Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Radical enzymatic peptide cyclization in natural product biosynthesis
Ziwei
Yao
and
Brandon I.
Morinaka
 *
*
Department of Pharmacy and Pharmaceutical Sciences, National University of Singapore, Singapore 117544, Singapore. E-mail: phambi@nus.edu.sg
First published on 16th February 2026
Abstract
Cyclic peptides are privileged scaffolds in natural product biosynthesis and drug discovery, valued for their structural diversity, metabolic stability, and potent biological activities. Radical-mediated enzymatic cyclization has emerged as a powerful biosynthetic strategy for constructing structurally complex and bioactive peptide natural products. This review surveys the growing repertoire of radical enzymes, including radical S-adenosylmethionine (rSAM) enzymes, cytochrome P450s, BURP-domain oxidases, and additional enzyme families that catalyze site-selective C–C, C–S, C–N, and C–O bond formation during peptide macrocyclization. Emphasis is placed on radical-driven cyclization reactions involved in natural product biosynthesis, covering both RiPPs and NRPS pathways. Examples of cyclic peptide natural products include vancomycin, arylomicin, telocidin, streptide, and nosiheptide illustrate the mechanistic and structural diversity of radical peptide cyclases. Key emerging themes include a diverse array of side-chain crosslinking, 3–4 residue macrocycles, and unique forms of chirality. This review also discusses advances in genome mining and mechanistic enzymology that continue to reveal new radical transformations and unusual enzyme mechanisms. Collectively, these insights highlight the role of radical enzymology in expanding the structural space of peptide-based natural products and its potential for applications in synthetic biology, drug discovery, and peptide engineering.
Introduction
Cyclic peptides represent one of the most chemically diverse and biologically significant classes of molecules in nature.1–5 These compounds are biosynthesized by various organisms including microorganisms, plants, and animals. They function as tools for communication, signaling, and defense.6 The structural complexity of peptide natural products arise from the enzymatic machinery that tailor their biosynthesis, yielding a vast array of scaffolds with unique functional groups and stereochemical features.7,8 Beyond their ecological roles, cyclic peptides have profoundly influenced drug discovery, providing templates for antibiotics (e.g., vancomycin), food preservative (e.g. nisin), anticancer agents (e.g., romidepsin, and immunosuppressants (e.g., cyclosporine).9–18 Their unparalleled chemical diversity vs their synthetic variations make cyclic peptide natural products a continuous source of inspiration for both synthetic chemists and pharmaceutical researchers.19,20The biosynthesis of peptide natural products generally occurs by two pathways. These include NRPs and RiPPs, both of which exhibit exceptional structural diversity and biological activity.2,21–23 In NRP biosynthesis, cyclization typically occurs during or immediately following peptide elongation, often catalyzed by specialized thioesterase domains or stand-alone cyclases that facilitate macrocyclization through ester, amide, or thioether bond formation.24–27 In contrast, RiPP cyclization generally takes place post-translationally, after the precursor peptide has been ribosomally produced and recognized by leader peptide-binding domains. This recognition enables modification enzymes such as radical S-adenosylmethionine (rSAM) enzymes, lanthionine synthetases, or P450s to install diverse intramolecular crosslinks that stabilize the mature core peptide structure.28–30 Among peptide-based natural products, cyclic peptides are especially notable for their macrocyclic frameworks, which confer enhanced stability, proteolytic resistance, and precise molecular interactions.1,31,32 Clinically important examples such as vancomycin and daptomycin underscore their therapeutic relevance, particularly in their widespread use as antibiotics.5 The chemical diversity of cyclic peptides is further expanded through post-translational modifications, including cross-links such as C–C, C–S, C–N and C–O bonds, which are critical for their bioactivity and structural rigidity.21,33–35
Two general strategies exist in nature to cyclize peptides, those that cyclize through paired electron mechanisms such as the formation of an amide or ester linkages by hydrolytic enzymes such as ligases or dehydration and cyclization by lanthionine synthetases. Alternatively, cyclization can occur by radical based mechanisms and predominantly involve crosslinking of two sidechains. The formation of side chain crosslinks involve chemistry at unactivated positions and therefore require higher energy intermediates to overcome the energy of activation. Unlike N- to C-terminal cyclization mechanisms, which typically rely on paired electron mechanisms, radical-mediated cyclization employs highly reactive radical intermediates to form new bonds.36,37 These intermediates, characterized by unpaired electrons, are transient and inherently unstable, requiring precise enzymatic control to direct their reactivity.30 Enzymes catalyzing radical-mediated peptide cyclization have evolved to stabilize and guide these reactive intermediates, enabling the formation of structurally complex and chemically diverse cyclic peptides that are often challenging to access by chemical synthesis.
Radical-mediated cyclization offers several distinct advantages over conventional enzymatic pathways. It enables the formation of unique chemical linkages that contribute to the structural novelty and functional diversity of cyclic peptides. The creation of unconventional macrocyclic architectures are the basis for new drugs with new modes of action. Interesting topologies including bridged and polycyclic systems have continued to be identified.38 Additionally, radical cyclization pathways promote stereochemical diversity, generating chiral centers and unconventional types of chirality that are difficult to achieve synthetically.37 The remarkable transformations achieved by radical enzymes are driven by finely evolved catalytic mechanisms. These enzymes often utilize cofactors to generate and stabilize radical intermediates, while their active sites are precisely designed to guide the radical species toward specific substrates, minimizing side reactions and ensuring regio- and stereoselectivity.39,40 The discovery and mechanistic understanding of the enzymes have not only expanded our understanding of radical chemistry in biological systems but also underscored their critical role in diversifying natural product scaffolds.
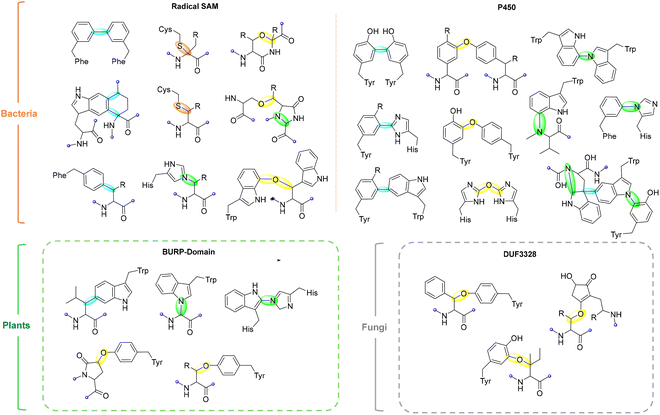
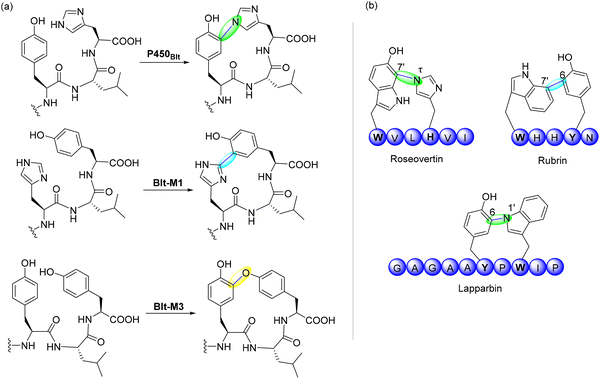
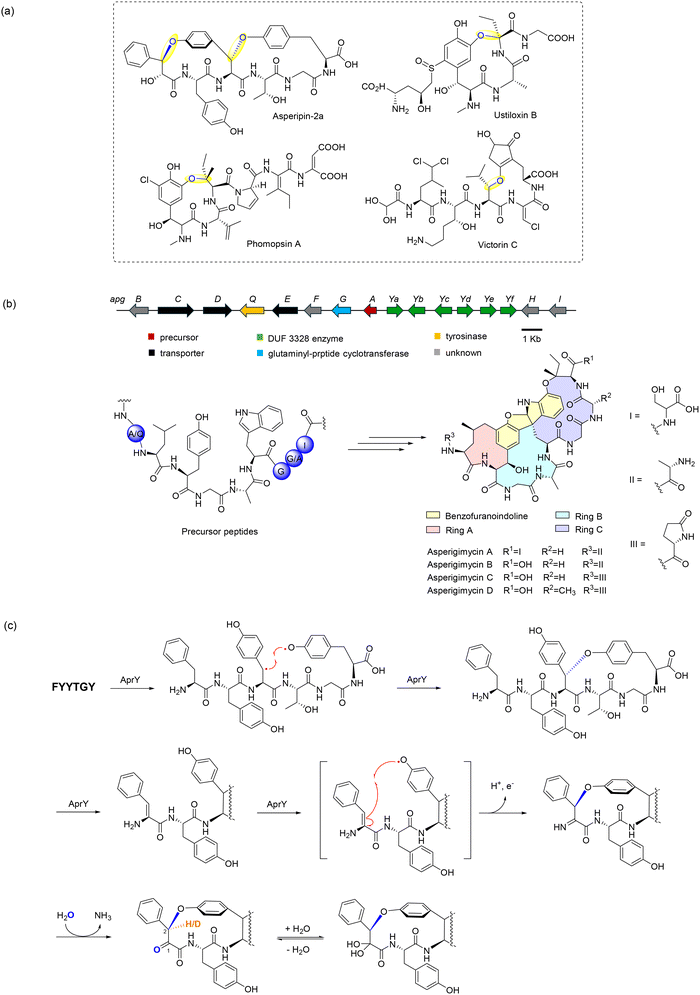
This review examines radical-mediated cyclization in peptides, focusing specifically on pathways where new covalent bonds form between amino acid residues, including backbone-to-backbone, backbone-to-side chain, and side chain-to-side chain linkages (Fig. 1). Cyclizations involving non-amino acid moieties or chemically modified side chains are excluded. We have also excluded examples of cyclization on a single residue although several examples have been reported.41–43 We highlight the broad landscape of radical peptide cyclization enzymes, which include bacterial rSAM enzymes, microbial cytochrome P450s, plant BURP-domain proteins, and fungal DUF-domain oxidases. Unraveling how these enzymes orchestrate radical-mediated transformations will allow us to harness their potential for the rational engineering of cyclic peptides, offering exciting opportunities for drug discovery and synthetic biology. The unique chemical transformations enabled by radical peptide cyclization not only enrich the structural diversity of cyclic peptides but also provide a valuable framework for designing novel therapeutic agents with enhanced bioactivity and stability.
1. Radical SAM enzymes
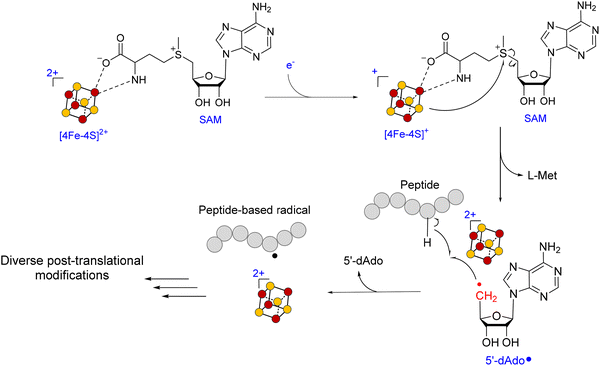
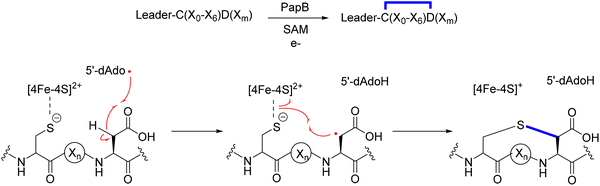
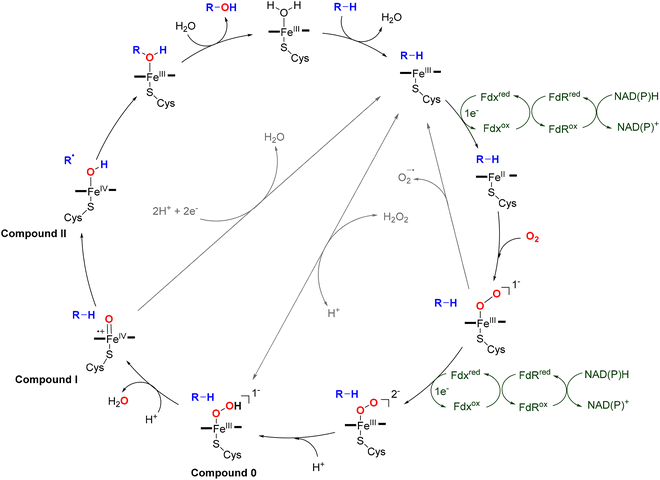
Radical S-adenosylmethionine (rSAM) enzymes constitute a vast and functionally diverse superfamily that catalyzes chemically challenging transformations across numerous biological pathways.44–50 These enzymes are characterized by a conserved CX3CX2C motif (rSAM domain) at their active site, which coordinates a [4Fe–4S] cluster essential for catalysis.46,51 Catalysis is initiated by a single-electron transfer from the [4Fe–4S] cluster to a bound SAM cofactor, triggering the reductive cleavage of the 5′ S–C bond. This reaction produces methionine and a highly reactive 5′-deoxyadenosyl radical (5′-dAdo˙), which subsequently abstracts a hydrogen atom from the substrate (Fig. 2).52 This step defines the initiation phase, where the first radical intermediate is generated. Immediately following hydrogen abstraction, the 5′-dAdo˙ is quenched to form 5′-deoxyadenosine (5′-dAdoH), which is a characteristic byproduct detected in most rSAM reactions and serves as a diagnostic marker of radical initiation. Recent studies have raised new questions about the initiation mechanism, including the possibility of organometallic intermediates and their roles in SAM activation.53–58 The resulting substrate-centered radical enters the propagation phafse, where it undergoes rearrangement, transfer, or addition to form new covalent bonds, enabling the construction of structurally complex products.59 During propagation, the radical often reacts with an electron-rich side chain, such as an aromatic ring (e.g., Trp in StrB and PqqE), a thiol (CteB, NxxcB),60 an alcohol (TqqB, HghC), or a backbone amide (HghB), resulting in diverse crosslink types.61–66 In some cases, radical propagation can proceed through multiple reactive intermediates, increasing pathway complexity and enabling the evolution of novel enzyme functions.67 Finally, the catalytic cycle is completed by termination, which restores a closed-shell product via oxidation or reduction of the radical intermediate. Termination can vary significantly across rSAM enzymes, ranging from classic two-electron oxidations to reductive quenching or even the absence of termination when stable radicals are the final products, such as in glycyl radical activation.67The generation and control of 5′-dAdo˙ allows rSAM enzymes to catalyze chemically challenging transformations that are otherwise thermodynamically or kinetically inaccessible. To date, more than 100 distinct enzymatic reactions have been attributed to this superfamily, including over 80 unique radical-mediated transformations, underscoring their exceptional catalytic versatility and evolutionary innovation.15,50,67 Through this radical-based mechanism, rSAM enzymes enable the functionalization of otherwise inert C–H bonds and the formation of structurally diverse linkages, including C–C, C–S, and C–N bonds.46,49,52,68,69
A notable subset of rSAM enzymes includes those containing additional structural domains, such as the SPASM or Twitch family of rSAM enzymes.70 The SPASM domain is named after four founding rSAM enzymes: the subtilosin A maturating enzyme AlbA, the PqqE protein involved in PQQ (pyrroloquinoline quinone) biosynthesis, the anaerobic sulfatase-maturating enzyme anSME, and the mycofactocin-associated enzyme MftC.44,45 This domain coordinates one or two auxiliary [4Fe–4S] clusters that expand catalytic versatility. The Twitch domain is a truncated variant that typically binds one auxiliary cluster.71 Together, these enzymes comprise >100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 predicted members and is distinguished by a conserved C-terminal domain that coordinates one (Twitch) or two (SPASM) auxiliary [4Fe–4S] clusters in addition to the canonical catalytic cluster.70 These auxiliary clusters are thought to facilitate multi-step electron transfer and stabilize reactive radical intermediates, enabling precise control over radical-mediated transformations and significantly broadening the catalytic repertoire of rSAM enzymes, particularly in complex or multi-step biosynthetic pathways.44,45,70 In some cases, auxiliary clusters may also serve as direct electron acceptors during oxidative termination, contributing to redox cycling within the enzyme itself.
000 predicted members and is distinguished by a conserved C-terminal domain that coordinates one (Twitch) or two (SPASM) auxiliary [4Fe–4S] clusters in addition to the canonical catalytic cluster.70 These auxiliary clusters are thought to facilitate multi-step electron transfer and stabilize reactive radical intermediates, enabling precise control over radical-mediated transformations and significantly broadening the catalytic repertoire of rSAM enzymes, particularly in complex or multi-step biosynthetic pathways.44,45,70 In some cases, auxiliary clusters may also serve as direct electron acceptors during oxidative termination, contributing to redox cycling within the enzyme itself.
The catalytic diversity of rSAM enzymes leads to products with a broad range of biological functions, including cofactor biosynthesis (e.g., biotin and molybdenum cofactor), DNA and RNA modifications, tRNA maturation, lipid metabolism, and the biosynthesis of antibiotics and other secondary metabolites.72 The last decade has revealed a significant number of rSAM enzymes involved in modifying RiPPs.28,29,44,45,48,49,69,73,74 In RiPP biosynthesis, rSAM enzymes catalyze a variety of modifications, such as the formation of thioether bridges in sactipeptides,75,76 introducing C–C cross-links in streptides,62 or facilitating backbone rearrangements that give rise to unique macrocyclic architectures.77 These transformations significantly expand the structural and functional diversity of RiPPs, underscoring the importance of rSAM enzymes in the maturation of bioactive peptides.
Genome mining has emerged as a powerful tool for uncovering novel rSAM enzymes and their associated chemistries, particularly within RiPP biosynthetic pathways. By leveraging sequence similarity networks (SSN), genome neighborhood analysis, and curated resources such as NCBI,78 UniProt,79 IMG-ABC,80 TIGRFAMs,81 and RadicalSAM.org,50 researchers have identified diverse rSAM-containing gene clusters that encode precursor peptides and associated tailoring enzymes. Both enzyme-first and product-first strategies, coupled with in vitro reconstitution or in vivo coexpression studies, have revealed numerous RiPP scaffolds, such as subtilosin A,82 darobactin,83 and triceptides22 each featuring unprecedented radical-mediated modifications. These efforts continue to expand our understanding of rSAM-catalyzed transformations and their functional relevance.
1.1. Cyclophane-containing RiPPs
Cyclophane-containing RiPPs represent a distinct and structurally diverse class of ribosomally synthesized and post-translationally modified peptides characterized by their rigid, often polycyclic, macrocyclic frameworks. These architectures arise through site-selective radical-mediated cyclizations, most notably catalyzed by SPASM-domain rSAM (rSAM/SPASM) enzymes, that forge C–C, C–S, C–N, or C–O linkages between unactivated side chains. These covalent crosslinks not only impart conformational rigidity and proteolytic resistance but also endow the peptides with potent biological activity and highly specific molecular recognition features. Such modifications frequently occur at late stages in the biosynthetic sequence, following initial leader peptide recognition and precursor binding, and are often tightly regulated by quorum sensing systems. This section highlights the growing diversity of cyclophane-containing RiPPs, focusing on representative families such as streptides, PQQ, triceptides, darobactins, dynobactins, nosiheptide, aminopyruvatides and elucidates the timing and enzymatic logic behind their radical-mediated macrocyclization.1.1.1.1. Streptide. Streptococcus species, prominent members of the human microbiome and opportunistic pathogens, harbor a diverse and underexplored repertoire of RiPP biosynthetic gene clusters (BGCs).73 These RiPPs are often quorum-sensing (QS)-regulated, enabling bacterial communication through post-translationally modified peptides. Bioinformatic surveys identified over 600 distinct QS-controlled rSAM-modified RiPP gene clusters in Streptococcal genomes, indicating a remarkable biosynthetic landscape. These RiPPs exhibit a range of C–C crosslinks, often catalyzed by rSAM/SPASM proteins, which contribute to structural rigidity, proteolytic stability, and bioactivity.73 Streptococcus-derived RiPPs can be categorized into several major groups based on their precursor peptide sequence motifs and crosslinking chemistry. The streptide, tryglysin, and ryptide families all feature C–C bond formations between aromatic and C(sp3) carbon centers, a hallmark of rSAM-dependent modifications.65,73,84,85 These biosynthetic pathways are often found adjacent to transporters and regulatory genes, reinforcing their role in quorum sensing and bacterial interaction.
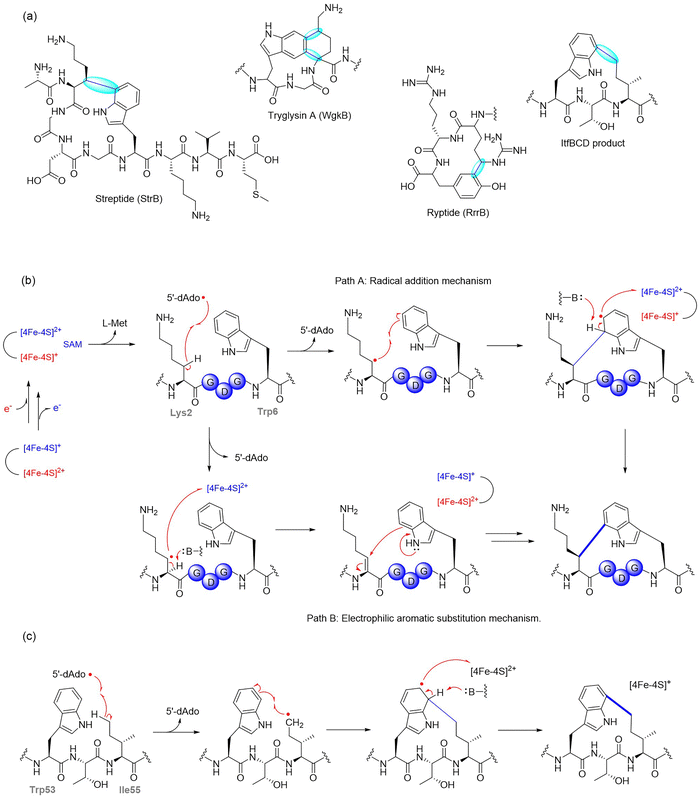
Streptide, a QS-regulated RiPP from Streptococcus thermophilus culture supernatants, features an unprecedented C–C bond between Lys2-Cβ and Trp6-C7.62 This crosslink is installed by StrB, a SPASM-domain rSAM enzyme, encoded within the str cluster, which also includes the precursor peptide StrA and a peptidase-transporter fusion protein StrC. StrB catalyzes this transformation through a radical-based mechanism initiated by reductive cleavage of SAM to generate the 5′-dAdo˙, which abstracts a hydrogen atom from Lys2-Cβ, generating a lysyl radical that subsequently attacks the indole C7 of Trp6 to form the crosslink. The chiral center generated at the Cβ of Lys was shown to adopt an R-configuration through total synthesis and stereochemical comparison of streptide diastereomers (Fig. 3b).62
SuiB and StrB are orthologous rSAM enzymes encoded in conserved str-like clusters across different Streptococcus species, sharing both sequence and functional similarity.86,87 Structural analysis of SuiB revealed three distinct functional domains: the rSAM core, a bridging domain, and a C-terminal SPASM domain that coordinates two auxiliary [4Fe–4S] clusters. The crystal structure provided key insights into the spatial organization of these clusters relative to the active site and revealed features likely involved in peptide substrate recognition and positioning during catalysis.63 The mechanism underlying Lys–Trp crosslink formation has been progressively elucidated through a combination of structural, biochemical, and spectroscopic studies. In 2018, Schramma et al. proposed two possible mechanisms: a canonical electrophilic aromatic substitution (EAS) pathway, involving oxidation of the lysyl radical to form an α,β-unsaturated intermediate followed by indole addition, and a radical electrophilic aromatic substitution (rEAS) pathway, in which the lysyl radical directly adds to the indole ring to generate a σ-radical intermediate (Fig. 3b). In the latter case, bond formation at the indole C7 position produces a cyclohexadienyl-type radical in which the addition site is sp3-hybridized and the aromaticity is disrupted. Although the unpaired electron is partially delocalized over the remaining π system, the defining feature is the sp3 center created by the new C–C σ bond, which classifies this species as a σ-radical. Through a combination of isotope labeling, substrate analogs, and site-directed mutagenesis, the EAS pathway was ruled out in favor of the rEAS model.88
This mechanistic proposal was later validated by Balo et al. in 2021, who used freeze-quench EPR spectroscopy to trap and characterize the Lys–Trp˙ radical intermediate, directly confirming its presence as an on-pathway species within the catalytic cycle.89 Further support for the rEAS mechanism came from isotopic labeling studies: when sidechain-deuterated lysine was incorporated into the precursor peptide, selective loss of the β-deuteron and formation of 5′-2H-5′-deoxyadenosine were observed, while the α-proton remained intact.89 These findings are consistent with site-specific hydrogen atom abstraction from Lys2-Cβ. In addition, the auxiliary [4Fe–4S] cluster AuxI was shown to act as the terminal oxidant responsible for indole rearomatization (Fig. 3b). Together, these results provide the first direct evidence for a radical electrophilic aromatic substitution mechanism for Lys-Trp crosslinking in RiPP biosynthesis and define the catalytic role of SPASM auxiliary clusters in the C–C coupling step.89
1.1.1.2. Tryglysin. Tryglysin A, found in Streptococcus ferus represents a distinct structural class of RiPPs, featuring an unprecedented tetrahydro[5,6]benzindole macrocycle, installed by the rSAM enzyme WgkB.85 Unlike streptide, which forms a single crosslink, WgkB introduces a more rigid bicyclic scaffold through two successive radical-mediated reactions. This enzyme catalyzes two sequential C–C couplings: Trp-C5 to Lys-Cδ (tryglysin A) and Trp-C6 and Lys-Cα (tryglysin A) (Fig. 3a). Notably, no single crosslink intermediates have been observed, suggesting that both modifications are introduced in a single catalytic cycle or that the enzyme exhibits enhanced affinity and/or specificity for the singly crosslinked products.85
Beyond its structural novelty, tryglysin A exhibits potent and selective antimicrobial activity. It effectively inhibits the growth of Streptococcus pneumoniae, a major human pathogen, with a minimal inhibitory concentration (MIC) below 100 nM, which is comparable to that of the broad-spectrum antibiotic ciprofloxacin.85 In contrast to ciprofloxacin, however, tryglysin A displays a narrow-spectrum profile, showing no inhibitory effect against 15 other streptococcal species or common human commensals. This selectivity highlights its potential as a targeted antimicrobial agent with reduced impact on the microbiota.90 Tryglysin A also exerts a self-inhibitory effect on its own producer strain, S. ferus, in a bacteriostatic rather than bactericidal manner. The absence of a dedicated immunity gene within the gene cluster and the reversible nature of the growth inhibition suggests a possible regulatory or signaling function. These observations, along with its species-specific activity, support a model in which tryglysin may act as both a niche-adapted antimicrobial and a quorum-sensing-linked ecological signal within streptococcal communities.90
1.1.1.3. Ryptides. The rrr BGC, originally identified in Streptococcus suis LSS38_2, consists of the shp/rgg system, a precursor peptide (RrrA), an rSAM enzyme (RrrB), and a transporter (RrrC). RrrB catalyzes the formation of a C–C bond between the Arg40-Cδ to Tyr42-C3 within the C-terminal RRY motif of the precursor, resulting in a three-residue cyclophane, termed Ryptide.91 Further studies revealed that this system exhibits substrate promiscuity, as the leader peptide is dispensable. However, any modifications to the core sequence were not tolerated.91 The catalytic mechanism of RrrB is proposed to follow the canonical rSAM radical pathway, where SAM cleavage produces 5′-dAdo˙, which abstracts a hydrogen from Arg-Cδ, generating an arginyl radical. This radical then undergoes nucleophilic attack on Tyr-C3, forming a stable C–C linkage.91 The presence of additional [4Fe–4S] clusters in RrrB likely contributes to the stabilization of the reactive intermediate, preventing side reactions and ensuring selective macrocyclization.
1.1.1.4. Trp crosslinked RiPP from Bacteroides. By employing a co-occurrence search strategy that links rSAM enzymes with transporters, Bushin et al. mapped approximately 15
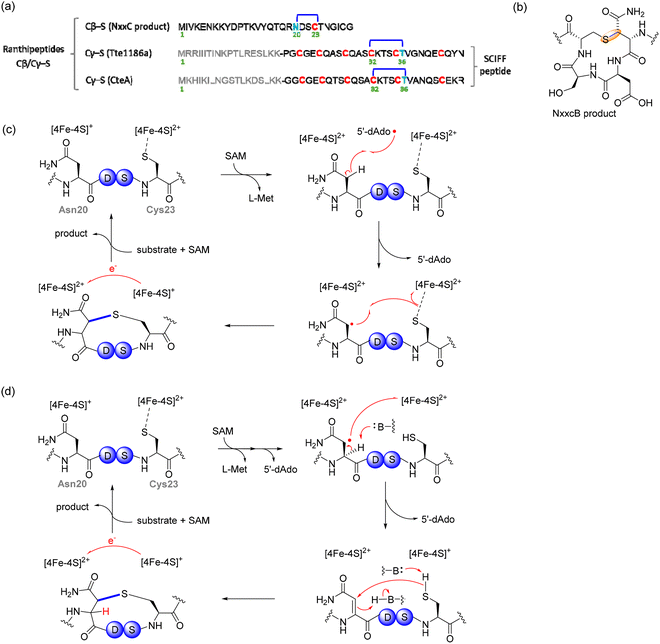
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 rSAM-RiPP BGCs across various microbial genomes, classifying them into 800 subfamilies.84,86 Through this approach, they identified a specific gene cluster from Bacteroides thetaiotaomicron, a prominent member of the human gut microbiome, encoding a YcaO enzyme (ItfB) and a rSAM/SPASM enzyme (ItfD), which catalyze two novel modifications: the installation of a backbone amidine and an unprecedented C–C crosslink between the Ile55-Cδ1 and Trp53-C7, forming a three-residue macrocycle (Fig. 3c).84 These findings were confirmed through in vivo and in vitro biochemical assays, including NMR and HR-MS/MS analysis, which provided mechanistic insights into these modifications.84 In particular, incorporation of uniformly deuterated Ile (2H10-Ile) at the crosslinking site led to a −4 Da mass shift in the product and generation of deuterated 5′-dAdo, implicating Ile55 as the site of hydrogen atom abstraction and supporting a radical-based mechanism for crosslink formation. The study not only expands the known enzymatic repertoire of RaS enzymes but also establishes a robust bioinformatic framework for the discovery of new RiPP natural products.
500 rSAM-RiPP BGCs across various microbial genomes, classifying them into 800 subfamilies.84,86 Through this approach, they identified a specific gene cluster from Bacteroides thetaiotaomicron, a prominent member of the human gut microbiome, encoding a YcaO enzyme (ItfB) and a rSAM/SPASM enzyme (ItfD), which catalyze two novel modifications: the installation of a backbone amidine and an unprecedented C–C crosslink between the Ile55-Cδ1 and Trp53-C7, forming a three-residue macrocycle (Fig. 3c).84 These findings were confirmed through in vivo and in vitro biochemical assays, including NMR and HR-MS/MS analysis, which provided mechanistic insights into these modifications.84 In particular, incorporation of uniformly deuterated Ile (2H10-Ile) at the crosslinking site led to a −4 Da mass shift in the product and generation of deuterated 5′-dAdo, implicating Ile55 as the site of hydrogen atom abstraction and supporting a radical-based mechanism for crosslink formation. The study not only expands the known enzymatic repertoire of RaS enzymes but also establishes a robust bioinformatic framework for the discovery of new RiPP natural products.
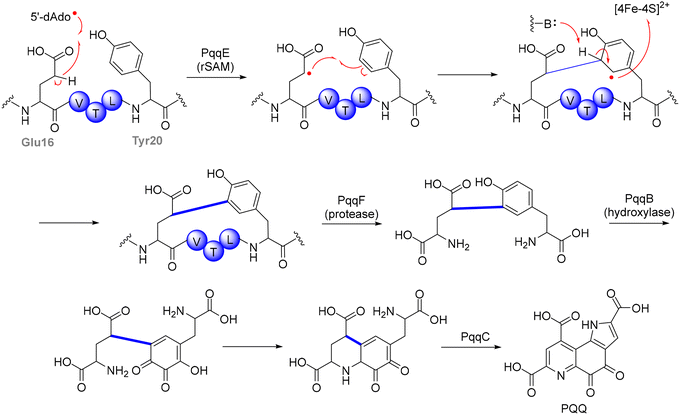
PqqE, a member of the SPASM-domain rSAM enzyme family, initiates PQQ biosynthesis through the reductive cleavage of SAM, generating a 5′-dAdo˙ (Fig. 4). This step was supported by isotope-labeling experiments using a PqqA variant with deuterium at the β-position of glutamate, which showed a significant deuterium kinetic isotope effect on 5′-deoxyadenosine (5′-dAdoH) formation, indicating that the 5′-dAdo˙ abstracts the β-hydrogen from glutamate. The resulting peptide-centered radical then undergoes regioselective coupling with the ortho-position of the tyrosine ring, yielding the core structure of PQQ.93 Unlike many RiPP-modifying rSAM enzymes, which directly recognize their precursor peptides via a leader sequence, PqqE requires PqqD, a specialized chaperone, to facilitate substrate positioning.96,97 Structural studies have revealed that PqqD forms a ternary complex with PqqA and PqqE, stabilizing the precursor and ensuring proper orientation of the reactive residues within the active site. This chaperone-assisted mechanism represents a distinct strategy in RiPP biosynthesis, analogous to the role of leader peptides in other rSAM-modified natural products. Following the PqqE-catalyzed C–C bond formation, the crosslinked PqqA-derived intermediate undergoes downstream processing steps involving proteolytic trimming and further maturation mediated by PqqB, before final conversion to PQQ through the oxygen-dependent oxidation catalyzed by PqqC. Biochemical and structural studies have identified PqqB as an Fe2+-dependent hydroxylase that catalyzes oxygen-insertion reactions on PqqA-derived intermediates, providing a missing link in the PQQ biosynthetic pathway, although the precise sequence and scope of these transformations in vivo remain under investigation.98,99
Beyond its primary [4Fe–4S] cluster for SAM cleavage, PqqE contains two auxiliary iron–sulfur clusters, characteristic of SPASM-domain enzymes, which are essential for radical stabilization and electron transfer during catalysis. The presence and nature of these clusters were elucidated through a multifaceted approach. X-ray crystallographic analysis of Methylorubrum extorquens PqqE revealed two Fe–S cluster-binding motifs in the C-terminal SPASM domain: AuxI and AuxII.100 While AuxII was shown to coordinate a canonical [4Fe–4S] cluster via three cysteines and a unique aspartate ligand (Asp319), AuxI exhibited a [2Fe–2S] configuration in the crystal structure.100 To complement the structural data, EPR spectroscopy using PqqE variants with mutated ligands enabled deconvolution of overlapping signals and assignment of distinct g-tensors corresponding to the RS, AuxI, and AuxII clusters.101 The AuxII [4Fe–4S] cluster was confirmed by its redox behavior and spin-relaxation properties, whereas low-potential reductants such as Ti(III) citrate or Eu(II)-DTPA were required to reduce the [4Fe–4S] state in AuxI, which otherwise adopts a [2Fe–2S] configuration under standard conditions. Further corroboration came from Mössbauer spectroscopy, which distinguished three separate F–S environments, and native mass spectrometry, which quantified metal–sulfur content and confirmed the coexistence of PqqE species harboring two [4Fe–4S] and one [2Fe–2S] cluster, as well as minor populations containing three [4Fe–4S] clusters.102 Site-directed mutagenesis of ligating residues (e.g., Cys → Ala, Asp → His) selectively disrupted cluster assembly, and reconstitution assays demonstrated that all three Fe–S clusters are essential for productive crosslinking activity, highlighting their functional relevance in radical initiation and stabilization.61 These additional clusters fine-tune the reaction specificity, guiding the radical-mediated crosslinking with high regio- and stereoselectivity.100 Spectroscopic and mutagenesis studies have demonstrated their critical role in modulating the reaction pathway and preventing side reactions that could lead to off-target modifications.61,100–102 Mechanistic studies have provided further insights into cooperative interactions within the biosynthetic complex. Hydrogen–deuterium exchange mass spectrometry (HDX-MS) has revealed dynamic conformational changes in the PqqA–PqqD–PqqE assembly, indicating that substrate binding induces long-range structural rearrangements that regulate catalytic efficiency.61 This cooperative allosteric control highlights a broader theme in RiPP biosynthesis, where structural scaffolds and auxiliary domains work in concert to direct enzymatic modifications with high precision.
As one of the few known RiPP-derived redox cofactors, PQQ biosynthesis expands the functional repertoire of rSAM-modified natural products. The combination of radical-mediated C–C bond formation, chaperone-assisted substrate recognition, and the involvement of auxiliary Fe–S clusters exemplify the remarkable versatility of rSAM enzymes in constructing complex molecular architectures. Understanding this biosynthetic logic provides new avenues for engineering bioinspired cofactors and leveraging rSAM chemistry to design synthetic RiPPs with tailored catalytic properties. The interplay between peptide-derived redox scaffolds and enzymatic radical chemistry continues to shape our understanding of both natural and engineered biochemical transformations.
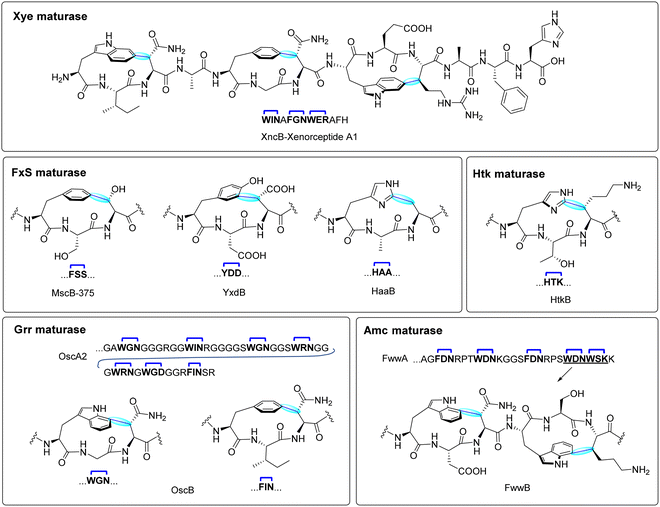
1.1.3.1. XYE Maturase system (xenorceptides). The Xye maturase system is named after the Xenorhabdus, Yersinia and Erwinia bacterial genera where the system is found.22 This system is usually composed of a precursor peptide (XyeA), rSAM/SPASM maturase XyeB responsible for crosslinking, an aspartic acid protease XyeC for peptide processing, a HlyD-type transporter (XyeD) for secretion, and a fused protease-transporter (XyeE). The precursor peptide XyeA contains approximately 60 residues, with a Gly–Gly motif separating the leader and core region, and a distinctive WxxxF/YxxWxxxx sequence at its C-terminus, which serves as the scaffold for radical-mediated crosslinking.22 The catalytic activity of XncB, an rSAM maturase from Xenorhabdus nematophila, was validated through heterologous expression with its cognate precursor XncA in E. coli. Functional characterization confirmed that XncB introduces three independent cyclophane crosslinks, linking Trp1-C6 to Asn3-Cβ, Phe5-C4 to Asn7-Cβ, and Trp8-C6 to Arg10-Cβ, forming a rigid macrocyclic framework (Fig. 5, top).22Interestingly, Han et al. conducted in vitro studies on PauB, a homologous maturase from Photorhabdus australis revealed these crosslinks occur independent and sequentially, with bond formation initiating at the N-terminal position, followed by the C-terminal, and finally the central crosslink.107 While a more recent study by Phan et al. further investigation into the biosynthetic flexibility of XncB revealed unexpected oxygen incorporation into the modified product, raising questions about alternative reaction pathways and enzymatic promiscuity within the Xye maturase system.108 Full-cluster expression studies of His6-SUMO-xncA + xncBCDE led to the detection of a minor +12 Da mass shift, which was localized via MS/MS analysis to Asn7, suggesting the presence of an oxygenated derivative. This modification was absent in non-aromatic X3 variants (N7R/K/S/A), indicating that oxygen incorporation may be linked to specific substrate properties or radical stability. The precise origin of this oxygen remains unclear. Potential mechanisms include hydroperoxide-mediated oxidation, radical rearrangement, or water addition via a dehydro amino acid intermediate.108 These findings highlight a previously unrecognized divergence in rSAM enzymology, where certain maturases, like XncB, may exhibit latent oxidative potential that extends beyond conventional C–C bond formation.
Mass spectrometry also revealed that XncB modifies cyclophanes in an unexpected order, differing from PauB. Instead of an N → C → central sequence, XncB modifies the C-terminal motif first, followed by the N-terminal, and finally the central ring.107,108 The underlying reason for this shift in reaction order between homologous enzymes remains an open question, though it may be attributed to differences in active site architecture, substrate positioning, or the influence of auxiliary Fe–S clusters on radical stabilization. These insights underscore the functional divergence of triceptide maturases, suggesting that fine-tuned enzymatic engineering could be leveraged to control modification order and expand the chemical space of RiPP-derived macrocycles.
1.1.3.2. Gly-rich repeat maturase systems. The Gly-rich repeat (Grr) maturase systems are primarily found in cyanobacteria, and encode a precursor peptide GrrA, a rSAM/SPASM enzyme GrrM, and an ABC transporter substrate-binding protein GrrP. These BGCs, such as those from Oscillatoriales cyanobacterium (osc), Lyngbya sp. PCC 8106 (lsc), and Geminocystis sp. NIES-3709 (gsc), direct the synthesis of precursor peptides ranging from 100 to 200 residues. Approximately half of each precursor corresponds to a C-terminal core region that is glycine-rich and interspersed with multiple Ω1–X2–X3 motifs, which serve as recognition sites for enzymatic modification (Fig. 5, bottom left).22
Unlike the Xye maturase system, which modifies compact precursors at specific sites, Grr-type maturases exhibit broad substrate promiscuity, catalyzing modifications at nearly all available motifs.109 Coexpression of His6-tagged precursors (native or engineered with trypsin cleavage sites) with their cognate maturases in E. coli led to −2 Da mass shifts detectable by LC-MS in almost every Ω1–X2–X3 motif, indicating widespread crosslinking activity.22 Detailed structural elucidation using a minimal OscA2 precursor containing two motifs (WGN and FIN) confirmed that OscB catalyzes the formation of a Trp-C7 to Asn-Cβ cyclophane as well as a Phe-C4 to X3-Cβ linkages (Fig. 5, bottom left).22 Substitutional flexibility at the X2 and X3 positions and tolerance for varied motif spacing further highlight the relaxed sequence specificity of OscB.109 Further functional studies revealed that OscB accepts a wide range of amino acids at the X2 position, including small, polar, and hydrophobic residues, while being more selective at X3—tolerating some substitutions but with reduced efficiency. Critically, only tryptophan and phenylalanine were accepted at the Ω1 position, underscoring the enzyme's specificity for aromatic radical initiation.109 In-depth mutational analysis also identified a key recognition region within the leader peptide, essential for OscB's catalytic activity. Notably, the enzyme demonstrated robust activity on substrates with significant variation in the number and spacing of Ω1–X2–X3 motifs, efficiently processing precursor peptides with C-terminal cores up to 47 residues and even engineered chimeric precursors extending to 80 residues. Despite this structural complexity, OscB consistently formed 3-residue cyclophanes, indicating precise control over ring topology and size.109
NMR-based conformational studies demonstrated that Trp-C7-modified products exhibit planar chirality, with the indole ring adopting an orientation comparable to the C6-modified Trp in xenorceptide. Specifically, in the Trp-C7-Asn-Cβ linkage, Trp-C7a is the highest priority carbon, and the pilot atom is Asn-Cα, corresponding to a 7M or 7Sp stereochemical configuration. These findings raise important questions regarding whether the observed stereocontrol arises from substrate pre-organization or enzyme-imposed constraints via the active site or auxiliary Fe–S cluster interactions. The physiological role of the extracellular substrate-binding protein (GrrP) and the fate of these highly modified peptides remain unknown.109
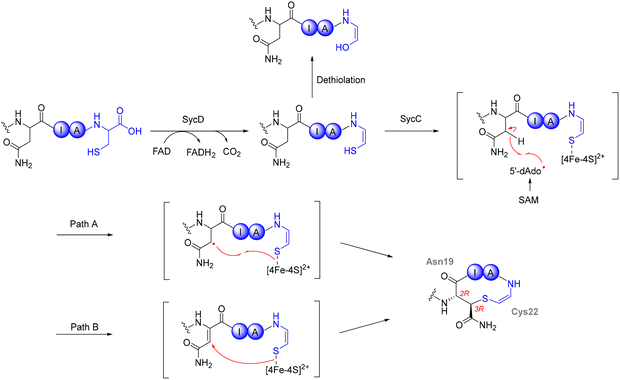
1.1.3.3. FxS Maturase systems. The FxS maturase systems are triceptide BGCs found in actinobacteria. These systems are characterized by a precursor peptide (FxsA) containing a C-terminal FxSxx motif, an rSAM/SPASM maturase (FxsB), and additional accessory proteins such as a tetratricopeptide repeat protein and mod_HExxH domain-containing protein.22,104,110 Initial studies on the maturase, MscB-375 from Micromonospora sp. demonstrated the enzymatic formation of a Phe-C4 to Ser-Cβ cross-link within this motif, highlighting the canonical reaction catalyzed by this family. Subsequent investigations into the FxS maturase system have expanded its known substrate scope and reaction diversity. The enzyme SjiB-399, exhibited notable catalytic promiscuity, forming not only canonical cyclophane products in vivo but also noncanonical oxidative modifications yielding formylglycine (FGly) and aminomalonate (Ama) in vitro.106 These findings suggest that certain FxS-type maturases may operate through mechanistically distinct pathways depending on reaction conditions. Additional studies using SjiB-375 have shown potential to incorporate aldehyde tags as an alternate strategy to formyl-Gly generating enzyme.111,112 This unique ability to catalyze oxidative side-chain modifications sets certain Fxs maturases apart from Xye and Grr systems, which predominantly form C–C cross-links. The discovery of YxdB, HaaB, WnsB and related maturases expanded the reactivity of FxsB-like enzymes to include all aromatic amino acids at the Ω1 position (Fig. 5, middle left).104,113
Additional substrate mutagenesis experiments showed that certain maturases exhibit relaxed specificity at the X3 position, allowing for the incorporation of alternative residues, which could influence regioselectivity of the crosslinking.113 The FxS and related maturases are a functionally diverse and evolutionarily distinct branch of radical-mediated RiPP maturation, expanding the known chemical space of triceptides. Understanding the enzymatic mechanism of triceptide biosynthesis provides a framework for engineering bioactive macrocyclic peptides, particularly in the development of structurally rigid and protease-resistant scaffolds with pharmaceutical relevance. By leveraging the inherent stereochemical control and regioselectivity of triceptide maturases, it may be possible to rationally design novel peptide-based therapeutics that exploit the unique binding properties and metabolic stability of cyclophane-containing RiPPs. The continued exploration of triceptide biosynthetic pathways, particularly in uncharacterized bacterial strains, is likely to uncover additional maturase subfamilies with novel enzymatic capabilities, further broadening the landscape of peptide-based natural product discovery.
1.1.3.4. Actinobacterial multiple cyclophane (Amc) maturase systems. Actinobacterial multiple cyclophane (Amc) maturases represent a distinct class of rSAM/SPASM enzymes involved in triceptide biosynthesis. Predominantly found in Actinobacteria, these enzymes catalyze multiple C(sp2)–Cβ(sp3) bond formations within a single precursor peptide, generating complex cyclophane architectures. Sequence similarity network (SSN) analyses revealed that Amc-type maturases are enriched in actinobacterial genomes, particularly in Streptomyces and Micromonospora, supporting their classification as a unique maturase subfamily.114
Bioinformatic profiling revealed over 500 unique Amc-associated precursors, many containing tandem or repetitive Ω1–X2–X3 motifs. These motifs frequently featured Trp or Phe as the aromatic crosslinking residue, arranged in clustered patterns that distinguish Amc precursors from those of Xye, Grr, or Fxs systems.114 Five major precursor families were identified, with some encoding up to 7–8 potential crosslinking sites.114 This suggested an unusually high macrocyclization potential encoded within a single gene product. The Fww system from Actinospira robinae served as a model for functional validation. Coexpression of the FwwB maturase with its precursor peptide FwwA in E. coli confirmed enzymatic formation of multiple cyclophanes, as evidenced by LC-MS/MS and NMR.114 Crosslinks were identified between Trp-C7 and either Asn-Cβ or Lys-Cβ in specific WDN and WSK motifs, with NMR analysis confirming planar chiral macrocycles analogous to those produced by OscB (Fig. 5, bottom right).22,114 Importantly, engineered variants revealed that FwwB modifies these sites in a partially processive and motif-selective manner, generally initiating at C-terminal motifs before proceeding upstream. Substrate mutagenesis showed that FwwB is relatively specific for Trp at the Ω1 position and tolerant of limited variation at X3, though certain changes can disrupt activity. Unlike some triceptide enzymes, FwwB does not require Aux I binding cysteines, suggesting it operates with a single auxiliary [4Fe–4S] cluster.114 Mutation of Glu280, a conserved active site residue, reduced catalytic activity and altered the number of rings formed, highlighting its role in tuning reactivity.114
Although the natural end-products of Amc BGCs have not been isolated, the functional capacity of FwwB demonstrates the potential of these enzymes to install multiple, stereochemically defined crosslinks in peptide scaffolds. Amc maturases therefore expand the diversity and biosynthetic logic of triceptide biosynthesis and represent a promising avenue for further exploration of these pathways.
1.1.3.5. YxRxHxRHxR-tail maturase system. The YxRxHxRHxR-tail maturase systems are encoded in diverse bacteria including cyanobacteria and enterobacteria. The precursor peptides in this family contain a canonical YxRxHxRHxR sequence in the core peptide at the C-terminus. Two types of these systems have been reported. The first type contains a precursor peptide (A), a rSAM/SPASM enzyme (B), an αKG- and Fe-dependent oxygenase (C), and in certain cases an αKG-HExxH oxygenase.104
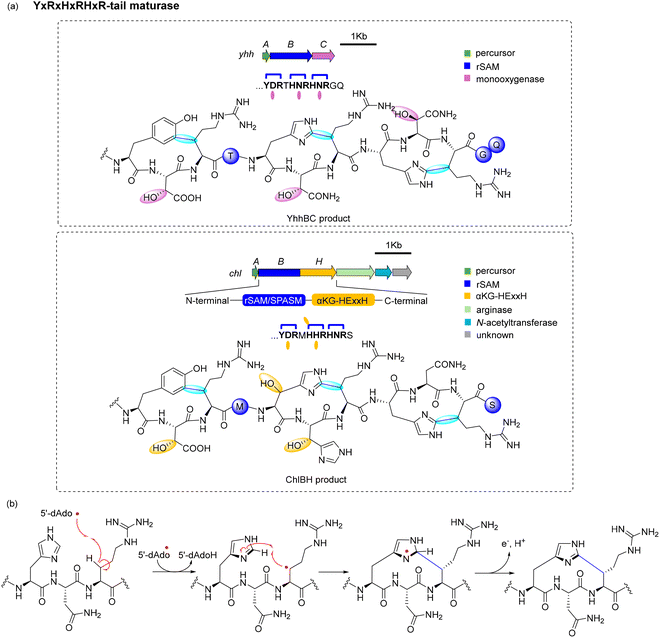
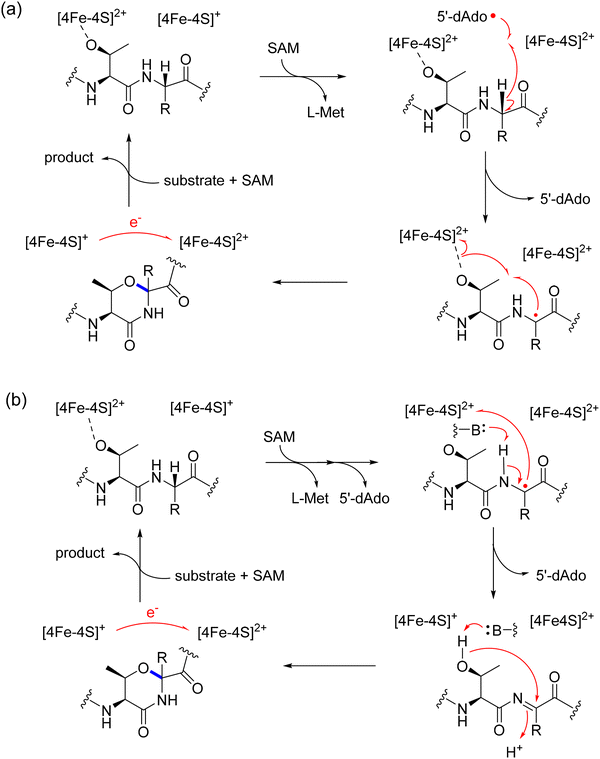
An example of this type is the yhh BGC from N. lactamica. In vivo coexpression of the rSAM/SPASM enzyme, YhhB with its precursor peptide YhhA in E. coli followed by MS/MS and NMR analysis identified the formation of three new types of C–C crosslinks.104 The rSAM/SPASM enzyme YhhB is a triceptide synthase that forms crosslinks on Tyr and His residues (Tyr1-C3 to Arg3-Cβ, His5-C2 to Arg7-Cβ, and His8-C2 to Arg10-Cβ).104 Triple expression of precursor peptide (YhhA), rSAM/SPASM enzyme (YhhB), and the associated oxygenase YhhC further introduced β-hydroxylations at the X2 residues (Asp2, Asn6, and Asn9), as evidenced by +16 Da mass shifts, modified spin systems in NMR, and Marfey's analysis confirming the L-erythro configuration (Fig. 6a, top).104
An example of the second type of YxRxHxRHxR-tail containing system was reported from Chlorogloeopsis sp. PCC 7702. This system contains a precursor peptide (ChlA), a fused rSAM/SPASM – αKG-HExxH oxygenase (ChlBH), arginase, N-acetyltransferase, and a protein of unknown function. ChlBH is a bifunctional enzyme comprising an rSAM/SPASM domain (B) and an αKG-dependent hydroxylase domain (H) (Fig. 6a, bottom).110 To understand the function of the fused protein, the rSAM/SPASM domain was studied first. Suarez et al. characterized that cyclophane formation occurs at Tyr1-C3 to Arg3-Cβ. These studies also showed that ChlB was promiscuous and can accept other amino acids at X2 and X3 positions. Subsequently, Morishita et al. studied the activity of the full length protein, ChlBH.110 These studies showed that ChlB catalyzes triceptide crosslinks on Tyr-C3 to Arg-Cβ, His5-C2 to Arg7-Cβ, and His8 to Arg-Cβ. The C-terminal HExxH domain which was proposed to act as a protease was found to catalyze β-hydroxylation at Asp2, His5, and His6. These studies suggested that modification begins in the rSAM/SPASM domain and traverses between the two domains for full modification. An X-ray crystal structure of the HExxH domain protein shows the core fold most closely resembles zincin protease but the remaining structure represents a new fold for αKG- and Fe-dependent oxygenases. These oxygenases are broadly encoded in triceptide and other RiPP pathways have been named αKG-HExxH family oxygenases.110
Recently, the crystal structure of the rSAM/SPASM domain ChlB in complex with variants of the precursor peptide (ChlA) revealed insights into substrate recognition and catalysis.110 The precursor peptide docks via its leader region to a helical linker connecting the two catalytic domains, allowing the core region to swing between the rSAM or oxygenase active sites.115 Biochemical and structural data revealed that ChlBH likely installs the modifications in a defined sequence: Cyclophanes are formed first on the YDR and HNR, followed by β-hydroxylation at Asp2 and His6, cyclization then occurs at the HHR motif, and finally β-hydroxylation at His5 (Fig. 6a, bottom).115 This ordered progression was confirmed by in vivo and in vitro assays using truncated ChlB variants (ChlBΔ375), LC-MS/MS, and mutational analysis of the substrate peptide.115
Crystal structures of ChlBΔ375 in complex with substrate analogs showed the precise arrangement of Arg Cβ in the active site for hydrogen abstraction by 5′-dAdo˙, and optimal orientation of the aromatic residue for C–C bond formation. These data also revealed a salt bridge between the Arg residue and Asp201 of ChlB that likely explains the conserved requirement for Arg at the X3 position in all three ΩXR motifs.115 Mechanistically, the rSAM domain initiates catalysis by reductive cleavage of SAM to generate the 5′-dAdo˙, which abstracts a hydrogen atom from the β-carbon of the X3 Arg residue. The resulting carbon-centered radical then undergoes regioselective addition to the ortho-position of the neighboring Tyr or His ring, forming a σ-radical intermediate that is rearomatized via electron transfer (Fig. 6b).115 Interestingly, ChlB lacks the canonical AuxI [4Fe–4S] cluster typical of SPASM domains. Instead, it contains only the [4Fe–4S]RS and AuxII clusters. Despite the absence of AuxI, the 12.5 Å distance between the reactive radical intermediate and the RS cluster is sufficient to allow direct electron transfer for radical quenching, circumventing the need for an auxiliary relay center.115 The lack of the AuxI cluster for appears to be a general trend for triceptide synthases.
1.1.3.6. Htk maturase system. The Htk maturase system, discovered in Pandoraea sp. XY-2, represents a recently characterized subfamily of triceptide synthases that catalyze radical-mediated C–C bond formation within HTK motifs. The BGC encodes a precursor peptide (HtkA) and a rSAM/SPASM maturase (HtkB), which together enable the formation of a macrocyclic crosslink between the His1-C2 and the Lys3-Cβ (Fig. 5, middle left).116 This transformation mirrors those catalyzed by other FxS-like triceptide maturases such as HaaB and YhhB but is distinguished by its substrate preferences and active site architecture.104,116 Functional characterization of HtkB was performed through heterologous expression of the HtkA/B pair in E. coli, followed by peptide purification, trypsin digestion, and LC-MS/MS analysis. A −2 Da mass shift localized to the HTK motif indicated successful macrocyclization. Structural elucidation by NMR confirmed the crosslink between His1-C2 and Lys3-Cβ and revealed the presence of planar chirality in the resulting cyclophane. The configuration was assigned as 2P based on NOESY correlations and Marfey's analysis, which confirmed that all residues retained the L-configuration.116 This configuration and substitution pattern resemble the HAA and HNR cyclophanes formed by other triceptide enzymes. To probe the substrate scope of HtkB, His1 was substituted with Phe and Trp. Coexpression with HtkB yielded detectable cyclized products, with the FTK variant forming a significant amount of product, confirmed by NMR to adopt a para-cyclophane topology. These results suggest that cyclophane topology may depend on both the identity of the aromatic residue and the properties of the flanking residues, with Phe-containing substrates favoring para- over meta-linkage.116 In addition, the double variant H1F/K3R retained minimal activity, indicating potential for broader substrate acceptance albeit with reduced efficiency.
A key insight from this study was the identification of a conserved active site residue, Asp214 in HtkB, which governs substrate specificity at the X3 position. Homology modeling and structural comparison with HaaB, a His1-C2 to Ala3-Cβ cyclophane synthase, revealed a positional equivalence between HtkB Asp214 and HaaB His204.116 Site-directed mutagenesis confirmed their roles: substitution of D214H in HtkB enhanced activity toward an X3 = Ala-containing substrate, while the reciprocal mutation H204D in HaaB conferred activity on Arg- and Lys-containing motifs.116 These findings demonstrate that single-residue changes can invert substrate preference, enabling rational reprogramming of substrate scope in triceptide synthases. This highlights the potential of triceptide maturases as a useful system for tuning rSAM enzyme reactivity and expanding the diversity of macrocyclic RiPPs.
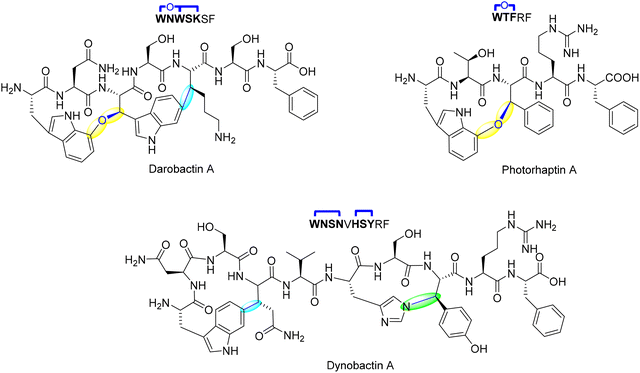
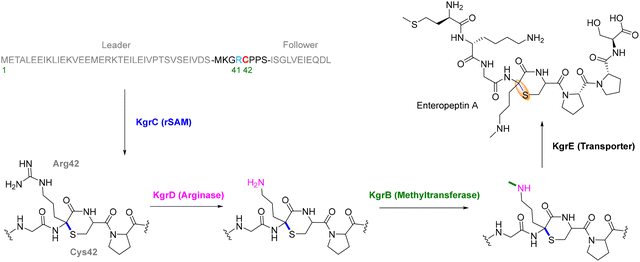
Darobactin, was discovered through a bioactivity-guided screening of 67 Photorhabdus and Xenorhabdus isolates, aimed at identifying compounds active against E. coli. While most isolates failed to inhibit growth, a concentrated extract from Photorhabdus khanii HGB1456 produced a small zone of inhibition. Guided by this activity, researchers isolated a novel compound with an m/z of 966.41047, later identified by HR-MS and NMR as a heptapeptide with an amino acid sequence of W1–N2–W3–S4–K5–S6–F7 (Fig. 8). NMR studies revealed this heptapeptide containing two unprecedented macrocyclic crosslinks.83 Genome mining confirmed a ribosomally synthesized origin from a silent operon encoding a rSAM enzyme (DarE), and deletion of the operon abolished compound production, validating its biosynthetic role. Darobactin features a heptapeptide scaffold stabilized by a C–O–C ether linkage between the indole of Trp and an adjacent residue, along with a C–C crosslink between Trp-C7 and Lys-Cβ.83 Unlike typical RiPP modifications, the ether bond is not derived from pre-existing functional groups but is post-translationally installed using molecular oxygen.120 This modification is catalyzed by DarE, an unprecedented oxygen-utilizing rSAM enzyme, expanding the known reactivity of this enzyme class.
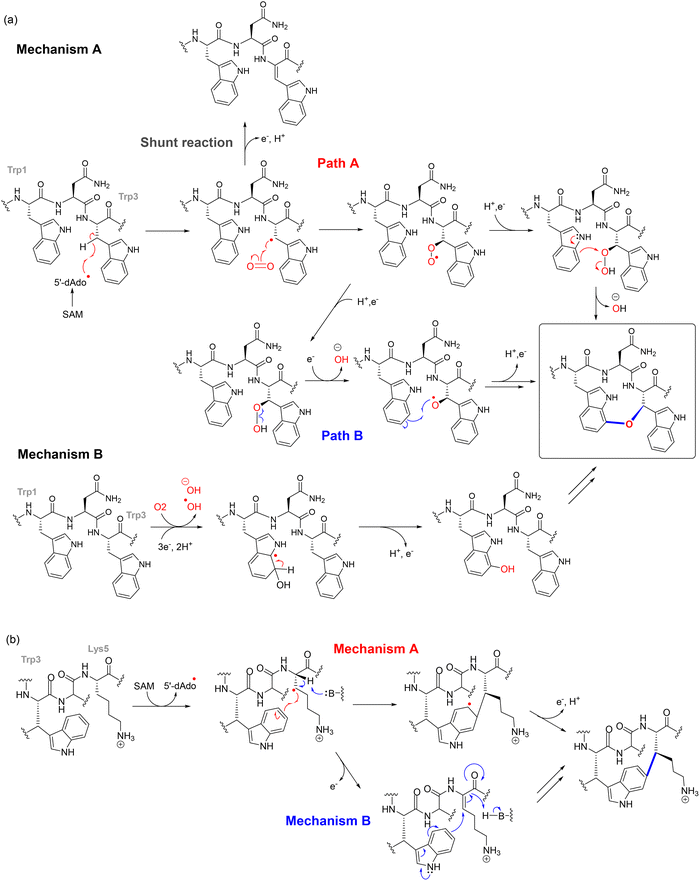
The mechanism of ether bond formation in darobactin biosynthesis has been elucidated through two complementary studies. Guo et al. demonstrated via in vitro reconstitution that the rSAM enzyme DarE is solely responsible for installing both the C–O–C and C–C crosslinks in the mature bicyclic scaffold.105 Initial mechanistic insights suggested that DarE-mediated ether formation proceeds via hydrogen abstraction from Trp3, followed by hydroxylation and subsequent coupling with the indole ring of Trp1. Notably, isotope labeling studies indicated that the oxygen atom in the ether linkage may originate from water.105 The term, daropeptides, was given to this class of cyclophane rSAM-RiPPs that contain an ether linkage as the defining transformation.
However, Nguyen et al. provided more definitive and physiologically relevant evidence that the ether oxygen is derived from molecular oxygen. In this updated model, DarE employs O2 as a cosubstrate, with the 5′-dAdo˙ radical generated from SAM cleavage initiating hydrogen abstraction from the peptide substrate. The resulting peptide radical undergoes oxygen insertion, ultimately leading to ether bond formation (Fig. 9). These findings firmly establish molecular oxygen, rather than water, as the true source of the oxygen atom in the darobactin ether linkage.120 In one plausible mechanism (Mechanism A) (Fig. 9a), the 5′-dAdo˙ abstracts the β-hydrogen from Trp3, forming a Cβ radical that reacts directly with O2, yielding a peroxyl radical. Subsequent electron and proton transfer, likely involving auxiliary clusters, converts this intermediate into a hydroperoxide (Fig. 9a). The ether crosslink may then be formed either by nucleophilic attack of the W1 indole ring onto the hydroperoxide (path a) or by homolytic O–O bond cleavage to generate an alkoxy radical that attacks the Trp1 indole (path b), completing ether formation.120 Alternatively, in Mechanism B (Fig. 9a), oxygen insertion may proceed via direct oxidation of the Trp1 indole ring by reactive oxygen species (ROS), such as hydroxyl radicals generated from O2 within the DarE active site. This would yield a 7-hydroxy-Trp1 intermediate, which could engage in ether bond formation either through radical coupling with a desaturated Trp3 or through interaction with the AUX1 cluster, similar to mechanisms observed in thioether-forming RiPP enzymes.120 This route could also explain the formation of the shunt product, where the Trp3-Cβ radical forms prior to successful Trp1 oxidation. However, DarE assays with hydrogen peroxide and superoxide, potential ROS sources, failed to produce the ether-linked product. Furthermore, in the absence of SAM and the resulting lack of 5′-deoxyadenosyl radical, no Trp1 oxidation was detected. This indicates that ROS alone are insufficient to drive the transformation. These observations make Mechanism B less likely, although it cannot be entirely excluded.120 Consistent with these findings, data from engineered substrates are consistent with substrate-controlled Cβ-radical lifetimes that favor ether formation for aromatic residues and C–C coupling for aliphatic residues. Moreover, DarE can install the ether and C–C macrocycles independently and at alternative positions, consistent with a substrate-controlled model for selectivity.121 Simultaneously, the C–C crosslink between Trp and Lys follows a canonical radical recombination mechanism, potentially via initial hydrogen abstraction from the Lys5-Cβ, generating a carbon-centered radical that recombines with the indole radical of Trp3. Alternatively, DarE may catalyze α,β-desaturation of Lys5, forming an electrophilic Michael acceptor that reacts with the W3 radical to complete the C–C linkage. The reaction is likely coordinated by the auxiliary Fe–S clusters of DarE, ensuring tight control over the radical chemistry and regioselectivity (Fig. 9b).81 These principles provide practical rules for design: tuning residue-3 electronics and binding pose can redirect DarE between ether and C–C outcomes without changing the enzyme.121
A study by Ma et al. on darobactin maturases, including the DarE ortholog PasB, showed that product outcome is controlled by the substrate sequence rather than by enzyme specificity alone.122 In this work, they discovered a new subfamily of daropeptides, from Photorhabdus asymbiotica, including photorhaptin A, which contain a single ether crosslink and lack C–C crosslinks typical of darobactin (Fig. 8).122 Through phylogenetic analysis and mutagenesis studies, the research demonstrated that daropeptide maturases exhibit intrinsic multifunctionality, being capable of catalyzing ether bond formation, C–C crosslinking, and serine oxidation. Notably, the Ω1–X2–Ω3 motif primarily promotes ether crosslink formation, whereas the Ω1–X2–X3 motif facilitates C–C crosslinking.122 These findings establish a substrate-controlled model of catalysis that underlies the structural diversity of daropeptides.
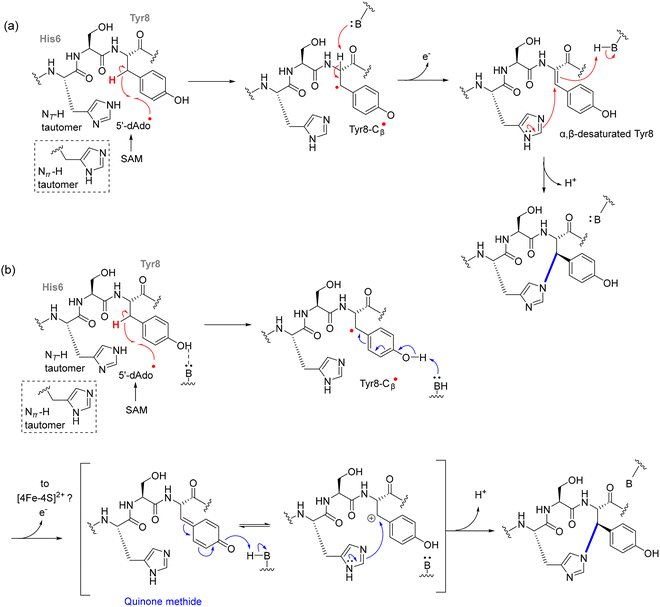
Although structurally distinct from darobactins, dynobactins share a common biosynthetic framework but introduce an unprecedented His–Tyr C–N crosslink in addition to a C–C linkage.103 The C–N bond formation, catalyzed by the rSAM enzyme DynA, represents a unique departure from previously known rSAM-RiPP cyclophane reactions. This transformation is initiated by Tyr-Cβ radical formation, facilitated by hydrogen abstraction via 5′-dAdo˙. Unlike conventional radical recombination pathways, the Tyr radical undergoes oxidation to form a p-quinone methide intermediate, which then reacts with His-Nτ via nucleophilic addition, completing the crosslink (Fig. 10).123 This mechanism diverges from typical radical-mediated crosslinks, where direct radical-radical recombination dominates. Instead, it resembles non-enzymatic quinone methide-mediated nucleophilic substitutions, indicating an expanded catalytic repertoire within rSAM enzymology. Support for the quinone methide mechanism comes from multiple experimental observations. Deuterium labeling at Tyr8 Hβ was transferred to 5′-dAdoH, directly implicating hydrogen abstraction at this position. In contrast, α,β-desaturation was excluded because no solvent deuterium was incorporated in D2O and α-deuterium was retained in the product. In addition, when His6 was replaced by Ala, DynA enabled neutral small nucleophiles such as imidazole, cysteamine, and cysteine to attack Tyr8-Cβ, producing covalent adducts that were labile under MS/MS, behavior consistent with capture of a short-lived quinone methide. The requirement for a Tyr phenol at position 8 and the detection of a His–Tyr crosslinked intermediate that precedes the final C–C bond further reinforce this stepwise pathway.123 Recent biochemical studies on DynA have demonstrated that the enzyme catalyzes the C–N crosslink in dynobactin A through a stepwise mechanism. It was found that the N–C bond between His6 and Tyr8 forms first, followed by the C–C crosslink between Trp1 and Asn4.124
Beyond their biosynthetic novelty, these pathways raise broader questions about the role of oxygen in rSAM enzyme function. Traditionally, rSAM enzymes were thought to be strictly anaerobic due to the oxygen sensitivity of [4Fe–4S] clusters, yet the oxygen-dependent etherification catalyzed by DarE challenges this paradigm. The ability of these enzymes to operate in oxidative environments suggests that some rSAM enzymes may have evolved to exploit oxygen-derived intermediates, a concept that warrants further investigation across other RiPP biosynthetic pathways. Moreover, the structural modularity of these peptides underscores the bioengineering potential of rSAM enzymes. The ability to install highly selective C–C, C–O–C, and C–N bonds within constrained peptide frameworks offers avenues for designing stable, protease-resistant macrocyclic peptides with improved pharmacological properties.
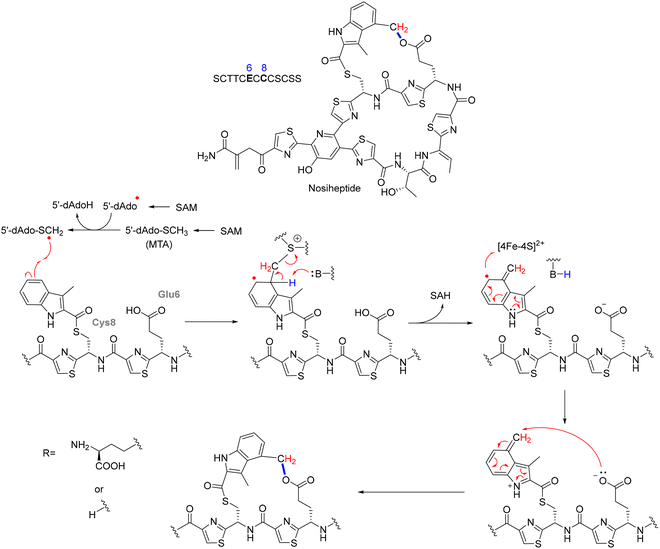
NosN's catalytic mechanism diverges from typical rSAM enzymes, which primarily catalyze reductive cleavage of SAM to generate a 5′-dAdo˙ for hydrogen abstraction. In the case of NosN, two molecules of SAM are simultaneously bound in the active site. One SAM undergoes canonical reductive cleavage to form the 5′-dAdo˙, which specifically abstracts a hydrogen atom from the methyl group of the second SAM molecule, generating a methylene radical.127 This reactive intermediate then adds to the C4 position of the MIA moiety (3-methyl-2-indolic acid) that is thioester-linked to Cys8 on the precursor peptide. The resulting radical intermediate is deprotonated at C4, yielding a radical anion that undergoes heterolytic C–S bond cleavage, eliminating 5′-thioadenosine and forming a reactive exocyclic methylene species. This electrophilic intermediate is subsequently trapped by the side-chain carboxylate of Glu6, forming the ester linkage that completes the side-ring system in nosiheptide. The reaction sequence, as outlined in Fig. 11 thus includes: (1) SAM cleavage → 5′-dAdo˙; (2) H-abstraction from methyl-SAM → methylene radical; (3) radical addition to MIA-C4; (4) elimination of 5′-thioadenosine; (5) Glu6 attack → ring closure (Fig. 11).127,128
The mechanistic proposal is supported by a comprehensive set of biochemical and structural studies. In vitro reconstitution of NosN with synthetic peptide mimics such as MIA-SNAC and tripeptide conjugates (e.g., MIA-S-3mer) demonstrated conversion to dimethylated products only in the presence of SAM and a reducing agent, such as dithionite or titanium(III) citrate.127 These reactions yielded two key byproducts, 5′-dAdoH and 5'-methylthioadenosine (MTA), instead of the typical S-adenosylhomocysteine (SAH), highlighting an unusual SAM fragmentation mechanism that is distinct from canonical SN2 methylation. Time-resolved LC-MS analysis and trapping experiments further captured SAM-indole adducts, consistent with a radical-based intermediate that precedes the formation of the exocyclic methylene group.127 Importantly, substrate analogs incapable of forming the side-ring or bearing rigid linkers led to accumulation of stalled intermediates, reinforcing the sequential nature of C1 transfer and macrocyclization. Complementary studies employing nosN-knockout mutants revealed the accumulation of a nosiheptide analogue lacking both the C4 methyl group and ester linkage, confirming the indispensability of NosN for complete side-ring assembly.129,130 Furthermore, analysis of substrate timing within the biosynthetic pathway indicated that NosN acts before NosO-mediated pyridine ring formation and likely prior to the NosD/E-catalyzed dehydrations, underscoring its early role in shaping the macrocyclic topology.125
NosN's action occurs in a precisely timed sequence, likely before pyridine ring formation catalyzed by NosO and before the dehydration of specific amino acids by NosD/E.125,130 Studies using peptide mimics have captured reaction-based SAM adducts, which provide evidence for an intermediate containing a delocalized radical species, supporting the proposed radical-based mechanism.126 Additionally, NosJ, NosI, and NosK are responsible for the activation and transfer of the MIA moiety to the core peptide, where NosK serves as an acyltransferase that appends MIA to Cys8 before NosN's intervention.125 The nosiheptide biosynthetic pathway exemplifies the divergent strategies employed by rSAM enzymes to catalyze complex macrocyclizations. The NosN-catalyzed reaction introduces an oxygen-linked methylation-coupled esterification step, which distinguishes it from other rSAM-mediated RiPP modifications such as C–C or C–S bond formation. This unique mode of modification not only expands the enzymatic repertoire of rSAM chemistry but also highlights new potential avenues for bioengineering thiopeptide derivatives with altered pharmacological properties.
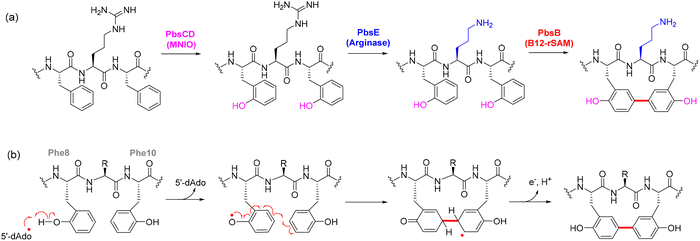
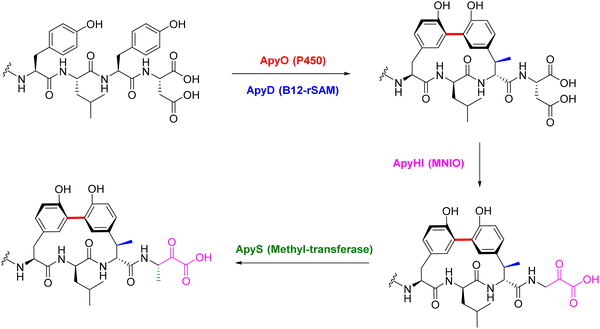
Recent studies have expanded the catalytic repertoire of B12-dependent rSAM enzymes beyond classical methylation reactions to include unprecedented carbon–carbon bond-forming processes.132 A novel macrocyclization strategy has been elucidated in Peribacillus simplex through the characterization of a BGC pbs that encodes a suite of three metalloenzymes acting sequentially to modify a ribosomally synthesized peptide.133 These enzymes include a multinuclear non-heme iron-dependent oxidative enzyme (MNIO, PbsC), an arginase (PbsE), and a B12-dependent rSAM enzyme (PbsB) (Fig. 12a). Together, they orchestrate the stepwise construction of a rigid macrocyclic framework via an unprecedented C–C bond formation between the ε2 positions of two ortho-hydroxylated aromatic residues derived from phenylalanine. Genome mining identified a conserved FHAFRF motif within the precursor peptides genes (pbsA1–A4), which are co-encoded with pbsCDE and the rSAM gene pbsB. Heterologous reconstitution of the pathway in Escherichia coli enabled the purification and structural characterization of key intermediates, including the bis-hydroxylated product (PbsA3-CD), the deguanidinated intermediate (PbsA3-CDE), and the fully cyclized macrocycle (PbsA3-BCDE) (Fig. 12a).133
The biosynthetic sequence begins with PbsC, in cooperation with its RiPP recognition element (RRE)-containing partner protein PbsD, catalyzing regioselective ortho-hydroxylation of Phe8 and Phe10 to yield ortho-tyrosine residues.133 This modification is a prerequisite for subsequent processing by PbsE, which selectively deguanidinates Arg9 to ornithine but only on the doubly hydroxylated intermediate.133 The final macrocyclization step is mediated by PbsB, which installs a biaryl C–C bond via a radical-based mechanism. It is proposed the [4Fe–4S] cluster of PbsB initiates catalysis by reductive cleavage of SAM to generate a 5′-dAdo˙. This radical abstracts a hydrogen atom from the phenolic hydroxyl group of one ortho-tyrosine, most likely at the Cδ1 position. This results in a stabilized tyrosyl-type radical intermediate. (Fig. 12b).133 NMR and HR-MS/MS analyses confirm regioselective activation at this site. The resulting tyrosyl radical displays significant spin density at the Cε2 position, facilitating intramolecular attack on the Cε2 carbon of the neighboring aromatic ring.133 This radical–radical coupling yields a σ-aromatic radical intermediate, which is subsequently rearomatized through electron and proton transfers.133 The final oxidation step is likely mediated by an auxiliary [4Fe–4S] cluster, similar to those found in other SPASM-domain rSAM enzymes such as StrB. This radical relay culminates in the formation of a rigid biaryl C–C bond that covalently links the two aromatic side chains and defines the macrocyclic architecture (Fig. 12b).133
Although the exact role of the B12 cofactor remains unresolved, given the absence of canonical B12-dependent chemistry such as methylation or alkyl cation transfer, it may participate as a redox mediator or serve a structural role in organizing the active site. Importantly, the requirement for ortho-hydroxylation prior to crosslinking underscores a dual mechanistic function: activation of the aromatic ring toward radical formation and direction of regioselectivity in the ensuing coupling. Notably, both PbsC and PbsB exhibit moderate substrate plasticity, with Phe substrates, PbsC-mediated ortho-hydroxylation is required and enables subsequent PbsE activity, whereas Tyr can be crosslinked by PbsB even without additional ortho-hydroxylation, with the Cε2–Cε2 regiochemistry maintained.133 Collectively, this pathway exemplifies a previously unrecognized catalytic paradigm for B12-rSAM enzymes, extending their reactivity to include aromatic C–C bond formation. The structural novelty of the resulting biaryl-linked macrocycles and the enzymatic precision with which they are assembled highlight the potential for leveraging such systems in the bioengineering of RiPP-based scaffolds with enhanced stability and therapeutic relevance.
1.2. Thioether-containing RiPPs
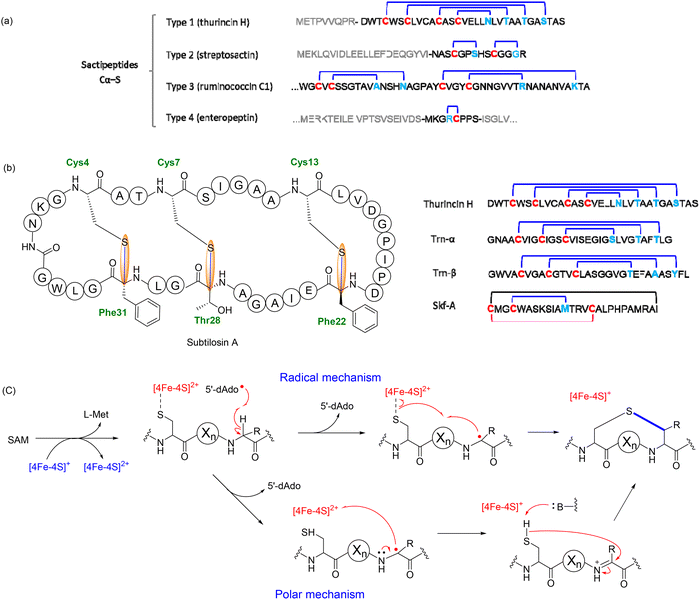
1.2.1.1. Type 1: nested topology. Among the four structural categories, Type 1 sactipeptides are the most extensively studied, particularly with regard to the enzymatic mechanisms underlying thioether bond formation. A well-characterized representative is subtilosin A, a prototypical sactipeptide from Bacillus subtilis that has served as a model for understanding rSAM-mediated cyclization. Subtilosin A is a 35-residue cyclic peptide featuring three Cα–S thioether cross-links.82 These bonds occur between the thiol side chains of cysteine residues and the α-carbons of phenylalanine and threonine, forming a constrained macrocyclic scaffold that is critical for its antimicrobial properties.82 These linkages are installed by AlbA, a rSAM/SPASM protein harboring three [4Fe–4S] clusters. Anaerobic reconstitution of AlbA yielded approximately 7.6 mol Fe and 7.7 mol S per mol of protein, consistent with three [4Fe–4S] centers.82 UV–visible spectra showed absorption features at 325 and 410 nm, characteristic of [4Fe–4S] proteins, which diminished upon chemical reduction.82 EPR spectroscopy further supported this assignment: reduced AlbA exhibited two distinct g-values (2.03 and 1.92) characteristic of [4Fe–4S]+ clusters, and a mutant lacking the canonical CxxxCxxC motif retained one of these signals, indicating an auxiliary cluster.82 Comparative structural modeling and sequence alignment identified seven conserved cysteines in AlbA's C-terminal domain, consistent with the coordination of two auxiliary clusters.82 Catalysis occurs at the [4Fe–4S]RS site. Reductive cleavage of SAM generats the 5′-deoxyadenosyl radical that abstracts a hydrogen atom from the Cα position of the acceptor amino acid, forming a carbon-centered radical. Rather than recombining directly with the donor thiol, this radical is oxidized via electron transfer to an auxiliary [4Fe–4S] cluster in the SPASM domain to give a ketoimine intermediate. The thioether bond is then formed through nucleophilic attack of a cysteine thiolate on this intermediate, completing the Cα–S linkage (Fig. 13b).82,143 This radical-initiated, polar-terminated pathway is consistent with mechanisms proposed for rSAM enzymes that contain SPASM auxiliary clusters, in which the SPASM domain functions as a redox relay and structural scaffold rather than the catalytic center.
Notably, AlbA catalyzes installation of three thioether bonds (Cys4 → Phe31, Cys7 → Thr28, Cys13 → Phe22) and acts sequentially, although the precise site order is not fully established (Fig. 13b).135 This consecutive behavior appears to be structurally enforced: thioether formation is leader-peptide dependent, and a C-terminal SPASM domain harboring an auxiliary [4Fe–4S] cluster carries out catalysis. The leader requirement and interaction of the precursor with the auxiliary cluster suggest substrate positioning during modification. This organization is consistent with vectorial, consecutive engagement of the core peptide, guiding sequential modification.82 This organization is consistent with vectorial, consecutive engagement of the core peptide, guiding sequential modification. This model is strongly supported by deuterium labeling experiments, which demonstrated that AlbA abstracts the Cα hydrogen from Phe31 during the cyclization step forming the Cys4–Phe31 bond.82 When peptides containing deuterated Phe31 were used as substrates, LC-MS analysis detected incorporation of deuterium into the 5′-dAdoH product, with labeling efficiency increasing from ∼20% under dithionite reduction to ∼50% under a physiological reducing system. In contrast, no labeling was observed when deuterium was introduced at the donor cysteine, ruling out hydrogen transfer between residues. These results support that AlbA directly targets the Cα–H of Phe31 and that Cα–H abstraction is rate-limiting for this bond-forming step (Fig. 13c).82,143
Additional examples of Type 1 sactipeptides underscore the structural and functional diversity enabled by this nested topology. SkfA, a 26-residue peptide from Bacillus subtilis, exemplifies a minimal scaffold containing a single Cα–S thioether bond and an intramolecular disulfide linkage (Fig. 13b).134 These features confer a compact, conformationally constrained architecture essential for its role in sporulation-associated cannibalism, where SkfA selectively lyses sibling cells to release nutrients. The Cα–S linkage is installed by the rSAM enzyme SkfB through Cα–H abstraction, as demonstrated by deuterium-labeled substrate studies that confirmed hydrogen transfer to 5′-dAdoH.141 In contrast, Thurincin H, produced by Bacillus thuringiensis SF361, adopts a rigid helical hairpin structure stabilized by four Cα–S thioether bridges. Rather than disrupting membranes, it induces dramatic morphological changes in Gram-positive bacteria such as Listeria monocytogenes, and has been evaluated as a potential food preservative due to its selective antimicrobial activity and exceptional physicochemical stability.137 Another nested sactipeptide system is thuricin CD from B. thuringiensis DPC6431, which comprises two synergistic peptides, Trn-α and Trn-β. Each peptide is approximately 30 residues long and contains three internal thioether cross-links that generate overlapping macrocyclic constraints.139 While individually only weakly active, the peptides function cooperatively to potently inhibit Clostridium difficile with remarkable specificity, sparing most commensal gut microbiota. The highly selective bioactivity and dual-component mechanism of thuricin CD exemplify the potential of nested sactipeptides as microbiome-sparing antibiotics.139
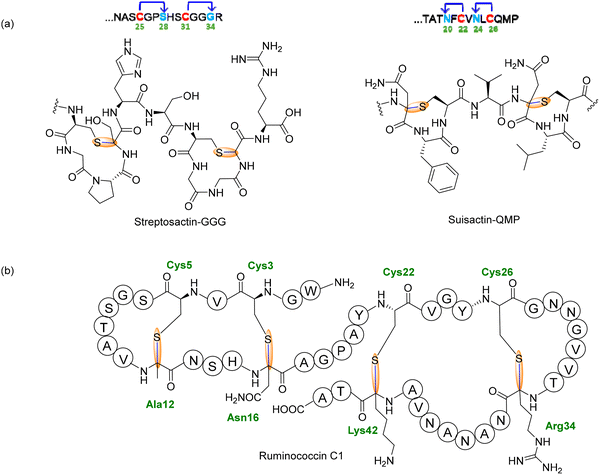
1.2.1.2. Type 2: unnested bicycle topology. Streptosactin, the first sactipeptide identified from Streptococcus thermophilus, exemplifies a novel Type 2 topology, defined by two non-overlapping Cα–S thioether macrocycles arranged in an unnested, sequential manner (Fig. 14a).144 Unlike Type 1 sactipeptides, which feature nested or interleaved crosslinks, the donor (Cys) and acceptor residues in Streptosactin alternate along the linear precursor, generating two discrete four-residue sactionine rings. This distinct architecture expands the topological diversity of sactipeptides and is encoded by the ggg BGC, prevalent in S. thermophilus and other streptococci in the human microbiome.144 Biosynthesis of streptosactin is catalyzed by GggB, a SPASM-domain rSAM enzyme. Heterologously expressed and anaerobically reconstituted GggB displayed spectroscopic features characteristic of rSAM enzymes: UV–vis absorbance at 325 and 410 nm consistent with at least one [4Fe–4S]2+ cluster, and EPR of the reduced protein showing signals for multiple [4Fe–4S]+ clusters with a minor [3Fe–4S]+ component.144 Quantitative analysis revealed ∼9 Fe and ∼8 S atoms per monomer, supporting the presence of a canonical rSAM cluster and two auxiliary clusters in the SPASM domain. HR-MS/MS localized Ser28 and Gly34 as the acceptor residues, with upstream conserved Cys25 and Cys31 as donors. Site-specific 13C-labeled peptide variants coupled with 1D and 2D NMR spectroscopy confirmed the formation of Cα–S thioether linkages at both modified sites.144 Mechanistic experiments using deuterated substrates showed that GggB abstracts the Cα hydrogen to initiate radical-mediated cyclization, transferring deuterium to 5′-dAdoH. N-Methylation of backbone amides inhibited crosslinking, implicating the amide N–H in the catalytic cycle.144 Following in vitro characterization, the mature 14-residue product, streptosactin, was identified in S. thermophilus culture supernatants, matching the enzymatic product by HR-MS and coelution. It exhibits narrow-spectrum, fratricidal activity against strains carrying the same BGC, including the producer itself, suggesting a quorum-sensing-regulated role in competence-associated microbial competition.144
Another representative of Type 2 sactipeptide system is encoded by the qmp gene cluster from Streptococcus suis YS56, which comprises the precursor peptide QmpA, a single SPASM-domain rSAM enzyme QmpB, and a transporter/protease QmpC.145In vitro reconstitution of QmpB with synthetic QmpA, SAM, and dithionite under anaerobic conditions produced a major species 2 Da lighter and a minor 4 Da lighter than the substrate in a ∼5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio, consistent with installation of one and two thioether linkages, respectively. HR-MS/MS analysis localized the modifications to N20–C22 and N24–C26, generating two 9-membered, non-nested macrocycles linked through α-thioether (sactionine) bridges (Fig. 14a).145 Comparison with β-thioether standards confirmed fragmentation behavior characteristic of α-linkages. Using uniformly 13C-labeled Asn at both acceptor sites, DEPT-HSQC spectra revealed diagnostic downfield shifts of the Cβ/Hβ signals (≈42.5/3.7 ppm), providing direct NMR evidence for Cα–S bond formation. Substitution of either cysteine (C22S or C26S) yielded singly cross-linked peptides, indicating that QmpB can cyclize both sites independently without strict order—in contrast to the sequential behavior of the type II sactionine synthase GggB in streptosactin formation. The mature doubly cross-linked product was proposed to be named suisactin. Although its biological activity remains unknown, the compact 9-membered dual-sactionine topology of QmpA expands the mechanistic and structural diversity of type II sactipeptides within the thioether-bond-forming RiPP superfamily.145
1 ratio, consistent with installation of one and two thioether linkages, respectively. HR-MS/MS analysis localized the modifications to N20–C22 and N24–C26, generating two 9-membered, non-nested macrocycles linked through α-thioether (sactionine) bridges (Fig. 14a).145 Comparison with β-thioether standards confirmed fragmentation behavior characteristic of α-linkages. Using uniformly 13C-labeled Asn at both acceptor sites, DEPT-HSQC spectra revealed diagnostic downfield shifts of the Cβ/Hβ signals (≈42.5/3.7 ppm), providing direct NMR evidence for Cα–S bond formation. Substitution of either cysteine (C22S or C26S) yielded singly cross-linked peptides, indicating that QmpB can cyclize both sites independently without strict order—in contrast to the sequential behavior of the type II sactionine synthase GggB in streptosactin formation. The mature doubly cross-linked product was proposed to be named suisactin. Although its biological activity remains unknown, the compact 9-membered dual-sactionine topology of QmpA expands the mechanistic and structural diversity of type II sactipeptides within the thioether-bond-forming RiPP superfamily.145
1.2.1.3. Type 3: bicyclic nested topology. Ruminococcin C1 (RumC1) (Fig. 14b), produced by Ruminococcus gnavus, exemplifies a structurally dense Type 3 sactipeptide characterized by a unique bicyclic nested topology.146,147 Unlike the simpler nested or unnested macrocycles seen in earlier sactipeptides, RumC1 adopts a compact double-hairpin fold stabilized by four Cα–S thioether crosslinks, forming two interlocked macrocyclic domains. This intricate topology not only imparts structural rigidity but also contributes significantly to its potent biological activity. Its gene cluster encodes multiple precursor peptides (RumC1–C5), all yielding active products.146,148–150 The mature RumC1 peptide contains four site-specific thioether bridges between Cys3–Asn16, Cys5–Ala12, Cys22–Lys42, and Cys26–Arg34. These crosslinks organize the peptide into two macrocyclic loops—an N-terminal domain linked through Cys3 and Cys5, and a C-terminal domain linked through Cys22 and Cys26—with overlapping regions that result in a nested configuration (Fig. 14b). Multidimensional NMR analysis confirmed the presence of these thioether bonds, revealing downfield shifts in α-carbon resonances and characteristic NOE contacts between cysteine β-protons and acceptor residue amide protons, consistent with Cα–S bond formation.146
The enzymatic installation of these crosslinks is catalyzed by the rSAM enzymes RumMc1 and RumMc2, both of which contain the canonical CxxxCxxC motif and a C-terminal SPASM domain housing auxiliary [4Fe–4S] clusters.148 Spectroscopic characterizations, including UV–vis and EPR analyses, together with iron and sulfide quantification, confirmed that holo-RumMc1 carries three [4Fe–4S] clusters, one serving as the SAM-binding RS cluster and the other two likely involved in substrate binding or electron transfer.148 Deuterium-labeling experiments provided direct mechanistic evidence for a radical-mediated process. Synthetic peptide substrates with Cα-deuterated alanine residues (e.g., Ala61-d4) demonstrated that RumMc2 initiates thioether bond formation via Cα–H abstraction, generating a Cα-centered radical that is subsequently quenched by a cysteine thiol.148 Mass spectrometry confirmed deuterium loss from the Cα, and concomitant production of 5′-dAdoH further supported SAM cleavage via the classical radical pathway.148 These findings are consistent with prior observations in other sactisynthases such as AlbA, reinforcing the mechanistic conservation across this enzyme family.
Functionally, RumC1 displays potent and selective bactericidal activity against Gram-positive pathogens, including multidrug-resistant Clostridium perfringens.149,150 Notably, it shows no toxicity toward human cell lines or intestinal tissues, highlighting its therapeutic potential. It's structural features confer RumC1 with high in vivo stability, therapeutic bioactivity, and selective bacterial targeting, making it a clinically promising RiPP.146
1.2.1.4. Type 4: thiomorpholine (minimal sactionine). Enteropeptins, a newly characterized family of sactipeptides from Enterococcus cecorum, have been proposed to represent Type 4 sactipeptides, featuring a minimal thioether macrocycle embedded within a largely unstructured peptide framework.142 Unlike previously described sactipeptides, which often contain multiple overlapping macrocycles, enteropeptins adopt a structurally simplified scaffold defined by a single six-membered Cα–S linkage forming a thiomorpholine ring. This distinctive architecture reflects a streamlined biosynthetic strategy while retaining biologically relevant activity. The BGC encodes a precursor peptide (KgrA) and three maturation enzymes: the rSAM enzyme KgrC, the Mn-dependent arginase homolog KgrD and an Fe–S-dependent methyltransferase KgrB. Of these, KgrC has been shown to catalyze the defining thioether crosslink, catalyzing a Cα–S bond between between a conserved arginine residue and the adjacent cysteine in the core peptide (Fig. 15).142In vitro reconstitution showed that KgrC alone converts KgrA to a product 2 Da lighter, consistent with thiomorpholine formation, and HR-MS/MS localized this modification to the conserved Arg residue and its adjacent Cys. The formation of this thiomorpholine macrocycle was confirmed by HR-MS/MS analysis, which pinpointed the crosslink to the α-carbon of Arg41 and the sulfur of Cys42.142 The downstream deguanidination and N-methylation steps were supported by HR-MS/MS together with detailed NMR analysis, corroborating both the α-thioether linkage and the presence of N-methylornithine in the enzymatic produc.142 Consistent with this assignment, KgrB-catalyzed methyl transfer was validated using S-(methyl-d3)-SAM, yielding the expected +3 Da mass shift and MS/MS localization of the deuterated methyl group to the ornithine side chain, ultimately leading to the mature enteropeptin A scaffold.142
Time-course assays monitoring the formation of 5′-dAdoH confirmed that Cα–H abstraction by 5′-dAdo˙ initiates the crosslinking reaction.142 The accumulation of 5′-dAdoH in the presence of SAM and dithionite, but not in control reactions lacking KgrB or reducing agent, confirmed that Cα–H abstraction by 5′-dAdo˙ initiates the crosslinking reaction.142 Additional substrate mutagenesis and analog testing further demonstrated the strict requirement for the reactive cysteine and α-carbon-bearing arginine for efficient cyclization, consistent with a radical-mediated mechanism. The proposed mechanism mirrors that of other sactipeptide synthases, wherein 5′-dAdo˙ abstracts a hydrogen from the α-carbon of the acceptor residue, forming a carbon-centered radical that is trapped by a nearby cysteine thiol to yield the thioether crosslink.142 Biologically, enteropeptin A inhibits the growth of Enterococcus faecalis, a close relative of its producer, suggesting a role in interspecies competition within microbial communities. This narrow-spectrum activity is quorum sensing-regulated and dependent on the presence of the thiomorpholine ring, as analogs lacking the Cα–S bond showed no antibacterial effect.142 Though structurally minimal, enteropeptins illustrate the functional and mechanistic versatility of rSAM enzymes in catalyzing peptide cyclization within reduced frameworks.
1.2.2.1. NxxC product. The unbiased nature of the genome mining from the Streptococcus rSAM-RiPPs by the Seyedsayamdost group also led novel types of thioether containing macrocycles. The NxxC system, originally identified from Streptococcus orisratti and Streptococcus porci, represents a unique subclass of rSAM-catalyzed thioether bond formation, in which a radical-mediated mechanism introduces a β-thioether cross-link between a Cys thiol and the upstream Asn-Cβ residue.60 This distinct type of modification led to the classification of these peptides as ranthipeptides, which are mechanistically and structurally differentiated from sactipeptides, which install α-thioether bonds termed sactionine bridges.75,134,135 Unlike sactipeptides, the Nxxc maturase (NxxcB) catalyzes β-thioetherification. While β-thioether linkages are a characteristic feature of lanthipeptides, where they are introduced heterolytically via an ATP-dependent dehydration step, forming dehydroalanine (Dha) or dehydrobutyrine (Dhb), followed by a Michael addition by Cys. Consequently, β-thioether bonds in lanthipeptide occur specifically at Ser/Thr residues and are chemically and mechanistically distinct from those in ranthipeptides.151,152
In contrast, NxxcB catalyzes radical-mediated β-thioetherification in its native precursor peptide, NxxcA, where Asn serves as the β-thioether acceptor, though alternative residues such as Ala, Gln, and Asp can also be accommodated.60 Functional characterization of NxxcB from Streptococcus orisratti and Streptococcus porci revealed its reliance on multiple auxiliary [4Fe–4S] clusters for radical generation and electron transfer. A proposed mechanistic model suggests that NxxcB functions similarly to sactisynthases by activating the cysteine thiol through chelation to an Fe–S cluster, a process supported by UV-vis absorption shifts upon incubation with NxxcA.60 Catalysis begins with the formation of a 5′-dAdo˙, which abstracts the β-hydrogen of Asn, generating a high-energy radical that subsequently recombines with the Fe–S-activated cysteine thiol, forming the C–S bond while reducing the Fe–S cluster (Fig. 16c).60 High-resolution MS and NMR analyses confirmed that NxxcB exclusively catalyzes Cβ–S bond formation, with no detectable α-thioether linkages, distinguishing it mechanistically from AlbA and other sactisynthases.60,64,76,82 Despite its broad acceptor tolerance, NxxcB does not process Ser or Thr in place of Cys. This restriction remains unexplained and has also been observed in other sactisynthases. Mutagenesis further demonstrated moderate substrate flexibility, allowing modifications at the acceptor site and expanding its potential biocatalytic applications.
1.2.2.2. SCIFF peptides. SCIFF peptides (Six Cysteines in Forty-Five Residues), originally proposed by Haft and Basu in 2011, were initially classified as sactipeptides due to the presence of conserved cysteine-rich motifs and their genomic association with rSAM enzymes.45 However, early attempts to isolate mature SCIFF products were unsuccessful. In vitro reconstitution experiments yielded only partially modified peptides, likely due to the presence of multiple unreacted cysteines and the limited efficiency or specificity of the modifying enzymes under the assay conditions. These technical challenges hindered full structural elucidation and left their chemical nature ambiguous for several years. Subsequent mechanistic and structural studies revealed that SCIFF peptides form thioether linkages at β or γ carbon positions, rather than at the α carbon as seen in canonical sactionine crosslinks.136,153 This structural distinction prompted their reclassification as ranthipeptides. A representative member is the Tte1186a peptide from Caldanaerobacter subterraneus, modified by the SPASM-domain rSAM enzyme Tte1186.64 Despite containing six cysteines, mass spectrometry and in vitro reconstitution demonstrated the selective formation of a single thioether bridge between Cys14 and Thr19 (Fig. 16a), indicative of strict site control. Mechanistic studies supported a radical initiation via SAM cleavage and a potential ketoimine intermediate, highlighting a polar-terminated mechanism facilitated by auxiliary [4Fe–4S] clusters.64
Building upon this foundational work, a closely related system was later identified in Clostridium thermocellum, providing a direct structural and mechanistic link between SCIFF peptides and the emerging ranthipeptide family. In this system, the precursor peptide CteA and its cognate rSAM enzyme CteB share high sequence similarity with Tte1186/Tte1186B and install a thioether cross-link at the conserved Cys14–Thr19 position.76,154 The mature peptide, known as thermocellin, is now classified as a ranthipeptide rather than a sactipeptide, due to its distinct S–Cγ thioether bond between a cysteine sulfur and the γ-carbon of threonine, which contrasts with the typical S–Cα linkages of sactipeptides.153 Structural characterization of CteB revealed a canonical (β/α)6 TIM barrel rSAM fold and a C-terminal SPASM domain containing two auxiliary [4Fe–4S] clusters, one of which presents an open coordination site that facilitates cysteine binding during catalysis.76 Co-crystal structures of CteB with SAM and a CteA-derived peptide fragment provided direct insights into the thioether bond formation mechanism: SAM cleavage generates a 5′-dAdo˙, which abstracts a hydrogen atom from the threonine γ-carbon to form a carbon-centered radical, which then couples with the pre-positioned cysteine thiol to form the S–Cγ linkage.76 These studies further emphasized that site-specific modification in ranthipeptides is dictated not by the number of cysteines, but by precise enzyme–substrate interactions and structural preorganization. Thermocellin thus represents a mechanistically and structurally distinct member of the ranthipeptide family, and preliminary assays have demonstrated antimicrobial activity against Gram-positive bacteria.136 While its detailed bioactivity profile remains to be fully elucidated, the unique thioether topology and conformational rigidity likely contribute to its chemical robustness and potential as a bioactive scaffold, in line with the functional roles proposed for other rSAM-modified RiPPs.
1.2.2.3. PapB family (CXXXD/E motif). The rSAM enzyme PapB, discovered in Paenibacillus polymyxa, defines a distinct class of thioether-forming maturases that install Cβ–S bonds between Cys and Asp/Glu residues within the precursor peptide PapA, giving rise to the characteristic CXXXD/E motif (Fig. 17).155 Belonging to the SPASM-domain subfamily of rSAM enzymes, PapB harbors three [4Fe–4S] clusters: the canonical RS cluster, responsible for SAM cleavage and radical initiation; an auxiliary cluster AC1, which binds and activates the cysteine thiol of the substrate; and a distal AC2, newly recognized as a conduit for electron transfer.156 Mechanistically, the reduced RS cluster generates a 5′-deoxyadenosyl radical that abstracts a hydrogen atom from the β-carbon of Asp or Glu, producing a substrate radical that couples to the cysteine thiolate bound at AC1 to form the Cβ–S linkage (Fig. 17).156 This transformation leaves one electron stored on AC1, which must be returned to the RS cluster to complete the catalytic cycle. Contrary to previous intramolecular models, recent studies revealed that PapB regenerates its active state through intermolecular electron transfer, in which the electron migrates from AC1 through AC2 to an external redox partner such as another PapB molecule, a flavin cofactor, or an unrelated rSAM enzyme, before returning to the RS cluster. This process constitutes an iso-mechanism redox isomerization, establishing a new paradigm for rSAM turnover and explaining how PapB can operate efficiently under mild FMN/NADPH reducing conditions without dedicated flavodoxin/reductase systems.156
Beyond its mechanistic novelty, PapB displays extraordinary substrate promiscuity. Beyond the native CXXXD motif, it accepts Glu as an alternative acceptor to form the corresponding Cys–Cγ linkage, tolerates wide spacing and diverse sequence contexts, and can process D-amino acids at or between the donor and acceptor positions. It catalyzes thioether formation even when the thiol or carboxylate side chains are extended, as in homocysteine or homoglutamate, and when the carboxylate is replaced by a tetrazole bioisostere. Selenocysteine can also serve as an effective donor.157,158 These conclusions are supported by systematic LC–MS/MS analyses that reveal the characteristic −2 Da mass shift, the absence of iodoacetamide alkylation, and fragmentation patterns diagnostic of a Cys–Asp/Glu or isosteric thioether cross-link.157 In parallel, PapB exhibits leader-independent catalysis: it efficiently cross-links substrates containing noncognate or truncated leaders and even those completely lacking leader sequences.158 Remarkably, the enzyme also macrocyclizes C-terminal CSANDA motifs appended to GLP-1-pathway therapeutics such as semaglutide, tirzepatide, and retatrutide, producing fully converted, alkylation-resistant thioether macrocycles. These features establish PapB as a versatile late-stage enzymatic tool for peptide macrocyclization in medicinal chemistry.158
Structural analysis of purified AviCamCys-containing peptides has confirmed the Z-geometry of the newly formed double bond, further highlighting the stereochemical control of the enzyme. Unlike other rSAM enzymes that process free cysteine thiols to generate thioether cross-links, SycC acts only after SycD (a LanD-like protein) mediated oxidative decarboxylation, demonstrating a sequential biosynthetic strategy involving both radical and flavoprotein chemistry.159 In the absence of SycC, the intermediate undergoes hydration-coupled dethiolation to form the shunt aldehyde. This two-step enzymatic process expands the functional landscape of rSAM-dependent RiPP modifications, providing a new mechanism for peptide macrocyclization at the C-terminal region.159 Given the widespread distribution of LanD–rSAM fusion systems in Actinobacteria and Firmicutes, it is likely that similar pathways remain undiscovered, further broadening the biosynthetic and functional diversity of thioether-modified RiPPs.50,162
1.3. Ether crosslinked peptides
The ether bond formation catalyzed by TqqB in the tqq biosynthetic pathway represents a novel rSAM-dependent C–O crosslinking reaction.65 The formation of diverse five-membered ring azoles, such as thiazoles and oxazoles, and their corresponding azolines (thiazolines and oxazolines), is well-documented across biosynthetic pathways. However, six-membered ring heterocycles in RiPPs are comparatively rare, with notable examples including pyridine-containing thiopeptides, the oxidized quinoline in PQQ, and the bicyclic structure in pantocin A.28,35,163–165The tqq cluster initially identified in Streptococcus suis,166 is not exclusive to streptococcal bacteria. Bioinformatic analysis revealed at least six homologous clusters, each encoding a precursor peptide (TqqA) with a conserved threonine residue, suggesting the existence of additional ether-linked RiPPs. This emerging family of RiPPs has been named rotapeptides (radical oxygen-to-alpha-carbon-linked peptides), defined by a unique aliphatic ether linkage between the hydroxyl group of threonine and the α-carbon of an adjacent amino acid, typically glutamine.65 The first characterized subgroup within the rotapeptides is the threoglucins, a set of 22 congeners discovered in S. suis, including Threoglucin A–R and four oxo-threoglucins, all sharing a central 1,3-oxazinane heterocycle.90
TqqB catalyzes the formation of an aliphatic ether bond between the hydroxyl group of a threonine residue and the α-carbon of an adjacent glutamine, leading to the formation of a 4-keto-1,3-tetrahydro-oxazine heterocycle.65 Two potential mechanisms have been proposed for this transformation: radical-mediated direct etherification and an iminium/oxazine route. In the first proposed mechanism, TqqB reductively cleaves SAM, generating a 5′-dAdo˙ that abstracts a hydrogen atom from Gln-Cα, producing a carbon-centered radical intermediate. This radical then undergoes direct recombination with the hydroxyl group of threonine, forming a C–O bond through a radical coupling event (Fig. 19a).65 This mechanism is analogous to rSAM-catalyzed C–C bond formation but instead involves a nucleophilic oxygen species, highlighting the enzyme's ability to control regioselective radical recombination.65 Alternatively, the Gln-Cα radical is deprotonated at the amide nitrogen to an iminium, Thr-Oγ adds to form a 1,3-oxazine intermediate, and oxidation/rearrangement furnishes the same ether product (Fig. 19b).65 The product is a 4-keto-1,3-tetrahydro-oxazine linkage between Thr-Oγ and Gln-Cα.65
Both proposed mechanisms underscore the unique reactivity of rSAM enzymes in RiPP biosynthesis. The catalytic activity of TqqB expands Nature's strategies for incorporating heterocyclic motifs into peptides, marking a significant addition to enzymatic macrocyclization pathways. The resulting threoglucins, characterized by their 1,3-oxazinane heterocycle, exhibit highly specific bioactivity against S. suis. Structure–activity studies reveal that both the heterocycle and a C-terminal Trp-Tyr dyad are essential for function.90 At low concentrations (0.5–4 µM), threoglucins induce bacteriostatic dormancy, enhancing bacterial survival under antibiotic stress, while higher doses (30 µM) are bactericidal, reducing viability to <1%. This dual role suggests they act as ecological mediators. They may promote persistence in harsh conditions or eliminate competitors when resources are abundant. Notably, their activity is exquisitely narrow-spectrum, with no effects on other streptococci or human cells, highlighting their potential as species-specific antimicrobials.90 Moreover, the identification of TqqB's ether bond-forming capability suggests that other uncharacterized rSAM enzymes may catalyze similar oxidative rearrangements, further broadening the functional landscape of radical-based transformations in natural product biosynthesis.65
1.4. C–N crosslinked peptides
Bicyclostreptins represent a subclass of rSAM-RiPPs, characterized by a unique macrocyclic β-ether linkage and an unprecedented C–N crosslink.66 These modifications are catalyzed by two rSAM enzymes, HghB and HghC, encoded in the hgh BGC identified in Streptococcus thermophilus and related species.66 Unlike typical RiPP macrocyclization strategies that target amino acid side chains, bicyclostreptin biosynthesis involves activation of a backbone amide nitrogen, marking the first known instance of radical chemistry facilitating an amide bond crosslink (Fig. 20).66 In the biosynthetic sequence, HghC first catalyzes the formation of a β-ether crosslink between a serine hydroxyl and a histidine Cβ, generating a 13-membered macrocycle. This intermediate is then further modified by HghB, which installs a C–N bond between the backbone amide nitrogen and an adjacent α-carbon, resulting in an imidazolidine-4-one heterocycle. This reaction fundamentally differs from typical radical-mediated crosslinking, as it necessitates the activation of an inherently stable amide bond, such a transformation has not been previously observed in RiPP biosynthesis.32,66,167 This transformation is mechanistically unprecedented in RiPP biosynthesis, as it involves the enzymatic activation of an amide N–H bond to form a covalent crosslink.66 The study have demonstrated that the bicyclostreptin pathway is tightly regulated by quorum sensing (QS) elements, suggesting that these modified peptides may function as bacterial signaling molecules or growth regulators.66 The mature bicyclostreptin peptides were found to exhibit selective bacteriostatic activity against S. thermophilus, implicating them in intra-species competition or population control. Given that bicyclostreptins are produced in nanomolar concentrations, their biological role may extend beyond direct antimicrobial activity to include regulatory interactions within microbial communities.2. Cytochrome P450 enzymes
Cytochromes P450 enzymes (P450s) constitute a vast superfamily of heme-thiolate monooxygenases, playing a significant role in diverse oxidative biochemical transformations.30,168,169 These enzymes derive their name from their characteristic spectroscopic shift, displaying an absorption maximum between 420 and 450 nm when complexed with carbon monoxide in their ferrous state.169,170 The radical-generating capability of cytochrome P450 enzymes originates from their conserved heme-thiolate prosthetic group, which cycles to Compound I (Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, porphyrin π-cation radical). P450 enzymes have been characterized from diverse biosynthetic pathways. More recently, P450s have emerged as versatile biocatalysts capable of mediating complex radical-based cyclization reactions in RiPP biosynthesis. Through their unique heme-dependent catalytic cycle, these enzymes generate high-valent oxo-ferryl intermediates that abstract hydrogen atoms from peptide substrates, creating radical species capable of undergoing controlled intramolecular crosslinking reactions.171
O, porphyrin π-cation radical). P450 enzymes have been characterized from diverse biosynthetic pathways. More recently, P450s have emerged as versatile biocatalysts capable of mediating complex radical-based cyclization reactions in RiPP biosynthesis. Through their unique heme-dependent catalytic cycle, these enzymes generate high-valent oxo-ferryl intermediates that abstract hydrogen atoms from peptide substrates, creating radical species capable of undergoing controlled intramolecular crosslinking reactions.171
Most P450s depend on electron transfer from redox partners to drive their catalytic cycle. In eukaryotic systems, this role is fulfilled by NADPH-cytochrome P450 oxidoreductase (CPR), whereas bacterial and mitochondrial P450s employ a ferredoxin reductase (FdR)/ferredoxin (Fdx) system.172–174 The catalytic cycle begins with substrate binding, which triggers the displacement of a coordinated water molecule from the resting ferric (Fe3+) heme iron, facilitating a shift to a high-spin state that primes the enzyme for reduction. A single electron transfer then reduces the heme iron from ferric (Fe3+) to ferrous (Fe2+), enabling molecular oxygen binding. A second electron transfer reduces the oxygen-bound ferrous intermediate, forming a ferric peroxo species that, upon protonation, generates the ferric hydroperoxo intermediate (commonly referred to as Compound 0).30,168,169,171,172,175–179
The cleavage of the O–O bond in Compound 0 leads to the formation of the highly reactive iron-oxo species, known as Compound I. This potent oxidant abstracts a hydrogen atom from the substrate, creating an iron(IV)-hydroxyl radical (Compound II) (Fig. 21). The resulting substrate radical undergoes a rapid oxygen rebound reaction, leading to hydroxylation or other oxidative modifications, thereby completing the catalytic cycle.30,168,171,172 This remarkable catalytic versatility enables P450s to participate in a wide range of biochemical pathways. Despite this broadly accepted framework, several intermediates, particularly Compound 0 and Compound II, have remained difficult to characterize experimentally, and alternative “shunt” pathways also exist under specific conditions. This remarkable catalytic versatility underpins the ability of P450s to participate in a wide range of biochemical pathways, catalyzing reactions that are often challenging to achieve through classical synthetic chemistry.30,173,174
Here, we focus on the role of P450 enzymes in radical-mediated cyclization within two major classes of natural peptides: NRPs and RiPPs. In peptide cyclization contexts, the substrate radical undergoes intramolecular recombination before the oxygen rebound step, diverting the canonical hydroxylation pathway toward bond-forming reactions.40,180 The spatial arrangement of reactive residues within the enzyme active site dictates the coupling geometry, enabling selective formation of biaryl ether (C–O), biaryl (C–C), or heteroatom-linked crosslinks.40 Recent structural studies of P450-peptide complexes reveal how substrate anchoring motifs position specific amino acid side chains near the heme center for sequential radical generation and coupling.181
2.1. P450 enzymes involved in NRPS biosynthesis
The oxidative cyclization of vancomycin begins with OxyB, which catalyzes the formation of a C–O bond between the phenolic rings of the C and D residues. This reaction is initiated by Compound I (Fig. 22b), an iron(IV)-oxo species that abstracts a hydrogen atom from the C-ring, generating a stabilized radical.184,185 A second oxidation event at the D-ring produces a phenoxy diradical, which recombines to form the first biaryl ether linkage. Nguy et al. have provided direct spectroscopic and kinetic evidence for the formation and functional relevance of Compound I in this transformation.185 Using a peroxide shunt strategy, OxyB was shown to form Compound I upon treatment with m-chloroperbenzoic acid (mCPBA), with its characteristic features at λ = 695 nm confirmed by stopped-flow UV–vis and freeze-quench EPR spectroscopy.185 This revealed the characteristic signal of an iron(IV)-oxo species coupled to a porphyrin radical. Furthermore, the reactivity of this transient intermediate was tested via sequential-mix stopped-flow kinetics, in which the decay rate of Compound I was directly proportional to peptide substrate concentration. The identity of the crosslinked monocyclic product was subsequently confirmed by HPLC-QToF-MS, establishing a direct link between Compound I decay and catalytic turnover.185 These results not only confirm the catalytic competence of Compound I in phenol coupling but also highlight the mechanistic departure from canonical oxygen rebound pathways. Following this, OxyA catalyzes a similar radical-mediated reaction, forming a second biaryl ether bond between the D and E rings, further rigidifying the heptapeptide structure.
Finally, OxyC catalyzes the ring A and ring B crosslink via a C–C coupling reaction, distinct from the previous etherifications.186 This step involves direct radical-radical coupling between aromatic carbons, creating the rigid polycyclic framework critical for vancomycin's antibacterial activity. The catalytic cycle of OxyC is proposed to initiate with the formation of Compound I, the canonical iron(IV)-oxo π-cation radical intermediate common to cytochrome P450 enzymes. This high-valent species abstracts a hydrogen atom from the phenolic hydroxyl group of the Hpg residue (ring A), yielding a phenoxy radical and generating Compound II (Fe(IV)–OH). The radical on Hpg delocalizes to the ortho position, which then undergoes radical phenol coupling with the Dpg ring (ring B), forming a direct biaryl C–C bond. Rearomatization of the resulting intermediate occurs through deprotonation, which is likely mediated by Compound II or an active-site base. This is followed by electron transfer that restores the heme iron to its ferric resting state, thereby completing the catalytic cycle (Fig. 22c).186
This mechanistic model is strongly supported by a comprehensive suite of in vitro experiments. Reconstitution of OxyC activity using synthetic heptapeptide substrates tethered to an X-domain carrier protein enabled the complete chemoenzymatic synthesis of vancomycin aglycone.186 Sequential enzymatic treatment with OxyB, OxyA, and OxyC yielded reaction products that exhibited cumulative mass losses consistent with the installation of the C–O and C–C crosslinks. High-resolution tandem mass spectrometry (HR-MS/MS) confirmed the expected structural changes, including the disappearance of diagnostic b- and y-type ions upon formation of the tricyclic scaffold. Moreover, isotopic labeling with deuterated Hpg provided critical mechanistic insight.186 OxyC-mediated crosslinking resulted in a characteristic 7 Da mass loss, corresponding to the abstraction of both a hydrogen and a deuteron.186 This observation is consistent with a phenol oxidation-initiated process rather than a classical diradical coupling. These results validate the catalytic role of Compound I and provide strong evidence for a refined mechanism involving electrophilic aromatic substitution.
In the glycopeptide-related natural product complestatin, a similar oxidative macrocyclization strategy is employed.187 Two P450 enzymes, ComI and ComJ, catalyze sequential crosslinks. The first is a biaryl C–C bond between 4-hydroxyphenylglycine (Hpg) and tryptophan, and the second is a C–O ether linkage. Together, these modifications construct a rigid bicyclic scaffold (Fig. 22a).188,189 As in vancomycin, these reactions likely proceed via iron(IV)-oxo-mediated generation of phenoxy radicals, which undergo regioselective ortho–ortho coupling.189 Structural studies and reconstitution experiments confirm that ComJ exhibits substrate flexibility and operates on peptides still tethered to the NRPS, with the crosslinks essential for binding bacterial peptidoglycan and inhibiting cell wall remodeling enzymes, offering activity even against vancomycin-resistant strains.189 Notably, complestatin also exhibits a broad range of pharmacological activities, including inhibition of HIV gp120–CD4 binding, anti-apoptotic and neuroprotective effects, and potent antibacterial activity against methicillin-resistant Staphylococcus aureus (MRSA), with a reported MIC of 2 µg mL−1.188,189
While early arylomycins showed potent activity against Gram-positive bacteria including Staphylococcus aureus and Staphylococcus epidermidis,190 their efficacy against Gram-negative pathogens was limited due to outer membrane permeability barriers and reduced SPase binding affinity.195 However, structure-guided optimization led to derivatives such as G0775, which not only penetrates the Gram-negative envelope but also forms a covalent adduct with the SPase active site via a novel oxadiazole warhead, effectively overcoming preexisting resistance mechanisms.196 G0775 exhibits potent in vitro and in vivo activity against multidrug-resistant ESKAPE pathogens, positioning arylomycin analogs as promising leads for new Gram-negative antibiotics.
The formation of the C4–N13 bond in teleocidins is proposed to follow a diradical coupling mechanism, facilitated by iron-oxo intermediates. The reaction is initiated by Compound I, an iron(IV)-oxo species, which abstracts a hydrogen atom from the N1 position of the indole ring, generating a stabilized indole radical. This is followed by a conformational shift in the substrate, bringing the amide N13 position closer to the iron center.210,213 A second hydrogen abstraction from the N13 amide nitrogen by Compound II (FeIV–OH species) results in the formation of a nitrogen-centered radical. The final C–N bond is established through diradical recombination between the C4 and N13 radicals, forming the fused indolactam ring system (Fig. 23b).210 This mechanism was elucidated through in vitro enzymatic assays using TleB and its homologue HinD.213,214 Radical clock experiments using an N13-cyclopropyl-substituted analogue resulted in ring-opened products, confirming the formation of a radical at N13. Substrates lacking the N1-hydrogen failed to undergo cyclization, highlighting the necessity of initial hydrogen abstraction at N1. Furthermore, X-ray crystal structures of TleB and HinD revealed that while N1 is ideally positioned (∼2.7–3.1 Å) for iron-mediated H-abstraction, N13 initially resides too far from the heme. However, a flipped substrate conformation observed in the HinD–substrate complex repositioned N13 within ∼3.1 Å of the heme iron, enabling the second abstraction and bond formation.213 These combined structural and biochemical insights strongly support a stepwise diradical mechanism underlying indolactam formation in teleocidin biosynthesis. Similar mechanisms have been proposed for LtxB in lyngbyatoxin biosynthesis and MpnC in pendolmycin biosynthesis, where P450 enzymes catalyze oxidative cyclization via radical intermediates.215 However, while TleB and LtxB are well-characterized, the mechanistic details of MpnC remain underexplored beyond bioinformatic predictions.
2.2. P450 enzymes involved in RiPP biosynthesis
Early Foundations in rSAM and RiPP Biosynthesis studies explored radical-mediated bond formations in RiPP pathways catalyzed by rSAM/SPASM enzymes, highlighting unique peptide cyclizations.37,143,179,216 These works set a precedent for studying unconventional bond formations in peptide biosynthesis (Fig. 24). While the broader roles of P450s in secondary metabolism have been studied for decades, their specific involvement in RiPP pathways represents a more recent area of investigation.The role of P450s in RiPPs began to gain systematic attention in the mid-to-late 2010s, with isolated reports on oxidative crosslinking, heterocyclization, and tailoring modifications in specific RiPP families. Several reviews published prior to and during the early 2020s have discussed the emerging functions of P450s in RiPP biosynthesis, helping to consolidate early discoveries and mechanistic insights.29,151,176,179,180,217–221 The P450 catalytic cycle facilitates RiPP cyclization through precisely controlled radical recombination events.170 Upon substrate binding, the heme iron undergoes sequential reduction and oxygen activation to form the high-valent Compound I (Fe4+![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O π-cation radical) species. In RiPP-modifying P450s, Compound I abstracts hydrogen atoms from aromatic amino acid side chains (typically tyrosine or tryptophan), generating delocalized substrate radicals that undergo intramolecular coupling rather than oxygen rebound.32,169,180 This radical termination pathway enables: (1) biaryl bond formation: C(sp2)–C(sp3) crosslinks between aromatic residues via radical recombination. (2) Heteroatom linkages: Formation of C–O or C–N bonds through oxygen/nitrogen-centered radical coupling. (3) Macrocycle size control: Precise positioning of reactive residues by the enzyme's architecture dictates ring sizes, typically ranging from 12 to 20 atoms. Notably, these reactions proceed without requiring peptidyl carrier protein (PCP) tethering. This requirement is a critical distinction from non-ribosomal peptide (NRP) cyclization systems and enables direct modification of ribosomally-synthesized precursor peptides and synthetic peptide substrates.178
O π-cation radical) species. In RiPP-modifying P450s, Compound I abstracts hydrogen atoms from aromatic amino acid side chains (typically tyrosine or tryptophan), generating delocalized substrate radicals that undergo intramolecular coupling rather than oxygen rebound.32,169,180 This radical termination pathway enables: (1) biaryl bond formation: C(sp2)–C(sp3) crosslinks between aromatic residues via radical recombination. (2) Heteroatom linkages: Formation of C–O or C–N bonds through oxygen/nitrogen-centered radical coupling. (3) Macrocycle size control: Precise positioning of reactive residues by the enzyme's architecture dictates ring sizes, typically ranging from 12 to 20 atoms. Notably, these reactions proceed without requiring peptidyl carrier protein (PCP) tethering. This requirement is a critical distinction from non-ribosomal peptide (NRP) cyclization systems and enables direct modification of ribosomally-synthesized precursor peptides and synthetic peptide substrates.178
Cihunamides, recently isolated from a Streptomyces sp. derived from a volcanic island, share highly homologous biosynthetic genes encoding a precursor peptide and a P450 enzyme, the P450-installed C–N crosslink between Trp1-C7 and Trp2-N1 and lack atropisomerism.226 An et al. proposed that Trp-Trp crosslinks should serve as the primary classification criterion, rather than final chemical structure. This led to the suggestion that atropitides should be merged into an expanded bitryptide family, where the presence of P450-catalyzed Trp-Trp linkages was the defining characteristic.226 This perspective was based on genomic similarities between tryptorubins and cihunamides, which shared highly homologous BGCs, including a precursor peptide and a dedicated P450 enzyme. The argument was that their biosynthetic commonalities outweighed structural differences such as atropisomerism, supporting a broader classification under bitryptides.227
However, a more recent study identified that bitryptides exhibit a conserved Trp–Trp crosslink motif (xWxxWx), which distinguishes them from other RiPP subclasses. Instead of viewing bitryptides as an overarching category, the authors proposed that bitryptides should be recognized as a specific subgroup within the larger Atropopeptide family. Atropopeptides, in this refined definition, are characterized not just by P450 modifications but also by conserved sequence motifs (such as KSLK in leader peptides) and distinct core peptide patterns, making them a broader RiPP class that includes bitryptides as a subset. The study revealed that Atropopeptide BGCs are distributed across different bacterial genera, primarily in Streptomyces and Xanthomonas.178,222 Biermann et al. also successfully heterologously expressed several Atropopeptide BGCs and elucidated their structures, including: Scabrirubin, Varsorubin B, and Laurentirubin B which is the most complex atropopeptide discovered to date.222
Experimental validation through heterologous expression of bytAO in Streptomyces coelicolor confirmed production of YYH, which contains a biaryl C–C linkage between the Tyr and His side chains. Following these findings, the first biarylitide from myxobacteria, termed myxarylin, was identified from Pyxidicoccus fallax.228 Myxarylin differs from Planomonospora-derived biarylitides in two key aspects: it incorporates an N-methyl group in place of N-acetylation, and it forms a C–N linkage between Tyr and His, reflecting distinct enzymatic processing and regioselectivity. This structural divergence highlights the catalytic flexibility of P450s and suggests evolutionary divergence in biarylitide biosynthesis across bacterial phyla.
Further mechanistic insights were obtained from studies on P450Blt, a BytO-like enzyme from Micromonospora sp. MW-13. This enzyme exhibits broad substrate tolerance and ability to catalyze Tyr-phenol-C to His-imidazole-N crosslinks, similar to myxarylin (Fig. 25a).230 Mutational analysis of its pentapeptide substrate (MRLYH) revealed that Met-1 and Arg-2 are critical for cyclization, while modifications at position 4 affected efficiency. Structural and computational studies further clarified P450Blt's regioselectivity.231 Recent work has expanded the utility of these P450 enzymes as biocatalysts through protein engineering. In a study by Treisman et al., active-site residues of P450Blt (A231, H234, and E238) were mutated to alter the substrate binding and cyclization profile. Engineered variants such as Blt-M1 (A231V-H234L) and Blt-M3 (H234L-E238N) enabled the efficient cyclization of alternative substrates like MRHLY and MRYLY, producing novel tripeptides with His–Tyr or Tyr–Tyr crosslinks (Fig. 25a).232 These engineered P450s achieved up to 85% conversion, and structural elucidation confirmed the formation of unconventional C–O linkages (e.g., Tyr-C3–Tyr-O), distinct from the natural C–C or C–N biaryl bonds.232 This work highlights the catalytic plasticity of biarylitide-type P450s and their potential for generating new macrocyclic peptide scaffolds with tailored properties. Genome mining has since uncovered diverse biarylitide BGCs, leading to the discovery of novel peptides like roseovertin (Trp-C7′ to His-Nτ), rubrin (Trp-C7′ to Tyr-C6), and lapparbin (Tyr-C6 to Trp-N1′), with unique crosslinking chemistries (Fig. 25b).175 Large-scale bioinformatics screening further identified newly crosslinked peptides from multiple P450 subclasses, many featuring N-terminal acylation or methylation, underscoring the expanding biochemical diversity of this RiPP family. These findings highlight the biocatalytic potential of P450 enzymes for engineering novel macrocyclic peptides.
Nocapeptin A, the major product from Nocardia terpenica, contains a C–N biaryl linkage formed between Trp2-N1 and Tyr3-C3. This modification is installed by NopF and was validated using stable isotope labeling, 2D NMR spectroscopy, and LC-MS/MS.236 The resulting product displays a rigid bicyclic architecture that extends beyond the classical macrolactam ring. In contrast, longipeptin A from L. tulufanense contains a Trp–Trp crosslink at analogous positions and undergoes additional tailoring reactions, including tryptophan hydroxylation and methionine S-methylation.237 The latter forms a sulfonium moiety, which represents the first example of this type of modification in RiPPs.236 Functionally, nocapeptin A displays modest antimicrobial activity against Micrococcus luteus with a minimum inhibitory concentration (MIC) of 16 µg mL−1. It does not exhibit cytotoxicity in the NCI-60 cancer cell line panel.236 Its structural rigidity, attributed to both the lasso conformation and the P450-mediated crosslink, contributes to its high resistance to heat and protease degradation. The identification of nocapeptins illustrates how P450 enzymes expand the chemical capabilities of RiPP biosynthetic pathways. This discovery suggests that additional lasso peptides containing diverse oxidative crosslinks are likely to be found through continued genome mining and biochemical exploration.131
Spectroscopic and MicroED studies confirmed the axial R chirality of the macrocyclic biaryl linkage, highlighting ApyO's ability to perform regio- and stereoselective radical phenol coupling. While this architecture mirrors that of arylomycin, it shares structural similarity with the macrocyclic product of the pbs system from Peribacillus simplex (see Section 2.1.8 and Fig. 12). In the pbs pathway from Peribacillus simplex, biaryl macrocyclization is not mediated by a P450, but by PbsB, a B12-dependent rSAM enzyme that operates via a 5′-deoxyadenosyl-radical-initiated pathway, yielding an ortho–ortho′ C–C bond between two ortho-hydroxylated Phe/Tyr residues after PbsC/D-mediated hydroxylation and PbsE-mediated deguanidination.133 While both ApyO and PbsB achieve a similar aromatic C–C bond, their catalytic strategies diverge fundamentally: ApyO proceeds through heme-mediated oxygen activation and phenoxyl radical coupling, whereas PbsB initiates via rSAM-mediated hydrogen abstraction and spin-delocalized radical attack.131 This comparison underscores a broader theme in RiPP biosynthesis: nature employs orthogonal enzymatic strategies to converge on similar structural outcomes. Both P450 and B12-rSAM enzymes can forge biaryl macrocycles, but they do so via distinct catalytic logics, cofactors, and substrate scopes. In the pbs pathway, biaryl macrocyclization is executed by PbsB, a B12-dependent rSAM enzyme, as demonstrated by biochemical reconstitution and mechanistic analysis. This juxtaposition with P450-mediated phenolic couplings highlights how orthogonal enzyme classes can converge on similar macrocyclic outcomes, underscoring nature's versatility in RiPP construction and the value of integrating biochemical and structural evidence when assigning enzyme function.
The most remarkable transformation was observed for SgrB, which formed gristide 834 bearing a His–His ether linkage generated by direct oxygen insertion. This modification was confirmed by a +16 Da mass increase and by diagnostic HMBC and NOESY correlations that established a C–O bridge between the imidazole rings of two histidine residues. Isotopic labeling and spectroscopic data showed that the inserted oxygen atom originates from molecular oxygen and is introduced through the canonical P450 catalytic cycle involving a high-valent Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (Compound I) species. The reaction thus represents the first example of a heme-dependent oxygen-insertion macrocyclization in RiPP biosynthesis. Mechanistically, this process differs fundamentally from the radical relay employed by the rSAM enzyme DarE, which mediates O-insertion in darobactin formation via Fe–S–cluster activation rather than a heme-oxo intermediate.83,120,234 Although their catalytic architectures differ, SgrB and DarE both achieve ether crosslink formation through molecular oxygen incorporation, representing convergent enzymatic solutions for C–O bond formation in RiPP macrocyclization.234 The discovery of these P450 enzymes expands the oxidative repertoire of RiPP biosynthesis beyond C–C and C–N couplings and establishes oxygen insertion as a new dimension of heme-dependent peptide diversification.
O (Compound I) species. The reaction thus represents the first example of a heme-dependent oxygen-insertion macrocyclization in RiPP biosynthesis. Mechanistically, this process differs fundamentally from the radical relay employed by the rSAM enzyme DarE, which mediates O-insertion in darobactin formation via Fe–S–cluster activation rather than a heme-oxo intermediate.83,120,234 Although their catalytic architectures differ, SgrB and DarE both achieve ether crosslink formation through molecular oxygen incorporation, representing convergent enzymatic solutions for C–O bond formation in RiPP macrocyclization.234 The discovery of these P450 enzymes expands the oxidative repertoire of RiPP biosynthesis beyond C–C and C–N couplings and establishes oxygen insertion as a new dimension of heme-dependent peptide diversification.
3. BURP domain proteins
The BURP-domain peptide cyclases (BpCs) form a lineage of post-translational peptide-modifying enzymes that generate macrocyclic products known as burpitides.238 Phylogenetic analyses reveal four subfamilies: USP (Unknown Seed Protein), RD22 (Responsive to Desiccation 22), PG1β (Polygalacturonase-Inhibiting Protein 1β), and BNM2 (Brassica napus Meristem 2), each distinguished by domain architecture and core peptide localization.74,239–241 These BURP domains share a conserved CHX10CHX25–27CHX25–26CH motif that organizes two opposing 2 × CH modules. Structural analyses have revealed a His2–Cu coordination typical of type II copper proteins on one side of the motif, with a potential second site located within 7 Å; whether both are simultaneously occupied and cooperate in O2 activation remains under investigation. This His-ligated copper environment, stabilized by nearby disulfide “staples,” represents an unusual feature among peptide-modifying metalloenzymes.240,242 BURP domain proteins typically consist of multiple modules, including an N-terminal hydrophobic domain signal peptide, a conserved segment, an optional repeat domain, and the C-terminal BURP domain itself.From a structural perspective, BURP-derived macrocycles encompass lyciumin-type C–N linkages, cyclopeptide alkaloids bearing Tyr–O–C ether bonds, stephanotic acid-type Trp–C–C scaffolds, and hibispeptin-type Tyr–C–C frameworks.243 Although these classes differ in ring topology and crosslink chemistry, they are all installed by only two mechanistically distinct enzyme families: fused and split burpitide cyclases, which form the basis of the discussion below.
Building on this classification, several representative molecules illustrate the breadth of BURP-installed side-chain macrocyclization. Moroidin from Celosia argentea contains both a Leu–Trp C–C and a Trp–His C–N linkage; hibispeptin B from Hibiscus syriacus features a Tyr–Leu C–C bond; and arabipeptin A from Coffea arabica carries a Leu–Tyr C–O ether crosslink catalyzed by a split BURP enzyme (Fig. 28).244–247 Despite their shared reliance on radical-mediated copper chemistry, BpCs display diverse expression patterns and biological roles: BNM2 is associated with microspore embryogenesis in Brassica, RD22 is induced by drought stress in Arabidopsis, and BURP domains frequently localize to the cell wall matrix, where they may contribute to intermolecular peptide crosslinking and structural stabilization.
3.1. Fused burpitide cyclases
A subset of BURP enzymes are encoded as fusion proteins that include both the modifying domain and the precursor peptide in the same polypeptide chain, enabling intramolecular processing. Recent structural and biochemical studies established that these fused BURP enzymes operate by a copper- and oxygen-dependent radical mechanism that is distinct from previously characterized RiPP cyclases. The crystal structure of Arachis hypogaea AhyBURP (a USP-type enzyme) uncovered an unprecedented BURP-domain fold, comprising a discontinuous β-barrel and β-sheet that enclose the core peptide above a His2–Cu site and a second potential Cu-binding module positioned less than 7 Å away in the structural model.242 Each active site contains a pair of His ligands stabilized by a Cys–Cys disulfide “staple”, forming two type II Cu sites per monomer. The enzyme functions as a homodimer, but each subunit acts autonomously on its own tethered core peptide, thus, to our knowledge, establishing AhyBURP as the first verified intramolecular (in cis) RiPP macrocyclase.242Mechanistically, AhyBURP catalyzes two sequential oxidations to yield first the monocyclic lyciumin I, followed by the bicyclic legumenin (Fig. 29). In the first step, a copper site is proposed to activate molecular oxygen to generate a reactive Cu–O species (e.g., Cu–superoxo and/or dicopper µ-oxyl formulations) that abstracts a hydrogen atom from Gly82-Cα, generating a carbon-centered radical. This radical couples with the Trp86-N1 to form the C(sp3)–N bond (lyciumin I).242 Radical-trapping experiments using CHANT (4-chloro-3,5-dinitrobenzyl hydroxylamine) provided support for transient radical intermediates localized to the Tyr81–Trp86 region, and isotope-labeling experiments are consistent with O2 serving as the terminal oxidant rather than a donor atom. In the second step, formation of the additional crosslink to afford legumenin is proposed to proceed via H-atom abstraction at Gln79, with Tyr84 implicated in this process based on Gln-d5 labeling (loss of a single deuterium) and the Y84A variant. While alternative mechanistic models have been proposed for BURP-domain enzymes, the scheme shown in Fig. 29 summarizes the key experimentally supported features common to current proposals.238,242
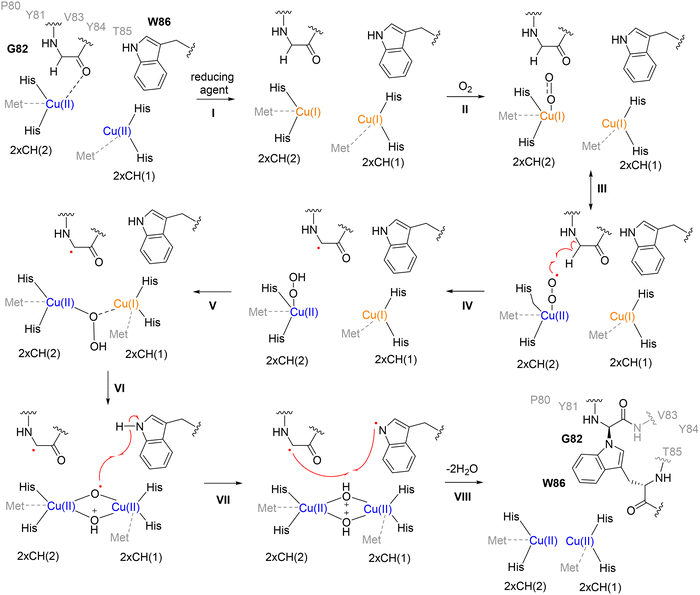 | ||
| Fig. 29 Proposed copper- and oxygen-dependent radical mechanism for AhyBURP-catalyzed macrocyclization. A reactive Cu–O species generated upon O2 activation is proposed to initiate hydrogen atom abstraction, leading to sequential C–N and C–O crosslink formation in lyciumin I and legumenin. Isotope-labeling data support O2 acting as the terminal oxidant without oxygen incorporation into the product, and catalytic turnover is supported by mild reductants such as ascorbate or glutathione. While alternative mechanistic models have been proposed for BURP-domain enzymes, the scheme shown highlights the key experimentally supported features common to current proposals.238,242 | ||
Catalytic turnover of fused BURP enzymes requires a mildly reducing environment; ascorbate or γ-glutamyl–cysteinyl–glycine (GSH) can support activity by maintaining the copper center(s) in a reduced state. An internal Cys104–Cys116 disulfide has been proposed to participate in electron or proton transfer during catalysis, although details of O2 binding/activation and the copper oxidation/coordination states throughout the reaction remain to be fully defined.242 Mutation of any coordinating histidine or disruption of the key disulfide abolishes macrocyclization, underscoring the essential roles of copper coordination and thiol redox dynamics.242
Building on these mechanistic foundations, recent large-scale transcriptome analyses across ≈8900 plant RNA-seq datasets revealed extensive diversification of BURP-type macrocyclases and their products, including newly characterized compounds such as glechomanin (Val–Trp C–C) and mercurialin (Phe–Tyr C–O). These results were reported in subsequent studies building on the initial structural and mechanistic insights (Fig. 28).248 These compounds retain the canonical Leu–Trp C–C linkage that defines the stephanotic acid scaffold but introduce novel secondary crosslinks catalyzed by BURP enzymes with distinct substrate specificities.248 Glechomanin from Glechoma hederacea features an additional Val–Trp C–C bond, while mercurialin from Mercurialis annua forms a Phe–Tyr C–O ether linkage, demonstrating that BURP-domain copper enzymes can mediate both C–C and C–O bond formations between aliphatic and aromatic side chains (Fig. 28).248 In Stellaria aquatica, a newly discovered moroidin variant (QLLVWRNH) exhibits a Trp–His C–N linkage typical of classic moroidins but displays enhanced antiproliferative activity against non-small-cell lung cancer cells (IC50 ≈ 1.4 µM), suggesting functional diversification driven by subtle sequence changes. Collectively, these findings underscore that BURP-domain macrocyclases constitute an evolutionarily convergent platform for oxidative side-chain macrocyclization, generating diverse scaffolds that span stephanotic acid, moroidin, and related plant RiPP families.248
3.2. Split burpitide cyclases
A second, widespread class of BURP enzymes functions in trans on separately encoded precursor peptides. These split BURP systems form conserved biosynthetic cassettes in plant genomes in which short precursor genes lie adjacent to one or more BURP homologs. In Coffea arabica, the split enzyme ArbB2 was shown by in vitro reconstitution to install the Leu–Tyr C–O ether bond of arabipeptin A using the cognate precursor ArbA2.249 The reaction requires copper ions for activity, and the modification is abolished in the presence of a copper chelator, consistent with a Cu-dependent oxidative mechanism as observed for other BURP-type cyclases.245 The corresponding arb locus encodes multiple precursor peptides and several BURP homologs, constituting a modular ensemble for generating macrocyclic peptide diversity.Mechanistic studies on ArbB2, together with structural analyses of fused BURP macrocyclases, indicate that split BURPs retain the conserved His-ligated di-copper motif and O2-dependent oxidative chemistry inferred for fused systems, but act as trans-acting, monomeric enzymes with true catalytic turnover on multiple peptide substrates.249 AlphaFold modeling and docking support a role for an N-terminal leader or recognition sequence upstream of the core peptide in guiding substrate binding, positioning aromatic residues from the core into a hydrophobic pocket adjacent to the copper active site for selective side-chain oxidation.249 Reductant-dependence experiments show that exogenous reducing agents are required to maintain activity and enable multiple turnovers, consistent with reductive regeneration of Cu(I) during catalysis.249 Sequential modification assays further demonstrate that split BURPs can process each core within multi-core precursors and can act on distinct precursor peptides, consistent with an expanded substrate scope.249 Comparative genomics, supported by transcriptome mining, reveals that split BURPs are broadly distributed across eudicots and are frequently co-clustered with stand-alone precursor genes encoding cores for arabipeptins, hibispeptins, cyclopeptide alkaloids, and moroidin-like burpitides.245,249,250 Orthologous split systems from Ceanothus americanus and Ziziphus jujuba are associated with cyclopeptide alkaloids such as frangulanine and homoamericine, whose 13–14-membered macrocycles feature tyrosine-derived C(sp3)–O–phenol ether crosslinks to β-hydroxy Leu or Ile residues.245,251 These data define split BURPs as trans-acting, copper- and oxygen-dependent macrocyclases that operate within a common Cu/O2-dependent oxidative paradigm proposed to underlie both fused and split burpitide cyclases, while providing greater catalytic flexibility and broader product diversity.238
Collectively, fused and split BURP macrocyclases illustrate an emerging Cu/O2-dependent radical strategy for oxidative side-chain macrocyclization. While fused enzymes such as AhyBURP act intramolecularly on tethered substrates, split systems like ArbB2 perform true catalytic turnovers on separate precursor peptides.179 Despite variations in architecture, both share the hallmark His-ligated copper site and generate structurally diverse macrocycles.
4. DUF3328 domain proteins
Fungal RiPPs have been identified for their potential therapeutic applications, including anticancer, antitubulin, antinematode, and immunosuppressant activities.218,221,252 Despite their importance, only a limited number of fungal RiPPs have been discovered, with most belonging to specific groups like cycloamanides, borosins, and dikaritins.221,253–258 Cycloamanides are cyclic peptides with thioether crosslinks formed through oxidative macrocyclization, examples include α-Amanitin and phallacidin derived from the mushroom Amanita bisporigera, but the enzymes responsible for their thioether macrocyclization remain unassigned (Fig. 30a).258–262 Borosins rely on self-processing enzymatic mechanisms for cyclization.253Beyond these families, a recently characterized DUF3328-associated branch, the asperigimycins A–D from Aspergillus flavus and A. oryzae, illustrates that multiple DUF3328 oxidases can be combined to build far more complex scaffolds (Fig. 30b).263 The apg gene cluster encodes a ribosomal precursor (ApgA) together with six DUF3328 enzymes (ApgYa–Yf) and an N-terminal glutaminyl cyclase (ApgG). Stepwise knockouts map the roles of individual DUF3328s in early macrocyclization, hydroxylation, and ether-ring construction, converging on a benzofuranoindoline core that is further elaborated into a heptacyclic framework.263 Deletion of apgA abolishes all products, confirming the RiPP origin, while biochemical assays establish ApgG as the enzyme installing pGlu at the N-terminus. Selected analogs display anticancer activity, and N-terminal lipidation enhances potency, underscoring DUF3328 enzyme cascades as a versatile platform for complex macrocycle assembly and activity tuning.263
Dikaritins constitute a growing class supported by extensive in vivo evidence: gene knockouts and heterologous expression implicate DUF3328 proteins as central to the biosynthesis of ustiloxins,254,255 phomopsins,257 asperipin-2a,256 victorins264 and epichloëcyclins265 (Fig. 30a). These peptides undergo post-translational oxidative modifications, leading to unique C–O crosslinks between Tyr/Phe residues and Cβ of nearby amino acids. For epichloëcyclins, the products are cyclic heptapeptides whose sequences and ring closure are defined by LC–MS/MS, but detailed crosslink patterns, 3D structures, and absolute stereochemistry remain to be fully resolved.265,266
A recent study provided the first in vitro biochemical characterization of AprY, a DUF3328 enzyme from the Aspergillus flavus asperipin-2a pathway. The study successfully reconstituted enzymatic function of AprY, demonstrating that it catalyzes two consecutive C–O crosslinking reactions in the core hexapeptide FYYTGY.267 This process installs two Cβ–O ether crosslinks between tyrosine and adjacent amino acids, enhancing the structural rigidity of the peptide. The first step involves hydrogen atom abstraction at Tyr3 and Tyr6, generating a diradical intermediate, which undergoes radical recombination to form a Tyr3–Tyr6 ether bond. The absence of proton exchange in deuterium-labeled assays supports a radical recombination mechanism rather than a Michael-type nucleophilic addition (Fig. 30c).267 The second crosslinking event involves Phe1 and Tyr3, initiated by oxidative desaturation of Phe1, forming an α,β-unsaturated intermediate. This intermediate undergoes further hydrogen atom abstraction, leading to a Phe1-centered radical, which recombines with Tyr3 to form the second ether crosslink. Finally, the α-amino group of Phe1 is oxidized to an α-ketone, confirmed by 18O incorporation from water, and subsequently stereoselectively reduced by AprR, a short-chain dehydrogenase/reductase, yielding the final asperipin-2a structure.267
Functional expansion: Copper-dependent halogenation within DUF3328. Beyond oxidative macrocyclization, DUF3328 enzymes also mediate copper-dependent C(sp3)–H halogenation: ApnU uses a binuclear Cu(II) center to iteratively halogenate otherwise inert aliphatic C–H bonds and, under O2/ascorbate, can also perform iodination and thiocyanation. Structural and spectroscopic data support a disulfide-linked homodimer with HXXHC-coordinated dicopper centers, broadening DUF3328 chemistry from macrocyclization to C–H functionalization.268
5. Multinuclear non-heme iron dependent oxidative enzymes (MNIOs)
MNIOs, formerly grouped as DUF692 proteins, are emerging peptide-tailoring oxidases that use multinuclear non-heme iron cofactors and molecular oxygen to initiate one-electron chemistry via Fe–O–mediated hydrogen-atom transfer. Across this large and mechanistically versatile family, several distinct oxidative outcomes have been described. The first involves cysteine-centered four-electron oxidations that generate local heterocycles such as oxazolone–thioamide and 5-thiooxazole units, as exemplified by MbnB/MbnC in methanobactin biosynthesis and BufB/BufC in bufferin maturation.269–271 The second encompasses cysteine β-carbon editing, which remodels a single residue without forming a ring, as seen in TglH/TglI.272–274 A third mode features oxidative cascades that culminate in inter-residue macrocyclization, typified by ChrH/ChrI, which form a thioether-linked macrocycle.272 Finally, several MNIOs target acidic residues: MovX-like enzymes catalyze oxidative cleavage of the Asn N–Cα bond to form terminal amides, while ApyH-type oxidases convert Asp to α-keto acids. These reactions collectively underscore the capacity of MNIOs to initiate radical transformations that remodel peptide backbones through oxygen-activated iron centers.269–271Among the characterized members, ChrH, together with its membrane partner ChrI, converts a linear precursor into chryseobasin through a radical initiated, iron dependent redox program that culminates in inter residue thioether macrocyclization (Fig. 31a).272 The proposed mechanism begins with Fe(II)–O2 activation at the active site, generating a superoxo-Fe(III) intermediate that abstracts the Cβ-hydrogen from Cys8, forming a Cβ-centered radical. This intermediate is thought to undergo further iron-mediated oxidation to a thioaldehyde, which may be intercepted intramolecularly by the adjacent Gly amide nitrogen to yield a transient β-lactam. A subsequent iron-catalyzed β-lactam cleavage, plausibly involving a high-valent Fe(IV)-oxo species, promotes carbonyl migration and gives rise to an imidazolidinedione core. A second cysteine residue (Cys5) then attacks the modified backbone to close the thioether macrocycle, and a final SAM-dependent thiomethylation completes the transformation (Fig. 31b). Comprehensive biochemical evidence supports this pathway. Heterologous coexpression of ChrH and its membrane partner ChrI in E. coli produced a +10 Da mass shift only in the presence of both proteins and iron. Site-directed mutagenesis of conserved cysteines disrupted product formation, confirming their essential catalytic roles.272 NMR spectra of the modified peptide (ChrA) showed loss of specific amide resonances and new cross-peaks consistent with an imidazolidinedione and a thioether macrocycle. Isotopic labeling with 13C-cysteine and 13CD3-SAM identified the rearrangement of cysteine-derived carbons and the methyl donor source, while in vitro reconstitution with Fe(II), O2, and SAM fully reproduced the transformation. Together, these results define ChrH as a multinuclear non-heme iron oxidase that channels radical intermediates into selective bond rearrangement and inter-residue cyclization—an archetypal MNIO example of radical enzymatic peptide cyclization.272
The MbnB/MbnC complex represents the best-characterized example of the MNIO family, defining the core oxidative chemistry of methanobactin biosynthesis. Acting on the ribosomally synthesized precursor MbnA, this heterodimeric enzyme catalyzes a four-electron oxidation of conserved cysteine residues to generate the hallmark oxazolone–thioamide motifs that coordinate copper in the mature peptide. Biochemical reconstitution established that MbnB, a TIM-barrel MNIO, requires MbnC for activity and that both Fe and O2 are essential cofactors.269,275 Structural, spectroscopic, and biochemical studies collectively indicate that the active cofactor is a mixed-valent Fe(II)/Fe(III) diiron center, rather than the previously suggested tri-iron cluster. Within the MbnB active site, the cysteine thiolate of MbnA directly coordinates the ferric ion, a feature observed crystallographically in MbnABC complexes and confirmed by electron-nuclear double resonance spectroscopy.270,271,276 Upon O2 binding to the ferrous site, a superoxo-Fe(III) intermediate is generated that abstracts a hydrogen atom from the Cβ of the cysteine residue, producing a substrate radical. This intermediate is subsequently oxidized to a thioaldehyde, which undergoes intramolecular attack by the adjacent amide nitrogen to yield a transient β-lactam species. Concurrently, the ferrous site is oxidized to a high-valent Fe(IV)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (ferryl) species that performs a second hydrogen-atom abstraction from the β-lactam, promoting its conversion into a thio-substituted cyclic imide.270,271 Nucleophilic attack by an enzyme residue and subsequent substitution or tautomerization steps lead to the formation of the oxazolone–thioamide moiety characteristic of methanobactins (Fig. 32). Together, these findings establish MbnB/MbnC as prototypical MNIOs that remodel cysteine side chains through O2-activated radical chemistry, expanding the oxidative and structural diversity achievable in ribosomally synthesized peptides.
O (ferryl) species that performs a second hydrogen-atom abstraction from the β-lactam, promoting its conversion into a thio-substituted cyclic imide.270,271 Nucleophilic attack by an enzyme residue and subsequent substitution or tautomerization steps lead to the formation of the oxazolone–thioamide moiety characteristic of methanobactins (Fig. 32). Together, these findings establish MbnB/MbnC as prototypical MNIOs that remodel cysteine side chains through O2-activated radical chemistry, expanding the oxidative and structural diversity achievable in ribosomally synthesized peptides.
Collectively, these studies support a mechanism in which the substrate Cys thiolate ligates the multinuclear iron center, O2 is activated at the site to effect Cβ–H abstraction and subsequent multi-electron steps, and intramolecular processing yields the oxazolone–thioamide unit that underpins high-affinity copper binding in mature methanobactins. Thus, MbnB/MbnC serve as prototypical MNIOs that remodel side chains to build local heterocycles shaping metal-binding sites, rather than forging new inter-residue rings.272,277,278
Conclusion
The study of radical peptide cyclization enzymes has expanded significantly, encompassing a broad range of enzymatic systems including rSAM enzymes, cytochrome P450s, BURP-domain proteins, and DUF-containing enzymes, all of which contribute to the chemical complexity and functional diversity of natural products. These enzymes facilitate a variety of bond-forming reactions, including C–C, C–S, C–N, and C–O linkages. These modifications enhance the structural stability, bioactivity, and pharmacokinetic properties of peptide-based natural products. A key area of future research lies in deciphering the structural and mechanistic intricacies of these diverse enzymes. While much progress has been made in understanding rSAM enzymology, further investigations are required to elucidate the catalytic potential of P450s, BURP-domain proteins, and DUF-family enzymes. Techniques such as time-resolved crystallography, spectroscopy to detect radicals, and computational prediction and modeling will be instrumental in revealing their underlying reaction mechanisms and engineering enzymes.In the context of peptide modification, a common theme among radical enzymes is their ability to catalyze oxidative reactions on both single residues as well as cross-linking reactions. Further exploration of radical enzymes that have been found to catalyze reactions on single residues may lead to new crosslinking reactions. Examples of this phenomenon were observed for MNIO (DUF692) enzymes and B12 dependent rSAM enzymes. Large superfamilies like αKG and Fe-dependent enzymes may be a fertile area for further exploration. While this review was focused on radical cyclization enzymes, these biosynthetic pathways also encode unique tailoring enzymes to functionalize the macrocycle further or to add additional rings by different enzyme families. Collectively, these paths for exploration will likely lead to chemically diverse peptide scaffolds as a source for next-generation therapeutics.
The discovery of these enzymatic systems has broad implications for drug discovery, particularly in addressing the urgent need for novel antibiotics. Many RiPPs, including darobactins, dynobactins have demonstrated activity against Gram-negative pathogens, making them valuable leads for next-generation antimicrobial agents. Additionally, the ability of radical peptide cyclization enzymes to introduce complex post-translational modifications provides new opportunities for bioengineering stable and functionally diverse therapeutic peptides. Another promising avenue involves harnessing these enzymes for biocatalysis and industrial applications. The regio- and stereoselectivity of radical-mediated transformations offers an innovative approach to peptide engineering, enabling the synthesis of structurally complex molecules with high efficiency. The integration of enzyme engineering, artificial intelligence-driven protein design, and genome mining will further accelerate the discovery of new catalytic functions and the development of novel biotechnological applications.
Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
B.I.M would like to thank the Ministry of Education for funding (A-8000449-00-00, A-0008495-00-00, A-8001694-00-00, A-8002393-00-00, A-8004144-00-00). Z.Y. would like to thank the Chinese Scholarship Council (CSC) for a PhD scholarship.References
- X. Ji, A. L. Nielsen and C. Heinis, Angew. Chem., Int. Ed., 2024, 63, e202308251 CrossRef CAS PubMed.
- A. K. Mishra, J. Choi, S. J. Choi and K. H. Baek, Molecules, 2017, 22, 688–697 CrossRef PubMed.
- D. J. Newman and G. M. Cragg, J. Nat. Prod., 2020, 83, 770–803 CrossRef CAS.
- E. D. Brown and G. D. Wright, Nature, 2016, 529, 336–343 CrossRef CAS PubMed.
- D. Ramirez-Rendon, A. K. Passari, B. Ruiz-Villafan, R. Rodriguez-Sanoja, S. Sanchez and A. L. Demain, Appl. Microbiol. Biotechnol., 2022, 106, 1855–1878 CrossRef CAS PubMed.
- L. Katz and R. H. Baltz, J. Ind. Microbiol. Biotechnol., 2016, 43, 155–176 CrossRef CAS.
- N. Chaachouay and L. Zidane, Drugs Drug-Candidates, 2024, 3, 184–207 CrossRef.
- A. H. Banday, N. u Azha, R. Farooq, S. A. Sheikh, M. A. Ganie, M. N. Parray, H. Mushtaq, I. Hameed and M. A. Lone, Phytochem. Lett., 2024, 59, 124–135 CrossRef CAS.
- X. Chen, S. Varghese, Z. Zhang, J. Du, B. Ruan, J. B. Baell and X. Liu, Eur. J. Med. Chem., 2024, 266, 116126 CrossRef CAS PubMed.
- E. Chain, H. W. Florey, A. D. Gardner, N. G. Heatley, M. A. Jennings, J. Orr-Ewing, A. G. Sanders and L. F. Peltier, Clin. Orthop. Relat. Res., 2005, 439, 23–26 CrossRef CAS PubMed.
- N. Kardos and A. L. Demain, Appl. Microbiol. Biotechnol., 2011, 92, 677–687 CrossRef CAS.
- T. M. Mekhail and M. Markman, Expert Opin. Pharmacother., 2002, 3, 755–766 CrossRef CAS PubMed.
- Y. H. Yang, J. W. Mao and X. L. Tan, Chin. J. Nat. Med., 2020, 18, 890–897 CAS.
- A. Rivera and J. Heitman, PLoS Pathog., 2023, 19, e1011056 CrossRef CAS.
- Y. J. Yoo, H. Kim, S. R. Park and Y. J. Yoon, J. Ind. Microbiol. Biotechnol., 2017, 44, 537–553 CrossRef CAS.
- E. Reich, Cancer Res., 1963, 23, 1428–1441 CAS.
- E. Reich and I. H. Goldberg, in Progress in Nucleic Acid Research and Molecular Biology, ed. J. N. Davidson and W. E. Cohn, Academic Press, 1964, vol. 3, pp. 183–234 Search PubMed.
- E. M. Bertino and G. A. Otterson, Expert Opin. Invest. Drugs, 2011, 20, 1151–1158 CrossRef CAS PubMed.
- G. M. Cragg and D. J. Newman, Biochim. Biophys. Acta, 2013, 1830, 3670–3695 CrossRef CAS.
- A. G. Atanasov, S. B. Zotchev, V. M. Dirsch, T. International Natural Product Sciences and C. T. Supuran, Nat. Rev. Drug Discovery, 2021, 20, 200–216 CrossRef CAS PubMed.
- G. A. Hudson and D. A. Mitchell, Curr. Opin. Microbiol., 2018, 45, 61–69 CrossRef CAS PubMed.
- T. Q. N. Nguyen, Y. W. Tooh, R. Sugiyama, T. P. D. Nguyen, M. Purushothaman, L. C. Leow, K. Hanif, R. H. S. Yong, I. Agatha, F. R. Winnerdy, M. Gugger, A. T. Phan and B. I. Morinaka, Nat. Chem., 2020, 12, 1042–1053 CrossRef CAS.
- C. Ongpipattanakul, E. K. Desormeaux, A. DiCaprio, W. A. van der Donk, D. A. Mitchell and S. K. Nair, Chem. Rev., 2022, 122, 14722–14814 CrossRef CAS PubMed.
- B. Pang, M. Wang and W. Liu, Nat. Prod. Rep., 2016, 33, 162–173 RSC.
- R. F. Little and C. Hertweck, Nat. Prod. Rep., 2022, 39, 163–205 RSC.
- C. J. White and A. K. Yudin, Nat. Chem., 2011, 3, 509–524 CrossRef CAS.
- F. Kopp and M. A. Marahiel, Nat. Prod. Rep., 2007, 24, 735–749 RSC.
- P. G. Arnison, M. J. Bibb, G. Bierbaum, A. A. Bowers, T. S. Bugni, G. Bulaj, J. A. Camarero, D. J. Campopiano, G. L. Challis, J. Clardy, P. D. Cotter, D. J. Craik, M. Dawson, E. Dittmann, S. Donadio, P. C. Dorrestein, K. D. Entian, M. A. Fischbach, J. S. Garavelli, U. Goransson, C. W. Gruber, D. H. Haft, T. K. Hemscheidt, C. Hertweck, C. Hill, A. R. Horswill, M. Jaspars, W. L. Kelly, J. P. Klinman, O. P. Kuipers, A. J. Link, W. Liu, M. A. Marahiel, D. A. Mitchell, G. N. Moll, B. S. Moore, R. Muller, S. K. Nair, I. F. Nes, G. E. Norris, B. M. Olivera, H. Onaka, M. L. Patchett, J. Piel, M. J. Reaney, S. Rebuffat, R. P. Ross, H. G. Sahl, E. W. Schmidt, M. E. Selsted, K. Severinov, B. Shen, K. Sivonen, L. Smith, T. Stein, R. D. Sussmuth, J. R. Tagg, G. L. Tang, A. W. Truman, J. C. Vederas, C. T. Walsh, J. D. Walton, S. C. Wenzel, J. M. Willey and W. A. van der Donk, Nat. Prod. Rep., 2013, 30, 108–160 RSC.
- M. Montalban-Lopez, T. A. Scott, S. Ramesh, I. R. Rahman, A. J. van Heel, J. H. Viel, V. Bandarian, E. Dittmann, O. Genilloud, Y. Goto, M. J. Grande Burgos, C. Hill, S. Kim, J. Koehnke, J. A. Latham, A. J. Link, B. Martinez, S. K. Nair, Y. Nicolet, S. Rebuffat, H. G. Sahl, D. Sareen, E. W. Schmidt, L. Schmitt, K. Severinov, R. D. Sussmuth, A. W. Truman, H. Wang, J. K. Weng, G. P. van Wezel, Q. Zhang, J. Zhong, J. Piel, D. A. Mitchell, O. P. Kuipers and W. A. van der Donk, Nat. Prod. Rep., 2021, 38, 130–239 RSC.
- M. C. Tang, Y. Zou, K. Watanabe, C. T. Walsh and Y. Tang, Chem. Rev., 2017, 117, 5226–5333 CrossRef CAS.
- T. Dang and R. D. Sussmuth, Acc. Chem. Res., 2017, 50, 1566–1576 CrossRef CAS PubMed.
- Y. Shi, Y. Xia, W. Gao, J. Wang, B. Shi and H. Wang, Nat. Prod. Rep., 2025, 42, 763–773 RSC.
- J. S. Davies, J. Pept. Sci., 2003, 9, 471–501 CrossRef CAS.
- T. Gulder and P. S. Baran, Nat. Prod. Rep., 2012, 29, 899–934 RSC.
- C. T. Walsh, Science, 2004, 303, 1805–1810 CrossRef CAS PubMed.
- S. H. Lee, H. Kyung, R. Yokota, T. Goto and T. Oe, Chem. Res. Toxicol., 2015, 28, 59–70 Search PubMed.
- P. Fang, W. K. Pang, S. Xuan, W. L. Chan and K. C. Leung, Chem. Soc. Rev., 2024, 53, 11725–11771 RSC.
- A. Gupta and J. K. Laha, Chem. Rec., 2023, 23, e202300207 CrossRef CAS PubMed.
- K. Chen and F. H. Arnold, Nat. Catal., 2020, 3, 203–213 CrossRef CAS.
- Y. Ye, H. Fu and T. K. Hyster, J. Ind. Microbiol. Biotechnol., 2021, 48, kuab021 CrossRef CAS.
- A. Kostenko, Y. Lien, A. Mendauletova, T. Ngendahimana, I. M. Novitskiy, S. S. Eaton and J. A. Latham, J. Biol. Chem., 2022, 298, 101881 CrossRef CAS PubMed.
- Y. Lien, J. C. Lachowicz, A. Mendauletova, C. Zizola, T. Ngendahimana, A. Kostenko, S. S. Eaton, J. A. Latham and T. L. Grove, ACS Chem. Biol., 2024, 19, 370–379 CrossRef CAS.
- J. K. Lewis, A. S. Jochimsen, S. J. Lefave, A. P. Young, W. M. Kincannon, A. G. Roberts, M. T. Kieber-Emmons and V. Bandarian, Biochemistry, 2021, 60, 3347–3361 CrossRef CAS PubMed.
- D. H. Haft, BMC Genomics, 2011, 12, 21 CrossRef CAS.
- D. H. Haft and M. K. Basu, J. Bacteriol., 2011, 193, 2745–2755 CrossRef CAS PubMed.
- J. B. Broderick, B. R. Duffus, K. S. Duschene and E. M. Shepard, Chem. Rev., 2014, 114, 4229–4317 CrossRef CAS.
- A. Benjdia, C. Balty and O. Berteau, Front. Chem., 2017, 5, 87 CrossRef PubMed.
- N. Mahanta, G. A. Hudson and D. A. Mitchell, Biochemistry, 2017, 56, 5229–5244 CrossRef CAS PubMed.
- Q. Guo and B. I. Morinaka, Curr. Opin. Chem. Biol., 2024, 81, 102483 CrossRef CAS PubMed.
- N. Oberg, T. W. Precord, D. A. Mitchell and J. A. Gerlt, ACS Bio Med Chem Au, 2022, 2, 22–35 CrossRef CAS.
- Y. Nicolet, Nat. Catal., 2020, 3, 337–350 CrossRef CAS.
- B. M. Hoffman, W. E. Broderick and J. B. Broderick, Annu. Rev. Biochem., 2023, 92, 333–349 CrossRef CAS.
- M. Horitani, K. Shisler, W. E. Broderick, R. U. Hutcheson, K. S. Duschene, A. R. Marts, B. M. Hoffman and J. B. Broderick, Science, 2016, 352, 822–825 CrossRef CAS PubMed.
- M. Dong, Y. Zhang and H. Lin, Biochemistry, 2018, 57, 3454–3459 CrossRef CAS.
- A. S. Byer, H. Yang, E. C. McDaniel, V. Kathiresan, S. Impano, A. Pagnier, H. Watts, C. Denler, A. L. Vagstad, J. Piel, K. S. Duschene, E. M. Shepard, T. P. Shields, L. G. Scott, E. A. Lilla, K. Yokoyama, W. E. Broderick, B. M. Hoffman and J. B. Broderick, J. Am. Chem. Soc., 2018, 140, 8634–8638 CrossRef CAS.
- M. B. Ho, R. J. Jodts, Y. Kim, A. McSkimming, D. L. M. Suess and B. M. Hoffman, J. Am. Chem. Soc., 2022, 144, 17642–17650 CrossRef CAS PubMed.
- P. H. Donnan and S. O. Mansoorabadi, J. Am. Chem. Soc., 2022, 144, 3381–3385 CrossRef CAS PubMed.
- R. J. Jodts, M. Wittkop, M. B. Ho, W. E. Broderick, J. B. Broderick, B. M. Hoffman and M. A. Mosquera, J. Am. Chem. Soc., 2023, 145, 13879–13887 CrossRef CAS.
- J. Wang, R. P. Woldring, G. D. Roman-Melendez, A. M. McClain, B. R. Alzua and E. N. Marsh, ACS Chem. Biol., 2014, 9, 1929–1938 CrossRef CAS.
- A. Caruso, L. B. Bushin, K. A. Clark, R. J. Martinie and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2019, 141, 990–997 CrossRef CAS.
- W. Zhu, A. T. Iavarone and J. P. Klinman, ACS Cent. Sci., 2024, 10, 251–263 CrossRef CAS.
- K. R. Schramma, L. B. Bushin and M. R. Seyedsayamdost, Nat. Chem., 2015, 7, 431–437 CrossRef CAS.
- K. M. Davis, K. R. Schramma, W. A. Hansen, J. P. Bacik, S. D. Khare, M. R. Seyedsayamdost and N. Ando, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 10420–10425 CrossRef CAS.
- N. A. Bruender, J. Wilcoxen, R. D. Britt and V. Bandarian, Biochemistry, 2016, 55, 2122–2134 CrossRef CAS PubMed.
- K. A. Clark, L. B. Bushin and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2019, 141, 10610–10615 CrossRef CAS PubMed.
- L. B. Bushin, B. C. Covington, K. A. Clark, A. Caruso and M. R. Seyedsayamdost, Nat. Chem. Biol., 2022, 18, 1135–1143 CrossRef CAS.
- M. W. Ruszczycky and H. W. Liu, Biochemistry, 2024, 63, 3161–3183 CrossRef CAS PubMed.
- J. A. Latham, I. Barr and J. P. Klinman, J. Biol. Chem., 2017, 292, 16397–16405 CrossRef CAS PubMed.
- C. S. Phan and B. I. Morinaka, Nat. Prod. Rep., 2024, 41, 708–720 RSC.
- T. A. Grell, P. J. Goldman and C. L. Drennan, J. Biol. Chem., 2015, 290, 3964–3971 CrossRef CAS.
- T. L. Grove, J. H. Ahlum, R. M. Qin, N. D. Lanz, M. I. Radle, C. Krebs and S. J. Booker, Biochemistry, 2013, 52, 2874–2887 CrossRef CAS PubMed.
- S. J. Booker and T. L. Grove, F1000 Biol. Rep., 2010, 2, 52 Search PubMed.
- K. A. Clark, L. B. Bushin and M. R. Seyedsayamdost, ACS Bio Med Chem Au, 2022, 2, 328–339 CrossRef CAS PubMed.
- D. Richter and J. Piel, Curr. Opin. Chem. Biol., 2024, 80, 102463 CrossRef CAS.
- L. Fluhe and M. A. Marahiel, Curr. Opin. Chem. Biol., 2013, 17, 605–612 CrossRef.
- T. L. Grove, P. M. Himes, S. Hwang, H. Yumerefendi, J. B. Bonanno, B. Kuhlman, S. C. Almo and A. A. Bowers, J. Am. Chem. Soc., 2017, 139, 11734–11744 CrossRef CAS.
- B. I. Morinaka, E. Lakis, M. Verest, M. J. Helf, T. Scalvenzi, A. L. Vagstad, J. Sims, S. Sunagawa, M. Gugger and J. Piel, Science, 2018, 359, 779–782 CrossRef CAS.
- N. A. O'Leary, M. W. Wright, J. R. Brister, S. Ciufo, D. Haddad, R. McVeigh, B. Rajput, B. Robbertse, B. Smith-White, D. Ako-Adjei, A. Astashyn, A. Badretdin, Y. Bao, O. Blinkova, V. Brover, V. Chetvernin, J. Choi, E. Cox, O. Ermolaeva, C. M. Farrell, T. Goldfarb, T. Gupta, D. Haft, E. Hatcher, W. Hlavina, V. S. Joardar, V. K. Kodali, W. Li, D. Maglott, P. Masterson, K. M. McGarvey, M. R. Murphy, K. O'Neill, S. Pujar, S. H. Rangwala, D. Rausch, L. D. Riddick, C. Schoch, A. Shkeda, S. S. Storz, H. Sun, F. Thibaud-Nissen, I. Tolstoy, R. E. Tully, A. R. Vatsan, C. Wallin, D. Webb, W. Wu, M. J. Landrum, A. Kimchi, T. Tatusova, M. DiCuccio, P. Kitts, T. D. Murphy and K. D. Pruitt, Nucleic Acids Res., 2015, 44, D733–D745 CrossRef PubMed.
- T. U. Consortium, Nucleic Acids Res., 2020, 49, D480–D489 CrossRef PubMed.
- M. Hadjithomas, I.-M. A. Chen, K. Chu, A. Ratner, K. Palaniappan, E. Szeto, J. Huang, T. Reddy, P. Cimermančič and M. A. Fischbach, mBio, 2015, 6, 00932 CrossRef.
- D. H. Haft, J. D. Selengut and O. White, Nucleic Acids Res., 2003, 31, 371–373 CrossRef CAS.
- L. Fluhe, T. A. Knappe, M. J. Gattner, A. Schafer, O. Burghaus, U. Linne and M. A. Marahiel, Nat. Chem. Biol., 2012, 8, 350–357 CrossRef PubMed.
- Y. Imai, K. J. Meyer, A. Iinishi, Q. Favre-Godal, R. Green, S. Manuse, M. Caboni, M. Mori, S. Niles, M. Ghiglieri, C. Honrao, X. Ma, J. J. Guo, A. Makriyannis, L. Linares-Otoya, N. Bohringer, Z. G. Wuisan, H. Kaur, R. Wu, A. Mateus, A. Typas, M. M. Savitski, J. L. Espinoza, A. O'Rourke, K. E. Nelson, S. Hiller, N. Noinaj, T. F. Schaberle, A. D'Onofrio and K. Lewis, Nature, 2019, 576, 459–464 CrossRef CAS.
- K. A. Clark and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2022, 144, 17876–17888 CrossRef CAS.
- B. E. Rued, B. C. Covington, L. B. Bushin, G. Szewczyk, I. Laczkovich, M. R. Seyedsayamdost and M. J. Federle, mBio, 2021, 12 Search PubMed.
- L. B. Bushin, K. A. Clark, I. Pelczer and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2018, 140, 17674–17684 CrossRef CAS PubMed.
- K. R. Schramma and M. R. Seyedsayamdost, ACS Chem. Biol., 2017, 12, 922–927 CrossRef CAS.
- K. R. Schramma, C. C. Forneris, A. Caruso and M. R. Seyedsayamdost, Biochemistry, 2018, 57, 461–468 CrossRef CAS PubMed.
- A. R. Balo, A. Caruso, L. Tao, D. J. Tantillo, M. R. Seyedsayamdost and R. D. Britt, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2101571118 CrossRef CAS PubMed.
- B. C. Covington and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2022, 144, 14997–15001 CrossRef CAS PubMed.
- A. Caruso, R. J. Martinie, L. B. Bushin and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2019, 141, 16610–16614 Search PubMed.
- I. Barr, J. A. Latham, A. T. Iavarone, T. Chantarojsiri, J. D. Hwang and J. P. Klinman, J. Biol. Chem., 2016, 291, 8877–8884 Search PubMed.
- W. Zhu and J. P. Klinman, Curr. Opin. Chem. Biol., 2020, 59, 93–103 Search PubMed.
- J. A. Duine, J. Biosci. Bioeng., 1999, 88, 231–236 CrossRef CAS PubMed.
- W. Zhu, A. M. Martins and J. P. Klinman, Methods Enzymol., 2018, 606, 389–420 CAS.
- J. A. Latham, A. T. Iavarone, I. Barr, P. V. Juthani and J. P. Klinman, J. Biol. Chem., 2015, 290, 12908–12918 Search PubMed.
- B. M. Wieckowski, J. D. Hegemann, A. Mielcarek, L. Boss, O. Burghaus and M. A. Marahiel, FEBS Lett., 2015, 589, 1802–1806 Search PubMed.
- E. M. Koehn, J. A. Latham, T. Armand, R. L. Evans III, X. Tu, C. M. Wilmot, A. T. Iavarone and J. P. Klinman, J. Am. Chem. Soc., 2019, 141, 4398–4405 Search PubMed.
- J. P. Klinman and F. Bonnot, Chem. Rev., 2014, 114, 4343–4365 Search PubMed.
- I. Barr, T. A. Stich, A. S. Gizzi, T. L. Grove, J. B. Bonanno, J. A. Latham, T. Chung, C. M. Wilmot, R. D. Britt, S. C. Almo and J. P. Klinman, Biochemistry, 2018, 57, 1306–1315 CrossRef CAS PubMed.
- L. Tao, W. Zhu, J. P. Klinman and R. D. Britt, Biochemistry, 2019, 58, 5173–5187 CrossRef CAS.
- W. Zhu, L. M. Walker, L. Tao, A. T. Iavarone, X. Wei, R. D. Britt, S. J. Elliott and J. P. Klinman, J. Am. Chem. Soc., 2020, 142, 12620–12634 CrossRef CAS.
- R. D. Miller, A. Iinishi, S. M. Modaresi, B. K. Yoo, T. D. Curtis, P. J. Lariviere, L. Liang, S. Son, S. Nicolau, R. Bargabos, M. Morrissette, M. F. Gates, N. Pitt, R. P. Jakob, P. Rath, T. Maier, A. G. Malyutin, J. T. Kaiser, S. Niles, B. Karavas, M. Ghiglieri, S. E. J. Bowman, D. C. Rees, S. Hiller and K. Lewis, Nat. Microbiol., 2022, 7, 1661–1672 Search PubMed.
- R. Sugiyama, A. F. L. Suarez, Y. Morishita, T. Q. N. Nguyen, Y. W. Tooh, M. Roslan, J. Lo Choy, Q. Su, W. Y. Goh, G. A. Gunawan, F. T. Wong and B. I. Morinaka, J. Am. Chem. Soc., 2022, 144, 11580–11593 CrossRef CAS.
- S. Guo, S. Wang, S. Ma, Z. Deng, W. Ding and Q. Zhang, Nat. Commun., 2022, 13, 2361 CrossRef CAS.
- S. Ma, H. Chen, H. Li, X. Ji, Z. Deng, W. Ding and Q. Zhang, Angew. Chem., Int. Ed., 2021, 60, 19957–19964 CrossRef CAS PubMed.
- Y. Han, S. Ma and Q. Zhang, Chin. Chem. Lett., 2023, 34 Search PubMed.
- C. S. Phan, L. Chang, T. Q. N. Nguyen, A. F. L. Suarez, X. H. Ho, H. Chen, I. Y. F. Koh and B. I. Morinaka, ACS Chem. Biol., 2024, 19, 855–860 CrossRef CAS.
- M. Purushothaman, L. Chang, R. J. Zhong and B. I. Morinaka, ACS Chem. Biol., 2024, 19, 1229–1236 CrossRef CAS PubMed.
- Y. Morishita, S. Ma, E. De La Mora, H. Li, H. Chen, X. Ji, A. Usclat, P. Amara, R. Sugiyama, Y. W. Tooh, G. Gunawan, J. Perard, Y. Nicolet, Q. Zhang and B. I. Morinaka, Nat. Chem., 2024, 16, 1882–1893 CrossRef CAS.
- S. Ma, H. Chen, S. Liu, X. Huang, T. Mo, W. Q. Liu, W. Zhang, W. Ding and Q. Zhang, Bioorg. Med. Chem. Lett., 2024, 101, 129653 CrossRef CAS PubMed.
- P. Wu, W. Shui, B. L. Carlson, N. Hu, D. Rabuka, J. Lee and C. R. Bertozzi, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 3000–3005 Search PubMed.
- A. F. L. Suarez, T. Q. N. Nguyen, L. Chang, Y. W. Tooh, R. H. S. Yong, L. C. Leow, I. Y. F. Koh, H. Chen, J. W. H. Koh, A. Selvanayagam, V. Lim, Y. E. Tan, I. Agatha, F. R. Winnerdy and B. I. Morinaka, ACS Chem. Biol., 2024, 19, 774–783 CrossRef CAS PubMed.
- C. S. Phan and B. I. Morinaka, ACS Chem. Biol., 2022, 17, 3284–3289 CrossRef CAS.
- J. Ruel, T. Q. N. Nguyen, Y. Morishita, A. Usclat, L. Martin, P. Amara, S. Kieffer-Jaquinod, M. C. Stefanoiu, E. de la Mora, B. I. Morinaka and Y. Nicolet, J. Am. Chem. Soc., 2025, 147, 16850–16863 CrossRef CAS PubMed.
- C. S. Phan and B. I. Morinaka, RSC Chem. Biol., 2024, 5, 1195–1200 Search PubMed.
- A. H. Khan, J. R. Haedar, V. Kiselov, V. Romanuks, G. Smits, S. Donadio and C. S. Phan, ACS Chem. Biol., 2025, 20, 259–265 Search PubMed.
- A. H. Khan, J. R. Haedar, A. Zīle and C.-S. Phan, Tetrahedron Chem., 2025, 14, 100128 CrossRef CAS.
- H. Kaur, R. P. Jakob, J. K. Marzinek, R. Green, Y. Imai, J. R. Bolla, E. Agustoni, C. V. Robinson, P. J. Bond, K. Lewis, T. Maier and S. Hiller, Nature, 2021, 593, 125–129 CrossRef CAS PubMed.
- H. Nguyen, I. D. Made Kresna, N. Bohringer, J. Ruel, E. Mora, J. C. Kramer, K. Lewis, Y. Nicolet, T. F. Schaberle and K. Yokoyama, J. Am. Chem. Soc., 2022, 144, 18876–18886 Search PubMed.
- A. M. Woodard, F. Peccati, C. D. Navo, G. Jimenez-Oses and D. A. Mitchell, J. Am. Chem. Soc., 2024, 146, 14328–14340 CrossRef CAS.
- S. Ma, W. Xi, S. Wang, H. Chen, S. Guo, T. Mo, W. Chen, Z. Deng, F. Chen, W. Ding and Q. Zhang, J. Am. Chem. Soc., 2023, 145, 22945–22953 CrossRef CAS PubMed.
- B. X. Nguyen, F. F. Gurusinga, U. Mettal, T. F. Schaberle and K. Yokoyama, J. Am. Chem. Soc., 2024, 146, 31715–31732 Search PubMed.
- Y. Han, H. Li, X. Li, W. Ding and Q. Zhang, CCS Chem., 2025, 7, 59–67 CrossRef CAS.
- E. D. Badding, T. L. Grove, L. K. Gadsby, J. W. LaMattina, A. K. Boal and S. J. Booker, J. Am. Chem. Soc., 2017, 139, 5896–5905 Search PubMed.
- B. Wang, J. W. LaMattina, S. L. Marshall and S. J. Booker, J. Am. Chem. Soc., 2019, 141, 5788–5797 CrossRef CAS PubMed.
- J. W. LaMattina, B. Wang, E. D. Badding, L. K. Gadsby, T. L. Grove and S. J. Booker, J. Am. Chem. Soc., 2017, 139, 17438–17445 Search PubMed.
- X. Ji, D. Mandalapu, J. Cheng, W. Ding and Q. Zhang, Angew. Chem., Int. Ed., 2018, 57, 6601–6604 Search PubMed.
- Y. Yu, L. Duan, Q. Zhang, R. Liao, Y. Ding, H. Pan, E. Wendt-Pienkowski, G. Tang, B. Shen and W. Liu, ACS Chem. Biol., 2009, 4, 855–864 CrossRef CAS PubMed.
- W. Ding, Y. Li, J. Zhao, X. Ji, T. Mo, H. Qianzhu, T. Tu, Z. Deng, Y. Yu, F. Chen and Q. Zhang, Angew. Chem., Int. Ed., 2017, 56, 3857–3861 Search PubMed.
- D. T. Nguyen, L. Zhu, D. L. Gray, T. J. Woods, C. Padhi, K. M. Flatt, D. A. Mitchell and W. A. van der Donk, ACS Cent. Sci., 2024, 10, 1022–1032 Search PubMed.
- J. Bridwell-Rabb, B. Li and C. L. Drennan, ACS Bio Med Chem Au, 2022, 2, 173–186 CrossRef CAS.
- C. Padhi, L. Zhu, J. Y. Chen, R. Moreira and W. A. van der Donk, bioRxiv, 2025, preprint DOI:10.1101/2025.04.04.647296.
- L. Fluhe, O. Burghaus, B. M. Wieckowski, T. W. Giessen, U. Linne and M. A. Marahiel, J. Am. Chem. Soc., 2013, 135, 959–962 CrossRef.
- Y. Chen, J. Wang, G. Li, Y. Yang and W. Ding, Front. Chem., 2021, 9, 595991 CrossRef CAS.
- G. A. Hudson, B. J. Burkhart, A. J. DiCaprio, C. J. Schwalen, B. Kille, T. V. Pogorelov and D. A. Mitchell, J. Am. Chem. Soc., 2019, 141, 8228–8238 CrossRef CAS PubMed.
- C. S. Sit, M. J. van Belkum, R. T. McKay, R. W. Worobo and J. C. Vederas, Angew. Chem., 2011, 123, 8877–8880 CrossRef.
- W.-T. Liu, Y.-L. Yang, Y. Xu, A. Lamsa, N. M. Haste, J. Y. Yang, J. Ng, D. Gonzalez, C. D. Ellermeier and P. D. Straight, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 16286–16290 CrossRef CAS PubMed.
- M. C. Rea, C. S. Sit, E. Clayton, P. M. O'Connor, R. M. Whittal, J. Zheng, J. C. Vederas, R. P. Ross and C. Hill, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 9352–9357 CrossRef CAS PubMed.
- K. E. Kawulka, T. Sprules, C. M. Diaper, R. M. Whittal, R. T. McKay, P. Mercier, P. Zuber and J. C. Vederas, Biochemistry, 2004, 43, 3385–3395 CrossRef CAS.
- N. A. Bruender and V. Bandarian, Biochemistry, 2016, 55, 4131–4134 CrossRef CAS.
- K. A. Clark, B. C. Covington and M. R. Seyedsayamdost, Nat. Chem., 2022, 14, 1390–1398 Search PubMed.
- A. Benjdia, A. Guillot, B. Lefranc, H. Vaudry, J. Leprince and O. Berteau, Chem. Commun., 2016, 52, 6249–6252 Search PubMed.
- L. B. Bushin, B. C. Covington, B. E. Rued, M. J. Federle and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2020, 142, 16265–16275 CrossRef CAS.
- A. Caruso and M. R. Seyedsayamdost, J. Org. Chem., 2021, 86, 11284–11289 Search PubMed.
- S. Chiumento, C. Roblin, S. Kieffer-Jaquinod, S. Tachon, C. Leprètre, C. Basset, D. Aditiyarini, H. Olleik, C. Nicoletti, O. Bornet, O. Iranzo, M. Maresca, R. Hardré, M. Fons, T. Giardina, E. Devillard, F. Guerlesquin, Y. Couté, M. Atta, J. Perrier, M. Lafond and V. Duarte, Sci. Adv., 2019, 5, eaaw9969 CrossRef CAS PubMed.
- A. Pujol, E. H. Crost, G. Simon, V. Barbe, D. Vallenet, A. Gomez and M. Fons, FEMS Microbiol. Ecol., 2011, 78, 405–415 CrossRef CAS.
- C. Balty, A. Guillot, L. Fradale, C. Brewee, B. Lefranc, C. Herrero, C. Sandstrom, J. Leprince, O. Berteau and A. Benjdia, J. Biol. Chem., 2020, 295, 16665–16677 Search PubMed.
- C. Roblin, S. Chiumento, O. Bornet, M. Nouailler, C. S. Muller, K. Jeannot, C. Basset, S. Kieffer-Jaquinod, Y. Coute, S. Torelli, L. Le Pape, V. Schunemann, H. Olleik, B. De La Villeon, P. Sockeel, E. Di Pasquale, C. Nicoletti, N. Vidal, L. Poljak, O. Iranzo, T. Giardina, M. Fons, E. Devillard, P. Polard, M. Maresca, J. Perrier, M. Atta, F. Guerlesquin, M. Lafond and V. Duarte, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 19168–19177 Search PubMed.
- L. Shamseddine, C. Roblin, I. Veyrier, C. Basset, L. De Macedo, A. Boyeldieu, M. Maresca, C. Nicoletti, G. Brasseur and S. Kieffer-Jaquinod, Iscience, 2023, 26, 107563 CrossRef CAS PubMed.
- H. Lee and W. A. van der Donk, Annu. Rev. Biochem., 2022, 91, 269–294 CrossRef CAS.
- J. D. Hegemann and R. D. Sussmuth, RSC Chem. Biol., 2020, 1, 110–127 RSC.
- Y. Chen, Y. Yang, X. Ji, R. Zhao, G. Li, Y. Gu, A. Shi, W. Jiang and Q. Zhang, Biotechnol. J., 2020, 15, e2000136 CrossRef PubMed.
- T. L. Grove, P. M. Himes, S. Hwang, H. Yumerefendi, J. B. Bonanno, B. Kuhlman, S. C. Almo and A. A. Bowers, J. Am. Chem. Soc., 2017, 139, 11734–11744 CrossRef CAS.
- K. A. S. Eastman, M. C. Mifflin, P. F. Oblad, A. G. Roberts and V. Bandarian, ACS Bio Med Chem Au, 2023, 3, 480–493 CrossRef CAS PubMed.
- K. A. S. Eastman, A. S. Jochimsen and V. Bandarian, J. Biol. Chem., 2023, 299, 105058 CrossRef CAS PubMed.
- K. A. S. Eastman, W. M. Kincannon and V. Bandarian, ACS Cent. Sci., 2022, 8, 1209–1217 CrossRef CAS PubMed.
- J. K. Pedigo, K. A. S. Eastman and V. Bandarian, ACS Bio Med Chem Au, 2025, 5, 1007–1015 CrossRef CAS.
- B. Cheng, J. Huang, Y. Duan and W. Liu, Angew. Chem., Int. Ed., 2023, 62, e202308733 CrossRef CAS.
- C. S. Sit, S. Yoganathan and J. C. Vederas, Acc. Chem. Res., 2011, 44, 261–268 CrossRef CAS PubMed.
- E. S. Grant-Mackie, E. T. Williams, P. W. R. Harris and M. A. Brimble, JACS Au, 2021, 1, 1527–1540 CrossRef CAS.
- Y. Gao, Y. Zhu, T. Awakawa and I. Abe, RSC Chem. Biol., 2024, 5, 293–311 RSC.
- B. J. Burkhart, C. J. Schwalen, G. Mann, J. H. Naismith and D. A. Mitchell, Chem. Rev., 2017, 117, 5389–5456 CrossRef CAS PubMed.
- S. A. Salisbury, H. S. Forrest, W. B. Cruse and O. Kennard, Nature, 1979, 280, 843–844 CrossRef CAS.
- M. Jin, L. Liu, S. A. Wright, S. V. Beer and J. Clardy, Angew. Chem., Int. Ed., 2003, 42, 2898–2901 CrossRef CAS PubMed.
- N. Fittipaldi, M. Segura, D. Grenier and M. Gottschalk, Future Microbiol., 2012, 7, 259–279 CrossRef CAS PubMed.
- P. M. Engelhardt, R. Keyzers and M. A. Brimble, Org. Biomol. Chem., 2024, 22, 8374–8396 RSC.
- X. Zhang and S. Li, Nat. Prod. Rep., 2017, 34, 1061–1089 RSC.
- A. Greule, J. E. Stok, J. J. De Voss and M. J. Cryle, Nat. Prod. Rep., 2018, 35, 757–791 RSC.
- K. D. Dubey and S. Shaik, Acc. Chem. Res., 2019, 52, 389–399 CrossRef PubMed.
- Z. Li, Y. Jiang, F. P. Guengerich, L. Ma, S. Li and W. Zhang, J. Biol. Chem., 2020, 295, 833–849 CrossRef.
- C. C. Chen, J. Min, L. Zhang, Y. Yang, X. Yu and R. T. Guo, ChemBioChem, 2021, 22, 1317–1328 CrossRef CAS.
- B. Meunier, S. P. de Visser and S. Shaik, Chem. Rev., 2004, 104, 3947–3980 CrossRef CAS PubMed.
- I. G. Denisov, T. M. Makris, S. G. Sligar and I. Schlichting, Chem. Rev., 2005, 105, 2253–2277 CrossRef CAS PubMed.
- Y. L. Hu, F. Z. Yin, J. Shi, S. Y. Ma, Z. R. Wang, R. X. Tan, R. H. Jiao and H. M. Ge, J. Am. Chem. Soc., 2023, 145, 27325–27335 CrossRef CAS.
- S. Kunakom, H. Otani, D. W. Udwary, D. T. Doering and N. J. Mouncey, J. Ind. Microbiol. Biotechnol., 2023, 50 Search PubMed.
- G. Zhong, ACS Bio Med Chem Au, 2023, 3, 371–388 CrossRef CAS.
- C. L. Liu, Z. J. Wang, J. Shi, Z. Y. Yan, G. D. Zhang, R. H. Jiao, R. X. Tan and H. M. Ge, Angew. Chem., Int. Ed., 2024, 63, e202314046 CrossRef CAS PubMed.
- S. K. Kandy, M. A. Pasquale and J. R. Chekan, Nat. Chem. Biol., 2025, 21, 168–181 CrossRef CAS.
- J. Liu, R. Liu, B. B. He, X. Lin, L. Guo, G. Wu and Y. X. Li, ACS Bio Med Chem Au, 2024, 4, 268–279 CrossRef CAS PubMed.
- R. Liu, H. Ren, G. Wu, B.-B. He, C. Pang, Z. Shen, J. Liu, Z.-M. Song, J. Zhou and Y.-X. Li, bioRxiv, 2024, preprint, DOI:10.1101/2024.10.06.616194.
- G. Yim, M. N. Thaker, K. Koteva and G. Wright, J. Antibiot., 2014, 67, 31–41 CrossRef CAS PubMed.
- B. Hadatsch, D. Butz, T. Schmiederer, J. Steudle, W. Wohlleben, R. Sussmuth and E. Stegmann, Chem. Biol., 2007, 14, 1078–1089 CrossRef CAS.
- H. S. Ali, R. H. Henchman and S. P. de Visser, Org. Biomol. Chem., 2020, 18, 4610–4618 RSC.
- A. K. L. Nguy, R. J. Martinie, A. Cai and M. R. Seyedsayamdost, J. Am. Chem. Soc., 2024, 146, 19629–19634 CrossRef CAS PubMed.
- C. C. Forneris and M. R. Seyedsayamdost, Angew. Chem., Int. Ed., 2018, 57, 8048–8052 Search PubMed.
- H.-T. Chiu, B. K. Hubbard, A. N. Shah, J. Eide, R. A. Fredenburg, C. T. Walsh and C. Khosla, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 8548–8553 CrossRef CAS PubMed.
- O. K. Park, H. Y. Choi, G. W. Kim and W. G. Kim, ChemBioChem, 2016, 17, 1725–1731 CrossRef CAS.
- A. Mollo, A. N. von Krusenstiern, J. A. Bulos, V. Ulrich, K. S. Åkerfeldt, M. J. Cryle and L. K. Charkoudian, RSC Adv., 2017, 7, 35376–35384 RSC.
- W. T. Liu, R. D. Kersten, Y. L. Yang, B. S. Moore and P. C. Dorrestein, J. Am. Chem. Soc., 2011, 133, 18010–18013 CrossRef CAS PubMed.
- J. Schimana, K. Gebhardt, A. Höltzel, D. G. Schmid, R. Süssmuth, J. Müller, R. Pukall and H. P. Fiedler, J. Antibiot., 2002, 55, 565–570 CrossRef CAS.
- P. A. Smith, M. E. Powers, T. C. Roberts and F. E. Romesberg, Antimicrob. Agents Chemother., 2011, 55, 1130–1134 Search PubMed.
- P. Kulanthaivel, A. J. Kreuzman, M. A. Strege, M. D. Belvo, T. A. Smitka, M. Clemens, J. R. Swartling, K. L. Minton, F. Zheng, E. L. Angleton, D. Mullen, L. N. Jungheim, V. J. Klimkowski, T. I. Nicas, R. C. Thompson and S. B. Peng, J. Biol. Chem., 2004, 279, 36250–36258 CrossRef CAS PubMed.
- H. Aldemir, S. Shu, F. Schaefers, H. Hong, R. Richarz, S. Harteis, M. Einsiedler, T. M. Milzarek, S. Schneider and T. A. M. Gulder, Chemistry, 2022, 28, e202103389 CrossRef CAS.
- D. G. Brown, T. L. May-Dracka, M. M. Gagnon and R. Tommasi, J. Med. Chem., 2014, 57, 10144–10161 CrossRef CAS PubMed.
- P. A. Smith, M. F. T. Koehler, H. S. Girgis, D. Yan, Y. Chen, Y. Chen, J. J. Crawford, M. R. Durk, R. I. Higuchi, J. Kang, J. Murray, P. Paraselli, S. Park, W. Phung, J. G. Quinn, T. C. Roberts, L. Rouge, J. B. Schwarz, E. Skippington, J. Wai, M. Xu, Z. Yu, H. Zhang, M. W. Tan and C. E. Heise, Nature, 2018, 561, 189–194 CrossRef CAS PubMed.
- M. Takashima, H. Sakai, R. Mori and K. Arima, Agric. Biol. Chem., 1962, 26, 660–678 CAS.
- S. Sakai, N. Aimi, K. Yamaguchi, Y. Hitotsuyanagi, C. Watanabe, K. Yokose, Y. Koyama, K. Shudo and A. Itai, Chem. Pharm. Bull., 1984, 32, 354–357 CrossRef CAS.
- S.-I. Sakai, Y. Hitotsuyanagi, K. Yamaguchi, N. Aimi, K. Ogata, T. Kuramochi, H. Seki, R. Hara, H. Fujiki, M. Suganuma, T. Sugimura, Y. Endo, K. Shudo and Y. Koyama, Chem. Pharm. Bull., 1986, 34, 4883–4886 CrossRef CAS PubMed.
- T. Yamashita, M. Imoto, K. Isshiki, T. Sawa, H. Naganawa, S. Kurasawa, B.-Q. Zhu and K. Umezawa, J. Nat. Prod., 1988, 51, 1184–1187 CrossRef CAS.
- H. H. Sun, C. B. White, J. Dedinas, R. Cooper and D. M. Sedlock, J. Nat. Prod., 1991, 54, 1440–1443 CrossRef CAS PubMed.
- S.-i Sakai, Y. Hitotsuyanagi, N. Aimi, H. Fujiki, M. Suganuma, T. Sugimura, Y. Endo and K. Shudo, Tetrahedron Lett., 1986, 27, 5219–5220 CrossRef CAS.
- J. H. Cardellina, F.-J. Marner and R. E. Moore, Science, 1979, 204, 193–195 CrossRef CAS PubMed.
- N. Aimi, H. Odaka, S.-I. Sakai, H. Fujiki, M. Suganuma, R. E. Moore and G. M. L. Patterson, J. Nat. Prod., 1990, 53, 1593–1596 CrossRef CAS.
- T. P. Zhou, Y. Fan, J. Zhang and B. Wang, ACS Bio Med Chem Au, 2025, 5, 16–30 Search PubMed.
- Y. Hitotsuyanagi, H. Fujiki, M. Suganuma, N. Aimi, S. Sakai, Y. Endo, K. Shudo and T. Sugimura, Chem. Pharm. Bull., 1984, 32, 4233–4236 CrossRef CAS PubMed.
- T. Awakawa and I. Abe, Org. Biomol. Chem., 2018, 16, 4746–4752 RSC.
- I. Abe, J. Antibiot., 2018, 71, 763–768 CrossRef CAS.
- T. Mori, Chem. Pharm. Bull., 2023, 71, 188–197 CrossRef CAS PubMed.
- J. Wei and Y. Liu, J. Chem. Inf. Model., 2021, 61, 3638–3648 CrossRef CAS PubMed.
- M. U. Huynh, M. C. Elston, N. M. Hernandez, D. B. Ball, S.-I. Kajiyama, K. Irie, W. H. Gerwick and D. J. Edwards, J. Nat. Prod., 2010, 73, 71–74 CrossRef CAS PubMed.
- D. J. Edwards and W. H. Gerwick, J. Am. Chem. Soc., 2004, 126, 11432–11433 CrossRef CAS.
- I. Morita, T. Mori and I. Abe, Chemistry, 2021, 27, 2963–2972 CrossRef CAS PubMed.
- F. He, T. Mori, I. Morita, H. Nakamura, M. Alblova, S. Hoshino, T. Awakawa and I. Abe, Nat. Chem. Biol., 2019, 15, 1206–1213 CrossRef CAS.
- J. Ma, D. Zuo, Y. Song, B. Wang, H. Huang, Y. Yao, W. Li, S. Zhang, C. Zhang and J. Ju, ChemBioChem, 2012, 13, 547–552 Search PubMed.
- A. Benjdia, L. Decamps, A. Guillot, X. Kubiak, P. Ruffié, C. Sandström and O. Berteau, J. Biol. Chem., 2017, 292, 10835–10844 CrossRef CAS.
- L. Padva, J. Gullick, L. J. Coe, M. H. Hansen, J. J. De Voss, M. Crusemann and M. J. Cryle, ChemBioChem, 2025, 26, e202400916 Search PubMed.
- S. C. Kessler and Y. H. Chooi, Nat. Prod. Rep., 2022, 39, 222–230 Search PubMed.
- T. Ozaki, A. Minami and H. Oikawa, J. Antibiot., 2023, 76, 3–13 Search PubMed.
- H. Kries, F. Trottmann and C. Hertweck, Angew. Chem., Int. Ed., 2024, 63, e202309284 Search PubMed.
- Q. Nie, C. Sun, S. Liu and X. Gao, Biochemistry, 2024, 63, 2948–2957 CrossRef CAS PubMed.
- F. Biermann, B. Tan, M. Breitenbach, Y. Kakumu, P. Nanudorn, Y. Dimitrova, A. S. Walker, R. Ueoka and E. J. N. Helfrich, Chem. Sci., 2024, 15, 17506–17523 RSC.
- P. Nanudorn, S. Thiengmag, F. Biermann, P. Erkoc, S. D. Dirnberger, T. N. Phan, R. Furst, R. Ueoka and E. J. N. Helfrich, Angew. Chem., Int. Ed., 2022, 61, e202208361 Search PubMed.
- T. P. Wyche, A. C. Ruzzini, L. Schwab, C. R. Currie and J. Clardy, J. Am. Chem. Soc., 2017, 139, 12899–12902 CrossRef CAS PubMed.
- S. H. Reisberg, Y. Gao, A. S. Walker, E. J. N. Helfrich, J. Clardy and P. S. Baran, Science, 2020, 367, 458–463 CrossRef CAS.
- J. S. An, H. Lee, H. Kim, S. Woo, H. Nam, J. Lee, J. Y. Lee, S. J. Nam, S. K. Lee, K. B. Oh, S. Kim and D. C. Oh, Angew. Chem., Int. Ed., 2023, 62, e202300998 CrossRef CAS PubMed.
- H. Nam, J. S. An, J. Lee, Y. Yun, H. Lee, H. Park, Y. Jung, K. B. Oh, D. C. Oh and S. Kim, J. Am. Chem. Soc., 2023, 145, 22047–22057 CrossRef CAS PubMed.
- J. J. Hug, N. A. Frank, C. Walt, P. Senica, F. Panter and R. Muller, Molecules, 2021, 26, 7483 CrossRef CAS.
- M. M. Zdouc, M. M. Alanjary, G. S. Zarazua, S. I. Maffioli, M. Crusemann, M. H. Medema, S. Donadio and M. Sosio, Cell Chem. Biol., 2021, 28(733–739), e734 Search PubMed.
- M. H. Hansen, A. Keto, M. Treisman, V. M. Sasi, L. Coe, Y. Zhao, L. Padva, C. Hess, V. Leichthammer, D. L. Machell, R. B. Schittenhelm, C. J. Jackson, J. Tailhades, M. Crüsemann, J. J. De Voss, E. H. Krenske and M. J. Cryle, ACS Catal., 2024, 14, 812–826 CrossRef CAS.
- Y. Zhao, E. Marschall, M. Treisman, A. McKay, L. Padva, M. Crusemann, D. R. Nelson, D. L. Steer, R. B. Schittenhelm, J. Tailhades and M. J. Cryle, Angew. Chem., Int. Ed., 2022, 61, e202204957 CrossRef CAS PubMed.
- M. Treisman, L. Coe, Y. Zhao, V. M. Sasi, J. Gullick, M. H. Hansen, A. Ly, V. Leichthammer, C. Hess, D. L. Machell, R. B. Schittenhelm, J. Hooper, C. J. Jackson, J. Tailhades, J. J. De Voss and M. J. Cryle, Org. Lett., 2024, 26, 1828–1833 CrossRef CAS PubMed.
- J. J. Hug, J. Dastbaz, S. Adam, O. Revermann, J. Koehnke, D. Krug and R. Muller, ACS Chem. Biol., 2020, 15, 2221–2231 CrossRef CAS.
- B. B. He, J. Liu, Z. Cheng, R. Liu, Z. Zhong, Y. Gao, H. Liu, Z. M. Song, Y. Tian and Y. X. Li, Angew. Chem., Int. Ed., 2023, 62, e202311533 CrossRef CAS PubMed.
- J. I. Tietz, C. J. Schwalen, P. S. Patel, T. Maxson, P. M. Blair, H. C. Tai, U. I. Zakai and D. A. Mitchell, Nat. Chem. Biol., 2017, 13, 470–478 CrossRef CAS PubMed.
- H. Saad, T. Majer, K. Bhattarai, S. Lampe, D. T. Nguyen, M. Kramer, J. Straetener, H. Brotz-Oesterhelt, D. A. Mitchell and H. Gross, Chem. Sci., 2023, 14, 13176–13183 RSC.
- S. E. Barrett and D. A. Mitchell, Trends Genet., 2024, 40, 950–968 CrossRef CAS PubMed.
- M. Noyon and S. Hematian, Trends Chem., 2024, 6, 649–655 CrossRef CAS.
- J. Hattori, K. A. Boutilier, M. L. Campagne and B. L. Miki, Mol. Gen. Genet., 1998, 259, 424–428 CrossRef CAS.
- R. D. Kersten and J. K. Weng, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, E10961–E10969 Search PubMed.
- J. Hattori, K. A. Boutilier, M. M. van Lookeren Campagne and B. L. Miki, Mol. Gen. Genet., 1998, 259, 424–428 CrossRef CAS PubMed.
- L. S. Mydy, J. Hungerford, D. N. Chigumba, J. R. Konwerski, S. C. Jantzi, D. Wang, J. L. Smith and R. D. Kersten, Nat. Chem. Biol., 2024, 20, 530–540 Search PubMed.
- J. R. Chekan, L. S. Mydy, M. A. Pasquale and R. D. Kersten, Nat. Prod. Rep., 2024, 41, 1020–1059 RSC.
- R. D. Kersten, L. S. Mydy, T. R. Fallon, F. de Waal, K. Shafiq, J. W. Wotring, J. Z. Sexton and J. K. Weng, J. Am. Chem. Soc., 2022, 144, 7686–7692 Search PubMed.
- S. T. Lima, B. G. Ampolini, E. B. Underwood, T. N. Graf, C. E. Earp, I. C. Khedi, M. A. Pasquale and J. R. Chekan, Angew. Chem., Int. Ed., 2023, 62, e202218082 CrossRef CAS PubMed.
- B.-S. Yun, I.-J. Ryoo, I.-K. Lee and I.-D. Yoo, Tetrahedron, 1998, 54, 15155–15160 CrossRef CAS.
- Y. Wang and M. M. Joullie, Chem. Rec., 2021, 21, 906–923 CrossRef CAS PubMed.
- X. Wang, K. Shafiq, D. A. Ousley, D. N. Chigumba, D. Davis, K. M. McDonough, L. S. Mydy, J. Z. Sexton and R. D. Kersten, Nat. Commun., 2025, 16, 4198 CrossRef CAS.
- S. T. Lima, M. A. Pasquale, M. Noyon, E. A. Clark, C. R. Laws, S. Hematian and J. R. Chekan, J. Am. Chem. Soc., 2025, 147, 20284–20293 CrossRef CAS.
- D. Kriger, M. A. Pasquale, B. G. Ampolini and J. R. Chekan, Beilstein J. Org. Chem., 2024, 20, 1548–1559 CrossRef CAS PubMed.
- E. Tuenter, V. Exarchou, A. Balde, P. Cos, L. Maes, S. Apers and L. Pieters, J. Nat. Prod., 2016, 79, 1746–1751 CrossRef CAS.
- G. A. Vignolle, R. L. Mach, A. R. Mach-Aigner and C. Derntl, BMC Genomics, 2020, 21, 258 CrossRef CAS PubMed.
- M. R. Quijano, C. Zach, F. S. Miller, A. R. Lee, A. S. Imani, M. Kunzler and M. F. Freeman, J. Am. Chem. Soc., 2019, 141, 9637–9644 CrossRef CAS PubMed.
- M. Umemura, N. Nagano, H. Koike, J. Kawano, T. Ishii, Y. Miyamura, M. Kikuchi, K. Tamano, J. Yu, K. Shin-ya and M. Machida, Fungal Genet. Biol., 2014, 68, 23–30 CrossRef CAS PubMed.
- Y. Ye, A. Minami, Y. Igarashi, M. Izumikawa, M. Umemura, N. Nagano, M. Machida, T. Kawahara, K. Shin-Ya, K. Gomi and H. Oikawa, Angew. Chem., Int. Ed., 2016, 55, 8072–8075 CrossRef CAS PubMed.
- N. Nagano, M. Umemura, M. Izumikawa, J. Kawano, T. Ishii, M. Kikuchi, K. Tomii, T. Kumagai, A. Yoshimi, M. Machida, K. Abe, K. Shin-Ya and K. Asai, Fungal Genet. Biol., 2016, 86, 58–70 CrossRef CAS.
- W. Ding, W. Q. Liu, Y. Jia, Y. Li, W. A. van der Donk and Q. Zhang, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 3521–3526 CrossRef CAS.
- Z. Wieczorek, I. Z. Siemion, M. K. Zimecki, E. Bolewska-Pedyczak and T. Wieland, Peptides, 1993, 14, 1–5 CrossRef CAS.
- H. Luo, S. Y. Hong, R. M. Sgambelluri, E. Angelos, X. Li and J. D. Walton, Chem. Biol., 2014, 21, 1610–1617 CrossRef CAS PubMed.
- H. Luo, H. E. Hallen-Adams, Y. Luli, R. M. Sgambelluri, X. Li, M. Smith, Z. L. Yang and F. M. Martin, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2201113119 CrossRef CAS.
- H. E. Hallen, H. Luo, J. S. Scott-Craig and J. D. Walton, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 19097–19101 CrossRef CAS PubMed.
- J. A. Pulman, K. L. Childs, R. M. Sgambelluri and J. D. Walton, BMC Genomics, 2016, 17, 1038 CrossRef PubMed.
- Q. Nie, F. Zhao, X. Yu, M. C. Madhusudhanan, C. Chang, S. Li, S. R. Chowdhury, B. Kille, A. Xu, R. Sharkey, C. Sun, H. Zeng, S. Liu, D. Zhou, X. Yu, K. Yang, S. A. C. Figueiredo, M. Zotova, Z. Hu, A. Y. Du, D. Guan, R. Tang, T. Treangen, J. Wang, P. N. Leao, Y. Gao, J. Chen, P. Liu, H. Renata and X. Gao, Nat. Chem. Biol., 2025, 21, 1938–1947 CrossRef CAS PubMed.
- S. C. Kessler, X. Zhang, M. C. McDonald, C. L. M. Gilchrist, Z. Lin, A. Rightmyer, P. S. Solomon, B. G. Turgeon and Y.-H. Chooi, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 24243–24250 CrossRef CAS.
- R. D. Johnson, G. A. Lane, A. Koulman, M. Cao, K. Fraser, D. J. Fleetwood, C. R. Voisey, J. M. Dyer, J. Pratt, M. Christensen, W. R. Simpson, G. T. Bryan and L. J. Johnson, Fungal Genet. Biol., 2015, 85, 14–24 CrossRef CAS PubMed.
- W. Zhang, N. T. Forester, P. Chettri, M. Heilijgers, W. J. Mace, E. Maes, Y. Morozova, E. R. Applegate, R. D. Johnson and L. J. Johnson, J. Agric. Food Chem., 2023, 71, 13965–13978 CrossRef CAS PubMed.
- C. Y. Chiang, M. Ohashi and Y. Tang, J. Am. Chem. Soc., 2025, 147, 8113–8117 CrossRef CAS PubMed.
- C. Y. Chiang, M. Ohashi, J. Le, P. P. Chen, Q. Zhou, S. Qu, U. Bat-Erdene, S. Hematian, J. A. Rodriguez, K. N. Houk, Y. Guo, J. A. Loo and Y. Tang, Nature, 2025, 638, 126–132 CrossRef CAS PubMed.
- G. E. Kenney, L. M. K. Dassama, M.-E. Pandelia, A. S. Gizzi, R. J. Martinie, P. Gao, C. J. DeHart, L. F. Schachner, O. S. Skinner, S. Y. Ro, X. Zhu, M. Sadek, P. M. Thomas, S. C. Almo, J. M. Bollinger, C. Krebs, N. L. Kelleher and A. C. Rosenzweig, Science, 2018, 359, 1411–1416 CrossRef CAS.
- C. Dou, Z. Long, S. Li, D. Zhou, Y. Jin, L. Zhang, X. Zhang, Y. Zheng, L. Li, X. Zhu, Z. Liu, S. He, W. Yan, L. Yang, J. Xiong, X. Fu, S. Qi, H. Ren, S. Chen, L. Dai, B. Wang and W. Cheng, Cell Res., 2022, 32, 302–314 CrossRef CAS.
- Y. J. Park, R. J. Jodts, J. W. Slater, R. M. Reyes, V. J. Winton, R. A. Montaser, P. M. Thomas, W. B. Dowdle, A. Ruiz, N. L. Kelleher, J. M. Bollinger, C. Krebs, B. M. Hoffman and A. C. Rosenzweig, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2123566119 CrossRef CAS.
- R. S. Ayikpoe, L. Zhu, J. Y. Chen, C. P. Ting and W. A. van der Donk, ACS Cent. Sci., 2023, 9, 1008–1018 CrossRef CAS PubMed.
- C. P. Ting, M. A. Funk, S. L. Halaby, Z. Zhang, T. Gonen and W. A. van der Donk, Science, 2019, 365, 280–284 CrossRef CAS.
- M. I. McLaughlin, Y. Yu and W. A. van der Donk, ACS Chem. Biol., 2022, 17, 930–940 CrossRef PubMed.
- G. E. Kenney and A. C. Rosenzweig, ACS Chem. Biol., 2012, 7, 260–268 CrossRef CAS.
- R. J. Jodts, M. B. Ho, R. M. Reyes, Y. J. Park, P. E. Doan, A. C. Rosenzweig and B. M. Hoffman, Biochemistry, 2024, 63, 1170–1177 CrossRef CAS PubMed.
- H.-w Liu and T. P. Begley, Comprehensive natural products III: chemistry and biology, Elsevier, 2020 Search PubMed.
- J. Y. Chen and W. A. van der Donk, Curr. Opin. Chem. Biol., 2024, 80, 102467 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |