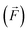Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Statistics makes a difference: machine learning adsorption dynamics of functionalized cyclooctyne on Si(001) at DFT accuracy
Hendrik
Weiske
a,
Rhyan
Barrett
a,
Ralf
Tonner-Zech
 a,
Patrick
Melix
a,
Patrick
Melix
 a and
Julia
Westermayr
a and
Julia
Westermayr
 *ab
*ab
aWilhelm Ostwald Institute for Physical and Theoretical Chemistry, Faculty of Chemistry, Leipzig University, Leipzig, Germany. E-mail: julia.westermayr@uni-leipzig.de
bCenter for Scalable Data Analytics and Artificial Intelligence Dresden/Leipzig, Leipzig, Germany
First published on 19th January 2026
Abstract
The interpretation of experiments on reactive semiconductor surfaces requires statistically significant sampling of configurational space by molecular dynamics, but conventional ab initio methods are limited due to prohibitive computational costs. Machine-learning interatomic potentials provide a promising solution, bridging the gap between the chemical accuracy of short ab initio molecular dynamics (AIMD) and the extensive sampling required to simulate experiment. Using ethinyl-functionalized cyclooctyne adsorption on Si(001) as a model system, we demonstrate that conventional AIMD undersamples the configurational space, resulting in discrepancies with scanning tunnelling microscopy and X-ray photoelectron spectroscopy data. To resolve these inconsistencies, we employ pre-trained equivariant message-passing neural networks, fine-tuned on only a few thousand AIMD snapshots, and integrate them into a “molecular-gun” workflow. This approach generates 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 independent trajectories more than 1000 times faster than AIMD. These simulations recover rare intermediates, clarify the competition between adsorption motifs, and reproduce the experimentally dominant on-top [2 + 2] cycloaddition structure. Our results show that fine-tuning of pre-trained foundational models enables statistically converged, chemically accurate simulations of bond-forming and bond-breaking events on complex surfaces, providing a scalable route to reconcile atomistic theory with experimental ensemble measurements in semiconductor functionalization.
000 independent trajectories more than 1000 times faster than AIMD. These simulations recover rare intermediates, clarify the competition between adsorption motifs, and reproduce the experimentally dominant on-top [2 + 2] cycloaddition structure. Our results show that fine-tuning of pre-trained foundational models enables statistically converged, chemically accurate simulations of bond-forming and bond-breaking events on complex surfaces, providing a scalable route to reconcile atomistic theory with experimental ensemble measurements in semiconductor functionalization.
1 Introduction
The functionalization of semiconductor surfaces, particularly silicon, offers a versatile means to tailor electronic, chemical, and mechanical properties.1–4 Cyclooctynes, widely used in strain-promoted click chemistry,5–11 serve as selective and reactive agents for Si(001) functionalization, enabling mild, covalent attachment while minimizing side reactions.7–9,12–19 Surface-sensitive experimental techniques such as scanning tunneling microscopy (STM) or X-ray photoelectron spectroscopy (XPS) provide rich detail on adsorption structures, coverage, and side reactions,19 yet they lack the temporal and atomistic resolution needed to observe transient intermediates and adsorption pathways required to resolve reaction kinetics. Moreover, ensemble-averaged spectroscopies yield information on overall surface composition and functional group identity, but fail to resolve site-specific energetics or orientation distributions. As a result, critical details, including the relative barriers for adsorption on the two non-equivalent dangling bonds of the Si(001) dimer (see Fig. 1), the influence of subsurface strain on cyclooctyne ring opening, and the lifetimes of metastable precursors, remain experimentally inaccessible.Computational approaches can complement experiment, but rely on computationally costly quantum-chemical calculations. As a consequence, studies are often limited to static analyses using density functional theory (DFT), which is usually the workhorse of such simulations.20,21 However, for capturing reaction kinetics and dynamical processes, molecular dynamics (MD) simulations are needed. Classical force fields, which offer a computationally viable solution, lack the ability to describe covalent bond formation and breaking. For some specific systems, reactive Force Fields (ReaxFF22,23) have been used in surface chemical studies. However, for statistically relevant sampling, these methods are also too demanding.24–27Ab initio molecular dynamics (AIMD), in principle, offers both reactivity and accuracy, yet its computational cost severely limits accessible timescales and statistical sampling.13,28–31 Recent work on ethinyl-functionalized cyclooctyne (ECCO, see Fig. 1) adsorption at Si(001) surfaces revealed a bottleneck: AIMD trajectories, even tens of picoseconds in length, can miss key binding modes observed experimentally, leading to discrepancies in predicted versus measured dominant adsorption geometries.19,32 Whether such mismatches stem from methodological limitations or from simple undersampling remains an open and critical question.
To address this question, we leverage machine learning (ML) to vastly accelerate surface MD simulations without compromising ab initio accuracy. Specifically, we fine-tune the foundational equivariant, message-passing atomic cluster expansion (MACE) model,33,34 MACE-MP-0, using our previous AIMD data for ECCO/Si(001),32 deploying a “molecular gun” strategy that generates thousands of statistically independent trajectories in a black-box fashion to simulate ultra-high-vacuum experiments (UHV) (for details see 2.1).
Therefore, simulations at near-DFT accuracy become accessible, while being multiple orders of magnitude faster.35 While the MACE-MP-0 architecture is widely adopted across molecular and materials applications,36–58 its suitability for surface fine-tuning with limited data,59–67 as demonstrated here, presents a practical solution to statistical convergence issues in surface chemistry.68–70
By fine-tuning a pre-trained MACE-MP-0 model34 with targeted ECCO/Si(001) AIMD snapshots,32 we remove the sampling bottleneck, enabling large-scale, chemically accurate simulations at affordable computational cost. Our machine learning molecular gun allows for detailed analysis of binding-site populations, desorption barriers, and ring-opening dynamics, placing the atomistic mechanism of ECCO adsorption in direct, quantitative correspondence with STM and XPS data.19
2 Computational details
To conduct ML-accelerated AIMD, we use the foundational MACE model for materials, MACE-MP-0,34 and fine-tune it on data obtained by some of us in a recent study.32 We therefore only briefly summarise the quantum-chemical reference simulations and the model architecture, referring to the cited publications for full details.2.1 AIMD reference simulations
The ab initio data for training were taken from previous work32,71 using DFT-based MD. Trajectories were generated using VASP 5.4.4,72–76 using the exchange–correlation functional by Perdew, Burke, and Ernzerhof (PBE)77,78 with the DFT-D3(BJ) dispersion correction scheme.79,80 The simulations were designed to model UHV deposition experiments, in which evaporated molecules impinge on a surface with finite kinetic energy. We therefore refer to this approach as the “molecular gun”. In this protocol, the Si(001) slab (Table S3 and Fig. S10) and the ECCO molecule were first equilibrated separately for 40 ps at 300 K in the NVT ensemble. From these trajectories, a configuration (coordinates and velocities) was extracted every 1 ps to sample thermally excited states of both subsystems. Ten such configurations served as initial states for subsequent MD runs, in which the molecule was accelerated towards the surface by adding a random downward and random ±x and ±y velocity component, mimicking the conditions of UHV deposition. The x and y contribution accounts for incoming angles in a randomised fashion. The velocity is rescaled to match 300 K before adding it to the molecule. We use this same strategy in our work to generate additional initial conditions and improve statistical sampling.In the reference data, dynamics were simulated in the NVT ensemble using a Nosé–Hoover thermostat81–83 at 300 K with a Nosé mass of 1.8.32
The complete AIMD dataset comprises approximately 327k frames: formed from ∼199k frames of the ECCO molecule and ∼128k frames of molecular gun runs (ECCO + Si(001)).71 All frames in which unphysical C–H bond fission occurred were removed, which was concluded to be the result of a too large time step in the underlying DFT data, surpassing the barrier that would be expected for the C–H fission. This was the case in two of the 10 AIMD trajectories.32 After randomising the remaining frames, every 25th configuration was selected to form the production machine learning dataset (coordinates, velocities, and energies), resulting in ∼13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 data points of ECCO on Si(001). As shown previously, only a fraction of all trajectories is enough to achieve good training results.84,85
000 data points of ECCO on Si(001). As shown previously, only a fraction of all trajectories is enough to achieve good training results.84,85
2.2 Machine learning MD
All MD simulations in this work were performed using the MD driver implemented in the atomic simulation environment (ASE).86 As initial configurations, we used the ten starting structures from the reference AIMD simulations,32 providing pre-equilibrated systems (see also Subsection 2.1). For each run, the position of the ECCO molecule in the x–y plane (parallel to the surface) was randomised. The distance between the surface atom plane and the ECCO centre of mass was fixed at 20 Å, corresponding to an approximate shortest atom–surface separation of 13 Å. To initiate motion towards the slab, a random velocity component was added to the initial DFT velocities of the ECCO molecule along the z-axis. The velocities of the slab atoms were kept unchanged from the AIMD frames. The simulations were propagated with a time step of 0.5 fs for 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 iterations, corresponding to a total simulation time of 10 ps. An NVE ensemble was employed,87 as the systems were pre-equilibrated at the target temperature in the DFT stage and the experimental surface-deposition process is intrinsically non-equilibrium.
000 iterations, corresponding to a total simulation time of 10 ps. An NVE ensemble was employed,87 as the systems were pre-equilibrated at the target temperature in the DFT stage and the experimental surface-deposition process is intrinsically non-equilibrium.
2.3 Machine learning
For machine learning, we employ the foundational messaging passing atomic cluster expansion (MACE)33 model, MACE-MP-0,34 which was originally trained on the Materials Project Trajectory (MPtrj) dataset.88 This dataset contains approximately 1.5 million configurations, primarily small periodic unit cells representing inorganic crystals with some molecular components.34 Notably, the MPtrj dataset contains limited surface-chemistry data, motivating the fine-tuning of MACE-MP-0 for improved data efficiency. Our fine-tuning approach assumes that knowledge gained from a large and diverse dataset of materials facilitates learning for new systems. Accordingly, the parameters of MACE-MP-0 were used to initialise the training of fine-tuned models. The model representation comprises of 128 scalar and 128 vectorial components. Fine-tuning was performed with a learning rate of 0.001 for 100 epochs to prevent overfitting, re-initialising the readout layers.89 Training employed a batch size of 16 across 8 NVIDIA A100-SXM4GPUs. A weighting factor of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 between energies and forces was applied during training in the loss function, where both contributions were computed using the mean squared error (MSE). The MSE of the energies was weighted by 1, and the MSE of the forces was weighted by 100, reflecting the greater importance of forces for MD simulations. Because of this weighting, the energies are considered less reliable in this work and only the forces are used to represent the correct dynamic behavior. Five percent of the 13k points are used for validation and 10k randomly selected structures from the total data points (excluding the 13k points used for training) are used for testing. All other architectural parameters were kept at their default values, matching those of the foundational MACE-MP-0 model.89 A fixed random seed of 24 was used for reproducibility. For comparison, we also trained MACE models from scratch using the same setup as the fine-tuned models, except for an increased training length of 1000 epochs.
100 between energies and forces was applied during training in the loss function, where both contributions were computed using the mean squared error (MSE). The MSE of the energies was weighted by 1, and the MSE of the forces was weighted by 100, reflecting the greater importance of forces for MD simulations. Because of this weighting, the energies are considered less reliable in this work and only the forces are used to represent the correct dynamic behavior. Five percent of the 13k points are used for validation and 10k randomly selected structures from the total data points (excluding the 13k points used for training) are used for testing. All other architectural parameters were kept at their default values, matching those of the foundational MACE-MP-0 model.89 A fixed random seed of 24 was used for reproducibility. For comparison, we also trained MACE models from scratch using the same setup as the fine-tuned models, except for an increased training length of 1000 epochs.
2.4 MD analysis
Trajectory analysis was performed using ASE modules.86,90 We further used an automated detection method for adsorption sites and modes. Structure visualizations were rendered using Blender91via our ASE-Blender interface.92 To evaluate the sampling density of the space above the Si(001) surface, a binning approach was conducted using NumPy v2.3.93 An xy-grid was created and, for each xy bin in the unit cell, the lowest occurring z-value of the centre of the cyclooctyne triple bond was stored for the respective set of trajectories. The spacing of the xy bins was set to one thousandth of the unit cell, corresponding to an area of 0.0015 Å2 per bin.3 Results and discussion
3.1 Machine learning
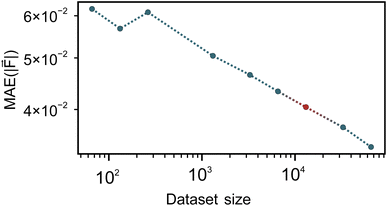
To ensure accurate machine learning interatomic potentials, we analysed the learning and training data distribution using learning curves and dimensionality reduction techniques, respectively.The learning curves for our fine-tuned models (Fig. 2) plot the force mean absolute error (MAE) on an independent test set, with respect to the DFT reference, against the number of training geometries on a log–log scale. The observed decay of the MAE is clearly linear, demonstrating that the fine-tuned model continues to benefit systematically from additional data. The energy and force errors for varying training set sizes are presented in detail for all models in Table S1 of the SI. Fine-tuning generally requires fewer epochs and, consequently, less training time than training models from scratch on the ECCO on Si(001) system (see also Table 1). Both models achieve lower errors than the non-fine-tuned foundational model. This is expected as the foundational model does not have knowledge on the data and is also trained on another reference method. Additionally, we find that models trained from scratch on our AIMD data achieve lower errors than the fine-tuned versions from the foundational model (1.75 × 10−3 eV and 2.97 × 10−2 eV Å−1 for energies and forces, respectively, compared to MAEs of 2.73 × 10−3 eV and 4.04 × 10−2 eV Å−1 for the fine-tuned models). This counter-intuitive result likely stems from differences between the MPtrj data and our target domain. The MPtrj dataset spans a much broader chemical space, containing bonding motifs and structures not directly relevant to our system. Fine-tuning adapts the model to our trajectories, but it begins from parameter values optimized for generalization across the MPtrj dataset. These values turn out to be less suitable for the narrower Si–C–H surface chemistry under study, compared with random initialization when training from scratch. Using a larger learning rate with a decay schedule may help the fine-tuned model escape the local minima associated with the pre-training, potentially achieving errors comparable to the model trained from scratch. In practice, this involves starting with a relatively high initial learning rate followed by exponential decay, where the learning rate is reduced after several epochs without improvement. However, this may come at the cost of losing much of the information gained from the MPtrj dataset. That said, it is important to note that a direct comparison of error metrics is not a good measure for evaluating performance in MD simulations. Machine learning potentials are, in essence, interpolators, and because the test set is drawn from the same underlying DFT trajectories as the training set, it contains very similar configurations. These metrics therefore do not reflect how the model behaves on new trajectories that sample regions of configuration space not well represented in the training data. For test set error metrics to more accurately reflect MD performance, a much more comprehensive set of reference configurations would be required, ideally covering the full space of potential trajectories. However, obtaining such coverage would require many expensive DFT trajectories and is therefore not practical. Nevertheless, the key advantage of fine-tuned models is that they retain knowledge from their pre-training, enabling broader transferability across chemical space. In our case, the strength lies in generalizing to unforeseen configurations not present in the training set that might be seen in a potential MD trajectory, making the model more suitable when generalizability and accurate observables are preferred over minimizing the error over a selection of predefined configurations. To support this, we compared MD simulations of unfunctionalized cyclooctyne at the Si(001) surface using both models, applying the same protocol. Remarkably, the fine-tuned model outperformed the model trained from scratch, resulting in a negligible number of unphysical cyclooctyne structures (see Section S4 of the SI for details). However, the foundational model used, MACE-MP-0, may not be an ideal starting point since it contains a large amount of inorganic bulk crystals not directly related to surface chemistry. Other models, such as the MACE-OMAT,94 which were trained on a vastly larger and more chemically diverse dataset, are expected to extrapolate better to surface systems; however, the underlying dataset still lacks explicit surface configurations. Future work in this area will likely focus on developing foundational models trained directly on surface and adsorbate data to improve transferability and reduce the data needed for task-specific fine-tuning.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 time steps. All timings in CPUh/GPUh, respectively. Training was performed on 8 GPUs and takes 1.1 h for fine-tuned models and 6.6 h for models trained from scratch
000 time steps. All timings in CPUh/GPUh, respectively. Training was performed on 8 GPUs and takes 1.1 h for fine-tuned models and 6.6 h for models trained from scratch
| Method | Full MDa | 1 MD step | 1000 MDs |
|---|---|---|---|
a 20.000 steps, extrapolated from 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 933 steps for DFT-MD.
b CPU: 20 Intel Haswell E5-2680v3 = 240 cores total.
c CPU: 1 AMD EPYC CPU 9334 = 1 core.
d GPU: NVIDIA H100-SXM5. 933 steps for DFT-MD.
b CPU: 20 Intel Haswell E5-2680v3 = 240 cores total.
c CPU: 1 AMD EPYC CPU 9334 = 1 core.
d GPU: NVIDIA H100-SXM5.
|
|||
| DFTb | 7.1 × 104 | 3.6 | 7.1 × 107 |
| MLc | 4.3 | 2.2 × 10−4 | 4.4 × 103 |
| MLd | 1.0 × 10−1 | 5.0 × 10−6 | 1.0 × 102 |
For final models, we use every 25th AIMD frame, resulting in 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 data points for training. To ensure that these ∼13
000 data points for training. To ensure that these ∼13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 configurations adequately span relevant reaction pathways and surface environments, we embedded both training and reference AIMD geometries into a low-dimensional manifold using principal component analysis (PCA) based on equivariant geometrical descriptors (Fig. 3). As shown, both datasets cover approximately the same space, demonstrating the completeness of our training dataset using only every 25th AIMD frame for training. Energy-scaled PCAs for all relevant parts of the dataset are presented in Fig. S6 (SI), indicating that reducing the production dataset size does not significantly reduce the chemical space or energy ranges covered.
000 configurations adequately span relevant reaction pathways and surface environments, we embedded both training and reference AIMD geometries into a low-dimensional manifold using principal component analysis (PCA) based on equivariant geometrical descriptors (Fig. 3). As shown, both datasets cover approximately the same space, demonstrating the completeness of our training dataset using only every 25th AIMD frame for training. Energy-scaled PCAs for all relevant parts of the dataset are presented in Fig. S6 (SI), indicating that reducing the production dataset size does not significantly reduce the chemical space or energy ranges covered.
 | ||
| Fig. 3 PCA of the full (red) and the production AIMD-datasets (blue). Equivariant features are used for the descriptors as inputs for PCA. | ||
3.2 ML driven MD simulations
To assess the role of statistics in MD simulations and to enable meaningful comparison with experiments, we performed 100, 1000, and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 trajectories using machine learning interatomic potentials, in contrast to the 10 trajectories feasible with full DFT-based AIMD. Both machine learning models were tested for their ability to reproduce relevant chemical events in the MD runs. As shown in Table 1, using machine learning models results in a dramatic reduction in computational time over AIMD of several orders of magnitude. At the same time, the training time of the fine-tuned model training is only 1.1 h on eight GPUs. For the model trained from scratch, increasing the number of epochs by a factor of 10 in comparison to the fine-tuned model, increases the training time by a factor of six to 6.6 h.
000 trajectories using machine learning interatomic potentials, in contrast to the 10 trajectories feasible with full DFT-based AIMD. Both machine learning models were tested for their ability to reproduce relevant chemical events in the MD runs. As shown in Table 1, using machine learning models results in a dramatic reduction in computational time over AIMD of several orders of magnitude. At the same time, the training time of the fine-tuned model training is only 1.1 h on eight GPUs. For the model trained from scratch, increasing the number of epochs by a factor of 10 in comparison to the fine-tuned model, increases the training time by a factor of six to 6.6 h.
To analyse whether improved statistics lead to larger sampling of configuration space during dynamics and new structures not observed in the 10 DFT-based AIMD, we analyse representative adsorption structures. These are illustrated in Fig. 4. Nine representative adsorption structures arise in our molecular gun simulations: 1 – on-top cyclooctyne (OT-CY), 2 – bridge cyclooctyne (BR-CY), 3 – on-top ethinyl (OT-ET), 4 – bridge ethinyl (BR-ET), 5 – double (DB), 6 – precursor (PC), 7 – sublayer (SL), 8 – sublayer double (SL-DB), and 9 – other.
Configurations 1–4 involve the molecule spanning two adjacent surface atoms, either via the cyclooctyne ring's triple bond (CY; 1 and 2) or the ethinyl group (ET; 3 and 4). The two surface atoms can be on the same Si dimer (on-top, OT; 1 and 3) or on neighboring dimers (bridge, BR; 2 and 4). Configuration 5 comprises states where both triple bonds react with the surface to form doubly bonded ECCO (DB). The precursor state (PC; 6) describes ECCO datively bound to a single Si atom. This state is observed in DFT data, where it is an important reaction intermediate.32 In sublayer (SL; 7) and sublayer-double (SL-DB; 8) structures, ECCO binds to a Si atom beneath the top layer, either singly or doubly. All other configurations (mainly both triple bonds in a datively bonded state) are grouped as other (9).
Fig. 5 shows the distribution of final ECCO adsorption sites on Si(001) for the fine-tuned model and the AIMD reference. The distributions obtained for the other discussed models are shown in Fig. S1. The main experimental adsorption mode corresponds to the on-top cyclooctyne (1), shown in yellow.19 The 10 DFT-based AIMDs fail to capture the experimentally dominant motif 1, while instead over-emphasizing the doubly bonded mode 5. The 1000 trajectories obtained using the fine-tuned MACE-MP-0 model are able to capture the experimental motif much better, recovering the statistics of the adsorption. Additionally, several adsorption configurations are found which correspond to intermediate states in the DFT trajectories. Notably, most of these structures are not observed experimentally since they are not stable enough to be observed under experimental conditions.
 | ||
| Fig. 5 Distribution of the binding modes at the end of each MD simulation for the fine-tuned MACE-MP-0 model and the AIMD reference. The binding mode distribution of the fine-tuned model is given in red and the AIMD reference in green. The “single adsorption configuration”19 observed in STM is highlighted in yellow.19 | ||
Not all MD trajectories yield covalent binding – non-productive outcomes are labelled: intact but floating ECCO (i), H-abstraction (ii), broken C–C bonds (iii, see Fig. S5, SI), and “explosions”, corresponding to total disintegration of the molecule due to insufficient training data in this region, (iv). As shown in Fig. S1a, the fine-tuned model best matches experiment. From-scratch models produce a high fraction (>30%) of desorbed intact ECCO (i), while MACE-MP-0 often breaks the C–C bond (∼20%; iii). The tendency of the MACE-MP-0 and from-scratch model yield unphysical or unproductive events, despite low MAEs, underscores that the fine-tuned model generalizes better and predicts realistic dynamics at scale. The increasing fraction of desorbed molecules predicted by the from-scratch model are considered unrealistic due to the experimental sticking coefficient of approximately one for the unfunctionalized cyclooctyne meaning that almost all molecules are expected to adsorb when hitting the surface.19 We do concede though that the models results may be less reliable once the dynamics moves too far outside of the training domain. Uncertainty measurements during key binding events could help increase the models trustworthiness.
Fig. S1b illustrates convergence of binding mode populations with increasing number of MD trajectories. Moving from 100 to 1000 machine learning based MD trajectories, there is a marked shift in the observed binding mode distribution and covered configurational space, as supported by PCA descriptor plots (see Fig. S7). The difference between 1000 and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 trajectories, however, is minor, indicating statistical convergence is reached after around 1000 independent simulations. The precise proportions of binding modes for 1000 and 10
000 trajectories, however, is minor, indicating statistical convergence is reached after around 1000 independent simulations. The precise proportions of binding modes for 1000 and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 runs are summarized in Table S2. In order to show the statistically unconverged results, four sets of 10 ML-trajectories are run (Fig. S12). This results in a varying distribution for each of the sets, showing the low number of trajectories to be a main contributor of the previous disagreement between experiment and theory.32
000 runs are summarized in Table S2. In order to show the statistically unconverged results, four sets of 10 ML-trajectories are run (Fig. S12). This results in a varying distribution for each of the sets, showing the low number of trajectories to be a main contributor of the previous disagreement between experiment and theory.32
Examining surface sampling, Fig. 6 displays the minimum distance between the cyclooctyne triple bond and the surface for all trajectories. DFT-based trajectories reach only a limited set of surface sites, reflecting the small sample. With 100 machine learning-driven simulations using the fine-tuned model, coverage expands, but many regions remain unsampled. For both 1000 and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 trajectories, all surface regions are visited, supporting the conclusion from Fig. S1b that 1000 runs suffice for statistical convergence (see Fig. S3 for mode-colored sampling, and Fig. S4 for a side view.)
000 trajectories, all surface regions are visited, supporting the conclusion from Fig. S1b that 1000 runs suffice for statistical convergence (see Fig. S3 for mode-colored sampling, and Fig. S4 for a side view.)
3.3 Preferred binding mode
The statistical convergence of binding mode distributions demonstrates that AIMD simulations based on only 10 DFT trajectories are insufficient to provide a realistic depiction of ECCO adsorption behavior. Notably, the most prominent double-adsorption structure (5) appears primarily because the PBE functional underestimates the ethinyl reaction barrier by approximately 0.1 to 0.2 eV, as previously shown.95 This underestimation, and the consequent increased likelihood for the ethinyl group to react with the surface, are therefore also inherited by our ML models.4 Conclusion
We have demonstrated that large-scale molecular dynamics sampling is essential to accurately reproduce experimental adsorption statistics for large molecules adsorbing on a surface at the example of ethinyl-functionalized cyclooctyne on Si(001). By leveraging equivariant message-passing neural network potentials, comparing models trained from scratch and fine-tuned models based on parameters of foundational models, we achieved over 103-fold speed-ups compared to conventional DFT-based molecular dynamics, enabling 103–104 trajectories to be run with moderate computational resources. Fine-tuning on a few thousand AIMD snapshots is critical to adapt foundational models for specific surface chemistry: without this step, important adsorption modes are missing in subsequent machine learning-driven MD. Moreover, fine-tuning requires only a handful of epochs, reducing both training times and data requirements, while also minimizing the risk of catastrophic forgetting compared to training from scratch.Our high-throughput based MDs uncover numerous new final states, such as mixed on-top/sublayer motifs, which are rare or completely absent in the few DFT-based runs. This comprehensive sampling shifts theoretical predictions towards the experimentally dominant on-top adsorption mode. Nevertheless, certain DFT-induced biases persist, in particular the tendency for the double-adsorption motif to appear due to the PBE underestimation of the ethinyl reaction barrier, which is inherited by the machine learning models. As the number of trajectories increases, surface sampling rapidly improves: at 100 runs, significant regions remain unsampled; at 1000, all key binding motifs are visited; at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, near-complete coverage of the surface is achieved.
000, near-complete coverage of the surface is achieved.
The statistical convergence of site populations between 1000 and 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 trajectories is minor, clearly indicating that poor statistical sampling – rather than deficiencies in ab initio theory – explains discrepancies between previous AIMD and experimental studies. Future work could further improve accuracy by employing Δ-learning, fine-tuning against higher-level quantum data, or selectively incorporating experimental observables to address residual DFT errors. Overall, our “machine-learning molecular gun” workflow provides a robust and scalable means to connect atomistic mechanisms with ensemble-level experiments, thereby guiding the rational design of surface-functionalized semiconductor devices. Due to the small fine-tuning required, this model can easily be extended to other surfaces and adsorbates with the potential to significantly increase the impact of modelling in computational surface dynamics.
000 trajectories is minor, clearly indicating that poor statistical sampling – rather than deficiencies in ab initio theory – explains discrepancies between previous AIMD and experimental studies. Future work could further improve accuracy by employing Δ-learning, fine-tuning against higher-level quantum data, or selectively incorporating experimental observables to address residual DFT errors. Overall, our “machine-learning molecular gun” workflow provides a robust and scalable means to connect atomistic mechanisms with ensemble-level experiments, thereby guiding the rational design of surface-functionalized semiconductor devices. Due to the small fine-tuning required, this model can easily be extended to other surfaces and adsorbates with the potential to significantly increase the impact of modelling in computational surface dynamics.
5 Outlook
This work is a mere first step into the combination of quantum-chemical surface chemistry with machine learning targeting a single molecule on a surface. In the future, this approach can be extended to include molecule–molecule interactions. However, on a DFT level, this is not possible at large scales because of the many possible arrangements of the molecules. Nevertheless, they can play a fundamental role in coverage dynamics, e.g., via reactivity reduction96,97 or a long-range steering effect,15 which has been investigated in a static fashion.The ability to obtain statistics in DFT-quality MD will enable completely new avenues in surface chemistry, functionalization, catalysis, thin-film growth, and related fields, enabling us to get the chemistry right and provide statistically relevant answers to experimental questions.
Author contributions
Hendrik Weiske: data curation, formal analysis, investigation, methodology, software, validation, visualization, writing – review & editing; Rhyan Barett: ML methodology, writing – editing; Ralf Tonner-Zech: conceptualization, funding acquisition, project administration, resources, supervision, writing – review & editing; Patrick Melix: data curation, formal analysis, methodology, project administration, software, supervision, writing – original draft preparation, review & editing; Julia Westermayr: funding acquisition, project administration, resources, supervision, writing – original draft preparation, review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data used to train ML models is freely available under the following link: https://doi.org/10.17172/NOMAD/2021.09.28-2.Supplementary information (SI): PDF and raw data. See DOI: https://doi.org/10.1039/d5dd00420a.
Code availability: The code used to conduct molecular dynamics with the MACE model is freely available via the ASE package. Instructions to train MACE models and foundational models using fine-tuning processes can be found in the corresponding manual. All scripts and outputs (excluding ML-MD trajectories due to size) produced in this project are available in the published raw data on zenodo: https://doi.org/10.5281/zenodo.17523493.
Acknowledgements
The authors thank L. Yang, S. Schumann, T. Oestereich, and D. Bitterlich for their help in preprocessing the data and setting up the MACE fine-tuning procedure. We thank Dr Fabian Pieck for providing the AIMD trajectories and discussion. This work was supported by the Deutsche Forschungsgemeinschaft (DFG) via SFB1083 “Structure and Dynamics of Internal Interfaces” and GRK 2721 “Hydrogen Isotopes 1,2,3H”. Computations for this work were done using resources of ZIH Dresden, NHR-PC2 Paderborn, CSC-Goethe Frankfurt, and the Leipzig University Computing Center. During manuscript preparation, the authors used the Perplexity AI writing assistant to refine language, improve clarity, and strengthen organization. AI suggestions were used to polish text, ensure consistency between sections, and enhance readability. All scientific data analysis, methodological development, interpretation, and content decisions were performed by the authors, who take full responsibility for the manuscript’s originality and accuracy.Notes and references
- R. A. Wolkow, Annu. Rev. Phys. Chem., 1999, 50, 413–441 CrossRef CAS PubMed.
- A. V. Teplyakov and S. F. Bent, J. Vac. Sci. Technol., A, 2013, 31, 050810 CrossRef.
- S. F. Bent, Surf. Sci., 2002, 500, 879–903 CrossRef CAS.
- J. S. Kachian, K. T. Wong and S. F. Bent, Acc. Chem. Res., 2010, 43, 346–355 CrossRef CAS PubMed.
- H. C. Kolb, M. G. Finn and K. B. Sharpless, Angew. Chem., Int. Ed., 2001, 40, 2004–2021 CrossRef CAS PubMed.
- N. Münster, P. Nikodemiak and U. Koert, Org. Lett., 2016, 18, 4296–4299 CrossRef PubMed.
- M. Reutzel, N. Münster, M. A. Lipponer, C. Länger, U. Höfer, U. Koert and M. Dürr, J. Phys. Chem. C, 2016, 120, 26284–26289 CrossRef CAS.
- T. Glaser, J. Meinecke, L. Freund, C. Länger, J.-N. Luy, R. Tonner, U. Koert and M. Dürr, Chem.–Eur. J., 2021, 27, 8082–8087 CrossRef CAS PubMed.
- T. Glaser, J. Meinecke, C. Länger, J.-N. Luy, R. Tonner, U. Koert and M. Dürr, ChemPhysChem, 2021, 22, 404–409 CrossRef CAS PubMed.
- P. Nalaoh, V. Clark, N. Arroyo-Currás and D. M. Jenkins, ACS Sens., 2025, 6039–6047 CrossRef CAS PubMed.
- M. Dürr, U. Höfer, U. Koert and R. Tonner-Zech, Acc. Chem. Res., 2025, 58, 2454–2465 CrossRef PubMed.
- G. Mette, M. Dürr, R. Bartholomäus, U. Koert and U. Höfer, Chem. Phys. Lett., 2013, 556, 70–76 CrossRef CAS.
- L. Pecher, S. Schmidt and R. Tonner, J. Phys. Chem. C, 2017, 121, 26840–26850 CrossRef CAS.
- L. Pecher, C. Schober and R. Tonner, Chem.–Eur. J., 2017, 23, 5459–5466 CrossRef PubMed.
- L. Pecher, S. Schmidt and R. Tonner, Beilstein J. Org. Chem., 2018, 14, 2715–2721 CrossRef CAS PubMed.
- L. Pecher and R. Tonner, Theor. Chem. Acc., 2018, 137, 48 Search PubMed.
- T. Glaser, C. Länger, J. Heep, J. Meinecke, M. G. Silly, U. Koert and M. Dürr, J. Phys. Chem. C, 2020, 124, 22619–22624 CrossRef CAS.
- T. Glaser, J. A. Peters, D. Scharf, U. Koert and M. Dürr, Chem. Mater., 2024, 36, 561–566 CrossRef CAS.
- C. Länger, J. Heep, P. Nikodemiak, T. Bohamud, P. Kirsten, U. Höfer, U. Koert and M. Dürr, J. Phys.: Condens. Matter, 2019, 31, 034001 CrossRef PubMed.
- R. J. Maurer, V. G. Ruiz, J. Camarillo-Cisneros, W. Liu, N. Ferri, K. Reuter and A. Tkatchenko, Prog. Surf. Sci., 2016, 91, 72–100 CrossRef CAS.
- R. J. Maurer, C. Freysoldt, A. M. Reilly, J. G. Brandenburg, O. T. Hofmann, T. Björkman, S. Lebègue and A. Tkatchenko, Annu. Rev. Mater. Res., 2019, 49, 1–30 CrossRef CAS.
- A. C. T. Van Duin, S. Dasgupta, F. Lorant and W. A. Goddard, J. Phys. Chem. A, 2001, 105, 9396–9409 CrossRef CAS.
- T. P. Senftle, S. Hong, M. M. Islam, S. B. Kylasa, Y. Zheng, Y. K. Shin, C. Junkermeier, R. Engel-Herbert, M. J. Janik, H. M. Aktulga, T. Verstraelen, A. Grama and A. C. T. Van Duin, npj Comput. Mater., 2016, 2, 15011 CrossRef CAS.
- X. Hu, J. Schuster and S. E. Schulz, J. Phys. Chem. C, 2017, 121, 28077–28089 CrossRef CAS.
- W. Zhu, H. Gong, Y. Han, M. Zhang and A. C. T. Van Duin, J. Phys. Chem. C, 2020, 124, 12512–12520 CrossRef CAS.
- J. Wen, T. Ma, W. Zhang, A. C. T. Van Duin and X. Lu, J. Phys. Chem. A, 2017, 121, 587–594 CrossRef CAS PubMed.
- N. Nayir, A. C. T. Van Duin and S. Erkoc, J. Phys. Chem. A, 2019, 123, 4303–4313 CrossRef CAS PubMed.
- R. Iftimie, P. Minary and M. E. Tuckerman, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 6654–6659 CrossRef CAS PubMed.
- M. R. Radeke and E. A. Carter, Annu. Rev. Phys. Chem., 1997, 48, 243–270 CrossRef CAS PubMed.
- D. Marx and J. Hutter, in Modern methods and algorithms of quantum chemistry, ed. J. Grotendorst, NIC, Jülich, 2000, pp. 301–449 Search PubMed.
- A. Groß, Curr. Opin. Electrochem., 2023, 40, 101345 CrossRef.
- F. Pieck and R. Tonner-Zech, Molecules, 2021, 26, 6653 CrossRef CAS PubMed.
- I. Batatia, D. P. Kovacs, G. Simm, C. Ortner and G. Csanyi, Adv. Neural Inf. Process. Syst., 2022, 11423–11436 Search PubMed.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon, W. J. Baldwin, F. Berger, N. Bernstein, A. Bhowmik, S. M. Blau, V. Cărare, J. P. Darby, S. De, F. Della Pia, V. L. Deringer, R. Elijošius, Z. El-Machachi, F. Falcioni, E. Fako, A. C. Ferrari, A. Genreith-Schriever, J. George, R. E. A. Goodall, C. P. Grey, P. Grigorev, S. Han, W. Handley, H. H. Heenen, K. Hermansson, C. Holm, J. Jaafar, S. Hofmann, K. S. Jakob, H. Jung, V. Kapil, A. D. Kaplan, N. Karimitari, J. R. Kermode, N. Kroupa, J. Kullgren, M. C. Kuner, D. Kuryla, G. Liepuoniute, J. T. Margraf, I.-B. Magdău, A. Michaelides, J. H. Moore, A. A. Naik, S. P. Niblett, S. W. Norwood, N. O'Neill, C. Ortner, K. A. Persson, K. Reuter, A. S. Rosen, L. L. Schaaf, C. Schran, B. X. Shi, E. Sivonxay, T. K. Stenczel, V. Svahn, C. Sutton, T. D. Swinburne, J. Tilly, C. van der Oord, E. Varga-Umbrich, T. Vegge, M. Vondrák, Y. Wang, W. C. Witt, F. Zills and G. Csányi, A foundation model for atomistic materials chemistry, arXiv, 2024, preprint, arXiv:2401.00096, DOI:10.48550/arXiv.2401.00096, https://arxiv.org/abs/2401.00096, Version Number: 2.
- W. G. Stark, C. Van Der Oord, I. Batatia, Y. Zhang, B. Jiang, G. Csányi and R. J. Maurer, Mach. Learn.: Sci. Technol., 2024, 5, 030501 Search PubMed.
- T. J. Giese, J. Zeng and D. M. York, J. Phys. Chem. B, 2025, 129, 5477–5490 CrossRef CAS PubMed.
- T. Shiota, K. Ishihara and W. Mizukami, Digital Discovery, 2024, 3, 1714–1728 RSC.
- L. L. Schaaf, B. Rhodes, M. Zick, S. Pugh, J. Hilliard, S. Sharma, C. Wade, P. Milner, G. Csanyi and A. Forse, AI for Accelerated Materials Design, 2024 Search PubMed.
- M. O. Sauer, P. M. Lyngby and K. S. Thygesen, Phys. Rev. Mater., 2025, 9, 074007 CrossRef CAS.
- V. Vanita, G. Mezzadra, C. Tealdi and O. Clemens, ACS Appl. Energy Mater., 2025, 8, 7562–7574 CrossRef CAS.
- P. Singh, A. M. K. R. and M. Dixit, ACS Appl. Electron. Mater., 2024, 6, 7065–7074 CrossRef CAS.
- J. Abdul Nasir, J. Guan, W. Jee, S. M. Woodley, A. A. Sokol, C. R. A. Catlow and A. Elena, Phys. Chem. Chem. Phys., 2025, 27, 19784–19796 RSC.
- A. Kabylda, B. Mortazavi, X. Zhuang and A. Tkatchenko, Adv. Funct. Mater., 2025, 35, 2417891 CrossRef CAS.
- T. Demeyere, T. Ellaby, M. Sarwar, D. Thompsett and C.-K. Skylaris, ACS Catal., 2025, 15, 5674–5682 CrossRef CAS.
- A. Loew, D. Sun, H.-C. Wang, S. Botti and M. A. L. Marques, npj Comput. Mater., 2025, 11, 178 CrossRef CAS.
- M. Cheng, C.-L. Fu, B. Yu, E. Rha, A. Chotrattanapituk, D. L. Abernathy, Y. Cheng and M. Li, A Foundation Model for Non-Destructive Defect Identification from Vibrational Spectra, arXiv, 2025, preprint, arXiv:2506.00725[cond-mat], DOI:10.48550/arXiv.2506.00725[cond-mat], http://arxiv.org/abs/2506.00725.
- C. Shen, S. Attarian, Y. Zhang, H. Zhang, M. Asta, I. Szlufarska and D. Morgan, SuperSalt: Equivariant Neural Network Force Fields for Multicomponent Molten Salts System, arXiv, 2024, preprint, arXiv:2412.19353, DOI:10.48550/arXiv.2412.19353, https://arxiv.org/abs/2412.19353, Version Number: 1.
- M. R. Schäfer, N. Segreto, F. Zills, C. Holm and J. Kästner, Apax: A Flexible and Performant Framework For The Development of Machine-Learned Interatomic Potentials, arXiv, 2025, preprint, arXiv:2505.22168, DOI:10.48550/arXiv.2505.22168, https://arxiv.org/abs/2505.22168, Version Number: 2.
- P. Novelli, G. Meanti, P. J. Buigues, L. Rosasco, M. Parrinello, M. Pontil and L. Bonati, Fast and Fourier Features for Transfer Learning of Interatomic Potentials, arXiv, 2025, preprint, arXiv:2505.05652, DOI:10.48550/arXiv.2505.0562, https://arxiv.org/abs/2505.05652, Version Number: 1.
- L. Hörmann, W. G. Stark and R. J. Maurer, npj Comput. Mater., 2025, 11, 196 CrossRef PubMed.
- D. Schwalbe-Koda, N. Govindarajan and J. B. Varley, Digital Discovery, 2025, 4, 234–251 RSC.
- S. Gupta, A. Rajan, E. Fako, T. J. F. Gonçalves, I. B. Müller, J. J. Varghese, A. Schäfer and S. De, J. Phys. Chem. C, 2025, 129, 3022–3033 CrossRef.
- E. Fako and S. De, Simple Heuristics for Advanced Sampling of Reactive Species on Surfaces, ChemRxiv, 2025, preprint DOI:10.26434/chemrxiv-2025-79nj4.
- X. Tian, A. Tosello Gardini, U. Raucci, H. Xiao, Y. Zhuo and M. Parrinello, Electrochemical Potential-Driven Water Dynamics Control CO2 Electroreduction at the Ag/H2O Interface, ChemRxiv, 2025, preprint DOI:10.26434/chemrxiv-2025-n41q2.
- J. Pitfield, M.-P. V. Christiansen and B. Hammer, Active Delta-learning with universal potentials for global structure optimization, arXiv, 2025, preprint, arXiv:2507.18485, DOI:10.48550/arXiv.2507.18485, https://arxiv.org/abs/2507.18485, Version Number: 1.
- M.-P. V. Christiansen and B. Hammer, J. Chem. Phys., 2025, 162, 184701 CrossRef CAS PubMed.
- A. Soyemi, K. Baral and T. Szilvasi, A Simple Iterative Approach for Constant Chemical Potential Simulations at Interfaces, arXiv, 2025, preprint, arXiv:2506.01050, DOI:10.48550/arXiv.2506.01050, https://arxiv.org/abs/2506.01050, Version Number: 1.
- L. Cvitkovich, F. Fehringer, C. Wilhelmer, D. Milardovich, D. Waldhör and T. Grasser, J. Chem. Phys., 2024, 161, 144706 CrossRef CAS PubMed.
- B. Deng, Y. Choi, P. Zhong, J. Riebesell, S. Anand, Z. Li, K. Jun, K. A. Persson and G. Ceder, npj Comput. Mater., 2025, 11, 9 CrossRef CAS.
- T. Rensmeyer, D. Kramer and O. Niggemann, On-the-Fly Fine-Tuning of Foundational Neural Network Potentials: A Bayesian Neural Network Approach, arXiv, 2025, preprint, arXiv:2507.13805[cs], DOI:10.48550/arXiv.2507.13805, http://arxiv.org/abs/2507.13805.
- M. Bertani and A. Pedone, J. Phys. Chem. C, 2025, 129, 12697–12709 CrossRef CAS.
- J. Hänseroth and C. Dreßler, Optimizing Machine Learning Potentials for Hydroxide Transport: Surprising Efficiency of Single-Concentration Training, arXiv, 2025, preprint, arXiv:2505.07580, DOI:10.48550/arXiv.2505.07580, https://arxiv.org/abs/2505.07580, Version Number: 1.
- H. Kaur, F. D. Pia, I. Batatia, X. R. Advincula, B. X. Shi, J. Lan, G. Csányi, A. Michaelides and V. Kapil, Faraday Discuss., 2025, 256, 120–138 RSC.
- A. M. Elena, P. D. Kamath, T. Jaffrelot Inizan, A. S. Rosen, F. Zanca and K. A. Persson, npj Comput. Mater., 2025, 11, 125 CrossRef CAS.
- Y. Lim, H. Park, A. Walsh and J. Kim, Matter, 2025, 8, 102203 CrossRef CAS.
- J. Steffen, J. Phys. Chem. C, 2025, 129, 13513–13531 CrossRef CAS.
- B. Focassio, L. P. M. Freitas and G. R. Schleder, ACS Appl. Mater. Interfaces, 2025, 17, 13111–13121 CrossRef CAS PubMed.
- O. Allam, M. Maghsoodi, S. S. Jang and S. D. Snow, ACS Appl. Mater. Interfaces, 2024, 16, 36215–36223 CrossRef CAS PubMed.
- N. Boulangeot, F. Brix, F. Sur and E. Gaudry, J. Chem. Theory Comput., 2024, 20, 7287–7299 CAS.
- X. Du, M. Liu, J. Peng, H. Chun, A. Hoffman, B. Yildiz, L. Li, M. Z. Bazant and R. Gómez-Bombarelli, ACS Cent. Sci., 2025, 11(9), 1558–1572 CrossRef CAS PubMed.
- F. Pieck, NOMAD dataset: Bonding and reactivity of an alkyne-functionalized cyclooctyne on Si(001) with quantitative electronic structure analysis, 2021, DOI:10.17172/NOMAD/2021.09.28-2.
- G. Kresse and J. Hafner, Phys. Rev. B: Condens. Matter Mater. Phys., 1993, 47, 558–561 CrossRef CAS PubMed.
- G. Kresse and J. Hafner, Phys. Rev. B: Condens. Matter Mater. Phys., 1994, 49, 14251–14269 CrossRef CAS PubMed.
- G. Kresse and J. Furthmüller, Phys. Rev. B: Condens. Matter Mater. Phys., 1996, 54, 11169–11186 CrossRef CAS PubMed.
- G. Kresse and J. Furthmüller, Comput. Mater. Sci., 1996, 6, 15–50 CrossRef CAS.
- G. Kresse and D. Joubert, Phys. Rev. B: Condens. Matter Mater. Phys., 1999, 59, 1758–1775 Search PubMed.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1997, 78, 1396 Search PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- S. Grimme, S. Ehrlich and L. Goerigk, J. Comput. Chem., 2011, 32, 1456–1465 CrossRef CAS PubMed.
- S. Nosé, J. Chem. Phys., 1984, 81, 511–519 CrossRef.
- W. G. Hoover, Phys. Rev. A, 1985, 31, 1695–1697 CrossRef PubMed.
- S. Nosé, Prog. Theor. Phys. Suppl., 1991, 103, 1–46 CrossRef.
- M. X. Tiefenbacher, B. Bachmair, C. G. Chen, J. Westermayr, P. Marquetand, J. C. B. Dietschreit and L. González, Digital Discovery, 2025, 4, 1478–1491 RSC.
- J. S. Smith, B. Nebgen, N. Lubbers, O. Isayev and A. E. Roitberg, J. Chem. Phys., 2018, 148, 241733 Search PubMed.
- A. H. Larsen, J. J. Mortensen, J. Blomqvist, I. E. Castelli, R. Christensen, M. Dułak, J. Friis, M. N. Groves, B. Hammer, C. Hargus, E. D. Hermes, P. C. Jennings, P. B. Jensen, J. Kermode, J. R. Kitchin, E. L. Kolsbjerg, J. Kubal, K. Kaasbjerg, S. Lysgaard, J. B. Maronsson, T. Maxson, T. Olsen, L. Pastewka, A. Peterson, C. Rostgaard, J. Schiøtz, O. Schütt, M. Strange, K. S. Thygesen, T. Vegge, L. Vilhelmsen, M. Walter, Z. Zeng and K. W. Jacobsen, J. Phys.: Condens. Matter, 2017, 29, 273002 Search PubMed.
- F. Bigi, M. Langer and M. Ceriotti, The dark side of the forces: assessing non-conservative force models for atomistic machine learning, arXiv, 2024, preprint, arXiv:2412.11569, DOI:10.48550/arXiv.2412.11569, https://arxiv.org/abs/2412.11569, Version Number: 5.
- B. Deng, P. Zhong, K. Jun, J. Riebesell, K. Han, C. J. Bartel and G. Ceder, Nat. Mach. Intell., 2023, 5, 1031–1041 CrossRef.
- Fine-tuning Foundation Models—mace 0.3.13 documentation, https://mace-docs.readthedocs.io/en/latest/guide/finetuning.htmlnaive-fine-tuning.
- P. Melix, F. Paesani and T. Heine, Adv. Theor. Simul., 2019, 2, 1900098 CrossRef CAS.
- B. O. Community, Blender – a 3D modelling and rendering package, 2018, http://www.blender.org.
- H. Weiske, F. Thiemann and P. Melix, Blender Import ASE, 2024, DOI:10.5281/zenodo.10776697.
- C. R. Harris, K. J. Millman, S. J. Van Der Walt, R. Gommers, P. Virtanen, D. Cournapeau, E. Wieser, J. Taylor, S. Berg, N. J. Smith, R. Kern, M. Picus, S. Hoyer, M. H. Van Kerkwijk, M. Brett, A. Haldane, J. F. Del Río, M. Wiebe, P. Peterson, P. Gérard-Marchant, K. Sheppard, T. Reddy, W. Weckesser, H. Abbasi, C. Gohlke and T. E. Oliphant, Nature, 2020, 585, 357–362 CrossRef CAS PubMed.
- L. Barroso-Luque, M. Shuaibi, X. Fu, B. M. Wood, M. Dzamba, M. Gao, A. Rizvi, C. L. Zitnick and Z. W. Ulissi, Open Materials 2024 (OMat24) Inorganic Materials Dataset and Models, arXiv, 2024, preprint, arXiv:2410.12771[cond-mat], DOI:10.48550/arXiv.2410.12771, http://arxiv.org/abs/2410.12771.
- L. Pecher and R. Tonner, Inorganics, 2018, 6, 17 CrossRef.
- T. Glaser, G. F. Nolte, T. Bohamud, P. Keller, M. G. Silly, H. Weiske, R. Tonner-Zech and M. Dürr, Angew. Chem., Int. Ed., 2025, e19990 Search PubMed.
- P. P. Wellmann, F. Pieck and R. Tonner-Zech, Chem. Mater., 2024, 36, 7343–7361 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |