 Open Access Article
Open Access ArticleStructure-guided machine learning for efficiency prediction of organic photovoltaics using experimentally informed molecular descriptors
JuHyun
Lee†
 a,
HyoJin
Ban†
a,
HyoJin
Ban†
 ad,
HyunIl
Seo
ad,
HyunIl
Seo
 b,
HangKen
Lee
b,
HangKen
Lee
 c,
Fiza
Arshad
c,
Fiza
Arshad
 ce and
DongWook
Kim
ce and
DongWook
Kim
 *a
*a
aDigital Chemistry Research Center, Korea Research Institute of Chemical Technology, South Korea. E-mail: dongwkim@krict.re.kr
bQuantumSoft Co., Inc., South Korea
cPhotoenergy Research Center, Korea Research Institute of Chemical Technology, South Korea
dDepartment of Computer Science and Engineering, Chungnam National University, Daejeon, South Korea
eAdvanced Materials and Chemical Engineering, University of Science and Technology (UST), Daejeon, South Korea
First published on 27th February 2026
Abstract
The efficiency of organic photovoltaics was estimated using a machine learning (ML) approach. We used the organic photovoltaics database built in-house by the Korea Research Institute of Chemical Technology. The dataset comprises reliable and representative experimental results for 1010 ternary organic solar cells (D1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D2
D2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A), obtained through repeated measurements. The data included 67 donors and 24 non-fullerene acceptors, device structures, donor/acceptor structures, donor-to-acceptor ratios, active-layer thicknesses, experimental conditions, and local symmetry. We fragmented the donors and acceptors using a self-developed method. A dataset was created by generating descriptors of the fragmented molecules and used to train various ML algorithms, including random forest, XGBoost, LightGBM, support vector regression, and multilayer perceptron. Model performance was evaluated using the coefficient of determination (R2). XGBoost showed the highest R2 of 0.849. The contributions of key features were interpreted using SHAP analysis. This paper presents an ML framework that combines molecular fragmentation and data-driven modeling.
A), obtained through repeated measurements. The data included 67 donors and 24 non-fullerene acceptors, device structures, donor/acceptor structures, donor-to-acceptor ratios, active-layer thicknesses, experimental conditions, and local symmetry. We fragmented the donors and acceptors using a self-developed method. A dataset was created by generating descriptors of the fragmented molecules and used to train various ML algorithms, including random forest, XGBoost, LightGBM, support vector regression, and multilayer perceptron. Model performance was evaluated using the coefficient of determination (R2). XGBoost showed the highest R2 of 0.849. The contributions of key features were interpreted using SHAP analysis. This paper presents an ML framework that combines molecular fragmentation and data-driven modeling.
1 Introduction
Organic photovoltaics (OPVs) have attracted considerable attention as a promising renewable energy technology owing to their low manufacturing cost, mechanical flexibility, and continuous improvements in power conversion efficiency (PCE).Since the first demonstration of OPV devices in 1986, significant progress has been achieved through advances in active-layer materials and device architectures.1 In particular, the introduction of donor–acceptor (D–A) bulk heterojunction (BHJ) structures enabled efficient exciton dissociation and charge transport, leading to substantial improvements in PCE.
Early efforts to enhance OPV performance relied heavily on the synthesis of novel donor and acceptor materials, as well as device-level optimization. Parallel to experimental developments, computational chemistry approaches based on density functional theory (DFT)2,3 and semi-empirical methods (e.g., AM1,4 PM6,5 and PM7 (ref. 6)) were employed to screen candidate materials and understand structure–property relationships. However, such approaches become computationally prohibitive when applied to large molecular libraries, limiting their applicability to high-throughput OPV material discovery.7–10
To overcome these limitations, data-driven approaches and machine learning (ML) techniques have increasingly been adopted to predict OPV performance. Early studies employed quantitative structure–property relationship (QSPR)11,12 models, followed by more advanced ML frameworks using experimentally measured or DFT13-derived descriptors. While these approaches demonstrated encouraging predictive capabilities, they often suffered from limited dataset sizes, computational bias, or reliance on idealized molecular representations.
More recently, fragmentation-based molecular representations have been introduced to reduce molecular complexity and improve model scalability. Wu et al.14 employed literature-derived fragmentation schemes to construct fingerprint-based ML models, whereas Kim et al.15 utilized synthesis-oriented functional group indexing to achieve improved prediction accuracy. Despite these advances, most existing studies remain focused on relatively simple donor–acceptor systems and are primarily based on literature-reported data,13,16–26 which are often biased toward high-efficiency devices and do not adequately reflect the diverse experimental conditions encountered in practical laboratories.
Machine-learning studies on ternary OPV systems have also emerged, with particular emphasis on D![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A1
A1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A2,27 configurations composed of small-molecule donors and acceptors. While these studies achieved meaningful success, their molecular representations are less suitable for polymer-rich systems, where repeating units, structural heterogeneity, and multiple donor components introduce additional complexity that is difficult to capture using conventional descriptors.
A2,27 configurations composed of small-molecule donors and acceptors. While these studies achieved meaningful success, their molecular representations are less suitable for polymer-rich systems, where repeating units, structural heterogeneity, and multiple donor components introduce additional complexity that is difficult to capture using conventional descriptors.
In this work, we focus on ternary OPV systems of the D1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D2
D2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A type, in which two polymeric donors are blended with a single acceptor. To the best of our knowledge, this study represents the first systematic ML investigation of ternary OPV devices involving multiple polymer donors. By leveraging a curated experimental database that includes both high- and low-efficiency devices, we mitigate literature bias and construct a more balanced representation of real experimental conditions.
A type, in which two polymeric donors are blended with a single acceptor. To the best of our knowledge, this study represents the first systematic ML investigation of ternary OPV devices involving multiple polymer donors. By leveraging a curated experimental database that includes both high- and low-efficiency devices, we mitigate literature bias and construct a more balanced representation of real experimental conditions.
To accurately describe such complex systems, we propose a fragment-based molecular representation based on chemically meaningful fragmentation. Fragment-level physicochemical descriptors are generated using RDKit and Stk, and local symmetry information is incorporated as a physically motivated proxy for molecular packing tendencies without explicitly modeling solid-state morphology. Redundant and irrelevant features are further removed through REF-based feature selection, enabling the construction of robust and transferable machine-learning models for ternary OPV systems.
Using this representation, we develop machine-learning models to predict power conversion efficiency and key device parameters. Interpretability analyses provide insights into structure–process–property relationships, highlighting the contributions of specific molecular fragments and device parameters to device performance (Fig. 1).
2 Results and discussion
2.1 Collection of experimental OPV data
The experimental data used in this study were collected over the past five years by the Photoenergy Research Center at the Korea Research Institute of Chemical Technology. In 2018, our Digital Chemistry Research team established an in-house OPV data platform to systematically accumulate, analyze, and visualize experimental data without reliance on external online tools (Fig. S1).The dataset consists of two main categories: materials and devices. The materials category contains chemical and structural information on donor and acceptor compounds, while the device category records OPV device architectures and electrode-related experimental results. ChemAxon's Marvin JS28 and JChem Microservices were used to convert compounds into MolV2000 format for structural representation, and CDK 2.8 was used to convert them into SMILES.29
In the device section, the platform stores detailed information on all OPV layers, including active-layer materials, additives, solvents, and processing conditions, which can be batch uploaded using standardized templates. Multiple experimental files can be uploaded simultaneously, enabling automated performance calculations and visualization. This automated workflow supports efficient integration and management of large-scale experimental datasets, thereby facilitating the application of artificial intelligence (AI) and machine learning (ML) techniques.
The dataset focuses on ternary organic photovoltaic systems composed of two donors and one acceptor. To accurately describe active-layer compositions, both the Donor1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Donor2 ratio and the overall (Donor1 + Donor2)
Donor2 ratio and the overall (Donor1 + Donor2)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Acceptor ratio are recorded, enabling precise and reproducible representation of multicomponent formulations (Fig. 2).
Acceptor ratio are recorded, enabling precise and reproducible representation of multicomponent formulations (Fig. 2).
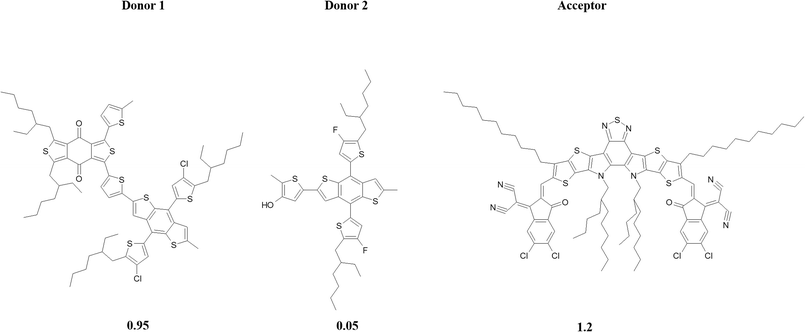 | ||
Fig. 2 Schematic representation of donor–donor–acceptor (D1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D2 D2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A) composition in a ternary organic photovoltaic system. A) composition in a ternary organic photovoltaic system. | ||
From the 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 430 devices stored on the established platform, non-fullerene data, data without missing values, and representative data selected by the experimenter from repeated experiments were selected. A dataset of 67 donors and 24 acceptors, totaling 1010 D–A combinations (Table S1), was utilized for machine learning. The data included the OPV device architectures, donor and acceptor material structures, D
430 devices stored on the established platform, non-fullerene data, data without missing values, and representative data selected by the experimenter from repeated experiments were selected. A dataset of 67 donors and 24 acceptors, totaling 1010 D–A combinations (Table S1), was utilized for machine learning. The data included the OPV device architectures, donor and acceptor material structures, D![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A ratios, device thicknesses, annealing conditions, and experimental PCE values (Fig. 3).
A ratios, device thicknesses, annealing conditions, and experimental PCE values (Fig. 3).
In this study, we fragmented large organic molecules into structural subunits and extracted the molecular descriptors for each fragment. Calculations were performed for each fragment to further understand the chemical properties of the polymer material. The device information, process conditions, and polymer ratio information (D![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) A) were added to the dataset. Furthermore, incorporating molecular symmetry, which has not been considered in previous studies, enables the extraction of structural characteristics that are not captured by conventional molecular descriptors. This approach enables the generation of diverse functions beyond device information and facilitates the processing of large molecules.
A) were added to the dataset. Furthermore, incorporating molecular symmetry, which has not been considered in previous studies, enables the extraction of structural characteristics that are not captured by conventional molecular descriptors. This approach enables the generation of diverse functions beyond device information and facilitates the processing of large molecules.
Because categorical features describing device structure and solvent type cannot be directly used in conventional machine learning models, they were converted into numerical values using label encoding. Integer labels were assigned as nominal identifiers without implying ordinal or physical meaning (Table S2). Missing values were imputed using the mode or median, depending on the feature.
2.2 Fragmentation of donors and acceptors
Most previous machine learning studies on organic photovoltaics (OPVs)30,31 relied on the polymer fingerprints of the donor and acceptor materials in the active layer. The materials used as the OPV donors and acceptors are often polymers, single molecules, or macromolecules. This makes it difficult to infer their physicochemical properties using molecular descriptors. To overcome this limitation, we employed a fragmentation strategy to extract fragment-specific properties from the intrinsic molecular structure and subsequently recombine them, enabling machine learning models to better reflect the chemical characteristics derived from the materials themselves.To digitally represent the donor and acceptor molecules in OPVs effectively, we developed our own fragmentation protocol. The fragmentation scheme was designed to simplify the polymer structures while retaining their essential chemical features, thereby producing independent fragments that contribute meaningfully to the overall molecular properties (Fig. 4).
Method 1: the backbone and side chains were explicitly separated and molecular descriptors were calculated independently for each structural unit.
Method 2 (scaffold-based fragmentation): the ring units that constitute the backbone are defined as scaffolds, and each scaffold represents a ring system connected by a single bond. Typically, a donor consists of two to eight scaffolds. For consistency, we standardized this number to four scaffolds. Each scaffold was further divided into a backbone and side chains.
(1) Ring center (RC): the center of symmetry of the FR system.
(2) Ring linker (RL): the ring units adjacent to the RC.
(3) Ring π-bridge (RP): the π-bridge ring bonded to the outer ring; in some cases, the RP is absent.
(4) π-Bridge (PB): the π-conjugated structure connecting the FR system and the end group; in some cases, the PB is considered a part of the FR system.
(5) End group (EG): terminal substituents located at the outermost positions of the acceptor molecule.
Through this fragmentation process, the acceptor molecule structure was defined as RC–RL–RP–PB–EG, centered on the central fused-ring system.
2.3 Extraction of molecular descriptors
The selection of appropriate molecular descriptors is a crucial step in developing reliable machine learning (ML) models. Considering computational efficiency and predictive performance, it is essential to employ descriptors that are both relevant and effective.Accordingly, descriptors for each fragment unit were extracted using the RDKit and Stk libraries. For both Method 1 and Method 2, 217 physicochemical descriptors were computed using RDKit, whereas seven additional energy-related properties were obtained from Stk. These descriptors describe a wide range of molecular characteristics, ranging from basic molecular properties to complex structural characteristics. The complete list and aggregation methods are provided in Table S3.
2.4 Knowledge-based feature selection
Some molecules did not contain certain fragments and were therefore zero-coded. To mitigate this gap effect, pairwise computations between descriptors were performed.
This allowed us to maintain the effectiveness of each fragment-specific descriptor while compensating for the gap effect in fragment-free regions, resulting in molecular descriptors that utilize partial structures. The detailed computational operations for each descriptor type are provided in SI Table S3.
In this paper, local symmetry is defined in a restricted sense as either two-fold rotational (C2) or mirror-plane (σv) symmetry, as illustrated in Fig. 5. For donor molecules, local symmetry describes the symmetry of side-chain groups with respect to the molecular backbone. Donors are categorized into four scaffolds, yielding four symmetry states per donor; when two donor components are present, this results in a total of eight donor symmetry states. For acceptor molecules, local symmetry characterizes the symmetry of the end groups with respect to the ring center, with an additional symmetry state assigned at the ring center, leading to twenty symmetry states in total (Table S4).
The symmetry descriptors employed in this study are molecular-level features derived from specific structural motifs, rather than explicit representations of solid-state packing. Although thin-film packing is governed by processing conditions and intermolecular interactions, these symmetry states capture structural characteristics that may indirectly relate to packing-relevant tendencies, while remaining general and scalable for machine-learning models.
2.5 Selection of machine learning algorithms
All machine-learning (ML) models were implemented within a unified Python framework using molecular descriptors generated with RDKit. XGBoost was selected as the primary model, and its performance was compared with random forest (RF), support vector regression (SVR), LightGBM (LGBM), and multilayer perceptron (MLP) models. Model performance was evaluated using the coefficient of determination (R2) under 5-fold cross-validation.The dataset comprised 1010 experimentally reported OPV devices and was split into training (80%) and test (20%) sets. To assess model robustness and minimize bias from a single data split, eight independent random seeds were used for repeated training and evaluation.
The initial feature space consisted of 1054 molecular descriptors, resulting in a high feature-to-sample ratio. To reduce overfitting and computational cost, recursive feature elimination (RFE) was applied, with feature selection guided by cross-validated R2. Feature removal was terminated once performance stabilized over consecutive iterations. Performance convergence was observed when approximately 90 descriptors remained (Fig. 6a). Within this region, the final feature set was selected by identifying the subset that achieved the highest cross-validated R2 while maintaining consistent descriptor composition across successive RFE steps. Based on this criterion, 70 descriptors were retained for the final models.
 | ||
| Fig. 6 (a) Recursive feature elimination (RFE); (b) train and test dataset distribution in OPV prediction. | ||
This reduction in feature dimensionality led to a substantial decrease in training time (from 101.7 s to 10.5 s) while maintaining comparable predictive performance (R2 of 0.848 and 0.839 before and after feature selection, respectively). The final descriptor sets for Methods 1 and 2 are summarized in Tables S5 and S6.
A Y-scrambling test was performed to assess dataset reliability. The original models achieved an average R2 of 0.81, whereas the scrambled models yielded R2 values close to zero (Fig. S8), indicating that the predictions arose from genuine structure–property relationships rather than spurious correlations. Kernel density estimation (KDE) analysis showed similar PCE distributions for the training (n = 808) and test (n = 202) sets, confirming statistically balanced data partitioning. The small positive density near PCE = 0 originates from the smoothing nature of KDE rather than the presence of zero-valued samples (Fig. 6b).
To assess multicollinearity among the selected descriptors, Pearson correlation analysis was performed on the final 70-feature set. The mean and median absolute correlation coefficients (|r|) were 0.344 and 0.295, respectively, indicating moderate overall correlations. Of the 2415 possible descriptor pairs, only 25 pairs (≈1.0%) exhibited very high correlation (|r| > 0.9), suggesting limited redundancy and minimal multicollinearity in the final feature set (Fig. 7).
| Random state | Method 1 | Method 2 |
|---|---|---|
| 0 | 0.879 | 0.841 |
| 42 | 0.854 | 0.885 |
| 150 | 0.782 | 0.842 |
| 500 | 0.892 | 0.841 |
| 790 | 0.847 | 0.758 |
| 1000 | 0.826 | 0.854 |
| 7500 | 0.881 | 0.830 |
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
0.829 | 0.820 |
| Average R2 | 0.849 ± 0.034 | 0.834 ± 0.034 |
The similar performance of the two strategies suggests that, for large donor–acceptor molecules where conventional RDKit descriptors are insufficient, fragmentation into smaller chemically meaningful units followed by recombination yields informative representations that enhance machine-learning training. Model robustness and generalization were further assessed using 5-fold cross-validation (Tables S8 and S9).
Table 4 summarizes the performance of the five machine-learning models using Method 1. XGBoost and RF achieved the highest average R2 values (0.849), followed by LGBM (0.812) and MLP (0.785), which exhibited moderate performance. SVR showed comparatively lower performance (0.718), consistent with its limited scalability in high-dimensional feature spaces.
| Random state | XGB | MLP | RF | SVR | LGBM |
|---|---|---|---|---|---|
| 0 | 0.879 | 0.760 | 0.877 | 0.713 | 0.831 |
| 42 | 0.854 | 0.811 | 0.869 | 0.725 | 0.828 |
| 150 | 0.782 | 0.770 | 0.796 | 0.657 | 0.800 |
| 500 | 0.892 | 0.845 | 0.881 | 0.818 | 0.841 |
| 790 | 0.847 | 0.735 | 0.838 | 0.657 | 0.812 |
| 1000 | 0.824 | 0.752 | 0.823 | 0.692 | 0.780 |
| 7500 | 0.881 | 0.866 | 0.874 | 0.813 | 0.853 |
10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
0.829 | 0.738 | 0.829 | 0.650 | 0.748 |
| Average R2 | 0.849 ± 0.034 | 0.785 ± 0.047 | 0.849 ± 0.029 | 0.670 ± 0.054 | 0.812 ± 0.032 |
Repeated experiments across eight random states yielded low standard deviations (0.029–0.054), indicating stable predictions with minimal sensitivity to data partitioning and a low risk of overfitting. These results are visualized using violin plots in Fig. 8.
Using the experimental dataset, we further evaluated the model predictions for PCE and additionally explored the applicability of the model to Jsc and FF as auxiliary targets. The distribution of PCE prediction errors approximated a Gaussian profile, indicating minimal systematic bias (Fig. 9a). Scatter plots comparing experimental and predicted values further illustrate the predictive capability of the model, yielding R2 values of 0.69 for Jsc and 0.71 for FF (Fig. 9b–d). The corresponding prediction errors, quantified by MAE and RMSE, are summarized in Table S10 for the XGBoost model evaluated on the test set.
In contrast, predictions for the open-circuit voltage (Voc) showed relatively low accuracy (R2 = 0.2–0.4). This limitation32–34 is attributed to the descriptor set used in this study, which primarily captures molecular and electronic structure information but does not explicitly account for interfacial disorder, charge–transfer state energetics, or nonradiative recombination processes.
Finally, a comparison with two previous studies on fragmentation methods highlights the methodological novelty of our approach (Table 5). Rather than directly comparing identical datasets, this analysis focuses on differences in molecular representation and feature construction strategies. Kim et al.15 used predefined, synthesis-oriented molecular fragments that efficiently organize structural diversity but treat fragments largely independently, whereas Wu et al.14 employed a function-driven fragmentation scheme based on the electron push–pull principle, emphasizing electronic roles with limited structural uniformity across molecular classes.
In contrast, our study adopts a chemically meaningful fragmentation framework that incorporates explicit local symmetry information and integrates fragment-level features into a unified molecular representation. Combined with experimentally measured device parameters and polymer-specific chemical descriptors, this approach simultaneously captures electronic, structural, and processing effects, leading to improved predictive accuracy and enhanced generalizability. Our model was evaluated using a held-out test set from an in-house experimental database, while the results from Kim et al.15 and Wu et al.14 were taken from their respective publications, each based on independently constructed datasets and evaluation protocols.
To further assess the generalization ability of the model, 130 data35–64 points independently collected from the literature and not used during model training were extracted and evaluated using the same XGBoost model. Despite the inherent heterogeneity of literature data, including variations in material combinations and experimental conditions, the model achieved an R2 of 0.62, an MAE of 2.11, and an RMSE of 2.63, indicating reasonable predictive performance on unseen data (Fig. S9).
2.6 Feature importance analysis using SHAP
To enhance interpretability and quantitatively assess the contribution of individual features to PCE predictions, we performed a SHAP (SHapley Additive exPlanations) analysis on the XGBoost model trained using Method 1.The SHAP analysis (Fig. 10) identifies key features governing PCE prediction, which are primarily associated with molecular stability, local electronic structure, and processing conditions, highlighting the multiscale nature of performance-determining factors in organic photovoltaic devices.
Among the top five features, RC_FpDensityMorgan1 shows the strongest impact, indicating that dense and electronically coherent core environments favor efficient charge transport and reduced energetic disorder. 1_fr_halogen further highlights the positive role of halogen substitution, consistent with enhanced intermolecular interactions and frontier orbital tuning. PB_GasteigerChargeMedian emphasizes the importance of balanced local charge distribution in promoting charge separation while suppressing recombination. The prominence of solution concentration (wt%) reflects its role as a key processing parameter that mediates thin-film formation and structure–property coupling, with its influence becoming significant in specific molecular contexts. RC_VSA_EState6 indicates that exposed electronic environments at the molecular core critically affect interfacial charge–transfer processes.
Beyond intrinsic molecular descriptors, several processing parameters including concentration, thermal annealing temperature, solvent type, device structure, and active-layer thickness also appear among the influential features. Although their average SHAP magnitudes are smaller, these parameters can become decisive depending on molecular structure, suggesting that they act as conditional modifiers of morphology, crystallinity, and charge–transport pathways. Notably, symmetry-related descriptors also contribute non-negligibly, implying that fragment-level local symmetry influences molecular packing regularity and orientational degeneracy.
Consistent with the SHAP results, the feature-importance rankings obtained from the XGBoost, RF, and LGBM models also highlight RC_FpDensityMorgan1 and 1_fr_halogen as major contributors (Fig. S10 and S11), supporting the reliability of the SHAP-based interpretation across different learning algorithms.
Overall, SHAP analysis demonstrates that PCE is governed by a synergistic interplay between molecular electronic structure, topology, symmetry, and processing conditions. The fragment-based, symmetry-aware descriptor framework effectively captures these coupled effects, providing both high predictive accuracy and physically interpretable structure–process property relationships.
3 Conclusions
In this study, we established an OPV data platform to systematically collect experimental data and developed machine-learning models for predicting device performance. Fragment-based representations of donor and acceptor molecules were constructed using two fragmentation strategies, and 224 physicochemical descriptors were generated using RDKit and Stk.Among five machine-learning models evaluated, XGBoost showed the best performance with an R2 of 0.849. Robust validation using five-fold cross-validation and multiple random states confirmed the reliability of the models. SHAP analysis identified local molecular symmetry as a key factor influencing PCE, highlighting its important role in OPV performance.
Future work will extend this framework by refining symmetry definitions and integrating external literature data to explore new high-efficiency donor–acceptor combinations.
Author contributions
J. L.: conceptualization, data curation, formal analysis, methodology, writing – original draft. H. B.: machine learning modeling, methodology, writing – review & editing. H. S.: code review, validation. H. L.: experimental investigation, sample preparation. F. A.: materials synthesis, experimental data analysis. D. K.: conceptualization, supervision, project administration. All authors contributed to manuscript review and revision.Conflicts of interest
The authors declare no competing interests.Data availability
Code availability: https://github.com/juhyun7749/OPV-ML-2025.The computational codes developed and used for this study are openly accessible via GitHub at [https://github.com/juhyun7749/OPV-ML-2025].
To ensure long-term preservation and reproducibility, a fixed version of the repository corresponding to the manuscript has been archived in Zenodo (DOI: https://doi.org/10.5281/zenodo.18688051).
All relevant datasets required to reproduce the analyses and figures presented in this article are included in the Zenodo archive.
The archived materials are provided under an open license to facilitate reuse and future research based on this work. The calculated molecular descriptors for the OPV donor and acceptor materials used in this study are openly available.
Supplementary information (SI): additional figures, model details, descriptor information, and dataset descriptions used in this study. See DOI: https://doi.org/10.1039/d5dd00496a.
References
- C. W. Tang, Two-layer organic photovoltaic cell, Appl. Phys. Lett., 1986, 48, 183–185, DOI:10.1063/1.96937.
- N. M. O'Boyle, C. M. Campbell and G. R. Hutchison, Computational design and selection of optimal organic photovoltaic materials, J. Phys. Chem. C, 2011, 115, 16200–16210, DOI:10.1021/jp202977x.
- C. Zanlorenzi and L. Akcelrud, L. Morphological and photophysical properties of polyfluorene copolymers containing quinoxaline units, J. Polym. Sci., Part B: Polym. Phys., 2017, 55, 919, DOI:10.1002/polb.24348.
- M. J. S. Dewar, E. G. Zoebisch, E. F. Healy and J. J. P. Stewart, Development and use of quantum mechanical molecular models. 76. AM1: a new general purpose quantum mechanical molecular model, J. Am. Chem. Soc., 1985, 107, 3902–3909, DOI:10.1021/ja00299a024.
- J. J. P. Stewart, Optimization of parameters for semiempirical methods v: modification of NDDO approximations and application to 70 elements, J. Mol. Model., 2007, 13, 1173–1213, DOI:10.1007/s00894-007-0233-4.
- J. J. P. Stewart, Optimization of parameters for semiempirical methods vi: more modifications to the NDDO approximations and re-optimization of parameters, J. Mol. Model., 2013, 19, 1–32, DOI:10.1007/s00894-012-1667-x.
- T. Fink, H. Bruggesser and J.-L. Reymond, Virtual exploration of the small-molecule chemical universe below 160 Daltons, Angew. Chem., 2005, 44, 1504–1508, DOI:10.1002/anie.200462457.
- J. L. Reymond, R. van Deursen, L. C. Blum and L. Ruddigkeit, Chemical space as a source for new drugs, MedChemComm, 2010, 1, 30–38, 10.1039/c0md00020e.
- T. Le Bahers, et al., Modeling dye-sensitized solar cells: from theory to experiment, J. Phys. Chem. Lett., 2013, 4, 1044–1050, DOI:10.1021/jz4001823.
- N. Martsinovich and A. Troisi, High-throughput computational screening of chromophores for dye-sensitized solar cells, J. Phys. Chem. C, 2011, 115, 11781–11792, DOI:10.1021/jp202827r.
- V. Venkatraman and B. K. Alsberg, Construction of thermally stable 3,6-disubstituted spiro-fluorene derivatives as host materials for blue phosphorescent organic light-emitting diodes, Dyes Pigm., 2015, 114, 69, DOI:10.1016/j.dyepig.2014.11.011.
- S. Tortorella, F. De Angelis and G. Cruciani, Quantitative structure-property relationship modeling of small organic molecules for solar cells applications 32, J. Chemom., 2018, e2957, DOI:10.1002/cem.2957.
- S. A. Lopez, et al., The Harvard organic photovoltaic dataset, Sci. Data, 2016, 3, 160086, DOI:10.1038/sdata.2016.86.
- Y. Wu, J. Guo, R. Sun and J. Min, Machine learning for accelerating the discovery of high-performance donor/acceptor pairs in non-fullerene organic solar cells, npj Comput. Mater., 2020, 6, 120, DOI:10.1038/s41524-020-00388-2.
- G.-H. Kim, C. Lee, K. Kim and D.-H. Ko, Novel structural feature-descriptor platform for machine learning to accelerate the development of organic photovoltaics, Nano Energy, 2023, 106, 108108, DOI:10.1016/j.nanoen.2022.108108.
- J. Hachmann, et al., The Harvard Clean Energy Project: large-scale computational screening and design of organic photovoltaics on the world community grid, J. Phys. Chem. Lett., 2011, 2, 2241–2251, DOI:10.1021/jz200866s.
- S. A. Lopez, B. Sanchez-Lengeling, J. de Goes Soares and A. Aspuru-Guzik, Design principles and top non-fullerene acceptor candidates for organic photovoltaics, Joule, 2017, 1, 857–870, DOI:10.1016/j.joule.2017.10.006.
- Z.-W. Zhao, M. del Cueto, Y. Geng and A. Troisi, Effect of increasing the descriptor set on machine learning prediction of small molecule-based organic solar cells, Chem. Mater., 2020, 32, 7777–7787, DOI:10.1021/acs.chemmater.0c02325.
- Z.-W. Zhao, M. del Cueto and A. Troisi, Limitations of machine learning models when predicting compounds with completely new chemistries: possible improvements applied to the discovery of new non-fullerene acceptors, Digit. Discov., 2022, 1, 595–610, 10.1039/D2DD00004K.
- H. Sahu, W. Rao, A. Troisi and H. Ma, Toward predicting efficiency of organic solar cells via machine learning and improved descriptors, Adv. Energy Mater., 2018, 8, 1801032, DOI:10.1002/aenm.201801032.
- Y. Wen, et al., Simultaneous optimization of donor/acceptor pairs and device specifications for nonfullerene organic solar cells, Nat. Commun., 2023, 14, 4524, DOI:10.1038/s41467-023-40032-3.
- H. Jang, et al., Deep learning-based design of high performance organic solar cells, ACS Appl. Mater. Interfaces, 2020, 12, 10346–10354, DOI:10.1021/acsami.9b21349.
- X. Ma, et al., Machine learning-assisted screening of polymer donor materials for high-performance non-fullerene organic solar cells, Adv. Sci., 2020, 7, 2001776, DOI:10.1002/advs.202001776.
- Q. Ling, High-Throughput Molecular Design of Donors and Non-Fullerene Acceptors for Organic Solar Cells Based on Convolutional Neural Networks, J. Chem. Inf. Model., 2025, 65, 10107–10123, DOI:10.1021/acs.jcim.5c01634.
- L. Zhu, The Key Descriptors for Predicting the Exciton Binding Energy of Organic Photovoltaic Materials, Angew. Chem., Int. Ed., 2025, 64, e202413913, DOI:10.1002/anie.202413913.
- P. Raccuglia, et al., Machine-learning-assisted materials discovery using failed experiments, Nature, 2016, 533, 73–76, DOI:10.1038/nature17439.
- H. Li Jin, Machine learning study of D:A1:A2 ternary organic solar cells, Org. Electron., 2024, 125, 106988, DOI:10.1016/j.orgel.2023.106988.
- ChemAxon, Marvin JS and JChem Microservices, ChemAxon Ltd., Budapest, Hungary, https://chemaxon.com Search PubMed.
- E. Willighagen, et al., The Chemistry Development Kit (CDK) v2.0: atom typing, depiction, molecular formulas, and substructure searching, J. Cheminf., 2017, 9, 33 Search PubMed.
- Z. Zhu, et al., Machine-learning-assisted exploration of new non-fullerene acceptors for high-efficiency organic solar cells, Cell Rep. Phys. Sci., 2024, 5, 102316, DOI:10.1016/j.xcrp.2024.102316.
- O. A. Álvarez-Gonzaga, U. A. Vergara-Beltran and J. I. Rodríguez, Machine learning models with different cheminformatics data sets to forecast the power conversion efficiency of organic solar cells, arXiv, 2024, preprint, 10.48550/arXiv.2410.23444 Search PubMed.
- T. Upreti, S. Wilken, H. Zhang and M. Kemerink, Slow relaxation of photogenerated charge carriers boosts open-circuit voltage of organic solar cells, J. Phys. Chem. Lett., 2021, 12, 9874–9881, DOI:10.1021/acs.jpclett.1c02235.
- M. Azzouzi, et al., Nonradiative energy losses in bulk-heterojunction organic photovoltaics, Phys. Rev. X, 2018, 8, 031055, DOI:10.1103/PhysRevX.8.031055.
- N. K. Elumalai and A. Uddin, Open circuit voltage of organic solar cells: an in-depth review, Energy Environ. Sci., 2016, 9, 391–410, 10.1039/C5EE02871J.
- X. Dong, et al., Large-Area Organic Solar Modules with Efficiency Over 14%, Adv. Funct. Mater., 2022, 32, 2110209, DOI:10.1002/adfm.202110209.
- W. Zhu, et al., Crystallography, Morphology, Electronic Structure, and Transport in Non-Fullerene/Non-Indacenodithienothiophene Polymer:Y6 Solar Cells, J. Am. Chem. Soc., 2020, 142, 14532–14547, DOI:10.1021/jacs.0c05560.
- R. Wang, et al., Rational Tuning of Molecular Interaction and Energy Level Alignment Enables High-Performance Organic Photovoltaics, Adv. Mater., 2019, 31(43), 1904215, DOI:10.1002/adma.201904215.
- R. Yu, et al., Improved Charge Transport and Reduced Nonradiative Energy Loss Enable Over 16% Efficiency in Ternary Polymer Solar Cells, Adv. Mater., 2019, 31(36), 1902302, DOI:10.1002/adma.201902302.
- X. Liu, et al., Non-Halogenated Polymer Donor-Based Organic Solar Cells with a Nearly 15% Efficiency Enabled by a Classic Ternary Strategy, ACS Appl. Energy Mater., 2021, 4(2), 1774–1783, DOI:10.1021/acsaem.0c02912.
- X. Xu, et al., Developing Wide Bandgap Polymers Based on Sole Benzodithiophene Units for Efficient Polymer Solar Cells, Chem.–Eur. J., 2020, 26(49), 11241–11249, DOI:10.1002/chem.202000951.
- M. H. Yang, et al., Roll-to-Roll Compatible Quinoxaline-Based Polymers toward High-Performance Polymer Solar Cells, J. Mater. Chem. A, 2020, 8(47), 25208–25216, 10.1039/D0TA09354H.
- P. Chao, et al., A Benzo[1,2-b:4,5-c′]dithiophene-4,8-Dione-Based Polymer Donor Achieving an Efficiency over 16%, Adv. Mater., 2020, 32(10), 1907059, DOI:10.1002/adma.201907059.
- X. Huang, et al., Novel Narrow Bandgap Terpolymer Donors Enable Record Performance for Semitransparent Organic Solar Cells Based on All-Narrow Bandgap Semiconductors, Adv. Funct. Mater., 2021, 32, 2108634, DOI:10.1002/adfm.202108634.
- Z. Wang, et al., High Power Conversion Efficiency of 13.61% for 1 cm2 Flexible Polymer Solar Cells Based on Patternable and Mass-Producible Gravure-Printed Silver Nanowire Electrodes, Adv. Funct. Mater., 2021, 3(4), 2007276, DOI:10.1002/adfm.202007276.
- X. Dong, et al., Large-Area Organic Solar Modules with Efficiency over 14%, Adv. Funct. Mater., 2022, 32(15), 2110209, DOI:10.1002/adfm.202110209.
- Y. Chen, et al., Asymmetric Alkoxy and Alkyl Substitution on Nonfullerene Acceptors Enabling High-Performance Organic Solar Cells, Adv. Energy Mater., 2021, 11(3), 2003141, DOI:10.1002/aenm.202003141.
- J. Yuan, et al., Patterned Blade Coating Strategy Enables Enhanced Device Reproducibility and Optimized Morphology of Organic Solar Cells, Adv. Energy Mater., 2021, 11(18), 2100098, DOI:10.1002/aenm.202100098.
- K.-E. Hung, et al., Non-Volatile Perfluorophenyl-Based Additive for Enhanced Efficiency and Thermal Stability of Nonfullerene Organic Solar Cells via Supramolecular Fluorinated Interactions, Adv. Energy Mater., 2022, 12(12), 2103702, DOI:10.1002/aenm.202103702.
- W.-Z. Fo, et al., Highly Efficient Binary Solvent Additive-Processed Organic Solar Cells by the Blade-Coating Method, Macromol. Chem. Phys., 2021, 222(17), 2100062, DOI:10.1002/macp.202100062.
- R. Ma, et al., Improving Open-Circuit Voltage by a Chlorinated Polymer Donor Endows Binary Organic Solar Cells Efficiencies over 17%, Sci. China: Chem., 2020, 63(3), 325–330, DOI:10.1007/s11426-019-9669-3.
- Z. Luo, et al., Altering Alkyl-Chain Branching Positions for Boosting the Performance of Small-Molecule Acceptors for Highly Efficient Nonfullerene Organic Solar Cells, Sci. China: Chem., 2020, 63(3), 361–369, DOI:10.1007/s11426-019-9670-2.
- J. Yuan, et al., Single-Junction Organic Solar Cell with over 15% Efficiency Using a Fused-Ring Acceptor with an Electron-Deficient Core, Joule, 2019, 3(4), 1140–1151, DOI:10.1016/j.joule.2019.01.004.
- K. Jiang, et al., Alkyl Chain Tuning of Small Molecule Acceptors for Efficient Organic Solar Cells, Joule, 2019, 3(12), 3020–3033, DOI:10.1016/j.joule.2019.09.010.
- F. Lin, et al., A Non-Fullerene Acceptor with Enhanced Intermolecular π-Core Interaction for High-Performance Organic Solar Cells, J. Am. Chem. Soc., 2020, 142(36), 15246–15251, DOI:10.1021/jacs.0c07083.
- D. Li, et al., Enhanced and Balanced Charge Transport Boosting Ternary Organic Solar Cells over 17% Efficiency, Adv. Mater., 2020, 32(34), 2002344, DOI:10.1002/adma.202002344.
- K. Li, et al., Ternary Blended Fullerene-Free Polymer Solar Cells with 16.5% Efficiency Enabled by a Higher-LUMO-Level Acceptor to Improve Film Morphology, Adv. Energy Mater., 2019, 9(33), 1901728, DOI:10.1002/aenm.201901728.
- K. Yu, et al., Achieving 18.14% Efficiency of Ternary Organic Solar Cells with an Alloyed Nonfullerene Acceptor, Small Struct., 2021, 2(11), 2100099, DOI:10.1002/sstr.202100099.
- M.-A. Pan, et al., 16.7%-Efficiency Ternary Blended Organic Photovoltaic Cells with PCBM as an Acceptor Additive to Increase Open-Circuit Voltage and Phase Purity, J. Mater. Chem. A, 2019, 7(36), 20713–20722, 10.1039/C9TA06929A.
- Y. Cho, et al., Guest-Oriented Non-Fullerene Acceptors for Ternary Organic Solar Cells with over 16.0% and 22.7% Efficiencies under One-Sun and Indoor Light, Nano Energy, 2020, 75, 104896, DOI:10.1016/j.nanoen.2020.104896.
- T. Wang, et al., Solution-Processed Polymer Solar Cells with over 17% Efficiency Enabled by an Iridium Complexation Approach, Adv. Energy Mater., 2020, 10(22), 2000590, DOI:10.1002/aenm.202000590.
- N. Nakao, et al., Pronounced Backbone Coplanarization by π-Extension in a Sterically Hindered Conjugated Polymer System Leads to Higher Photovoltaic Performance in Non-Fullerene Solar Cells, ACS Appl. Mater. Interfaces, 2021, 13(47), 56420–56429, DOI:10.1021/acsami.1c17199.
- G. Zhang, et al., Naphthalenothiophene Imide-Based Polymer Exhibiting over 17% Efficiency in Organic Solar Cells, Joule, 2021, 5(4), 931–944, DOI:10.1016/j.joule.2021.02.003.
- X. Xu, et al., Single-Junction Polymer Solar Cells with 16.35% Efficiency Enabled by a Platinum(II) Complexation Strategy, Adv. Mater., 2019, 31(29), 1901872, DOI:10.1002/adma.201901872.
- N. Nakao, et al., Triphenyleno[1,2-c:7,8-c′]bis([1,2,5]thiadiazole) as a V-Shaped Electron-Deficient Unit to Construct Wide-Bandgap Amorphous Polymers for Efficient Organic Solar Cells, ACS Appl. Mater. Interfaces, 2021, 13(47), 56420–56429, DOI:10.1021/acsami.1c19708.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |








