Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
When machine learning models learn chemistry II: applying WISP to real-world examples
Kerrin
Janssen
a,
Jan M.
Wollschläger
 b,
Jonny
Proppe
*a and
Andreas H.
Göller
b,
Jonny
Proppe
*a and
Andreas H.
Göller
 *c
*c
aTU Braunschweig, Institute of Physical and Theoretical Chemistry, Gauss Str 17, 38106 Braunschweig, Germany. E-mail: j.proppe@tu-braunschweig.de
bBayer AG, Pharmaceuticals, R&D, Machine Learning Research, 13353 Berlin, Germany
cBayer AG, Pharmaceuticals, R&D, Computational Molecular Design, 42096 Wuppertal, Germany. E-mail: andreas.goeller@bayer.com
First published on 9th February 2026
Abstract
In our previous work, we introduced WISP (Workflow for Interpretability Scoring using matched molecular Pairs), which enables users to quantitatively assess the performance of explainability methods for machine learning models. In this work, we focus on more complex tasks, such as yield prediction, pKi values for inhibition of coagulation Factor Xa and AMES mutagenicity, where the explanations of the predicted property need to capture more intricate interaction patterns between structural motifs of either the reaction partner, the protein or DNA. Expanding upon part I of the “When Machine Learning Learns Chemistry” series, we demonstrate additional functionalities of the WISP workflow. Alongside the model and descriptor—agnostic atom attributor, WISP integrates a SHAP-based and an RDKit-based attribution method, enabling the comparison of multiple explainability approaches, as demonstrated on the Factor Xa dataset. This work also showcases WISP's capability to evaluate explanations for classification tasks such as AMES mutagenicity. The application of WISP to the AMES mutagenicity dataset reveals that the respective machine learning model fails to learn the underlying chemical relationships, instead relying primarily on numerical correlations. When applied to the yield dataset, WISP highlights specific cases where explainability methods that usually perform well fail to provide meaningful insights. This highlights WISP's ability to detect such limitations in trained models, providing valuable insights to guide targeted improvements in model development and data quality.
1 Introduction
By providing meaningful explanations for predictions of machine learning models, these models can be leveraged to guide molecular design decisions in the life sciences.1–3 This is often achieved in the form of heatmaps that highlight the contributions of individual molecular atoms, indicating whether they influence a property of interest positively or negatively (Fig. 1).4–9 Such visualizations can be valuable to both machine learning experts and non-experts.5,10 For example, in computer-aided synthesis planning, such explanations can help identify functional groups that lower reaction yields, allowing chemists to replace them with more favorable alternatives.11,12 In drug design, they can pinpoint molecular regions that reduce bioactivity or increase toxicity, which can then be modified or removed to improve a compound's properties.13,14A key limitation of current explainability methods in chemistry is that single-atom attributions often lack chemical plausibility, making it difficult to trust or interpret the explanations meaningfully. For example, highlighting a single carbon atom within a phenyl ring provides no actionable insight, as this does not reflect the delocalized electronic nature of the system or inform any chemically sensible design choices a chemist could make. To address this issue, we proposed WISP (a Workflow for Interpretability Scoring using matched molecular Pairs) to evaluate XAI attribution schemes.15 To evaluate the atom attributions produced by the explainability methods, we applied the concept of matched molecular pairs (MMPs). By comparing atom attributions specifically on the variable and constant parts of MMPs, this approach ensures that any structural change assessed is chemically meaningful and interpretable. Since modifications within an MMP are inherently chemically plausible, this method provides a robust basis for validating whether atomic contributions align with established chemical structure–property relationships.16
In part I of the “When Machine Learning Models Learn Chemistry” series, we introduced WISP as a model- and descriptor-agnostic tool to quantify the chemical explainability capabilities of machine learning models.15 Using datasets with well-defined physicochemical properties, we demonstrated how WISP can help identify robust models that are capable of providing reliable explanations for unseen data.15
In this work, we apply WISP to tasks relevant for computer-aided synthesis planning and drug discovery, demonstrating its application to more complex endpoints. We highlight the implications of these findings for model and dataset development and discuss how insights into explainability can inform decision-making in life sciences.
2 Methods
2.1 Datasets
To evaluate the explainability performance on a dataset relevant to computer-aided synthesis planning, we used a dataset containing liquid chromatography area percent (LCAP) yields.11,17 These yields stem from a mild, functional-group-tolerant synthesis protocol developed by Felten et al., which involves the carboxylation of 1,3-azoles followed by an amide coupling step.17 With 264 LCAP yields, this is the smallest dataset used in this work (Table 1).For the domain of drug design, we employed two datasets: one consisting of 1068 pKi values for inhibition of coagulation Factor Xa, and another containing 6130 AMES mutagenicity measurements (Table 1).18,19
For the preprocessing, we applied the same standardizing protocol as in our previous work:15
• Filter molecules to maximally 1000 atoms.
• Enumerate up to 10 tautomers for canonicalization.
• Retain only the largest fragment per SMILES.
• Perform normalization and sanitization.
• Remove SMILES duplicates with conflicting experimental target values; keep one entry when targets agree.
This uniform preprocessing ensures that all molecular representations are standardized and comparable.
2.2 WISP specifications
In this work, we also made use of several additional features included in WISP. The general workflow of WISP is described in detail in part I of this series. It includes a preprocessing step followed by model training, supporting both various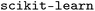 models and
models and 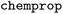 models. Subsequently, attributions are calculated and evaluated using matched molecular pairs (MMPs).
models. Subsequently, attributions are calculated and evaluated using matched molecular pairs (MMPs).
WISP supports the evaluation of model explanations for classification tasks. To this end, it integrates a training routine that combines Morgan fingerprints with a random forest classifier, which we applied here to the AMES dataset. To enable the same explainability assessment as for regression tasks, the model outputs are provided as class probabilities. The maximum heatmap color intensity is set to 0.7 in this case.
In the Python version of WISP, unlike in the web application, users can also include a pre-trained model within the workflow. This option is particularly valuable for machine learning users with specific requirements for model training that fall outside the scope of WISP's built-in routines. We demonstrate this feature using the LCAP dataset.
In addition to the atom attributor, WISP integrates two further explainability methods.15 While these methods are less generally applicable than the atom attributor, their integration allows for a direct comparison of different explainability approaches within the WISP framework.
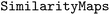 module as
module as 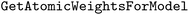 .24 This approach developed by Riniker and Landrum is model-agnostic and works with both Morgan and RDK fingerprints.24 The method determines atom attributions by identifying the fingerprint bits set by the atom of interest, removing those bits, and recalculating the model prediction using the modified fingerprint.24 One limitation of this method is that potential bit collisions can influence the attributions, which may partly explain why this explainability approach sometimes fails to capture the effects of specific functional groups.24 Bit collisions occur when multiple structural motifs are hashed into the same bit in the fingerprint.
.24 This approach developed by Riniker and Landrum is model-agnostic and works with both Morgan and RDK fingerprints.24 The method determines atom attributions by identifying the fingerprint bits set by the atom of interest, removing those bits, and recalculating the model prediction using the modified fingerprint.24 One limitation of this method is that potential bit collisions can influence the attributions, which may partly explain why this explainability approach sometimes fails to capture the effects of specific functional groups.24 Bit collisions occur when multiple structural motifs are hashed into the same bit in the fingerprint.
3 Results and discussion
3.1 Model performances
In this study, we employed the machine learning model previously trained and evaluated on a dataset of 264 liquid chromatography area percent (LCAP) yields, published by Felten et al.11,17The models for Factor Xa and AMES were trained entirely within the WISP workflow, resulting in models based on Morgan fingerprints and a random forest architecture (Table 2).18,19
| Property of interest | Model type | r 2 | MAE | RMSE | AEmax | Model source |
|---|---|---|---|---|---|---|
| LCAP yield | RDK fingerprint; Bayesian ridge | 0.69 | 7.4 | 9.9 | 23.6 | 11 |
| Factor Xa | Morgan fingerprint; random forest | 0.63 | 0.52 | 0.73 | 2.43 | WISP |
| Property of interest | Model type | Accuracy | Precision | Recall | CM | Model source |
|---|---|---|---|---|---|---|
| AMES | Morgan fingerprint; random forest | 0.80 | 0.79 | 0.80 | 406![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 141 141 |
WISP |
110![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 568 568 |
3.2 LCAP yield
Yield prediction plays an important role in computer-aided synthesis planning (CASP).25 It helps conserve resources and significantly reduces time requirements.25Since this model is trained on RDKit path-based fingerprints, the atom attribution method and the RDKit attribution method can be applied. However, the WISP workflow does not include an evaluation of train-test dependency when a pre-trained model is provided, as is the case here.
For this model, the atom-based explanations over the entire molecule yield a relatively high correlation with the predicted differences, with an r2 of 0.92 (Table 3). However, this correlation drops significantly when considering only the variable part of the MMP, as shown in Table 3. This drop in performance is primarily due to MMPs where a nitrile group is replaced with a methyl ester (Fig. 2, middle), which deviates notably from the previously observed correlation of the other MMP attributions with the predictions. However, such an imbalance does not seem to affect other transformations, such as the exchange of a methoxy group for a hydrogen atom (Fig. 2, bottom), where the MMP exchange shows no apparent influence on the prediction, a trend that is consistently reflected in the explainability results. These findings suggest that, while uncritical for most transformations, the model does not capture the underlying chemistry of specific transformations and their influence on reaction yield. This demonstrates how WISP can highlight regions where the model's explanations may lack chemical validity, providing valuable insight into the extent to which these explanations should inform design decisions. Moreover, this information can be used to systematically improve models or datasets in areas where the current model fails to capture the underlying chemistry, ultimately supporting the development of more robust and reliable predictive models in the future.
| Attribution method | r 2 whole molecule to pred. | r 2 variable part to pred. | r 2 whole molecule to LCAP yield | Std constant part | Accuracy variable part to pred. |
|---|---|---|---|---|---|
| Atom attributions | 0.92 | 0.51 | 0.25 | 3.02 | 0.89 |
| RDKit attributions | 0.87 | 0.37 | 0.10 | 2.78 | 0.70 |
Another limitation of the atom-based attribution is its lack of alignment with actual experimental outcomes. The r2 between the ΔattributionsMMP of the whole molecule and the experimental yield reaches only 0.25, which cannot be considered a meaningful correlation (Table 3). This implies that the model's explanations are unsuitable for guiding experimental design in general. Therefore, the explainability methods are only capable of capturing the predictions to a limited extent and fail to generalize to the true property of interest. This suggests that the model did not learn the underlying chemistry but instead relied mainly on numerical correlations in the input data. The small standard deviation in the constant part suggests that the model has captured consistent patterns, though not the underlying chemistry itself. Given that the LCAP dataset is purely combinatorial, this result is not entirely unexpected. All these aspects highlight the inherent difficulty of yield prediction as a task.26
Looking into the classifying results, we feel that qualitatively, the atom attribution method still appears to perform well (Table 3). This is primarily due to the fact that most of the ΔattributionsMMP and the corresponding prediction differences fall within the negative range. As a result, even flawed regions are classified as false negatives, which artificially inflates the accuracy. In effect, these instances contribute positively to the overall accuracy despite introducing underlying attribution errors. This issue could potentially be mitigated by employing larger or more balanced datasets.
The quantitative performance of the RDKit attribution method is similarly poor as that of the atom attributor (Table 3). In this case, the lack of correlation is further emphasized by the clear separation of different MMP transformations, which prevents the formation of a coherent explainability–prediction relationship. The lower region of the correlation plot (Fig. SI-1b) corresponds to transformations where a bromine atom is replaced with hydrogen (Fig. 2, top); the central region includes nitrile-to-methyl ester transformations (Fig. 2, middle); and the upper region reflects methoxy-to-hydrogen substitutions (Fig. 2, bottom). As with the RDKit-based attribution method, the atom attributor fails to differentiate sufficiently between the two molecules involved in the nitrile-to-methyl ester transformation. Visual inspection of the attributions reveals that the variable part of the MMP receives low attribution values (Fig. 2, middle). The underlying cause is the sparse bit coverage in the fingerprint representation of these bonds, with only 36 bits for 12-2-43 and 38 for 13-2-43. In contrast, other bonds are associated with 70 to 369 bits. This indicates that an attribution value close to zero in the RDKit method does not necessarily imply a low contribution to the model's explanation but may instead reflect a sparsity in fingerprint coverage.
This demonstrates how WISP can be applied to external models to evaluate their explainability and to identify instances where the model fails to capture the underlying chemistry. However, it is important to note that the dataset used in this analysis comprises only 46 MMPs (Table 1), which limits the statistical robustness and generalizability of the conclusions drawn. Accordingly, the size of this dataset may indicate a lower limit for the practical application of WISP. However, the exact requirement on dataset size will vary based on the specific structural characteristics of the dataset in question.
3.3 Factor Xa
In adaptation of the work by Harren et al., we evaluated the explainability methods on a dataset of bioactivity values for coagulation Factor Xa.22 As the original dataset was not publicly available, the authors recommended the dataset used by Bailey et al. as a comparable alternative.18When evaluating the qualitative performance of the explainability methods, all approaches perform overall comparably (Table 4). For whole-molecule explanations, the SHAP attributor shows a slight advantage, whereas the RDKit attributor performs best when focusing on the variable part.
| Attribution method | Whole molecule training set | Variable part training set | Whole molecule test set | Variable part test set |
|---|---|---|---|---|
| Atom attributions | 0.91 | 0.84 | 0.76 | 0.72 |
| RDKit attributions | 0.89 | 0.84 | 0.80 | 0.80 |
| SHAP attributions | 0.94 | 0.80 | 0.85 | 0.74 |
To obtain a quantitative measure, we correlate the Δ in MMP attributions with the Δ in model predictions using the squared Pearson correlation coefficient. Across all three attribution methods, the explanations for the whole molecule are well-correlated with the predictions, showing high squared Pearson correlation coefficients: r2 = 0.91 (atom), r2 = 0.89 (RDKit), and r2 = 0.93 (SHAP) (Table 5, r2 whole molecule to pred.). The performance of the explanations for the training values is similarly strong (Table 5). For the variable part of the MMPs, the r2 drops to 0.66 for the atom attributor and to 0.65 for the RDKit attributor, and to 0.57 for SHAP (Table 5). Nonetheless, these values remain relatively high in comparison to other endpoints evaluated in this and the previous study.15 The standard deviation in the attributions of the constant part is 15.78 for the atom attributor and 15.17 for the RDKit attributor—placing them in the bottom compared to previous models. The SHAP attributor achieves a notably lower standard deviation of 7.36. Although some strong outliers are visible in the data, the overall trend suggests reliable and consistent performance of the explainability methods across attribution types.
In Fig. 3, the heatmaps for structure 873, generated by all three explainability methods, are shown. This structure corresponds to compound 3 in the study by Harren et al.22 The predicted pKi of our model for this compound is 6.28 (weak activity), which falls between the values predicted by the models in Harren et al. (6.20 and 6.38).22 This trend is also reflected in the heatmaps: the isopropyl group and the tertiary nitrogen are indicated as having a neutral influence by both the atom attributor and the RDKit attributor, while the SHAP attributor suggests a slight positive influence for the isopropyl group (Fig. 3). Since the azetidine motif is generally less favored compared to larger substituents in this position, the model explanations appear to capture this subtle structural context well, in contrast to the heatmaps presented by Harren et al. for the larger dataset.22
 | ||
| Fig. 3 Comparison of the heatmaps for structure 873 from the Factor Xa dataset. This molecule is part of the training set and has an experimental pKi of 6.08 and a predicted pKi of 6.27. | ||
Another example is an MMP discussed by Harren et al.22 In this case, the variable part is an exchange of a bromine atom for an iodine atom (Fig. 4). The structure containing the bromine has the higher pKi, while the structure with the iodine shows a lower pKi. This difference should be reflected in the attributions and, consequently, in the heatmaps. Indeed, the bromine is highlighted as having a positive influence, whereas the iodine shows a neutral influence (Fig. 4). This result contrasts with the findings of Harren et al., where their model explanation did not correctly attribute the variable part of this MMP, despite using their better-performing model for the analysis.22
This is also reflected in the quantitative assessment of the explainability methods on the Factor Xa dataset, where the atom attributor achieves the highest squared Pearson correlation coefficient for property of interest explanations on the training set across all datasets examined in this study (Table 5). This outcome further validates both the atom attributor and the WISP workflow, demonstrating strong alignment between the qualitative insights from the heatmaps and the quantitative results—both of which are supported by the experimental reference data. Additionally, this analysis highlights the critical importance of careful dataset curation: if specific functional group effects are to be learned by the model, the necessary variations must be well represented in the underlying data.
3.4 AMES
The AMES mutagenicity dataset represents a complex challenge, since the model must learn to recognize non-additive toxicophores—structural motifs that are known to cause mutagenicity but whose effects do not simply add up linearly.27 As AMES is the only classification task in this study, it poses an additional challenge to the WISP framework. Because for a classification problem, there are four discrete types of property changes associated with each MMP (A, B): A0 ↦ B0(Δ = 0), A1 ↦ B1(Δ = 0), A0 ↦ B1(Δ = 1), A1 ↦ B0(Δ = −1), but only three of these are distinguishable in the plot (Fig. SI-2 and SI-3a, b). This is because both in the case of two negatives and two positives, the same difference (Δ = 0) is obtained. From an application perspective, these MMP types include pairs where the transformation leads from mutagenic to non-mutagenic (Δ = −1), from non-mutagenic to mutagenic (Δ = 1), or results in no change in mutagenicity (Δ = 0). Consequently, the original binary classification problem with discrete targets ytarget ∈ {0, 1} and predictions ypred ∈ {0, 1}, once transformed into the MMP framework of WISP, turns into an ordinary regression. As we judged squared errors to remain sensible within this setting (e.g. a −1/1 mistake is four-times more costly compared to a 0/1 mistake), it was decided to remain evaluating with the r2 on the predicted probability output of the model. A further advantage of this approach is its consistency with the regression endpoints, allowing us to compare the explainability performances on whole molecule and variable part (Table 5) to the other observations made.However, these distinctions disappear when examining the test set, which already indicates that the model may not generalize well. The variance in the constant part is in the same range as for the Factor Xa dataset, with a standard deviation of 14.61 for the atom attributor and 13.82 for the RDKit attributor (Table 5). On the test set, the atom attributor shows generally good explainability performance. However, the explanations do not appear to correlate strongly with the property of interest values (Table 5). The same holds for the RDKit attributor, although its overall performance on the test set is noticeably lower except for the standard deviation in the constant part (Table 5).
When examining example MMPs (Fig. 5), it becomes evident that the explainability methods fail to correctly localize the specific structural changes within the respective MMPs. This observation aligns with the high standard deviation found in the constant part, indicating inconsistent attributions. Given that aliphatic halides are well-known toxicophores for AMES mutagenicity,27 one would expect the model to consistently highlight these regions, as shown in Fig. 5 (top MMP). However, the low correlation coefficient on the mutagenicity itself already suggests that the model struggles to capture and predict the relevant toxicophores, resulting in poor generalization to unseen data. The same holds true for aromatic amines (Fig. 5, middle and bottom), which are also established mutagenic motifs.27 This stands in contrast to the work by Vangala et al., whose Grad-CAM-based explainability approach correctly highlighted these key fragments.28 Accurately identifying such critical substructures would be essential for enabling clear, chemistry-guided decision-making.
 | ||
| Fig. 5 Comparison of the heatmaps for different MMPs from the AMES mutagenicity dataset, generated using the atom attributor within WISP and scaled uniformly for consistency. | ||
4 Overall trends
WISP allowed us to quantify the capabilities of the different explainability methods considered in this work (Table 5). Our results show that the newly developed, model- and descriptor-agnostic atom attributor overall performs comparably—and especially more universally—than the SHAP or RDKit attributors.The analysis of WISP with respect to the Crippen log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P dataset in part I demonstrates how effective these explanations can be:15 the performance of the explainability of the variable part of the MMPs on the training set reaches an r2 of 0.79 and 0.77 on the test set. The machine learning model can explain unseen data with an r2 of up to 0.64 with respect to the whole molecule. This indicates that the explainability methods are already capable of addressing the challenge of explaining future data, but there is still significant room for improvement in model quality. In general, explaining unseen data remains challenging: only two of the eight evaluated experiments (Crippen log
P dataset in part I demonstrates how effective these explanations can be:15 the performance of the explainability of the variable part of the MMPs on the training set reaches an r2 of 0.79 and 0.77 on the test set. The machine learning model can explain unseen data with an r2 of up to 0.64 with respect to the whole molecule. This indicates that the explainability methods are already capable of addressing the challenge of explaining future data, but there is still significant room for improvement in model quality. In general, explaining unseen data remains challenging: only two of the eight evaluated experiments (Crippen log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and solubility) show potential for successful application to future data when considering the whole molecule. This is a new finding compared to the work of Jiménez-Luna et al., where no method was able to color previously unseen activity cliffs correctly.29 However, for many of the tested endpoints, the explanations for predictions on the training set perform well.
P and solubility) show potential for successful application to future data when considering the whole molecule. This is a new finding compared to the work of Jiménez-Luna et al., where no method was able to color previously unseen activity cliffs correctly.29 However, for many of the tested endpoints, the explanations for predictions on the training set perform well.
Through the insights provided by WISP, users can also evaluate the predictive performance of their models by identifying regions where the model fails to learn the structure–property relationship accurately. This is demonstrated for the LCAP yield dataset (Section 3.2) and enables users to actively improve the model by including relevant data, thereby enhancing both predictive performance and explainability.
Based on the data presented in Table 5, we developed an action guide to help interpret the outputs of WISP (Fig. 6). Here, the correlation of each metric with the r2 on the test set provides perspective on its impact. An example correlation is shown in the SI (Fig. SI-4). This metric's impact was incorporated into the action guide (Fig. 6). Since the action guide is derived from the datasets used in this study, the cutoffs should not be interpreted as strict thresholds but rather as general guidelines. Users are advised to adjust these values according to the specific requirements of their application.
The primary indicator of explainability quality is the underlying model performance, as discussed in the first part of this series.15 When the model demonstrates sufficient predictive quality, gaps in explainability performance can reveal specific model limitations, as exemplified by the LCAP yield dataset (Section 3.2). In general, we recommend evaluating the dataset at hand using a combination of the different metrics provided. The standard deviation of the constant part itself appears to be a qualitative indicator of explainability performance. When ranking the experiments by this metric on the training set, the resulting order aligns well with the qualitative impressions given by the heatmaps. This indicates that if WISP calculates a standard deviation of the constant part around 5 or higher, the resulting heatmaps should be interpreted with caution—particularly when the r2 value shows that the explainability performance on the property of interest is also low. Each metric focuses on a different aspect of explainability performance—whether it quantifies how well the model explains its predictions or the property of interest, and whether it assesses the whole molecule, the variable part, or the constant part. This combination can help to identify cases where the explanations may be unreliable, allowing WISP to flag such datasets and support more robust decision-making.
5 Conclusion
By systematically applying WISP to a diverse range of datasets—including the Crippen log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, experimental log
P, experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, aqueous solubility, LCAP yield, Factor Xa pKi, and AMES mutagenicity—we demonstrated that the performance of model explanations can be systematically quantified and benchmarked for both simple and complex structure–property relationships using matched molecular pairs. The generally applicable atom attributor performs comparably to less widely applicable methods like RDKit's SimilarityMaps and SHAP in many scenarios, providing robust, transferable explanations.
P, aqueous solubility, LCAP yield, Factor Xa pKi, and AMES mutagenicity—we demonstrated that the performance of model explanations can be systematically quantified and benchmarked for both simple and complex structure–property relationships using matched molecular pairs. The generally applicable atom attributor performs comparably to less widely applicable methods like RDKit's SimilarityMaps and SHAP in many scenarios, providing robust, transferable explanations.
Furthermore, WISP exposes the limitations of current machine learning models in capturing subtle functional group effects, especially in challenging tasks like mutagenicity or bioactivity prediction, where non-additive and context-dependent interactions are critical.
Importantly, our findings emphasize that explaining predictions for unseen data remains a significant challenge: while on several datasets the explainability methods perform well on training data, only a few demonstrate the consistency and generalizability required for trustworthy predictions on new molecules. This underlines the necessity for rigorous dataset curation and continuous improvement of both models and explanation techniques.
Author contributions
JMW and AHG: conceptualization, KJ and JMW: data curation, KJ: formal analysis, JP and AHG: funding acquisition, KJ: investigation, KJ and JMW: methodology, JP and AHG: project administration, JP and AHG: resources, KJ and JMW: software, JMW, JP and AHG: supervision, KJ: validation, KJ: visualization, KJ: writing – original draft, JMW, JP and AHG: writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
All code associated with this work, including the URL to the public web application, is available on GitHub at https://github.com/kerjans/ml-XAI. An archived version of the complete codebase is additionally accessible via Zenodo at https://doi.org/10.5281/zenodo.17055142.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00399g.
Acknowledgements
JP acknowledges funding by Germany's joint federal and state program supporting early-career researchers (WISNA) established by the Federal Ministry of Research, Technology and Space (BMFTR). The authors acknowledge Philipp Held (TU Braunschweig) for providing explanatory Jupyter Notebooks on WISP. In preparing this work, the authors used ChatGPT (OpenAI) to refine wording and provide coding assistance. All AI-generated material was thoroughly reviewed and edited by the authors, who assume full responsibility for the accuracy and integrity of the final publication.References
- E. N. Muratov, J. Bajorath, R. P. Sheridan, I. V. Tetko, D. Filimonov, V. Poroikov, T. I. Oprea, I. I. Baskin, A. Varnek, A. Roitberg, O. Isayev, S. Curtalolo, D. Fourches, Y. Cohen, A. Aspuru-Guzik, D. A. Winkler, D. Agrafiotis, A. Cherkasov and A. Tropsha, Chem. Soc. Rev., 2020, 49, 3525–3564 RSC.
- L. Zhang, J. Tan, D. Han and H. Zhu, Drug Discovery Today, 2017, 22, 1680–1685 CrossRef PubMed.
- K. Jorner, A. Tomberg, C. Bauer, C. Sköld and P.-O. Norrby, Nat. Rev. Chem., 2021, 5, 240–255 CrossRef CAS PubMed.
- J. Jiménez-Luna, M. Skalic, N. Weskamp and G. Schneider, J. Chem. Inf. Model., 2021, 61, 1083–1094 CrossRef.
- C. Humer, H. Heberle, F. Montanari, T. Wolf, F. Huber, R. Henderson, J. Heinrich and M. Streit, J. Cheminf., 2022, 14, 21 Search PubMed.
- A. Chatzimparmpas, R. M. Martins, I. Jusufi and A. Kerren, Inf. Visual., 2020, 19, 207–233 Search PubMed.
- P. Polishchuk, J. Chem. Inf. Model., 2017, 57, 2618–2639 CrossRef CAS PubMed.
- Y. Sushko, S. Novotarskyi, R. Körner, J. Vogt, A. Abdelaziz and I. V. Tetko, J. Cheminf., 2014, 6, 48 Search PubMed.
- G. Marcou, D. Horvath, V. Solov’ev, A. Arrault, P. Vayer and A. Varnek, Mol. Inf., 2012, 31, 639–642 CrossRef CAS PubMed.
- M. Matveieva and P. Polishchuk, J. Cheminf., 2021, 13, 41 CAS.
- K. Janssen and J. Proppe, J. Chem. Inf. Model., 2025, 65, 1862–1872 CrossRef CAS PubMed.
- M. Eckhoff, J. V. Diedrich, M. Mücke and J. Proppe, J. Phys. Chem. A, 2024, 128, 343–354 CrossRef CAS PubMed.
- J. Jiménez-Luna, F. Grisoni and G. Schneider, Nat. Mach. Intell., 2020, 2, 573–584 CrossRef.
- L. Rosenbaum, G. Hinselmann, A. Jahn and A. Zell, J. Cheminf., 2011, 3, 11 CAS.
- K. Janssen, J. M. Wollschläger, J. Proppe and A. H. Göller, Digital Discovery, 2025 CrossRef , submitted..
- C. Tyrchan and E. Evertsson, Comput. Struct. Biotechnol. J., 2017, 15, 86–90 CrossRef CAS PubMed.
- S. Felten, C. Q. He, M. Weisel, M. Shevlin and M. H. Emmert, J. Am. Chem. Soc., 2022, 144, 23115–23126 CrossRef CAS PubMed.
- M. Bailey, S. Moayedpour, R. Li, A. Corrochano-Navarro, A. Kötter, L. Kogler-Anele, S. Riahi, C. Grebner, G. Hessler, H. Matter, M. Bianciotto, P. Mas, Z. Bar-Joseph and S. Jager, Deep Batch Active Learning for Drug Discovery, bioArXiv, 2023 DOI:10.1101/2023.07.26.550653.
- K. Hansen, S. Mika, T. Schroeter, A. Sutter, A. Ter Laak, T. Steger-Hartmann, N. Heinrich and K.-R. Müller, J. Chem. Inf. Model., 2009, 49, 2077–2081 CrossRef CAS PubMed.
- H. Heberle, L. Zhao, S. Schmidt, T. Wolf and J. Heinrich, J. Cheminf., 2023, 15, 2 Search PubMed.
- G. P. Wellawatte, H. A. Gandhi, A. Seshadri and A. D. White, J. Chem. Theory Comput., 2023, 19, 2149–2160 CrossRef CAS PubMed.
- T. Harren, H. Matter, G. Hessler, M. Rarey and C. Grebner, J. Chem. Inf. Model., 2022, 62, 447–462 CrossRef CAS PubMed.
- S. M. Lundberg and S.-I. Lee in Advances in neural information processing systems, ed. I. Guyon, U. V. Luxburg, S. Bengio, H. Wallach, R. Fergus, S. Vishwanathan and R. Garnett, Curran Associates, Inc., 2017, vol. 30 Search PubMed.
- S. Riniker and G. A. Landrum, J. Cheminf., 2013, 5, 43 CAS.
- P. Schwaller, A. C. Vaucher, T. Laino and J.-L. Reymond, Machine Learning: Science and Technology, 2021, 2, 015016 Search PubMed.
- V. Voinarovska, M. Kabeshov, D. Dudenko, S. Genheden and I. V. Tetko, J. Chem. Inf. Model., 2024, 64, 42–56 CrossRef CAS PubMed.
- J. Kazius, R. McGuire and R. Bursi, J. Med. Chem., 2005, 48, 312–320 CrossRef CAS PubMed.
- S. R. Vangala, S. R. Krishnan, N. Bung, R. Srinivasan and A. Roy, J. Chem. Inf. Model., 2023, 63, 5066–5076 CrossRef CAS.
- J. Jiménez-Luna, M. Skalic and N. Weskamp, J. Chem. Inf. Model., 2022, 62, 274–283 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |