Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Deep learning-enabled discovery of low-melting-point ionic liquids
Gaopeng
Ren
a,
Austin M.
Mroz
 ab,
Frederik
Philippi
ab,
Frederik
Philippi
 a,
Tom
Welton
a,
Tom
Welton
 a and
Kim E.
Jelfs
a and
Kim E.
Jelfs
 *a
*a
aDepartment of Chemistry, Imperial College London, White City Campus, London, W12 0BZ, UK. E-mail: k.jelfs@imperial.ac.uk
bI-X Centre for AI in Science, Imperial College London, White City Campus, London, W12 0BZ, UK
First published on 21st January 2026
Abstract
Ionic liquids (ILs) are salts that are liquids at ambient conditions (typically below 373 K) and are known for their many unique properties, including low volatility and high thermal stability. Despite the promise of ILs, their targeted design is challenging for several reasons, including (i) the vast number of candidate ions that could be synthesised as components of an IL, (ii) the lack of predictive methods to determine what ion combinations will yield ILs with desired melting points, and (iii) experimentally known ILs possess limited chemical diversity. In this work, we present a data-driven framework for designing novel low-melting-point ILs. We model ILs as bipartite graphs and apply a link prediction algorithm to identify promising cation–anion pairs, expanding the collected IL database more than 30-fold, while prioritising low melting points. To further explore chemical space, we trained variational auto-encoders (VAEs) to generate new IL candidates through learning a latent space that enables modelling the data distribution. A thermodynamics-inspired classification model is subsequently employed to filter out ILs predicted to melt above 373 K. Finally, molecular dynamics simulations validate our approach, confirming that 18 out of 20 generated ILs have melting points below 373 K.
1 Introduction
Ionic liquids (ILs) are liquids composed entirely of ions, and of particular interest are ILs with low melting points (typically below 373 K).1 ILs often exhibit a range of unique properties, including high thermal stability, negligible vapour pressure, non-inflammability, and wide electrochemical windows.2,3 More intriguingly, by combining different cations and anions, the properties of ILs can be tailored for specific applications, making them highly versatile.4 As such, ILs have emerged as promising alternatives to conventional organic solvents.5,6 Beyond their use as solvents, ILs have also found application as functional materials in various domains, including batteries,7 pharmaceuticals,8 and catalysis.9Due to the extensive range of possible cation–anion combinations, the chemical space of ILs is vast, and therefore it would not be possible to screen all possible ILs experimentally.10 Further, the complex, and often non-linear, structure–property relationships limit the utility of design rules for informed IL design, and frequently means that experiments are performed in a trial-and-improvement manner. Computational design methods provide a tractable alternative to experimental efforts. The computational design of ILs typically involves two key components: proposing new IL systems and developing models to calculate their properties. A straightforward method for constructing IL systems is via high-throughput screening and candidate enumeration. Here, new combinations of known cations and anions are enumerated, resulting in a large number of possible ILs.11,12 To further expand the IL chemical space, some studies decompose cations and anions into cores and substituents (for example, side chains) and recombine them to create new structures.13–18 These approaches allow the creation of extensive IL libraries, which can then be screened using conventional simulations or property prediction models.
Two widely used methods for calculating IL properties are density functional theory (DFT) and molecular dynamics (MD) simulations. Although these techniques can provide accurate results, such calculations are typically computationally expensive and often unsuitable for high-throughput screening. Recently, machine learning (ML) models have emerged as an efficient alternative for IL property prediction.5,19–23 These data-driven models can learn structure–property relationships from data, and predict various IL properties, such as melting point,20,21,24 viscosity,25,26 heat capacity,27 and CO2 solubility.28,29 By combining IL construction strategies with property prediction models, there is the potential to identify ILs optimised for specific applications.
Despite the promise of property prediction models, the IL chemical space in the aforementioned workflows is limited, as ILs are constructed by combining existing cations and anions or manually combining different molecular substituents onto ions. In recent years, generative ML models have emerged as a powerful tool for exploring chemical space.30,31 The goal of generative models is to create new samples that closely resemble existing ones.32 Variational auto-encoders (VAEs) are among the most popular generative methods. In VAEs, input data is encoded into a latent space that follows a normal distribution, and the decoder reconstructs the data from the latent space. These models have been widely applied to molecular generation, with examples including ChemicalVAE,33 GraphVAE,34 and JT-VAE.35 These VAEs can efficiently learn smooth, continuous latent spaces that capture molecular features, enabling the generation of novel, diverse, and valid molecular structures. Yet, these VAEs are mainly focused on small, organic molecules, for which there are extensive datasets available. Applying VAEs to molecular materials like ILs poses significant challenges, particularly due to data scarcity. Indeed, the number of known ILs is limited. For instance, the NIST ILThermo database36 contains only ∼3000 unique ILs, far fewer than the 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 examples typically required to train robust VAE models.37 Furthermore, IL datasets that include experimental properties are even scarcer. This greatly hinders the development of robust ML models for accurate IL property prediction.
000 examples typically required to train robust VAE models.37 Furthermore, IL datasets that include experimental properties are even scarcer. This greatly hinders the development of robust ML models for accurate IL property prediction.
There are several methods that have been introduced to address the issue of data scarcity. For example, Liu et al.38,39 employed particle swarm optimization to simultaneously optimise a property prediction model and a VAE. They introduced a normative score to guide the VAE towards generating valid ILs and a property score to conditionally generate ILs with desired properties. However, this approach does not fully resolve the data scarcity issue, as no additional data or external information is introduced into the workflow. As a result, the IL chemical space to be explored will remain fairly limited. Another strategy to mitigate the data scarcity issue is transfer learning. Specifically, since large datasets of general organic compounds are available, the VAE model can be pre-trained on such datasets and then fine-tuned using the smaller IL datasets. However, Beckner et al.40 found that while transfer learning improves the generation of valid molecules, the dominance of neutral organic compounds in the training data reduces sample efficiency, as the majority of generated molecules are neutral, not the desired ions. Furthermore, neither the optimisation-based nor the transfer learning methods typically explicitly consider the melting points of generated ILs. As a result, they do not guarantee the production of ILs with low melting points, limiting practical application. In addition to post-processing and transfer learning, data augmentation is another commonly used technique to address data scarcity. For ILs, a straightforward approach is to combine all available cations and anions to expand the dataset, given the sparsity of existing cation–anion pairs (Fig. 1). However, not all of these combinations would produce ILs with low melting points. For example, the salt, NaCl, composed of Na+ as the ‘IL cation’ and Cl− as the ‘IL anion’ has a melting point up to 801 °C, which is far from that of typical ILs. Another problem with all combinations is the low training efficiency. The training examples will increase significantly, while the unique ion structures do not increase. Additionally, the generated examples will include many high-melting-point cation–anion combinations.
Here, we proposed a link prediction method to address the issue of data scarcity in using VAEs for IL generation. We formulated the task of identifying suitable cation–anion pairs as a link prediction problem, which predicts whether two nodes in a network are likely to be connected.41 In the IL scenario, this task aims to identify new cation–anion combinations that are likely to form low-melting-point ILs. Using this method, we generated over 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 new cation–anion combinations, expanding the collected database more than 30-fold. A VAE was then trained on the expanded IL database, and achieved good performance in terms of novelty, validity, uniqueness, and reconstruction accuracy. However, the generated ILs are not inherently guaranteed to have low melting points. To address this, we developed a thermodynamics-based classification model to filter out undesirable systems likely to have melting points above 373 K. Finally, we validated the expanded ILs from link prediction and the generated ILs from the VAE by performing MD simulations to calculate their melting points. The results showed that the IL samples from both link prediction and the VAE exhibited low melting points, with most below 373 K, demonstrating the effectiveness of our workflow.
000 new cation–anion combinations, expanding the collected database more than 30-fold. A VAE was then trained on the expanded IL database, and achieved good performance in terms of novelty, validity, uniqueness, and reconstruction accuracy. However, the generated ILs are not inherently guaranteed to have low melting points. To address this, we developed a thermodynamics-based classification model to filter out undesirable systems likely to have melting points above 373 K. Finally, we validated the expanded ILs from link prediction and the generated ILs from the VAE by performing MD simulations to calculate their melting points. The results showed that the IL samples from both link prediction and the VAE exhibited low melting points, with most below 373 K, demonstrating the effectiveness of our workflow.
2 Methods
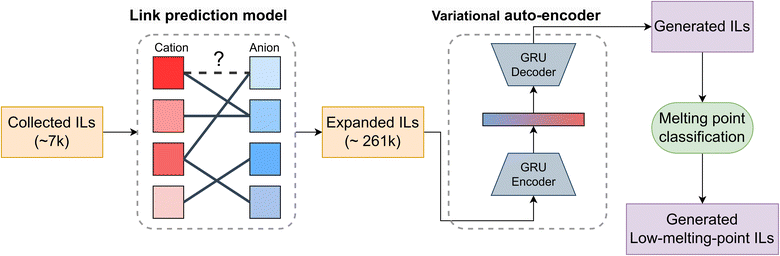
Our workflow for low-melting-point IL generation is shown in Fig. 2. First, we collected ILs from both the literature and open-source databases. These ILs were then expanded into a much larger dataset using a link prediction model, which enhanced the robustness of a VAE model for generating novel, valid, and diverse ILs. Next, a melting point classification model was employed for post-filtering. Finally, the generated ILs were validated through MD simulations. The data and code are available at https://github.com/fate1997/ILGen-overall.2.1 Data set construction
To construct a large and diverse IL dataset, we extracted IL structures from the NIST ILThermo database36 and several previous publications.42–46 All IL data points were standardised using canonical SMILES strings, and duplicates were removed. The collected database comprises 7508 unique ILs, featuring 3223 unique cations and 510 unique anions. While not all the ILs in the database are low-melting-point ILs, all IL data points were preserved to ensure a large and diverse chemical space for IL design. Moreover, as shown in the melting point distribution (Fig. S1), the database still exhibits relatively low melting points, with a mean value of 361 K, which is lower than that of typical salts. In this work, melting point considerations were addressed after the generation process.2.2 Link prediction model to expand IL chemical space
The distribution of cations and anions in the collected IL database is illustrated in Fig. 1. Each data point indicates a cation–anion combination present in the collected database. From this figure, it is evident that the collected database exhibits significant sparsity, with only a few cations and anions appearing frequently. Among anions, Br− and [NTf2]− are the most common, while for cations, [bmim]+, [P6,6,6,14]+, [emim]+, and [Ch]+ are the most common (the structures of these common ions are shown in Fig. 1). Nearly half of the collected database (3100 out of 7507) contains one of these common ions, while other cations and anions are heavily underexplored. To address this imbalance, we developed a link prediction model that aims to find the most plausible cation–anion combinations.Link prediction aims to identify potential links based on existing ones. In the context of ILs, a link could be interpreted as a cation–anion combination that can form an IL. As shown in Fig. 2, we constructed a bipartite graph using the cations and anions in the collected IL database as heterogeneous nodes. We used extended-connectivity fingerprints (ECFP) descriptors47 as the initial node embeddings. Then, a graph neural network (GNN), GraphSAGE,48 was applied to update the node embeddings iteratively. The computation process of GNNs consists of two main phases: (i) an aggregation phase to gather information from node neighbours, and (ii) an update phase to update node embeddings according to the gathered information. In the GraphSAGE model, the node embeddings are updated through:
 | (1) |
 are node embeddings with d dimensions.
are node embeddings with d dimensions.
By stacking multiple GNN layers, nodes can incorporate information from increasingly distant neighbours, enriching their representations. To determine the likelihood of a cation–anion pair forming an IL, we computed the dot product similarity of their updated embeddings, followed by a Sigmoid activation function to output a probability:
| sij = σ(xiTxj), | (2) |
Before training, we split the existing links into training, validation, and test sets with a ratio of 85![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5
5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10. Negative samples were generated by randomly selecting non-existent links. To ensure a balanced dataset, the number of negative samples is equivalent to the number of existing links. Note that the non-existent links may include combinations capable of forming ILs, but this probability is likely low, given the very small ratio of existing ILs to non-existent ILs (0.4%). During training, we optimised the GNN using cross-entropy loss between the predicted probabilities and the true link labels. We set the number of training epochs to 100, and used the Adam49 optimiser with a learning rate of 0.01 to optimise the model parameters. To expand the collected IL database as much as possible, we conducted multiple iterative training rounds. In each round, the newly predicted links were added to the existing links, progressively increasing the database.
10. Negative samples were generated by randomly selecting non-existent links. To ensure a balanced dataset, the number of negative samples is equivalent to the number of existing links. Note that the non-existent links may include combinations capable of forming ILs, but this probability is likely low, given the very small ratio of existing ILs to non-existent ILs (0.4%). During training, we optimised the GNN using cross-entropy loss between the predicted probabilities and the true link labels. We set the number of training epochs to 100, and used the Adam49 optimiser with a learning rate of 0.01 to optimise the model parameters. To expand the collected IL database as much as possible, we conducted multiple iterative training rounds. In each round, the newly predicted links were added to the existing links, progressively increasing the database.
2.3 VAE generation
We trained a VAE to generate novel ILs. A VAE is composed of an encoder and a decoder. The encoder processes the input data and maps it to a latent representation, which follows a normal distribution. The encoder will first calculate a mean µ and a variance σ2, and then calculate the latent representation by using the reparameterization trick:| z = µ + σ·ε, | (3) |
 . The decoder then reconstructs the input data from the latent representation. After training the VAE, we can generate new data by sampling random noise from a standard normal distribution and passing it through the decoder.
. The decoder then reconstructs the input data from the latent representation. After training the VAE, we can generate new data by sampling random noise from a standard normal distribution and passing it through the decoder.
Here, we applied gated recurrent units (GRUs) for both the encoder and the decoder. SMILES strings are used to represent the cations and anions that comprise the ILs. First, we built a vocabulary by collecting the unique characters from the SMILES strings in the database and adding special tokens (i.e., start, end, and padding). Each SMILES string can be encoded to a series of integers that represent the indices of characters in the vocabulary. These integer sequences are padded to a maximum length of 256 to ensure uniform input size and are further projected to high-dimensional embeddings by an embedding layer. The GRU encoder takes these embeddings as inputs. Here, the encoder and decoder each consist of three GRU layers, and the dimension of the latent representations is set to 128.
The training objective of a VAE is to maximise the evidence lower bound (ELBO) which consists of two components: the reconstruction loss 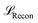 to regulate the recovered data to be close to the initial input data, and the Kullback–Leibler (KL) divergence to encourage the latent representation follows a standard normal distribution. To stabilise the training process (specifically to avoid the KL vanishing), we adopted the loss function from Fu et al.:50
to regulate the recovered data to be close to the initial input data, and the Kullback–Leibler (KL) divergence to encourage the latent representation follows a standard normal distribution. To stabilise the training process (specifically to avoid the KL vanishing), we adopted the loss function from Fu et al.:50
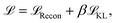 | (4) |
2.4 Melting point classification
Since neither the link prediction model nor the VAE prediction model explicitly considers the melting point, the generated ILs may not necessarily be low-melting-point ILs. To develop a post-filtering model based on melting point information, we used the IL melting point database from Venkatraman et al.51. Notably, that database was compiled from diverse sources, and some data points may be approximate. However, since our study here focuses solely on melting point classification, no additional data cleaning was performed. This database contains 2208 ILs labelled by their melting points, and was used to train a linear classification model capable of distinguishing between low-melting-point ILs and higher melting systems. In this model, we first labelled the data in the IL melting point database as 1 if the melting point is below 373 K and 0 otherwise. Instead of relying on standard descriptors like ECFP, we carefully selected three specific descriptors known to have a strong correlation with the melting point of ILs. These three descriptors are the lattice Gibbs energy,52 the solvent-accessible surface area (SASA), and molecular flexibility based on functional group counts.53 These descriptors are closely related to the Gibbs free energy of fusion and play an important role in determining the melting points of IL. Further details can be found in Section S2 of the SI.2.5 MD validation
We used MD simulations to validate the melting point classification model and verify that the novel ILs possess melting points lower than 373 K. We modified the workflow from Karu et al.54 to achieve a highly automated MD workflow. In the original workflow, the initial configuration of IL systems was generated using Packmol.55 Then the solid phase structures were generated with the help of eight predefined coulombic potential wells and six NVT simulations, corresponding to two types of ionic lattices and four lattice vector ratios. Annealing simulations were then performed from 175 K to 475 K using GROMACS software.56 Finally, the melting point was obtained by a regression analysis of the diffusion coefficient, which was estimated from the mean square displacement for every consecutive 10 ps. While this duration is insufficient to obtain accurate diffusion coefficients, this approach can reveal the relative change in the diffusion coefficient during annealing, and the melting point can be identified as the intersection of the fitted lines. In our revised workflow, the input data consists of a series of generated SMILES strings. First, we converted the SMILES string into a 3D structure using xTB calculations.57 Next, the input files for GROMACS were generated using the Sobtop package.58 To enhance the method's performance in the high-melting-point region, we increased the equilibration time before beginning the annealing simulations and raised the final temperature to 600 K. We used the General AMBER Force Field (GAFF)59 for the MD workflow, as it has previously been successfully applied to ILs.60 During annealing simulations, approximate self-diffusion coefficients (D) were recorded and analysed by plotting ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D versus 1/T. Assuming Arrhenius behaviour with a constant diffusion activation energy within each phase,61–63 two linear regimes corresponding to the solid and liquid states were identified. The intersection of these regimes was taken as the melting temperature.64 Further details of the MD validation process are provided in Section S3 of the SI. The code for the MD simulation is available at https://github.com/fate1997/MD4IL.
D versus 1/T. Assuming Arrhenius behaviour with a constant diffusion activation energy within each phase,61–63 two linear regimes corresponding to the solid and liquid states were identified. The intersection of these regimes was taken as the melting temperature.64 Further details of the MD validation process are provided in Section S3 of the SI. The code for the MD simulation is available at https://github.com/fate1997/MD4IL.
3 Results and discussion
3.1 Link prediction
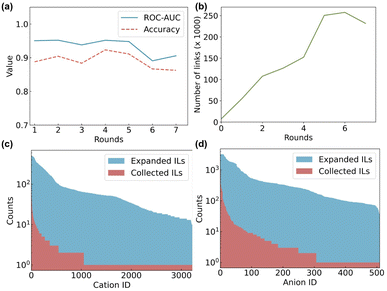
To progressively grow the collected IL dataset, we conducted seven iterations of training and evaluation for the link prediction task. The performance of the link prediction model was measured by accuracy and the area under the receiver operating characteristic curve (ROC-AUC), which indicates how well the model distinguishes between positive and negative links, with higher values meaning better performance. Performance metrics and the number of predicted links in each round are shown in Fig. 3a and b, respectively. The model demonstrated consistently strong performance during the first five rounds, achieving an accuracy of approximately 0.90 and an ROC-AUC of around 0.95. There is a plateau in the number of predicted links after the fifth round, Fig. 3b. This slight decrease in performance suggests that five rounds are optimal for expanding the IL dataset; this balances both high predictive performance and affords a substantial increase in predicted links. The collected database contained only 7508 ILs, but after five rounds of link prediction, this number expanded significantly to 262![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 076. To further analyse the dataset expansion, we visualised the frequency of occurrence of cations and anions in both the collected and expanded IL datasets, Fig. 3c and d. In the collected IL database, the distributions of cations and anions were highly imbalanced, with only a few species being dominant. After expansion, the distributions became more uniform while still maintaining a trend similar to the collected dataset.
076. To further analyse the dataset expansion, we visualised the frequency of occurrence of cations and anions in both the collected and expanded IL datasets, Fig. 3c and d. In the collected IL database, the distributions of cations and anions were highly imbalanced, with only a few species being dominant. After expansion, the distributions became more uniform while still maintaining a trend similar to the collected dataset.
To further validate the performance of the link prediction model, we randomly selected 10 ILs predicted by the model and conducted MD simulations to calculate their melting points. As shown in Fig. 4 and S6, 6 out of the 10 new ILs had predicted melting points less than 373 K, while a further 3 had melting points near 373 K (within 15 K deviation). Only 1 of the 10 ILs had a relatively high melting point (560 K). These results demonstrate that the link prediction approach is an effective method for identifying potential cation–anion combinations with low melting points, even without a filtering step to check the melting point prediction. This effectiveness can be attributed to the ability of GNNs to capture and transfer knowledge across different nodes, enabling the link prediction model to recognise intrinsic patterns that determine whether a cation–anion combination can form an IL. Finally, the expanded IL database (virtual database) contains a large number of ILs with low melting points, which is essential for training an effective generative model for IL generation.
3.2 VAE generation
In this work, we represent an IL as an SMILES string in the format “{cation}.{anion}”. The expanded ILs serve as inputs, which are expressed as SMILES strings. The performance of generative models can be assessed using four key metrics: validity, uniqueness, novelty, and reconstruction accuracy. Validity measures the percentage of sampled molecules that are syntactically correct, meaning they follow proper SMILES grammar and can be parsed by RDKit.65 Novelty is defined as the percentage of valid molecules that do not appear in the training dataset. Uniqueness refers to the proportion of valid molecules that are distinct, without duplicates in the sampled set. In this study, we randomly sampled 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ILs for evaluation. Unlike the above metrics, reconstruction accuracy is assessed based on the test set. It represents the percentage of SMILES strings from the test set that the model can successfully reconstruct. To evaluate the reconstruction accuracy of the VAE model, we excluded 10% of the total data points for use as a separate test set.
000 ILs for evaluation. Unlike the above metrics, reconstruction accuracy is assessed based on the test set. It represents the percentage of SMILES strings from the test set that the model can successfully reconstruct. To evaluate the reconstruction accuracy of the VAE model, we excluded 10% of the total data points for use as a separate test set.
Here, we evaluate the generation performance of the VAE model using three different input datasets: the collected IL database (VAE-collected), the expanded IL database generated through link prediction (VAE-link), and a dataset containing all possible cation–anion combinations (VAE-all). The results are summarised in Table 1. The VAE-collected model demonstrates high novelty but suffers from low validity and reconstruction accuracy. This limitation arises because of the small dataset, making it challenging for the VAE model to learn the correct SMILES grammar effectively. In contrast, the VAE-all model exhibits strong performance in terms of validity, uniqueness, and reconstruction accuracy, but has very low novelty, indicating that most generated ILs are already present in the training dataset. Among the three models, VAE-link performs well across all metrics, showing strong validity and reconstruction accuracy. The high validity and reconstruction accuracy of the VAE-link model suggest that this model effectively learns the correct SMILES grammar, enabling the generation of valid molecular structures. However, the novelty of the VAE-link is slightly lower than that of the VAE-collected, primarily because the expanded IL database does not introduce new ion structures. Despite this minor decrease in novelty, the VAE-link model excels in sample efficiency, defined as the number of novel valid ILs generated among all the sampled SMILES strings. It significantly outperforms both the VAE-collected and the VAE-all in this regard, demonstrating its effectiveness in generating diverse and novel IL structures.
| Model | Data points | Validity | Unique | Novelty | Recon. |
|---|---|---|---|---|---|
| VAE-collected | 7507 | 31% | 95% | 89% | 11% |
| VAE-all | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 640 640![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 507 507 |
97% | 100% | 10% | 93% |
| VAE-link | 262![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 076 076 |
84% | 99% | 78% | 90% |
3.3 Post filter and MD validation
To maintain the diversity of generated ILs from VAE-link, we did not restrict the training examples to low-melting-point ILs. Instead, we applied a post-filtering step using a melting point classification model. This model was trained on the IL melting point database,51 with 80% of the data used for training and the remaining 20% for testing. We found that a simple linear model performs well on this task when using carefully selected descriptors, including lattice Gibbs energy, flexibility, and SASA, which are essential in determining melting points (as shown in Section S2 in the SI). The melting point classification model achieves an accuracy of 81%, a recall of 87%, and a precision of 84%. This performance is comparable to, and in some cases better than, previously reported ML-based classification models.51 These results indicate strong generalisability, likely due to the simplicity of the model and its grounding in physical principles. This generalisability is crucial for effectively filtering the generated ILs.Using the melting point classification model, we divided the generated ILs from the VAE-link model into two categories: generated low-melting-point ILs and generated high-melting systems. To compare the distribution of generated ILs with the collected ILs, we selected 2000 novel generated ILs and 2000 ILs from the collected database.51 We then computed the ECFP features for both sets and applied t-SNE to reduce the dimensionality for visualisation. As shown in Fig. 5a, the chemical space of the generated low-melting-point ILs largely overlaps with that of the collected ones, indicating that the generated low-melting-point ILs are more diverse, while still covering the original distribution. In contrast, the chemical space visualisation of higher melting systems (Fig. 5b) shows that the number of generated higher melting systems (175) is significantly smaller than that of the collected higher melting systems (875). Furthermore, 90% of generated ILs are classified as low-melting-point ILs. These results suggest that the expanded IL dataset enhances the diversity of generated ILs while ensuring a preference for those with low melting points.
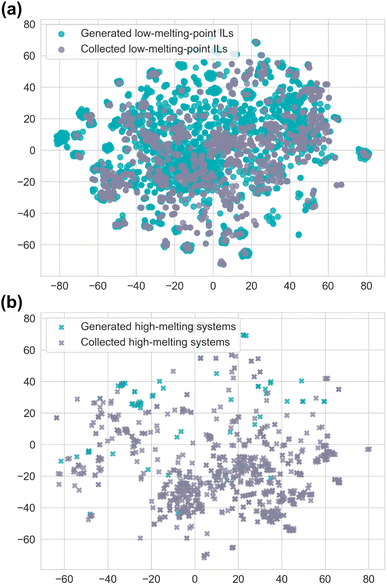 | ||
| Fig. 5 t-SNE plots of (a) low-melting-point ILs (<373 K) and (b) high-melting systems (>373 K) in both the generated and collected IL datasets. | ||
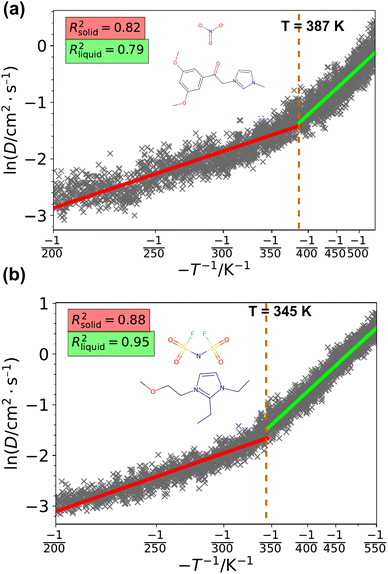
We employed MD simulations to estimate the melting points of novel ILs. To validate the MD workflow, we tested it on 20 ILs with experimentally known melting points (ranging from 246 K to 421 K) and found it aligns well with the experimental values (Fig. S5). We then extended the simulations to additional ILs, including newly expanded and generated ones. Representative figures illustrating the determination of melting points are presented in Fig. 6. The melting points were identified as the intersection of the fitted solid-phase and liquid-phase lines. All the plots to determine the melting point are shown in Section S4 and S5 in the SI.
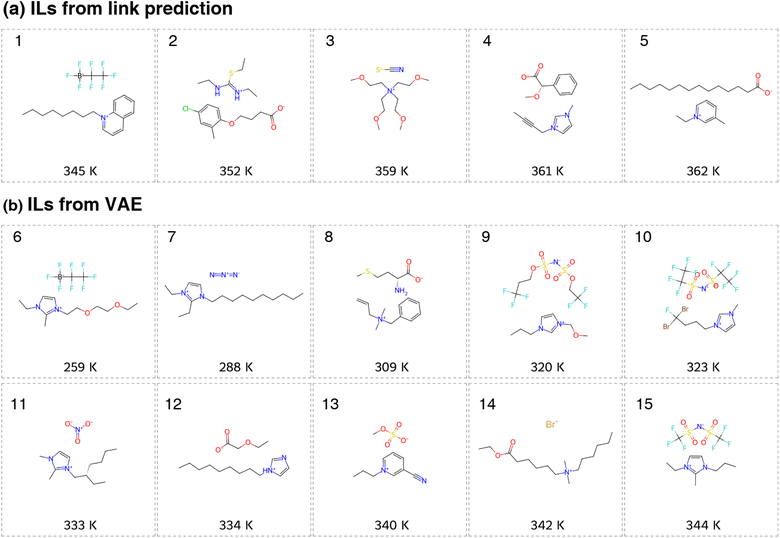
We randomly selected 20 generated ILs from the VAE-link model and estimated their melting points using the MD workflow. The structures and predicted melting points of 10 generated low-melting-point ILs are shown in Fig. 4, with more examples provided in Fig. S9. Our analysis shows that the generated ILs exhibit low melting points, with 18 out of 20 classified as low-melting-point ILs (melting points below 373 K). The remaining 2 ILs have melting points (385 K) slightly above this threshold, but do not exceed it significantly. The average melting point of the 20 generated ILs is 339 K, which is lower than the average melting point of the IL melting point database (361 K). Several factors contribute to these results. First, the link prediction model effectively identifies combinations of cation–anion that are likely to have low melting points. Second, the VAE model generates structures similar to those in the input dataset, ensuring that the produced cations and anions resemble those found in the collected IL database. Third, the melting point classification model accurately differentiates between low-melting-point ILs and higher melting systems by leveraging thermodynamic knowledge.
The low melting points of the newly designed ILs (both from link prediction and VAE) can be rationalised based on the intermolecular interactions between ion pairs. In particular, localised and directional interactions, such as hydrogen bonds, tend to lower melting points significantly.66 For example, ILs composed of imidazolium cations and amide anions (IL 9, 10, 15 in Fig. 4) exhibit extensive hydrogen-bond networks involving C–H and N–H bonds, which explains their low melting points.67 Hydrogen bonding is also present in IL examples 1, 3–8, 11–14, also contributing to reduced melting points.68–71 Another key factor is the anion size: larger anions weaken ion–ion interactions, which also leads to lower melting points.52,72 Large anions are observed in IL examples 1, 4–6, 8–10, and 15, further explaining their reduced melting behaviour.
4 Conclusion
We present a deep learning-enabled workflow for discovering novel low-melting-point ILs. The associated code can be found in https://github.com/fate1997/ILGen-overall. To address data scarcity in the IL field, we leveraged link prediction and developed a GNN model to identify likely low-melting-point cation–anion combinations based on existing ILs. Our results demonstrate that the link prediction model effectively identifies such pairs, expanding the collected IL database by 30-fold through iterative predictions, and finally obtained 262![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 076 ILs. The expanded dataset enables the effective training of a VAE with improved sample efficiency. The generated ILs are further screened using a melting point filter, to test whether their melting points are predicted to be below 373 K. Finally, MD simulations validated that 90% of generated ILs meet this criterion. We believe our strategy can also facilitate the discovery of other molecular mixtures, such as deep eutectic solvents. However, there are still limitations to our approach; the expansion of the database has not increased the diversity of the individual molecular components, which may restrict exploration of the IL chemical space. Additionally, synthetic feasibility, an important factor for the practical application of low-melting-point ILs, was not explicitly considered in this study. Future work will focus on broadening the range of ion structures to enhance their diversity, while considering synthetic accessibility and the prediction of other properties, beyond the melting point, of relevance to the IL community. In parallel, we will pursue the synthesis of selected predicted novel ILs and experimentally validate their melting points.
076 ILs. The expanded dataset enables the effective training of a VAE with improved sample efficiency. The generated ILs are further screened using a melting point filter, to test whether their melting points are predicted to be below 373 K. Finally, MD simulations validated that 90% of generated ILs meet this criterion. We believe our strategy can also facilitate the discovery of other molecular mixtures, such as deep eutectic solvents. However, there are still limitations to our approach; the expansion of the database has not increased the diversity of the individual molecular components, which may restrict exploration of the IL chemical space. Additionally, synthetic feasibility, an important factor for the practical application of low-melting-point ILs, was not explicitly considered in this study. Future work will focus on broadening the range of ion structures to enhance their diversity, while considering synthetic accessibility and the prediction of other properties, beyond the melting point, of relevance to the IL community. In parallel, we will pursue the synthesis of selected predicted novel ILs and experimentally validate their melting points.
Author contributions
G. R. designed the workflow, conducted experiments, and analysed the results. A. M. M. contributed to the project design and execution. F. P. and T. W. contributed to project conceptualisation and expertise in ILs. K. E. J. developed the research question and supervised the project. G. R. drafted the manuscript, and all authors contributed to the final version.Conflicts of interest
There are no conflicts of interest to declare.Data availability
All the data, code and models used to generate the results are available in the Github repository, https://github.com/fate1997/ILGen-overall and Zenodo: https://doi.org/10.5281/zenodo.17957179. The molecular dynamics workflow for melting point estimation is available in the Github repository, https://github.com/fate1997/MD4IL and Zenodo: https://doi.org/10.5281/zenodo.15736993.Supplementary information (SI) is available: melting point distribution in the database, melting point classification model, MD for melting point estimation, generated ILs from link prediction, generated ILs from VAE, and melting point distribution of generated ILs. See DOI: https://doi.org/10.1039/d5dd00282f.
Acknowledgements
G. R. and F. P. acknowledge Imperial College London for funding through the President's PhD Scholarships. K. E. J. acknowledges the AI for Chemistry: AIchemy hub for funding (EPSRC grant EP/Y028775/1 and EP/Y028759/1). A. M. M. is supported by the Eric and Wendy Schmidt AI in Science Postdoctoral Fellowship, a Schmidt Sciences program.References
- Z. Lei, B. Chen, Y.-M. Koo and D. R. MacFarlane, Chem. Rev., 2017, 117, 6633–6635 CrossRef PubMed
.
- R. D. Rogers and K. R. Seddon, Science, 2003, 302, 792–793 CrossRef PubMed
.
- G. Kaur, H. Kumar and M. Singla, J. Mol. Liq., 2022, 351, 118556 CrossRef CAS
.
- Z. Song, J. Chen, J. Cheng, G. Chen and Z. Qi, Chem. Rev., 2024, 124, 248–317 CrossRef CAS PubMed
.
- S. Koutsoukos, F. Philippi, F. Malaret and T. Welton, Chem. Sci., 2021, 12, 6820–6843 RSC
.
- S. N. A. Shafie, N. A. H. Md Nordin, S. M. Racha, M. R. Bilad, M. H. D. Othman, N. Misdan, J. Jaafar, Z. A. Putra and M. D. H. Wirzal, J. Mol. Liq., 2022, 358, 119192 CrossRef CAS
.
- X. Tang, S. Lv, K. Jiang, G. Zhou and X. Liu, J. Power Sources, 2022, 542, 231792 CrossRef CAS
.
- W. Zhuang, K. Hachem, D. Bokov, M. Javed Ansari and A. Taghvaie Nakhjiri, J. Mol. Liq., 2022, 349, 118145 CrossRef CAS
.
- B. Karimi, M. Tavakolian, M. Akbari and F. Mansouri, ChemCatChem, 2018, 10, 3173–3205 CrossRef CAS
.
- D. C. Weis and D. R. MacFarlane, Aust. J. Chem., 2012, 65, 1478–1486 CrossRef CAS
.
- K. Paduszyński, M. Królikowski, M. Zawadzki and P. Orzeł, ACS Sustainable Chem. Eng., 2017, 5, 9032–9042 CrossRef
.
- M. Taheri, R. Zhu, G. Yu and Z. Lei, Chem. Eng. Sci., 2021, 230, 116199 CrossRef CAS
.
- H. Matsuda, H. Yamamoto, K. Kurihara and K. Tochigi, Fluid Phase Equilib., 2007, 261, 434–443 CrossRef CAS
.
- S. E. McLeese, J. C. Eslick, N. J. Hoffmann, A. M. Scurto and K. V. Camarda, Comput. Chem. Eng., 2010, 34, 1476–1480 CrossRef CAS
.
- J. Wang, Z. Song, H. Cheng, L. Chen, L. Deng and Z. Qi, ACS Sustainable Chem. Eng., 2018, 6, 12025–12035 CrossRef CAS
.
- V. Venkatraman, S. Evjen, K. C. Lethesh, J. J. Raj, H. K. Knuutila and A. Fiksdahl, Sustainable Energy Fuels, 2019, 3, 2798–2808 RSC
.
- J. Wang, X. Tang, Z. Qi, T. Xu, T. Zou, Y. Bie, D. Wang and Y. Liu, ACS Sustainable Chem. Eng., 2022, 10, 2248–2261 CrossRef CAS
.
- C. H. Huang and S. T. Lin, J. Chem. Inf. Model., 2023, 63, 7711–7728 CrossRef CAS PubMed
.
- W. Wang, T. Yang, W. H. Harris and R. Gómez-Bombarelli, Chem. Commun., 2020, 56, 8920–8923 RSC
.
- K. Low, R. Kobayashi and E. I. Izgorodina, J. Chem. Phys., 2020, 153, 104101 Search PubMed
.
- H. Feng, L. Qin, B. Zhang and J. Zhou, ACS Omega, 2024, 9, 16016–16025 CrossRef CAS PubMed
.
- J. G. Rittig, K. Ben Hicham, A. M. Schweidtmann, M. Dahmen and A. Mitsos, Comput. Chem. Eng., 2023, 171, 108153 CrossRef CAS
.
- J. Sun, Y. Sato, Y. Sakai and Y. Kansha, J. Cleaner Prod., 2023, 414, 137695 CrossRef CAS
.
- Z. Dai, L. Wang, X. Lu and X. Ji, Green Energy Environ., 2024, 9, 1802–1811 CrossRef CAS
.
- H. Moslehi, S. M. Hosseini, M. Pierantozzi, M. M. Alavianmehr and B. Haghighi, J. Mol. Liq., 2023, 383, 122004 CrossRef CAS
.
- Y. Ding, M. Chen, C. Guo, P. Zhang and J. Wang, J. Mol. Liq., 2021, 326, 115212 Search PubMed
.
- S. Li, H. He and Y. Wang, Ind. Eng. Chem. Res., 2024, 63, 15257–15265 Search PubMed
.
- Y. Tian, X. Wang, Y. Liu and W. Hu, J. Mol. Liq., 2023, 383, 122066 CrossRef CAS
.
- F. Yan, M. Lartey, K. Jariwala, S. Bowser, K. Damodaran, E. Albenze, D. R. Luebke, H. B. Nulwala, B. Smit and M. Haranczyk, J. Phys. Chem. B, 2014, 118(47), 13609–13620 CrossRef CAS PubMed
.
- Y. Du, A. R. Jamasb, J. Guo, T. Fu, C. Harris, Y. Wang, C. Duan, P. Liò, P. Schwaller and T. L. Blundell, Nat. Mach. Intell., 2024, 6, 589–604 CrossRef
.
- J. Zhou, A. Mroz and K. E. Jelfs, Digital Discovery, 2023, 2, 1925–1936 RSC
.
- F. Faez, Y. Ommi, M. S. Baghshah and H. R. Rabiee, IEEE Access, 2021, 9, 106675–106702 Search PubMed
.
- R. Gómez-Bombarelli, J. N. Wei, D. Duvenaud, J. M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T. D. Hirzel, R. P. Adams and A. Aspuru-Guzik, ACS Cent. Sci., 2018, 4, 268–276 CrossRef PubMed
.
-
M. Simonovsky and N. Komodakis, Artificial Neural Networks and Machine Learning – ICANN 2018, Cham, 2018, pp. 412–422 Search PubMed
.
-
W. Jin, R. Barzilay and T. Jaakkola, Proceedings of the 35th International Conference on Machine Learning, 2018, pp. 2323–2332 Search PubMed
.
- Q. Dong, C. D. Muzny, A. Kazakov, V. Diky, J. W. Magee, J. A. Widegren, R. D. Chirico, K. N. Marsh and M. Frenkel, J. Chem. Eng. Data, 2007, 52, 1151–1159 CrossRef CAS
.
- M. A. Skinnider, R. G. Stacey, D. S. Wishart and L. J. Foster, Nat. Mach. Intell., 2021, 3, 759–770 Search PubMed
.
- X. Liu, J. Chu, Z. Zhang and M. He, Mater. Des., 2022, 220, 110888 Search PubMed
.
- X. Liu, J. Chu, S. Huang, A. Li, S. Wang and M. He, ACS Sustainable Chem. Eng., 2023, 11, 8978–8987 Search PubMed
.
- W. Beckner, C. Ashraf, J. Lee, D. A. Beck and J. Pfaendtner, J. Phys. Chem. B, 2020, 124, 8347–8357 Search PubMed
.
- D. Liben-Nowell and J. Kleinberg, J. Am. Soc. Inf. Sci., 2007, 58, 1019–1031 Search PubMed
.
- K. Paduszyński, Ind. Eng. Chem. Res., 2019, 58, 5322–5338 CrossRef
.
- K. Paduszyński, Ind. Eng. Chem. Res., 2019, 58, 17049–17066 CrossRef
.
- K. Paduszyński, Ind. Eng. Chem. Res., 2021, 60, 5705–5720 CrossRef
.
- Z. Song, H. Shi, X. Zhang and T. Zhou, Chem. Eng. Sci., 2020, 223, 115752 Search PubMed
.
- J. Han, M. Li, N. Tian, C. Liu, Y. Zhang, Z. Ji and X. Sun, Fluid Phase Equilib., 2023, 565, 113675 CrossRef CAS
.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef CAS PubMed
.
-
W. L. Hamilton, R. Ying and J. Leskovec, Proceedings of the 31st International Conference on Neural Information Processing Systems, Red Hook, NY, USA, 2017, pp. 1025–1035 Search PubMed
.
-
D. P. Kingma and J. Ba, Proceedings of the 3rd International Conference on Learning Representations, ICLR, 2015 Search PubMed
.
-
H. Fu, C. Li, X. Liu, J. Gao, A. Celikyilmaz and L. Carin, Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (Long and Short Papers), Minneapolis, Minnesota, 2019, pp. 240–250 Search PubMed
.
- V. Venkatraman, S. Evjen, H. K. Knuutila, A. Fiksdahl and B. K. Alsberg, J. Mol. Liq., 2018, 264, 318–326 CrossRef CAS
.
- I. Krossing, J. M. Slattery, C. Daguenet, P. J. Dyson, A. Oleinikova and H. Weingärtner, J. Am. Chem. Soc., 2006, 128, 13427–13434 CrossRef CAS PubMed
.
- F. Philippi, D. Rauber, O. Palumbo, K. Goloviznina, J. McDaniel, D. Pugh, S. Suarez, C. C. Fraenza, A. Padua, C. W. M. Kay and T. Welton, Chem. Sci., 2022, 13, 9176–9190 RSC
.
- K. Karu, F. Elhi, K. Põhako-Esko and V. Ivaništšev, Appl. Sci., 2019, 9, 5367 CrossRef CAS
.
- L. Martínez, R. Andrade, E. G. Birgin and J. M. Martínez, J. Comput. Chem., 2009, 30, 2157–2164 CrossRef PubMed
.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef
.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 CrossRef CAS PubMed
.
- Sobtop, http://sobereva.com/soft/Sobtop, Online, accessed 26-November-2023.
- J. Wang, W. Wang, P. A. Kollman and D. A. Case, J. Mol. Graphics Modell., 2006, 25, 247–260 CrossRef CAS PubMed
.
- K. G. Sprenger, V. W. Jaeger and J. Pfaendtner, J. Phys. Chem. B, 2015, 119, 5882–5895 CrossRef CAS PubMed
.
-
D. M. Ruthven, Principles of adsorption and adsorption processes, John Wiley & Sons, 1984 Search PubMed
.
- S. Berkowicz and F. Perakis, Phys. Chem. Chem. Phys., 2021, 23, 25490–25499 RSC
.
-
R. B. Bird, W. E. Stewart and E. N. Lightfoot, Transport phenomena, John Wiley & Sons, New York, 2nd edn, 2002 Search PubMed
.
- R. Lin, P.-L. Taberna, S. Fantini, V. Presser, C. R. Pérez, F. Malbosc, N. L. Rupesinghe, K. B. K. Teo, Y. Gogotsi and P. Simon, J. Phys. Chem. Lett., 2011, 2, 2396–2401 CrossRef CAS
.
- RDKit: Open-source cheminformatics, http://www.rdkit.org, Online, accessed 11-April-2024.
- T. Peppel, C. Roth, K. Fumino, D. Paschek, M. Köckerling and R. Ludwig, Angew. Chem., Int. Ed., 2011, 50, 6661–6665 CrossRef CAS PubMed
.
- K. Fumino, T. Peppel, M. Geppert-Rybczyńska, D. H. Zaitsau, J. K. Lehmann, S. P. Verevkin, M. Köckerling and R. Ludwig, Phys. Chem. Chem. Phys., 2011, 13, 14064–14075 Search PubMed
.
- K. Dong, S. Zhang, D. Wang and X. Yao, J. Phys. Chem. A, 2006, 110, 9775–9782 CrossRef CAS PubMed
.
- Y. Ma, Y. Liu, H. Su, L. Wang and J. Zhang, J. Mol. Liq., 2018, 255, 176–184 CrossRef CAS
.
- S. Schneider, T. Hawkins, M. Rosander, J. Mills, A. Brand, L. Hudgens, G. Warmoth and A. Vij, Inorg. Chem., 2008, 47, 3617–3624 CrossRef CAS PubMed
.
- P. A. Hunt, C. R. Ashworth and R. P. Matthews, Chem. Soc. Rev., 2015, 44, 1257–1288 RSC
.
- H. Markusson, J.-P. Belières, P. Johansson, C. A. Angell and P. Jacobsson, J. Phys. Chem. A, 2007, 111, 8717–8723 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |