Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Data augmentation in a triple transformer loop retrosynthesis model
Yves
Grandjean
 ,
David
Kreutter
,
David
Kreutter
 and
Jean-Louis
Reymond
and
Jean-Louis
Reymond
 *
*
Department of Chemistry, Biochemistry and Pharmaceutical Sciences, University of Bern, Freiestrasse 3, 3012 Bern, Switzerland. E-mail: jean-louis.reymond@unibe.ch
First published on 21st January 2026
Abstract
Reactions in the US Patent Office (USPTO) are biased towards a few over-represented reaction types, which potentially limits their usefulness for computer-assisted synthesis planning (CASP). To obtain an equilibrated dataset, we applied retrosynthesis templates to USPTO molecules as products (P) to generate starting materials (SM). We then used transformer T2 from our recently reported triple transformer loop (TTL) retrosynthesis model to predict reagents (R) for the SM → P reaction. Finally, we validated the prediction by requesting a high confidence prediction (>95%) for the prediction of P from SM + R by TTL transformer T3. We generated up to 5000 reactions per template, resulting in 27.5m validated fictive reactions covering the chemical space of the original USPTO dataset. To exemplify the use of this dataset, we demonstrate that a single-step retrosynthesis transformer model trained on a template equilibrated subset of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 fictive reactions outperforms the corresponding model trained on USPTO reactions only.
374 fictive reactions outperforms the corresponding model trained on USPTO reactions only.
Introduction
The challenge of computer-assisted synthesis planning (CASP) consists of training neural networks and related models with data on organic reactions to automatically propose possible retrosyntheses of any molecule of interest.1–21 One of the key current limitations in this field is the dataset of reactions available for training, which is based on data extracted from the publicly available US Patent Office (USPTO).22 For instance, classifying reactions in this dataset by reaction templates, as discussed below, reveals a typical power-law distribution in which the majority of USPTO reactions correspond to a small number of templates, while many templates only possess very few examples. This imbalance reflects a bias towards reaction types frequently used in the patent literature, as well as the fact that some reaction types are simply rare and only documented sparsely, including in the primary literature.This relative sparsity of reaction data has been addressed by data augmentation using SMILES randomization23 and more directly by applying reaction templates (abstracted transformation rules encoded in SMARTS or SMIRKS) to molecules from various sources to generate fictive reactions that are then added to the training data to augment CASP tool performance.24–26 The decision to apply a reaction template to a molecule and/or to accept or reject the generated fictive reaction relies on molecular similarity between the molecule or generated reaction and the database examples from which the template was originally extracted.
Herein we report a new data augmentation approach to enrich datasets in poorly represented reaction types by combining the use of reaction templates with transformer models inspired by our recently reported triple transformer loop (TTL) single-step retrosynthesis model.27,28 In the TTL, a product molecule (P) is first tagged at hypothetical atoms with a changing environment to form a series of labeled products (P*), each corresponding to a different bond disconnection. For each P*, a first transformer T1* predicts a starting material (SM), a second transformer T2 predicts reagents (R) from the output of T1*, and a third transformer T3 predicts the product P from the combined outputs of T1* and T2. The reaction is validated if the confidence score (CS) is higher than a chosen threshold, usually CS > 95%. In the data augmentation approach reported here, we do not use T1* but instead generate fictive reactions by applying reaction templates to P to generate a corresponding SM, followed by transformer T2 to predict R. To validate the reaction, we use the atom-mapping information to tag atoms with environmental changes in SM to form a labeled SM*, and request CS > 95%, a value previously found to efficiently select valid reactions,27,28 for the prediction of P by a transformer T3* trained to predict P from SM* + R (Fig. 1a).
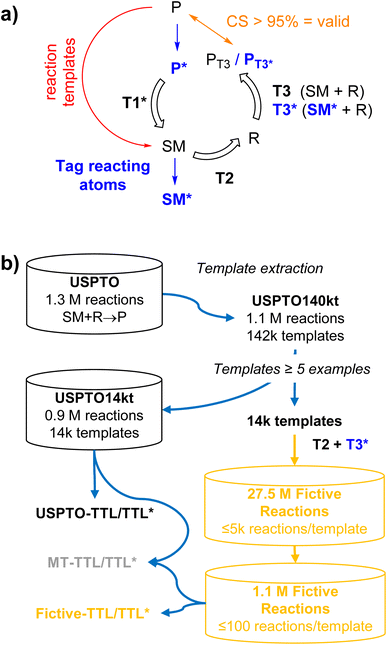 | ||
Fig. 1 Data augmentation in a triple transformer loop and evaluation for single-step retrosynthesis. (a) Details of the triple transformer loops TTL and TTL*. In the previously described TTL,27 the reactive atoms of a product molecule P are first tagged to produce P*. Transformer T1 then predicts the starting materials SM from P*, transformer T2 predicts the reagents R necessary to convert the predicted SM to P, and transformer T3 predicts the product P from the predicted SM + R. In the modified TTL* reported here, the atoms with environmental changes in SM are additionally tagged to produce SM*, and transformer T3* is trained to predict P from SM* + R. In both TTL and TTL*, the reaction SM + R → P or SM* + R → P is validated if the product predicted by T3 or T3* (PT3 or PT3*) is identical to P with a confidence score >95%. (b) 27.5m fictive reactions were generated by applying 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 templates with at least 5 examples, extracted from USPTO, to USPTO molecules as products (P), to generate starting materials (SM), and using transformers T2 to predict reagents (R) and transformer T3* to validate the fictive reactions, up to 5000 reactions per template. For evaluation, TTL models were trained with USPTO or fictive reactions. 024 templates with at least 5 examples, extracted from USPTO, to USPTO molecules as products (P), to generate starting materials (SM), and using transformers T2 to predict reagents (R) and transformer T3* to validate the fictive reactions, up to 5000 reactions per template. For evaluation, TTL models were trained with USPTO or fictive reactions. | ||
To obtain a template equilibrated dataset of fictive reactions, we apply our data augmentation approach to 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 reaction templates with at least 5 examples in the USPTO dataset to generate up to 5000 reactions per template, resulting in a dataset of 27.5m fictive reactions including reagents (SM + R → P). To test the effect of a template-equilibrated dataset on single-step retrosynthesis, we train a TTL using a subset of this dataset consisting of 1
024 reaction templates with at least 5 examples in the USPTO dataset to generate up to 5000 reactions per template, resulting in a dataset of 27.5m fictive reactions including reagents (SM + R → P). To test the effect of a template-equilibrated dataset on single-step retrosynthesis, we train a TTL using a subset of this dataset consisting of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 fictive reactions containing up to 100 reactions per template, to match the size of the original USPTO dataset, and compare its performance to that of a similar model trained on the USPTO dataset, and to that of a model trained with both datasets simultaneously by multi-task learning, which we have found previously to work well for reaction prediction models (Fig. 1b).29–31 Indeed, we find that our template-equilibrated dataset of fictive reactions leads to significant improvements in template-averaged single-step retrosynthesis performance.
374 fictive reactions containing up to 100 reactions per template, to match the size of the original USPTO dataset, and compare its performance to that of a similar model trained on the USPTO dataset, and to that of a model trained with both datasets simultaneously by multi-task learning, which we have found previously to work well for reaction prediction models (Fig. 1b).29–31 Indeed, we find that our template-equilibrated dataset of fictive reactions leads to significant improvements in template-averaged single-step retrosynthesis performance.
Results and discussion
Reaction dataset selection and generation of fictive reactions
We used the United States Patent and Trademark Office (USPTO) chemical reaction dataset,22,32 in the modified version listing 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 266
266![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 734 USPTO reactions with a single product (P) and between two and ten starting materials (SM).33 From this dataset, template extraction for both radius 0 (r0) and radius 1 (r1) templates in SMARTS format was performed using the rxnutils package34–36 and succeeded for 1
734 USPTO reactions with a single product (P) and between two and ten starting materials (SM).33 From this dataset, template extraction for both radius 0 (r0) and radius 1 (r1) templates in SMARTS format was performed using the rxnutils package34–36 and succeeded for 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100
100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 773 reactions, a dataset here named USPTO140kt. Templates were then standardized using the templatecorr package,37,38 which resulted in 141
773 reactions, a dataset here named USPTO140kt. Templates were then standardized using the templatecorr package,37,38 which resulted in 141![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 584 unique r1 templates, corresponding to between 1 and 24
584 unique r1 templates, corresponding to between 1 and 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 523 reactions per template (blue line, Fig. 2a).
523 reactions per template (blue line, Fig. 2a).
We focused the study on the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 r1 templates with at least five example reactions in the USPTO140kt dataset, corresponding to 934
024 r1 templates with at least five example reactions in the USPTO140kt dataset, corresponding to 934![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 688 reactions (84.9% of the USPTO140kt dataset, hereafter designated as USPTO14kt). These templates corresponded to up to 10 tags, with almost 50% of templates containing two tags. The distribution of these templates to be used for fictive reaction generation was somewhat narrower than in the entire USPTO140kt dataset because many templates with three or more tagged atoms have fewer than five examples (Fig. 2b). For each of the 14
688 reactions (84.9% of the USPTO140kt dataset, hereafter designated as USPTO14kt). These templates corresponded to up to 10 tags, with almost 50% of templates containing two tags. The distribution of these templates to be used for fictive reaction generation was somewhat narrower than in the entire USPTO140kt dataset because many templates with three or more tagged atoms have fewer than five examples (Fig. 2b). For each of the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 reaction templates, we searched the full USPTO dataset for all molecules matching the SMARTS template of the product (P). We then processed each matching molecule by applying the retrosynthesis template to obtain the corresponding starting materials (SM), and transformer T2 of our previously reported TTL27 to generate possible reagents R.
024 reaction templates, we searched the full USPTO dataset for all molecules matching the SMARTS template of the product (P). We then processed each matching molecule by applying the retrosynthesis template to obtain the corresponding starting materials (SM), and transformer T2 of our previously reported TTL27 to generate possible reagents R.
Early attempts to validate the resulting fictive reactions SM + R → P by applying transformer T3 of our original TTL to predict P from SM + R with a high confidence score (CS > 95%) resulted in a very low validation rate, which was caused by sensitivity to structural changes in the molecules that were unrelated to the reacting functional groups and often trivial (e.g. ethyl vs. methyl in a site remote from the reactive site). Fortunately, we found that the validation rate could be increased by identifying atoms with environmental changes in the predicted SM using RXNmapper39 to obtain a labeled SM* and using a modified transformer T3* trained with the USPTO140kt reactions to predict P from SM* + R.
We applied the above procedure to each of the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 templates until a maximum of 5000 reactions had been validated for each template. In total, approximately 60 million SM + R precursor pairs were produced by T1 and T2, 38.5 million of which produced the correct P when subjected to T3*. A subset of 27.5 million of these had a confidence score above 95%, covering 13
024 templates until a maximum of 5000 reactions had been validated for each template. In total, approximately 60 million SM + R precursor pairs were produced by T1 and T2, 38.5 million of which produced the correct P when subjected to T3*. A subset of 27.5 million of these had a confidence score above 95%, covering 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 953 (99.5%) of the 14
953 (99.5%) of the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 templates. In this dataset, only 692 templates had fewer reaction examples than in USPTO140kt, while most templates had more reaction examples than in USPTO140kt (red line, Fig. 2a). In view of training a retrosynthesis model, we selected a maximum of 100 reactions per template to form an equilibrated dataset of 1
024 templates. In this dataset, only 692 templates had fewer reaction examples than in USPTO140kt, while most templates had more reaction examples than in USPTO140kt (red line, Fig. 2a). In view of training a retrosynthesis model, we selected a maximum of 100 reactions per template to form an equilibrated dataset of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 validated fictive reactions. In this case, 12
374 validated fictive reactions. In this case, 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 285 of the templates had fewer examples than in USPTO140kt, 1518 templates had more examples than in USPTO140kt, and 150 templates had the same number of examples as in USPTO140kt (orange line, Fig. 2a).
285 of the templates had fewer examples than in USPTO140kt, 1518 templates had more examples than in USPTO140kt, and 150 templates had the same number of examples as in USPTO140kt (orange line, Fig. 2a).
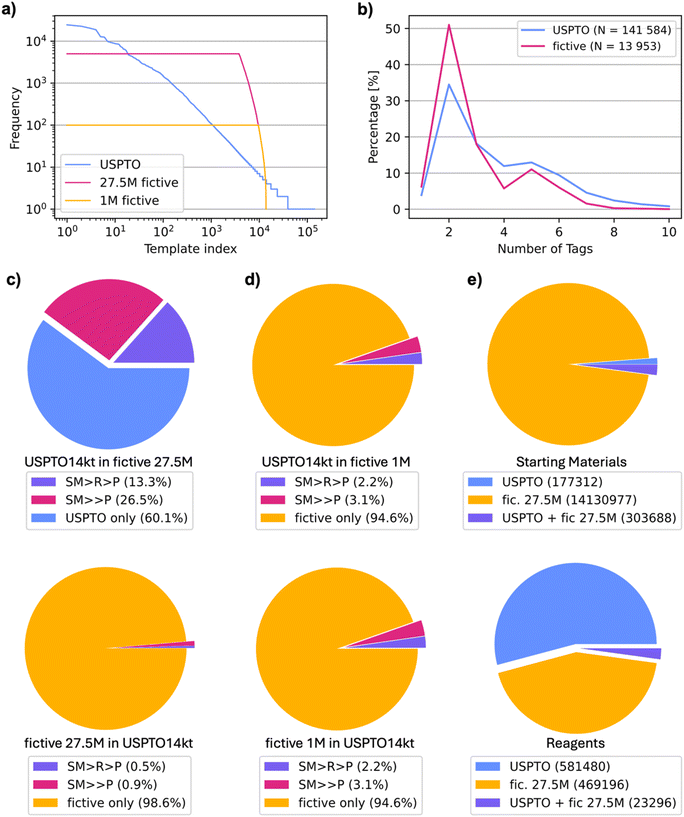
Further comparison of our 27.5m fictive reactions with the USPTO14kt dataset showed that our procedure had regenerated 39.9% of the USPTO14kt dataset when considering SM → P and 13.3% when considering SM + R → P. However, due to their number, most of the 27.5m fictive reactions (98.6% SM → P, 99.5% SM + R → P) were novel compared to USPTO14kt (Fig. 2c). Furthermore, the overlap between USPTO14kt and the 1m subset of our fictive reactions amounted to 6.3% of USPTO14kt (5.4% of the 1m fictive subset) for SM → P reactions and 2.6% of USPTO14kt (2.2% of the 1m fictive subset) for SM + R → P (Fig. 2d). In terms of starting materials, 303![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 688 of the 481
688 of the 481![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 (63.1%) of the SM in USPTO14kt had been regenerated by our fictive reaction generation procedure; however, 14
000 (63.1%) of the SM in USPTO14kt had been regenerated by our fictive reaction generation procedure; however, 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 130
130![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 977 (96.7%) of the SM in the 27.5m fictive reaction dataset were novel compared to USPTO14kt. For reagents R, the procedure had generated 469
977 (96.7%) of the SM in the 27.5m fictive reaction dataset were novel compared to USPTO14kt. For reagents R, the procedure had generated 469![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 196 new reagents, while only 23
196 new reagents, while only 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 296 (3.9%) of the 604
296 (3.9%) of the 604![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 776 reagents in USPTO14kt appeared in the fictive reaction dataset, reflecting the selection of templates as well as the effect of transformer T2 in predicting the most probable reagents (Fig. 2e).
776 reagents in USPTO14kt appeared in the fictive reaction dataset, reflecting the selection of templates as well as the effect of transformer T2 in predicting the most probable reagents (Fig. 2e).
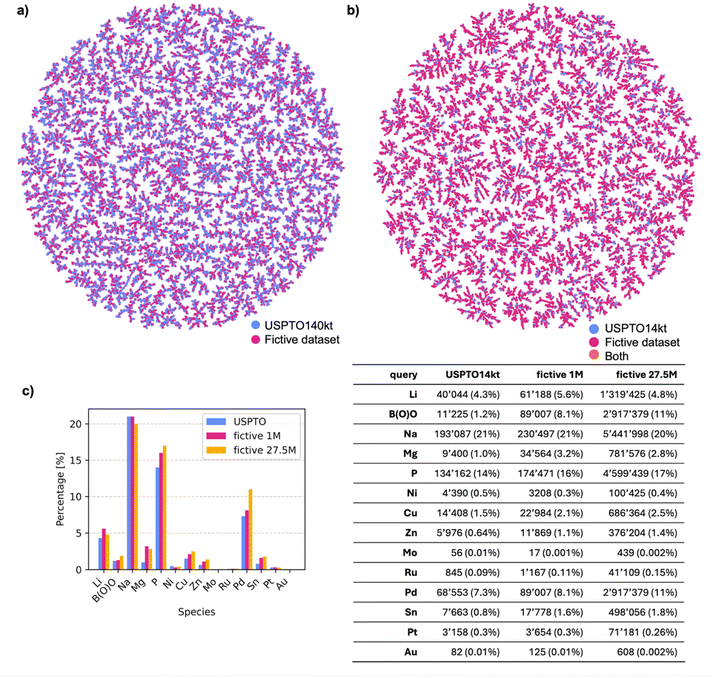
A closer comparison of USPTO and fictive reactions using a TMAP layout,40 computed for SM → P reactions using the differential reaction fingerprint DRFP as a similarity measure,41 showed that the generated reactions covered a similar chemical space to the original USPTO140kt (Fig. 3a). Similarly, although the vast majority of SM in the fictive reactions were novel compared to USPTO14kt, a TMAP layout using the substructure fingerprint MHFP6 as a similarity measure,42 showed that the fictive SM covered the space of USPTO14kt more broadly but in a similar manner (Fig. 3b). Indeed, the overall reaction types remained broadly comparable, as revealed by an analysis of reagents. For instance, sodium (Na) was present in approximately one fifth of the reactions in both USPTO14kt and the fictive datasets, reflecting mostly ester hydrolysis reactions (Fig. 3c). Phosphorus, present mostly in triphenylphosphine (metal-catalyzed processes) and olefination reagents (phosphoranes), increased slightly in the fictive reactions compared to USPTO14kt. Similarly, metals such as magnesium, lithium and zinc used in organometallic processes, as well as other relatively rare elements (Sn, Pt, Ru, and Au), increased significantly in fictive reactions, reflecting the effect of template equilibration.
Taken together, these analyses showed that our data augmentation approach combining templates and TTL transformers allowed us to produce a large, template equilibrated reaction dataset covering a chemical space comparable to the source data.
Impact of fictive reactions on template-averaged retrosynthesis performance
To assess the practical value of our fictive reaction dataset, we evaluated our approach on the single-step retrosynthesis task in our TTL model. This task is particularly well-suited for testing an equilibrated dataset, as it corrects the overrepresentation of highly frequent transformations while enriching underrepresented reactions, and template-free retrosynthesis models are especially sensitive to such distributional imbalances.33 For our comparative evaluation, we trained a reference TTL, here named USPTO-TTL, using the dataset of 934![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 688 USPTO14kt reactions corresponding to the 14
688 USPTO14kt reactions corresponding to the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 templates, with a train
024 templates, with a train![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) validation
validation![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) test set ratio of 80
test set ratio of 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, grouping reactions using a common template to avoid data leakage. Using the same procedure, we trained a second TTL, here named fictive-TTL, using the dataset of 1
10, grouping reactions using a common template to avoid data leakage. Using the same procedure, we trained a second TTL, here named fictive-TTL, using the dataset of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 fictive reactions, splitting training, validation and test sets with reactions derived from the templates assigned to the corresponding sets in the USPTO-TTL training. Finally, we trained a model using both reaction datasets by multi-task learning, here named MT-TTL. In each case, we also trained models in which the forward validation transformer T3* used starting material SM* with labeled atoms with a change in the environment, labeled USPTO-TTL*, fictive-TTL* and MT-TTL* (Fig. 1b).
374 fictive reactions, splitting training, validation and test sets with reactions derived from the templates assigned to the corresponding sets in the USPTO-TTL training. Finally, we trained a model using both reaction datasets by multi-task learning, here named MT-TTL. In each case, we also trained models in which the forward validation transformer T3* used starting material SM* with labeled atoms with a change in the environment, labeled USPTO-TTL*, fictive-TTL* and MT-TTL* (Fig. 1b).
To compare the different retrosynthesis models, we measured the single-step round-trip accuracy (RTA), averaged per reaction and starting with the product with tagged atoms. The RTA was introduced by Schwaller et al.43 and tests the ability of a retrosynthesis model to propose a valid retrosynthetic operation on a product molecule, rather than the ability to reproduce the same retrosynthetic operation as recorded in the test dataset. In addition, we also computed the RTA averaged per template (TA-RTA) to obtain an estimate across different reaction templates, independent of the number of examples per template.
The performances of the different models on the USPTO14kt dataset, which is dominated by a small number of highly populated reaction templates, dropped from approximately 82% on a per reaction basis (RTA) to approximately 65% when averaged per template (TA-RTA) across all three TTLs (Table 1, upper left). For this dataset, the USPTO-TTL performed best in terms of RTA but was overtaken by the fictive-TTL in terms of TA-RTA, while the MT-TTL was in between, reflecting the favorable effect of a template-equilibrated dataset on model performance. A similar effect was visible in the three TTLs*, whose performance was generally higher, taking advantage of starting materials with tagged atoms (dropped from >90% per reaction to ∼80% per template, Table 1, upper right). In this case, however, the fictive-TTL* surpassed the USPTO-TTL* in both RTA and TA-RTA.
| USPTO-TTL | Fictive-TTL | MT-TTL | USPTO-TTL* | Fictive-TTL* | MT-TTL* | ||
|---|---|---|---|---|---|---|---|
| USPTO14kt RTA | Top-1 | 82.8 | 78.6 | 82.7 | 90.4 | 91.9 | 91.5 |
| Top-2 | 84.5 | 0.9 | 84.5 | 90.8 | 92.3 | 91.8 | |
| Top-3 | 85.3 | 82.0 | 85.2 | 90.9 | 92.5 | 92.0 | |
| USPTO14kt TA-RTA | Top-1 | 62.6 | 69.0 | 65.2 | 78.2 | 83.4 | 81.7 |
| Top-2 | 64.7 | 71.3 | 66.9 | 78.5 | 83.8 | 82.1 | |
| Top-3 | 65.8 | 72.4 | 67.8 | 78.7 | 84.1 | 82.2 | |
| Fictive RTA | Top-1 | 64.3 | 78.5 | 70.7 | 84.1 | 92.4 | 89.9 |
| Top-2 | 66.4 | 80.5 | 72.6 | 85.5 | 92.7 | 90.1 | |
| Top-3 | 67.5 | 81.4 | 73.5 | 85.6 | 92.9 | 90.2 | |
| Fictive TA-RTA | Top-1 | 59.3 | 73.9 | 66.0 | 79.9 | 88.4 | 85.4 |
| Top-2 | 61.4 | 76.0 | 67.8 | 80.2 | 88.8 | 85.7 | |
| Top-3 | 62.6 | 76.8 | 68.7 | 80.4 | 89.0 | 85.8 | |
However, performances on the fictive reaction dataset, which contains the same number of reactions per template, were similar on a per-reaction (RTA) and on a per template (TA-RTA) basis (Table 1, lower half). On this dataset, models trained with USPTO14kt data only (USPTO-TTL and USPTO-TTL*) clearly suffered from the uneven composition of training data with respect to templates, performing ∼60% as TTL and ∼80% as TTL* compared to ∼74% and ∼88% for the corresponding models trained with fictive reactions (fictive-TTL and fictive-TTL*). Again, the MT-TTL and MT-TTL* performed in between the two other models.
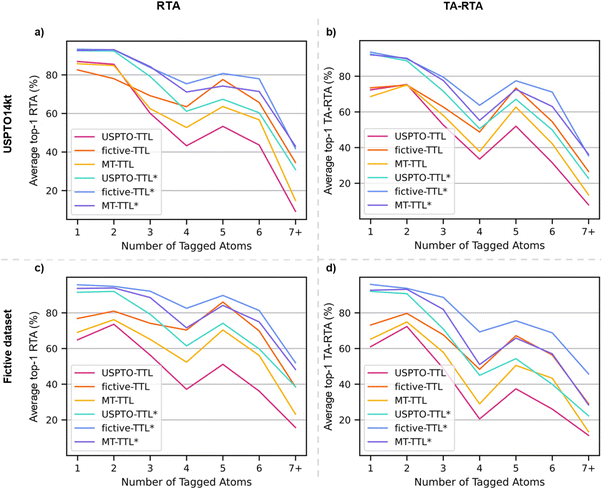
For both the UPSTO and the fictive reactions, performances were highest with the TTLs trained using fictive reactions (fictive-TTL and fictive-TTL*), reflecting the advantage of a template-equilibrated dataset for model training. There was no performance increase with the models trained on both datasets simultaneously (MT-TTL and MT-TTL*). In all cases, using starting materials with tagged atoms provided a strong performance advantage, with the model fictive-TTL* performing best across both test sets on a per reaction and on a per template basis. The same trends appeared when analyzing performance as a function of the number of tagged atoms, serving as an indication of reaction complexity (Fig. 4a and b). For all models, the RTA and TA-RTA dropped at four tagged atoms and strongly decreased for reactions with seven or more tagged atoms. The curves followed the same trend as the number of reactions in the datasets as a function of tagged (reacting) atoms (Fig. 2b). This trend might therefore simply reflect the influence of dataset size, although increasing reaction complexity might also play a role.
Conclusion
In this work, we presented a new data augmentation approach allowing for the enrichment of all reactivities present in the source dataset. Our approach consisted of generating starting materials (SM) by applying 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 r1 reaction templates extracted from the USPTO dataset to USPTO molecules as products (P) and applying transformer models for reagent (R) and product (P) prediction to validate up to 5000 fictive reactions SM + R → P per template. We used the confidence score of transformer models trained on the data as a filter to select meaningful transformations based on established knowledge. By this approach, we obtained a large dataset of 27.5m fictive reactions that covers and expands USPTO14kt's chemical space. A template-equilibrated dataset of 1
024 r1 reaction templates extracted from the USPTO dataset to USPTO molecules as products (P) and applying transformer models for reagent (R) and product (P) prediction to validate up to 5000 fictive reactions SM + R → P per template. We used the confidence score of transformer models trained on the data as a filter to select meaningful transformations based on established knowledge. By this approach, we obtained a large dataset of 27.5m fictive reactions that covers and expands USPTO14kt's chemical space. A template-equilibrated dataset of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 validated fictive reactions containing up to 100 reactions per template was used to evaluate the impact of equilibrated datasets on the single-step retrosynthesis task. We showed that the per-template round-trip accuracy of the non-augmented TTL can be significantly improved by using fictive, template-equilibrated data, and even more so by replacing the forward reaction prediction T3 model with a forward-tag validation model T3*. The fictive reaction dataset presented here could be useful to evaluate different retrosynthesis models, evaluate classification performance or other tasks related to reaction SMILES. Furthermore, the data augmentation scheme can be applied to better exploit the information contained in other open-source datasets.
374 validated fictive reactions containing up to 100 reactions per template was used to evaluate the impact of equilibrated datasets on the single-step retrosynthesis task. We showed that the per-template round-trip accuracy of the non-augmented TTL can be significantly improved by using fictive, template-equilibrated data, and even more so by replacing the forward reaction prediction T3 model with a forward-tag validation model T3*. The fictive reaction dataset presented here could be useful to evaluate different retrosynthesis models, evaluate classification performance or other tasks related to reaction SMILES. Furthermore, the data augmentation scheme can be applied to better exploit the information contained in other open-source datasets.
Methods
Datasets and template extraction
We used the United States Patent and Trademark Office (USPTO) chemical reaction dataset,22,32 in the modified version by Thakkar et al.,33 which is limited to reactions with a single product (P) and between 2 and 10 starting materials (SM) and reagents (R). Retrosynthetic reaction templates from this dataset were extracted in SMARTS format with radii 0 and 1 using the rxnutils package.34–36 A radius 0 template only carries information concerning atoms whose environment changed throughout the reaction, whereas radius 1 templates also include information for the atoms connected to them. The extraction yielded 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 663 unique template hashes for radius 0 and 262
663 unique template hashes for radius 0 and 262![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 266 for radius 1. To standardize the syntax of radius 1 templates, we employed the templatecorr package,37,38 which requires both radii 0 and 1 templates. The hierarchical correction algorithm uses subgraph isomorphisms on templates sharing the same general template (r0). If several higher radius templates are found to be equivalent, they are all rewritten as the most general and exclusive pattern (we used the published method following the available tutorials as detailed in https://github.com/hesther/templatecorr). The standardization resulted in the dataset here named USPTO140kt featuring 1
266 for radius 1. To standardize the syntax of radius 1 templates, we employed the templatecorr package,37,38 which requires both radii 0 and 1 templates. The hierarchical correction algorithm uses subgraph isomorphisms on templates sharing the same general template (r0). If several higher radius templates are found to be equivalent, they are all rewritten as the most general and exclusive pattern (we used the published method following the available tutorials as detailed in https://github.com/hesther/templatecorr). The standardization resulted in the dataset here named USPTO140kt featuring 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100
100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 773 reactions corresponding to 141
773 reactions corresponding to 141![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 584 unique radius 1 templates. Further constraining this dataset to templates with at least five occurrences left the dataset here named USPTO14kt, containing 934
584 unique radius 1 templates. Further constraining this dataset to templates with at least five occurrences left the dataset here named USPTO14kt, containing 934![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 688 reactions from 14
688 reactions from 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 radius 1 templates.
024 radius 1 templates.
Generating fictive reactions
The generation of fictive reactions started with a pool of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 505
505![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 837 molecules collected from USPTO, split into 1000 subsets. For each of the above-mentioned 14
837 molecules collected from USPTO, split into 1000 subsets. For each of the above-mentioned 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 radius 1 templates in SMARTS format, we searched the 1000 subsets of USPTO molecules in random order for molecules matching the product (P) of the template. For each matching molecule, we applied the template to generate the corresponding starting materials (SM) and used transformer T2 of our previously reported TTL28 to obtain reagents (R). We then labeled atoms with an environmental change in the template-generated SM using RNXmapper39 to obtain SM* with tagged atoms, labeled with a specific token (“!”) placed next to each of the said atoms as described before.27 Finally, we predicted P using transformer T3*, trained to predict P from SM* + R with tagged reactant atoms using USPTO140kt (split into 990
024 radius 1 templates in SMARTS format, we searched the 1000 subsets of USPTO molecules in random order for molecules matching the product (P) of the template. For each matching molecule, we applied the template to generate the corresponding starting materials (SM) and used transformer T2 of our previously reported TTL28 to obtain reagents (R). We then labeled atoms with an environmental change in the template-generated SM using RNXmapper39 to obtain SM* with tagged atoms, labeled with a specific token (“!”) placed next to each of the said atoms as described before.27 Finally, we predicted P using transformer T3*, trained to predict P from SM* + R with tagged reactant atoms using USPTO140kt (split into 990![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 391 reactions for training, 55
391 reactions for training, 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 278 for validation, and 55
278 for validation, and 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 104 for testing), and retained the fictive reaction if the confidence score of T3* exceeded 95%. Confidence scores of OpenNMT44,45 models have been developed and used in previous work by Kreutter et al.31
104 for testing), and retained the fictive reaction if the confidence score of T3* exceeded 95%. Confidence scores of OpenNMT44,45 models have been developed and used in previous work by Kreutter et al.31
The above procedure was repeated for each of the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 r1 templates until up to 5000 fictive reactions had been validated or all USPTO molecules matching the product side of the template had been tested as products. The procedure succeeded for 13
024 r1 templates until up to 5000 fictive reactions had been validated or all USPTO molecules matching the product side of the template had been tested as products. The procedure succeeded for 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 953 (99.5%) of the 14
953 (99.5%) of the 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 024 r1 templates and resulted in 27.5m fictive reactions. A smaller template-equilibrated subset of 1
024 r1 templates and resulted in 27.5m fictive reactions. A smaller template-equilibrated subset of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 fictive reactions was obtained by collecting up to 100 reactions per template.
374 fictive reactions was obtained by collecting up to 100 reactions per template.
Performance evaluation
The possible use of our fictive reactions was evaluated by measuring the single-step retrosynthesis performance of TTL models trained with either USPTO14kt or the similarly sized, smaller subset of fictive reactions mentioned above, or with both sets together in multitask learning using each 0.5 weight coefficient using the OpenNMT library.44,45 Datasets were split so that all reactions, from both the USPTO14kt subset and the fictive reaction subset, belonging to the same reaction template would be in the same split (training, validation or test set), for an overall 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10. TTL transformer models were trained using our previously reported procedures,27,28 whereby the validation transformer T3 was trained either with unlabeled reactions (USPTO-TTL, fictive-TTL and MT-TTL) or with reactions featuring labeled reactive atoms in SM* (USPTO-TTL*, fictive-TTL* and MT-TTL*). The six different models were compared with the round-trip accuracy metric (RTA), measuring the frequency with which the product (P) is regenerated by the TTL among the list of top-N predictions, averaged across all reactions (RTA), or averaged per template (TA-RTA).
10. TTL transformer models were trained using our previously reported procedures,27,28 whereby the validation transformer T3 was trained either with unlabeled reactions (USPTO-TTL, fictive-TTL and MT-TTL) or with reactions featuring labeled reactive atoms in SM* (USPTO-TTL*, fictive-TTL* and MT-TTL*). The six different models were compared with the round-trip accuracy metric (RTA), measuring the frequency with which the product (P) is regenerated by the TTL among the list of top-N predictions, averaged across all reactions (RTA), or averaged per template (TA-RTA).
Author contributions
YG designed and carried out the study and wrote the paper, DK designed the study and gave technical guidance, and JLR designed and supervised the study and wrote the paper.Conflicts of interest
The authors declare that they have no competing interests.Data availability
Code availability: the rxnutils package used for template extraction is available at https://github.com/MolecularAI/reaction_utils. The templatecorr package used to correct r1 templates is available at https://github.com/hesther/templatecorr. The code of the enrichment framework is available at https://github.com/yvsgrndjn/USPTO_balance.The USPTO version from Thakkar et al. can be found in their Zenodo repository.33,46 The dataset of fictive reactions created in this work (27.5m reactions) is available on Zenodo at https://doi.org/10.5281/zenodo.13120462. The equilibrated fictive dataset of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 097
097![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 374 reactions with up to 100 reactions per reaction template is available at https://doi.org/10.5281/zenodo.17301372. TMAPs from Fig. 3 are available as interactive plots on Zenodo: https://zenodo.org/records/17300855.
374 reactions with up to 100 reactions per reaction template is available at https://doi.org/10.5281/zenodo.17301372. TMAPs from Fig. 3 are available as interactive plots on Zenodo: https://zenodo.org/records/17300855.
Acknowledgements
This work was supported financially by the Swiss National Science Foundation, Grant no. 200020_207976. Calculations were performed on UBELIX (http://www.id.unibe.ch/hpc), the HPC cluster at the University of Bern.References
- E. J. Corey, General Methods for the Construction of Complex Molecules, Pure Appl. Chem., 1967, 14(1), 19–38, DOI:10.1351/pac196714010019.
- E. J. Corey and W. T. Wipke, Computer-Assisted Design of Complex Organic Syntheses, Science, 1969, 166(3902), 178–192, DOI:10.1126/science.166.3902.178.
- D. A. Pensak and E. J. Corey, LHASA—Logic and Heuristics Applied to Synthetic Analysis, in Computer-assisted organic synthesis, 1977, pp. 1–32, DOI:10.1021/bk-1977-0061.ch001.
- E. J. Corey, A. K. Long and S. D. Rubenstein, Computer-Assisted Analysis in Organic Synthesis, Science, 1985, 228(4698), 408–418, DOI:10.1126/science.3838594.
- P. Y. Johnson, I. Burnstein, J. Crary, M. Evans and T. Wang, Designing an Expert System for Organic Synthesis, in Expert System Applications in Chemistry, ACS Symposium Series, American Chemical Society, 1989, vol. 408, pp. 102–123, DOI:10.1021/bk-1989-0408.ch009.
- W.-D. Ihlenfeldt and J. Gasteiger, Computer-Assisted Planning of Organic Syntheses: The Second Generation of Programs, Angew Chem. Int. Ed. Engl., 1996, 34(23–24), 2613–2633, DOI:10.1002/anie.199526131.
- J. Law, Z. Zsoldos, A. Simon, D. Reid, Y. Liu, S. Y. Khew, A. P. Johnson, S. Major, R. A. Wade and H. Y. Ando, Route Designer: A Retrosynthetic Analysis Tool Utilizing Automated Retrosynthetic Rule Generation, J. Chem. Inf. Model., 2009, 49(3), 593–602, DOI:10.1021/ci800228y.
- C. D. Christ, M. Zentgraf and J. M. Kriegl, Mining Electronic Laboratory Notebooks: Analysis, Retrosynthesis, and Reaction Based Enumeration, J. Chem. Inf. Model., 2012, 52(7), 1745–1756, DOI:10.1021/ci300116p.
- A. Bøgevig, H.-J. Federsel, F. Huerta, M. G. Hutchings, H. Kraut, T. Langer, P. Löw, C. Oppawsky, T. Rein and H. Saller, Route Design in the 21st Century: The ICSYNTH Software Tool as an Idea Generator for Synthesis Prediction, Org. Process Res. Dev., 2015, 19(2), 357–368, DOI:10.1021/op500373e.
- S. Szymkuć, E. P. Gajewska, T. Klucznik, K. Molga, P. Dittwald, M. Startek, M. Bajczyk and B. A. Grzybowski, Computer-Assisted Synthetic Planning: The End of the Beginning, Angew. Chem., Int. Ed., 2016, 55(20), 5904–5937, DOI:10.1002/anie.201506101.
- A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, Ł. Kaiser and I. Polosukhin, Attention Is All You Need, in Advances in Neural Information Processing Systems 30, ed. I. Guyon, U. V. Luxburg, S. Bengio, H. Wallach, R. Fergus, S. Vishwanathan and R. Garnett, Curran Associates, Inc., 2017, pp. 5998–6008 Search PubMed.
- C. W. Coley, R. Barzilay, T. S. Jaakkola, W. H. Green and K. F. Jensen, Prediction of Organic Reaction Outcomes Using Machine Learning, ACS Cent. Sci., 2017, 3(5), 434–443, DOI:10.1021/acscentsci.7b00064.
- M. H. S. Segler, M. Preuss and M. P. Waller, Planning Chemical Syntheses with Deep Neural Networks and Symbolic AI, Nature, 2018, 555(7698, 7698), 604–610, DOI:10.1038/nature25978.
- P. Schwaller, T. Gaudin, D. Lányi, C. Bekas and T. Laino, “Found in Translation”: Predicting Outcomes of Complex Organic Chemistry Reactions Using Neural Sequence-to-Sequence Models, Chem. Sci., 2018, 9(28), 6091–6098, 10.1039/C8SC02339E.
- A. A. Lee, Q. Yang, V. Sresht, P. Bolgar, X. Hou, J. L. Klug-McLeod and C. R. Butler, Molecular Transformer Unifies Reaction Prediction and Retrosynthesis across Pharma Chemical Space, Chem. Commun., 2019, 55(81), 12152–12155, 10.1039/C9CC05122H.
- F. Strieth-Kalthoff, F. Sandfort, M. H. S. Segler and F. Glorius, Machine Learning the Ropes: Principles, Applications and Directions in Synthetic Chemistry, Chem. Soc. Rev., 2020, 49(17), 6154–6168, 10.1039/C9CS00786E.
- S. Genheden, A. Thakkar, V. Chadimová, J.-L. Reymond, O. Engkvist and E. Bjerrum, AiZynthFinder: A Fast, Robust and Flexible Open-Source Software for Retrosynthetic Planning, J. Cheminform., 2020, 12(1), 70, DOI:10.1186/s13321-020-00472-1.
- A. Thakkar, S. Johansson, K. Jorner, D. Buttar, J.-L. Reymond and O. Engkvist, Artificial Intelligence and Automation in Computer Aided Synthesis Planning, React. Chem. Eng., 2021, 6(1), 27–51, 10.1039/D0RE00340A.
- P. B. R. Hartog, A. M. Westerlund, I. V. Tetko and S. Genheden, Investigations into the Efficiency of Computer-Aided Synthesis Planning, J. Chem. Inf. Model., 2025, 65(4), 1771–1781, DOI:10.1021/acs.jcim.4c01821.
- L. Long, R. Li and J. Zhang, Artificial Intelligence in Retrosynthesis Prediction and Its Applications in Medicinal Chemistry, J. Med. Chem., 2025, 68(3), 2333–2355, DOI:10.1021/acs.jmedchem.4c02749.
- Z. Tu, S. J. Choure, M. H. Fong, J. Roh, I. Levin, K. Yu, J. F. Joung, N. Morgan, S.-C. Li, X. Sun, H. Lin, M. Murnin, J. P. Liles, T. J. Struble, M. E. Fortunato, M. Liu, W. H. Green, K. F. Jensen and C. W. Coley, ASKCOS: Open-Source, Data-Driven Synthesis Planning, Acc. Chem. Res., 2025, 58(11), 1764–1775, DOI:10.1021/acs.accounts.5c00155.
- D. M. Lowe, Extraction of Chemical Structures and Reactions from the Literature, 2012 Search PubMed.
- I. V. Tetko, P. Karpov, R. Van Deursen and G. Godin, State-of-the-Art Augmented NLP Transformer Models for Direct and Single-Step Retrosynthesis, Nat. Commun., 2020, 11(1), 5575, DOI:10.1038/s41467-020-19266-y.
- M. E. Fortunato, C. W. Coley, B. C. Barnes and K. F. Jensen, Data Augmentation and Pretraining for Template-Based Retrosynthetic Prediction in Computer-Aided Synthesis Planning, J. Chem. Inf. Model., 2020, 60(7), 3398–3407, DOI:10.1021/acs.jcim.0c00403.
- X. Wu, Y. Zhang, J. Yu, C. Zhang, H. Qiao, Y. Wu, X. Wang, Z. Wu and H. Duan, Virtual Data Augmentation Method for Reaction Prediction, Sci. Rep., 2022, 12(1), 17098, DOI:10.1038/s41598-022-21524-6.
- X. Zhang, Y. Mo, W. Wang and Y. Yang, Retrosynthesis Prediction Enhanced by In-Silico Reaction Data Augmentation, arXiv, 2024, preprint, arXiv:2402.00086, DOI:10.48550/arXiv.2402.00086.
- D. Kreutter and J.-L. Reymond, Multistep Retrosynthesis Combining a Disconnection Aware Triple Transformer Loop with a Route Penalty Score Guided Tree Search, Chem. Sci., 2023, 14(36), 9959–9969, 10.1039/D3SC01604H.
- D. Kreutter and J.-L. Reymond, Chemoenzymatic Multistep Retrosynthesis with Transformer Loops, Chem. Sci., 2024, 15(43), 18031–18047, 10.1039/D4SC02408G.
- D. Kreutter and J.-L. Reymond, Chemoenzymatic Multistep Retrosynthesis with Transformer Loops, ChemRxiv, 2024, preprint, DOI:10.26434/chemrxiv-2024-svr99.
- G. Pesciullesi, P. Schwaller, T. Laino and J.-L. Reymond, Transfer Learning Enables the Molecular Transformer to Predict Regio- and Stereoselective Reactions on Carbohydrates, Nat. Commun., 2020, 11(1), 4874, DOI:10.1038/s41467-020-18671-7.
- D. Kreutter, P. Schwaller and J.-L. Reymond, Predicting Enzymatic Reactions with a Molecular Transformer, Chem. Sci., 2021, 12(25), 8648–8659, 10.1039/D1SC02362D.
- D. Lowe, Chemical Reactions from US Patents (1976–Sep 2016), 2017, DOI:10.6084/m9.figshare.5104873.v1.
- A. Thakkar, A. C. Vaucher, A. Byekwaso, P. Schwaller, A. Toniato and T. Laino, Unbiasing Retrosynthesis Language Models with Disconnection Prompts, ACS Cent. Sci., 2023, 9(7), 1488–1498, DOI:10.1021/acscentsci.3c00372.
- C. Kannas, A. Thakkar, E. Bjerrum and S. Genheden, Rxnutils – A Cheminformatics Python Library for Manipulating Chemical Reaction Data, ChemRxiv, 2022, preprint, DOI:10.26434/chemrxiv-2022-wt440-v2.
- MolecularAI/Reaction_utils, 2024. https://github.com/MolecularAI/reaction_utils (accessed 2024-05-28).
- rxnutils documentation—ReactionUtils 1.5.0 documentation, https://molecularai.github.io/reaction_utils/(accessed 2024-05-28).
- E. Heid, J. Liu, A. Aude and W. H. Green, Influence of Template Size, Canonicalization, and Exclusivity for Retrosynthesis and Reaction Prediction Applications, J. Chem. Inf. Model., 2022, 62(1), 16–26, DOI:10.1021/acs.jcim.1c01192.
- hesther. Hesther/Templatecorr, 2023, https://github.com/hesther/templatecorr (accessed 2024-05-28).
- P. Schwaller, B. Hoover, J.-L. Reymond, H. Strobelt and T. Laino, Extraction of Organic Chemistry Grammar from Unsupervised Learning of Chemical Reactions, Sci. Adv., 2021, 7(15), eabe4166, DOI:10.1126/sciadv.abe4166.
- D. Probst and J.-L. Reymond, Visualization of Very Large High-Dimensional Data Sets as Minimum Spanning Trees, J. Cheminform., 2020, 12(1), 12, DOI:10.1186/s13321-020-0416-x.
- D. Probst, P. Schwaller and J.-L. Reymond, Reaction Classification and Yield Prediction Using the Differential Reaction Fingerprint DRFP, Digit. Discov., 2022, 1(2), 91–97, 10.1039/D1DD00006C.
- D. Probst and J.-L. Reymond, A Probabilistic Molecular Fingerprint for Big Data Settings, J. Cheminform., 2018, 10(1), 66, DOI:10.1186/s13321-018-0321-8.
- P. Schwaller, R. Petraglia, V. Zullo, V. H. Nair, R. A. Haeuselmann, R. Pisoni, C. Bekas, A. Iuliano and T. Laino, Predicting Retrosynthetic Pathways Using Transformer-Based Models and a Hyper-Graph Exploration Strategy, Chem. Sci., 2020, 11(12), 3316–3325, 10.1039/c9sc05704h.
- G. Klein, Y. Kim, Y. Deng, J. Senellart and A. Rush, Open, N. M. T: Open-Source Toolkit for Neural Machine Translation, in Proceedings of ACL 2017, System Demonstrations, ed. M. Bansal and H. Ji, Association for Computational Linguistics, Vancouver, Canada, 2017, pp. 67–72 Search PubMed.
- OpenNMT. GitHub. https://github.com/OpenNMT (accessed 2024-05-29).
- A. Thakkar, A. Vaucher, A. Byekwaso, P. Schwaller, A. Toniato and T. Laino, Disconnection Labelled Reaction Data, 2022, DOI:10.5281/zenodo.7101695.
| This journal is © The Royal Society of Chemistry 2026 |