtures. However, achieving such mapping remains challenging due to extensive signal overlap and low resolution of 1H spectra measured using benchtop NMR spectrometers. Therefore, realizing the mapping to 1H spectra when applying the NMR fingerprinting method to high-field NMR spectrometers could be a goal of future work.
In scenarios where HSQC acquisition is impractical, e.g. when minimal measurement time is required, the DSM-based approach cannot be applied, as the HSQC information is essential to establish the set-based structure of the NMR input. In such cases, our previous SVC-based NMR fingerprinting approach,36 which operates solely on 1D NMR data, can be used.
Evaluation on experimental test data for unseen pure components demonstrates the excellent performance of the DSM in predicting the structural groups of pure components, significantly exceeding our previous NMR fingerprinting approach, which was based on an SVC. To demonstrate its applicability to unknown mixtures, we have applied the DSM to NMR spectra of test mixtures measured using a simple benchtop NMR device. The results show remarkable agreement with the true mixture compositions, demonstrating the potential of the DSM-based NMR fingerprinting method for efficient automated mixture analysis, including in low-field NMR settings, which provides the basis for thermodynamic modeling of unknown mixtures via group-contribution methods.45,47
Despite the high-quality predictions, the current method still has limitations. Most importantly, it is restricted to structural groups consisting of only carbon, oxygen, and hydrogen. Expanding the range of structural groups poses challenges due to the increasing overlap in their spectral ranges, which can, in principle, be mitigated by combining multiple spectroscopic techniques, as demonstrated for Fourier-transform infrared (FT-IR) spectroscopy.56,57 However, the flexible architecture of the DSM allows for the incorporation of additional NMR data, e.g., from heteronuclear multiple bond correlation (HMBC) NMR, which can enhance distinguishability and enable the inclusion of further structural groups, such as those containing nitrogen, without requiring spectroscopic methods beyond NMR in future work. Furthermore, the successful integration of synthetic data in this work suggests that augmenting the training data with synthetic NMR spectra can further improve the model's predictive capabilities for new structural groups with limited available data.
Author contributions
Jens Wagner: data curation, investigation, methodology, software, validation, visualization, writing – original draft. Kerstin Münnemann: funding acquisition, resources, writing – review & editing. Thomas Specht: conceptualization, methodology, writing – review & editing. Hans Hasse: funding acquisition, resources, supervision, writing – review & editing. Fabian Jirasek: conceptualization, funding acquisition, resources, supervision, writing – review & editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
Data for this article, including the NMR spectra data, training scripts, and the final trained model, are available at Zenodo. The archived version corresponding to the manuscript is available at https://doi.org/10.5281/zenodo.18310430. The most recent version is available at https://doi.org/10.5281/zenodo.17597708.Supplementary information (SI): generation of synthetic NMR data; data distribution in the 1H and 13C spectrum; sensitivity studies; comparison of DSM with support vector classification; experimental methods; augmented data set, training scripts, and final model. See DOI: https://doi.org/10.1039/d5dd00490j.
Acknowledgements
We gratefully acknowledge financial support by the Carl Zeiss Foundation in the projects ‘Process Engineering 4.0’ and ‘Halocycles’, as well as by DFG in the Research Grant (project number 462456621), the Core Facility ‘LASE-MR’ (project number 537627671), and the research training group ‘WERA’ (project number 503479768).References
- M. Elyashberg, K. Blinov, S. Molodtsov, Y. Smurnyy, A. J. Williams and T. Churanova, Computer-assisted methods for molecular structure elucidation: realizing a spectroscopist’s dream, J. Cheminf., 2009, 1(1), 3 Search PubMed.
- D. C. Burns, E. P. Mazzola and W. F. Reynolds, The role of computer-assisted structure elucidation (CASE) programs in the structure elucidation of complex natural products, Nat. Prod. Rep., 2019, 36, 919–933 RSC.
- M. Elyashberg and D. Argyropoulos, Computer Assisted Structure Elucidation (CASE): Current and future perspectives, Magn. Reson. Chem., 2020, 59, 669–690 CrossRef PubMed.
- Z. Huang, M. S. Chen, C. P. Woroch, T. E. Markland and M. W. Kanan, A framework for automated structure elucidation from routine NMR spectra, Chem. Sci., 2021, 12, 15329–15338 RSC.
- M. Valli, H. M. Russo, A. C. Pilon, M. E. F. Pinto, N. B. Dias, R. T. Freire, I. Castro-Gamboa and V. d. S. Bolzani, Computational methods for NMR and MS for structure elucidation I: software for basic NMR, Phys. Sci. Rev., 2019, 4(10) DOI:10.1515/psr-2018-0108.
- B. Sridharan, S. Mehta, Y. Pathak and U. D. Priyakumar, Deep Reinforcement Learning for Molecular Inverse Problem of Nuclear Magnetic Resonance Spectra to Molecular Structure, J. Phys. Chem. Lett., 2022, 13, 4924–4933 CrossRef CAS PubMed.
- M. Alberts, F. Zipoli and A. C. Vaucher, Learning the Language of NMR: Structure Elucidation from NMR spectra using Transformer Models, ChemRxiv, 2023, preprint, DOI:10.26434/chemrxiv-2023-8wxcz.
- F. Hu, M. S. Chen, G. M. Rotskoff, M. W. Kanan and T. E. Markland, Accurate and Efficient Structure Elucidation from Routine One-Dimensional NMR Spectra Using Multitask Machine Learning, ACS Cent. Sci., 2024, 10, 2162–2170 CrossRef CAS PubMed.
- X. Tan, A transformer based generative chemical language AI model for structural elucidation of organic compounds, J. Cheminf., 2025, 17(1) DOI:10.1186/s13321-025-01016-1.
- M. Alberts, N. Hartrampf and T. Laino, Automated Structure Elucidation at Human-Level Accuracy via a Multimodal Multitask Language Model, ChemRxiv, 2025, preprint, DOI:10.26434/chemrxiv-2025-q80r9.
- R. Behrens, E. Kessler, K. Münnemann, H. Hasse and E. von Harbou, Monoalkylcarbonate formation in the system monoethanolamine–water–carbon dioxide, Fluid Phase Equilib., 2019, 486, 98–105 CrossRef CAS.
- D. Bellaire, H. Kiepfer, K. Münnemann and H. Hasse, PFG-NMR and MD Simulation Study of Self-Diffusion Coefficients of Binary and Ternary Mixtures Containing Cyclohexane, Ethanol, Acetone, and Toluene, J. Chem. Eng. Data, 2020, 65, 793–803 CrossRef CAS.
- J.-N. Dumez, NMR methods for the analysis of mixtures, Chem. Commun., 2022, 58, 13855–13872 RSC.
- Y. Lee, Y. Matviychuk, B. Bogun, C. S. Johnson and D. J. Holland, Quantification of mixtures of analogues of illicit substances by benchtop NMR spectroscopy, J. Magn. Reson., 2022, 335, 107138 CrossRef CAS PubMed.
- M. Lin and M. J. Shapiro, Mixture Analysis by NMR Spectroscopy, Anal. Chem., 1997, 69, 4731–4733 CrossRef CAS.
- Y. Lu, F. Hu, T. Miyakawa and M. Tanokura, Complex Mixture Analysis of Organic Compounds in Yogurt by NMR Spectroscopy, Metabolites, 2016, 6, 19 CrossRef PubMed.
- Y. Matviychuk, E. Steimers, E. von Harbou and D. J. Holland, Bayesian approach for automated quantitative analysis of benchtop NMR data, J. Magn. Reson., 2020, 319, 106814 CrossRef CAS PubMed.
- Y. Matviychuk, E. Steimers, E. von Harbou and D. J. Holland, Improving the accuracy of model-based quantitative nuclear magnetic resonance, Magn. Reson., 2020, 1, 141–153 CrossRef PubMed.
- Y. Matviychuk, E. von Harbou and D. J. Holland, An experimental validation of a Bayesian model for quantification in NMR spectroscopy, J. Magn. Reson., 2017, 285, 86–100 CrossRef CAS PubMed.
- M. I. Osorio-Garcia, D. M. Sima, F. U. Nielsen, U. Himmelreich and S. Van Huffel, Quantification of magnetic resonance spectroscopy signals with lineshape estimation, J. Chemom., 2011, 25, 183–192 CrossRef CAS.
- A. A. Smith, INFOS: spectrum fitting software for NMR analysis, J. Biomol. NMR, 2017, 67, 77–94 CrossRef CAS PubMed.
- S. Sokolenko, T. Jézéquel, G. Hajjar, J. Farjon, S. Akoka and P. Giraudeau, Robust 1D NMR lineshape fitting using real and imaginary data in the frequency domain, J. Magn. Reson., 2019, 298, 91–100 CrossRef CAS PubMed.
- Z. Zhou, X. Liao, X. Qiu, Y. Zhang, J. Dong, X. Qu and D. Lin, NMRformer: A Transformer-Based Deep Learning Framework for Peak Assignment in 1D 1H NMR Spectroscopy, Anal. Chem., 2025, 97, 904–911 CrossRef CAS PubMed.
- N. Schmid, S. Bruderer, F. Paruzzo, G. Fischetti, G. Toscano, D. Graf, M. Fey, A. Henrici, V. Ziebart, B. Heitmann, H. Grabner, J. Wegner, R. Sigel and D. Wilhelm, Deconvolution of 1D NMR spectra: A deep learning-based approach, J. Magn. Reson., 2023, 347, 107357 CrossRef CAS PubMed.
- G. Fischetti, N. Schmid, S. Bruderer, G. Caldarelli, A. Scarso, A. Henrici and D. Wilhelm, Automatic classification of signal regions in 1H Nuclear Magnetic Resonance spectra, Front. Artif. Intell., 2023, 5, 1116416 CrossRef PubMed.
- N. Schmid, M. Wanner, G. Fischetti, A. Henrici, M. Meshkian, S. Bruderer, R. M. Füchslin, B. Heitmann, J. D. Wegner and R. K. O. Sigel, A Chemistry-Informed Deep Learning Model for Next-Generation Automated Analysis of 1H NMR Spectra, ChemRxiv, 2026, preprint, DOI:10.26434/chemrxiv.10001683/v1.
- A. F. Tawfike, C. Viegelmann and R. Edrada-EbelMetabolomics Tools for Natural Product Discovery, Humana Press, 2013, pp. 227–244 Search PubMed.
- A. Bakiri, J. Hubert, R. Reynaud, S. Lanthony, D. Harakat, J.-H. Renault and J.-M. Nuzillard, Computer-Aided 13C NMR Chemical Profiling of Crude Natural Extracts without Fractionation, J. Nat. Prod., 2017, 80, 1387–1396 CrossRef CAS PubMed.
- A. Bruguière, S. Derbré, J. Dietsch, J. Leguy, V. Rahier, Q. Pottier, D. Bréard, S. Suor-Cherer, G. Viault, A.-M. Le Ray, F. Saubion and P. Richomme, MixONat, a Software for the Dereplication of Mixtures Based on 13C NMR Spectroscopy, Anal. Chem., 2020, 92, 8793–8801 CrossRef PubMed.
- M. Elyashberg, Identification and structure elucidation by NMR spectroscopy, TrAC, Trends Anal. Chem., 2015, 69, 88–97 CrossRef CAS.
- Y. Jin, J.-J. Wang, F. Xu, X. Ji, Z. Gao, L. Zhang, G. Ke; R. Zhu and W. E. NMR-Solver: Automated Structure Elucidation via Large-Scale Spectral Matching and Physics-Guided Fragment Optimization, arXiv, 2025, preprint arXiv:2509.00640, https://arxiv.org/abs/2509.00640 Search PubMed.
- B. Yuan, C. Zhang, C. Ji, G. Liu, X. Li, S. Gong, X. Huang, A. Shen, X. Li and Y. Liu, HSQCid: A Powerful Tool for Paving the Way to High-Throughput Structural Dereplication of Natural Products Based on Fast NMR Experiments, Anal. Chem., 2025, 97, 3227–3235 CrossRef CAS PubMed.
- L. Yao, M. Yang, J. Song, Z. Yang, H. Sun, H. Shi, X. Liu, X. Ji, Y. Deng and X. Wang, Conditional Molecular Generation Net Enables Automated Structure Elucidation Based on 13C NMR Spectra and Prior Knowledge, Anal. Chem., 2023, 95, 5393–5401 CrossRef CAS PubMed.
- K. P. C. Vollhardt and N. E. Schore, Organic Chemistry: Structure and Function, 8th edn, Macmillan Learning, 2018 Search PubMed.
- T. Specht, K. Münnemann, H. Hasse and F. Jirasek, Automated Methods for Identification and Quantification of Structural Groups from Nuclear Magnetic Resonance Spectra Using Support Vector Classification, J. Chem. Inf. Model., 2021, 61, 143–155 CrossRef CAS PubMed.
- T. Specht, J. Arweiler, J. Stüber, K. Münnemann, H. Hasse and F. Jirasek, Automated nuclear magnetic resonance fingerprinting of mixtures, Magn. Reson. Chem., 2023, 62, 286–297 CrossRef PubMed.
- E. L. Ulrich, et al., BioMagResBank, Nucleic Acids Res., 2007, 36, D402–D408 CrossRef PubMed.
- S. Kuhn and N. E. Schlörer, Facilitating quality control for spectra assignments of small organic molecules: nmrshiftdb2 – a free in-house NMR database with integrated LIMS for academic service laboratories, Magn. Reson. Chem., 2015, 53, 582–589 CrossRef CAS PubMed.
- Daylight Theory Manual, Version 4.9. Daylight Chemical Information Systems, Inc., Aliso Viejo, CA. https://www.daylight.com/dayhtml/doc/theory/index.html, Last accessed: 13.12. 2024.
- T. Specht, K. Münnemann, H. Hasse and F. Jirasek, Rational method for defining and quantifying pseudo-components based on NMR spectroscopy, Phys. Chem. Chem. Phys., 2023, 25, 10288–10300 RSC.
- F. Jirasek, J. Burger and H. Hasse, Method for Estimating Activity Coefficients of Target Components in Poorly Specified Mixtures, Ind. Eng. Chem. Res., 2018, 57, 7310–7313 CrossRef CAS.
- F. Jirasek, J. Burger and H. Hasse, NEAT—NMR Spectroscopy for the Estimation of Activity Coefficients of Target Components in Poorly Specified Mixtures, Ind. Eng. Chem. Res., 2019, 58, 9155–9165 CrossRef CAS.
- F. Jirasek, J. Burger and H. Hasse, Application of NEAT for determining the composition dependence of activity coefficients in poorly specified mixtures, Chem. Eng. Sci., 2019, 208, 115161 CrossRef CAS.
- T. Specht, K. Münnemann, F. Jirasek and H. Hasse, Estimating activity coefficients of target components in poorly specified mixtures with NMR spectroscopy and COSMO-RS, Fluid Phase Equilib., 2020, 516, 112604 CrossRef CAS.
- J. Wagner, Z. Romero, K. Münnemann, T. Specht, F. Jirasek and H. Hasse, Thermodynamic modeling of poorly specified mixtures using NMR fingerprinting and group-contribution equations of state, Fluid Phase Equilib., 2025, 596, 114446 CrossRef CAS.
- F. Jirasek, J. Burger and H. Hasse, Application of NEAT for the simulation of liquid–liquid extraction processes with poorly specified feeds, AIChE J., 2019, 66 Search PubMed.
- T. Specht, H. Hasse and F. Jirasek, Predictive Thermodynamic Modeling of Poorly Specified Mixtures and Applications in Conceptual Fluid Separation Process Design, Ind. Eng. Chem. Res., 2023, 62, 10657–10667 CrossRef CAS.
- E. Wagstaff, F. B. Fuchs, M. Engelcke, M. A. Osborne and I. PosnerUniversal Approximation of Functions on Sets, 2021; preprint arXiv:2107.01959, https://arxiv.org/abs/2107.01959.
- M. Zaheer, S. Kottur, S. Ravanbakhsh, B. Poczos, R. Salakhutdinov and A. SmolaDeep Sets, 2017, preprint arXiv:1703.06114, https://arxiv.org/abs/1703.06114.
- M. M. Bronstein, J. Bruna, T. Cohen and P. VeličkovićGeometric Deep Learning: Grids, Groups, Graphs, Geodesics, and Gauges, 2021; preprint arXiv:2104.13478, https://arxiv.org/abs/2104.13478.
- RDKit: Open-Source Cheminformatics. https://www.rdkit.org, Last accessed: 13.12.2024.
- L. Patiny, H. Musallam, A. Bolaños, M. Zasso, J. Wist, M. Karayilan, E. Ziegler, J. C. Liermann and N. E. Schlörer, NMRium: Teaching nuclear magnetic resonance spectra interpretation in an online platform, Beilstein J. Org. Chem., 2024, 20, 25–31 CrossRef CAS PubMed.
- T. D. W. Claridge, High-Resolution NMR Techniques in Organic Chemistry; Elsevier, Amsterdam, Netherlands, 2016 Search PubMed.
- M. Soelch, A. Akhundov, P. van der Smagt and J. Bayer, Artificial Neural Networks and Machine Learning – ICANN 2019: Theoretical Neural Computation, Springer International Publishing, 2019, pp. 444–457 Search PubMed.
- A. Paszke, et al., PyTorch: An Imperative Style, High-Performance Deep Learning Library. 2019, preprint arXiv:1912.01703, https://arxiv.org/abs/1912.01703.
- J. A. Fine, A. A. Rajasekar, K. P. Jethava and G. Chopra, Spectral deep learning for prediction and prospective validation of functional groups, Chem. Sci., 2020, 11, 4618–4630 RSC.
- G. Lee, H. Shim, J. Cho and S.-I. Choi, Machine-Learning Approach to Identify Organic Functional Groups from FT-IR and NMR Spectral Data, ACS Omega, 2025, 10, 12717–12723 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |