Open Access Article
Open Access ArticlePhotoswitchable imaging contrast agents as an emerging frontier in precision bioimaging
Xiaoqiong
Tan†
ad,
Qiankun
Ni†
b,
Ting
Luo
b,
Ki Taek
Nam
f,
Hong-Bo
Cheng
 *ae,
Paul J.
Dyson
*ae,
Paul J.
Dyson
 *e,
Xing-Jie
Liang
*e,
Xing-Jie
Liang
 *b and
Juyoung
Yoon
*b and
Juyoung
Yoon
 *cg
*cg
aState Key Laboratory of Organic-Inorganic Composites, Beijing Laboratory of Biomedical Materials, College of Materials Science and Engineering, Beijing University of Chemical Technology, Beijing 100029, P. R. China. E-mail: chenghb@mail.buct.edu.cn
bCAS Key Laboratory for Biomedical Effects of Nanomaterials and Nanosafety, CAS Center for Excellence in Nanoscience, National Center for Nanoscience and Technology, Beijing 100190, P. R. China. E-mail: liangxj@nanoctr.cn
cDepartment of Chemistry and Nanoscience, Ewha Womans University, Seoul 03760, Republic of Korea. E-mail: jyoon@ewha.ac.kr
dDepartment of Biomedical Engineering, College of Future Technology, Peking University, Beijing 100871, P. R. China
eInstitute of Chemical Sciences and Engineering, École Polytechnique Fedérale de Lausanne (EPFL), Lausanne CH-1015, Switzerland. E-mail: paul.dyson@epfl.ch
fSeverance Biomedical Science Institute, Brain Korea 21 PLUS Project for Medical Science, College of Medicine, Yonsei University, Seoul 03760, Republic of Korea
gGraduate Program in Innovative Biomaterials Convergence, Ewha Womans University, Seoul 03760, Korea
First published on 17th February 2026
Abstract
Photoswitchable molecules exhibit reversible photoisomerization between distinct isomeric states under alternating light irradiation, possessing unique photophysical properties. Derived from these molecules, photoswitchable imaging contrast agents (PICAs) offer significant advantages over conventional agents: (1) background suppression through state-switching algorithms; (2) continuous imaging capability enabled by high fatigue resistance; (3) rapid signal modulation minimizing acquisition time. A comprehensive analysis of PICA advancements and persistent challenges across bioimaging applications is imperative. This review systematically evaluates several representative photoswitchable molecules as PICAs and proposes innovative molecular design and assembly strategies to address the limitations hindering the widespread implementation of PICAs in bioimaging, such as hydrophobicity, short-wavelength excitation, and suboptimal emission performance. Additionally, recent advances in PICAs for fluorescence, phosphorescence, photoacoustic, magnetic resonance and Raman imaging are summarized, highlighting their significant contributions to improving bioimaging quality and precision. Finally, an outlook is provided on the intersection of PICA design and property optimization for propelling both preclinical physiological research and clinical diagnostic and monitoring technologies.
1. Introduction
As a bridge connecting molecular mechanisms and macroscopic physiological properties, bioimaging provides an indispensable tool in preclinical physiological research, medical diagnosis, and drug development. The core challenge in bioimaging technology is achieving a balance among molecular specificity, spatiotemporal resolution, and biocompatibility in complex biological systems. Imaging contrast agents (ICAs) have made considerable strides in bioimaging, driven by advances in synthetic and assembly techniques that enhance their capabilities for biological analysis and disease diagnosis. Photoswitchable molecules, which undergo reversible structural transition between trans and cis forms or between ring-opened and ring-closed configurations resulting in significant changes in fluorescence or absorption spectra upon irradiation at two distinct wavelengths, serve as the foundation for developing photoswitchable ICAs (PICAs).1 Owing to their distinctive photoactive and photo-controllable physical properties, PICAs outperform conventional ICAs in several key ways, offering a revolutionary solution to overcome current limitations in bioimaging.2–5 First, the switchable states of PICAs can effectively enhance imaging contrast and eliminate imaging background noise through specific algorithms, while most conventional ICAs emit a single and “always on” signal.6–8 Second, the high fatigue resistance of PICAs facilitates continuous and stable imaging,9,10 contrasting with some conventional dyes, such as indocyanine green (ICG), which experience rapid photobleaching even under short irradiation times. Third, the fast photo-response of PICAs promotes instantaneous switching between two different signals, drastically shortening imaging time.11–13 Additionally, rational molecular design, such as group modification, structural adjustment, and chemical conjugation, along with advanced assembly strategies, has further expanded the bioimaging applications of PICAs.14–18 Further advancements could aim at improving their stability, hydrophilicity, biocompatibility, targeted delivery, deep penetration capability under long-wavelength excitation, emission performance and photoisomerization efficiency.19–23To date, an increasing number of studies have demonstrated the utility of PICAs across diverse imaging modalities, including fluorescence imaging (FI), phosphorescence imaging (PI), photoacoustic imaging (PAI), magnetic resonance imaging (MRI), and Raman imaging (RI) (Fig. 1). Despite these developments, a comprehensive review summarizing the role of PICAs in various bioimaging techniques is still lacking. Therefore, a systematical evaluation of recent progress in PICA-based bioimaging is both timely and necessary.
This review systematically evaluates several representative photoswitchable molecules functioning as PICAs and elucidates their underlying photoisomerization mechanisms. It also describes diverse optimization strategies focused on addressing current challenges that hinder the broader applications of PICAs across different bioimaging modalities. Finally, an outlook on this emerging field at the intersection of PICA design and performance optimization is presented. By consolidating these advancements, this review aims to support both preclinical physiological research and clinical diagnostic and monitoring technologies for various diseases.
2. Photoswitchable molecules as photoswitchable imaging contrast agents
The photoswitchable (photochromic) phenomenon was first described by Y. Hirshberg in the 1950s, who proposed that the reversible structural transformation of dianthrone under ultraviolet (UV) irradiation was related to photo-responsive switching behavior.24,25 Photochromism is now typically defined as the photo-induced reversible switching between two isomers, each displaying distinct physical properties when exposed to specific wavelengths of electromagnetic radiation, such as UV, visible (Vis), or infrared light.26 Consequently, these molecules are designated as photoswitchable or photochromic molecules.The reversible transition of photoswitchable molecules is driven by various mechanisms, including ring-opening/closing reaction, trans-to-cis or cis-to-trans isomerization, intramolecular proton transfer, intramolecular electron transfer, and cycloaddition.27,28 Photoswitchable molecules have been extensively studied for preclinical physiological research, disease diagnosis and monitoring. These photoswitchable molecules and their derivatives function as PICAs themselves or as switches that regulate the photophysical properties of additional contrast agents. This section discusses several representative photoswitchable molecules either served directly as PICAs or used to construct PICAs, including azobenzene (Azo), spiropyran (SP), diarylethene (DAE), and photoswitchable proteins, with particular elucidation on their photoswitching processes and mechanisms (Fig. 2).
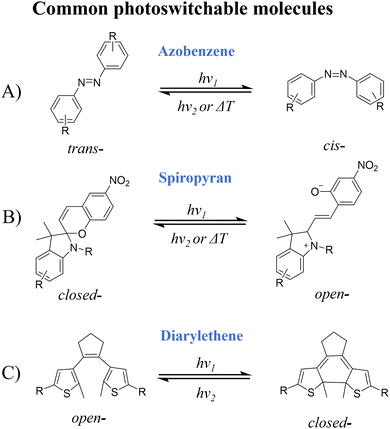 | ||
| Fig. 2 Common photoswitchable molecules concluding (A) azobenzene, (B) spiropyran, (C) Diarylethene and their photoisomerization behavior. | ||
2.1. Azobenzene
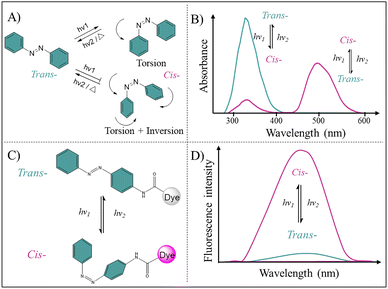
Azo is the most prominent photoswitchable molecule and has been studied extensively. Azo exists as the thermodynamically stable trans configuration at room temperature. It undergoes trans-to-cis isomerization upon UV irradiation and can reversibly return to the trans isomer through Vis light irradiation, slow thermal relaxation, or heating (Fig. 2A).29 This trans-to-cis photoisomerization proceeds by torsion around N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bond or torsion accompanied with in-plane inversion (Fig. 3A).30,31 The photoswitching behavior of Azo results in distinct light absorption profiles: the trans isomer absorbs light irradiation below 400 nm, while the cis isomer exhibits a red-shifted absorption above 440 nm, attributable to the occurrence of a n–π* transition (Fig. 3B).32 Considering these unique optical properties, combined with its structural tunability and inherent stability, Azo holds great potential for developing advanced PICAs.33,34
N bond or torsion accompanied with in-plane inversion (Fig. 3A).30,31 The photoswitching behavior of Azo results in distinct light absorption profiles: the trans isomer absorbs light irradiation below 400 nm, while the cis isomer exhibits a red-shifted absorption above 440 nm, attributable to the occurrence of a n–π* transition (Fig. 3B).32 Considering these unique optical properties, combined with its structural tunability and inherent stability, Azo holds great potential for developing advanced PICAs.33,34
Because of its intrinsic non-fluorescent nature, Azo is typically employed as a building block to develop Azo-based PICAs, through covalent conjugation or non-covalent interaction with fluorescent dyes or other imaging contrast agents. In general, photoisomerization of Azo regulates the fluorescence intensity or wavelength of adjacent dyes through altering their electronic environment and rigidity constraints (Fig. 3C). In its trans state, Azo functions as an energy acceptor, quenching dye fluorescence through Förster resonance energy transfer (FRET) effect (Fig. 3D).35,36 When Azo transforms to the cis state under UV light irradiation, the conjugated system is disrupted, thus enhancing the molecular dipole moment and polarity. This change eliminates the absorption-emission spectral overlap between Azo and dye, restoring the dye's emission. Conversely, Vis light irradiation reverts Azo to its trans isomer, re-establishing the conjugated system to recover the non-fluorescence state. Moreover, the trans-to-cis isomerization increases molecular steric hindrance, disrupting the planar, conjugated structure of trans-Azo dye. This alleviates its aggregation-caused quenching (ACQ) effect and promotes aggregation-induced emission (AIE) behavior. Due to the distinct spatial conformation change, trans-cis isomerization of Azo has been widely employed to reversibly modulate macromolecular or supramolecular assembly structures for precise control over their emission properties.37–39
However, the broader application of Azo-based PICAs in bioimaging is still hindered by their intrinsic hydrophobicity, short-wavelength excitation, and fast thermal relaxation. The second section will elaborate on the optimization strategies through structural engineering and functional modification to address these challenges.
2.2. Spiropyran
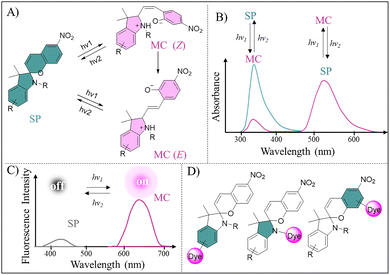
SP represents another promising candidate for photoswitchable bioimaging, featuring a distinct photoisomerization mechanism from Azo. SP maintains a thermal-stable ring-closed form in its native state, with UV absorption (<400 nm). Upon appropriate light irradiation, SP undergoes reversible photochemical cleavage of the C–O bond to open the pyran ring, transforming into the merocyanine (MC) isomer (Fig. 2B).40 Moreover, MC exhibits cis–trans (Z/E) isomerism at its C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond, with the trans (E) configuration typically being thermodynamically favored arising from decreased steric strain (Fig. 4A). Notably, the SP → MC isomerization generates an extended π-conjugated system with charge separation, resulting in a large electric dipole moment.41 This unique structural rearrangement red-shifts the absorption spectral to the 500–600 nm region (Fig. 4B) and dramatically enhances fluorescence enhancement from a weakly emissive state (low intensity at <450 nm) to a strong fluorescent one (high intensity at >600 nm) (Fig. 4C). Conversely, the MC → SP isomerization occurs through spontaneous thermal relaxation in the dark, heating, or Vis light exposure, with reduced absorption and fluorescence.41,42 The reversible photoisomerization of SP enables precise control over the on/off state of fluorescence, or facilitates switching between multiple emission wavelengths for monochromatic or multicolor FI.43,44 In addition to serving as a direct photoswitchable fluorophore, SP provides three common modification sites for covalent dye conjugation, thereby allowing modulation of the fluorescence of adjacent dyes via FRET (Fig. 4D).
C bond, with the trans (E) configuration typically being thermodynamically favored arising from decreased steric strain (Fig. 4A). Notably, the SP → MC isomerization generates an extended π-conjugated system with charge separation, resulting in a large electric dipole moment.41 This unique structural rearrangement red-shifts the absorption spectral to the 500–600 nm region (Fig. 4B) and dramatically enhances fluorescence enhancement from a weakly emissive state (low intensity at <450 nm) to a strong fluorescent one (high intensity at >600 nm) (Fig. 4C). Conversely, the MC → SP isomerization occurs through spontaneous thermal relaxation in the dark, heating, or Vis light exposure, with reduced absorption and fluorescence.41,42 The reversible photoisomerization of SP enables precise control over the on/off state of fluorescence, or facilitates switching between multiple emission wavelengths for monochromatic or multicolor FI.43,44 In addition to serving as a direct photoswitchable fluorophore, SP provides three common modification sites for covalent dye conjugation, thereby allowing modulation of the fluorescence of adjacent dyes via FRET (Fig. 4D).
As an important photoswitchable molecule developed after SP, spirooxazine (SO) has emerged as a research hotspot in the frontier fields such as super-resolution imaging, target labeling, and in vivo tracking, due to its unique photophysical properties and excellent stability. Sharing similar structure with SP, SO also exhibits reversible photoisomerization where UV light irradiation cleaves the C–O bond to open the oxazine ring, and Vis light irradiation triggers reversible ring closing through an electrocyclization reaction that reforms the C–O bond (Fig. 5A).40 Unlike SP, which contains only oxygen atom in the pyran ring, SO contains both oxygen and nitrogen atoms in the heterocycle, along with an extended π-conjugated system. As a result, SO has longer half-life and higher photostability compared to SP. The thermal stability of SO in its ring-opened form lays the foundation for its application in time-resolved imaging.45–49 Furthermore, SO exhibits excellent fatigue resistance, withstanding over 104 switching cycles without significant photodegradation, far surpassing SP that typically achieves fewer than 103 cycles.50,51 This superior performance primarily stems from the electronic effect and steric hindrance effect of its oxazine ring, which together effectively suppress photobleaching.52
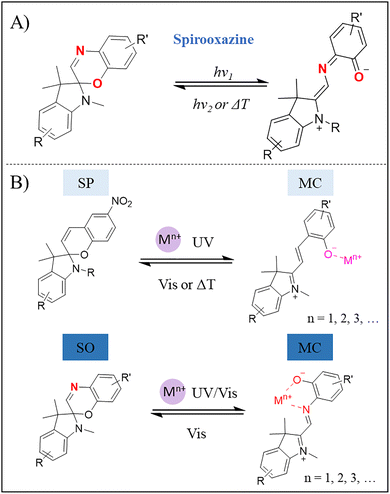 | ||
| Fig. 5 (A) The ring-opening/ring-closing reactions of SO. (B) Responsiveness of SP and SO to metal ions. | ||
Apart from functioning as photoswitchable fluorescent probes, both SP and SO perform superior potential for biosensing. This depends on the distinct responsiveness of their molecular structures: the oxygen atom in the pyran/oxazine ring enables sensitive detection of metal ions, while the reactive C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond allows for the recognition of various analytes (e.g., CN−, H2S, SO2).53,54 Metal ion binding promotes the UV light-induced ring-opening, stabilizes the resultant ring-opened isomer and acts as an auxochrome with unique absorption and fluorescence signals (Fig. 5B). The coordination affinity of the oxygen and nitrogen atoms in SP or SO varies considerably among different metal ions, such as Li+, Na+, K+, Mg2+, Cu2+, Ca2+, Fe2+, Fe3+, Zn2+, and Ag+. The coordination of metal ions alters the geometry and π-conjugation extent of SO, leading to distinct fluorescence signals upon external stimulation. This property positions SO as a promising probe for detecting diverse metal ions in living organisms.55,56
C bond allows for the recognition of various analytes (e.g., CN−, H2S, SO2).53,54 Metal ion binding promotes the UV light-induced ring-opening, stabilizes the resultant ring-opened isomer and acts as an auxochrome with unique absorption and fluorescence signals (Fig. 5B). The coordination affinity of the oxygen and nitrogen atoms in SP or SO varies considerably among different metal ions, such as Li+, Na+, K+, Mg2+, Cu2+, Ca2+, Fe2+, Fe3+, Zn2+, and Ag+. The coordination of metal ions alters the geometry and π-conjugation extent of SO, leading to distinct fluorescence signals upon external stimulation. This property positions SO as a promising probe for detecting diverse metal ions in living organisms.55,56
Despite ongoing molecular optimization of SP and SO, its applications in bioimaging remain underexplored, primarily due to several persistent challenges: (1) excitation wavelengths are largely confined to the UV-Vis region (<600 nm); (2) enhanced thermal stability may paradoxically limit the photoisomerization efficiency; (3) suboptimal biocompatibility hinders effective cellular uptake; and (4) low fluorescence quantum yield. In the second section, the corresponding solutions strategies will be discussed.
2.3. Diarylethene
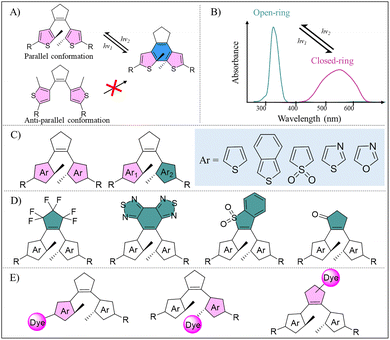
DAE has a core structure consisting of two aromatic rings (typically thiophene or benzothiophene units) bridged by the ethylene bond in cyclopentene. Upon irradiation with altering lights, DAE undergoes reversible photocyclization, transforming from an unconjugated, ring-opened isomer to a conjugated, ring-closed isomer (Fig. 2C).57 The ring-opened DAE isomer exists in two conformational states: parallel and anti-parallel (Fig. 6A). Notably, photoisomerization occurs exclusively in the anti-parallel conformation, whereas the parallel conformation remains photochemically inactive.57,58 This conformational dependence is a critical consideration in the molecular design and application of DAE.UV light exposure dynamically rearranges the π-electron cloud of the vinyl double bond and the π–π conjugated system in the DAE skeleton. According to the Woodward–Hoffmann rules, the photocyclization of the ring-opened DAE proceeds by the photochemical conrotatory electrocyclization of its central 6π-electron 1,3,5-hexatriene system to form a new C–C bond.59 This process results in the isomerization from the ring-opened state to the ring-closed state to augment intramolecular conjugation, consequently red-shifting the absorption spectrum (λmax from ∼300 nm to ∼550 nm) along with visible color change (Fig. 6B). Conversely, irradiation with Vis light or near-infrared (NIR) light triggers the ring-opening reaction of DAE, cleaving the newly formed C–C bond and reverting the molecule to its initial ring-opened state.
Diverse molecular design strategies have been proposed to precisely tune the photoswitching properties of DAE. The reversible photocyclization activity is governed by the core architecture of DAE, where the two core aryl groups offer the foundation for electron conjugation. Their electronic properties and steric hindrance determine the overall stability and reaction activity of the molecule. DAE derivatives exist in either symmetric configuration (with identical aryl ring type) or asymmetric configuration (with distinct aryl ring type) (Fig. 6C). Typical aryl ring units include thiophene, benzothiophene, sulfonyl-substituted thiophene, thiazole, and oxazole. Asymmetric DAE commonly improves photoisomerization efficiency and photostability, yet their synthetic complexity is a challenge. Furthermore, the structure of the vinyl bridge plays a crucial role in modulating the photoswitching performance of DAE, such as photoisomerization efficiency, chemical stability, fatigue resistance, and thermal half-life (Fig. 6D).
In addition, DAE derivatives display high value in bioimaging because of their exceptional thermal stability (both isomers are indefinitely stable at room temperature in the darkness) and fatigue resistance (allowing hundreds to thousands of switching cycles without pronounced degradation). These properties render DAE particularly suitable for long-term FI, where they support repeated on/off toggling and diverse imaging modalities.20,60,61
As a molecular switch, DAE regulates the fluorescence emission of conjugated dyes via a FRET process.62 Fluorophores are typically attached to one or both terminal thiophene units of the DAE core, which allows effective fluorescence modulation through a photocyclization-induced quenching mechanism (Fig. 6E). Moreover, DAE can be co-encapsulated with fluorophores into nanoparticles via supramolecular interactions, thereby achieving FRET-based reversible fluorescence switching for bioimaging.12 However, similar to other photoswitchable molecules, DAE faces inherent challenges in bioimaging applications, such as short-wavelength activation, hydrophobicity, and weak emission. The main strategy for optimizing the properties of DAE is molecular engineering through the introduction of functional groups, such as: (1) heterocyclic aryl groups to red-shift absorption; (2) fluorophores to improve emission; and (3) hydrophilic moieties to increase solubility in biological media.10,14,63–66
2.4. Other photoswitchable molecules
In addition to the development based on classical molecule scaffolds, other photoswitchable molecules, such as rhodamine, dihydropyrene (DHP), hexaarylbiimidazole (HABI), and donor–acceptor Stenhouse adduct (DASA), have been explored as PICAs for photoswitchable bioimaging.The photoswitchable reaction of rhodamine amides was first proposed by Knauer and Gleiter in the 1970s, a discovery that has since spurred the development of numerous photoswitchable rhodamine derivatives.67,68 The photoswitching mechanism of rhodamine depends on the reversible cleavage and reformation of the spirocyclic bond, which interconverts between a fluorescent zwitterionic (ring-opened) state and a non-fluorescent spirolactone (ring-closed) state. Rhodamine offers advantages over conventional fluorescent probes in super-resolution imaging, including high fluorescence quantum yield and excellent photochemical stability.9 However, the short lifetime of its fluorescent ON-state and inherent switching stochasticity fundamentally constrain the extension of temporal imaging windows and precise spatiotemporal coordination, necessitating a trade-off between spatiotemporal super-resolution performance. A novel rhodamine derivative, Rh-Gly, was developed with remarkable fluorescence brightness and prolonged ON-state stability under UV light irradiation.69 The presence of a carboxyl group in Rh-Gly stabilizes its zwitterionic ring-opened conformation via intramolecular hydrogen bonding. This yields a prolonged ON-state lifetime (68.5 ms) and enhanced fluorescence brightness (3.57 × 104 photons per s), enabling simultaneous high spatial resolution (∼50 nm) and temporal resolution (10 s) in live-cell super-resolution imaging.
HABI is another class of photoswitchable molecule characterized by the reversible dimerization of 2,4,5-triphenylimidazolyl radical (TPIR).70–73 Upon 365 nm light irradiation, HABI dissociates into TPIR radicals, which can subsequently revert to HAPI through thermal transformation.74 HABI has attracted increasing attention due to its rapid photo-responsive behavior and robust fatigue resistance. Owing to this reversible dimerization involving bond cleavage and reformation, HABI has been covalently attached to BODIPY dye to modulate their fluorescence.75
Compared to conventional UV-triggered photoswitchable molecules, negative photoswitchable molecules such as DHP and DASA undergo photoisomerization upon Vis/NIR light irradiation. This reduces the phototoxicity to cells or living organisms, rendering them more suitable for prolonged imaging. Owing to the rigid planar structure and extended π-system, DHP absorbs at longer wavelength (>600 nm) than conventional photoswitchable molecules.76,77 Its negative photoswitching behavior involves a 6π-electrocyclic rearrangement that disrupts the π-conjugation, yielding a decolorized cyclopentadiene upon Vis/NIR light irradiation. However, photoisomerization under illumination exceeding 750 nm remains challenging. One viable synthetic strategy is to introduce donor and acceptor groups at the 2,7- or 4,9-substituted positions of DHP, which red-shift the absorption spectrum into the NIR region reaching 900 nm.78 In addition, DASA represents a new class of negative photoswitchable molecules with a D–π–A architecture. This structure allows reversible photoisomerization between a colored (open) form and a colorless (closed) form upon Vis-NIR light (490–700 nm) irradiation.79
Despite the development of numerous novel photoswitchable molecules, their applications in bioimaging remain limited and require further exploration.
2.5. Photoswitchable proteins
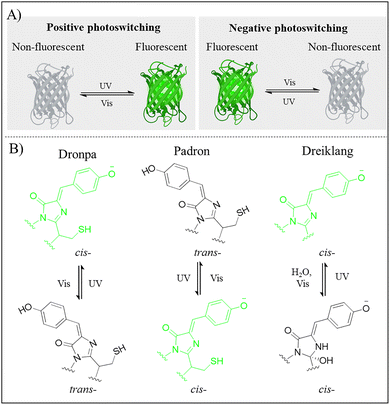
The advent of reversibly photoswitchable fluorescent proteins (RSFPs) has revolutionized bioimaging, particularly by super-resolution microscopy to visualize subcellular dynamics at the nanoscale in living systems.80–82 The first RSFP, Dronpa, was discovered by Ando et al. in bacterial studies.83 Since then, a wide variety of RSFPs have been rapidly developed to meet the specific demands of advanced bioimaging modes. Under alternating light irradiation at two distinct wavelengths (typically UV light and Vis light), RSFPs undergo reversible positive or negative photoswitching between fluorescent and non-fluorescent states (Fig. 7A).84 This is primarily driven by light-induced conformational change of the chromophore in RSFPs, though it may also involve protonation or other complex photophysical processes.85Fig. 7B illustrates the photoswitching mechanisms of three classical RSFPs (Dronpa, Padron, and Dreiklang) based on distinct structural changes of chromophores: cis–trans isomerization, trans–cis isomerization, and hydration/dehydration reaction, respectively. The chromophore of Dronpa follows by a negative photoswitching mechanism: Vis light irradiation (e.g., 488 nm) transforms it from a fluorescent deprotonated cis-state to a non-fluorescent protonated trans-state, whereas UV or blue light irradiation (e.g., 405 nm) reverses this process to restore the fluorescence.86 Padron, a Dronpa variant, maintains the same chromophore structure but displays positive photoswitching process: UV or blue light irradiation (e.g., 405 nm) triggers trans-to-cis isomerization and deprotonation, consequently activating bright green fluorescence at 522 nm.87 Conversely, Vis light (e.g., 488 nm) returns the chromophore to a non-fluorescent protonated trans-state. Dreiklang is another RSFP that holds a distinctive photoswitching mechanism. Unlike the cis–trans isomerization typical of many RSFPs, Dreiklang relies on a reversible hydration/dehydration reaction occurring at a specific carbon atom in the imidazolidinone ring.88 Specifically, 405 nm light irradiation triggers a hydration reaction that converts the chromophore from a fluorescent state to a non-fluorescent state, while 365 nm light irradiation drives a dehydration reaction to restore the fluorescence. However, most RSFPs are limited by intrinsic photophysical properties, including finite switching cycles, low photon output, and short-wavelength-light activation, which collectively compromise their applications in live-cell super-resolution imaging and in vivo imaging.Over the past decade, considerable efforts have been directed toward engineering improved RSFPs for super-resolution and in vivo imaging. The Padron-derived mutant, Konihoor, exhibited rapid switching kinetics, enhanced photostability, and high quantum yields, enabling successful super-resolution imaging in living cells via reversible saturable optical fluorescence transition (RESOLFT).7 Moreover, Kohinoor showed high photostability with minimal fluorescence decay per switching cycle (∼0.1%, compared to Padron's 2.5%). It also achieved 17-fold more cycles than Padron (340 cycles versus 20 cycles) before 50% intensity loss, demonstrating accelerated switching kinetics. Kohinoor exhibited substantially higher quantum yield than Padron in both states: the open state (Φon = 0.02) and closed state (Φoff = 0.15) exceeded Padron's corresponding values (Φon ≈ 0.003, Φoff ≈ 0.04) by 6.7-fold and 3.75-fold, respectively. These improvements enabled biocompatible live-cell imaging at ultralow laser intensity (0.004 J cm−2).
Most RSFPs are activated by UV light, raising concerns over phototoxicity and scattering effect. Therefore, there is a demand for the development of RSFPs excited by longer wavelength. Several red RSFPs, including rsCherry, rsCherryRev, and rsTagRFP, have been used as PICAs for super-resolution FI under Vis light.89 More recently, three novel RSFPs derived from FusionRed (a red-emitting RSFP) enable reversible switching upon the irradiation of green light (510 nm) and orange light (590 nm).90 These rsFusionReds exhibited outstanding fatigue resistance (>1000 photoswitching cycles) and a rapid photoswitching rate (58.3 ± 0.4 ms), facilitating nanoscale live-cell imaging. Their operation at wavelengths beyond 500 nm also enabled the RESOLFT implementation with mitigated DNA damage and photodamage. Furthermore, the advantages of NIR light have directed increasing attention toward NIR fluorescence proteins for light-controlled cellular and in vivo imaging.91 Advancements in the photoacoustic properties of NIR-induced photoswitchable proteins have also motivated studies exploring NIR RSFPs for high-resolution PAI.92,93
Despite progress in developing biologically safe and genetically encodable NIR RSFPs, their broad bioimaging adoption remains constrained by some shortcomings, such as low brightness, poor photostability, insufficient fatigue resistance, and complex genetic circuitry.48,89 Future efforts should address these challenges to enhance the overall photoswitching performance of NIR RSFPs.
In brief, these representative classical photoswitchable molecules, including Azo, SP, DAE, and RSFPs, have demonstrated superior potential for development into PICAs. Nonetheless, their widespread adoption in bioimaging confronts considerable challenges. The photoisomerization of most photoswitches is driven by short-wavelength light, which falls outside the optical window of biological tissues. To overcome this limitation, further molecular modifications are necessary to red-shift their absorption toward longer wavelength. Strategies for elongating the π-conjugated structure or introducing appropriate fluorophores are viable. Alternatively, integrating photoswitchable molecules with upconversion materials enables excitation by NIR light. Since most photoswitchable molecules are inherently hydrophobic, incorporating hydrophilic moieties can improve biocompatibility.28,94 Additionally, coupling photoswitchable molecules with other functional materials is a promising route to tailor specific physicochemical properties.95,96
3. Optimization strategies on photoswitchable imaging contrast agents
Despite considerable improvements in photoswitchable molecule design, some challenges continue to impede the practical application of PICAs in bioimaging, particularly for in vivo use. These include: (1) poor stability in complex biological environments hinders prolonged imaging; (2) hydrophobicity-induced aggregation impairs photoswitching sensitivity and raises safety concerns; (3) dependence on short-wavelength excitation limits their effectiveness in whole-organism and deep-tissue imaging; (4) inadequate emission performance; and (5) low photoisomerization efficiency diminishes imaging resolution. Molecular design strategies, such as functional group modification, structural adjustment, and chemical conjugation, have been employed to improve the stability, water solubility, and optical performance of photoswitchable molecules. Alternatively, intermolecular assembly via hydrophilic-hydrophobic interactions, electrostatic forces, host-guest complexes, or π–π stacking provides a complementary method to address these challenges.In this section, the challenges are categorized, and an exploration of how molecular design and assembly strategies can address them to optimize the properties of PICAs for bioimaging applications is provided (Fig. 8).
 | ||
| Fig. 8 Schematic illustration of optimization strategies on PICAs aimed at improving stability, water solubility, Vis/NIR excitation, emission properties, and photoisomerization efficiency. | ||
3.1. Strengthening the stability
The stability of PICAs, encompassing chemical stability, thermal stability, photostability, and fatigue resistance, is essential for their application in bioimaging. However, these properties are often compromised under harsh external conditions. First, the chemical stability of PICAs is particularly challenged by complex biological milieus. For example, enzyme-mediated modification or reduction can occur in Azo molecules, which deactivates their photoswitching ability.97,98 Rapid thermal relaxation also favors the transition of PICAs back to the ground state, while photobleaching resulting from continuous irradiation hampers long-term stable imaging. Furthermore, repeated photoswitching cycles necessitates PICAs with robust fatigue resistance.Significant progress has been made in improving the chemical stability, photostability, and thermal half-life of PICAs to facilitate high-quality photoswitching imaging via structural modification.20 Chemical modification with certain groups on photoswitchable molecule itself is a viable strategy.99,100 First, the introduction of hydrogen bonds with specific functional groups (e.g. hydroxyl, carboxyl, and nitrogen heterocycle) can markedly enhance the thermal/photo-stability. For example, a series of Azo derivatives with phenyl rings substituted by pyrazole rings at different positions, significantly extends the thermal half-life from hours to years (Fig. 9A).101 These Azo derivatives containing pyrazole rings produce intramolecular C–H⋯N hydrogen bonds between the pyrazole rings further stabilizes the cis form. Second, introducing strong electron-withdrawing groups lowers the molecular oxidation potential and inhibits the photo-oxidation reaction, leading to the improved chemical stability of photoswitching molecules. Taking DAE derivatives as an example, the use of perfluorocyclopentenes as a vinyl bridge moiety with strong electron-withdrawing and steric characteristics markedly enhances their thermal stability and photobleaching resistance. While non-fluorinated DAEs only have 20–30 photoswitching cycles, fluorinated DAEs possess high fatigue resistance with hundreds to thousands of photoswitching cycles.102 In addition, perfluorocyclopentene-contained DAEs exhibit thermal half-lives of their ring-closed isomers reaching several years.103
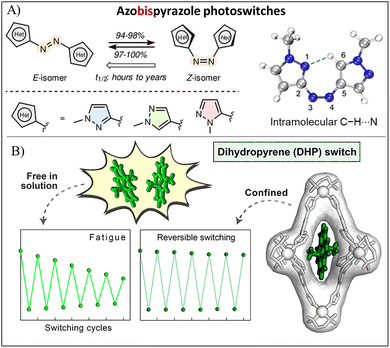 | ||
| Fig. 9 (A) Photoswitchable isomerization of dipyrazole-substituted Azo derivatives with an intramolecular C–H⋯N hydrogen bond between the two pyrazole rings. Reproduced with permission.101 Copyright 2021, Wiley-VCH. (B) Enhanced fatigue resistance of DHP confined within a PdII6L4 coordination cage. Reproduced with permission.19 Copyright 2020, American Chemical Society. | ||
Supramolecular assembly between photoswitchable molecules and biomaterials or synthetic materials offers a straightforward approach to improve their chemical stability and fatigue resistance while preserving their intrinsic photophysical properties. For example, peptide/protein-based assemblies can provide a confined environment that protects Azo compounds from enzymatic damage and inhibit cis–trans thermal relaxation, thereby enhancing fluorescence emission.37,104 Assembly strategies have been demonstrated to confine photoswitchable molecules within a limited space, which prevents molecular aggregation and improves fatigue resistance.22 Furthermore, molecular confinement successfully avoids the quenching of photoswitchable molecules. For example, DHP was encapsulated within a PdII6L4 coordination cage, significantly improving its fatigue resistance (Fig. 9B).19 When exposed to Vis light irradiation, DHP underwent isomerization to form the CPD isomer, which was stabilized in a conformation similar to its initial state due to the molecular confinement effect. After UV light irradiation or thermal relaxation in the dark, the CPD isomer converted back to DHP, suggesting reversible photoswitching within the coordination cage. Following encapsulation in the cage, DHP maintained a rapid photoswitching response after 10 switching cycles, with its attenuation rate decreasing from 28% to 8%, indicating enhanced fatigue resistance.
3.2. Improving the water solubility
The inherent hydrophobicity of many photoswitchable molecules restricts their applications in imaging within cells and organisms. Hydrophobicity-induced molecular aggregation can impair the photoswitching properties and increase potential safety risks. A common countermeasure is to functionalize photoswitchable molecules with hydrophilic groups, such as sulfonate groups, carboxylic acid groups, quaternary amine groups, polyethylene glycol, and water-soluble cavitands.4,10,20,105–111 In one study, DAE was modified with a short tetra(ethylene glycol) methyl ether (mTEG) group.10 The incorporation of the neutral mTEG group significantly improved the water solubility of the PICAs, facilitating cellular uptake and biosafety. Additionally, this modification increased the on-off contrast in FI under reversible photoswitching conditions. The introduction of quaternary ammonium salts onto the DAE structure also increases its hydrophilicity.4,112–114 Naren et al. synthesized a water-soluble DAE derivative with two quaternary amine substituents for cellular FI (Fig. 10A).4 Separately, Hell et al. engineered two novel hydrophilic PICAs by functionalizing DAE derivatives with either four or eight carboxylic acid groups (Fig. 10B).20 Both PICAs exhibited high aqueous solubility, robust photostability, and reversible photoswitching behavior, rendering them effective for labeling cellular structures. While hydrophilic modification can reduce the hydrophobicity of photoswitchable molecules, entirely inhibiting molecular aggregation in aqueous media remains difficult. To overcome this limitation, Kim et al. designed a water-soluble DAE-based dendritic nanocluster covalently linked to Cy5.5.10 The dendritic architecture effectively mitigated molecular aggregation while showing reversibly photoswitchable NIR fluorescence (Fig. 10C). Using confocal microscopy, NIR fluorescence signal was detected in living HeLa cells and zebrafish following the internalization of dendritic nanocluster. Irradiation with 365 nm UV light reverted DAE to the ring-closed state (absorption band within 450–750 nm), leading to efficient FRET quenching of the Cy5.5 NIR fluorescence at 700 nm (excitation at 630 nm). Subsequent exposure to 532 nm visible light restored the NIR fluorescent signal to its initial level. | ||
| Fig. 10 (A) Structure of a DAE derivative functionalized with two quaternary amine substituents. Reproduced with permission.4 Copyright 2008, American Chemical Society. (B) Structure and photoisomerization of a hydrophilic DAE derivative engineered with eight carboxylic acid groups. Reproduced with permission.20 Copyright 2016, Wiley-VCH. (C) Structure of a water-soluble DAE-based dendritic nanocluster. Reproduced with permission.10 Copyright 2022, Wiley-VCH. (D) Assembly of hydrophilic DAE-based nanoparticles that exhibit photoswitchable fluorescence emission and photoacoustic signal. Reproduced with permission.63 Copyright 2018, Springer Nature. (E) Enhanced solubility of hydrophobic SP (6) via host-guest assembly in a flexible coordination cage (4). Reproduced with permission.121 Copyright 2018, Springer Nature. | ||
However, even with hydrophilic functionalization, some photoswitchable molecules still show poor aqueous solubility, thereby impeding their applications in bioimaging. This limitation can be addressed by assembling photoswitches with amphiphilic molecules through non-covalent interactions.12,63 Qi et al. used a nanoprecipitation method to develop a photoswitchable nanomicelle based on amphiphilic lipid-PEG2000 (Fig. 10D).63 During assembly process, DAE was loaded into the lipid layer through hydrophobic interactions to form the core, while the hydrophilic PEG in the outer layer supported colloidal stability. This photoswitchable nanomicelle allowed dual-mode FI and PAI to guide tumor resection in mice. In addition to lipid-based systems, amphiphilic macrocycles have been used to assemble hydrophobic photoswitchable molecules.115–117 Notably, supramolecular host–guest assembly with amphiphilic macrocyclic compounds (having hydrophilic surface and hydrophobic cavity), including cucurbit[7]uril, sulfonatocalix[4]arene, and cyclodextrin, or with large-cavity coordination cages, can markedly improve the hydrophilicity of photoswitchable molecules.118–120 For instance, hydrophobic SP 6, which is poorly soluble in aqueous media, became highly water-soluble upon forming the host-guest complex 6⊂4 with flexible coordination cage 4 (Fig. 10E).121
3.3. Extending the activation wavelength to the visible/near-infrared region
Currently, most photoswitchable molecules require UV light irradiation for photoisomerization. This short-wavelength-dependent activation imposes some critical challenges for bioimaging: (1) limited tissue penetration – UV light is extensively absorbed by biological tissues as a result of the high extinction coefficient of biomolecules (such as nucleic acids and proteins) within the UV spectrum region. Therefore, activation is confined to superficial cells or tissues, accompanied by rapid energy dissipation over micrometer-scale depth. (2) Phototoxicity – long-term UV light exposure induces oxidative stress via reactive oxygen species (ROS) production, along with direct DNA damage (e.g., thymine dimer formation). These side effects compromise cell viability during live imaging, generally manifesting as reduced proliferation or apoptosis of target cells. (3) High autofluorescence background – UV excitation triggers intrinsic autofluorescence from endogenous components, such as NADH (450–500 nm) and collagen (305–450 nm). This interference degrades the signal-to-noise ratio (SNR), necessitating higher excitation intensities or complex spectral filtering to distinguish target signals. Therefore, it is crucial to develop photoswitchable molecules responsive to Vis/NIR light for in vivo bioimaging. Current approaches to red-shift the activation wavelength of PICAs can be categorized into four main aspects: chemical modification of electron-donating or electron-accepting groups; design of negatively photoswitchable molecules; integration with upconversion nanoparticles (UCNPs); and utilization of two-photon excitation technology.122–125Constructing a donor–acceptor (D–A) structure by chemical modification of electron-donating or electron-accepting groups on the skeleton of photoswitchable molecules offers a promising approach to red-shift the absorption wavelength.99,126–129 Electron-donating moieties (such as aromatic groups,101 heterocyclic groups,130,131 and ammonium ions21,132) can extend the π-conjugation system of photoswitchable molecules and lower the HOMO–LUMO energy gap. Woolley et al. is dedicated to modifying Azo derivatives with electron-donating moieties to red-shift absorption wavelength.133–135 After substitution with ortho or meta-methoxy groups, p-amino-substituted Azo derivatives were converted to azonium ions, resulting in a pronounced red shift in absorption toward the NIR and far-red spectral regions (>700 nm) for trans–cis photoisomerization (Fig. 11A).21 Moreover, the formation of intramolecular H-bond between the ortho-methoxy groups and the azonium protons further stabilized the azonium ions. Upon red light irradiation, trans–cis photoisomerization disrupted intramolecular H-bonds, which generated neutral cis isomer and slowed thermal relaxation. This azonium ion-based PICA provides a promising alternative for NIR-excited photoswitchable imaging in vivo.
 | ||
| Fig. 11 (A) Structure of Azo derivatives (azonium ions) bearing ortho or meta-methoxy and para-amino substituents on the phenyl rings. Reproduced with permission.21 Copyright 2015, Royal Society of Chemistry. (B) Construction of a DASA-conjugated polymer and its photoswitching behavior upon irradiation (λ = 550 nm). Reproduced with permission.140 Copyright 2018, Wiley-VCH. (C) Schematic illustration of NIR laser-regulated assembly and disassembly of a NIR-II Azo-based nanoprobe. Reproduced with permission.149 Copyright 2018, Wiley-VCH. (D) Two-photon-induced SP → MC photoswitching behavior and fluorescence emission. Reproduced with permission.153 Copyright 2011, American Chemical Society. | ||
Negatively photoswitchable molecules differ fundamentally from conventional photoswitches: they undergo photoisomerization upon Vis/NIR light irradiation and return to the ground state through thermal relaxation in the dark or upon short-wavelength light irradiation.45,136,137 DASA is a prominent class of such negative photochromic molecules.138,139 The initial DASA adopts an elongated triene structure with strong color and hydrophobicity, while Vis light (550 nm) irradiation converts it back to a colorless, hydrophilic, cyclized cyclopentenone isomer. Based on the characteristics of DASA, Senthilkumar et al. developed a negative photoswitchable polymer nanoparticle as a delivery vehicle by conjugating DASA to the terminal ends of a poly(p-phenylene vinylene) structure (Fig. 11B).140 Upon Vis light irradiation, DASA converted to a ring-closed cyclopentenone isomer, inducing the disassembly of the nanoparticles and facilitating drug release. Structural modification of traditional photoswitchable molecules represents a general approach for generating negatively photoswitchable variants.141,142 Irie et al. introduced an oxidized thiophene ring at the 2-position of the thiophene ring in DAE to extend the π-conjugation length of the ring-opened DAE.143 The resultant negatively photoswitchable DAE derivative underwent photocyclization upon irradiation with Vis light (λ > 430 nm) and then a ring-opening reaction under UV light irradiation, with a red-shifted absorption profile.
Another strategy is integrating UCNPs capable of converting Vis/NIR light into shorter-wavelength emission, which spectrally overlaps with the absorption required for photoswitchable molecules.144 UCNPs are primarily composed of inorganic materials represented by rare-earth fluorides and oxides. Characterized by a large Stokes shift, UCNPs absorb NIR light (typically at 808 nm and 980 nm) and upconvert this energy into UV or Vis emission.145–148 To broaden the window of PICAs for in vivo bioimaging, it is desirable to further red-shift the excitation wavelength into the second NIR region (NIR-II, 1000–1700 nm), because NIR-II light provides deeper tissue penetration capacity and reduced background interference. Zhang et al. designed a novel photoswitchable UCPN (ps-UCNP) system for NIR-II-excited photoisomerization of Azo (Fig. 11C).149 Specifically, trans-Azo-modified UCNPs were assembled with cyclodextrin-modified nanoparticles through host–guest interactions. Upon 980 nm light irradiation, the UCNPs emitted UV light to drive the isomerization of the trans-Azo to its cis-isomer, leading to nanoparticle disassembly.
In addition to chemical modification or structural adjustment that red-shifts the absorption of photoswitchable molecules into the Vis/NIR region, two-photo excitation offers an efficient route to operate photoswitching with NIR light.150 Two-photon excitation is a nonlinear process in which molecules simultaneously absorb two photons to reach an excited state with higher energy.151 Generally, two-photon excitation occurs in the NIR range, while emission falls in the Vis range.152 There is growing interest in developing PICAs that can be activated using two-photon Vis/NIR light irradiation.153–155 Zhu et al. proposed that two-photon excitation using NIR light facilitated the photoisomerization of the SP to the MC isomer, accompanied by red fluorescence emission, comparable to that achieved through single-photon excitation with UV light (Fig. 11D).153 In another study, Azo was covalently attached to a two-photon-absorbing unit, specifically a triarylamine antenna, which enabled NIR light-driven (750 nm) isomerization from the cis isomer to the trans form.156
3.4. Enhancing the emission performance
High emission intensity at long-wavelength region of PICAs is crucial for effective bioimaging. PICAs can benefit from flexible molecular design strategies based on structure–property relationships or host-gust assembly.Introducing electron-donating groups to extend the π-conjugation system has been shown to red-shift the absorption, while enhancing emission intensity or broadening emission wavelength range. Irie et al. synthesized a series of DAE derivatives and found that, the ring-closed derivatives with phenyl or thiophene substituents displayed high fluorescence emission (Fig. 12A).157 These substituents function as strong electron donor, while the central benzobis(thiadiazole) bridge acts as efficient electron acceptor, resulting in the formation of a D–π–A structure along with an intramolecular charge transfer (ICT) process. During ICT process, electron transfers from the donor (D) to the acceptor (A) to activate a polarized excited state and suppress non-radiative decay. The nonradiative decay rate constant (knr) of substituted DAE in the ring-closed state dramatically reduced from 6.5 × 108 s−1 for the non-substituted structure to 4.3 × 107 s−1. Eventually, the substitution strategy significantly improved the fluorescence quantum yield (ΦF) of ring-closed DAE from ∼0.2 to ∼0.9. In addition to improved fluorescence quantum yield, extending the π-conjugation system has been demonstrated to endow photoswitchable molecules with dual-color FI capability.158
 | ||
| Fig. 12 (A) Structures of DAE derivatives functionalized with various aromatic groups. Reproduced with permission.157 Copyright 2011, American Chemical Society. (B) Schematic illustration of a sterically hindered DAE derivative that exhibits AIE behavior by preventing π–π stacking in the aggregated state. Reproduced with permission.163 Copyright 2020, Wiley-VCH. (C) Enhanced emission of DAE@CB7 through host-guest assembly strategy. Reproduced with permission.22 Copyright 2022, American Chemical Society. | ||
Most small-molecule photoswitches are hydrophobic, thereby tending to aggregate in aqueous media to cause ACQ that diminishes both photoswitching and emission capacities. Introducing hydrophilic groups20 and regulating steric hindrance (e.g., rigid planar structures and AIE motifs) are effective approaches to prevent aggregation.159–162 Zhu et al. synthesized a novel DAE derivative through functionalizing it with two bulky benzothiophene groups, which showed characteristic AIE behavior, including restriction of intramolecular vibration (RIV) and rotation (RIR), as well as intermolecular stacking (Fig. 12B).163 The steric effect generated a rigid aggregation structure, decreasing the nonradiative decay rate to enhance the fluorescence emission of ring-opened DAE. Upon UV light irradiation, the AIE-type DAE underwent a cyclization reaction to cause fluorescent quenching owing to reduced steric hindrance and increased π–π stacking, which eventually resulted in negligible emission intensity caused by ACQ. Therefore, modulating the nonradiative decay rate and aggregation state of PICAs provides a promising tool for improving the fluorescence emission performance.
Host–guest supramolecular assembly is capable to modulate the spatial confinement, environmental polarity, energy transfer, and dynamic reversibility of photoswitchable molecules, leading to an enhanced emission performance.22,164,165 For example, encapsulation by CB7 reduced the interaction between DAE and water molecules, inhibited non-radiative transitions, increased the ΦF from 0.40 to 0.63, and prolonged the fluorescence lifetime (Fig. 12C).22 The nonradiative decay rate constants of DAE and DAE@CB7 were calculated to be knr = 3.3 × 108 s−1 and 1.6 × 108 s−1, respectively, indicating the enhanced emission property.
In photoswitching imaging, FRET not only turns the on/off state of fluorescence,61 but also manipulates fluorescence properties through two distinct interaction pathways between the photoswitchable molecule and the fluorophore: (1) enhancing the emission intensity of photoswitchable molecule via energy transformation from fluorophore; (2) red-shifting the emission wavelength of photoswitchable molecule by activating the fluorescence of fluorophore (Fig. 13). FRET is a non-radiative energy transfer process that needs spatial coupling between the donor and acceptor.166,167 As shown in Fig. 13A, FRET proceeds from fluorophore (as a donor, D) to photoswitchable molecule (photoswitch, as an acceptor, A) when the emission spectra of the fluorophore overlaps with the absorption spectra of the isomerized photoswitch at off-state.168 In this way, the fluorescence of donor is quenched while the emission of photoswitch is activated, generating dual color fluorescence signals. On the contrary, when the emission spectra of the photoswitch at on-state overlaps with the absorption spectra of fluorophore, FRET occurs from the photoswitch (now as donor) to the fluorophore (now as acceptor) (Fig. 13B). The second FRET process enables fluorophore activation through energy transfer from the photoswitch, thereby both enhancing the overall fluorescence intensity169 and red-shifting the emission from the Vis region to the NIR region15 for improved bioimaging.
3.5. Improving the photoisomerization efficiency
Photoisomerization is a defining characteristic of PICAs distinctive from conventional contrast agents. Photoisomerization efficiency serves as a critical criterion for assessing the potential of PICAs in photoswitchable bioimaging. Photoswitchable molecules proceed a reversible isomerization process wherein they populate an excited singlet state upon light irradiation and subsequently return to the ground state. However, during this sequence, competing pathways, such as non-radiative decay and intermolecular energy/charge transfer, inevitably dampen photoisomerization efficiency. In addition, poor thermal stability of the two ground-state isomers further attenuates the apparent quantum yield. Molecular design optimization and supramolecular assembly have been shown to overcome these limitations, thereby strengthening the practical viability of photoswitching molecules as PICAs for bioimaging.A promising method is introducing electron-donating groups (such as amino and methoxy) or electron-accepting groups (such as nitro and cyano), to promote the D–A character of photoswitchable molecules and reduce excitation energy required for photoisomerization. Conventional Azo compounds typically display insufficient trans → cis photoisomerization efficiency, with a low quantum yield of approximately 10–30%.31 Modifying a pair of D–A substituents (such as methoxy/cyano and amino/nitro) on both sides of the Azo markedly decreased the excitation energy and raised the photoisomerization quantum yield to 35% (Fig. 14A).23 In addition, extending the π-conjugated system with electron-rich groups enlarges the π-electron delocalization range, which can also reduce the excitation energy to improve the photoisomerization efficiency.170 For example, replacing one phenyl ring with thiophene generated a lone-pair (S atom)⋯π (Azo) interaction between the S atom and the Azo system, strengthening ICT from the thiophene donor to the phenyl acceptor. This change raised the quantum yield to 40% for trans → cis and 65% for cis → trans photoisomerization.171
 | ||
| Fig. 14 (A) Excitation energy of Azo derivatives functionalized with various electron donor groups. Reproduced with permission.23 Copyright 2020, American Chemical Society. (B) Photoswitching behavior of anti-parallel DAE functionalized with two benzothiophene substituents. Reproduced with permission.163 Copyright 2020, Wiley-VCH. (C) Stable host-guest interaction between CD and SP/MC. Reproduced with permission.176 Copyright 2015, American Chemical Society. | ||
Incorporating steric groups into the skeleton of photoswitchable molecules possess a twist conformation and increases steric hindrance, thereby enhancing photoisomerization efficiency.172 For instance, DAE generally has a photoisomerization (ring-opened → ring-closed) quantum yield below 50%, owing to the coexistence of parallel (photoinactive) and antiparallel (photoactive) conformations. After introducing large steric-hindrance groups (e.g., benzothiophene) on both sides of the DAE skeleton, the obtained twisted structure constrained non-radiative pathways such as intramolecular rotation and vibration, which in turn compelled the molecule to preferentially adopt the anti-parallel conformation (Fig. 14B).163 This symmetric benzothiophene-based DAE promoted cyclization reaction, raising the photoisomerization quantum yield to 88% for ring-closed state and 90% for ring-opened state.
Supramolecular assembly strategy via non-covalent interactions (including hydrophobicity, hydrogen bonding, π–π stacking, and host–guest recognition) between photoswitchable molecules and large-cavity host molecules, tailors the aggregation state of photoswitchable molecules, thus leading to enhanced photoisomerization efficiency.173–175 For example, hydrophobic SP could insert into the cavity of cyclodextrin (CD) by host–guest interaction, resulting in a stable assembly complex (Fig. 14C).176 Due to the stabilization of MC state by CD, the half-life of MC state prolonged and the emission intensity was enhanced.
4. PICAs for bioimaging systems
Bioimaging technologies are crucial for the early diagnosis and treatment of diseases because they enable sensitive and precise detection of changes in biological and pathological signals.177–179 Bioimaging also contribute fundamentally to understanding the structure of living organisms and elucidating their underlying physiological functions.180 However, achieving high accuracy and resolution in biological imaging remains challenging owing to the complexity of environment. PICAs, with their dynamic signal modulation and controllable performance, can reduce background interference and improve the quality of photo-controllable bioimaging. Recent advances in fluorescence microscopy have driven the widespread use of PICAs in fluorescence sensing and imaging, particularly in super-resolution imaging.81,163,181 Additionally, PICAs have expanded into other imaging modalities, such as PI,164 PAI,63 MRI,182 and RI (Table 1).14| Photoswitchable imaging modality | Photoswitching moiety | Photoswitching light (nm) | Half-life of isomer (t1/2) | Photoswitching speed | Photoswitching quantum yield | Photoswitching ratio | Photoswitching cycle | Imaging object | Ref. | |
|---|---|---|---|---|---|---|---|---|---|---|
| Molecule type | Original name | |||||||||
| FI | RSFP | GMars-Q | 405/488 | 120 min of off-state | 2 ms for on-switching/24 ms for off-switching | — | — | >50 | Various cell structures and organelles with fwhm of ∼80 nm | 197 |
| RSFP | rsFusionRed variants | 510/590 | >20–40 min | — | — | — | >1000 | F-Tractin of U2OS cells with fwhm of <80 nm | 90 | |
| SP | DSA-2SP | 365 or 302/561 | >24 h | — | — | — | >10 | Cylindrical micelles formed with fwhm of ∼50 nm | 195 | |
| DAE | PMI-3DTE | 302/532 | — | 2–5 s for ring-closing/2 s for ring-opening | 0.459 of closed-state/0.007 of open-state | 3026 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
Polymer film with fwhm of 36 nm | 5 | |
| DAE | DAE@CB7 | 355/485 | — | — | 0.35 of closed-state/9.2 × 10−3 of open-state | — | 2560 | Microtubules of Cos7 cells with fwhm of 70–90 nm | 22 | |
| SP | SP-PCL | <420/>420 | — | 1 s for ring opening | — | — | >6 | HeLa cells with sub-50 nm resolution | 198 | |
| SP | SP-CE | 300–400/Vis | — | — | — | — | — | Li+ detection in HeLa cells and zebrafish | 106 | |
| SP | Ly-NT-SP | UV/Vis | — | — | — | — | >3 | SO2 detection in lysosome during heat shock | 210 | |
| SP | NpG@HSA | 365/550 | — | — | — | — | >5 | Endogenous β-Gal detection in SKOV-3 cells and WI-38 cells with fwhm of <80 nm | 16 | |
| SP | SPMA | 302/525 | — | — | — | — | >5 | A549 cells | 224 | |
| DAE | DBTEO | 365/>420 | — | — | — | — | >4 | RAW264.7 cells with dual-color fluorescence and fwhm of 64 nm | 12 | |
| SP | BOSA-SP | 365/524 | — | — | — | — | >5 | HeLa cells with three-color fluorescence and fwhm of 31 nm | 225 | |
| PI | DAE | DTE-C3⊂CB[8]@Cy5 | 365/>600 | — | — | — | — | >10 | A549 cells | 15 |
| DAE | DTE-TP/CB[8] | 365/>600 | — | — | — | — | >6 | A549 cells | 164 | |
| PAI | RSFP | RpBphP1 | 730–790/630–690 | ∼210 s | 1.56 s−1 | — | ∼4.3 | >10 | U87 tumor in a mouse kidney or liver with 34 times contrast enhancement | 92 |
| RSFP | DrBphP-PCM | 630–690/730–790 | — | 0.54 s−1 | — | — | >5 | Brain, kidney, and liver tumors in mice | 93 | |
| RSFP | BphP1 | 620–680/740–800 | — | — | — | — | >7 | 4T1 tumor in mice | 255 | |
| DAE | DTE-TPECM | 365/610 | — | — | — | — | >10 | 4T1 tumor in mice | 63 | |
| Azo | wsT-AzHCy | — | <10 ps | — | — | — | — | Liver, kidney and spleen in mice | 17 | |
| Azo | UCNPs@PAA-Azo | 365/450 | — | — | — | — | — | Nude mouse | 260 | |
| MRI | SP | Gd-SPDO3A | 365/Vis | — | — | — | — | — | — | 269 |
| SP | spiropyran-DO3A-Gd | 532/— | — | 200–300 ms for ring-closing | — | — | >13 | — | 271 | |
| Azo | Azopyridine functionalized Ni-porphyrin | 500/435 | — | — | — | — | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
— | 18 | |
| Azo | Bacteriochlorin 11 | 720/505 | — | — | — | — | >100 | — | 277 | |
| Azo | Azoimidazole functionalized Ni-II porphyrins | 365/— | — | — | — | — | — | Phantom | 276 | |
| Azo | KMR-Az | 365/— | — | — | — | — | — | — | 278 | |
| SP | CLADIO-NH-SP | UV/Vis | — | — | — | — | — | — | 279 | |
| RI | DAE | DAE620 | 375/640 | — | 515 ns for ring-closing, 437 ns for ring-opening | — | >25 | >8 | HeLa cells | 285 |
| DAE | DTE-alkyne derivatives | 360/633 | — | — | — | >100 | HeLa cells | 14 | ||
| DAE | Asymmetric DAE polymers | 365(405)/640 | — | <200 µs for ring-closing | — | — | >100 | U2OS cells | 286 | |
4.1. Fluorescence imaging
FI is the most widely used imaging modality owing to its noninvasive nature, high spatiotemporal resolution, operational simplicity, and capacity for real-time monitoring. FI has demonstrated a remarkable success in diverse domains, including the elucidation of cellular pathophysiological processes, early-stage disease diagnosis, and real-time intraoperative imaging guidance.183–185 Compared to conventional fluorescent probes, PICAs offer several key advantages: (1) reversible on-off switching enables enhanced imaging resolution by breaking the optical diffraction limit; (2) multiple switching cycles statistically eliminate random noise for improved SNR; (3) selective illumination of regions of interest minimizes nonspecific background signals; and (4) on-demand light activation mitigates photobleaching and phototoxicity, permitting long-term and dynamic tracking. Consequently, PICAs have been employed as effective alternatives to conventional fluorescent probes in super-resolution imaging, biosensing, dual/multiple-color imaging, and multiplexed imaging.RSFPs are suitable for RESOLFT-based dynamic imaging owing to their superior live-cell compatibility and switching cyclability, as well as lower light dose requirement for saturation transition. Testa's group developed a multi-sheet RESOLFT microscope to overcome the limitations of conventional light-sheet fluorescence microscopy (LSFM) in subcellular imaging.196 The technology employes wide-field illumination to activate all RSFPs, followed by sinusoidal interference-patterned depletion light pulse to confine on-state RSFPs to a tightly defined sheet. This allows emission sheets with a thickness far below the diffraction limit (<100 nm)—10 times thinner than conventional excitation sheets. By parallelizing the imprinting of multiple sheets and rapid readout, the technique achieved fast volumetric imaging at 1–2 Hz with a spatial resolution below 250 nm. It required only 10–20 switching cycles and is compatible with a wide range of RSFPs. Using a classical RSFP, rsEGFP (N205S), multi-sheet RESOLFT succeeded in high-resolution live-cell 3D tracking of subcellular dynamics, including actin dynamics, cell division, and virus-like particle movement. Future research can focus on incorporating other modulation patterns and image rotators to further improve spatial resolution and expanding its applications to 3D culture and multicellular organisms.
The performance of RESOLFT nanoscopy relies on multiple properties of RSFPs, including switching rate, photostability, brightness, and residual fluorescence in their off state. It is challenging to achieve optimal performance due to inherent trade-offs between these properties. For instance, enhanced brightness often compromises switching rate, and a lower off-switching rate increases the signal-to-background ratio (SBR). To reach a balance, Sun et al. developed a novel monomeric RSFP, namely GMars-Q, for long-term RESOLFT imaging in living cells (Fig. 15A).197 GMars-Q was activated by 405 nm light and switched off by 488 nm light, respectively. Significantly, the off-state residual fluorescence of GMars-Q was 6–9-fold lower than that of rsEGFP2 and N205S, two classical reversibly photochromic fluorescent proteins, enabling significantly reduced SNR and improved imaging contrast. Moreover, GMars-Q exhibited higher fatigue resistance and SBR compared with N205S. These improvements may stem from an unusual biphasic photobleaching feature. When tagged with the KDEL targeting sequence, GMars-Q achieved stable live-cell observation of endoplasmic reticulum tubulation for 2 minutes, which was an order of magnitude longer than that achieved by rsEGFP (N205S) at equivalent resolution. This study provides a methodological advancement for long-term super-resolution imaging in living cells. Nevertheless, green-emitting RSFPs generally need violet-blue light for photoswitching, which potentially causes scattering effect and phototoxicity. Red-shifted RSFPs with strong fatigue resistance and rapid switching rate are needed. Testa et al. engineered three green photo-activated red-emitting rsFusionRed variants (rsFusionRed1, rsFusionRed2, and rsFusionRed3) via site-specific amino acid mutagenesis.90 Compared to switching under short light irradiation (405 or 488 nm), all variants could be efficiently switched on through 510 nm green light irradiation, exhibiting slightly slower on-switching kinetics but lower fatigue. Upon 590 nm orange light irradiation, rsFusionRed3 showed substantially faster off-switching kinetics—30-fold faster than rsTagRFP (487 ± 2 ms), and 6-fold faster than rsCherryRev1.4. Importantly, the improved off-switching kinetics of rsFusionRed3 did not compromise their fatigue resistance (>1000 consecutive switching cycles) and thermostability (half-life, >20–40 min). Using RESOLFT nanoscopy, rsFusionRed3 tagged with either F-Tractin or vimentin enabled live-cell visualization of actin networks or vimentin filaments with full width at half-maximum (fwhm) of ∼80 nm under low intensity irradiation (∼1 kW cm−2). This demonstrates the excellent potential of rsFusionRed variants in ref-shifted non-invasive imaging in vivo by RESOLFT nanoscopy.
 | ||
| Fig. 15 (A) Reversible photoswitching behavior and super-resolution image of GMars-Q. Reproduced with permission.197 Copyright 2016, American Chemical Society. (B) Intramolecular energy transfer in DSA-2MC and super-resolution imaging via photoswitching between DSA-2MC and DSA-2SP. Reproduced with permission.195 Copyright 2017, American Chemical Society. (C) Photoswitching mechanism and super-resolution fluorescence images of PMI-3DTE. Reproduced with permission.5 Copyright 2014, Springer Nature. | ||
In addition to RSFPs, small-molecule photoswitches also display superior capacity for super-resolution imaging owing to their chemical tunability and photophysical diversity, which can be tailored through precise synthesis and modification based on structure–property relationships. However, as mentioned above, most small-molecule photoswitches tend to aggregate in aqueous solutions as a result of molecular stacking and hydrophobicity, causing ACQ effect to diminish their photoswitching behavior. To address this issue, Qi et al. coupled two SP units with an AIE moiety, 9,10-distyrylanthrance (DSA), to develop a photoswitchable SP derivative (DSA-2SP).195 To improve its water solubility, DSA-2SP was loaded into a nanomicelle assembled from amphiphilic polymer, polystryrene-block-poly(ethylene oxide). The nonplanar molecular structure of DSA offers large free space to prevent molecular stacking, which together with intramolecular hydrogen bonds, support reversible photoisomerization between DSA-2SP and DSA-2MC upon altering irradiation with 302 nm UV light and 561 nm Vis light. Notably, intramolecular energy transfer from the DSA donor to the MC acceptor resulted in a bright red fluorescence at 670 nm with a fwhm of ∼49.5 nm for efficient photoswitchable super-resolution imaging (Fig. 15B). Zhu et al. coupled a single perylenemonoimide (PMI) molecule with three DAE units via covalent oxygen bridge to develop a trident photoswitchable probe (PMI-3DTE). (Fig. 15C).5 In this design, PMI offers high fluorescence quantum yield and photostability, while DAE functions as an efficient switch with excellent photoisomerization behavior. The three-dimensional structure of this trident photoswitchable probe efficiently prevents the ACQ effect. Upon 2–5 s of 302 nm UV light irradiation, >95% of PMI-3DTE isomerized to its ring-closed state. This isomerization, combined with energy transfer from PMI to the ring-closed DAE, triggered rapid fluorescence quenching, with an efficiency close to 100% and a photoswitchable fluorescence quenching ratio in the photostationary state exceeding 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. In addition, under alternating irradiation with UV light and Vis light, the probe showed superior fatigue resistance over 10
000. In addition, under alternating irradiation with UV light and Vis light, the probe showed superior fatigue resistance over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 switching cycles, holding great potential for super-resolution bioimaging.
000 switching cycles, holding great potential for super-resolution bioimaging.
Assembly strategies have been demonstrated to improve the performance of small-molecule photoswitches for super-resolution imaging, such as photostability, water solubility, and photoisomerization capacity. Hell et al. developed a supramolecular assembly probe (DAE@CB7), in which DAE was encapsulated in cucurbit[7]uril (CB7) via the host–guest interactions, for super-resolution RESOLFT imaging.22 DAE@CB7 exhibited rapid, reversible photoisomerization upon altering irradiation of 355 nm light and 485 nm light, and increased fluorescence quantum yield of 0.63 compared to 0.40 for unimolecular DAE. Possibly ascribing to reduced aggregation in aqueous solution, DAE@CB7 demonstrated robust fatigue resistance that endured over 2560 switching cycles before half-bleaching, whereas unimolecular DAE persisted only 8 switching cycles. After conjugation with targeting antibodies, the probe succeeded in labelling cellular microtubules and photo-controllable super-resolution imaging by RESOLFT, with a 70–90 nm resolution. Dual-color fluorescence switching of PICAs offers another approach to achieve super-resolution. In addition to photoswitchable performance, the aqueous biocompatibility of assembles is crucial for their use in bioimaging. Liu et al. designed a biodegradable nanoparticle probe (SP-PCL) for super-resolution FI through the assembly of poly(ε-caprolactone) terminated with a SP unit via ring-opening polymerization.198 Upon altering irradiation of 365 nm UV light and >420 nm Vis light, the nanoparticle probe underwent reversible photoisomerization between the SP-PCL state and the MC state, thereby achieving fluorescence switching between green emission at 530 nm and red emission at 650 nm. Owing to the lipophilic nature of PCL, the probe was readily internalized into HeLa cells, resolving cellular structures with sub-50 nm resolution by FI.
The utilization of PICAs for detecting biomarkers such as metal ions, specific gases, and proteins, has gained growing interest in exploring cellular processes and disease mechanisms.202–208 SP has been widely used to detect metal ions, particularly alkali metal ions, owing to the chelating ability of its colored MC isomer. In one study, SP was functionalized with aza-12-crown-4 to constructed a red-emitting probe (SP-CE) for Li+ detection (Fig. 16A).106 Once encountering Li+, the crown ether of SP-CE rapidly chelated with it, resulting in the photoisomerization of SP to the colored MC state without the requirement for light irradiation. The resultant phenolate ion in MC also chelated with Li+ to further stabilize the complex, which significantly enhanced the red fluorescence emission to achieve accurate Li+ sensing (Fig. 16B). Following exposure to Vis light, MC reverted to SP and released Li+. This photoswitchable sensor subsequently succeeded in Li+ imaging in living HeLa cells and zebrafish (Fig. 16C), indicating the potential of SP for metal ion biosensing in vivo.
 | ||
| Fig. 16 (A) The isomerization of SP-CE is triggered by Li+ binding and is reversed by Vis light irradiation. Reproduced with permission.106 Copyright 2021, Elsevier. (B) Absorption spectra and fluorescence intensity of SP-CE in Li+ solution. Reproduced with permission.106 Copyright 2021, Elsevier. (C) Red fluorescence images of SP-CE in zebrafish. Reproduced with permission.106 Copyright 2021, Elsevier. (D) A lysosome-targeted, SP-derived PICA (Ly-NT-SP) for SO2 sensing. Reproduced with permission.210 Copyright 2020, American Chemical Society. (E) A SP-based PICA (NpG@HSA) enabling super-resolution detection of β-Gal in cells. Reproduced with permission.16 Copyright 2020, American Chemical Society. | ||
Beyond chelating with metal ions, the MC isomer contains a polarized unsaturated C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond that serves as a nucleophilic sites, enabling Michael addition reactions with nucleophiles such as sulfur dioxide (SO2), hydrogen sulfide (H2S), and sulfite ion (SO32−).209–212 Yin et al. synthesized a SP-based fluorescent probe, Ly-NT-SP, for detecting SO2 in lysosomes (Fig. 16D).210 Ly-NT-SP consisted of a lysosome-targeted group, a fluorophore (naphthalimide with green fluorescence), and a SP moiety. After UV light irradiation, Ly-NT-SP isomerized to the Ly-NT-MC with a weak red-shifted emission at 630 nm. The C
C bond that serves as a nucleophilic sites, enabling Michael addition reactions with nucleophiles such as sulfur dioxide (SO2), hydrogen sulfide (H2S), and sulfite ion (SO32−).209–212 Yin et al. synthesized a SP-based fluorescent probe, Ly-NT-SP, for detecting SO2 in lysosomes (Fig. 16D).210 Ly-NT-SP consisted of a lysosome-targeted group, a fluorophore (naphthalimide with green fluorescence), and a SP moiety. After UV light irradiation, Ly-NT-SP isomerized to the Ly-NT-MC with a weak red-shifted emission at 630 nm. The C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C–C
C–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N+ fragment of MC could react with SO2 to generate a adduct that emitted strong green fluorescence at 535 nm. Using this sensor, the overproduced SO2 in lysosomes (induced by heat shock) was successfully visualized with a detection limit of 0.47 µM. Notably, tissue imaging showed that increased SO2 alleviated small intestine injury caused by heat shock via modulating oxidative stress. Additionally, Tian's group reported an amphiphilic photoswitchable molecule through modifying the naphthalimide-SP conjugate with a hydrophilic D-galactose (Fig. 16E).16 This molecule could assemble into a nanomicelle sensor for SO32− detection in living cells. Subsequently, the group functionalized the similar naphthalimide–MC conjugate with a β-galactosidase (β-Gal)-responsive galactose unit to quench the red fluorescence emission (620 nm) of MC. The resulting molecule was bound to human serum albumin (HSA) through host-guest interactions, for increased green fluorescence emission at 520 nm as well as better biocompatibility. In the presence of β-Gal, the shielding unit was cleaved to expose the free phenol motif, thereby restoring the red fluorescence emission and photoisomerization function of the system. The reversible photoswitchable behavior of the sensor enabled on/off photoblinking for STORM imaging, which precisely visualized the subcellular localization of β-Gal in different cell lines. Collectively, these studies demonstrate the significance of small-molecule photoswitchable biosensors for detecting biomarkers related to physiological homeostasis or pathological processes, such as metal ions, small molecules, and enzymes.213–217
N+ fragment of MC could react with SO2 to generate a adduct that emitted strong green fluorescence at 535 nm. Using this sensor, the overproduced SO2 in lysosomes (induced by heat shock) was successfully visualized with a detection limit of 0.47 µM. Notably, tissue imaging showed that increased SO2 alleviated small intestine injury caused by heat shock via modulating oxidative stress. Additionally, Tian's group reported an amphiphilic photoswitchable molecule through modifying the naphthalimide-SP conjugate with a hydrophilic D-galactose (Fig. 16E).16 This molecule could assemble into a nanomicelle sensor for SO32− detection in living cells. Subsequently, the group functionalized the similar naphthalimide–MC conjugate with a β-galactosidase (β-Gal)-responsive galactose unit to quench the red fluorescence emission (620 nm) of MC. The resulting molecule was bound to human serum albumin (HSA) through host-guest interactions, for increased green fluorescence emission at 520 nm as well as better biocompatibility. In the presence of β-Gal, the shielding unit was cleaved to expose the free phenol motif, thereby restoring the red fluorescence emission and photoisomerization function of the system. The reversible photoswitchable behavior of the sensor enabled on/off photoblinking for STORM imaging, which precisely visualized the subcellular localization of β-Gal in different cell lines. Collectively, these studies demonstrate the significance of small-molecule photoswitchable biosensors for detecting biomarkers related to physiological homeostasis or pathological processes, such as metal ions, small molecules, and enzymes.213–217
In addition to small-molecule photoswitches, RSFPs have been used as genetically encoded indicators (GEIs) for photo-controllable sensing of specific metabolites and ions. This mechanism depends on photoisomerization-induced signal changes in the readout moiety of GEIs following target binding. Stiel's team constructed a prototype photoswitchable GEI (rsGCaMP) based on a common Ca2+ indicator GCaMP5G, which enables reversible cis/trans photoisomerization in its Ca2+-bound form upon altering irradiation with 405 nm and 488 nm light.11 The key sensing function stems from interaction between the β-barrel structure of rsGCaMP and its calcium receptor moiety. rsGCaMP exhibited a fluorescent quantum yield of 9.2% and robust fatigue resistance at various pH values. It achieved super-resolution imaging of Ca2+ concentration in cultured cells and visualization of Ca2+ distribution in the endoplasmic reticulum, as well as photoacoustic Ca2+ imaging in HeLa cells implanted in mice. Notably, the universality of this photoswitchable molecular sensor concept was validated by successful construction of RSFP-based maltose sensor and dopamine sensor, offering a new approach for mapping analytes at the nanoscale and in live animal tissues. Subsequently, Dedecker's team engineered a series of GCaMP variants for cellular activity sensing, by retaining a Ca2+-sensitive moiety (CaM) domain and a CaM-binding peptide (RS20) while introducing photochromic mutations.218 GCaMP6s-Q was selected as the best variant with higher molecular brightness and Ca2+ response. Since Ca2+ concentration affects the overall fluorescence intensity and photochromic properties of the sensor, the team proposed a photochromism-enabled absolute quantification (PEAQ) method for Ca2+ detection. PEAQ correlated the fluorescence intensity changes before and after light irradiation with Ca2+ concentration and established a calibration correlation curve, enabling accurate determination of absolute Ca2+ concentration by simply measuring the fluorescence intensity. This strategy addressed the limitation of conventional photoswitchable probes that rely on complete switching cycles, thereby significantly lowering the required light dosage and shortening imaging acquisition time.
However, the key challenge is that spectral overlap between photoswitchable molecules and fluorescent dyes often induces FRET process, resulting in selective fluorescence quenching of the donor molecules. Therefore, fluorescent dyes featuring large Stokes shifts minimize spectral overlap with photoswitchable molecules by significantly separating their excitation and emission bands. A dual-color photoswitchable nanoparticle was engineered by integrating a blue-emitted (λem = 498 nm, ΦF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) =
= ![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.55) DAE (DBTEO) at closed state and a large Stokes shift dye (3-(1-phenyl-1H-phenanthro[9,10-d]imidazol-2-yl)naphthalen-2-ol, HPNIC) with absorption at
0.55) DAE (DBTEO) at closed state and a large Stokes shift dye (3-(1-phenyl-1H-phenanthro[9,10-d]imidazol-2-yl)naphthalen-2-ol, HPNIC) with absorption at![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 337 nm and emission at 590 nm (ΦF = 0.11) (Fig. 17A).12 This design achieved 100% switching on/off of blue emission from DBTEO upon alternating irradiation at 365 nm (on state) and >420 nm (off state). The persistent orange emission of HPNIC enabled long-term nanocarrier trafficking analysis in living cells, while the photoswitchable blue fluorescence of DBTEO provided spatiotemporally precise target localization via UV/vis light control (Fig. 17B).
337 nm and emission at 590 nm (ΦF = 0.11) (Fig. 17A).12 This design achieved 100% switching on/off of blue emission from DBTEO upon alternating irradiation at 365 nm (on state) and >420 nm (off state). The persistent orange emission of HPNIC enabled long-term nanocarrier trafficking analysis in living cells, while the photoswitchable blue fluorescence of DBTEO provided spatiotemporally precise target localization via UV/vis light control (Fig. 17B).
 | ||
| Fig. 17 (A) Construction of a DBTEO-based nanoparticle and its corresponding normalized absorption/emission spectra, as well as structure of DBTEO and HPNIC. Reproduced with permission.12 Copyright 2019, Springer Nature. (B) Dual-color photoswitchable FI in living cells by using F127-encapsulated nanoparticles containing DBTEO-based nanoparticles, featuring emission at 498 nm (closed DBTEO) and 590 nm (HPNIC). Reproduced with permission.12 Copyright 2019, Springer Nature. (C) Photoswitchable dual-color FI using SP-based nanoparticles in living cells. Reproduced with permission.224 Copyright 2015, American Chemical Society. (D) Schematic illustration of three-color fluorescence switching (green/red/yellow) of BOSA-SP and corresponding photoswitchable FI in HeLa cells. Reproduced with permission.225 Copyright 2022, Wiley-VCH. | ||
Conversely, an alternative strategy for dual-color PICAs exploits the FRET effect between a photoswitchable molecule and spectrally coupled fluorescent dye.44,87,158,221–223 Jiang et al. synthesized a photoswitchable amphiphilic polymer nanoparticle that contains conjugated SP and BODIPY for FRET-mediated dual-color FI with high contrast in living cells (Fig. 17C).224 Upon exposure to Vis light, BODIPY emitted green fluorescence during cell imaging, while SP remained nonfluorescent. When UV light was applied, the green fluorescence disappeared, and red fluorescence emerged owing to the FRET process from BODIPY to the MC isomer.
Compared to multi-component PICA systems, multicolor imaging that employs a single photoswitchable molecule can efficiently suppress intermolecular energy transfer (e.g., FRET) or electron transfer through confining all optical transitions within one molecule skeleton. In one study, SP was covalently with a fluorophore (distyryl anthracene, BOSA) via a flexible ester chain to develop a single-component probe for photoswitchable three-color FI (Fig. 17D).225 This probe displayed wavelength-dependent color modulation through the synergistic interaction between SP/MC photoisomerization and energy acceptor (BOSA). Upon 405 nm light irradiation, SP underwent ring-opening reaction to form MC, activating FRET effect from the BOSA to MC. This change red-shifted the emission from 520 nm (green) to 620 nm (red). Reversal to the SP state was achieved by 532 nm light irradiation, which terminated the FRET effect to recover green fluorescence. Notably, 800 nm two-photon excitation can directly trigger a yellow fluorescence (at 570 nm) of BOSA. To enhance aqueous solubility, BOSA-SP was encapsulated within amphiphilic F127 polymer-based nanoparticles to achieve efficient cellular uptake. This nanosystem programmed triple-fluorescence switching (green/yellow/red) in confocal microscopy, allowing significant suppression of cellular autofluorescence interference and high-contrast live-cell imaging with improved spatial resolution (fwhm = 31 nm).
However, realizing multiple-color fluorescence switching via a single photoswitchable molecule faces persistent challenges, such as inefficient switching, poor photostability, and low fluorescence quantum yield. Innovative strategies on molecular design hold promise to overcome these challenges, such as developing integrated chromophore systems combining complementary photoswitching mechanisms, devising stable covalent immobilization schemes, and refining energy transfer networks to suppress non-radiative decay. Progress in this field will provide improved spatiotemporal precision for multicolor super-resolution imaging, thereby achieving real-time visualization of intricate biomolecular interactions and cellular dynamics with minimal background noise.
Marriott et al. firstly employed OLID technology to isolate the characteristic fluorescence signal of photoswitchable probes (such as nitrospirobenzopyran and Dronpa) from background or overall fluorescence.230 By modulating the fluorescent and non-fluorescent states of photoswitchable probes and performing cross-correlation analysis with the switching waveform, OLID generates correlation images that can discriminate these probes from conventional fluorophores. This approach achieved high-contrast imaging, as evidenced in zebrafish neurons labeled with Dronpa, where the SBR was improved approximately 5-fold. In contrast, SAFIRe relies on the amplitude of fluorescence modulation induced by a secondary light source, which depopulates the dark state of the photoswitchable probe.233
Jullien et al. developed OPIOM, a technique using periodically modulated monochromatic illumination to drive oscillations in photoswitchable fluorophores between their two states.231 Through phase-sensitive detection and resonance matching of illumination parameters and photoswitching kinetics, OPIOM isolates the specific fluorescence signal from overall out-of-phase fluorescence. In HEK293 cells co-expressing membrane-localized EGFP and nuclear-localized Dronpa-3, OPIOM rapidly removed EGFP interference to extract the Dronpa-3 signal. Similarly, in Dronpa-3-labeled zebrafish embryos, OPIOM also filtered out yolk autofluorescence and proved compatibility with light sheet fluorescence microscopy for 3D image reconstruction of the Dronpa-3 signal. OPIOM provides an effective approach for selective imaging of photoswitchable fluorescent probes. However, a key issue is its suboptimal photoswitching rate and extent. Jullien et al. developed speed OPIOM by optimizing RSFP switching using two antiphase-modulated light sources (Fig. 18A).234 This approach doubled the signal intensity of OPIOM while reducing the imaging time to the millisecond level that is compatible with image acquisition rate of up to 50 Hz. Additionally, speed OPIOM effectively suppressed autofluorescence and scattered light, enhancing the contrast between photoswitchable fluorophores and spectrally overlapping non-photoactivatable fluorophores or ambient light by a factor of 102–103. The technique also enabled discrimination of spectrally similar RSFPs (Dronpa, Dronpa-2, Dronpa, Kohinoor, and Padron) through modulated resonant illumination. In HeLa cells, speed OPIOM achieved real-time monitoring of Kohinoor-tagged ERK2 nuclear translocation in response to epidermal growth factor stimulation. Notably, speed OPIOM clearly distinguished Dronpa-2 (nucleus), EGFP (cell membrane), Padron (mitochondria), and Dronpa (Golgi) in U2OS cells, overcoming the spectral limitations of multiplexed imaging (Fig. 18B). This technology was subsequently implemented in macroscale fluorescence and fiber-optic epifluorescence imaging.235,236 Further improvements in temporal resolution and multiplexing capabilities could be achieved by developing brighter and faster-switching fluorescent probes.
 | ||
| Fig. 18 (A) Schematic illustration of the speed OPIOM method. Reproduced with permission.234 Copyright 2017, Springer Nature. (B) Independent speed OPIOM images showing the distribution of four spectrally similar fluorescent proteins in HeLa cells. Localization: H2B-Dronpa-2 (nucleus), Mito-Padron (mitochondria), GTS-Dronpa (Golgi), and Lyn11-EGFP (cell membrane). Reproduced with permission.234 Copyright 2017, Springer Nature. (C) Schematic illustration of the NEEMO approach. Reproduced with permission.238 Copyright 2024, Elsevier. (D) NEEMO image showing the distribution of four differently targeted fluorescent proteins expressed in HeLa cells. Reproduced with permission.238 Copyright 2024, Elsevier. | ||
The aforementioned methods demand the photoswitching response rate to match the temporal resolution of instruments, which is intrinsically constrained by the camera's exposure and readout time in widefield imaging. To address this constraint, Dedecker's team proposed a novel intra-exposure excitation modulation (NEEMO) technique to distinguish spectrally overlapping fluorophores.237 Through rapid modulation of excitation light, NEEMO induced photoisomerization in photoswitchable fluorophores while acquiring time-integrated fluorescence (Fig. 18C).238 Using merely 2–4 captured images, the technique successfully differentiated bacteria individually labeled with distinct fluorescent proteins (EGFP, ffDronpa, ffDronpaF, and ffDronpa2F). Afterwards, the team developed an extended NEEMO approach to discriminate these four differently targeted fluorescent proteins expressed in HeLa cells, achieving fluorophore assignment accuracy of 80–95% (Fig. 18D).238 Unlike conventional methods, NEEMO bypasses direct kinetic analysis and instead leverages solely the modulation rate of the light source, allowing multiplexed imaging based on broad temporal dynamics.
Current dynamic-contrast FI generally employs a single temporal dimension, necessitating prolonged acquisition time and characteristic time span to distinguish multiple photoswitchable fluorophores. Jullien's team developed light-tunable time-gated reading-out of photocycles for multiplexed FI (LIGHTNING), a technique that substantially increases the number of distinguishable and identifiable RSFPs.239 LIGHTNING capitalizes on the primary light- and thermally-driven reactions of RSFPs, exploiting the dependence of photochemical reaction rate on light intensity to extract rich, RSFP-specific kinetic information. By applying multi-color illumination sequences at varying intensities, multiple characteristic times of RSFPs are collected, parameterizing the evolution of their fluorescence signals into a LIGHTNING kinetic fingerprint for each RSFP. LIGHTNING successfully distinguished 20 out of 22 spectrally similar green RSFPs in solution. Moreover, when tested on 16 Escherichia coli (E. coli) strains, each expressing a distinct RSFP, it accurately identified 9 strains with 93% accuracy. These results suggest that LIGHTNING can reliably discriminate fluorescent labels in complex samples, underscoring its potential for chromatic-aberration-free, multiplexed FI without the requirement for special optics or extensive data processing.
FRET offers an effective tool for understanding molecular interaction mechanisms, but its use in multiplexed imaging is limited by the difficult establishment of multiple donor–acceptor FRET pairs due to spectral overlap. Dedecker et al. proposed a strategy to distinguish spectrally similar FRET pairs by harnessing the photochromic parameters of the donor.240 This technique relies on cyclically switching the donor between fluorescent and non-fluorescent states by cyclic light irradiation, followed by calculating the photoswitching ratio from fluorescence changes of the donor. By constructing a photochromic FRET biosensor for cAMP-dependent protein kinase (PKA) activity (rsAKARev) and a spectrally overlapping but non-photochromic FRET biosensor for extracellular signal-regulated kinase (ERK) activity (EKARev), simultaneous detection of both kinases in the same cell was achieved. Additionally, combining this separation strategy with a red-shifted calcium sensor (RCaMP) enabled triple measurement of PKA, ERK, and calcium dynamics, demonstrating the efficacy of this method for FRET-based multiplexed imaging.
4.2. Phosphorescence imaging
PI is achieved by detecting the release of stored photons from phosphors after illumination ceases. The emission lifetime of phosphorescence (milliseconds to seconds) is significantly longer than that of fluorescence (nanoseconds), effectively suppressing autofluorescence interference with improved SNR.241–245 Currently, PI often suffers from weak signal intensity and environmental quenching under biological environments. Moreover, most phosphors emit at short wavelengths, limiting their applications in deep-tissue imaging.246 Although photoswitchable molecules lack intrinsic PI ability, they can spatiotemporally modulate phosphor performance by reversible on-off switching.Recent progress has centered on developing photoswitchable phosphorescent probes with red-shifted emission via phosphorescence resonance energy transfer (PRET). Unlike FRET, PRET enable energy transfer from a triplet state to a singlet state, thus extending the lifetime of conventional phosphors. Leveraging this mechanism, Liu's team constructed a series of DAE-derived supramolecular assemblies for NIR PI.117,164,247–250 For instance, a DAE derivative (DTE-C3) modified with binary phenylpyridinium units was assembled with cucurbit[8]uril (CB[8]) to generate a photoswitchable phosphorescent probe (DTE-C3⊂CB[8]) (Fig. 19A).15 In this system, the host-guest complex between CB[8] and DTE, described as a “single molecular dual-fold”, exhibited strong room-temperature phosphorescence. Additionally, the FRET interaction between ring-opened DTE and co-encapsulated Cy5 extended the phosphorescence peak from 600 to 673 nm, improving penetration depth. Upon UV or Vis light irradiation, the phosphorescence signal was switched off/on owing to the isomerization between the non-luminescent ring-closed DTE and luminescent ring-opened DTE, which was further tested for photoswitchable PI in cells (Fig. 19B). On the basis, DTE-C3 was functionalized through substituting the terminal bromine atoms with 5-methyl thiophene moieties, followed by assembly with CB[8] (Fig. 19C).164 The optimized probe demonstrated extended phosphorescence emission at 700 nm, achieving photo-controllable NIR PI of lysosomes in living cells. The NIR dye IR780 was incorporated to further extended the emission wavelength to 817 nm through the PRET effect (Fig. 19D). Collectively, photoswitchable phosphorescent probes offer a potential alternative for deep-tissue imaging with spatiotemporal precision.
 | ||
| Fig. 19 (A) Structure of DAE-C3 and its supramolecular assembly exhibiting photoswitchable phosphorescence at 673 nm. Reproduced with permission.15 Copyright 2022, Wiley-VCH. (B) Photoswitchable PI in living cells under UV/Vis irradiation. Reproduced with permission.15 Copyright 2022, Wiley-VCH. (C) Photoswitchable phosphorescence of DAE-TP-based supramolecular assembly (DTE-TP/CB[8]) at 700 nm. Reproduced with permission.164 Copyright 2022, Wiley-VCH. (D) Phosphorescence spectra of DTE-TP/CB[8]. Reproduced with permission.164 Copyright 2022, Wiley-VCH. | ||
4.3. Photoacoustic imaging
PAI combines the high sensitivity and contrast of optical imaging with the superior spatial resolution of ultrasound, circumventing the depth limitation typical of optical imaging (∼1 mm).251,252 In PAI, biological samples are typically illuminated with a nanosecond-pulsed laser. The molecules within the samples absorb the light energy and convert it into thermal energy, causing a localized temperature increase that generates a photoacoustic signal.253 However, endogenous chromophores in blood, such as hemoglobin and melanin, introduce background interference, which severely undermine detection sensitivity and resolution. This highlights the necessity for developing high-contrast PAI probes.254 Benefiting from the photoswitching behavior, PICAs can effectively eliminate background signals by subtracting the PAI signals respectively acquired in the on and off states.92,93 Furthermore, the controllable switching between these two distinct isomeric states considerably improves detection sensitivity and the robustness of image reconstruction.Genetically encoded photochromic chromoproteins, notably bacterial phytochromes (BphPs), offer advantages due to their high photostability, harmless non-invasiveness, and ability to be expressed by tissue-specific promoters. BphPs consist of a photosensory core module (PCM) and various effector domains. BphPs bind to biliverdin IXα chromophore, enabling differential PAI because of its two isomeric states, Pfr (ground state) and Pr. Reversible switching between these two states is mediated by cis–trans isomerization of the D-ring around the C15![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C16 double bond. Wang's group developed a NIR photochromic BphP from Rhodopseudomonas palustris (referred to as RpBphP1).92 The ground-state RpBphP1 (on state) transited to the Pr isomer (off state) upon illumination with 780 nm light, and reversed back upon illumination with 630 nm light or through natural relaxation with a half-life of ∼210 s (Fig. 20A). The robust and repeatable photoswitching could be maintained over many cycles without photobleaching. In scattering media, the PAI signal amplitude of RpBphP1 at a depth of 10 mm was 10 times higher than that of rsTagRF, a convention RSFP that can be toggled between 440 nm and 570 nm light illumination. This enhancement is attributed to the ∼200-nm red-shifted absorption and twofold higher extinction coefficient of RpBphP1. Differential PAI of RpBphP1 showed 21-fold increase in contrast-to-noise ratio (CNR) compared with on-state imaging alone. U87 human glioblastoma cells were engineered to express RpBphP1 for monitoring tumor growth in mice. After 20 switching cycles, differential PAI could clearly indicate kidney tumors at depths of up to 8 mm and brain tumors at ∼2 mm depth, with an average CNR of 20. Furthermore, PAI provided accurate longitudinal tracking of both primary hepatic tumor progression and tumor metastases over a one-month period (Fig. 20B). This study provides groundbreaking guidance for noninvasive multi-contrast deep-seated PAI. Subsequently, the same group developed another NIR photochromic BphP from Deinococcus radiodurans (referred to as DrBphP-PCM).93 DrBphP-PCM exclusively preserved the 55-kDa PCM region, resulting in improved folding efficiency and a 2.3-fold higher expression level in U87 cells than RpBphP1 (Fig. 20C). Along with significantly increased NIR absorption and photoswitching contrast, DrBphP-PCM demonstrated a 10-fold enhancement in CNR compared to RpBphP1. Through single-impulse panoramic photoacoustic computed tomography, brain, kidney, and liver tumors expressing DrBphP-PCM were successfully detected in vivo with a resolution of 125 µm. Therefore, reducing the size of protein-based probes offers a potential pathway to resolution enhancement.
C16 double bond. Wang's group developed a NIR photochromic BphP from Rhodopseudomonas palustris (referred to as RpBphP1).92 The ground-state RpBphP1 (on state) transited to the Pr isomer (off state) upon illumination with 780 nm light, and reversed back upon illumination with 630 nm light or through natural relaxation with a half-life of ∼210 s (Fig. 20A). The robust and repeatable photoswitching could be maintained over many cycles without photobleaching. In scattering media, the PAI signal amplitude of RpBphP1 at a depth of 10 mm was 10 times higher than that of rsTagRF, a convention RSFP that can be toggled between 440 nm and 570 nm light illumination. This enhancement is attributed to the ∼200-nm red-shifted absorption and twofold higher extinction coefficient of RpBphP1. Differential PAI of RpBphP1 showed 21-fold increase in contrast-to-noise ratio (CNR) compared with on-state imaging alone. U87 human glioblastoma cells were engineered to express RpBphP1 for monitoring tumor growth in mice. After 20 switching cycles, differential PAI could clearly indicate kidney tumors at depths of up to 8 mm and brain tumors at ∼2 mm depth, with an average CNR of 20. Furthermore, PAI provided accurate longitudinal tracking of both primary hepatic tumor progression and tumor metastases over a one-month period (Fig. 20B). This study provides groundbreaking guidance for noninvasive multi-contrast deep-seated PAI. Subsequently, the same group developed another NIR photochromic BphP from Deinococcus radiodurans (referred to as DrBphP-PCM).93 DrBphP-PCM exclusively preserved the 55-kDa PCM region, resulting in improved folding efficiency and a 2.3-fold higher expression level in U87 cells than RpBphP1 (Fig. 20C). Along with significantly increased NIR absorption and photoswitching contrast, DrBphP-PCM demonstrated a 10-fold enhancement in CNR compared to RpBphP1. Through single-impulse panoramic photoacoustic computed tomography, brain, kidney, and liver tumors expressing DrBphP-PCM were successfully detected in vivo with a resolution of 125 µm. Therefore, reducing the size of protein-based probes offers a potential pathway to resolution enhancement.
 | ||
| Fig. 20 RSFP-based PICAs for PAI. (A) Structure of RpBphP1 and photoisomerization behavior of the photoswitchable moiety (BV chromophore) between Pfr and Pr states. Reproduced with permission.92 Copyright 2016, Springer Nature. (B) Deep-tissue PAI of RpBphP1 in the kidney region of nude mouse. Reproduced with permission.92 Copyright 2016, Springer Nature. (C) Structure and PAI signals of the small-size RSFP, DrBphP-PCM. Reproduced with permission.93 Copyright 2018, Springer Nature. (D) Schematic illustration of tumor-homing bacteria engineered with RSFPs for photoswitchable PAI on tumor. Reproduced with permission.258 Copyright 2022, The National Academy of Sciences. | ||
The heterogeneous distribution in endogenous biliverdin poses a challenge for BphP-mediated PAI of specific tissues or organs. Kasatkina et al. employed a Cre recombinase-dependent method to express BphP1 with temporal and spatial targeting.255 In BphP1-Cre transgenic mice, differential PAI of BphP1 efficiently eliminated blood background noise, and indicated distinct BphP1 expression patterns across organs, such as higher signals in liver and spleen compared to stomach. The intracellular BphP1 concentration within these organs was estimated at ∼100 nM. This system was also successfully applied for non-invasive imaging of developing embryos and longitudinal tracking of viral-mediated gene expression in the kidney and liver. These results demonstrate the high sensitivity and deep penetration of PAI achieved by Cre-dependent BphP1 expression. In this study, beyond its PAI role, BphP1 also functioned as an optogenetic promoter to activate target gene expression.
Targeted delivery of exogenous BphPs or their encoding genes represents an alternative strategy. Chen et al. engineered a protein assembly, DrBphP-CBD, by fusing the photosensory module of DrBphP with a collagen-binding domain (CBD).256,257 DrBphP-CBD exhibited high affinity for type I and III collagen, and preferentially accumulated and retained in collagen-rich tumors following intravenous administration. This construct maintained superior reversible photoswitching properties, including high photoswitching contrast, photobleaching resistance, and an absorption coefficient ratio (Pfr/Pr) of 11 at 760 nm. In aqueous media, it achieved a 20-fold increase in the SNR of differential PAI compared to the on-state Pfr. Ten hours after administration, DrBphP-CBD demonstrated approximately 7-, 10.6-, and 2.5-fold higher differential PAI signals than DrBphP in MDA-MB-231, MC38, and 4T1 solid tumor models, respectively. The differential accumulation of DrBphP-CBD across tumor models is attributed to higher collagen expression in MDA-MB-231 and MC38 than 4T1. Compared to non-photoswitchable IR780 nanoparticles, DrBphP-CBD significantly enhanced imaging contrast and background discrimination for real-time dynamic monitoring and accurate quantitative analysis. This study advances photoswitchable phytochrome-based PAI imaging and provides a novel strategy for precision tumor diagnosis. Notably, although the endogenous blood protein-derived CBD may reduce immunogenicity, the tissue toxicity of DrBphP-CBD still needs more comprehensive evaluation. In a study, tumor-homing Escherichia coli (E. coli, MG1655) was reprogrammed to express a genetically encoded BphP, mDrBphP-PCMm/F469W (Fig. 20D).258 mDrBphP-PCMm/F469W maintained most of the superior photoswitching properties of BphPs, while exhibiting 1.75-fold and 4.2-fold higher photoswitching contrast compared to DrBphP-PCM and RpBphP1, respectively. After being intravenously administrated to 4T1 tumor-bearing mice, E. coli preferentially migrated toward the solid tumor and stated to proliferate. The tumor-tropic accumulation of E. coli enabled differential PAI of deep-seated tumor tissues with significantly enhanced imaging contrast. In contrast, only negligible PAI signals were observed in normal tissues or organs, demonstrating the remarkable tumor-targeting ability of E. coli for PAI probe delivery. Collectively, spatiotemporally selective PAI of BphPs is expected to benefit early tumor detection and tumor progression monitoring.
PAI probes based on organic photoswitchable molecules surpass BphPs in chemical tunability and diversity, while offering simpler screening and preparation advantages. In one study, two (1-(4-(1,2,2-triphenylvinyl)phenyl)ethylidene)malononitrile units were attached to the terminal positions of a DTE core (DTE-TPECM).63 In its ring-closed form, the planar conjugated structure formed by the two thiophene rings promoted intramolecular energy transfer and interactions, redirecting energy dissipation toward thermal relaxation for efficient PAI. To boost biocompatibility in vivo, DTE-TPECM was assembled with an amphiphilic lipid-PEG2000 into nanoparticles (DTE-TPECM NPs). These nanoparticles showed photoacoustic intensity 2.0-fold and 1.8-fold higher than that of methylene blue and polymer nanoparticles, respectively, and markedly enhanced resistance to reactive oxygen and nitrogen species compared to ICG. Following conjugation with the targeting peptide YSAYPDSVPMMS, DTE-TPECM NPs preferentially enriched in tumors that overexpress the EphA2 protein. In 4T1-tumor-bearing mice, intravenous injection of the peptide-modified DTE-TPECM NPs successfully achieved precise PAI of deep-seated tumors to guide tumor resection. On the other hand, upon 610 nm light irradiation for 5 minutes, these nanoparticles underwent photoisomerization to the ring-opened state, thereby restoring fluorescence to indicate postoperative tumor residues.
However, most organic photoswitchable molecules require high-energy light (e.g., UV light) for isomerization, restricting their applicability in deep-tissue differential PAI. To overcome this issue, UCNPs have been utilized to convert low-energy NIR light into UV energy for the reversible photoisomerization of small-molecule photoswitches. One study designed a photoswitchable hybrid nanoprobe by integrating a dithienylethene-containing β-diketone (3ThacacH, exhibiting strong photoacoustic signal under the ring-closed state) and a core-shell structured UCNP.259 The UCNP consisted of a non-doped NaYF4 shell, and a NaYF4 nanocrystal core co-doped with ytterbium (Yb) for NIR light harvesting and thulium (Tm) for UV emission. The UCNP surface was coated with an amphiphilic polymer (PSMA-M2070) to form a hydrophobic matrix layer for loading 3ThacacH. Upon 980 nm light irradiation, the UCNP efficiently converted NIR light to 365 nm UV emission, which spectrally overlaps with the absorption of the ring-opened 3ThacacH, thus driving its photocyclization to the closed state. This mechanism enabled reversible photoacoustic signal switching: 980 nm light switched the signal on, while 640 nm light returned the system to the ring-opened state to turn it off. The nanoprobe showed a photoacoustic on/off signal ratio of ∼10, and maintained a CNR over 3 in hemoglobin solution even at a low concentration of 0.05 nM, indicating its superior ability to suppress background interference by differential imaging. When used to HeLa cell labeling, nanoprobe-based differential imaging by 20 switching cycles achieved a detection sensitivity of 5 × 103 cells in vitro and 104 subcutaneously implanted cells in vivo, a two-order-of-magnitude increase over conventional photoacoustic contrast agents such as ICG. Notably, employing low-energy 980 nm light irradiation to switch on PAI minimized phototoxicity and extended imaging depth to ∼1.8 cm. In another study, analogous NaYF4 UCNPs were functionalized with a photoswitchable Azo-grafted polymer (PAA-Azo) via ligand exchange method (Fig. 21A).260 Upon 808 nm light irradiation, UCNPs simultaneously emitted 365 nm UV light and 475 nm blue light. This dual-wavelength output not only triggered reversible trans–cis–trans isomerization of PAA-Azo to produce vibrational energy and heat, but also transformed it into a liquid state with a higher thermal expansion coefficient. These effects synergistically amplified photoacoustic signal by sixfold compared to uncoated UCNPs (Fig. 21B). The probe allowed high-resolution PAI of mouse femora artery and rapid lymph node detection, clearly visualizing tissue structure and function. Moreover, by combining the benefits of PAI and NIR-II imaging, the probe supported real-time, precise guidance during lymph node resection.
 | ||
| Fig. 21 Photoswitchable molecule-based PICAs for PAI. (A) Principal concept for photoacoustic signal enhancement through the construction of Azo-UCNP-based polymer nanoparticles. Reproduced with permission.260 Copyright 2019, Wiley-VCH. (B) Dual photoacoustic and NIR-II imaging of lymph nodes in mice using Azo-UCNP-based polymer nanoparticles. Reproduced with permission.260 Copyright 2019, Wiley-VCH. (C) Structures of Azo-cyanine dyes. Reproduced with permission.17 Copyright 2024, Wiley-VCH. (D) In vivo PAI showing the biodistribution of wsT-AzHCy in nude mice. Reproduced with permission.17 Copyright 2024, Wiley-VCH. | ||
In addition, molecular switches have been integrated with existing organic photoacoustic contrast agents to address their limitations in PAI, such as photostability and nonlinear responses to excitation intensity caused by slow S1 → S0 electronic relaxation. Azo, with its non-fluorescent emission and picosecond S1 → S0 relaxation halftime, serves as an excellent energy acceptor for nonradiative heat dissipation.261,262 Furthermore, Gambhir's work demonstrated the complex Azo BHQ3 as a strong photoacoustic emitter.263 However, no water-soluble Azo derivatives have been available in the ideal NIR window for PAI. Therefore, Thorn-Seshold et al. developed a series of azohemicyanines by grafting the Azo switch onto the middle or terminal position of the NIR cyanine dyes (strong absorption to enter S1 state) for enhanced photoswitchable PAI (Fig. 21C).17 Benefiting from ultrafast vibrational relaxation (<10 ps, >100-fold faster than ICG), most synthetic azohemicyanines with NIR absorption demonstrated > 10-fold higher photostability compared to ICG. Furthermore, the ultrafast vibrational relaxation enabled multiple S1 → S0 → S1 excitation cycles within a single laser pulse, significantly enhancing the photoacoustic signal generation efficacy of these probes to over 3 times that of ICG. The improved signal longevity and loudness effectively addressed the nonlinear response of cyanine photoacoustic dyes to illumination intensity, thus achieving higher photoacoustic signal output for improved resolution and lower background. Among all azohemicyanin, the hydrophilic thiophenylazo-bridged probe (designated as wsT-AzHCy, absorption peak at 729 nm) showed the optimal potential for in vivo PAI, with >1900-fold enhanced photostability and 3.2-fold signal intensity. Two hours after intravenous administration of wsT-AzHCy, PAI successfully separated the probe signal from endogenous background signal, suggesting the sufficient survival time of wsT-AzHCy for practical applications (Fig. 21D). Furthermore, the photoacoustic signal was predominantly observed in the liver, spleen, and gut, but absent in the kidneys—a promising indication for future molecular imaging. This work proposes a novel chemical concept that leverages molecular switches to accelerate the S1 → S0 relaxation of photoacoustic contrast agents for improved PAI.
4.4. Magnetic resonance imaging
Magnetic resonance imaging (MRI) is a noninvasive, non-ionizing radiation, and multi-dimensional imaging modality.264 The signal amplification of MRI contrast agents stems from paramagnetic centers that perturb proton relaxation kinetics. Specifically, hydration-shell 1H nuclei under an external applied field experience magnetic field distortion, which enhances signal amplitude via modulating T1/T2 (longitudinal/transverse) relaxation time.265 In T1-weighted imaging, short T1 contrast agents (e.g. Gad-type) produce bright signals; conversely, in T2-weighted imaging, long T2 contrast agents (e.g. Iron-type) enhances MRI intensity. Since the initial clinical approval of gadopentetate dimeglumine (Gd-DTPA) in 1988, over 200 million doses of MRI contrast agents have been administered globally.266 However, the suboptimal relaxivity of current MRI contrast agents often requires high doses for adequate image differentiation.Over the past decades, photo-activated MRI contrast agents have enabled spatiotemporally precise signal modulation and enhanced proton relaxivity.267,268 Unlike unidirectional photocage agents that are limited to single-use contrast release, PICAs support repeatable signal switching, thereby facilitating longitudinal MRI tracking and higher SNR. Upon light irradiation, photoisomerization modulates proton relaxivity by two primary mechanisms: (1) photoisomerization of photoswitchable moiety changes the symmetry of the primary hydration shell around paramagnetic centers (e.g. Gd3+ and Ni2+), which directly governs proton relaxivity; (2) light-triggered conversion of the metal coordination geometry (when photoswitchable molecule as a ligand) induces the switching from a diamagnetic state to a paramagnetic state, thereby amplifying proton relaxivity.
Photoswitchable MRI probes primarily employs SP or Azo as photoswitching moieties, leveraging their pronounced conformation changes to dynamically modulate MRI signals. Since 2007, Louie et al. have systematically studied SP-based PICAs for MRI.269–273 The photoisomerization between the SP state and the MC state perturbs the molecular microenvironment to modulate water-proton relaxivity. In their initial study, they designed a photoswitchable MRI contrast agent (Gd-SPDO3A) through coordinating SP with Gd-DO3A, enabling reversible photoswitching of MRI signals (Fig. 22A).269 Under dark-adapted equilibration, coordination between Gd3+ and the phenolate oxygen promoted the SP → MC isomerization and stabilized the metastable MC form in Gd-MCDO3A, thereby generating an absorption peak at 486 nm. Upon Vis light irradiation, the MC → SP photoisomerization increased the relaxivity (r1) of Gd-MCDO3A by 21.2% (from 3.72 to 2.93 mM−1 s−1). This increase resulted from the shortened distance between the indoline unit of SP and Gd3+ through enhanced electrostatic interaction, which restricted water access to the paramagnetic center—Gd3+ ion.
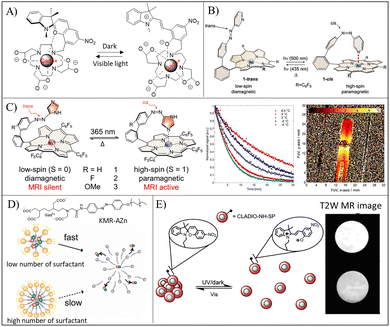 | ||
| Fig. 22 (A) Photoisomerization of Gd-SPDO3A (left) and Gd-MCDO3A (right). Reproduced with permission.269 Copyright 2007, Royal Society of Chemistry. (B) Light-induced coordination and de-coordination between azopyridine and Ni2+ for magnetic signal switching between diamagnetism and paramagnetism via trans–cis isomerization of azopyridine. Reproduced with permission.18 Copyright 2011, American Association for the Advancement of Science. (C) Schematic illustration of the photoisomerization of azoimidazole-based PICAs and time-dependent thermal MRI mapping. Reproduced with permission.276 Copyright 2021, American Chemical Society. (D) Structure of amphiphilic Azo derivatives (KMR-AZn) and KMR-AZn-based micelles. Reproduced with permission.278 Copyright 2012, Wiley-VCH. (E) Photo-induced aggregation of CLADIO-NH-SP with reduced T2 MRI signal. Reproduced with permission.279 Copyright 2010, American Chemical Society. | ||
Currently, the in vivo application of photoswitchable MRI agents is limited by potential phototoxicity under high-intensity illumination. Louie's group developed a SP-based MRI contrast agent that undergoes MC → SP isomerization upon irradiation with low-intensity Vis light.271 Specifically, just 1 min of exposure to blue LED light (λ = 465 nm, <2.37 × 1012 photons s−1) converted MC-DO3A-Gd into SP-DO3A-Gd, decreasing its r1 relaxivity from 2.93 to 2.63 mM−1 s−1. Notably, this light intensity matches the emission level of luciferase-luciferin systems, indicating the feasibility of bioluminescence-driven photoisomerization. Although luciferase emission alone failed to trigger the MC → SP isomerization, this approach retains potential for future endogenously activated MRI.
Unlike SP-based MRI agents that attenuate MRI signals by disrupting the interaction between water and paramagnetic center, Azo-based MRI agents exhibit enhanced paramagnetism. Their trans → cis photoisomerization enables switching between diamagnetic state and paramagnetic state via light-driven coordination-induced spin-state switching (LD-CISSS). Herges’ group has been dedicated to developing Azo-based photoswitchable MRI probes and utilizing trans-cis isomerization to modulate the spin state of coordinated transition metals.3,18,274–276 For instance, they designed an Azo-based PICA (1-trans) in which trans-to-cis isomerization of the phenylazopyridine (PAP) modulates axial coordination between the pyridyl nitrogen at the Azo terminal and the square-planar Ni-porphyrin, thereby driving the spin transition from low-spin (S = 0) to high-spin (S = 1) (Fig. 22B).18 Irradiation with 500 nm light converted the diamagnetic 1-trans isomer to the paramagnetic 1-cis isomer with a yield of ∼65%. Conversely, exposure to 435 nm light reversed the isomerization process with a yield of ∼90%. This reversible photoswitching persisted for >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cycles, indicating the outstanding fatigue resistance of PAP-based PICAs. Notably, the magnetic moment of the 1-cis isomer was 2.989 ± 0.084 B.M., consistent with paramagnetic Ni2+ complexes (expected range: 2.8 to 3.4 B.M.), while the resultant 1-trans isomer upon blue light irradiation exhibited diamagnetism behavior with a negligible magnetic moment (µ ≈ 0 B.M.). To improve clinical translation potential, the group further presented an optimized design of a molecular spin-state switch that is water-soluble and switchable in the far-red and NIR regions.277 By engineering porphyrins into substituted dihydroporphyrin (chlorin) or tetrahydroporphyrin (bacteriochlorin), the newly synthesized PICAs demonstrated enhanced photoswitching performance: cis ↔ trans (also diamagnetic ↔ paramagnetic) switching efficiency reached 72–78%, while spin-state switching efficiency in aqueous systems raised from 19% to 41%. Moreover, both chlorin and bacteriochlorin probes exhibited a blue-shift absorption and reversible photoisomerization upon long-wavelength illumination: 505 nm light irradiation triggered cis-to-trans conversion, whereas 620/720 nm light irradiation induced trans-to-cis switching. Such NIR-activatable switching offers an attractive translational prospect for deep-tissue MRI diagnosis.
000 cycles, indicating the outstanding fatigue resistance of PAP-based PICAs. Notably, the magnetic moment of the 1-cis isomer was 2.989 ± 0.084 B.M., consistent with paramagnetic Ni2+ complexes (expected range: 2.8 to 3.4 B.M.), while the resultant 1-trans isomer upon blue light irradiation exhibited diamagnetism behavior with a negligible magnetic moment (µ ≈ 0 B.M.). To improve clinical translation potential, the group further presented an optimized design of a molecular spin-state switch that is water-soluble and switchable in the far-red and NIR regions.277 By engineering porphyrins into substituted dihydroporphyrin (chlorin) or tetrahydroporphyrin (bacteriochlorin), the newly synthesized PICAs demonstrated enhanced photoswitching performance: cis ↔ trans (also diamagnetic ↔ paramagnetic) switching efficiency reached 72–78%, while spin-state switching efficiency in aqueous systems raised from 19% to 41%. Moreover, both chlorin and bacteriochlorin probes exhibited a blue-shift absorption and reversible photoisomerization upon long-wavelength illumination: 505 nm light irradiation triggered cis-to-trans conversion, whereas 620/720 nm light irradiation induced trans-to-cis switching. Such NIR-activatable switching offers an attractive translational prospect for deep-tissue MRI diagnosis.
MRI signals depend on multiple interdependent parameters (e.g., contrast agent concentration, temperature sensitivity, and relaxation time). Body temperature is a critical biomarker in clinical diagnosis for tissue inflammation, metabolic disorders, and tumors. It is important to separating temperature contribution to the generated MRI signals through independent quantification of these parameters. However, current methods impose technical barriers to hinder clinical translation. Herges et al. designed a series of azoimidazole-based photoswitchable MRI probes to assess time-dependent thermal MRI mapping, superseding conventional static relaxation time measurement (Fig. 22C).276 Upon 365 nm light irradiation, cis-PICAs (∼60% yield) showed accelerated thermal relaxation, reverting completely to the trans configuration within seconds to minutes at room temperature. The thermo-driven cis–trans isomerization of the azoimidazole triggered dissociation of the imidazole ligand from Ni2+, resulting in a spin-state switching in the Ni-porphyrin from paramagnetic (MRI-detectable) to diamagnetic (MRI-silent). This succeeded in the acquirement of MRI signals with standard clinical sequences by exploiting the intrinsic correlation between temperature and kinetic parameters (reaction rate and t1/2), which is independent of probe concentration. These thermo-responsive MRI PICAs enabled high-resolution temperature mapping (130 × 130 µm) with ± 0.22 °C accuracy in vitro through monitoring the thermal relaxation kinetics of cis-Azo.
In addition, photoswitchable molecules enable dual-level MRI signal modulation: (1) molecular-level signal control by photoswitchable metal-ligand coordination; and (2) nanoscale regulation between aggregated state and dispersed state by photoisomerization. Compared with free molecular agents, nanoscale contrast agents demonstrate significantly higher MRI signal intensity due to increased local concentration of paramagnetic centers and restricted molecular motion upon molecular aggregation. To improve photoswitchable MRI contrast, a series of nanomicelle probes (KMR-AZn) were developed through the self-assembly of an engineered amphiphilic molecule that couples a hydrophobic Azo derivative with hydrophilic Gd-DTPA, while also encapsulating a hydrophobic drug substitute, Nile Red (Fig. 22D).278 Upon 365 nm UV irradiation, the Azo derivative underwent trans-to-cis photoisomerization, evidenced by attenuation of the 350 nm absorption peak. This conversion triggered Nile Red release, indicating photo-controllable disassembly of the aggregated architecture. KMR-AZn exhibited high longitudinal relaxivity (14.5–16.5 mM−1 s−1), surpassing clinical Gd-DTPA (4.1 mM−1 s−1) by 3–4-fold. Following UV irradiation, r1 value decreased from 15.3 to 12.9 mM−1 s−1 for KMR-AZ6 and from 16.5 to 14.7 mM−1 s−1 for KMR-AZ8, demonstrating reversible MRI signal photoswitching.
Similar to T1-weighed MRI, the MC → SP isomerization also decreases the signal intensity of T2-weighed MRI. Louie et al. designed a photoswitchable MRI probe based on SP-modified iron oxide nanoparticles (CLADIO-NH-SP) to modulate the T2 relaxation time (Fig. 22E).279 CLADIO-NH-SP was initially incubated in the darkness for >1 h to ensure complete SP → MC photoisomerization. Upon Vis light irradiation, the absorption at 563 nm decreased concomitant with the nanoparticle hydrodynamic diameter transiting from 200 nm in the dispersed state (CLADIO-NH-MC) to 520 nm in the aggregated state (CLADIO-NH-SP). After 2-hour dark incubation, CLADIO-NH-SP gradually depolymerized, accompanied by increased absorption at 563 nm and restoration of the CLADIO-NH-MC with dispersed size distribution, indicating reversible SP ↔ MC photoisomerization. CLADIO-NH-MC exhibited a T2 relaxation time of 37.09 ± 3.09 ms in the darkness, while CLADIO-NH-SP upon Vis light irradiation showed a shortened T2 relaxation time of 24.55 ± 1.86 ms. This change in magnetic response stems from enhanced interparticle magnetic coupling by nanoparticle aggregation. In T2-weighted imaging, Vis light exposure induced a 15% signal decline from 249 ± 5 to 211 ± 3 arbitrary units, correlating with reduced T2 relaxation time. This photoswitchable probe holds promise as an MRI gene reporter for deep-tissue luciferase expression system.
4.5. Raman imaging
Raman scattering is an inelastic scattering phenomenon where photons interact with molecules, resulting in energy exchange that reveals molecular vibrational information.272,273 RI acquires high-resolution maps of the chemical composition and molecular structure of a sample by spatially tracking its Raman scattering signals. This technique supports both label-free and targeted small-molecule imaging, and exhibits superior multiplexing capability because Raman spectral linewidths are ∼50 times narrower than those of FI.280,281 Stimulated Raman scattering (SRS) overcomes the low sensitivity limitation of spontaneous RI, while electronic pre-resonance SRS (epr-SRS) further improves sensitivity to 250 nM and multiplexity of up to 24 colors.226,282 Nevertheless, conventional Raman probes suffer from static signals and are susceptibility to photodegradation during prolonged illumination, which undermines imaging stability. By contrast, photoswitchable Raman probes can dynamically alter their vibrational signatures via photoisomerization, generating spectral shift that improves imaging contrast and multiplexing capability. Additionally, the reversible on/off switching of Raman signals allows spatiotemporally controlled dynamic imaging.Small-molecule Raman probes offer distinct advantages such as single-wavelength excitation for analyzing multiple vibrational modes, exceptional photostability, and reaction specificity, but suffer from relatively low sensitivity. Gao's team designed LysoAzo-pH, an Azo-based resonance Raman pH probe for lysosomal pH sensing (Fig. 23A).283 The intrinsic Raman signal of LysoAzo-pH was 104 times greater than that of a conventional alkyne-based Raman reporter molecule, 5-ethynyl-2′-deoxyuridine. By substituting electron-donating and -withdrawing groups on the Azo moiety, simultaneous fluorescence quenching and resonance-enhanced Raman scattering was achieved, with an enhancement factor reaching approximately 103. The protonation/deprotonation-induced vibrational fingerprint shift of the Azo group enabled precise visualization of proton distribution dynamics in autophagic cells, achieving a sensitivity of <0.2 pH units. In addition, Ji et al. constructed photochromic vibrational probes for SRS imaging by modifying both ends of a DTE scaffold with strong Raman-tag acetylene groups (Fig. 23B).14 For the representative probe (DTE-Ph), UV light irradiation (365 nm) induced ring-closing photoisomerization of the conjugated DTE moiety, resulting in a 20 cm−1 red shift of the narrow acetylene Raman peak. Subsequent visible light irradiation (633 nm) completely converted DTE-pH to the ring-opened isomer, restoring the origin sharp Raman peak at 2214 cm−1. The reversible photoisomerization of DTE-Ph enabled high-contrast SRS imaging with a switching ratio of 50%. When used for targeting mitochondria, DTE-Ph achieved light-controlled “drawing” and “erasing” of cells, as well as dual-label multiplexed imaging (Fig. 23C). Furthermore, DTE-Ph exhibited excellent fatigue resistance (>100 switching cycles), promoting long-term tracking of mitochondrial dynamics. However, SRS beam-caused photobleaching of this probe, likely due to two-photon excited cycloreversion, remained a limitation in bioimaging. Further optimization of molecular design should focus on enhance photostability and imaging sensitivity.
 | ||
| Fig. 23 (A) Raman spectra at varying concentrations and RI of Azo-based PICA (LysoAzo-pH) in lysosomes. Reproduced with permission.287 Copyright 2021, American Chemical Society. (B) Spontaneous and stimulated Raman signal response in the photoisomerization of DTE-alkyne derivatives (DTE-TMS and DTE-Ph). Reproduced with permission.14 Copyright 2021, Springer Nature. (C) Live-cell painting and erasing with SRS microscopy using DTE-Ph-Mito. Reproduced with permission.14 Copyright 2021, Springer Nature. (D) Principle and design of RESORT microscopy. Reproduced with permission.285 Copyright 2023, American Association for the Advancement of Science. (E) Normal SRS imaging and RESORT imaging of live HeLa cells at 1503 cm−1 (Raman on-resonance). Reproduced with permission.285 Copyright 2023, American Association for the Advancement of Science. (F) Chemical structure of 8 DAE derivatives with their normalized SRS spectra in both the open and closed forms. Reproduced with permission.286 Copyright 2024, Springer Nature. | ||
Wei's team proposed a general design strategy for EPR-SRS probes that exploits molecular electronic and vibrational coupling to switch the molecular electronic state among four different configurations, thereby enabling photoswitchable control of ground-state EPR-SRS signal.284 Unlike small molecules, photoswitchable proteins possess intrinsic biocompatibility and markedly higher fatigue resistance. For example, the NIR photoswitchable DrBphP-PCM could shift its electronic resonance toward or away from the pump photon energy. After switching to the on state (Pfr state) under 690 nm light, DrBphP-PCM generated a Raman signal 200-fold stronger than that of 5-ethynyl-2′-deoxyuridine. In addition, it displayed outstanding stability (>40 switching cycles) and detection sensitivity below 1 µM, demonstrating its suitability for bioimaging based on this universal EPR-SRS strategy.
Traditional vibrational microscopy techniques possess certain limitations, such as requiring cell fixation and high-power illumination. Ozeki's team combined photoswitchable SRS with RESOLFT technology to develop reversible saturable optical Raman transitions (RESORT) microscopy, an approach that enabled super-resolution vibrational imaging (Fig. 23D).285 RESORT exploits a low-power (microwatt level) continuous-wave laser to modulate the SRS detection of photoswitchable Raman probes (under competitive conditions, only the photoswitchable probes at the center of the donut-shaped beam focus remain activating). The team designed a photoswitchable DAE derivative (DAE620) that reversibly toggles between on/off states by alternating irradiation with 375 nm light and 640 nm light, thereby activating or depleting Raman signals. Its ring-closed isomer displayed a strong SRS peak at 1503 cm−1, with an intensity 25 times higher than that of its ring-opened isomer. Significantly, DAE620 showed excellent fatigue resistance (>20 switching cycles) and ultrafast photoswitching kinetics (time constant, τ < 1 µs), providing critical foundation for RESORT imaging. In DAE620-labeled HeLa cells, RESORT reached a spatial resolution of 151 nm, approximately double that of conventional SRS imaging, surpassing the optical diffraction limit and revealing nanoscale organelle details in living cells (Fig. 23E). Moreover, RESORT demonstrated high contrast, chemical specificity, and minimal phototoxicity ascribing to its low-power irradiation. With the expanded library of photoswitchable Raman probes, RESORT is expected to propel highly multiplexed super-resolution bioimaging.
At present, research on reversible photo-controllable multiplexed SRS imaging remains limited. Hu's group developed a super-multiplexed Carbow palette (Carbow-switch) consisting of 16 photoswitchable polyyne-based Raman probes. These probes were built on an asymmetric DAE backbone and incorporated molecular design strategies such as polyyne conjugation, heterocyclic incorporation, and terminal functionalization (Fig. 23F).286 Their absorption peaks lied within a favorable range (560–720 nm), allowing efficient SRS excitation and signal amplification. Particularly, DAE derivatives based on the 1-methylindole skeleton displayed outstanding performance, including rapid photoswitching kinetics under low-power irradiation (microwatt level), high light-to-signal conversion yield (>70%, up to 93%), and robust fatigue resistance (>95% after 100 switching cycles). Reversible photoisomerization of these probes (triggered by alternating illumination with 405 nm light and 640 nm light) resulted in a significant Raman shift between 12–24 cm−1 and SRS signal enhancement up to 90-fold. With negligible spectral crosstalk and high signal resolution, Carbow-switch showed great potential for high-resolution multiplexed imaging. By targeted modifications, the team achieved photo-controllable 6-color and 9-color SRS imaging of specific organelles in living cells, and successfully observed dynamic changes of mitochondria under oxidative stress, lysosomal changes during stress granule formation, and intercellular organelle transport. Carbow-switch opens up a new avenue for studying subcellular dynamics and interactions in living cells with high spatiotemporal precision and multiplexing ability.
5. Conclusions
Photoswitchable molecules undergo reversible photoisomerization upon light irradiation at two distinct wavelengths, allowing precision bioimaging through tunable photophysical properties. These molecules serve as intrinsic PICAs or modulators for conventional ICAs, offering unique advantages: (1) background suppression via state-switching algorithms; (2) high fatigue resistance for stable long-term imaging; and (3) instantaneous signal control shortening acquisition time.Small-molecule photoswitches occupy a larger share in PICA market. They exhibit superior chemical tunability, enabling precise control over physicochemical parameters such as absorption/emission wavelengths, photoswitching rates, hydrophilicity, and targeting property through synthesis and modification. Concurrently, their high stability and diverse photoswitching mechanisms allow adaptation to multiple imaging modalities. Azo exhibits rapid trans–cis isomerization for dynamic imaging, but suffer from thermal instability and UV light-dependent photoisomerization. Strategies such as introducing steric hindrance groups and metal coordination can stabilize the cis isomer. The thermal relaxation of Azo plays a dual role: although it impedes fluorescence switching for super-resolution imaging, it contributes to photoacoustic signal generation.260 Moreover, light-controlled coordination of Azo to Ni2+ enables reversible switching between paramagnetic and diamagnetic MRI signals.18 The temperature-responsive relaxation of cis-Azo has been leveraged for 3D temperature MRI.276 Concurrently, the inherent molecular vibrational energy of Azo scaffold enhances RI signals.288,289
SP exhibits bright fluorescence in its MC state, which underpins super-resolution and dual-color imaging.195,224 Its multiple stimuli-responsive (ions/pH/temperature) ability expands biosensing applications,290 and reversible metal coordination allows MRI signal modulation. However, the slow switching kinetics and susceptibility to photodegradation require further electronic and steric optimization.291,292
DAE offers exceptional thermal stability and fatigue resistance, rendering it suitable for super-resolution and PI.22,107,164 The NIR absorption of its closed form supports PAI,63,66 while alkyne-functionalized derivatives enable photoswitchable RI.14 However, the relatively low cyclization quantum yield and limited magnetic moment variation impede its utility in MRI, necessitating further molecular design optimization.
In contrast, RSFPs are expressed in specific organelles or proteins via gene transfection, offering high biocompatibility and suitability for long-term dynamic imaging. They have been extensively utilized in precise single-molecule and cellular tracking, as well as protein-protein interaction research by super-resolution imaging.293–295 Recent studies has focused on improving photoswitching efficiency, photostability, and fluorescence quantum yield of RSFPs.296,297 RSFPs with NIR absorption demonstrate superior photoacoustic conversion capacity, enabling high-contrast PAI at tissue depth up to 8 cm.92,298 However, the instability and complex structure of RSFPs hinder their combination with exogenous contrast agents (e.g., inorganic nanoparticles, organic dyes, and metal chelates). Therefore, their broad implementation in PI, MRI, and RI still faces technical obstacles.
Recent progress has yielded numerous photoswitchable molecules that surpass classical counterparts mentioned above, greatly enriching the molecular repertoire for super-resolution imaging.299,300 However, photoswitchable imaging is still predominantly constrained to fluorescent technique. Advancing this field forward requires the development of next-generation PICAs capable of supporting multimodal applications. In clinic, accurate disease diagnosis and assessment often necessitates the integration of multiple imaging modalities, which can be achieved by the spatiotemporally programmable signal control via PICAs. For example, certain PICAs combine NIR absorption and fluorescence switching, enabling simultaneous photoswitchable PAI and FI.63,92 Similarly, coupling fluorescent photoswitchable molecules with magnetic metals facilitates dual-mode MRI and FI, where the on/off states of fluorescence and MRI signals are controlled independently.271,273 Notably, PICAs supporting concurrent imaging modes can benefit patients by reducing the requirement for separate administration of single-function contrast agents. Beyond imaging, rationally designed PICAs can also modulate cellular metabolism, enabling simultaneous disease diagnosis and therapeutic intervention. For instance, some PICAs serve as activatable photosensitizers for controlled generation of ROS—a key mediator in cellular damage, aging, and tumor progression.47 Furthermore, photoswitchable molecules can function as optogenetic actuators to reversibly regulate ion-channel gating, thereby precisely manipulating cellular behavior.294 Future efforts should prioritize the development of photoswitchable systems that combine multimodal imaging with diagnostics and therapeutic ability.
Critical limitations still persist in the design of photoswitchable molecules, including structural instability, hydrophobicity, short-wavelength excitation, and suboptimal photophysical properties. While functional group modification and structural adjustment offer partial solutions, they often trade off other essential properties, owing to the inherent spatial constraint of small molecule frameworks. Self-assembly strategies present a facile and scalable alternative to tackle these issues, albeit at the cost of losing precise molecular-level switching control, which potentially compromising imaging accuracy.
Another challenge is the application of PICAs in living systems. Developing PICAs with red-shifted excitation is promising to address phototoxicity and poor tissue penetration. Equally important is controlling their in vivo distribution. Modification with targeting groups and employing nanomaterial-based delivery systems can enhance the accumulation of PICAs at interested sites, thereby improving imaging accuracy while mitigating off-target side effects to healthy tissues. However, the potential toxicity from the introduced targeting groups or carriers should be thoroughly evaluated to promote the clinical translation of PICAs.
In summary, PICAs are rapidly evolving. With ongoing optimization and deep learning-directed predictive design, next-generation PICAs hold promise to revolutionize precision bioimaging across diverse modalities.
Conflicts of interest
No conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This study was supported by the National Natural Science Foundation of China (NSFC) (Grant No. 22371012, 82430067, and 32030060) and the National Key Research & Development Program of China (Grant No. 2021YFA1201000 and 2023YFC2605000). The authors also acknowledge support from the Major Project of Guangzhou National Laboratory (Grant No. GZNL2024A03010), the Key Grant of the Beijing Natural Science Foundation (Grant No. L258028), and the Science Fund for Creative Research Groups of Nature Science Foundation of Hebei Province (Grant No. B2021201038). H. B. C. received financial support from the China Scholarship Council (CSC) (Grant No. 202306880018). J. Y. thanks the Nano & Material Technology Development Program through the National Research Foundation of Korea (NRF) funded by Ministry of Science and ICT (Grant No. RS-2024-00407093) and the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (Grant No. 2022R1A2C3005420).Notes and references
- L. Wang and Q. Li, Chem. Soc. Rev., 2018, 47, 1044–1097 RSC.
- A. M. Rice, C. R. Martin, V. A. Galitskiy, A. A. Berseneva, G. A. Leith and N. B. Shustova, Chem. Rev., 2020, 120, 8790–8813 CrossRef CAS PubMed.
- M. Dommaschk, M. Peters, F. Gutzeit, C. Schutt, C. Nather, F. D. Sonnichsen, S. Tiwari, C. Riedel, S. Boretius and R. Herges, J. Am. Chem. Soc., 2015, 137, 7552–7555 CrossRef CAS PubMed.
- G. Naren, W. Larsson, C. Benitez-Martin, S. Li, E. Perez-Inestrosa, B. Albinsson and J. Andreasson, Chem. Sci., 2021, 12, 7073–7078 RSC.
- C. Li, H. Yan, L. X. Zhao, G. F. Zhang, Z. Hu, Z. L. Huang and M. Q. Zhu, Nat. Commun., 2014, 5, 5709 CrossRef CAS PubMed.
- M. Hofmann, C. Eggeling, S. Jakobs and S. W. Hell, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 17565–17569 CrossRef CAS PubMed.
- D. K. Tiwari, Y. Arai, M. Yamanaka, T. Matsuda, M. Agetsuma, M. Nakano, K. Fujita and T. Nagai, Nat. Methods, 2015, 12, 515–518 CrossRef CAS PubMed.
- T. J. Chozinski, L. A. Gagnon and J. C. Vaughan, FEBS Lett., 2014, 588, 3603–3612 CrossRef CAS PubMed.
- J. Fölling, V. Belov, R. Kunetsky, R. Medda, A. Schönle, A. Egner, C. Eggeling, M. Bossi and S. W. Hell, Angew. Chem., Int. Ed., 2007, 119, 6382–6386 CrossRef.
- H. Y. Jung, B. Kim, M. H. Jeon and Y. Kim, Small, 2022, 18, e2103523 CrossRef PubMed.
- K. Mishra, J. P. Fuenzalida-Werner, F. Pennacchietti, R. Janowski, A. Chmyrov, Y. Huang, C. Zakian, U. Klemm, I. Testa, D. Niessing, V. Ntziachristos and A. C. Stiel, Nat. Biotechnol., 2022, 40, 598–605 CrossRef CAS PubMed.
- D. Kim, K. Jeong, J. E. Kwon, H. Park, S. Lee, S. Kim and S. Y. Park, Nat. Commun., 2019, 10, 3089 CrossRef PubMed.
- J. Chen, W. Zhong, M. Xue, H. Wang, M. Yu, P. Zhang and P. Yi, Polym. Chem., 2017, 8, 6520–6526 RSC.
- J. Ao, X. Fang, X. Miao, J. Ling, H. Kang, S. Park, C. Wu and M. Ji, Nat. Commun., 2021, 12, 3089 CrossRef CAS PubMed.
- C. Wang, X. K. Ma, P. Guo, C. Jiang, Y. H. Liu, G. Liu, X. Xu and Y. Liu, Adv. Sci., 2022, 9, e2103041 CrossRef PubMed.
- X. Chai, H. H. Han, A. C. Sedgwick, N. Li, Y. Zang, T. D. James, J. Zhang, X. L. Hu, Y. Yu, Y. Li, Y. Wang, J. Li, X. P. He and H. Tian, J. Am. Chem. Soc., 2020, 142, 18005–18013 CrossRef CAS PubMed.
- M. Müller, N. Liu, V. Gujrati, A. Valavalkar, S. Hartmann, P. Anzenhofer, U. Klemm, A. Telek, B. Dietzek-Ivanšić, A. Hartschuh, V. Ntziachristos and O. Thorn-Seshold, Angew. Chem., Int. Ed., 2024, 63, e202405636 CrossRef PubMed.
- S. Venkataramani, U. Jana, M. Dommaschk, F. D. Sönnichsen, F. Tuczek and R. Herges, Science, 2011, 331, 445–448 CrossRef CAS PubMed.
- M. Canton, A. B. Grommet, L. Pesce, J. Gemen, S. Li, Y. Diskin-Posner, A. Credi, G. M. Pavan, J. Andréasson and R. Klajn, J. Am. Chem. Soc., 2020, 142, 14557–14565 CrossRef CAS PubMed.
- B. Roubinet, M. L. Bossi, P. Alt, M. Leutenegger, H. Shojaei, S. Schnorrenberg, S. Nizamov, M. Irie, V. N. Belov and S. W. Hell, Angew. Chem., Int. Ed., 2016, 55, 15429–15433 CrossRef CAS PubMed.
- M. Dong, A. Babalhavaeji, M. J. Hansen, L. Kálmán and G. A. Woolley, Chem. Commun., 2015, 51, 12981–12984 RSC.
- D. Kim, A. Aktalay, N. Jensen, K. Uno, M. L. Bossi, V. N. Belov and S. W. Hell, J. Am. Chem. Soc., 2022, 144, 14235–14247 CrossRef CAS PubMed.
- F. Aleotti, A. Nenov, L. Salvigni, M. Bonfanti, M. M. El-Tahawy, A. Giunchi, M. Gentile, C. Spallacci, A. Ventimiglia, G. Cirillo, L. Montali, S. Scurti, M. Garavelli and I. Conti, J. Phys. Chem. A, 2020, 124, 9513–9523 CrossRef CAS PubMed.
- M. Natali and S. Giordani, Chem. Soc. Rev., 2012, 41, 4010–4029 RSC.
- G. Kortüm, W. Theilacker and V. Braun, J. Chem. Phys, 1955, 23, 1723–1724 CrossRef.
- J. Zhang, Q. Zou and H. Tian, Adv. Mater., 2012, 25, 378–399 CrossRef PubMed.
- H. Nie, J. L. Self, A. S. Kuenstler, R. C. Hayward and J. Read de Alaniz, Adv. Opt. Mater., 2019, 7, 1900224 CrossRef.
- I. Yildiz, E. Deniz and F. M. Raymo, Chem. Soc. Rev., 2009, 38, 1859–1867 Search PubMed.
- H. Zhou, C. Xue, P. Weis, Y. Suzuki, S. Huang, K. Koynov, G. K. Auernhammer, R. Berger, H.-J. Butt and S. Wu, Nat. Chem., 2017, 9, 145–151 CrossRef CAS PubMed.
- A. Cembran, F. Bernardi, M. Garavelli, L. Gagliardi and G. Orlandi, J. Am. Chem. Soc., 2004, 126, 3234–3243 CrossRef CAS PubMed.
- C.-W. Chang, Y.-C. Lu, T.-T. Wang and E. W.-G. Diau, J. Am. Chem. Soc., 2004, 126, 10109–10118 Search PubMed.
- G. A. Woolley, Chem. Soc. Rev., 2011, 40, 4422–4437 RSC.
- H. B. Cheng, S. Zhang, J. Qi, X. J. Liang and J. Yoon, Adv. Mater., 2021, 33, e2007290 Search PubMed.
- H.-J. Yu, H. Wang, F.-F. Shen, F.-Q. Li, Y.-M. Zhang, X. Xu and Y. Liu, Small, 2022, 18, 2201737 CrossRef CAS PubMed.
- H. Jin, F. Liu, P. Wu, Z. Sun, P. Sui, Y. Cao, Y. Zhou and S. Lin, Small, 2025, 21, 2408147 Search PubMed.
- G. Park, C. Ettles, M. Charles and R. H. E. Hudson, J. Photoch. Photobio. A, 2023, 441, 114653 CrossRef CAS.
- K. Punia, D. Britton, K. Hüll, L. Yin, Y. Wang, P. D. Renfrew, M. L. Gilchrist, R. Bonneau, D. Trauner and J. K. Montclare, Soft Matter, 2023, 19, 497–501 Search PubMed.
- D. Chao, Y. Yang, X. Jia and E. B. Berda, Dyes Pigm., 2019, 167, 77–82 CrossRef CAS.
- H. Xiong, K. A. Alberto, J. Youn, J. Taura, J. Morstein, X. Li, Y. Wang, D. Trauner, P. A. Slesinger, S. O. Nielsen and Z. Qin, Nano Research, 2023, 16, 1033–1041 Search PubMed.
- G. Berkovic, V. Krongauz and V. Weiss, Chem. Rev., 2000, 100, 1741–1754 Search PubMed.
- R. Klajn, Chem. Soc. Rev., 2014, 43, 148–184 RSC.
- A. D. Pugachev, I. V. Ozhogin, M. B. Lukyanova, B. S. Lukyanov, I. A. Rostovtseva, I. V. Dorogan, N. I. Makarova, V. V. Tkachev, A. V. Metelitsa and S. M. Aldoshin, Spectrochim. Acta, Part A, 2020, 230, 118041 CrossRef CAS PubMed.
- Y. Wu, C. Yin, W. Zhang, J. Chao and F. Huo, Dyes Pigm., 2021, 193, 109467 Search PubMed.
- Z. Tian, W. Wu, W. Wan and A. D. Q. Li, J. Am. Chem. Soc., 2009, 131, 4245–4252 Search PubMed.
- G. Copley, J. G. Gillmore, J. Crisman, G. Kodis, C. L. Gray, B. R. Cherry, B. D. Sherman, P. A. Liddell, M. M. Paquette, L. Kelbauskas, N. L. Frank, A. L. Moore, T. A. Moore and D. Gust, J. Am. Chem. Soc., 2014, 136, 11994–12003 CrossRef CAS PubMed.
- Y. Xiong, A. Vargas Jentzsch, J. W. M. Osterrieth, E. Sezgin, I. V. Sazanovich, K. Reglinski, S. Galiani, A. W. Parker, C. Eggeling and H. L. Anderson, Chem. Sci., 2018, 9, 3029–3040 Search PubMed.
- T. Zhang, X.-Y. Lou, X. Li, X. Tu, J. Han, B. Zhao and Y.-W. Yang, Adv. Mater., 2023, 35, 2210551 CrossRef CAS PubMed.
- A. T. Frawley, V. Wycisk, Y. Xiong, S. Galiani, E. Sezgin, I. Urbancic, A. Vargas Jentzsch, K. G. Leslie, C. Eggeling and H. L. Anderson, Chem. Sci., 2020, 11, 8955–8960 RSC.
- A. T. Frawley, K. G. Leslie, V. Wycisk, S. Galiani, D. Shrestha, C. Eggeling and H. L. Anderson, ChemPhysChem, 2023, 24, e202300125 Search PubMed.
- H. Xia, K. Xie and G. Zou, Molecules, 2017, 22, 2236 Search PubMed.
- V. K. Seiler, N. Tumanov, K. Robeyns, J. Wouters, B. Champagne and T. Leyssens, Crystals, 2017, 7, 84 CrossRef.
- K. Schöller, S. Küpfer, L. Baumann, P. M. Hoyer, D. de Courten, R. M. Rossi, A. Vetushka, M. Wolf, N. Bruns and L. J. Scherer, Adv. Funct. Mater., 2014, 24, 5194–5201 Search PubMed.
- Y. Wang, T. Lan, N. Ji, Q. Meng and W. He, Dyes Pigm., 2025, 238, 112654 Search PubMed.
- C. Gong, P. Zhang, Y. Liu, R. Chen, K. Huang, W. Yang, L. Gao, M. Xu, X. Yang, Y. Liu, Q. Liu, B. Yan, B. Zhao and G. Jiang, J. Hazard. Mater., 2025, 493, 138343 Search PubMed.
- X. Tan, Y. Tang, T. Yang, G. Dai, C. Ye, J. Meng and F. Li, Anal. Chem., 2023, 95, 3309–3316 CrossRef CAS PubMed.
- J.-T. Ye, L. Wang, H.-Q. Wang, Z.-Z. Chen, Y.-Q. Qiu and H.-M. Xie, RSC Adv., 2017, 7, 642–650 RSC.
- M. Irie, Chem. Rev., 2000, 100, 1685–1716 CrossRef CAS PubMed.
- H. Tian and S. Yang, Chem. Soc. Rev., 2004, 33, 85–97 RSC.
- C. Shi, R. Zhu, K. Guo, L. Zhang, Z. Liu, X. Liu, Q. Ai and X. Hu, Adv. Opt. Mater., 2023, 11, 2301844 Search PubMed.
- T. Fukaminato, T. Hirose, T. Doi, M. Hazama, K. Matsuda and M. Irie, J. Am. Chem. Soc., 2014, 136, 17145–17154 CrossRef CAS PubMed.
- Z. Zheng, H. Hu, Z. Zhang, B. Liu, M. Li, D.-H. Qu, H. Tian, W.-H. Zhu and B. L. Feringa, Nat. Photonics, 2022, 16, 226–234 Search PubMed.
- T. Fukaminato, T. Doi, N. Tamaoki, K. Okuno, Y. Ishibashi, H. Miyasaka and M. Irie, J. Am. Chem. Soc., 2011, 133, 4984–4990 CrossRef CAS PubMed.
- J. Qi, C. Chen, X. Zhang, X. Hu, S. Ji, R. T. K. Kwok, J. W. Y. Lam, D. Ding and B. Z. Tang, Nat. Commun., 2018, 9, 1848 CrossRef PubMed.
- M. Estrader, J. Salinas Uber, L. A. Barrios, J. Garcia, P. Lloyd-Williams, O. Roubeau, S. J. Teat and G. Aromí, Angew. Chem., Int. Ed., 2017, 56, 15622–15627 CrossRef CAS PubMed.
- K.-X. Qin, Y.-S. Su, M.-Q. Zhu and C. Li, ChemBioChem, 2024, 25, e202400326 CrossRef CAS PubMed.
- X. Tan, Y. Wang, H. Li, Y. Duan, B. Wen, J. Zhao, H. Kim, J. Y. Lee, L. Zhou, H.-B. Cheng and J. Yoon, Small, 2025, 21, 2409027 CrossRef CAS PubMed.
- K.-H. Knauer and R. Gleiter, Angew. Chem., Int. Ed. Engl., 1977, 16, 113 CrossRef.
- K. Li, Y. Xiang, X. Wang, J. Li, R. Hu, A. Tong and B. Z. Tang, J. Am. Chem. Soc., 2014, 136, 1643–1649 CrossRef CAS PubMed.
- Z. Ye, H. Yu, W. Yang, Y. Zheng, N. Li, H. Bian, Z. Wang, Q. Liu, Y. Song, M. Zhang and Y. Xiao, J. Am. Chem. Soc., 2019, 141, 6527–6536 CrossRef CAS PubMed.
- H. Miyasaka, Y. Satoh, Y. Ishibashi, S. Ito, Y. Nagasawa, S. Taniguchi, H. Chosrowjan, N. Mataga, D. Kato, A. Kikuchi and J. Abe, J. Am. Chem. Soc., 2009, 131, 7256–7263 Search PubMed.
- Y. Kobayashi, T. Katayama, T. Yamane, K. Setoura, S. Ito, H. Miyasaka and J. Abe, J. Am. Chem. Soc., 2016, 138, 5930–5938 CrossRef CAS PubMed.
- S. Honda and T. Toyota, Nat. Commun., 2017, 8, 502 Search PubMed.
- Y. Liu, Y. Yang, D. Shi, M. Xiao, L. Jiang, J. Tian, G. Zhang, Z. Liu, X. Zhang and D. Zhang, Adv. Mater., 2019, 31, 1902576 Search PubMed.
- Z. Chen, Y. Sun, X. Wang, W. Zhang and Z. Zhang, Macromol. Rapid Commun., 2023, 44, 2300198 CrossRef CAS PubMed.
- Q. X. Hua, B. Xin, Z. J. Xiong, W. L. Gong, C. Li, Z. L. Huang and M. Q. Zhu, Chem. Commun., 2017, 53, 2669–2672 RSC.
- R. H. Mitchell, Eur. J. Org. Chem., 1999, 2695–2703 CrossRef CAS.
- R. H. Mitchell, T. R. Ward, Y. Chen, Y. Wang, S. A. Weerawarna, P. W. Dibble, M. J. Marsella, A. Almutairi and Z.-Q. Wang, J. Am. Chem. Soc., 2003, 125, 2974–2988 CrossRef CAS PubMed.
- K. Klaue, W. Han, P. Liesfeld, F. Berger, Y. Garmshausen and S. Hecht, J. Am. Chem. Soc., 2020, 142, 11857–11864 Search PubMed.
- S. Dubuis, A. Dellai, C. Courdurié, J. Owona, A. Kalafatis, L. Vellutini, E. Genin, V. Rodriguez and F. Castet, J. Am. Chem. Soc., 2023, 145, 10861–10871 CrossRef CAS PubMed.
- K. I. Willig, W. Wegner, A. Muller, V. Calvet-Fournier and H. Steffens, Cell Rep., 2021, 35, 109192 Search PubMed.
- X. Zhang, X. Chen, Z. Zeng, M. Zhang, Y. Sun, P. Xi, J. Peng and P. Xu, ACS Nano, 2015, 9, 2659–2667 CrossRef CAS PubMed.
- G. U. Nienhaus, Angew. Chem., Int. Ed., 2012, 51, 1312–1314 Search PubMed.
- R. Ando, H. Mizuno and A. Miyawaki, Science, 2004, 306, 1370–1373 CrossRef CAS PubMed.
- M. Andresen, M. C. Wahl, A. C. Stiel, F. Gräter, L. V. Schäfer, S. Trowitzsch, G. Weber, C. Eggeling, H. Grubmüller, S. W. Hell and S. Jakobs, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 13070–13074 CrossRef CAS PubMed.
- A. Acharya, A. M. Bogdanov, B. L. Grigorenko, K. B. Bravaya, A. V. Nemukhin, K. A. Lukyanov and A. I. Krylov, Chem. Rev., 2017, 117, 758–795 CrossRef CAS PubMed.
- S. Habuchi, R. Ando, P. Dedecker, W. Verheijen, H. Mizuno, A. Miyawaki and J. Hofkens, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 9511–9516 CrossRef CAS PubMed.
- M. Andresen, A. C. Stiel, J. Folling, D. Wenzel, A. Schonle, A. Egner, C. Eggeling, S. W. Hell and S. Jakobs, Nat. Biotechnol., 2008, 26, 1035–1040 CrossRef CAS PubMed.
- T. Brakemann, A. C. Stiel, G. Weber, M. Andresen, I. Testa, T. Grotjohann, M. Leutenegger, U. Plessmann, H. Urlaub, C. Eggeling, M. C. Wahl, S. W. Hell and S. Jakobs, Nat. Biotechnol., 2011, 29, 942–947 CrossRef CAS PubMed.
- D. M. Shcherbakova, O. M. Subach and V. V. Verkhusha, Angew. Chem., Int. Ed., 2012, 51, 10724–10738 CrossRef CAS PubMed.
- F. Pennacchietti, E. O. Serebrovskaya, A. R. Faro, I. I. Shemyakina, N. G. Bozhanova, A. A. Kotlobay, N. G. Gurskaya, A. Boden, J. Dreier, D. M. Chudakov, K. A. Lukyanov, V. V. Verkhusha, A. S. Mishin and I. Testa, Nat. Methods, 2018, 15, 601–604 CrossRef CAS PubMed.
- K. G. Chernov, T. A. Redchuk, E. S. Omelina and V. V. Verkhusha, Chem. Rev., 2017, 117, 6423–6446 Search PubMed.
- J. Yao, A. A. Kaberniuk, L. Li, D. M. Shcherbakova, R. Zhang, L. Wang, G. Li, V. V. Verkhusha and L. V. Wang, Nat. Methods, 2016, 13, 67–73 Search PubMed.
- L. Li, A. A. Shemetov, M. Baloban, P. Hu, L. Zhu, D. M. Shcherbakova, R. Zhang, J. Shi, J. Yao, L. V. Wang and V. V. Verkhusha, Nat. Commun., 2018, 9, 2734 CrossRef PubMed.
- J. Zhang, J. Wang and H. Tian, Mater. Horiz., 2014, 1, 169–184 Search PubMed.
- J.-L. Zhao, M.-H. Li, Y.-M. Cheng, X.-W. Zhao, Y. Xu, Z.-Y. Cao, M.-H. You and M.-J. Lin, Coord. Chem. Rev., 2023, 475, 214918 CrossRef CAS.
- P. Hong, N.-H. Xie, K. Xiong, J. Liu, M.-Q. Zhu and C. Li, J. Mater. Chem. A, 2023, 11, 5703–5713 RSC.
- A. A. Beharry, L. Wong, V. Tropepe and G. A. Woolley, Angew. Chem., Int. Ed., 2011, 50, 1325–1327 CrossRef CAS PubMed.
- K.-T. Chung, J. Environ. Sci. Health, Part C, 2016, 34, 233–261 CrossRef CAS PubMed.
- P. Hong, J. Liu, K.-X. Qin, R. Tian, L.-Y. Peng, Y.-S. Su, Z. Gan, X.-X. Yu, L. Ye, M.-Q. Zhu and C. Li, Angew. Chem., Int. Ed., 2024, 63, e202316706 CrossRef CAS PubMed.
- Z. Zhang, W. Wang, P. Jin, J. Xue, L. Sun, J. Huang, J. Zhang and H. Tian, Nat. Commun., 2019, 10, 4232 Search PubMed.
- Y. He, Z. Shangguan, Z.-Y. Zhang, M. Xie, C. Yu and T. Li, Angew. Chem., Int. Ed., 2021, 60, 16539–16546 CrossRef CAS PubMed.
- K. Uno, M. L. Bossi, M. Irie, V. N. Belov and S. W. Hell, J. Am. Chem. Soc., 2019, 141, 16471–16478 CrossRef CAS PubMed.
- T. I. Lansakara, F. Tong, C. J. Bardeen and A. V. Tivanski, Nano Lett., 2020, 20, 6744–6749 CrossRef CAS PubMed.
- L. Zhang, S. Jeong, J. Lee, J. Kim, J. S. Lee, J. Park, J. Hong, J.-H. Eom, H. Kim, Y. M. Rhee, H. Lee and H.-S. Lee, Small, 2024, 20, 2401480 CrossRef CAS PubMed.
- S. He, S. Schog, Y. Chen, Y. Ji, S. Panitz, W. Richtering and R. Göstl, Adv. Mater., 2023, 35, 2305845 CrossRef CAS PubMed.
- J. Kang, E. Li, L. Cui, Q. Shao, C. Yin and F. Cheng, Sens. Actuators B Chem., 2021, 327, 128941 CrossRef CAS.
- B. Roubinet, M. Weber, H. Shojaei, M. Bates, M. L. Bossi, V. N. Belov, M. Irie and S. W. Hell, J. Am. Chem. Soc., 2017, 139, 6611–6620 CrossRef CAS PubMed.
- K. Uno, A. Aktalay, M. L. Bossi, M. Irie, V. N. Belov and S. W. Hell, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2100165118 CrossRef CAS PubMed.
- S. H. Um, H. J. Kim, D. Kim, J. E. Kwon, J. W. Lee, D. Hwang, S. K. Kim and S. Y. Park, Dyes Pigm., 2018, 158, 36–41 CrossRef CAS.
- S. Qi, X. Lu, W. Mei, G. Gu, W. Li and A. Zhang, Nanoscale, 2023, 15, 18053–18067 RSC.
- H. Kong, X. Xie, Y. Bao, F. Zhang, L. Bian, K. Cheng, Y.-D. Zhao and J. Xia, Angew. Chem., Int. Ed., 2025, 64, e202419538 CrossRef CAS PubMed.
- Y. Zou, T. Yi, S. Xiao, F. Li, C. Li, X. Gao, J. Wu, M. Yu and C. Huang, J. Am. Chem. Soc., 2008, 130, 15750–15751 CrossRef CAS PubMed.
- S. Qiu, A. T. Frawley, K. G. Leslie and H. L. Anderson, Chem. Sci., 2023, 14, 9123–9135 RSC.
- J. Chen-Wu, D. B. Guzmán-Ríos, P. Remón, J. A. González-Delgado, A. J. Martínez-Martínez, F. Nájera, J. F. Arteaga and U. Pischel, Adv. Mater., 2023, 35, 2300536 CrossRef CAS PubMed.
- S. Song, H. Zhang and Y. Liu, Acc. Mater. Res., 2024, 5, 1109–1120 CrossRef CAS.
- W.-L. Guan, J.-F. Chen, J. Liu, B. Shi, H. Yao, Y.-M. Zhang, T.-B. Wei and Q. Lin, Coor. Chem. Rev., 2024, 507, 215717 CrossRef CAS.
- X. Y. Dai, Y. Y. Hu, Y. Sun, M. Huo, X. Dong and Y. Liu, Adv. Sci., 2022, e2200524 CrossRef PubMed.
- X. Y. Dai, Y. Y. Hu, Y. Sun, M. Huo, X. Dong and Y. Liu, Adv. Sci., 2022, e2200524 CrossRef PubMed.
- S. Wan, Z. Ma, C. Chen, F. Li, F. Wang, X. Jia, W. Yang and M. Yin, Adv. Funct. Mater., 2016, 26, 353–364 CrossRef CAS.
- T. Takeshita, A. Yano and M. Hara, ChemistrySelect, 2017, 2, 11288–11292 CrossRef CAS.
- D. Samanta, D. Galaktionova, J. Gemen, L. J. W. Shimon, Y. Diskin-Posner, L. Avram, P. Král and R. Klajn, Nat. Commun., 2018, 9, 641 CrossRef PubMed.
- J.-X. Wang, C. Li and H. Tian, Coord. Chem. Rev., 2021, 427, 213579 CrossRef CAS.
- L. Li, Y.-T. Yu, N.-N. Zhang, S.-H. Li, J.-G. Zeng, Y. Hua and H. Zhang, Coord. Chem. Rev., 2024, 500, 215526 CrossRef CAS.
- K. Hakouk, O. Oms, A. Dolbecq, J. Marrot, A. Saad, P. Mialane, H. El Bekkachi, S. Jobic, P. Deniard and R. Dessapt, J. Mater. Chem. C, 2014, 2, 1628–1641 RSC.
- C.-C. Ko and V. W.-W. Yam, Acc. Chem. Res., 2018, 51, 149–159 Search PubMed.
- S. Samanta, A. A. Beharry, O. Sadovski, T. M. McCormick, A. Babalhavaeji, V. Tropepe and G. A. Woolley, J. Am. Chem. Soc., 2013, 135, 9777–9784 Search PubMed.
- Z. Li, X. Zeng, C. Gao, J. Song, F. He, T. He, H. Guo and J. Yin, Coord. Chem. Rev., 2023, 497, 215451 Search PubMed.
- S. Lin, K. G. Gutierrez-Cuevas, X. Zhang, J. Guo and Q. Li, Adv. Funct. Mater., 2020, 31, 2007957 CrossRef.
- F. Li, M. Li, Y. Shi, X. Bian, N. Lv, S. Guo, Y. Wang, W. Zhao and W.-H. Zhu, Chem. Sci., 2025, 16, 14270–14277 RSC.
- M. Dong, A. Babalhavaeji, S. Samanta, A. A. Beharry and G. A. Woolley, Acc. Chem. Res., 2015, 48, 2662–2670 CrossRef CAS PubMed.
- C. Li, K. Xiong, Y. Chen, C. Fan, Y. L. Wang, H. Ye and M. Q. Zhu, ACS Appl. Mater. Interfaces, 2020, 12, 27651–27662 CrossRef CAS PubMed.
- S. Samanta, A. Babalhavaeji, M.-X. Dong and G. A. Woolley, Angew. Chem., Int. Ed., 2013, 52, 14127–14130 CrossRef CAS PubMed.
- M. Dong, A. Babalhavaeji, C. V. Collins, K. Jarrah, O. Sadovski, Q. Dai and G. A. Woolley, J. Am. Chem. Soc., 2017, 139, 13483–13486 CrossRef CAS PubMed.
- D. Wu, M. Dong, C. V. Collins, A. Babalhavaeji and G. A. Woolley, Adv. Opt. Mater., 2016, 4, 1402–1409 CrossRef CAS.
- A. A. Beharry and G. A. Woolley, Chem. Soc. Rev., 2011, 40, 4422–4437 RSC.
- S. Aiken, R. J. L. Edgar, C. D. Gabbutt, B. M. Heron and P. A. Hobson, Dyes Pigm., 2018, 149, 92–121 CrossRef CAS.
- V. A. Barachevsky, Rev. J. Chem., 2017, 7, 334–371 Search PubMed.
- M. M. Lerch, W. Szymański and B. L. Feringa, Chem. Soc. Rev., 2018, 47, 1910–1937 RSC.
- C. A. Reyes, H. J. Lee, C. Karanovic and E. Picazo, Nat. Commun., 2024, 15, 5533 CrossRef CAS PubMed.
- T. Senthilkumar, L. Zhou, Q. Gu, L. Liu, F. Lv and S. Wang, Angew. Chem., Int. Ed., 2018, 57, 13114–13119 CrossRef CAS PubMed.
- Z.-X. Zhang, F.-Q. Bai, L. Li and H.-X. Zhang, New J. Chem., 2015, 39, 1634–1642 Search PubMed.
- R. Li, B. Mou, M. Yamada, W. Li, T. Nakashima and T. Kawai, Molecules, 2024, 29, 2007957 Search PubMed.
- T. Fukaminato, M. Tanaka, L. Kuroki and M. Irie, Chem. Commun., 2008, 3924–3926 RSC.
- T. Yang, Q. Liu, J. Li, S. Pu, P. Yang and F. Li, RSC Adv., 2014, 4, 15613–15619 RSC.
- V. Marturano, J. Kozlowska, A. Bajek, M. Giamberini, V. Ambrogi, P. Cerruti, R. Garcia-Valls, J. M. Montornes and B. Tylkowski, Coord. Chem. Rev., 2019, 398, 213013 Search PubMed.
- J. Kim, S. Lee, Y. Jeong, K. Kim, K. Nam, H. Jin, Y. Choi, H.-J. Kim, H. Ryu, K. H. Kim, J.-I. Kim, J. Park, J. Joo and J.-H. Park, Adv. Mater., 2025, 37, 2502739 CrossRef CAS PubMed.
- X. Zhu, Q. Su, W. Feng and F. Li, Chem. Soc. Rev., 2017, 46, 1025–1039 Search PubMed.
- Y. Mi, H. B. Cheng, H. Chu, J. Zhao, M. Yu, Z. Gu, Y. Zhao and L. Li, Chem. Sci., 2019, 10, 10231–10239 Search PubMed.
- M. Zhao, B. Li, P. Wang, L. Lu, Z. Zhang, L. Liu, S. Wang, D. Li, R. Wang and F. Zhang, Adv. Mater., 2018, 30, e1804982 CrossRef PubMed.
- C. Benitez-Martin, S. Li, A. Dominguez-Alfaro, F. Najera, E. Pérez-Inestrosa, U. Pischel and J. Andréasson, J. Am. Chem. Soc., 2020, 142, 14854–14858 Search PubMed.
- G. S. He, L.-S. Tan, Q. Zheng and P. N. Prasad, Chem. Rev., 2008, 108, 1245–1330 Search PubMed.
- F. Helmchen and W. Denk, Nat. Methods, 2005, 2, 932–940 CrossRef CAS PubMed.
- M.-Q. Zhu, G.-F. Zhang, C. Li, M. P. Aldred, E. Chang, R. A. Drezek and A. D. Q. Li, J. Am. Chem. Soc., 2011, 133, 365–372 CrossRef CAS PubMed.
- K. Mutoh, K. Yamamoto and J. Abe, Photoch. Photobio. Sci., 2022, 21, 1445–1458 Search PubMed.
- L. Lin, Z. Zhang, Z. Lu, Y. Guo and M. Liu, J. Phys. Chem. A, 2016, 120, 7859–7864 CrossRef CAS PubMed.
- J. Moreno, M. Gerecke, L. Grubert, S. A. Kovalenko and S. Hecht, Angew. Chem., Int. Ed., 2016, 55, 1544–1547 CrossRef CAS PubMed.
- K. Uno, H. Niikura, M. Morimoto, Y. Ishibashi, H. Miyasaka and M. Irie, J. Am. Chem. Soc., 2011, 133, 13558–13564 CrossRef CAS PubMed.
- X. Hu, B. Fang, P. Li, J. Wang, J. Li, Y. Dai and M. Yin, Adv. Funct. Mater., 2025, 35, 2423793 CrossRef CAS.
- J. He, H. Zhao, H. Wu, Y. Yang, Z. Wang, Z. He and G. Jiang, Phys. Chem. Chem. Phys., 2021, 23, 17939–17944 RSC.
- Z. Xiong, X. Zhang, L. Liu, Q. Zhu, Z. Wang, H. Feng and Z. Qian, Chem. Sci., 2021, 12, 10710–10723 RSC.
- Y. Su, X. Li, D. Zheng, J. Andreásson, H. Wang, L. Yu, J. Chen, J. Ma and Y. Fang, Chem. Sci., 2025, 16, 23121–23128 Search PubMed.
- A. T. Muhammed Munthasir, P. Rani, P. Dhanalakshmi, S. Geremia, N. Hickey and P. Thilagar, Inorg. Chem., 2025, 64, 5878–5892 Search PubMed.
- H. Yang, M. Li, C. Li, Q. Luo, M. Q. Zhu, H. Tian and W. H. Zhu, Angew. Chem., Int. Ed., 2020, 59, 8560–8570 Search PubMed.
- C. Wang, Y. H. Liu and Y. Liu, Small, 2022, e2201821 Search PubMed.
- R. Zhang, Z. Lei, Z. Yu, Y. Chen and Y. Liu, Adv. Sci., 2025, 12, e07090 CrossRef CAS PubMed.
- L. Wu, C. Huang, B. P. Emery, A. C. Sedgwick, S. D. Bull, X.-P. He, H. Tian, J. Yoon, J. L. Sessler and T. D. James, Chem. Soc. Rev., 2020, 49, 5110–5139 Search PubMed.
- N. Asadi-Zaki, H. Mardani, H. Roghani-Mamaqani and F. Wang, Coord. Chem. Rev., 2024, 500, 215518 CrossRef CAS.
- Y. Xiong, P. Rivera-Fuentes, E. Sezgin, A. Vargas Jentzsch, C. Eggeling and H. L. Anderson, Org. Lett., 2016, 18, 3666–3669 Search PubMed.
- Z.-M. Yue, S.-M. Zhai, B.-Y. Wei, B.-X. Zhao and Z.-M. Lin, Dyes Pigm., 2025, 233, 112544 Search PubMed.
- J. Wu, L. Kreimendahl and J. L. Greenfield, J. Am. Chem. Soc., 2025, 147, 17549–17554 Search PubMed.
- C. Slavov, C. Yang, A. H. Heindl, H. A. Wegner, A. Dreuw and J. Wachtveitl, Angew. Chem., Int. Ed., 2020, 59, 380–387 Search PubMed.
- R. Yang, Y. Jiao, B. Wang, B. Xu and W. Tian, J. Phys. Chem. Lett., 2021, 12, 1290–1294 CrossRef CAS PubMed.
- S. Dutta Choudhury, Langmuir, 2022, 38, 14819–14826 CrossRef CAS PubMed.
- G. Das, T. Prakasam, N. Alkhatib, R. G. AbdulHalim, F. Chandra, S. K. Sharma, B. Garai, S. Varghese, M. A. Addicoat, F. Ravaux, R. Pasricha, R. Jagannathan, N. I. Saleh, S. Kirmizialtin, M. A. Olson and A. Trabolsi, Nat. Commun., 2023, 14, 3765 CrossRef CAS PubMed.
- Z. Ye, Z. Yang, L. Wang, L. Chen, Y. Cai, P. Deng, W. Feng, X. Li and L. Yuan, Angew. Chem., Int. Ed., 2019, 58, 12519–12523 CrossRef CAS PubMed.
- V. Dryza and E. J. Bieske, J. Phys. Chem. C, 2015, 119, 14076–14084 CrossRef CAS.
- T. Pan, D. Lu, H. Xin and B. Li, Light Sci. Appl., 2021, 10, 124 Search PubMed.
- H. Xu, D. Kim, Y.-Y. Zhao, C. Kim, G. Song, Q. Hu, H. Kang and J. Yoon, Adv. Mater., 2024, 36, 2402806 CrossRef CAS PubMed.
- J. Huang, J. Li, Y. Lyu, Q. Miao and K. Pu, Nat. Mater., 2019, 18, 1133–1143 CrossRef CAS PubMed.
- J. Du, S. Yang, Y. Qiao, H. Lu and H. Dong, Biosens Bioelectron., 2021, 191, 113478 Search PubMed.
- D. Hu, Z. Tian, W. Wu, W. Wan and A. D. Q. Li, J. Am. Chem. Soc., 2008, 130, 15279–15281 CrossRef CAS PubMed.
- L. Z. Cai, Q. S. Chen, C. J. Zhang, P. X. Li, M. S. Wang and G. C. Guo, J. Am. Chem. Soc., 2015, 137, 10882–10885 CrossRef CAS PubMed.
- S. Wang, W. X. Ren, J. T. Hou, M. Won, J. An, X. Chen, J. Shu and J. S. Kim, Chem. Soc. Rev., 2021, 50, 8887–8902 RSC.
- H. Li and J. C. Vaughan, Chem. Rev., 2018, 118, 9412–9454 CrossRef CAS PubMed.
- M. Koch and V. Ntziachristos, Annu. Rev. Med., 2016, 67, 153–164 Search PubMed.
- B. Huang, M. Bates and X. Zhuang, Annu. Rev. Biochem., 2009, 78, 993–1016 CrossRef CAS PubMed.
- Z. Yang, A. Sharma, J. Qi, X. Peng, D. Y. Lee, R. Hu, D. Lin, J. Qu and J. S. Kim, Chem. Soc. Rev., 2016, 45, 4651–4667 Search PubMed.
- Y. Xu, R. Xu, Z. Wang, Y. Zhou, Q. Shen, W. Ji, D. Dang, L. Meng and B. Z. Tang, Chem. Soc. Rev., 2021, 50, 667–690 Search PubMed.
- J. Kwon, M. S. Elgawish and S. H. Shim, Adv. Sci., 2022, 9, e2101817 CrossRef PubMed.
- H. Chang, M. Zhang, W. Ji, J. Chen, Y. Zhang, B. Liu, J. Lu, J. Zhang, P. Xu and T. Xu, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 4455–4460 Search PubMed.
- M. Minoshima and K. Kikuchi, J. Biol. Inorg. Chem., 2017, 22, 639–652 Search PubMed.
- B. Wang, M. Xiong, J. Susanto, X. Li, W.-Y. Leung and K. Xu, Angew. Chem., Int. Ed., 2022, 61, e202113612 CrossRef CAS PubMed.
- Q. Qi, W. Chi, Y. Li, Q. Qiao, J. Chen, L. Miao, Y. Zhang, J. Li, W. Ji, T. Xu, X. Liu, J. Yoon and Z. Xu, Chem. Sci., 2019, 10, 4914–4922 RSC.
- H.-l D. Lee, S. J. Lord, S. Iwanaga, K. Zhan, H. Xie, J. C. Williams, H. Wang, G. R. Bowman, E. D. Goley, L. Shapiro, R. J. Twieg, J. Rao and W. E. Moerner, J. Am. Chem. Soc., 2010, 132, 15099–15101 CrossRef CAS PubMed.
- Q. Qi, C. Li, X. Liu, S. Jiang, Z. Xu, R. Lee, M. Zhu, B. Xu and W. Tian, J. Am. Chem. Soc., 2017, 139, 16036–16039 CrossRef CAS PubMed.
- A. Bodén, D. Ollech, A. G. York, A. Millett-Sikking and I. Testa, Nat. Methods, 2024, 21, 882–888 Search PubMed.
- S. Wang, X. Chen, L. Chang, R. Xue, H. Duan and Y. Sun, ACS Nano, 2016, 10, 9136–9144 CrossRef CAS PubMed.
- M.-Q. Zhu, G.-F. Zhang, Z. Hu, M. P. Aldred, C. Li, W.-L. Gong, T. Chen, Z.-L. Huang and S. Liu, Macromolecules, 2014, 47, 1543–1552 CrossRef CAS.
- Y. Kim, J. Gonzales and Y. Zheng, Small, 2021, 17, e2004988 CrossRef PubMed.
- C. Xu and K. Pu, Nat. Rev. Bioeng., 2024, 2, 425–441 CrossRef CAS.
- M. Li, J. Zhao, H. Chu, Y. Mi, Z. Zhou, Z. Di, M. Zhao and L. Li, Adv. Mater., 2019, 31, 1804745 CrossRef CAS PubMed.
- D. Yi, J. Zhao and L. Li, Angew. Chem., Int. Ed., 2021, 60, 6300–6304 CrossRef CAS PubMed.
- W. Yu, H. Jin, C. Tang, J. Du and Z. Zhang, Br. J. Pharmacol., 2018, 175, 1114–1125 Search PubMed.
- H. Zhao, G. Patterson and P. Schuck, Biophys. J., 2016, 110, 347a Search PubMed.
- W. N. Harrington, M. V. Novoselova, D. N. Bratashov, B. N. Khlebtsov, D. A. Gorin, E. I. Galanzha and V. P. Zharov, Sci. Rep., 2019, 9, 12439 Search PubMed.
- Z. Qiao, H. Qi, H. Zhang, Q. Zhou, N. Wei, Y. Zhang and K. Wang, Anal. Chem., 2020, 92, 1934–1939 Search PubMed.
- N. Wang, J. Wang, S. Huang, C. Zheng, S. Cui and S. Pu, Anal. Chim. Acta, 2026, 1382, 344848 CrossRef CAS PubMed.
- Y. Wu, Z. Gong, X. Wu, Y. Huang, L. Sun, H. Ding, Y.-L. Zeng, C. Fan, G. Liu and S. Pu, Anal. Chim. Acta, 2025, 1359, 344114 Search PubMed.
- W. Zhang, F. Huo and C. Yin, Org. Lett., 2019, 21, 5277–5280 CrossRef CAS PubMed.
- W. Zhang, F. Huo, Y. Yue, Y. Zhang, J. Chao, F. Cheng and C. Yin, J. Am. Chem. Soc., 2020, 142, 3262–3268 CrossRef CAS PubMed.
- Y. Hong, P. Zhang, H. Wang, M. Yu, Y. Gao and J. Chen, Sens. Actuators, B, 2018, 272, 340–347 CrossRef CAS.
- Y. Fu, X. Zhang, J. Liu, G. Qian, Z. P. Xu and R. Zhang, J. Mater. Chem. B, 2022, 10, 3366–3374 Search PubMed.
- X. Xie, G. Mistlberger and E. Bakker, J. Am. Chem. Soc., 2012, 134, 16929–16932 CrossRef CAS PubMed.
- X. Chai, H. H. Han, Y. Zang, J. Li, X. P. He, J. Zhang and H. Tian, Beilstein J. Org. Chem., 2019, 15, 2380–2389 CrossRef CAS PubMed.
- Y. Fu, H. H. Han, J. Zhang, X. P. He, B. L. Feringa and H. Tian, J. Am. Chem. Soc., 2018, 140, 8671–8674 CrossRef CAS PubMed.
- J. Zhang, Y. Fu, H. H. Han, Y. Zang, J. Li, X. P. He, B. L. Feringa and H. Tian, Nat. Commun., 2017, 8, 987 CrossRef PubMed.
- M. Yu, W. Zhao, F. Ni, Q. Zhao and C. Yang, Adv. Opt. Mater., 2022, 10, 2102437 Search PubMed.
- F. Bierbuesse, A. C. Bourges, V. Gielen, V. Mönkemöller, W. Vandenberg, Y. Shen, J. Hofkens, P. Vanden Berghe, R. E. Campbell, B. Moeyaert and P. Dedecker, Nat. Commun., 2022, 13, 1850 Search PubMed.
- L. Liu, S. Wang, C.-H. Zhang, J. Jiang, Y. Gao, P. Zhang, R. Zeng and J. Chen, Dyes Pigm., 2021, 191, 109370 Search PubMed.
- M. Bates, B. Huang, G. T. Dempsey and X. Zhuang, Science, 2007, 317, 1749–1753 Search PubMed.
- M. Deng, Y. Zhang, Y. Wang and S. Jiang, J. Mater. Chem. C, 2020, 8, 15697–15704 Search PubMed.
- W. Zhong, X. Zeng, J. Chen, Y. Hong, L. Xiao and P. Zhang, Polym. Chem., 2017, 8, 4849–4855 RSC.
- Q. Yan, Z. Qiao, W. Zhao, J. Ren, Y. Wang, W. Yang and S. Wang, J. Mater. Chem. C, 2022, 10, 7024–7030 Search PubMed.
- J. Chen, W. Zhong, Y. Tang, Z. Wu, Y. Li, P. Yi and J. Jiang, Macromolecules, 2015, 48, 3500–3508 CrossRef CAS.
- R. Yang, X. Ren, L. Mei, G. Pan, X.-Z. Li, Z. Wu, S. Zhang, W. Ma, W. Yu, H.-H. Fang, C. Li, M.-Q. Zhu, Z. Hu, T. Sun, B. Xu and W. Tian, Angew. Chem., Int. Ed., 2022, 61, e202117158 CrossRef CAS PubMed.
- L. Wei, Z. Chen, L. Shi, R. Long, A. V. Anzalone, L. Zhang, F. Hu, R. Yuste, V. W. Cornish and W. Min, Nature, 2017, 544, 465–470 CrossRef CAS PubMed.
- R. Chouket, A. Pellissier-Tanon, A. Lemarchand, A. Espagne, T. Le Saux and L. Jullien, Chem. Sci., 2020, 11, 2882–2887 RSC.
- T. Niehörster, A. Löschberger, I. Gregor, B. Krämer, H.-J. Rahn, M. Patting, F. Koberling, J. Enderlein and M. Sauer, Nat. Methods, 2016, 13, 257–262 CrossRef PubMed.
- A. M. Valm, S. Cohen, W. R. Legant, J. Melunis, U. Hershberg, E. Wait, A. R. Cohen, M. W. Davidson, E. Betzig and J. Lippincott-Schwartz, Nature, 2017, 546, 162–167 CrossRef CAS PubMed.
- G. Marriott, S. Mao, T. Sakata, J. Ran, D. K. Jackson, C. Petchprayoon, T. J. Gomez, E. Warp, O. Tulyathan, H. L. Aaron, E. Y. Isacoff and Y. Yan, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 17789–17794 CrossRef CAS PubMed.
- J. Querard, T.-Z. Markus, M.-A. Plamont, C. Gauron, P. Wang, A. Espagne, M. Volovitch, S. Vriz, V. Croquette, A. Gautier, T. Le Saux and L. Jullien, Angew. Chem., Int. Ed., 2015, 54, 2633–2637 CrossRef CAS PubMed.
- J.-C. Hsiang, A. E. Jablonski and R. M. Dickson, Acc. Chem. Res., 2014, 47, 1545–1554 CrossRef CAS PubMed.
- Y.-C. Chen, A. E. Jablonski, I. Issaeva, D. Bourassa, J.-C. Hsiang, C. J. Fahrni and R. M. Dickson, J. Am. Chem. Soc., 2015, 137, 12764–12767 CrossRef CAS PubMed.
- J. Quérard, R. Zhang, Z. Kelemen, M.-A. Plamont, X. Xie, R. Chouket, I. Roemgens, Y. Korepina, S. Albright, E. Ipendey, M. Volovitch, H. L. Sladitschek, P. Neveu, L. Gissot, A. Gautier, J.-D. Faure, V. Croquette, T. Le Saux and L. Jullien, Nat. Commun., 2017, 8, 969 CrossRef PubMed.
- R. Zhang, R. Chouket, M.-A. Plamont, Z. Kelemen, A. Espagne, A. G. Tebo, A. Gautier, L. Gissot, J.-D. Faure, L. Jullien, V. Croquette and T. Le Saux, Light Sci. Appl., 2018, 7, 97 CrossRef PubMed.
- R. Zhang, R. Chouket, A. Tebo, M.-A. Plamont, Z. Kelemen, L. Gissot, J.-D. Faure, A. Gautier, V. Croquette, L. Jullien and T. Saux, Optica, 2019, 6, 972–980 CrossRef CAS.
- H. Valenta, S. Hugelier, S. Duwé, G. Lo Gerfo, M. Müller, P. Dedecker and W. Vandenberg, Biophys. Rep., 2021, 1, 100026 CAS.
- H. Valenta, F. Bierbuesse, R. Vitale, C. Ruckebusch, W. Vandenberg and P. Dedecker, Talanta, 2024, 269, 125397 CrossRef CAS PubMed.
- R. Chouket, A. Pellissier-Tanon, A. Lahlou, R. Zhang, D. Kim, M.-A. Plamont, M. Zhang, X. Zhang, P. Xu, N. Desprat, D. Bourgeois, A. Espagne, A. Lemarchand, T. L. Saux and L. Jullien, Nat. Commun., 2022, 13, 1482 CrossRef CAS PubMed.
- T. Roebroek, W. Vandenberg, F. Sipieter, S. Hugelier, C. Stove, J. Zhang and P. Dedecker, Nat. Commun., 2021, 12, 2005 CrossRef CAS PubMed.
- B. Chang, D. Li, Y. Ren, C. Qu, X. Shi, R. Liu, H. Liu, J. Tian, Z. Hu, T. Sun and Z. Cheng, Nat. Biomed. Eng., 2022, 6, 629–639 Search PubMed.
- W. Zhao, Z. He and B. Z. Tang, Nat. Rev. Mater., 2020, 5, 869–885 CrossRef CAS.
- J. Huang, L. Su, C. Xu, X. Ge, R. Zhang, J. Song and K. Pu, Nat. Mater., 2023, 22, 1421–1429 CrossRef PubMed.
- C. Xu, J. Huang, Y. Jiang, S. He, C. Zhang and K. Pu, Nat. Biomed. Eng., 2023, 7, 298–312 CrossRef CAS PubMed.
- L. Yang, M. Zhao, W. Chen, J. Zhu, W. Xu, Q. Li, K. Pu and Q. Miao, Angew. Chem., Int. Ed., 2024, 63, e202313117 CrossRef CAS PubMed.
- F. Xiao, H. Gao, Y. Lei, W. Dai, M. Liu, X. Zheng, Z. Cai, X. Huang, H. Wu and D. Ding, Nat. Commun., 2022, 13, 186 CrossRef CAS PubMed.
- X.-Y. Dai, M. Huo and Y. Liu, Nat. Rev. Chem., 2023, 7, 854–874 CrossRef CAS PubMed.
- Y.-Y. Hu, X.-Y. Dai, X. Dong, M. Huo and Y. Liu, Angew. Chem., Int. Ed., 2022, 61, e202213097 CrossRef CAS.
- Y. Zhang, C. Zhang, Y. Chen, J. Yu, L. Chen, H. Zhang, X. Xu and Y. Liu, Adv. Opt. Mater., 2022, 10 Search PubMed.
- H.-J. Wang, W.-W. Xing, H.-Y. Zhang, W.-W. Xu and Y. Liu, Adv. Opt. Mater., 2022, 10, 2201178 Search PubMed.
- X. L. Dean-Ben, S. Gottschalk, B. Mc Larney, S. Shoham and D. Razansky, Chem. Soc. Rev., 2017, 46, 2158–2198 Search PubMed.
- C. Liu, X. Gong, R. Lin, F. Liu, J. Chen, Z. Wang, L. Song and J. Chu, Theranostics, 2016, 6, 2414–2430 CrossRef CAS PubMed.
- A. B. E. Attia, G. Balasundaram, M. Moothanchery, U. S. Dinish, R. Bi, V. Ntziachristos and M. Olivo, Photoacoustics, 2019, 16, 100144 CrossRef PubMed.
- H. B. Cheng, Y. Li, B. Z. Tang and J. Yoon, Chem. Soc. Rev., 2020, 49, 21–31 RSC.
- L. A. Kasatkina, C. Ma, M. E. Matlashov, T. Vu, M. Li, A. A. Kaberniuk, J. Yao and V. V. Verkhusha, Nat. Commun., 2022, 13, 2813 CrossRef CAS PubMed.
- S. Chen, K. Li, X. Chen, S. Lei, J. Lin and P. Huang, Sci. Adv., 2024, 10, eadn8274 CrossRef CAS PubMed.
- K. Mishra, M. Stankevych, P. Fuenzalida-Werner Juan, S. Grassmann, V. Gujrati, Y. Huang, U. Klemm, R. Buchholz Veit, V. Ntziachristos and C. Stiel Andre, Sci. Adv., 2020, 6, eaaz6293 Search PubMed.
- R. Gao, F. Liu, W. Liu, S. Zeng, J. Chen, R. Gao, L. Wang, C. Fang, L. Song, A. C. Sedgwick, J. L. Sessler, J. Chu, F. Yan and C. Liu, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2121982119 CrossRef CAS.
- C. Liu, X. Zheng, T. Dai, H. Wang, X. Chen, B. Chen, T. Sun, F. Wang, S. Chu and J. Rao, Angew. Chem., Int. Ed., 2022, 61, e202116802 CrossRef CAS PubMed.
- S. He, J. Song, J. Liu, L. Liu, J. Qu and Z. Cheng, Adv. Opt. Mater., 2019, 7, 1900045 CrossRef.
- H. M. D. Bandara and S. C. Burdette, Chem. Soc. Rev., 2012, 41, 1809–1825 Search PubMed.
- J. Garcia-Amorós, B. Maerz, M. Reig, A. Cuadrado, L. Blancafort, E. Samoylova and D. Velasco, Chem. Eur. J., 2019, 25, 7726–7732 CrossRef PubMed.
- J. Levi, S. R. Kothapalli, T.-J. Ma, K. Hartman, B. T. Khuri-Yakub and S. S. Gambhir, J. Am. Chem. Soc., 2010, 132, 11264–11269 CrossRef CAS PubMed.
- M. Weiger and K. P. Pruessmann, Prog. Nucl. Magn. Reson. Spectrosc., 2019, 114–115, 237–270 CrossRef CAS PubMed.
- C. Fu, Y. Yu, X. Xu, Q. Wang, Y. Chang, C. Zhang, J. Zhao, H. Peng and A. K. Whittaker, Prog. Polym. Sci., 2020, 108, 101286 CrossRef CAS.
- D. Hao, T. Ai, F. Goerner, X. Hu, V. M. Runge and M. Tweedle, Magn. Reson. Imaging, 2012, 36, 1060–1071 CrossRef PubMed.
- J. Wahsner, E. M. Gale, A. Rodríguez-Rodríguez and P. Caravan, Chem. Rev., 2019, 119, 957–1057 CrossRef CAS.
- I. M. Welleman, C. L. F. van Beek, I. Belcin, A. M. Schulte, R. A. J. O. Dierckx, B. L. Feringa, H. H. Boersma and W. Szymański, Smart Mol., 2024, 2, e20230029 CrossRef CAS PubMed.
- C. Tu and A. Y. Louie, Chem. Commun., 2007, 1331–1333, 10.1039/b616991k.
- C. Tu, E. A. Osborne and A. Y. Louie, Tetrahedron, 2009, 65, 1241 CrossRef CAS PubMed.
- K. Kruttwig, D. R. Yankelevich, C. Brueggemann, C. Tu, N. L'Etoile, A. Knoesen and A. Y. Louie, Molecules, 2012, 17, 6605–6624 CrossRef CAS PubMed.
- M. Gao, B. Shen, J. Zhou, R. Kapre, A. Y. Louie and J. T. Shaw, ACS Omega, 2020, 5, 14759–14766 CrossRef CAS.
- B. Shen, M. Gao, F. C. Franco, Jr., R. Kapre, J. Zhou, X. Li, J. Garcia, J. T. Shaw and A. Y. Louie, J. Org. Chem., 2020, 85, 7333–7341 CrossRef CAS PubMed.
- S. Thies, H. Sell, C. Schutt, C. Bornholdt, C. Nather, F. Tuczek and R. Herges, J. Am. Chem. Soc., 2011, 133, 16243–16250 CrossRef CAS PubMed.
- G. Heitmann, C. Schütt, J. Gröbner, L. Huber and R. Herges, Dalton Trans., 2016, 45, 11407–11412 RSC.
- V. Wellm, J. Groebner, G. Heitmann, F. D. Sonnichsen and R. Herges, Angew. Chem., Int. Ed., 2021, 60, 8220–8226 CrossRef CAS PubMed.
- V. Wellm, C. Nather and R. Herges, J. Org. Chem., 2021, 86, 9503–9514 CrossRef CAS PubMed.
- Y. Heta, K. Kumaki, H. Hifumi, D. Citterio, A. Tanimoto and K. Suzuki, Photochem. Photobiol., 2012, 88, 876–883 CrossRef CAS PubMed.
- E. A. Osborne, B. R. Jarrett, C. Tu and A. Y. Louie, J. Am. Chem. Soc., 2010, 132, 5934–5935 CrossRef CAS PubMed.
- Q. Jin, X. Fan, C. Chen, L. Huang, J. Wang and X. Tang, Anal. Chem., 2019, 91, 3784–3789 CrossRef CAS PubMed.
- S. Stewart, R. J. Priore, M. P. Nelson and P. J. Treado, Annu. Rev. Anal. Chem., 2012, 5, 337–360 CrossRef CAS.
- R. C. Prince, R. R. Frontiera and E. O. Potma, Chem. Rev., 2017, 117, 5070–5094 CrossRef CAS PubMed.
- Y. Tang, X. Chen, S. Zhang, Z. J. Smith and T. Gao, Anal. Chem., 2021, 93, 15659–15666 CrossRef CAS PubMed.
- D. Lee, C. Qian, H. Wang, L. Li, K. Miao, J. Du, D. M. Shcherbakova, V. V. Verkhusha, L. V. Wang and L. Wei, J. Chem. Phys., 2021, 154, 135102 CrossRef CAS PubMed.
- J. Shou, A. Komazawa, Y. Wachi, M. Kawatani, H. Fujioka, S. J. Spratt, T. Mizuguchi, K. Oguchi, H. Akaboshi, F. Obata, R. Tachibana, S. Yasunaga, Y. Mita, Y. Misawa, R. Kojima, Y. Urano, M. Kamiya and Y. Ozeki, Sci. Adv., 2023, 9, eade9118 CrossRef CAS PubMed.
- Y. Yang, X. Bai and F. Hu, Nat. Commun., 2024, 15, 2578 CrossRef CAS PubMed.
- Y. Tang, X. Chen, S. Zhang, Z. J. Smith and T. Gao, Anal. Chem., 2021, 93, 15659–15666 CrossRef CAS.
- Y. Tang, Y. Zhuang, S. Zhang, Z. J. Smith, Y. Li, X. Mu, M. Li, C. He, X. Zheng, F. Pan, T. Gao and L. Zhang, ACS Cent. Sci., 2021, 7, 768–780 CrossRef CAS PubMed.
- Y. Shen, G. Xue, Y. Dai, S. M. Quintero, H. Chen, D. Wang, F. Miao, F. Negri, Y. Zheng and J. Casado, Nat. Commun., 2021, 12, 6262 CrossRef CAS PubMed.
- A. A. Ali, R. Kharbash and Y. Kim, Anal. Chim. Acta, 2020, 1110, 199–223 CrossRef CAS PubMed.
- M. Du and C. Li, Adv. Mater., 2024, 36, 2408484 CrossRef CAS PubMed.
- H.-W. Chien, C.-H. Yang, M.-T. Tsai and T.-L. Wang, J. Photochem. Photobiol., A, 2020, 392, 112303 CrossRef CAS.
- R. S. Gormal, P. Padmanabhan, R. Kasula, A. T. Bademosi, S. Coakley, J. Giacomotto, A. Blum, M. Joensuu, T. P. Wallis, H. P. Lo, S. Budnar, J. Rae, C. Ferguson, M. Bastiani, W. G. Thomas, E. Pardon, J. Steyaert, A. S. Yap, G. J. Goodhill, M. A. Hilliard, R. G. Parton and F. A. Meunier, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 30476–30487 Search PubMed.
- F. Pennacchietti, J. Alvelid, R. A. Morales, M. Damenti, D. Ollech, O. S. Oliinyk, D. M. Shcherbakova, E. J. Villablanca, V. V. Verkhusha and I. Testa, Nat. Commun., 2023, 14, 8402 Search PubMed.
- K. H. Rainey and G. H. Patterson, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 864–873 Search PubMed.
- J. Lee, S. Lai, S. Yang, S. Zhao, F. A. Blanco, A. C. Lyons, R. Merino-Urteaga, J. F. Ahrens, N. A. Nguyen, H. Liu, Z. Liu, G. G. Lambert, N. C. Shaner, L. Chen, K. F. Tolias, J. Zhang, T. Ha and F. St-Pierre, Nat. Commun., 2025, 16, 3241 Search PubMed.
- V. Adam, K. Hadjidemetriou, N. Jensen, R. L. Shoeman, J. Woodhouse, A. Aquila, A.-S. Banneville, T. R. M. Barends, V. Bezchastnov, S. Boutet, M. Byrdin, M. Cammarata, S. Carbajo, N. Eleni Christou, N. Coquelle, E. De la Mora, M. El Khatib, T. Moreno Chicano, R. Bruce Doak, F. Fieschi, L. Foucar, O. Glushonkov, A. Gorel, M. L. Grünbein, M. Hilpert, M. Hunter, M. Kloos, J. E. Koglin, T. J. Lane, M. Liang, A. Mantovanelli, K. Nass, G. Nass Kovacs, S. Owada, C. M. Roome, G. Schirò, M. Seaberg, M. Stricker, M. Thépaut, K. Tono, K. Ueda, L. M. Uriarte, D. You, N. Zala, T. Domratcheva, S. Jakobs, M. Sliwa, I. Schlichting, J.-P. Colletier, D. Bourgeois and M. Weik, ChemPhysChem, 2022, 23, e202200192 CrossRef CAS PubMed.
- P. Vetschera, K. Mishra, J. P. Fuenzalida-Werner, A. Chmyrov, V. Ntziachristos and A. C. Stiel, Anal. Chem., 2018, 90, 10527–10535 CrossRef CAS.
- Q. Qi, Y. Liu, V. Puranik, S. Patra, Z. Svindrych, X. Gong, Z. She, Y. Zhang and I. Aprahamian, J. Am. Chem. Soc., 2025, 147, 16404–16411 CrossRef CAS PubMed.
- G.-e Go, U. Jeong, H. Park, S. Go and D. Kim, Angew. Chem., Int. Ed., 2024, 63, e202405246 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |