 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Increasing trustworthiness of machine learning-based drug sensitivity prediction with a multivariate random forest approach
Lisa-Marie
Rolli
 *ab,
Lea
Eckhart
*ab,
Lea
Eckhart
 acd,
Lutz
Herrmann
acd,
Lutz
Herrmann
 b,
Andrea
Volkamer
b,
Andrea
Volkamer
 b,
Hans-Peter
Lenhof
b,
Hans-Peter
Lenhof
 a and
Kerstin
Lenhof
a and
Kerstin
Lenhof
 acde
acde
aChair for Bioinformatics, Center for Bioinformatics, Saarland University, Saarland Informatics Campus, Saarland, 66123, Germany. E-mail: lisa-marie.rolli@uni-saarland.de
bChair for Data Driven Drug Design, Center for Bioinformatics, Saarland University, Saarland Informatics Campus, Saarland, 66123, Germany
cIntegrative Bioinformatics Group, Department of Medical Bioinformatics, University Medical Center Göttingen, Georg-August-University Göttingen, Lower Saxony, Germany
dCAIMed – Lower Saxony Center for Artificial Intelligence and Causal Methods in Medicine, Göttingen, Germany
eComputational Biology Group, Department of Biosystems Science and Engineering, ETH Zürich, Klingelbergstrasse 48, Basel, 4056, Switzerland
First published on 25th March 2026
Abstract
Ensuring the trustworthiness of machine learning (ML) models in high-stake applications is crucial. One such application is predicting anti-cancer drug sensitivity, where ML models are built with the final goal of integrating them into treatment recommendation systems for personalized medicine. Here, we propose a trustworthy multivariate random forest method MORGOTH, available in our package ‘morgoth’. Besides standard regression and classification functions, MORGOTH allows for the simultaneous optimization of regression and classification tasks via a joint splitting criterion. Additionally, it provides a graph representation of the random forest to address model interpretability, and a cluster analysis of the leaves to measure the dissimilarity of new inputs from the training data to account for its reliability and robustness. In total, MORGOTH provides a comprehensive approach that unites simultaneous regression and classification, interpretability, reliability, and robustness in a single framework. While our package is broadly applicable, we demonstrate its capabilities for anti-cancer drug sensitivity prediction by a comprehensive large-scale study on the Genomics of Drug Sensitivity in Cancer (GDSC) database. We trained single-drug as well as multi-drug models. In either case, MORGOTH clearly outperforms state-of-the-art neural network approaches. Moreover, we highlight an evaluation issue for multi-drug models and demonstrate that single-drug models consistently outperform them when evaluated fairly.
1 Introduction
Personalized medicine aims to identify the most suitable treatment for a given patient based on their characteristics, e.g., clinical or molecular profiles. This is particularly difficult for cancer treatment because of the high level of heterogeneity between patients, tumors and even within the same tumor.1,2 Consequently, the development of machine learning (ML)-based decision support systems for cancer therapy has garnered attention for many years.1–9 To help with this task, supervised ML models can be trained to estimate the treatment efficacy of a compound based on molecular data of samples. Ideally, such models could be used to recommend drug treatments for a patient based on the estimated drug response. Currently, there is not yet enough real patient data available to train ML models for this task, which is why the models are typically trained on cancer cell line data.10When using ML for such critical tasks, trustworthiness is particularly important. In a recently published review, Lenhof et al.5 analyzed 36 articles in the field of anti-cancer drug sensitivity prediction with a focus on three aspects of trustworthiness:
(1) Performance, which is the overall correctness of the predictions for (test) data measured by common metrics such as mean-squared error (regression) or Matthews correlation coefficient (classification).
(2) Reliability, which is the degree of trust one can have in a specific prediction, especially for samples with unknown response, typically assessed via uncertainty estimation or p-values.5,11–13
(3) Interpretability, which describes how understandable the model and its results are to humans.5
Concerning performance, Li et al.14 showed that simple models, e.g., random forests (RFs), are often competitive to complex models such as different kinds of (deep) neural networks (NN) for anti-cancer drug sensitivity prediction. Similar observations have also been made by Wissel et al.15 for survival prediction, where statistical models, in particular also a tree-ensemble based method, outperformed the NNs. Generally, tree-based methods are particularly interesting as they have been shown to have state-of-the-art performance for tabular data.16–18 Despite the predominant focus in the literature on improving model performance, the evaluation for anti-cancer drug sensitivity prediction often overlooks the prioritization task (i.e., the sorting of drugs by predicted treatment efficacy for a specific tumor or cell line).5,19 This would, however, be desirable since this is the goal of a treatment recommendation system.
In terms of reliability, Lenhof et al.5 found that only two approaches (partially) address the demand for it: Fang et al.20 present a partially reliable approach based on a quantile regression forest-method. Yet, they did not guarantee specific confidence levels, which is however, highly desirable. In contrast, the conformal prediction method as introduced in the reliable SAURON-RF publication by Lenhof and Eckhart et al.,21 provides rigorous certainty guarantees for independent and identically distributed (i.i.d.) data.
The review also finds that the term ‘interpretability’ is not well-defined but used intuitively, while there exist several connotations of what is meant by this term.5 For this reason, the review introduces a taxonomy of different interpretability types and categorized the current drug sensitivity literature accordingly. This analysis revealed that some interpretability types are not explored at all (i.e., model-, sample-, and concept-based explainability), while most approaches categorized as interpretable only partially comply.5
In this manuscript, we address the gaps identified in our literature review in terms of performance, reliability and interpretability.
Since RFs have been identified as one of the best approaches for drug sensitivity prediction,22 we decided to extend beyond our own RF-based approach reliable SAURON-RF.21 Our novel approach is called MORGOTH (Multivariate classificatiOn and Regression increasinG trustwOrTHiness) and it extends SAURON-RF in the following aspects:
(1) Explicit multi-task learning: SAURON-RF performs a simultaneous classification and regression, while optimizing only the regression task during training. As we observed that this ‘implicit’ multi-task learning considerably increased both, the regression and classification performance, we now investigate the effect of explicitly integrating both, regression and classification task, in the objective function (‘explicit’ multi-task learning). To this end, we implemented a flexible splitting criterion function for MORGOTH, which is a weighted linear combination of two error measures: one for regression and one for classification.
(2) RF graph representation: we implemented a novel representation for RFs, where the traces of the samples through the forest are condensed in a graph. Each node represents a visited feature and each edge in the graph corresponds to a used edge (i.e., path from one internal node to its respective child node) in the forest. The resulting graph visually highlights the most important parts of the RF, thus allowing for a feature- and concept-based interpretability of our approach.
(3) Cluster analysis: we propose a distance-based test for the applicability domain of our model, i.e., the domain that is covered by the training samples.23 To this end, we score how well the test samples fit to the most similar training samples. This might serve as an explanation for inaccurate or uncertain predictions for badly fitting samples. Consequently, the cluster analysis provides a sample-based interpretability5 of both, the model predictions as well as the reliability. By identifying out-of-distribution samples, the cluster analysis not only enhances interpretability but also strengthens the robustness of the approach (cf. Lenhof et al.24 for a definition of robustness within the ML realm).
MORGOTH is implemented in Python and our package can easily be installed via pip. Thus, it can be applied to other ML datasets than the one that we use in our study. In this article, we demonstrate the capabilities of our approach by applying it to the Genomics of Drug Sensitivity in Cancer (GDSC) database to predict drug sensitivity of cancer cell lines. It has been claimed that multi-drug models for anti-cancer drug sensitivity prediction outperform single-drug approaches.25,26 Thus, we did not only develop drug-specific models but also a multi-drug model by incorporating drug features during the training process. To identify suitable drug features, we analyzed 16 models from the drug sensitivity literature that use drug features. It turned out that 5 out of 16 use physicochemical properties while 7 out of 16 employ molecular fingerprints (cf. SI Table 1). We decided to abstain from employing more sophisticated approaches, as it has been repeatedly questioned that more complex descriptors show superior performance in comparison to conventional fingerprints or physicochemical properties.27–29 Therefore, we solely use structural (MACCS and Morgan) fingerprints and physicochemical properties. Our analyses show that the typical evaluation strategies between the single-drug and the multi-drug settings differ, which leads to seemingly better results for multi-drug models. This finding supports the study by Codicè et al.,30 who show that typical evaluation strategies can be fooled due to dataset biases that are present in most drug response datasets, including the GDSC. Performing a fair evaluation, we demonstrate that single-drug models actually outperform multi-drug models. Moreover, we show that in both settings, MORGOTH outperforms state-of-the-art deep neural networks in terms of performance.
2 Materials and methods
2.1 Data acquisition
In our study, we employed the GDSC database Release 8.3 from June 2020.31–33 The GDSC contains drug response measurements for around 500 anti-cancer drugs as well as different data types to characterize the tested cell lines, such as gene expression, mutation, or copy number data. Since gene expression has been shown to be the most informative data type for drug sensitivity prediction,34 we downloaded the RMA normalized gene expression values. Moreover, we used the raw drug response data from the GDSC to calculate the CMax viability, which is a recently introduced drug sensitivity measure by Lenhof and Eckhart et al.21 The CMax viability is defined as the relative viability that corresponds to the peak plasma concentration that was measured, when administering the drug in the maximum clinically recommended dose.19,21 Consequently, the CMax viability is a continuous value in [0, 1], where a lower value indicates a higher sensitivity to the compound, s.t. the CMax viability is comparable across drugs. Thus, the CMax viability can be used to mitigate common problems observed in conventional measures such as IC50 or AUC. We also derived a binary CMax value as described in,21 here 0 corresponds to ‘resistant’ and 1 to ‘sensitive’.Using the drug names from the GDSC, we queried the ChEMBL database Version 33 from May 2023 (ref. 35 and 36) to obtain the SMILES representation of the drugs.37 Based on the SMILES, we calculated different commonly used (cf. SI Table 1) drug feature types using RDKit Version 2023.3.2:38
• MACCS fingerprints39 of length 166.
• Morgan fingerprints40 with radius 3 and length 2048.
• The 209 RDKit physicochemical properties41
2.2 Multivariate classificatiOn and regression increasinG trustwOrTHiness (MORGOTH)
With our novel approach MORGOTH, we built upon our own method reliable SAURON-RF. SAURON-RF simultaneously performs classification and regression by optimizing a weighted mean-squared-error during the training process. The weights are based on the class labels, which are thus, implicitly used during the optimization, but there is no explicit optimization of the classification task. Furthermore, the continuous and discrete response for each sample is stored in the leaf nodes s.t. a prediction for both tasks can be casted. Moreover, SAURON-RF enables feature-based interpretation by providing feature importance scores and the reliability of the approach is assessed using conformal prediction (CP).21 However, our novel approach, MORGOTH, is more flexible and trustworthy: It offers the selection between several criteria that can be used to split the internal nodes of the decision trees and it is not only possible to perform a joint classification and regression, but also each task separately (cf. Section 2.2.1). In addition to the feature importance scores and the integration of a CP framework that were already provided by SAURON-RF, we implemented a novel RF graph representation, which can be used to identify important features and connections between features (cf. Section 2.2.2). Moreover, MORGOTH offers the possibility to perform a cluster analysis to score how well the test samples fit the training samples, on which the respective prediction is based to account for applicability to the new samples (cf. Section 2.2.3).MORGOTH is an RF and, consequently, it is an ensemble of decision trees.42 Since we want to offer the possibility to simultaneously optimize for both, classification and regression, we propose a novel splitting criterion function, which is a weighted linear combination of an error function for classification and an error function for regression. Similarly, Ishwaran et al.44 implemented a multivariate splitting rule for multivariate RFs in R. Their implementation uses a fixed normalized composite score, i.e., the user cannot gauge the relative importances of the classification and regression task. In contrast, our method introduces a tunable weighting parameter λ that allows practitioners to explicitly adjust the relative importance of classification versus regression objectives. This flexibility is crucial in real-world scenarios where one task may dominate the decision-making process or where domain-specific priorities exist. For example, in drug sensitivity prediction, the classification labels (e.g., ‘sensitive’ and ‘resistant’) are often derived from continuous response values (e.g., CMax viability).3,21 For samples with a response far from this threshold, optimizing regression error naturally improves classification accuracy. However, for samples close to the threshold, classification becomes critical to correctly assign them to the appropriate category. As most of the samples are relatively far from the threshold, practitioners may choose to emphasize regression (e.g., MSE) over classification (e.g., Gini impurity).
Throughout this manuscript, we use Gini-impurity (classification) and MSE (regression); the weighting factor λ is tuned as a hyperparameter in a 5-fold cross-validation (cf. Section 2.3). Moreover, to counteract class imbalance, we weight our samples with the ‘simple weights’ introduced by Lenhof et al.2. The simple weights weight the training samples that belong to the majority class with 1. The samples that belong to the minority class are assigned with a higher weight, which is the ratio of majority and minority class in the training set. However, other error functions and sample weights are also available for MORGOTH.
Since we consider error functions, the output of the two functions should lie in [0, ∞), where larger values indicate a higher error. However, the ranges of the error functions may differ. Therefore, we use the values of the error functions that are obtained at the root node of the respective decision tree to scale the values to the same range. Since during the training the values of MSE and Gini-impurity are minimized, we expect the values obtained at the root node to be the maximum values that we obtain in the tree, i.e., we scale the values in the range [0, 1]. Let MSEroot and Giniroot be the root node error values. Then, the combined error function (SCλ) calculated on the subset of the data Zv, which is assigned to a node v in the current tree, is defined as follows:
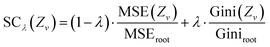 | (1) |
The factor λ balances the importance of the classification error versus the regression error. While for λ ∈ (0.5, 1], the classification task is considered more important during the training, for λ ∈ [0, 0.5), the regression task is more important. For λ = 0 or λ = 1, eqn (1) is, except for the constant scaling term, equivalent to the ordinary regression and classification error, respectively.
Using eqn (1), we can find the best split of node v into a left and right child node vl and vr by maximizing the improvement in error introduced by the split. The improvement of a split s that splits the data Zv into Zvl and Zvr can be expressed as:
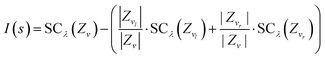 | (2) |
MORGOTH provides two general possibilities constructing graphs:
(1) We can construct one graph for individual samples.
(2) We can construct one graph for all samples in a given set, e.g., the whole training/test set or the set containing all ‘sensitive’/‘resistant’ samples.
After constructing these graphs, it is possible to apply different standard graph algorithms for further analyses to calculate, e.g., difference graphs (cf. Section 3.2).
Now, we first explain how we can construct the graph for the first case and then how this information is used for the second case.
Each internal node v in the forest is associated with a feature Xv and the corresponding threshold used to split the samples. With L(v) we denote the level of node v defined as the number of edges between v and the root node, i.e., the level of the root node is 0. When propagating a sample through the forest, we track all internal nodes [v1, …, vl] and all edges between the internal nodes that we visit. This information is finally used to fill two lists:
(1) A list of all features that we visit at the internal nodes.
(2) A list of all the directed edges between features, i.e., for each visited edge we note the features corresponding to the connected nodes and store the information as a directed connection between the parent and the child feature.
Moreover, the entries in both lists are weighted: For each feature Xi in the first list, we calculate its average level L′(Xi). Each feature Xi is weighted by the average level of all visited nodes that use Xi for splitting:
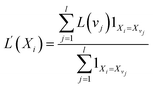 | (3) |
The indicator variable 1Xi=Xvj is one iff node vj uses feature Xi for splitting. To weight the directed edges in the second list, we count how often we used an edge that connected the same parent and child feature. The final graph for a specific test sample consists of the features as nodes and the connections between features as edges weighted as described above.
When building a graph for a group of samples, e.g., the whole test set, we again need the two lists that are filled as described above. The only difference to the sample-specific setting is that we now consider all features and edges that have been visited by any sample in the group. Thus the weights are also calculated across all samples. Notably, the ranges for the average feature rank will not differ to the sample-specific graphs, but the edge weights will be higher since we count the number of uses per edge. To make comparison between sample-specific and group-graphs easier, we additionally offer the possibility to average the edge weights by the number of samples in the group.
After constructing the graphs one can apply standard graph algorithms for further analyses. For example, one can calculate difference graphs between the graphs of two (groups of) samples (cf. Section 3.2).
CP compares whether the prediction for a sample conforms to the predictions for previously seen samples with respect to a score. In the cluster analysis, we directly compare the similarity between the samples based on their features, which might give us complementary information to explain uncertain predictions, i.e., empty sets, sets containing both classes, or large intervals. Intuitively, we would expect that these uncertain predictions are more likely to be cast for unseen samples that are dissimilar to the training set. Moreover, one expects relatively poor predictions compared to samples that are similar to the training instances. However, neither the RF itself nor the CP algorithm directly provide means to measure the (dis)similarity between test and training instances based on feature-values. Moreover, it is known that ML models can be over-confident for samples that deviate from the original training distribution.45 Therefore, in addition to the CP functionality, we implemented a cluster analysis, which scores how well an input sample fits to the most similar training samples in terms of features. This type of out-of-distribution analysis introduces robustness to our framework,24 while making the conformal prediciton more interpretable.
To this end, we leverage the fact that RFs can be interpreted as an adaptive nearest neighbor procedure:46 Each leaf can be regarded as a cluster of similar training samples. For a specific sample xt the overall cluster Xt, in which xt ends up is the union of the training samples in all reached leaves. To score how well a specific sample xt fits to the associated cluster Xt, we calculated a special case of the silhouette score. The silhouette score is commonly used to quantify how well a clustering algorithm works.43 It is based on the distance between samples of the same cluster (intra-cluster distance) and between samples of different clusters (inter-cluster distance). In this special case, we considered Xt as one cluster and xt as the ‘other’ cluster. Due to the bootstrap sampling per tree, one sample can be included within the cluster several times. Thus, our model calculates the silhouette score in two versions: with and without duplicates in the training sample cluster. Let x1, …, xn be the elements of Xt (with or without the duplicates). Let d be a distance measure, then the average pairwise distance within Xt given by:
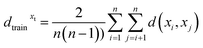 | (4) |
Moreover, dtestxt is the mean distance between the test sample xt and all training samples x1, …, xn:
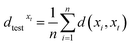 | (5) |
Then, the silhouette score is defined as
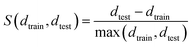 | (6) |
The silhouette score will lie in [−1, 1]. Here, negative values indicate that the test sample fits better to the cluster than the average training sample, positive values indicate that the test sample fits worse. If the value is (close to) 0, the test sample fits equally well to Xt as the training samples in the cluster fit to each other on average.
MORGOTH provides several distance functions for the cluster analysis: cosine,47 Euclidean,48 Pearson,49 and Spearman50 distance as well as rank magnitude.49 Note that Pearson, Spearman, and rank magnitude are correlation measures with values in [−1, 1].49 The distances are obtained by subtracting the respective value from 1. According to Jaskowiak et al.,49 the best measure to cluster gene expression values is Pearson distance. Thus, we focus on this function for our analyses.
2.3 Evaluation strategy and experimental settings
We use the data as described above as input to train different ML models. Generally, we distinguish between 2 scenarios:(1) Single-drug models: one model per drug.
(2) Multi-drug models: one model for all drugs.
We performed a hyperparameter tuning on the training set with a 5-fold cross validation on the training data with random folds and retrained each model for the best-performing combination on the entire training set (details in SI).
To select features – for the training and the CV – we used the feature selection (FS) by Kwak and Choi51 that is based on the minimum redundancy maximum relevance (MRMR) principle. This method is one of the best performing dimensionality reduction techniques for drug sensitivity prediction on the GDSC dataset, as shown by Eckhart et al.22. In particular, we aim to identify the features with the highest mutual information to the response (maximum relevance) while avoiding features with a high mutual information to other already selected features (minimum redundancy).22 Using this FS, we selected 50 genes per drug.
As a reference, we include a dummy baseline by training MORGOTH on permuted data instead of the real data. The models trained on the permuted data are then applied to the actual test set and the performance is reported. As the models are trained on randomly permuted data, we expect that they should not be able to learn properly, which should be reflected in poor performance. We consider two permutation strategies: (1) the models are trained on the correct response but with a permuted feature matrix, i.e., the entries of each column (feature) were randomly permutated, so that the column-wise order was randomized, and (2) the models are trained on the correct feature matrix but with a permuted response. Furthermore, we investigate whether the explicit optimization of both, regression and classification, in the objective function improved the prediction performance, by a comparison of single-drug instances of MORGOTH with SAURON-RF trained on exactly the same data. Moreover, we will compare the performance of MORGOTH to deep neural network approaches as described in the following.
To benchmark our single-drug models, we decided to use the state-of-the-art deep neural network by Sharifi-Noghabi et al.,53 which is called ‘MOLI’ (Multi-Omics Late Integration). MOLI requires three input omics types to characterize the cell lines (gene expression, copy number data, and mutations) and outputs a binary drug response.53 MOLI is trained as single-drug model. Thus, we compare this approach with the MORGOTH single-drug models when trained on the same drugs and cell lines. To this end, we reimplemented the MOLI network according to the description of Sharifi-Noghabi et al.53. The details of our reimplementation can be found in the SI. Then, we trained 25 single-drug models using the same training and test data as described above. Note that in addition to gene expression, MOLI also receives the required additional omics types, which we obtained from the GDSC.
 | ||
| Fig. 2 Feature matrix for the multi-drug setting. For each sample (drug-cell line combination), we have gene expression values und multiple drug characteristics as features. | ||
As for the single-drug models, we performed a hyperparameter tuning for the multi-drug model using a 5-fold CV on the training set (cf. SI for details). For the multi-drug model, we selected 150 genes and 150 drug features, i.e., 300 features in total, since this was a good hyperparameter choice in our previous large-scale benchmarking.22 To select the features, we, first removed all features with constant values across all samples, and then pre-selected the 500 genes and 500 drug features with the highest estimated mutual information to the response. The mutual information was estimated using the python package scikit-learn.43 The final 150 genes and 150 drug features were selected using the algorithm by Kwak and Choi.51
When evaluating the performance of the multi-drug model, there are generally two possibilities: we can (1) either calculate the performance on the whole test set or (2) on drug-specific parts of the test set. While the first option is the one, which is commonly evaluated and reported for multi-drug models,30 the second one would be needed to ensure a fair comparison to the single-drug models. In particular, Codicè et al.30 show that the first option, which they refer to as global evaluation, can be misleading since the models may show seemingly good scores due to a bias in the data set caused by the different response ranges covered by the drugs. However, we provide both evaluations for all our experiments. Notably, since we want to compare the multi-drug model with the single-drug models, we evaluated the single-drug models also not only per drug but also using the global evaluation strategy. To this end, we concatenated all drug-specific test sets to one large test set and assessed the performance in terms of MCC and PCC across all compounds. Similar to the single-drug baseline models, we also added baseline multi-drug models that were trained on permuted data and then applied to the original test sets: (1) only the cell line features were permuted column-wise as described for the single-drug models in Section 2.3.1, (2) only the drug features were permuted column-wise, (3) only the response was permuted. Moreover, the multi-drug model was also benchmarked with a deep neural network.
2.4 Implementation
We fully implemented MORGOTH in Python 3.55 The package is called ‘morgoth’ and can be installed via pip. The open source libraries that are used in our implementation can be found in the SI.3 Results
In this section, we discuss the results of our analyses. First, we focus on evaluating the performance of our novel approach for the single-drug and multi-drug instances, also in comparison to the two reimplemented DNN models. Please note that we employ different data sets for our analyses (cf. Description Section 2.3 and Table 2 in SI) since we wanted to ensure fair comparisons to our competitors and these methods might require additional filtering. In particular, we trained MORGOTH on the respective data set to have a fair comparison to each competitor, which causes slight variations in the different performance plots. Then, we analyze the trustworthiness of our model. Here, we first show how to interpret the RF graph, a visual summary of the forest's most important components. Next, we demonstrate how our approach enables reliable drug prioritization per cell line using CP. Finally, we present a cluster analysis that identifies test samples that differ from the training data, helping to explain uncertain or incorrect predictions.3.1 Performance
We evaluate model performance in terms of Matthews correlation coefficient (MCC) for classification and Pearson correlation coefficient (PCC) for regression. First, we show the evaluation for the single-drug models, followed by the multi-drug model.Fig. 4 shows the comparison between MORGOTH and SAURON-RF.
When trained on the same data, MORGOTH is on par with SAURON-RF: the mean MCCs over the drugs are 0.36 (MORGOTH) and 0.37 (SAURON-RF) and the mean PCCs are 0.54 (MORGOTH) and 0.52 (SAURON-RF). Since differences in performance between SAURON-RF and MORGOTH can solely be traced back to the novel objective function (cf. Section 2.2.1), which is responsible for the explicit multi-task learning, we conclude that the combination of MSE and Gini-impurity (explicit multi-task learning) did not improve upon only MSE with class-dependent sample-weights (implicit multi-task learning). Notably, the objective function (eqn (1)) depends on the weighting factor λ ∈ [0, 1] that weights the importance of the regression and the classification task in eqn (1). If λ is set to 0, we only optimize regression (i.e., the same as SAURON-RF) and if it is 1, we only optimize classification.
To understand how the model used lambda, we investigated the chosen values. For each of the 25 single-drug models, we tuned λ as a hyperparameter using a 5-fold CV on the training data set. In particular, we tested λ ∈ {0, 0.25, 0.5, 0.75, 1}. The value for λ that was used in the final model was the one with the highest value for PCC + MCC on the validation set averaged over all CV iterations. Fig. 5, shows the final values of λ for the 25 single-drug models. We can see that for the majority of the compounds (15 out of 25), λ was chosen in (0,1) meaning that the combination of classification and regression task in the splitting criterion function improved the performance on the validation set for these compounds. Thus, we conclude that the combined splitting criterion has apparently improved the performance slightly for some splits but does not bring significant improvement on our dataset compared to the implicit multi-task learning. Yet, we want to emphasize that even if MORGOTH did not improve upon SAURON-RF in terms of performance, it provides more trustworthiness-related properties, i.e., the cluster analysis and the graph representation.
 | ||
| Fig. 5 Distribution of the selected value for the parameter λ in the splitting criterion function. λ was tuned as hyperparameter in a 5-fold CV for each of the 25 investigated compounds. | ||
Fig. 6 shows the test set performance in terms of MCC for MOLI53 and MORGOTH. Notably, since MOLI is a single-drug model, we compare it with the single-drug instances of MORGOTH. However, we also evaluated the performance when combining all drug-specific test sets to a single large one (across drug evaluation strategy). MORGOTH outperforms MOLI in the per-drug evaluation, i.e., MCC of 0.36 (MORGOTH) and 0.26 (MOLI), while the MCCs across drugs are similar, i.e., 0.62 (MORGOTH) and 0.59 (MOLI).
We compared the performance of the multi-drug MORGOTH model to the modified version of DeepDR that we introduced in Section 2.3.2. Note that both models were trained using the same cell lines and drugs (cf. Section 2.3.2). Fig. 8 shows the performance in terms of PCC for modified DeepDR vs. MORGOTH. While the performance is on par when calculating the PCC on the whole test set (blue points), we observe that MORGOTH outperforms modified DeepDR when evaluating the performance per drug (red points). Notably, in our publication of reliable SAURON-RF, we already demonstrated that single-drug SAURON-RF also outperforms DeepDR, when evaluating per drug.21
Moreover, we investigate the performance of our multi-drug MORGOTH model compared to our single-drug MORGOTH models. Note that here, we compare the 23 out of the 25 drugs, for which we could calculate drug features, and also use the cell-blind split described in Section for both single- and multi-drug models (cf. 2.3.1 for the splitting).
Fig. 9 summarizes the MCC (classification) and PCC (regression) for the single- and multi-drug models evaluated per drug and across the whole test set (across drug evaluation strategy). We can conclude that both models perform clearly better than the random baseline. Moreover, the single drug models outperform the multi-drug model in either evaluation strategy (per drug, across drug) and task (classification, regression). The mean MCC over the single-drug models is 0.34, while it is only 0.24 for the multi-drug model. Evaluated across all drugs, the single-drug models achieve an MCC of 0.62 and the multi-drug model of 0.49. For the regression, we observe a mean PCC of 0.52 in the per drug evaluation for the single-drug models and 0.41 for the multi-drug model. The single-drug models have a PCC of 0.83, when evaluated across all drugs, while the multi-drug model solely reaches 0.65. The same trend can also be observed, when evaluating the regression task using the coefficient of determination R2 (cf. SI Section 5). Our analysis aligns very well with the findings by Codicè et al.,30 who showed that the evaluation of the PCC across all drugs artificially leads to high correlation values. As mentioned above, multi-drug models are often evaluated on the whole test set, while single-drug models are usually evaluated per drug. Since the typical evaluation strategies in literature differ, the multi-drug models are often believed to be superior, while we show that this is not the case.
To further analyze why the multi-drug models performance is worse than expected, we investigate the weighting factor λ from eqn (1) and compare its values to those chosen ones for the single-drug models. In contrast to the single-drug models, where the majority of the models used both, MSE and Gini-impurity in their splitting criterion function, we observe that the multi-drug model only employs the MSE, i.e., λ is set to 0. Further investigation is needed to determine whether this effect is reproducible, under what conditions it arises, and what its implications are.
Moreover, we investigate the importance of the drug features since they are one of the main differences between the data that we used to train the single- and the multi-drug models. Surprisingly, when evaluating the feature importances of the multi-drug model, we observe that only 3 of the 150 drug features have an importance greater than 0 (cf.Table 1) indicating that the other features are noninformative. These 3 features share around 40% of the overall features importance. Notably, all 3 important features are physicochemical properties, although 144 of the 150 chosen drug features are structural fingerprint bits. This observation aligns well with the findings of Sultan et al.27 for molecular property prediction: they show that models with access to physicochemical properties perform better than models that are trained only on the structure, implying that the physicochemical features are more informative for drug characterization.
| Feature name | Score | Description |
|---|---|---|
| VSA_EState7 | 0.19 | Sum over the electrotopological-state indices (cf. Kier and Hall56) of all atoms with a contribution to the van der Waals surface area (VSA) x (cf. Labute57) s.t. 6.07 ≤ x < 6.45 (cf. Landrum58 for more details) |
| FractionCSP3 | 0.13 | The fraction of C atoms that are SP3 hybridized41 |
| Ipc | 0.09 | Information content for polynomial coefficients as defined by Bonchev and Trinajstić59 based on the hydrogen-supressed molecule graph |
Fig. 10 shows a 3-dimensional scatter plot of the drugs in the space spanned by the three features. The drugs are colored in red, when the majority of the cell lines is annotated as ‘resistant’ and blue if the majority of the cell lines is ‘sensitive’. In the plot, we observe that the compounds can be separated in groups with the same majority class using the drug features. Thus, we suspect that the model is using these groups to cast predictions. Intuitively, we would expect that sharing information for groups of similar compounds helps the multi-drug model to have a better prediction performance than the single-drug models. However, we observed that the multi-drug model performed worse than the single-drug models. Therefore, we conclude that the drug features complicated the drug sensitivity prediction task for the model. Overall, this might indicate that the drug features are not suited for this task.
 | ||
| Fig. 10 Three-dimensional scatter plot, where each point is a drug and the x-, y-, and z-axis show the three drug features from Table 1. The drugs are colored according to whether the majority of their tested cell line is sensitive (blue) or resistant (red). | ||
3.2 RF graph
To exemplify the RF graph results, we randomly sampled a drug from the 25 drugs that we considered for the single-drug models: Irinotecan. In the following, we investigate three different settings:(1) The complete test set graph of the randomly chosen drug Irinotecan (Fig. 11a).
(2) The graph of a randomly chosen single test cell line TYK-nu (COSMIC ID: 909774) for Irinotecan (Fig. 11b).
(3) The difference graphs between (1) and (2) (Fig. 11c and d).
Fig. 11 shows the respective graphs for the different settings. Notably, we only show the edges that have been used at least 4 times for the sample-specific graph (setting 2) or 3 times the number of test samples for the whole test set graph (setting 1) to make the visualization clearer. After deleting the edges accordingly, we excluded all nodes without in- or outgoing edges since we found that they generally have an average level higher than two. This indicates that they are neither often visited by the sample(s) (no or low weight edges) nor are they often used early in the tree to split the data (high level). Thus, we conclude that they are not particularly important, but rather make the visualization more complex, which is why we decided to remove them.
To interpret the graphs biologically, we map a priori knowledge to the observed graph visualizations. To this end, we queried the PharmacoDB database (release 1.1.1)60 and the COSMIC database (version v.100 from May 2024).61 PharamcoDB contains a list of genes that are known biomarkers for the efficacy of more than 700 compounds, including 17 out of our 25 investigated drugs. The COSMIC database provides information on the cell lines, particularly the mutated genes.
When comparing the first graph (whole test set) with the biomarker list from PharmacoDB, we observe that the three genes with the lowest average level (PPIC, SQSTM1, and SLAIN1) are also known biomarkers of Irinotecan. Thus, we conclude that our model did not only capture this biological relationship, but the graph is also able to visualize this. However, not only the color but also the degree (number of in-and outgoing edges) may be an indication of importance. From the three nodes with the highest degree (PPIC, SLAIN1, and ARHGEF6) two are known biomarkers, while we do not know whether there is a biological explanation for the third one.
The second graph (cf.Fig. 11b) shows the sample-specific graph for the test cell line TYK-nu, which is an ovary carcinoma cell line. To interpret the second graph, we used the mutation list from the COSMIC database and we found that from the 50 selected features for Irinotecan, five genes are mutated in TYK-nu:61 CSNK1G2, BCAT1, EPHX1, ASPHD2, and ARHGEF1. BCAT1 and ASPHD2 were already present in the overall test set graph (cf.Fig. 11a) while the other 3 features have a degree of 0 and are thus not shown in the graph.
When looking at the graph, we can already see that some edges differ between the test set graph and the sample specific graph. This indicates that the connections between the respective nodes differ between the investigated test cell line and the whole test set. This may be caused by mutations in the test cell line, which are not present in the other cell lines in the test set. To investigate this further, we constructed difference graphs. A difference graph from graph A – graph B contains all edges (and the connected nodes) that are present in A but not in B. Consequently, graph difference is not a symmetric operation s.t. There exist two difference graphs per graph pair: A, B and B, A. To calculate the difference graphs, we employed the Python library networkx,62 which is based on undirected unweighted graphs. Thus, in this step the edge direction and weight are eliminated.
Interestingly, in Fig. 11c, we observe that one of the edges connects SQSTM1 with ASPHD2, which is one of the mutated genes in TYK-nu. Possibly, the edge was more important in the overall test set compared to TYK-nu because of the mutation present in TYK-nu. Moreover, one of the edges in the graph 11d connects SQSTM1 with a gene from the mutation list of TYK-nu: EPHX1. Therefore, a possible conclusion could be that there is a biological relationship between the mutations in the cell line and the gene SQSTM1. However, in both graphs are also edges for which we found no literature evidence, and experimental validations would be needed to comprehensively interpret our results.
3.3 Conformal prediction
Conformal prediction (CP) is a versatile framework that can be applied on top of almost any machine learning model to enhance its reliability by providing certainty guarantees for its predictions, based on (un)certainty scores, for data assumed to be i.i.d.63 In particular, it provides sets (classification) or intervals (regression) that are guaranteed to contain the true response with at least a user-specified certainty. This can be especially interesting for high-risk applications of ML models, e.g., toxicity or drug sensitivity prediction,21,64 where such a certainty can support risk-aware decision making.Here we show the results for the application of CP with a 90% certainty guarantee to our new method, MORGOTH. For classification, we employ the two scoring functions that showed promising results for SAURON-RF:2 the true class and the Mondrian score. For regression, we use a quantile regression-based score, which is explained in our previous publication.21
Fig. 12 shows the results of the single-drug models with and without the application of CP evaluated for the classification task. Since we consider binary classification in this article, CP has four possible outcomes: {0}, {1}, {}, and {0, 1}. In the latter case, we order the classes by prediction probability (i.e., the first class is the one that is predicted, when not applying CP) s.t. there are actually 5 cases that we consider. A prediction is only classified as a true or false positive/negative after applying CP if CP results in a prediction set containing a single class. Thus, CP can reduce the number of true/false positives/negatives by outputting either an empty set or a set containing multiple classes (rather than a single class set). When comparing the number of false positives (FP) and negatives (FN) predicted by MORGOTH with the remaining single class predictions after CP, we observe that the true class score reduces the number of the original false predictions by almost 40%. However, the number of correctly predicted samples is only reduced by around 25%. For the Mondrian score, we obtain a reduction of the original false predictions around 65%. The number of original correct predictions, however, is only reduced by 30%. Comparing the Mondrian and the true class score, we observe that using the Mondrian score, the model generally casts less single class predictions. Thus, it reduces both, the number of correct and false predictions, more than the true class score. To evaluate the suitability of a score, one typically also calculates the CP efficiency (cf.21,64). The CP efficiency is defined as the ratio of single class predictions among all samples. Thus, the Mondrian score shows overall a lower efficiency (61%) than the true class score (78%), which means that the true class score outputs more meaningful (single class) predictions. Yet, the Mondrian score outputs less false predictions.
Fig. 13 shows the results for the single-drug models with and without CP evaluated for the regression task. For regression, the efficiency is measured by the interval size, where narrow intervals are desirable.21 We calculate two values to measure the interval size, i.e., the absolute and the relative interval size (cf.Fig. 13). While the first is the absolute size of the intervals normalized to the CMax viability range [0, 1], the latter is the size relative to the spanned range of the training set responses. We observe that the interval sizes are relatively large, i.e., relative interval size of 0.7 and absolute interval size of 0.63. Large interval sizes indicate that the model was uncertain in its prediction and could, therefore, not predict smaller intervals while fulfilling the certainty guarantee. Thus, from the large intervals, we can conclude that the learning task was difficult for the model.
The results for both, classification and regression, are similar to the results for reliable SAURON-RF.21
While the preceding analysis is just based on a randomly chosen example cell line, we also evaluated the prioritization task across all cell lines in the test set (cf.Fig. 15). Fig. 15 A shows the evaluation of the classification task evaluated per cell line across all 25 compounds. We observe that MORGOTH without CP reaches a mean MCC of 0.59 over all investigated cell lines, i.e., it can identify effective treatments. For regression (cf.Fig. 15B), the model reaches an PCC between predicted and actual CMax viability of 0.85, i.e., it can also sort drugs by efficiency. After applying CP for regression, the SCC between actual response and the upper interval boundary is 0.73. The observation of the relatively low SCC is consistent with our findings for reliable SAURON-RF.10 Together with the large prediction intervals (cf.Fig. 13), this suggests that a better conformity score for regression would be desirable. When calculating the percentage of correct predictions in the effective drug list, we can observe that MORGOTH without CP reaches a mean of 83.33%. Applying the true class CP score raises this value to 93.75% (Mondrian: 83.97%). However, when investigating whether the most effective drug has been identified, we observe that MORGOTH without CP can identify it in 93% of the cases, while applying CP lowers this number, i.e., 80% for true class score and 40% using Mondrian score. This observation indicates that the model has a certainty for this prediction lower than 90% since this is the guarantee that is fulfilled by applying CP. Thus, it would be important to improve the model s.t. it is less uncertain.
We conclude that MORGOTH is well suited to prioritize compounds for cell lines. Especially, using the true class score, we can reach a high precision in the effective drug list. However, the ranking using the upper CP interval boundaries should be improved.
3.4 Cluster analysis
As described above, we introduce a cluster analysis, which scores how well an unknown sample fits to the most similar training samples (i.e., the applicability domain). To this end, we calculate the silhouette score between the sample of interest and the training samples that are assigned to the same RF leaves. Intuitively, we would expect that a poorly fitting sample will lead to a high model uncertainty and/or a high error. Thus, we analyzed whether there is a relationship between poorly fitting samples (i.e., high silhouette score), high model uncertainty, and poor performance.Fig. 16 shows a scatter plot, where each point is a test sample of the single-drug models (cf. Section 3.1.1). On the y-axis, we show the squared deviation between predicted and actual CMax viability (i.e., a measure for regression performance) and on the x-axis the silhouette score. Moreover, the points (test samples) are colored indicating whether the model was certain (single class prediction) or not (set with both classes or empty set) using CP at a 90% certainty level using the true class score. To calculate the silhouette score, as defined in eqn (6), we need a distance function. Although MORGOTH offers the selection between various distance functions, we focus on one distance function following the observations and recommendations by Jaskowiak et al.49. We transform the PCC between the features of two samples into a distance by taking one minus the respective value. We refer to this distance as Pearson distance (PD).
By fitting a linear least squares regression line between these two values, we can investigate whether there is a correspondence between bad performance and high silhouette score that we would intuitively expect. We can see that the expectation is fulfilled for PD since the line has a positive slope.
We observe that the model is always certain for samples with a silhouette score lower than 0. We performed a one-sided Wilcoxon rank sum test to investigate whether the uncertain samples have statistically significant (p-value < 0.05) higher silhouette scores than the certain samples. The test reveals that the uncertain samples have higher values with a p-value of 1.54 × 10−25. Thus, for PD, our expectation that the silhouette score corresponds to some extent to model uncertainty is fulfilled. However, we must acknowledge that this correspondence is not perfect and there are samples with a high silhouette score, for which the model is certain. This might be caused by the fact that the distance function, on which the silhouette score is based, is not perfect.
4 Discussion
Although there is a plethora of ML approaches to predict anti-cancer drug sensitivity, the trustworthiness of these models is often not considered during model development.5 However, in such high-risk application cases, building trustworthy models is crucial. In this article, we, therefore, propose a novel approach that is not only performant but also implements two other aspects of trustworthiness that we consider exceptionally important, i.e., interpretability and reliability.Our novel approach is a multivariate RF for simultaneous classification and regression called MORGOTH. To increase the trustworthiness of our approach compared to other approaches, MORGOTH provides two novel functionalities, i.e., an RF graph representation to visualize the representative parts of the forest and a cluster analysis that scores how well unseen samples fit to the training samples that are assigned to the same RF leaves. To demonstrate its capabilities, we applied MORGOTH to the GDSC dataset and built not only single-drug but also multi-drug models. In our evaluation, we show how these new trustworthiness-related functionalities can make the models more reliable and interpretable. Moreover, in both settings (single- and multi-drug) MORGOTH tremendously outperforms state-of-the-art neural networks in terms of performance. We also showed that in a fair comparison the multi-drug MORGOTH model is outperformed by its single-drug counterpart.
In the following, we discuss the potential shortcomings in our analysis regarding the data we used, our feature selection, and our implementation. We place particular emphasis on aspects that are critical to ensuring the trustworthiness of our approach.
4.1 Data
We trained single-drug and multi-drug models on the GDSC dataset. For both settings, we split the data ‘cell-blind’, i.e., the cell lines from the training and test set are disjoint across all drugs. This simulates the application of our approach in personalized medicine, where all drugs are included in the training set, but the omics profiles of the cell lines (or patients) are not. Our evaluation did not only reveal that our model has state-of-the-art performance, but it can even increase the trustworthiness in the predictions, as the model performs well for the cell-blind scenario, which should approximate the application of our approach for personalized medicine.While our model demonstrated strong predictive performance on the GDSC dataset, we did not conduct cross-dataset validation using external resources such as the Cancer Cell Line Encyclopedia (CCLE).65 Future work should include such evaluations to assess the robustness and generalizability of our approach across independent datasets.
To characterize the cell lines, we solely used gene expression values as inputs for all our models. However, it might be desirable to use multi-omics data. In particular, discrete mutation or CNV data also have the advantage that they are easier to interpret than continuous gene expression values. However, gene expression is the most informative data type for predicting anti-cancer drug sensitivity, while other omics types like copy number variations (CNV) or mutations do not necessarily improve the predictive performance.34 Similar observations have been made by Wissel et al.66 for survival predictions for cancer, where the integration of noninformative or noisy omics types may even decrease the performance.
Furthermore, the multi-drug model relies on drug features. Our analysis of feature importance scores revealed that only three physicochemical properties impacted predictions of our model, while other features, such as MACCS and Morgan fingerprints, were not considered important. As shown by Sultan et al.27 the physicochemical properties are also one of the most informative data types when predicting molecular properties. Intuitively, we think that predicting general molecular properties (e.g., the solubility of a compound) should be an easier task for an ML model than predicting drug sensitivity for specific cell lines. However, in their analyses on various datasets for molecular property prediction, Sultan et al.27 show that neither with the simple nor with the complex drug representations the ML models reach R2 values above 0.3 for most of the investigated datasets. Therefore, we conclude that the development of novel drug representations is essential.29
We trained our models to predict the drug response for cell lines. However, the final goal would be to use such models in a treatment recommendation system s.t. it is crucial to transfer the task from cell lines to real patients.67 In particular, training on real cancer/patient data would increase the trustworthiness of the model. However, since it is typically more difficult to gather real patient data instead of model system-derived data10 – making labeled patient-data even more scarce – it might be beneficial to further investigate transfer learning algorithms for RFs as proposed by Segev et al.68. A further important consideration when developing models for clinical application is the representation of prior medical and biological knowledge. In particular, embedding a priori knowledge into the model ensures that important biological concepts are captured, which makes the model more trustworthy.
4.2 Feature selection and hyperparameter tuning
The FS algorithm that we used as well as the selected number of features (300) for the multi-drug model are based on the results of our previous benchmarking.22 However, we did not optimize it for the multi-drug model, which might, however, help to increase its performance. Moreover, Strobl et al.69 showed that combining continuous and discrete features may bias the selection of features for splits in the RF. Thus, combining the binary structural fingerprints with the continuous physicochemical properties and gene expression values may be the reason why our RF did only consider 3 drug features. Consequently, it could be beneficial to implement the unbiased variable selection algorithm by Strobl et al.69 for MORGOTH.For the hyperparameter tuning we used a 5-fold CV. We did not perform a grid search to extensively test all possible hyperparameter combinations, but tuned each hyperparameter individually. Thus, it is possible that another (better) hyperparameter combination was not tested in our hyperparameter tuning. To identify better hyperparameter combinations, it would be beneficial to run a grid search or apply Bayesian optimization.70
4.3 Random forest graph
To increase the interpretability of MORGOTH, we propose a graph visualization of the most important parts of the forest. To this end, the features in the forest are depicted as nodes, which are connected by directed edges indicating the transition from a parent to a child node in the forest. The features obtain their average rank as importance indicator. The edges are weighted by the number of times they are used by the sample(s). MORGOTH provides sample-specific graphs and graphs for groups of samples.To make the visualizations clearer, we currently exclude all edges below a certain frequency (cf. Section 3.2). This frequency-based selection of shown edges could result in structurally preferred edges (at the top of the tree) being overrepresented while edges further down are not represented, although they may be interesting. Thus, it might be advisable to find a more suitable way of identifying important edges. To this end, a reference distribution could be created for each edge for its usage, and then the observed frequency could be tested to see if it is significantly more often used.
4.4 Implementation of the multivariate random forest
While there is an R package (randomForestSRF71) that supports multivariate classification and regression, our splitting criterion function is more flexible, as we offer a tunable weight for the individual tasks. In particular, Fig. 5 shows that only 7 of 25 models chose λ = 0.5 as weight. Thus, flexible weighting improved the model performance. Moreover, randomForestSRF does not support the trustworthiness-related properties which are one of the main contributions of our package.As we fully implemented MORGOTH in Python, it has a higher runtime than other RF implementations in C++ or Cython, e.g., sklearn.43 However, the runtime can be optimized in future work, e.g., by integrating C Code.
The implementation of MORGOTH is modular s.t. the available functionalities can be extended easily. Most of the functionalities that are already implemented can be independently switched on and off (e.g., the construction of RF graphs or the cluster analysis), which can be useful to save runtime. Furthermore, in this article, we only used MORGOTH for simultaneous classification and regression. However, it is also possible to obtain reliable predictions for either of the tasks. In particular, MORGOTH is not only applicable to drug sensitivity data but to any supervised ML dataset.
5 Conclusion
When developing ML models, it is essential to not only consider the performance of the model but also other trustworthiness-related properties, such as reliability or interpretability. This particularly applies for high-risk-application cases, such as anti-cancer drug sensitivity prediction. In this article, we propose a novel approach, called MORGOTH, which is a multivariate RF for classification and regression with additional functionality rendering it reliable and interpretable. Our analyses on the GDSC dataset demonstrate that MORGOTH shows state-of-the-art performance in both tasks, i.e., classification and regression. In addition, we show that our model is reliable, because of the CP framework that allows the specification of certainty levels. Moreover, MORGOTH offers a cluster analysis that can be used to explain poor predictions and model uncertainty (assessed using CP) for specific samples, increasing both, reliability and interpretability. To further enhance the interpretability, MORGOTH provides not only feature importance scores but also a new visualization for RF. Although we evaluated our model on the GDSC database, we want to emphasize that MORGOTH is applicable to any supervised ML data set to obtain trustworthy predictions.Author contributions
K. L. and L. E. conceived the study. K. L., L. E., and L.-M. R. curated the data. L.-M. R. implemented the software and conducted the experiments. L.-M. R., L. H., and K. L. implemented the deep neural network architectures for benchmarking. L.-M. R. wrote the original draft of the manuscript. K. L., H.-P. L., and A. V. supervised the study. All authors analyzed the data, evaluated the results, and edited the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Our study was carried out using publicly available data from the GDSC at https://www.cancerrxgene.org/ with Release 8.3 from June 2020. The preprcoessed dataset as well as an example workflow can be found at zenodo https://doi.org/10.5281/zenodo.17977413. The code for MORGOTH can be found at https://github.com/volkamerlab/MORGOTH. Moreover, we published it as a pip installable package on PyPi https://pypi.org/project/morgoth/. The version of the code employed for this study is Version 1.1. Supplementary information: on the data, feature selection and hyperparameter tuning as well as the implementation. See DOI: https://doi.org/10.1039/d5dd00284b.Acknowledgements
We thank Nico Gerstner for fruitful discussions on trustwothy ML. This work was performed in the context of the European RADAR project and has received funding from the the European Union’s Horizon program under grant agreement No. 101178148. Views and opinions expressed are, however, those of the author(s) only and do not necessarily reflect those of the European Union. Neither the European Union nor the granting authority can be held responsible for them. This work is partially supported by the Ministry of Science and Culture of Lower Saxony through funds from the program zukunft.niedersachsen of the Volkswagen Foundation for the CAIMed – Lower Saxony Center for Artificial Intelligence and Causal Methods in Medicine’ project (grant no. ZN4257).References
- K. Lenhof, et al. , Bioinformatics, 2021, 37, 3881–3888 Search PubMed.
- K. Lenhof, L. Eckhart, N. Gerstner, T. Kehl and H.-P. Lenhof, Sci. Rep., 2022, 12, 13458 CrossRef CAS PubMed.
- T. Knijnenburg, et al. , Sci. Rep., 2016, 6, 36812 Search PubMed.
- C. De Niz, R. Rahman, X. Zhao and R. Pal, Algorithms, 2016, 9, 77 Search PubMed.
- K. Lenhof, L. Eckhart, L.-M. Rolli and H.-P. Lenhof, Briefings Bioinf., 2024, 25(5), bbae379 CrossRef CAS PubMed.
- I. Cortes-Ciriano, L. H Mervin and A. Bender, Curr. Pharm. Des., 2016, 22, 6918–6927 Search PubMed.
- G. Riddick, H. Song, S. Ahn, J. Walling, D. Borges-Rivera, W. Zhang and H. A. Fine, Bioinformatics, 2011, 27, 220–224 Search PubMed.
- R. Rahman, K. Matlock, S. Ghosh and R. Pal, Sci. Rep., 2017, 7, 1–11 CAS.
- K. Yanagisawa, et al. , Int. J. Mol. Sci., 2020, 21, year CrossRef PubMed.
- K. Lenhof, Machine learning-based anti-cancer drug treatment optimization, PhD thesis, Saarland University, 2024 Search PubMed.
- G. Nicora, M. Rios, A. Abu-Hanna and R. Bellazzi, J. Biomed. Inf., 2022, 127, 103996 CrossRef PubMed.
- M. Kukar and I. Kononenko, Machine Learning: ECML 2002: 13th European Conference on Machine Learning Helsinki, Finland, 2002, pp. 219–231 Search PubMed.
- V.-L. Nguyen, S. Destercke, M.-H. Masson and E. Hüllermeier, 27th International Joint Conference on Artificial Intelligence, IJCAI, 2018, pp. 5089–5095 Search PubMed.
- Y. Li, D. E. Hostallero and A. Emad, Bioinformatics, 2023, 39, btad390 CrossRef CAS PubMed.
- D. Wissel, N. Janakarajan, A. Grover, E. Toniato, M. R. Martínez and V. Boeva, bioRxiv, preprint, 2022, pp. 2022–11 Search PubMed.
- L. Grinsztajn, E. Oyallon and G. Varoquaux, Adv. Neural Inf. Process. Syst., 2022, 35, 507–520 CrossRef.
- V. Borisov, T. Leemann, K. Seßler, J. Haug, M. Pawelczyk and G. Kasneci, IEEE Trans. Neural Networks Learn. Syst., 2024, 35, 7499–7519 Search PubMed.
- R. Shwartz-Ziv and A. Armon, Inform. Fusion, 2022, 81, 84–90 CrossRef.
- L. Eckhart, K. Lenhof, L. Herrmann, L.-M. Rolli and H.-P. Lenhof, iScience, 2025, 28, 112622 Search PubMed.
- Y. Fang, P. Xu, J. Yang and Y. Qin, PloS One, 2018, 13, e0205155 CrossRef PubMed.
- K. Lenhof, L. Eckhart, L.-M. Rolli, A. Volkamer and H.-P. Lenhof, Sci. Rep., 2024, 14, 1–19 CrossRef PubMed.
- L. Eckhart, K. Lenhof, L.-M. Rolli and H.-P. Lenhof, Briefings Bioinf., 2024, 25, bbae242 CrossRef PubMed.
- M. Mathea, W. Klingspohn and K. Baumann, Mol. Inform., 2016, 35, 160–180 CrossRef CAS PubMed.
- K. Lenhof, L.-M. Rolli, L. Buhr, S. Roth, R. Binkyte-Sadauskiene, S. Schicktanz, M. Fritz, A. Volkamer and N. Beerenwinkel, The trustworthiness landscape in machine learning: a conceptual guide with applications in medicine, zenodo, 2025, DOI:10.5281/zenodo.17591544.
- K. Lee, D. Cho, J. Jang, K. Choi, H.-O. Jeong, J. Seo, W.-K. Jeong and S. Lee, Briefings Bioinf., 2023, 24, bbac504 CrossRef PubMed.
- M. P. Menden, F. Iorio, M. Garnett, U. McDermott, C. H. Benes, P. J. Ballester and J. Saez-Rodriguez, PLoS One, 2013, 8, e61318 CrossRef CAS PubMed.
- A. Sultan, M. Rausch-Dupont, S. Khan, O. Kalinina, A. Volkamer and D. Klakow, arXiv, preprint arXiv:2503.03360, 2025, DOI:10.48550/arXiv.2503.03360.
- D. Jiang, Z. Wu, C.-Y. Hsieh, G. Chen, B. Liao, Z. Wang, C. Shen, D. Cao, J. Wu and T. Hou, J. Cheminf., 2021, 13, 1–23 Search PubMed.
- X. An, X. Chen, D. Yi, H. Li and Y. Guan, Briefings Bioinf., 2021, 23, bbab393 CrossRef PubMed.
- F. Codicè, C. Pancotti and C. Rollo, J. Cheminform., 2025, 17, 33 CrossRef PubMed.
- M. Garnett, et al. , Nature, 2012, 483, 570–575 CrossRef CAS PubMed.
- W. S. Institute, GDSC Database, News, https://www.cancerrxgene.org/news, Online; accessed 24-March-2024 Search PubMed.
- W. Yang, et al. , Nucleic Acids Res., 2012, 41, D955–D961 CrossRef PubMed.
- J. Costello, et al. , Nat. Biotechnol., 2014, 32, 1202–1212 CrossRef CAS PubMed.
- D. Mendez, A. Gaulton, A. P. Bento, J. Chambers, M. De Veij, E. Félix, M. P. Magariños, J. F. Mosquera, P. Mutowo, M. Nowotka, M. Gordillo-Marañón, F. Hunter, L. Junco, G. Mugumbate, M. Rodriguez-Lopez, F. Atkinson, N. Bosc, C. J. Radoux, A. Segura-Cabrera, A. Hersey and A. R. Leach, Nucleic Acids Res., 2018, 47, D930–D940 CrossRef PubMed.
- M. Davies, M. Nowotka, G. Papadatos, N. Dedman, A. Gaulton, F. Atkinson, L. Bellis and J. P. Overington, Nucleic Acids Res., 2015, 43, W612–W620 CrossRef CAS PubMed.
- D. Weininger, J. Chem. Inf. Comput. Sci., 1988, 28, 31–36 CrossRef CAS.
- G. Landrum, et al., RDKit: version 2023.3.2, 2023, DOI:10.5281/zenodo.8053810, http://www.rdkit.org.
- A. Capecchi, D. Probst and J.-L. Reymond, J. Cheminf., 2020, 12, 1–15 Search PubMed.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef CAS PubMed.
- G. Landrum et al. , RDKit Documentation – rdkit.Chem.Descriptors module, E-mail: https://www.rdkit.org/docs/source/rdkit.Chem.Descriptors.html, Online; accessed 19-May-2024 Search PubMed.
- G. James, D. Witten, T. Hastie, R. Tibshirani and J. Taylor, Tree-based methods, An introduction to statistical learning: with applications in python, Springer International Publishing, Cham, 2023, pp. 331–366 Search PubMed.
- F. Pedregosa, et al. , J. Mach. Learn. Res., 2011, 12, 2825–2830 Search PubMed.
- H. Ishwaran, F. Tang, M. Lu and U. B. Kogalur, randomForestSRC: multivariate splitting rule vignette, 2021, https://randomforestsrc.org/articles/mvsplit.html, https://randomforestsrc.org/articles/mvsplit.html, Online; accessed 02-December-2025 Search PubMed.
- E. Heim, O. Wright and D. Shriver, arXiv, preprint arXiv:2503.00563, 2025, DOI:10.48550/arXiv.2503.00563.
- Y. Lin and Y. Jeon, J. Am. Stat. Assoc., 2006, 101, 578–590 CrossRef CAS.
- F. Rahutomo, T. Kitasuka, M. Aritsugiet al., The 7th international student conference on advanced science and technology ICAST, 2012, p. 1 Search PubMed.
- L. Wang, Y. Zhang and J. Feng, IEEE Trans. Pattern Anal. Mach. Intell., 2005, 27, 1334–1339 Search PubMed.
- P. A. Jaskowiak, R. J. Campello and I. G. Costa, BMC Bioinf., 2014, 1–17 Search PubMed.
- C. Croux and C. Dehon, Stat. Methods Appl., 2010, 19 CrossRef.
- N. Kwak and C.-H. Choi, IEEE Trans. Neural Netw., 2002, 13, 143–159 CrossRef CAS PubMed.
- Q. Liu, Z. Hu, R. Jiang and M. Zhou, Bioinformatics, 2020, 36, i911–i918 CrossRef CAS PubMed.
- H. Sharifi-Noghabi, O. Zolotareva, C. C. Collins and M. Ester, Bioinformatics, 2019, 35, i501–i509 CrossRef CAS PubMed.
- Y.-C. Chiu, H.-I. H. Chen, T. Zhang, S. Zhang, A. Gorthi, L.-J. Wang, Y. Huang and Y. Chen, BMC Med. Genomics, 2019, 12, 143–155 CrossRef PubMed.
- G. Van Rossum and F. Drake, Python 3 Reference Manual, CreateSpace, Scotts Valley, CA, 2009 Search PubMed.
- L. B. Kier and L. H. Hall, Pharm. Res., 1990, 7, 801–807 CrossRef CAS PubMed.
- P. Labute, J. Mol. Graphics Modell., 2000, 18, 464–477 CrossRef CAS PubMed.
- G. Landrum, What are the VSA descriptors?, E-mail: https://greglandrum.github.io/rdkit-blog/posts/2023-04-17-what-are-the-vsa-descriptors.html, Online; accessed 07-August-2024 Search PubMed.
- D. Bonchev and N. Trinajstić, J. Chem. Phys., 1977, 67, 4517–4533 CrossRef CAS.
- P. Smirnov, V. Kofia, A. Maru, M. Freeman, C. Ho, N. El-Hachem, G.-A. Adam, W. Ba-alawi, Z. Safikhani and B. Haibe-Kains, Nucleic Acids Res., 2017, 46, D994–D1002 CrossRef PubMed.
- Z. Sondka, N. B. Dhir, D. Carvalho-Silva, S. Jupe, Madhumita, K. McLaren, M. Starkey, S. Ward, J. Wilding, M. Ahmed, J. Argasinska, D. Beare, M. S. Chawla, S. Duke, I. Fasanella, A. G. Neogi, S. Haller, B. Hetenyi, L. Hodges, A. Holmes, R. Lyne, T. Maurel, S. Nair, H. Pedro, A. Sangrador-Vegas, H. Schuilenburg, Z. Sheard, S. Y. Yong and J. Teague, Nucleic Acids Res., 2023, 52, D1210–D1217 CrossRef PubMed.
- A. Hagberg, P. Swart and D. S Chult, Exploring network structure, dynamics, and function using NetworkX, Los alamos national lab.(lanl), los alamos, nm (united states) technical report, 2008 Search PubMed.
- A. Angelopoulos and S. Bates, arXiv, preprint arXiv:2107.07511, 2021, DOI:10.48550/arXiv.2107.07511.
- A. Morger, M. Mathea, J. H. Achenbach, A. Wolf, R. Buesen, K.-J. Schleifer, R. Landsiedel and A. Volkamer, J. Cheminform., 2020, 12, 24 CrossRef CAS PubMed.
- J. Barretina, G. Caponigro, N. Stransky, K. Venkatesan, A. A. Margolin, S. Kim, C. J. Wilson, J. Lehár, G. V. Kryukov and D. Sonkin, et al. , Nature, 2012, 483, 603–607 CrossRef CAS PubMed.
- D. Wissel, D. Rowson and V. Boeva, Cell Rep. Methods, 2023, 3 CAS.
- D. Maeser, W. Zhang, Y. Huang and R. S. Huang, Curr. Opin. Struct. Biol., 2024, 84, 102745 CrossRef CAS PubMed.
- N. Segev, M. Harel, S. Mannor, K. Crammer and R. El-Yaniv, IEEE Trans. Pattern Anal. Mach. Intell., 2016, 39, 1811–1824 Search PubMed.
- C. Strobl, A.-L. Boulesteix, A. Zeileis and T. Hothorn, BMC Bioinf., 2007, 8, 1–21 CrossRef PubMed.
- J. Wu, X.-Y. Chen, H. Zhang, L.-D. Xiong, H. Lei and S.-H. Deng, J. Electron. Sci. Technol., 2019, 17, 26–40 CrossRef.
- H. Ishwaran and U. Kogalur, Fast Unified Random Forests for Survival, Regression, and Classification, RF-SRC, 2025 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |













