Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A fully differentiable pore network for digital reconstruction of porous media
Michael
McKague
 ,
Mohammad
Mehrnia
,
Mohammad
Mehrnia
 ,
Mohammad Amin
Sadeghi
,
Mohammad Amin
Sadeghi
 and
Jeff
Gostick
and
Jeff
Gostick
 *
*
Department of Chemical Engineering, University of Waterloo, ON, Canada. E-mail: jgostick@uwaterloo.ca
First published on 18th February 2026
Abstract
Pore network models are useful for efficiently studying transport processes in porous materials at the pore scale. However, constructing accurate pore network representations is not always straight forward as it involves either complicated image processing or tedious calibration of network properties to experimental data. While network extraction tools are much more accessible today, access to high resolution X-ray tomography images is not nearly as widespread. Researchers of porous materials without access to high resolution microtomography images must therefore rely on calibrating a pore network by manually adjusting pore sizes and constantly checking that it satisfies the properties of the actual material. This paper presents a new strategy for fitting a regular lattice-based pore network to experimental data by using automatic differentiation for gradient descent optimization. The optimization was demonstrated on a data set of carbonate, sandstone, and sandpack materials for which images and experimental information were both available. The optimization of a 103 pore network took on average 21 and a half minutes on a GPU and the resulting networks matched the porosimetry curves and permeability in all directions of the actual material very closely. The optimization was then tested on real experimental data in which a final loss of 9.7 × 10−4 was achieved. In addition, a workflow was developed for obtaining a stochastic network of arbitrary size from the fitted network. This required a description of the highly specific spatial arrangement of pore sizes found by the optimization, for which a Gaussian Process model was trained.
1 Introduction
Pore networks are widely used to study the interplay between pore structure and transport processes. Their main appeal is computational efficiency obtained by discretizing the pore space into a representative network with pore bodies as nodes connected by throats as edges. However, simplifying a real material with such an abstract representation can result in inaccurate predictions if not done correctly and therefore special attention must be given when generating pore networks to ensure accurate results.1–3Networks can be obtained either by network extraction from images4 or by fitting to experimental data.5,6 The latter approach was used by McKague et al., who were able to predict the performance of a capacitive electrode by fitting the parameters of a Weibull pore size distribution to Mercury Intrusion Porosimetry (MIP) data.5 Similarly, Gostick et al. modelled the gas diffusion layer of a fuel cell as a cubic network of pores and determined its geometric properties by fitting to both porosimetry and gas permeability data of the actual material.6 Network extractions are particularly appealing and have gained much interest from researchers because of how they map a pore network directly to the void space of an entire three-dimensional image.7 The various approaches for network extraction can be categorized as either medial axis,8–11 maximal ball,12,13 or watershed segmentation.14 Today, network extractions are commonplace and the development of open-source network extractions, such as those available in  ,15 have made these tools widely accessible to researchers.
,15 have made these tools widely accessible to researchers.
Despite these advancements, there are still three challenges to the use of network extractions. Firstly, and foremostly, suitable images are time consuming and costly to obtain, and the equipment is not widely available. Secondly, the images are generally a small subsection of a large sample, so are not necessarily representative, and moreover they represent just a single sample so multiple realizations are not possible. Lastly, network extractions are often excessive as they require complicated image processing steps while a simple cubic lattice network is often sufficient to perform simulations or calculations. For instance, one can use simple cubic networks to study the structure–performance relationship in devices such as fuel cells,16,17 flow batteries,18,19 or capacitive deionization5 to name a few.
On the other hand, fitting a pore network to experimental data is not as trivial as it may appear. Firstly, while researchers may use the bundle of tubes model to obtain pore sizes directly from capillary pressure data,20,21 it is well known that these sizes are the size of the smallest constriction.22 In pore network models, the smallest constriction is the throat not the pore. Therefore, without any information about the actual pore–throat size relationship, researchers must use a trial-and-error approach to guess the pore sizes of the network until the simulated porosimetry curve matches the experimental data. Secondly, to construct a more accurate network it is necessary to match effective properties of the real material that may include permeability or effective diffusivity.6,23 While this already adds a layer of complexity when choosing a pore size distribution, it is especially complicated knowing that transport properties can be largely affected by the spatial correlation of pore sizes24 which is difficult to know without having access to high resolution images. Ultimately, trial-and-error is not feasible, and one must turn to computerized optimization techniques.25
Optimization methods can be classified as either gradient or non-gradient descent. The later includes genetic algorithms which have been successfully used for the optimization of porous materials for specific applications.26–31 The drawback to non-gradient descent methods is the large number of different pore network realizations and evaluations that are required every iteration. Gradient descent, on the other hand, works by optimizing the properties of a single network rather than having multiple constructions and searching to find the best one. To the best of our knowledge, however, gradient descent, has not been used in the context of pore network modeling. To calculate the gradient numerically using traditional approaches requires applying small perturbations to the properties of a single pore followed by model evaluations for each pore in the network. This explains why Sharqawy intentionally avoided gradient-descent optimization when he selected the use of non-linear programming methods that do not require taking the gradient.32
Automatic Differentiation (AD) is a compelling alternative to numerical differentiation because of how it solves the gradient exactly and efficiently for high dimensional spaces.33 It works on the principle that any numerical computation can be stored in memory as a computational graph of nodes each representing an elementary operation of known derivatives.34 The drawback of this approach is that it becomes quite complicated to code, but thanks to its integration with machine learning libraries such as  ,35 AD has become a mainstream tool. Crucially, AD has a reverse mode option that is especially powerful for optimization problems because it can compute the partial derivative for all input variables with respect to one output variable in a single pass.33 In the case of pore networks, there are many pore and/or throat properties (i.e. input variables) that control just a handful of possible macroscopic properties (i.e. output variables) and therefore, automatic differentiation, particularly the reverse mode, is a highly promising route for generating statistical pore networks. Note that it would be possible to use the optimization capabilities of the JAX library to fit the parameters of a given statistical pore (and throat) size distribution, but in this work, we focused on adjusting individual pore diameters directly since this is a unique capability provided by the auto-differentiation ability of JAX. Moreover, this approach does not require any assumption about the true pore size distribution, which is not something normally known without access to volumetric images.
,35 AD has become a mainstream tool. Crucially, AD has a reverse mode option that is especially powerful for optimization problems because it can compute the partial derivative for all input variables with respect to one output variable in a single pass.33 In the case of pore networks, there are many pore and/or throat properties (i.e. input variables) that control just a handful of possible macroscopic properties (i.e. output variables) and therefore, automatic differentiation, particularly the reverse mode, is a highly promising route for generating statistical pore networks. Note that it would be possible to use the optimization capabilities of the JAX library to fit the parameters of a given statistical pore (and throat) size distribution, but in this work, we focused on adjusting individual pore diameters directly since this is a unique capability provided by the auto-differentiation ability of JAX. Moreover, this approach does not require any assumption about the true pore size distribution, which is not something normally known without access to volumetric images.
In this work, automatic differentiation is used for the first time to construct pore networks that match MIP and permeability data. This paper starts with a discussion of the methodology on how the network is optimized, the development of differentiable porosimetry and flow simulators, as well as an analysis of the computational performance. Next, the optimization is tested using data simulated from a set of 13 images that includes sandstone, carbonate, and sandpack materials followed by using real experimental data of a Berea sandstone obtained from the work by Churchel et al.36 Finally, a Gaussian Process model was trained to capture the spatial correlations of a fitted network, enabling the construction of larger networks with similar properties.
2 Optimization strategy
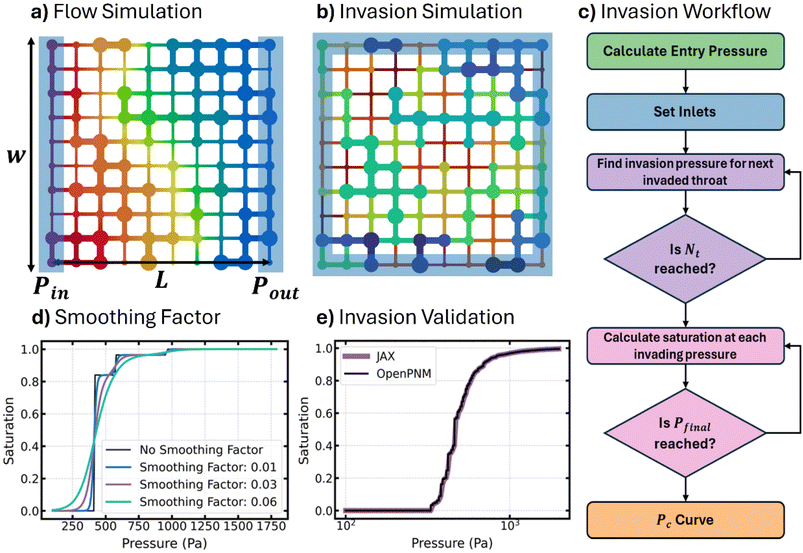
When constructing a pore network from experimental data, it is common to make some basic assumptions about the network prior to optimization.5–7 These assumptions often involve the network's size, shape, connectivity, or geometry of the pores/throats to name a few. Therefore, we started with a network based on some basic assumptions, and this network will be improved over the course of optimization. The network's connectivity was assumed to be a cubic lattice where each pore has a coordination number of 6 except for the sides, edges, and corners of the network, which have coordination numbers of 5, 4, and 3 respectively. Arranging pores in this way meant that the spacing between each pore was fixed across the entire network but could be used as a single free parameter from one optimization to the next. The size of the network was chosen by balancing computational efficiency with statistically representative sample size. Fig. S1 of the SI shows how a 103 network is large enough to statistically represent the pore size distribution of a Berea sandstone. Therefore, a moderately sized network of 103 pores was chosen for optimization. The geometry of the pores was assumed to be spheres connected by cylindrical throats. Fig. 1a shows the pore network with its assumed spheres and cylinders geometry. A closer look at the spheres and cylinders geometry model shows how the throat aspect ratio (ξ), which will be discussed further, is the ratio between the throat diameter and the diameter of the smallest connected pore.After deciding on network structure and geometry, the optimization procedure was applied. Fig. 1b shows the optimization process starting with choosing an initial guess for all parameter weights (w0), choosing the learning rate (lr) for gradient descent, and setting the maximum number of steps to take (nmax) prior to completing the optimization. The so called “parameter weights” are the geometric properties of the network assuming that all topological properties (i.e. coordination number) remain fixed. The two geometric properties that make up parameter weights are the pore diameter and throat aspect ratio. Together, these properties can be combined into a single array, wn, with a length equal to the number of pores plus the number of throats, containing both pore and throat size information. After setting up the parameters for optimization, the initial guess of parameter weights is passed to the loss function and automatic differentiation is used to calculate the gradient of the loss function with respect to each weight. It should be mentioned that since we have many parameters but a single output, using the reverse AD mode is perfectly suited as it calculates the entire gradient at a computational cost on the same order as just one loss function evaluation. The loss function involves running both a flow and an invasion simulation before calculating the combined loss from simulation results and the target data. Sections 2.1 and 2.2 describe the development of both flow and invasion simulations in more detail.
The loss function, which quantifies how closely the current network predicts the target values, was composed of the following terms. First, the saturation loss, Sloss, is the sum of squared errors (SSE) between the model saturation, S, and the target saturation, Starget, from experimental data. The saturation loss was calculated using eqn (1) where n is the number of observations. Since the model saturation and the target saturation may vary in size, linear interpolation was used to evaluate the model saturation at the same invasion pressures as the target saturation.
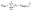 | (1) |
Second, the permeability loss, Kloss, is the mean squared error (MSE) between the modelled permeability, K, and the target or experimental permeability, Ktarget. In this work, the permeability in all three spatial directions was considered and therefore, the average loss of all three directions was used. Eqn (2) shows how the permeability loss was calculated, where n is 3 for each of the three spatial coordinates.
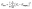 | (2) |
The loss function can also be used to impose constraints onto the final network design by adding penalty terms. In the present case the following two constraints were imposed:
1. No two pore bodies can overlap and,
2. Each throat must have a diameter that is smaller than the minimum diameter of it's two connected pores.37
These two constraints were accomplished by setting pore diameters to a scale factor (<1) of the lattice spacing and second, by optimizing throat aspect ratios instead of throat sizes directly. In this way, if all weights (i.e. pore diameters and aspect ratios) are maintained between 0 and 1, it is guaranteed that no pore bodies overlap since each pore cannot exceed one unit of spacing, and no throat diameter will ever exceed the minimum diameter of its connected pores because the aspect ratio is always less than one. To impose this constraint, eqn (3) was used to calculate the penalty loss where n is the total number of weights (i.e. the number of pores plus the number of throats). Notice that no penalty is added if the weight is greater than 0.01 and less than 0.99 to maintain a small safety margin away from these physical limits.
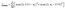 | (3) |
Finally, the loss function, f, was defined as a weighted sum of all loss terms including the permeability, saturation, and penalty loss terms just described, yielding a scalar output. Eqn (4) shows the resulting loss function which includes additional weights, α, β, and γ, for the saturation, permeability, and penalty loss terms respectively. These weights were chosen to give the saturation and permeability loss terms a similar order of magnitude to start while a large weight was assigned to the penalty loss term to ensure strict adherence to the rule that weights should be between 0 and 1. Specifically, α was chosen to be 10, β was chosen to be 1, and γ was chosen to be 1000.
| f = αSloss + βKloss + γfpenalty | (4) |
After defining the loss function, gradient descent was used to minimize the loss function given by eqn (4). The  package
package  , which is a numerical differential equation solver developed for working with
, which is a numerical differential equation solver developed for working with  , was used to perform gradient descent. Please see the SI for a more detailed description and code snippet on how exactly
, was used to perform gradient descent. Please see the SI for a more detailed description and code snippet on how exactly  was used for gradient descent optimization.
was used for gradient descent optimization.
2.1 Flow
Pore network models solve flow problems by calculating a hydraulic conductance for each pore–throat–pore conduit based on assumed geometrical shapes (in this case spheres and cylinders) and then performing a mass balance around each pore in the network. The resulting mass balance equations are solved simultaneously as a set of linear algebraic equations. The mass balance is calculated around each pore i in the network using eqn (5) assuming incompressible flow at steady state:38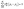 | (5) |
The hydraulic conductance, Ghij, is calculated assuming a resistor-in-series model of pore–throat elements with well-defined analytical solutions. Eqn (6) shows how to calculate the conductance for an entire pore–throat–pore conduit connecting pores i and j.
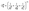 | (6) |
Eqn (7) is used to calculate the hydraulic conductance of a conduit element with arbitrary shape where λhi is the hydraulic size factor of element i and µ is the dynamic viscosity of the fluid. To calculate the hydraulic size factor, eqn (8) is used generally for any shape of varying cross-sectional area, Ai(x), assuming negligible inertial loss.39 The specific polar moment of inertia is given by  which is calculated by
which is calculated by 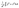 . The complete analytical solution for a spheres and cylinders geometry is somewhat involved, and therefore it is included in the SI.
. The complete analytical solution for a spheres and cylinders geometry is somewhat involved, and therefore it is included in the SI.
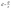 | (7) |
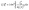 | (8) |
 was used to solve the system of equations using its implementation of a sparse linear solver which should be mentioned was under the experimental module in
was used to solve the system of equations using its implementation of a sparse linear solver which should be mentioned was under the experimental module in  at the time of this work. Therefore,
at the time of this work. Therefore,  's implementation was tested and results showed good agreement in terms of accuracy to
's implementation was tested and results showed good agreement in terms of accuracy to  's sparse linear solver. Fig. 2a shows the solution of a flow simulation on a 10 by 10 network of pores using
's sparse linear solver. Fig. 2a shows the solution of a flow simulation on a 10 by 10 network of pores using  . The solution for a similar but larger 103 network was solved for using an applied pressure difference of 1Pa using
. The solution for a similar but larger 103 network was solved for using an applied pressure difference of 1Pa using  and
and  solvers. The solutions of both solvers were compared and the average difference was 3.02 × 10−10.
solvers. The solutions of both solvers were compared and the average difference was 3.02 × 10−10.
After solving the flow problem, the permeability of the pore network was calculated using Darcy's law (eqn (9)) where Q is the flow rate (m3 s−1) and ΔP is the pressure drop (Pa). Fig. 2a shows a 2D schematic of the domain with the boundary pores highlighted in blue. The boundary pores on the left-hand side are assigned some inlet pressure, Pin, while the boundary pores on the right-hand side are assigned some outlet pressure, Pout. The length of the domain, L, was taken as the distance from the centre of the inlet and outlet pores while the width and height (not shown in Fig. 2a) spanned the entire domain and was used to calculate the cross-sectional area, A.
 | (9) |
The permeability can be calculated for each of the three spatial coordinates by solving the flow problem three times, once for each direction, so the above procedure was repeated using the front-back and top-bottom faces.
2.2 Porosimetry
Porosimetry is often used to characterize the pore size distribution of porous materials experimentally.40 It works by injecting a non-wetting liquid (e.g. mercury) into a porous material and recording the volume invaded at each discrete pressure step. This procedure can be simulated on a pore network quite easily using percolation algorithms.41 Existing code (e.g. in )42 is not auto-differentiable and therefore, we wrote a custom percolation algorithm in
)42 is not auto-differentiable and therefore, we wrote a custom percolation algorithm in  as described below.
as described below.
Fig. 2c is a flow chart showing the procedure that was used to simulate invasion. First, the entry pressure of each throat was calculated using the Washburn equation,43 given by eqn (10), where σ is the surface tension of the invading fluid, θ is the contact angle, and Dt is the throat diameter. The properties of mercury were assumed for the invasion such that the surface tension and contact angle are 0.4791 N m−1 and 140° respectively. The Washburn equation is commonly used when simulating MIP data44 and as such is suitable for highly non-wetting invading phases in most materials. It relates the entry pressure of a throat to its diameter by assuming the throat is cylindrical which, as discussed earlier in Section 2 is consistent with this work.
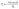 | (10) |
It should be noted that wettability distributions, such as those discussed by Garfi et al. and Foroughi et al.,45,46 could also be considered if capillary pressure data were available using invading fluids other than mercury. In this case, the mercury intrusion data would provide information about the pore and throat sizes, while the additional data could be used in a second optimization process to adjust the distribution of contact angles used in eqn (10) while holding the pore and throat sizes constant.
After calculating the throat entry pressure, the inlet pores were selected. Fig. 2b shows an example of an invasion simulation on a 10 by 10 network of pores where the inlet pores are highlighted in blue. The inlet pores represent the initial invading front from which invasion occurs. Starting with the uninvaded throat that has the lowest entry pressure (or largest diameter), it is set to invaded and its neighboring pore is added to the invading front. This process, which occurs one throat at a time, is known as invasion percolation and is performed by iterating through each throat in the network as shown by the purple loop in Fig. 2b. Porosimetry is more akin to an ordinary percolation process however, but it is possible to obtain an ordinary percolation result from an invasion percolation simulation if we also note the maximum pressure that has been reached at each step in the invasion process. This way it is possible to find all pores and throats which will be invaded at a given pressure step by thresholding according to this value.47
Once the invasion pressure, pinv, for each throat was known, the pink loop shown in Fig. 2b is used to calculate the saturation for a predefined set of invading pressures. At each pressure, p, the saturation is calculated by adding the total volume of pores and throats that are invaded and dividing by the total volume of pores and throats, as shown in eqn (11):
 | (11) |
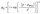 | (12) |
The sigmoid function attempts to model a stepwise function as a smooth curve from 0 to 1 but it's smoothness can be controlled by the smoothing factor, β. A higher smoothing factor decreases the steepness while a smaller smoothing factor increases the steepness in the transition from 0 to 1. Fig. 2d shows the effect of the smoothing factor on the saturation curve for a simple network of four pores attached sequentially from largest to smallest. Observe how increasing the smoothing factor decreases the rapid change in saturation that occurs during invasion.
In the present work we focused on modifying the quasi-static displacement algorithm so that porosimetry experiments could be simulated in a differentiable way. It might be feasible to use dynamic displacement algorithms such as those discussed by Joekar-Niasar and Hassanizadeh,48 which are potentially differentiable by default. It should be stressed however, that the quasi-static simulations are much more computationally efficient, yet they already represent the slowest step in the current workflow.
Finally, after writing a fully differentiable invasion simulator in  , the simulation was validated by comparing to
, the simulation was validated by comparing to  .42Fig. 2e shows the comparison of the newly developed simulation in
.42Fig. 2e shows the comparison of the newly developed simulation in  (purple) to the porosimetry simulation module currently available in
(purple) to the porosimetry simulation module currently available in  version 3.5.0 (black) for a 103 network of pores on a cubic lattice. For the comparison, no smoothing factor was used, and the resulting sum of squared errors was negligibly small. Therefore, it was concluded that our
version 3.5.0 (black) for a 103 network of pores on a cubic lattice. For the comparison, no smoothing factor was used, and the resulting sum of squared errors was negligibly small. Therefore, it was concluded that our  version of a porosimetry simulator was valid for this work.
version of a porosimetry simulator was valid for this work.
2.3 Computational efficiency
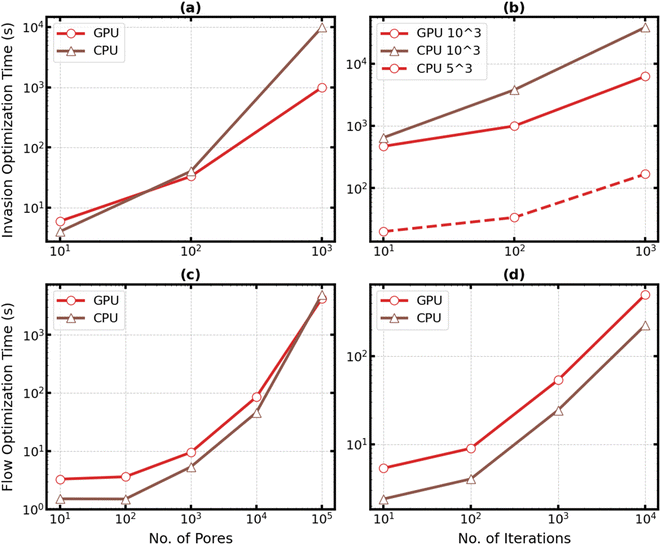
Next, we evaluated the computational efficiency of the optimizer by measuring the time required to optimize saturation and permeability losses separately. These measurements were performed on both CPU and GPU devices. The CPU used was an Intel Core i5-1135G7 processor rated at 2.40 GHz while the GPU used was Nvidia's Quadro RTX 8000 with 48 GB GDDR6 memory. The optimization time depends on both the network size and the number of iterations; therefore, we examined the effect of each parameter on the total optimization time.Fig. 3 shows the time it took to optimize flow and invasion problems. Here, plots (a) and (b) show the time measured to optimize invasion while plots (c) and (d) show the time measured to optimize flow. Plots (a) and (c) show the effect of network size while plots (b) and (d) show the effect of the number of iterations on computation time. When studying the effect of network size, a hundred iterations were used, while, when studying the effect of the number of iterations, a network with 103 pores was used.
After studying the optimization speed and plotting the results, we were able to make several observations. First, we observed that the time it took to optimize the flow problem was roughly a hundred times faster than the time it took to optimize the invasion problem. In fact, it took just 9.5 seconds to optimize the flow problem compared to 992 seconds (or 16 and a half minutes) to optimize the invasion problem for a 103 network and 100 iterations. Second, increasing the number of iterations initially causes only a moderate increase in computation time. In fact, increasing the number of iterations from 10 to 100 increased the optimization time by just 2X. This happens because there is significant overhead cost of compiling the code including the computational graph necessary for automatic differentiation. It is important to mention that while  uses Accelerated Linear Algebra (XLA) to compile efficient machine code at runtime, XLA is well known to slow down compilation. Finally, the GPU was leveraged to improve computational efficiency. For a 103 network, a speed up of 10 times was observed compared to the same operation on the CPU. However, the GPU did not speed up the computational performance in all cases, particularly when optimizing the flow problem. In fact, the GPU did not start to show any signs of improvement until using a network with 105 pores. This is consistent with what we can expect for sparse linear solvers.49–51 However, as mentioned previously, optimizing the flow problem was computationally much more efficient than optimizing the invasion problem, and therefore, the GPU was used in the rest of this work for all network optimizations.
uses Accelerated Linear Algebra (XLA) to compile efficient machine code at runtime, XLA is well known to slow down compilation. Finally, the GPU was leveraged to improve computational efficiency. For a 103 network, a speed up of 10 times was observed compared to the same operation on the CPU. However, the GPU did not speed up the computational performance in all cases, particularly when optimizing the flow problem. In fact, the GPU did not start to show any signs of improvement until using a network with 105 pores. This is consistent with what we can expect for sparse linear solvers.49–51 However, as mentioned previously, optimizing the flow problem was computationally much more efficient than optimizing the invasion problem, and therefore, the GPU was used in the rest of this work for all network optimizations.
3 Application to real data
3.1 Data set
A data set of images from Dong and Blunt12 were used to test the optimization. The data set included 13 different rock samples including 1 Berea, 2 carbonate, 9 sandstone, and 1 sandpack. The SIcontains detailed information on all the samples in the data set but, for the sake of brevity, we will focus primarily on three of the samples, mainly Berea, carbonate 2 (C2), and sandpack 1 (A1) materials, which were all structurally quite different from each other. Fig. 4 shows digital renderings of the three samples from the data set that will be the primary highlights of this work. The Berea, C2, and A1 samples had image sizes of 4003, 4003, and 3003 voxels with corresponding resolutions of 5.35 µm, 5.35 µm, and 3.85 µm, respectively. The porosity of each of the samples were 19.6, 16.8, and 42.9 percent, in the same order.Simulations were performed directly on the images in the data set to obtain the experimental data we might expect for each of the samples. To start, the permeabilities were measured on each of the samples in the x, y, and z directions using the Lattice Boltzmann Method (LBM). The LBM permeability values recorded here were retrieved from the work by Yi et al.52 but were confirmed to be accurate by performing our own LBM simulations using the MPLBM-UT package.53 Second, porosimetry data was obtained by simulating mercury invasion on the extracted networks using the watershed algorithm. Fig. 4 shows the porosimetry and permeability data determined this way for the Berea, C2, and A1 samples.
It was important to ensure that our image-based simulations resulted in reasonably accurate results with respect to actual experimental data. To confirm our simulation results, we relied on the Berea sample which has been well studied in the literature.20,21,36,54 One such experimental study by Churchel et al. was quite comprehensive as it showed that the permeability and porosimetry data can vary widely from one sample of Berea to another.36 While this work did not have experimental data with the exact characteristics of our Berea sample, this work did identify a relationship between breakthrough pressure and permeability that we used to confirm our permeability-porosimetry data pair. Fig. S3 of the SI shows how our simulated data compares exceptionally well to the empirical relationship by Churchel et al. when we used the equivalent diameter to simulate mercury invasion. Lastly, it is worth mentioning that a real porosimeter can operate over a large range of pressures finding multiple pore size distributions across different length scales while pore network models are often fitted to just one scale. Therefore, truncating the porosimetry data prior to optimization may be necessary to fit the pore network appropriately. We demonstrate this in Section 3.3 when we apply our optimization to real experimental data.
While the drawback of using image-based simulations is obvious here in that we are not using real experimental data, the advantage is that we can use the pore sizes from the images for comparison to the fitted network after optimization. To find the pore sizes of the image, a network was extracted from each of the samples using the SNOW algorithm.14 Finding the actual pore sizes in this way provides insights into the optimizers performance that would otherwise be unavailable without access to images. Fig. 4 shows the extracted network overlaid onto the digital renderings for Berea, C2, and A1 samples.
3.2 Application to data set
The data set of images and their corresponding simulated data was used by the optimizer to construct pore networks with matching flow and invasion properties. This is where the power of automatic differentiation was leveraged, by using AD to calculate the gradient for optimization. This required a number of parameters to be predefined including an initial guess, learning rate, number of iterations, pore-to-pore spacing, and smoothing factor (as introduced in Section 2.2).Table 1 summarizes the optimized parameters (i.e. pore diameter and throat aspect ratio) along with their initial guess and range. First, the initial guess for pore diameter is assumed to follow a Weibull distribution with shape parameter k and scale parameter λ, obtained from the bundle of tubes model. The pore diameters are allowed to range anywhere from zero to lattice spacing represented by Lc in the table. The lattice spacing was selected as the largest pore size obtained from the bundle of tubes model. This ensures that all sizes of the capillary pressure curve can be modelled while preventing pores from overlapping. Second, the initial guess for throat aspect ratio is assumed to be normally distributed with a mean of 0.4 and a standard deviation of 0.05. These distribution parameters were chosen arbitrarily as it was assumed that we have no other prior knowledge to inform a better initial guess. The reader is encouraged to read the discussion on the effect of the initial guess in the SI. The throat aspect ratio was allowed to range from 0 to 1.
Next, hyperparameters for gradient descent were selected by balancing numerical stability and speed. To determine an appropriately sized learning rate, we looked at a distribution of gradients (see Fig. S10 in SI) and saw that a learning rate of 1 would make reasonably sized adjustments to optimized parameters upon each iteration. Then, by monitoring the loss upon each successive iteration (see Fig. S14 in SI) we selected 300 iterations as both convergent and efficient. It was also found that gradient clipping was useful for preventing extreme changes in optimized parameters from one iteration to the next. With a learning rate of 1, gradients larger than 0.1 represent at least a 10% change in pore or throat sizes. Such large updates during a single iteration were observed to push parameters outside the physically meaningful training range [0, 1]. Therefore, while the magnitude of most gradients is less than 0.1 (see Fig. S10 in SI), gradient clipping at −0.1 and 0.1 was used to avoid excessively large negative or positive gradients. Finally, the smoothing factor was selected. The smoothing factor was selected by studying its effect on the final loss. Fig. S11 in the SI shows the effect of smoothing factor on the final loss for the Berea sample. A smoothing factor of 0.4 was selected because it minimized the final loss. Note that significant changes in the smoothing factor (approximately ±25%) resulted in small variations of the final loss except for small smoothing factors where gradient estimation becomes increasingly difficult. Once these hyperparameters were set they were left unchanged for all 13 samples in the data set to demonstrate their ranging applicability.
Fig. 5 shows the results after optimizing the geometry of three different pore networks to fit data for (a) Berea, (b) C2, and (c) A1 samples. The resulting permeabilities and saturation curves from the initial guess or bundle of tubes model are shown in blue while the target data is presented in green. It is important to draw attention to just how inaccurate the bundle of tubes model is at estimating pore sizes. The initial values of the loss function (eqn (4)) for the Berea, C2, and A1 samples were 2.19, 1.05, and 2.47 respectively. After performing gradient descent-based optimization, pore and throat sizes were found that compared well to the target data. See how for each of the samples, Berea, C2, and A1, that the purple saturation curve matches the target saturation very well, and likewise for flow where the permeability after optimization matches exceptionally well in all three spatial coordinates. Quantitatively speaking, the final loss for Berea, C2, and A1 samples are now 4.5 × 10−4, 3.3 × 10−4, and 1.0 × 10−3 respectively. As for computational speed, the optimization took on average just 21 and a half minutes for 300 iterations on a single RTX8000 GPU. The reader is referred to the SI where Fig. S4–S6 show a complete set of results for all samples in the data set. As far as performance, the results for all 13 samples were comparable to the results shown here with an average loss of 9.2 × 10−4.
While the objective of the fitted networks is to match MIP and anisotropic permeability data, the availability of images means we can compare the fitted pore sizes to the pore size distribution obtained from network extraction. The bottom row of Fig. 5 shows the resulting pore size distributions of the fitted network (purple), network extraction (blue), and the bundle of tubes model (blue line). In all cases the fitted networks have pores size distributions which are the same range as both the bundle of tubes and extracted network, but the shapes of the distribution disagree. For instance the Berea and S5 samples both have more large pores than the other two distributions.
With regards to shape, look at the Berea sample. The fitted pore size distribution is multi-modal and has many pores that are larger than the pore sizes obtained by network extraction. This informs us that the guess for throat aspect ratio is too small. Therefore, we can suspect that the resulting pore size distribution is largely impacted by the initial guess. To test this, a new initial guess for throat aspect ratio was tried that had a Weibull distribution of shape 5 and scale 1. Fig. S13 in the SI shows how the resulting pore size distribution has fewer large pores. Therefore, an accurate initial guess is important. Unfortunately, without access to images or other informed prior knowledge, it is impossible to know a good initial guess and therefore, it may be helpful to augment the initial guess with more readily available SEM images or other prior knowledge.
Some exceptions to pore sizes landing outside of the expected range can be explained. First, at the small end of the range, the network extraction is constrained by the image resolution as the smallest pore diameter it can detect is on the order of a few voxels, while the optimizer is free to fit diameters as small as it may without invoking a penalty as mentioned in Section 2. Second, at the upper end of the range, the fitted sizes are constrained by the spacing of the network while there is effectively no limitation to the upper end of pore sizes obtained from an extracted network.
To conclude, while the optimizer fits a network with pore sizes in the range of the actual sizes, the shape of the distribution can vary widely depending on the starting guess. To remedy this it would be possible to include the deviation from a given statistical pore (and throat) size distribution in the loss function, but in this work, we focused on adjusting individual pore diameters directly since this is a unique capability provided by the auto-differentiation ability of JAX. This has the added benefit of incorporating spatial correlations into the network automatically, which is essential for materials with anisotropic permeability coefficients. Moreover, the present approach does not require any assumption about the true pore size distribution, which is not something normally known. Probably the most robust way to constrain the pore size distribution is to include additional physical information in the loss function such as tortuosity and porosity of the network, both of which can be obtained experimentally. This is outside the scope of the present work however, which focuses on demonstrating the viability of auto-differentiation to construct pore networks.
3.3 Application to experimental data
Finally, the optimizer is tested using real experimental data. The experimental data for a Berea sandstone was retrieved from the work by Churchel et al.36 to demonstrate. The Berea 5 sample was selected with a porosity of 26.1% and permeability of 1168mD. This sample was selected because it represents roughly average characteristics of a Berea sandstone. Fig. 6 shows the results before and after fitting the network to experimental data. In Fig. 6a, the raw porosimetry data (black) is compared to the target porosimetry data (green) that was obtained after pre-processing. Pre-processing is necessary because pore networks can only model one scale at a time when in fact real porous materials may have pore sizes across multiple scales. In the literature, dual networks are used to model a material's entire range of pore sizes.55–57 Here, the data was pre-processed by first truncating at a maximum capillary pressure corresponding to a minimum pore size of 5.3 µm and, on the lower end, truncating at a minimum capillary pressure corresponding to a maximum pore size of 68.1 µm. Second, the saturation data was normalized from 0 to 1 because the experimental data did not exceed 85% of the total pore volume. While this is likely due to trapping or isolated pore regions, the pore network model is perfectly connected and assumed to operate at perfect vacuum (i.e. no trapping of wetting phase), and therefore, a correction was made to normalize the data from 0 to 1. Fig. 6b and c show the results of fitting a network to the experimental data. In the case of the experimental data by Churchel et al. the permeability in only one direction was measured and so the permeability in all directions was fitted to that value. It took 20 minutes and 48 seconds to achieve a final loss of 9.7 × 10−4. All parameters used for optimization including learning rate, smoothing factor, number of iterations, etc. are the same as what was used in Section 3.2. Fig. 6d shows a final look at the network fitted to real experimental data. | ||
| Fig. 6 A pore network is constructed from real experimental data of a Berea sandstone made available by Churchel et al.36 Plot (a) shows the real experimental data that was truncated prior to optimization while plots (b) and (c) show the porosimetry and permeability data before and after fitting the network. Image (d) visualizes the fitted network. | ||
3.4 Generating a stochastic network
In the previous section, we showed how gradient descent optimization can be used to fit the geometrical properties of a pore network to available data. The computational demands of the optimization process dictated that the process was done on a relatively small network. It is of interest however, to generate other realizations, especially larger ones, from the fitted geometric properties. However, as we know from past literature24 and as we will show in this section, the pore size distribution is not enough for generating new realizations with the same flow and invasion properties of the network. The reason being is that the geometrical properties of the fitted pore network can be correlated spatially and therefore, generating other accurate realizations relies on learning these spatial correlations along with matching the pore and throat size distributions. Therefore, in this section we present an approach that uses the geometrical properties obtained from fitting a smallish network, in this case a 103 network of pores, to a stochastic network of any size or shape. The developed approach relies on Gaussian kernel density estimator (KDE) for estimating the probability density function of pore size and aspect ratio distributions, sampling from those distributions, and training a Gaussian Process model to predict those properties based on their spatial location.Gaussian KDE is a non-parametric method for estimating the probability density function of a random variable (e.g. pore size) based on a finite set of samples.58 Unlike histograms, which bin data into discrete intervals, KDE provides a continuous estimate of the probability density function (pdf). Gaussian KDE was chosen for this work because of its ability to fit a pdf to a multimodal distribution similar to what was observed in Section 3.2. It works by using a kernel density estimator (i.e. KDE). In general, for a sample of n observations {xi}i=1n, the kernel density estimate of the probability density function at point x, ![[p with combining circumflex]](https://www.rsc.org/images/entities/i_char_0070_0302.gif) (x), is given by eqn (13) where n is the number of observations, h is the bandwidth controlling smoothness, and K is the kernel function.
(x), is given by eqn (13) where n is the number of observations, h is the bandwidth controlling smoothness, and K is the kernel function.
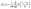 | (13) |
Selection of the kernel function, K, is important for determining the shape of the “bumps” centred at each xi point. In this work, we use the gaussian kernel given by eqn (14) where u is the input to the kernel function, in this case,  .
.
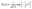 | (14) |
Meanwhile, Gaussian Process or GP is a non-parametric Bayesian approach to regression, modeling distributions over functions.59 The underlying assumption of GP is that function values have a joint multivariate gaussian distribution. The prior prediction of a GP is defined as f(x) ∼ GP(m(x), k(x, x′)) where m(x) is the mean function, often set to zero, and k(x, x′) is the covariance (kernel) function. The choice of kernel strongly impacts the shape of the functions the GP model predicts. In this work, we chose a Radial Basis Function (RBF) kernel given by eqn (15) where l is the length scale and σ2 is the signal variance.
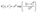 | (15) |
It is common to combine kernels together by multiplying or adding them because each kernel enforces a particular structure of the modelled function. In this work, we combined a constant kernel with the radial basis function and used regression to find all the hyperparameters that best fit our GP model.
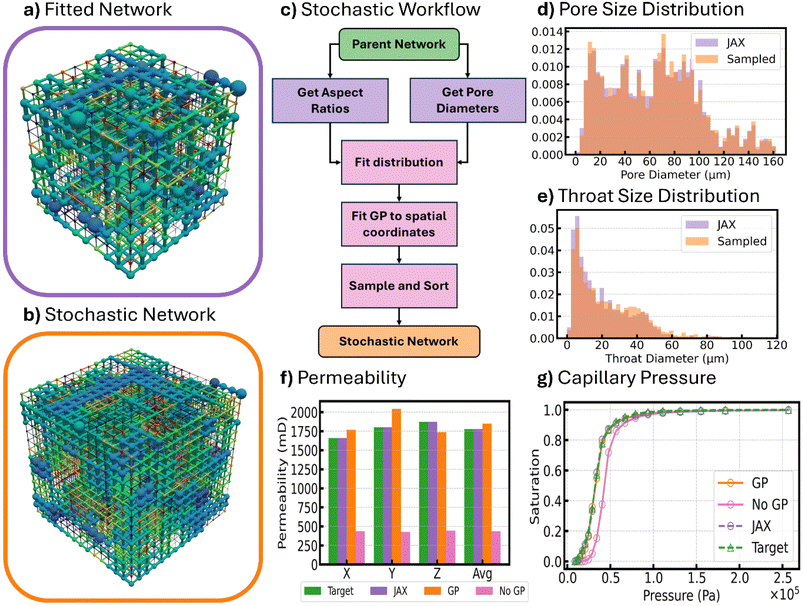
This process of using Gaussian KDE and Gaussian Process to generate a new network of arbitrary size was attempted for the Berea, C2, and A1 samples in the data set. Fig. 7 shows the results of obtaining a scaled-up network with 153 pores from the geometric properties and their spatial correlations of the parent network with 103 pores that was previously fitted to experimental data. The parent network resulting from gradient descent optimization is shown in Fig. 7a while the scaled network is shown in Fig. 7b. Although the scaled network has a different shape, its pore sizes and in particular the spatial distribution of pore sizes is strikingly similar. This result was made possible by the combination of Gaussian KDE and Gaussian Process.
The flow chart in Fig. 7c shows the workflow for obtaining a scaled-up network from the fitted parent network. First, the pore sizes and aspect ratios from the parent network were extracted and distributions were fitted to these using Gaussian KDE. At this point, the bandwidth is chosen, and while there are different correlations for determining an appropriate bandwidth such as Scott60 which helps as a guide to prevent overfitting, we selected a small bandwidth of just 0.01 (compared to Scott's estimate which was roughly 0.2) because ultimately overfitting was not as much of a concern as it was to have a representative pore network with very similar geometric property distributions. After fitting a distribution, using Gaussian KDE, any number of samples (i.e. any number of pore/throat sizes) can be sampled from the distribution and used as the geometric properties of the new and enlarged network. After sampling, Gaussian Process is used to identify to what pore or throat these sizes should be assigned. It is important that the GP model is trained on the same spatial domain that will be used to predict pore sizes of the new network. Therefore, prior to training, the spatial coordinates of pores in the fitted network are scaled to fit between 0 and 1 in the x, y, and z coordinates. Then, prior to predicting pore sizes of the new network, the spatial coordinates of the enlarged network are also scaled to fit within the same space. After scaling the coordinates, two GP models were trained. The first GP model was trained on the pore sizes and their spatial coordinates taken from the fitted network while the second GP model was trained on the throat aspect ratios and the coordinates of the two pores connected to each throat. Training a GP model to predict the spatial locations of the aspect ratios was found to be more difficult due to how throats can orient themselves in three different spatial directions. To get around this problem, it was found that training on the set of pore coordinates each throat connects, was the best way to predict throat sizes spatially. Justification for this is given by Fig. S15 in the SI. Training GP models on this size of data was relatively fast as it took less than 30 seconds to train both models on an Intel Core i7-12700KF CPU using data from a 1000 pore and 2700 throat network. The hyperparameters used to fit the two GP models including signal variance and length scale are recorded in Table S2 of the SI. After fitting GP models, pore diameters and throat aspect ratios were sampled, but instead of using these sizes directly, their size was used to sort the properties sampled using gaussian KDE. In this way, the distribution of geometric properties of the parent network are maintained while at the same time maintaining their spatial correlation.
Finally, Fig. 7 compares the properties of the scaled network using GP to target properties of the actual Berea sample. In Fig. 7f, the permeability of the scaled network using GP is presented as the orange bar while the target and fitted permeabilities are shown as green and purple bars. The average permeability of the scaled network is only 4.1% off from the target. Compare that to the average without using GP to account for spatial correlations and the absolute percent difference increases to 73%. The impact of considering spatial correlations is also noticed in the predicted saturation curve from porosimetry simulations on spatially and not spatially correlated networks shown in Fig. 7g. Here, the orange curve indicates the saturation of the network with spatially correlated properties using GP, while the pink curve is the network without spatially correlated properties and therefore did not use GP. The effect of considering spatial correlations obviously improves the fit and in fact, the SSE improved from 5.1 × 10−1 to 4.6 × 10−3 after considering spatial correlations. The impact of spatial correlations is obvious here and continues to reinforce the idea established early in the literature by Bryant et al. that randomly assigning pore sizes can give erroneous prediction of properties especially permeability.24 That is why it was so important in this work to develop an approach that captures spatial correlations after computationally expensive optimization to be able to scale the network as necessary.
4 Conclusions
In this paper, we presented a novel approach for constructing a pore network directly from experimental data, without relying on high-resolution X-ray tomography, SEM, or FIB imaging. The main highlights of this work were the use of automatic differentiable and GPU compatible flow and invasion porosimetry solvers, a proven workflow for constructing a pore network from MIP and permeability data that uses gradient descent-based optimization, and finally a workflow for stochastic network generation of any size/shape that trains a Gaussian Process model to learn spatial correlations. This work builds on past literature which has developed optimization methods that use non-gradient based approaches,28,32 and is the only work, to the best of our knowledge, that has attempted the use of automatic differentiation to optimize a pore network.While novel advancements were made in this work, there are some limitations of the proposed optimization strategy worth discussing to help guide future work. First, the optimization depends on the use of a number of parameters including initial guess, learning rate, smoothing factor, and number of iterations. It was observed that these parameters, in particular the smoothing factor used for invasion algorithms, learning rate, and initial guess, strongly influence the final solution. A more detailed study on some of these parameters and their effect on optimization would be helpful to understand parameter selection better. In particular, how these parameters effect prediction of the actual pore size distribution, the final loss, and the resulting spatial correlation should be investigated in more detail. Second, this work focused on generating regular networks with fixed coordination number among other simplifications. While it is common to assume coordination number when calibrating pore networks to experimental data6,7,28 there is much interest in generating more realistic networks as many works treat coordination number as a free parameter.30,61,62 Therefore, it is recommended that future works will attempt to use gradient descent to construct irregular networks, or perhaps networks with further reduced simplifications, including the spheres and cylinders geometry assumed here. Third and finally, is the extent of computational resources used. The source of some of the many advantages observed by JAX is the XLA compilation it used to write efficient machine code. One of the drawbacks, however, of using XLA is its potentially long compilation time especially observed for large networks. This is why, a small network of just 103 pores was used for optimization in this work. If it is possible to write more efficiently compiled code or make use of other high performance computational resources such as TPUs, it may become practical to optimize larger networks.
To conclude, the recent development of  ,35 a high-performance computing package that allows for easy creation of automatic differentiable functions, was the main motivation behind taking a second look at using gradient descent for pore network optimization. The results suggest exceptional capability of gradient descent optimization to meet specific design objectives, while considering a pore's spatial relationship with respect to its neighbours. Therefore, it is strongly recommended that automatic differentiation continue to be explored as a route for optimizing pore networks in the future.
,35 a high-performance computing package that allows for easy creation of automatic differentiable functions, was the main motivation behind taking a second look at using gradient descent for pore network optimization. The results suggest exceptional capability of gradient descent optimization to meet specific design objectives, while considering a pore's spatial relationship with respect to its neighbours. Therefore, it is strongly recommended that automatic differentiation continue to be explored as a route for optimizing pore networks in the future.
Author contributions
Michael McKague: methodology, software, validation, data Curation, visualization, writing – original draft, writing – review & editing. Mohammad Mehrnia: conceptualization, methodology, software, writing – review & editing. Mohammad Amin Sadeghi: conceptualization. Jeff Gostick: supervision, resources, funding acquisition, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
The source code used in this work can be found here: https://doi.org/10.5281/zenodo.18612631.The images used in this study were made available by researchers at Imperial College London. The DOIs for the images are: https://doi.org/10.6084/m9.figshare.1153794, https://doi.org/10.6084/m9.figshare.1189257, https://doi.org/10.6084/m9.figshare.1189258, https://doi.org/10.6084/m9.figshare.1189274, https://doi.org/10.6084/m9.figshare.1189275, https://doi.org/10.6084/m9.figshare.1189276, https://doi.org/10.6084/m9.figshare.1189280, https://doi.org/10.6084/m9.figshare.1189281, https://doi.org/10.6084/m9.figshare.1189282, https://doi.org/10.6084/m9.figshare.1189284, https://doi.org/10.6084/m9.figshare.1189285, https://doi.org/10.6084/m9.figshare.1189286, https://doi.org/10.6084/m9.figshare.1189255
Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00455a.
Acknowledgements
This work was supported by the Natural Sciences and Engineering Research Council of Canada (NSERC) grant RGPIN-2023-03741; and the Azzam-Dullien Endowed Professorship donated to the University of Waterloo by Mohamed Azzam.References
- C. Shi, H. Janssen, C. Shi and H. Janssen, Improved Algorithms for Stochastic Pore Network Generation for Porous Materials, Transp. Porous Media, 2025, 152(5), 1–27, DOI:10.1007/S11242-025-02174-4.
- J. Zhao, F. Qin, D. Derome, Q. Kang and J. Carmeliet, Improved pore network models to simulate single-phase flow in porous media by coupling with lattice Boltzmann method, Adv. Water Resour., 2020, 145, 103738, DOI:10.1016/J.ADVWATRES.2020.103738.
- X. Miao, K. M. Gerke and T. O. Sizonenko, A new way to parameterize hydraulic conductances of pore elements: A step towards creating pore-networks without pore shape simplifications, Adv. Water Resour., 2017, 105, 162–172, DOI:10.1016/J.ADVWATRES.2017.04.021.
- Z. A. Khan, M. Agnaou, M. A. Sadeghi, A. Elkamel and J. T. Gostick, Pore Network Modelling of Galvanostatic Discharge Behaviour of Lithium-Ion Battery Cathodes, J. Electrochem. Soc., 2021, 168, 070534, DOI:10.1149/1945-7111/ac120c.
- M. McKague, H. Fathiannasab, M. Agnaou, M. A. Sadeghi and J. Gostick, Extending pore network models to include electrical double layer effects in micropores for studying capacitive deionization, Desalination, 2022, 535, 115784, DOI:10.1016/j.desal.2022.115784.
- J. T. Gostick, M. A. Ioannidis, M. W. Fowler and M. D. Pritzker, Pore network modeling of fibrous gas diffusion layers for polymer electrolyte membrane fuel cells, J. Power Sources, 2007, 173, 277–290, DOI:10.1016/j.jpowsour.2007.04.059.
- Q. Xiong, T. G. Baychev and A. P. Jivkov, Review of pore network modelling of porous media: Experimental characterisations, network constructions and applications to reactive transport, J. Contam. Hydrol., 2016, 192, 101–117, DOI:10.1016/j.jconhyd.2016.07.002.
- W. B. Lindquist, S. M. Lee, D. A. Coker, K. W. Jones and P. Spanne, Medial axis analysis of void structure in three-dimensional tomographic images of porous media, J. Geophys. Res. Solid Earth, 1996, 101, 8297–8310, DOI:10.1029/95JB03039.
- Z. Liang, M. A. Ioannidis and I. Chatzis, Geometric and Topological Analysis of Three-Dimensional Porous Media: Pore Space Partitioning Based on Morphological Skeletonization, J. Colloid Interface Sci., 2000, 221, 13–24, DOI:10.1006/JCIS.1999.6559.
- Z. Jiang, K. Wu, G. Couples, M. I. J. van Dijke, K. S. Sorbie and J. Ma, Efficient extraction of networks from three-dimensional porous media, Water Resour. Res., 2007, 43, W12S03, DOI:10.1029/2006WR005780.
- M. McKague, H. Fathiannasab, M. A. Sadeghi and J. Gostick, MAGNET: Medial Axis Guided Network Extraction Tool, InterPore J., 2025, 2, IPJ011225–IPJ011227, DOI:10.69631/G47X8W91.
- H. Dong and M. J. Blunt, Pore-network extraction from micro-computerized-tomography images, Phys. Rev. E, 2009, 80, 036307, DOI:10.1103/PHYSREVE.80.036307/FIGURES/18/MEDIUM.
- D. Silin and T. Patzek, Pore space morphology analysis using maximal inscribed spheres, Physica A, 2006, 371, 336–360, DOI:10.1016/J.PHYSA.2006.04.048.
- J. T. Gostick, Versatile and efficient pore network extraction method using marker-based watershed segmentation, Phys. Rev. E, 2017, 96, 023307, DOI:10.1103/PHYSREVE.96.023307/FIGURES/13/MEDIUM.
- J. Gostick, Z. Khan, T. Tranter, M. Kok, M. Agnaou, M. Sadeghi and R. Jervis, PoreSpy: A Python Toolkit for Quantitative Analysis of Porous Media Images, J. Open Source Softw., 2019, 4, 1296, DOI:10.21105/joss.01296.
- M. Aghighi, M. A. Hoeh, W. Lehnert, G. Merle and J. Gostick, Simulation of a full fuel cell membrane electrode assembly using pore network modeling, J. Electrochem. Soc., 2016, 163, F384–F392, DOI:10.1149/2.0701605jes.
- M. Rebai and M. Prat, Scale effect and two-phase flow in a thin hydrophobic porous layer. Application to water transport in gas diffusion layers of proton exchange membrane fuel cells, J. Power Sources, 2009, 192, 534–543, DOI:10.1016/J.JPOWSOUR.2009.02.090.
- R. van Gorp, M. van der Heijden, M. Amin Sadeghi, J. Gostick and A. Forner-Cuenca, Bottom-up design of porous electrode by combining a genetic algorithm and a pore network model, Chem. Eng. J., 2022, 139947, DOI:10.1016/J.CEJ.2022.139947.
- M. A. Sadeghi, M. Aganou, M. Kok, M. Aghighi, G. Merle, J. Barralet and J. Gostick, Exploring the Impact of Electrode Microstructure on Redox Flow Battery Performance Using a Multiphysics Pore Network Model, J. Electrochem. Soc., 2019, 166, A2121–A2130, DOI:10.1149/2.0721910jes.
- O. R. Cardoso and R. de C. Balaban, Comparative study between Botucatu and Berea sandstone properties, J. South Am. Earth Sci., 2015, 62, 58–69, DOI:10.1016/J.JSAMES.2015.04.004.
- J. H. Jin, J. Kim, J. Y. Lee and Y. M. Oh, Correlative multiple porosimetries for reservoir sandstones with adoption of a new reference-sample-guided computed-tomographic method, Sci. Rep., 2016, 6, 1–10, DOI:10.1038/SREP30250;TECHMETA=123;SUBJMETA=301,354,357,431,445,639,704;KWRD=NANOPARTICLES,PETROLOGY.
- G. Mason and N. R. Morrow, Capillary behavior of a perfectly wetting liquid in irregular triangular tubes, J. Colloid Interface Sci., 1991, 141, 262–274, DOI:10.1016/0021-9797(91)90321-X.
- A. Fathiganjehlou, E. A. J. F. Peters, K. A. Buist and J. A. M. Kuipers, Pore Network Modeling of Intraparticle Transport Phenomena Accompanied by Chemical Reactions, Ind. Eng. Chem. Res., 2024, 63, 17662–17678, DOI:10.1021/ACS.IECR.4C01727/ASSET/IMAGES/LARGE/IE4C01727_0015.JPEG.
- S. L. Bryant, P. R. King and D. W. Mellor, Network model evaluation of permeability and spatial correlation in a real random sphere packing, Transp. Porous Media, 1993, 11, 53–70, DOI:10.1007/BF00614635.
- N. Baishnab, E. Herron, A. Balu, S. Sarkar, A. Krishnamurthy and B. Ganapathysubramanian, 3D multiphase heterogeneous microstructure generation using conditional latent diffusion models, Digital Discovery, 2025, 3175–3190, 10.1039/D5DD00159E.
- C. Yu, W. Chen, J. Li and S. Wang, An Efficient Method for Generating a Super-Sized and Heterogeneous Pore-Throat Network Model of Rock, Appl. Sci., 2025, 15, 1047–15, DOI:10.3390/APP15031047.
- Z. Zhao, Y. D. Shou and X. P. Zhou, A novel digital extraction approach of pore network models from carbonates inspired by quantum genetic optimization techniques, Acta Geotech., 2024, 19, 3805–3820, DOI:10.1007/S11440-024-02310-2/FIGURES/10.
- S. Mufti and A. Das, Optimization-based pore network modeling approach for determination of hydraulic conductivity function of granular soils, Int. J. Numer. Anal. Methods GeoMech., 2024, 48, 4035–4056, DOI:10.1002/NAG.3826.
- L. Xu, X. Liu and L. Liang, A pore network model reconstruction method via genetic algorithm, J. Nat. Gas Sci. Eng., 2014, 21, 907–914, DOI:10.1016/j.jngse.2014.09.038.
- A. Raoof and S. Majid Hassanizadeh, A new method for generating pore-network models of porous media, Transp. Porous Media, 2010, 81, 391–407, DOI:10.1007/S11242-009-9412-3.
- M. van der Heijden, G. Szendrei, V. de Haas and A. Forner-Cuenca, A versatile optimization framework for porous electrode design, Digital Discovery, 2024, 3, 1292–1307, 10.1039/D3DD00247K.
- M. H. Sharqawy, Construction of pore network models for Berea and Fontainebleau sandstones using non-linear programing and optimization techniques, Adv. Water Resour., 2016, 98, 198–210, DOI:10.1016/J.ADVWATRES.2016.10.023.
- C. C. Margossian and C. C. Charles Margossian, A review of automatic differentiation and its efficient implementation, WIREs Data Min. Knowl. Discov., 2019, 9, e1305, DOI:10.1002/WIDM.1305.
- A. G. Baydin, B. A. Pearlmutter, A. A. Radul and J. M. Siskind, Automatic Differentiation in Machine Learning: a Survey, J. Mach. Learn. Res., 2018, 18, 1–43 Search PubMed , http://jmlr.org/papers/v18/17-468.html.
- J. Bradbury, R. Frostig, P. Hawkins, M. J. Johnson, C. Leary, D. Maclaurin, G. Necula, A. Paszke, J. VanderPlas, S. Wanderman-Milne and Q. Zhang, JAX: composable transformations of Python+NumPy programs, 2018. http://github.com/jax-ml/jax Search PubMed.
- P. L. Churchel, P. B. French, J. C. Shaw, L. L. Schramm, Rock properties of Berea sandstone, Baker dolomite, and Indiana limestone, 1991, pp, 431–446, DOI:10.2118/21044-MS.
- N. A. Idowu and M. J. Blunt, Pore-scale modelling of rate effects in waterflooding, Transp. Porous Media, 2010, 83, 151–169, DOI:10.1007/S11242-009-9468-0/METRICS.
- M. A. Sadeghi, M. Agnaou, J. Barralet and J. Gostick, Dispersion modeling in pore networks: A comparison of common pore-scale models and alternative approaches, J. Contam. Hydrol., 2019, 103578, DOI:10.1016/j.jconhyd.2019.103578.
- M. Akbari, D. Sinton and M. Bahrami, Viscous flow in variable cross-section microchannels of arbitrary shapes, Int. J. Heat Mass Tran., 2011, 54, 3970–3978, DOI:10.1016/J.IJHEATMASSTRANSFER.2011.04.028.
- H. Giesche, Mercury Porosimetry: A General (Practical) Overview, Part. Part. Syst. Char., 2006, 23, 9–19, DOI:10.1002/PPSC.200601009.
- A. Hunt, R. Ewing and B. Ghanbarian, Percolation Theory for Flow, Porous Media, 2014, pp. 880 , DOI:10.1007/978-3-319-03771-4.
- J. Gostick, M. Aghighi, J. Hinebaugh, T. Tranter, M. A. Hoeh, H. Day, B. Spellacy, M. H. Sharqawy, A. Bazylak, A. Burns, W. Lehnert and A. Putz, OpenPNM: A Pore Network Modeling Package, Comput. Sci. Eng., 2016, 18, 60–74, DOI:10.1109/MCSE.2016.49.
- E. W. Washburn, The Dynamics of Capillary Flow, Phys. Rev., 1921, 17, 273, DOI:10.1103/PhysRev.17.273.
- A. Rodríguez de Castro, M. Agnaou, A. Ahmadi-Sénichault and A. Omari, Numerical porosimetry: Evaluation and comparison of yield stress fluids method, mercury intrusion porosimetry and pore network modelling approaches, Comput. Chem. Eng., 2020, 133, 106662, DOI:10.1016/J.COMPCHEMENG.2019.106662.
- G. Garfi, C. M. John, M. Rücker, Q. Lin, C. Spurin, S. Berg and S. Krevor, Determination of the spatial distribution of wetting in the pore networks of rocks, J. Colloid Interface Sci., 2022, 613, 786–795, DOI:10.1016/J.JCIS.2021.12.183.
- S. Foroughi, B. Bijeljic, Q. Lin, A. Q. Raeini and M. J. Blunt, Pore-by-pore modeling, analysis, and prediction of two-phase flow in mixed-wet rocks, Phys. Rev. E, 2020, 102, 023302, DOI:10.1103/PhysRevE.102.023302.
- B. K. Primkulov, S. Talman, K. Khaleghi, A. Rangriz Shokri, R. Chalaturnyk, B. Zhao, C. W. Macminn and R. Juanes, Quasistatic fluid-fluid displacement in porous media: Invasion-percolation through a wetting transition, Phys. Rev. Fluids, 2018, 3, 104001, DOI:10.1103/PHYSREVFLUIDS.3.104001/INVASION_160_FAST.MP4.
- V. Joekar-Niasar and S. M. Hassanizadeh, Analysis of fundamentals of two-phase flow in porous media using dynamic pore-network models: A review, Crit. Rev. Environ. Sci. Technol., 2012, 42, 1895–1976, DOI:10.1080/10643389.2011.574101.
- J. J. Dongarra, I. S. Duff, D. C. Sorensen and H. A. van der Vorst, Numerical linear algebra for high-performance computers, 1998, p. 342 Search PubMed.
- D. B. Kirk and W. W. Hwu, Programming Massively Parallel Processors: A Hands-on Approach, Morgan Kaufmann Publishers Inc., San Francisco, CA, USA, 1st edn, 2010 Search PubMed.
- Y. Saad, Iterative Methods for Sparse Linear Systems, Iterative Methods for Sparse Linear Systems, 2003, DOI:10.1137/1.9780898718003.
- Z. Yi, M. Lin, W. Jiang, Z. Zhang, H. Li and J. Gao, Pore network extraction from pore space images of various porous media systems, Water Resour. Res., 2017, 53, 3424–3445, DOI:10.1002/2016WR019272.
- J. E. Santos, A. Gigliotti, A. Bihani, C. Landry, M. A. Hesse, M. J. Pyrcz and M. Prodanović, MPLBM-UT: Multiphase LBM library for permeable media analysis, SoftwareX, 2022, 18, 101097, DOI:10.1016/J.SOFTX.2022.101097.
- W. M. Mahmud, Rate-Controlled Mercury Injection Experiments to Characterize Pore Space Geometry of Berea Sandstone, E3S Web Conf., 2023, 366 DOI:10.1051/E3SCONF/202336601016.
- T. Bultreys, L. Van Hoorebeke and V. Cnudde, Multi-scale, micro-computed tomography-based pore network models to simulate drainage in heterogeneous rocks, Adv. Water Resour., 2015, 78, 36–49, DOI:10.1016/J.ADVWATRES.2015.02.003.
- A. Mehmani and M. Prodanović, The effect of microporosity on transport properties in porous media, Adv. Water Resour., 2014, 63, 104–119, DOI:10.1016/J.ADVWATRES.2013.10.009.
- C. D. Tsakiroglou, M. A. Ioannidis, E. Amirtharaj and O. Vizika, A new approach for the characterization of the pore structure of dual porosity rocks, Chem. Eng. Sci., 2009, 64, 847–859, DOI:10.1016/J.CES.2008.10.046.
- B. W. Silverman, Density estimation: For statistics and data analysis, Density Estimation: For Statistics and Data Analysis, 2018, pp. 1–175, DOI:10.1201/9781315140919/DENSITY-ESTIMATION-STATISTICS-DATA-ANALYSIS-BERNARD-SILVERMAN/RIGHTS-AND-PERMISSIONS.
- C. E. Rasmussen and C. K. I. Williams, Gaussian Processes for Machine Learning, Gaussian Processes for Machine Learning, 2005, DOI:10.7551/MITPRESS/3206.001.0001.
- D. W. Scott, Multivariate density estimation: Theory, practice, and visualization: Second edition, Multivariate Density Estimation: Theory, Practice, and Visualization, 2nd edn, 2015, ppp. 1–360, DOI:10.1002/9781118575574.
- P. Čapek, V. Hejtmánek, L. Brabec, A. Zikánová and M. Kočiřík, Network modelling of capillary pressure curves, permeability, and diffusivity, Chem. Eng. Sci., 2007, 62, 5117–5122, DOI:10.1016/J.CES.2007.01.011.
- Z. Jiang, M. I. J. van Dijke, K. Wu, G. D. Couples, K. S. Sorbie and J. Ma, Stochastic Pore Network Generation from 3D Rock Images, Transp. Porous Media, 2012, 94, 571–593, DOI:10.1007/S11242-011-9792-Z/METRICS.
| This journal is © The Royal Society of Chemistry 2026 |