Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Machine learning-driven molecular engineering of nucleic acids
Qien
Shi
a,
Hui
Lv
b,
Fei
Wang
a,
Chunhai
Fan
 *ac and
Mingqiang
Li
*ac and
Mingqiang
Li
 *a
*a
aState Key Laboratory of Synergistic Chem-Bio Synthesis, School of Chemistry and Chemical Engineering, Frontiers Science Center for Transformative Molecules, New Cornerstone Science Laboratory, Zhang Jiang Institute for Advanced Study, National Center for Translational Medicine, Shanghai Jiao Tong University, Shanghai, 200240, China. E-mail: limingqiang@sjtu.edu.cn; fanchunhai@sjtu.edu.cn
bInstitute of Materiobiology, College of Sciences, Shanghai University, Shanghai, 200444, China
cInstitute of Molecular Medicine, Shanghai Key Laboratory for Nucleic Acids Chemistry and Nanomedicine, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200127, China
First published on 12th March 2026
Abstract
Molecular engineering has played a pivotal role in biomedical fields, driving significant advancements in gene therapy, disease diagnosis, and biosensing. However, nucleic acid molecular engineering faces various challenges including vast design spaces, complex structure–function relationships, lengthy application validation cycles, and inefficient optimization processes. Machine learning (ML), with its superior pattern recognition, multidimensional data integration, and automated optimization capabilities, offers a unique opportunity to construct predictive models of sequence-structure–function relationships, thereby enabling a paradigm shift from empirically driven to data-driven approaches. This review systematically surveys recent progress in ML applications across three major domains: nucleic acid structure construction, performance modulation, and application expansion. It also explores core challenges such as data quality, model interpretability, and experimental validation efficiency, along with potential resolution strategies. These insights are poised to propel nucleic acid molecular engineering from static structure prediction toward dynamic behavior simulation, and from single-molecule design to complex system engineering, guiding future directions in hybrid ML-quantum models and expanded applications to non-canonical nucleic acids for transformative innovation in biomedicine, environmental monitoring, and information technology.
Key learning points(1) Fundamentals of ML algorithms applied to nucleic acid sequence-structure–function relationships.(2) Strategies for nucleic acid structure construction using ML, from primary sequences to three-dimensional models. (3) ML-driven modulation of nucleic acid properties, editing tools, and functional elements for enhanced performance. (4) Applications of ML-enhanced nucleic acids in diagnostics, therapeutics, and information processing. (5) Challenges like data quality and interpretability, with future directions in hybrid ML-quantum models. |
Introduction
Nucleic acid molecular engineering1–4 has seen expanding applications in biomedicine and information sciences, largely owing to its unique advantages in molecular diagnostics,5 drug development,6–8 biosensors,9,10 molecular computing, and information storage,11 including high programmability, self-assembly capability, biocompatibility, etc. (Box 1). As an interdisciplinary technology, this field achieves efficient solutions from disease detection to gene editing through precise manipulation of DNA and RNA sequences and structures. For instance, clustered regularly interspaced short palindromic repeats (CRISPR)-based gene editing technologies12–14 and aptamer diagnostic platforms15,16 have demonstrated significant potential in clinical practice. Moreover, progress in nucleic acid molecular engineering is particularly prominent in information storage, where DNA serves as a high-density, long-term stable medium with theoretical capacity far exceeding traditional silicon-based technologies.11,17 Currently, researchers are actively exploring the potential of nucleic acids in molecular computing and nanodevices, such as DNA origami techniques and nucleic acid logic circuits, paving new paths for next-generation computing architectures.18–22 Despite substantial advancements, major barriers remain for further development. These include the time-consuming nature of nucleic acid structure construction, the lack of rational performance design, and insufficient scalability in emerging applications.10,23–25 These challenges not only hinder large-scale implementation but also pose urgent demands for deeper advancements in synthetic biology and precision medicine.26–28 By overcoming these obstacles, nucleic acid molecular engineering is poised to profoundly reshape the landscapes of biotechnology and information technology in the coming decades.
Box 1: Nucleic acid molecular engineering and its unique featuresThe inherent challenges and limitations of traditional biological and informational molecular engineering have prompted researchers to develop and utilize nucleic acid molecules (such as DNA and RNA) for more precise and controllable design of molecular systems that surpass natural structures and functions, termed nucleic acid molecular engineering in this review.Compared to traditional molecular engineering, nucleic acid molecular engineering possesses several unique features: High programmability. The sequence-specific and programmable nature of nucleic acids enables precise design of complex nanostructures. By optimizing DNA or RNA sequences, intermolecular interactions can be predicted and modulated to assemble nanoscale components with predefined shapes and functions. Self-assembly capability. Nucleic acid molecules spontaneously assemble via base pairing, forming complex structures such as DNA origami or DNA bricks. This mechanism simplifies the construction process of nanostructures and enhances design reproducibility and precision. Biocompatibility. As inherent components of living organisms, nucleic acids typically exhibit excellent biocompatibility, suitable for biomedical applications like drug delivery and biosensors, thereby reducing immune responses and toxicity risks. Multifunctionality. Nucleic acid nanostructures can integrate multiple functions, including molecular recognition, catalysis, signal transduction, and regulation. These properties are enhanced through sequence optimization and chemical modifications, showing broad potential in detection, therapy, and materials science. Modifiability. Nucleic acid molecules can be enhanced in stability, specificity, and functionality through various chemical modifications. For example, fluorescent labeling, drug conjugation, or integration of functional molecules can elevate their application potential. Dynamic tunability. Nucleic acid structures can undergo dynamic transitions in response to environmental stimuli (such as temperature, pH, or ion concentration), laying the foundation for developing intelligent responsive materials. Information processing capability. Beyond structural construction, nucleic acids possess information storage and computing functions. Sequence encoding enables molecular-level information processing, a unique attribute unattainable by other nanomaterials. |
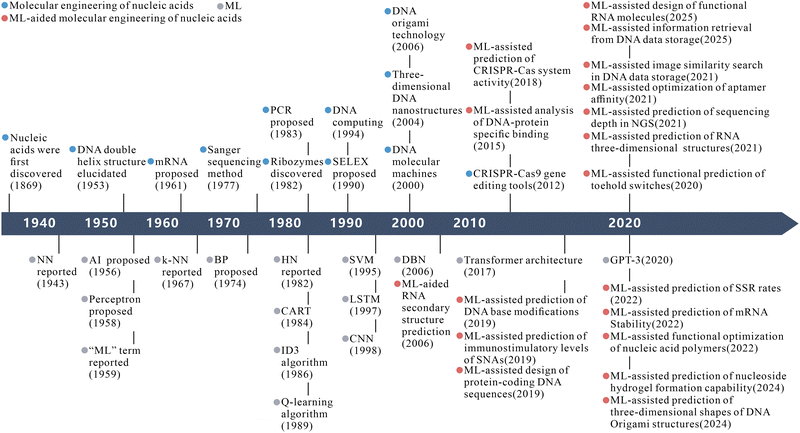
ML, as a core branch of artificial intelligence (AI), empowers systems with autonomous decision-making and predictive capabilities by extracting patterns and regularities from data, profoundly reshaping research paradigms across multiple disciplines. Based on learning paradigms, ML can be categorized into supervised learning, unsupervised learning, semi-supervised learning, and reinforcement learning, which are respectively suited for labeled data-driven prediction tasks, discovery of intrinsic data structures, partially labeled scenarios, and dynamic decision optimization.29–31 Within ML architectures, neural networks (NNs) stand out. They offer powerful hierarchical feature learning capabilities and serve as core tools for complex tasks.32,33 Specifically, graph neural networks (GNNs) excel in modeling network data.34 Long short-term memories (LSTMs) are adept at processing sequential data.35 Variational autoencoders (VAEs) advance generative models.36 Transformer architectures lead with breakthrough performance in natural language processing and cross-modal tasks.37–40 Beyond NNs, traditional methods such as random forests (RF), support vector machine (SVM), and regression models remain widely used for analyzing small- to medium-scale datasets due to their efficiency and interpretability.41–43 These models are based on statistical inference, kernel methods, or ensemble learning principles, each with distinct advantages and complementary coexistence. In the field of nucleic acid molecular engineering, ML demonstrates unique potential to significantly accelerate time-consuming tasks while enhancing insights into complex system behaviors (Fig. 1).44–46 By integrating multimodal data with high-performance computing, ML is opening innovative pathways for nucleic acid research.47,48
Compared to traditional nucleic acid molecular engineering, which relies on laborious experimental iterations and theoretical modeling, ML has significantly accelerated the engineering process of nucleic acid molecules through its easily deployable model architectures and an increasingly rich ecosystem of user-facing ML/AI software implementations and benchmarks (Box 2), together with powerful data-driven capabilities.49–51 Specifically, ML applications in this field have expanded to multiple core dimensions (Table 1). In structure construction, algorithms fusing DNNs and generative models have substantially improved RNA secondary and three-dimensional structure prediction accuracy (Fig. 2a).44,52,53 In performance modulation, deep learning (DL)-based methods efficiently predict sequences with specific functions. Examples include CRISPR guide RNA optimization (Fig. 2b).54–56 In application expansion, ML-enabled high-throughput screening has advanced disease diagnostics and nucleic acid drug development (Fig. 2c).57–59 These achievements stem from ML's unique advantages in handling high-dimensional data and complex molecular interactions.30,60 Notably, the maturity and reliability of these approaches are not uniform across domains. ML is currently most established as an auxiliary tool for structural analysis and design assistance, serves as a powerful optimizer for performance modulation within well-characterized experimental regimes, and is only beginning to enable proof-of-concept advances in application expansion that still require extensive validation. From a practical standpoint, a useful rule of thumb for nucleic acid engineering is to start with supervised learning whenever sufficiently large labelled datasets are available for the task of interest, because such models provide direct quantitative predictions and well-understood evaluation metrics. Unsupervised and deep generative models are best suited when the goal is to discover latent structure in sequence or structural data, cluster related molecules, or generate novel variants in the absence of exhaustive labels. Semi-supervised methods can be advantageous when many sequences are available but only a subset has been characterized experimentally, allowing unlabeled data to regularize and enrich the learned representations. Reinforcement learning, in turn, is currently most appropriate for sequential design problems, where an agent iteratively edits a sequence and receives a reward only after simulating or measuring the resulting structure. However, ML applications in nucleic acid molecular engineering still face several challenges, particularly in areas like liquid-phase regulation of DNA reaction networks (i.e., solution-phase strand-displacement cascades and DNA computing circuits whose behavior is dominated by coupled kinetics and environmental factors) and complex structure design, where its adoption remains relatively limited, highlighting bottlenecks in data dependency, model generalization, and computational efficiency.61,62 These limitations provide ample room for future technological innovations, especially in multiscale modeling and interdisciplinary data integration.63 This review aims to systematically outline the latest advancements, key challenges, and potential opportunities in nucleic acid molecular engineering, with a focus on how emerging ML technologies can more efficiently drive innovation and applications in this field.
| Category | Methods | Advantages | Disadvantages | Applications |
|---|---|---|---|---|
| Supervised learning | • Linear models (e.g., linear regression, logistic regression)101 | • Easy to implement and interpret for simpler models | • Inability to capture highly complex nonlinear relationships in basic linear models | • Nucleic acid-protein interaction prediction104–106 |
| • Distance-based models (e.g., k-NN)102 | • Well-defined objectives leading to accurate predictions on labeled data | • Heavy reliance on large amounts of high-quality labeled data | • Nucleic acid structure prediction52,61,75 | |
| • SVM43 | • High interpretability, especially for linear and tree-based models | • Limited generalization if training data is biased or insufficient | • Gene editing efficiency prediction55,56,107 | |
| • Tree-based models (e.g., Decision Trees, RF)41 | • Clear evaluation criteria using metrics like accuracy, precision, recall, and F1-score | • Failure to discover latent patterns | • Nucleic acid species and modification prediction82,108,109 | |
| • NNs (e.g., CNNs for images, transformers for various tasks)103 | • Risk of overfitting, particularly in deep neural networks (DNNs) | • Aptamer and nucleic acid switch design57,72,110 | ||
| • Sensitivity to data quality and noise | • Nucleic acid structure classification and reconstruction111–113 | |||
| • DNA information processing84,85,114 | ||||
| • Nucleic acid detection99,115,116 | ||||
| • Design of nucleic acid composite nanomaterials87,91,93 | ||||
| • Prediction of nucleic acid physicochemical properties49,117,118 | ||||
| • Nucleic acid bioproperty prediction97,119,120 | ||||
| Unsupervised learning | • Clustering analysis (e.g., K-means clustering, hierarchical clustering)121 | • Uncovers intrinsic data structures and patterns without requiring labeled data | • Opaque interpretability of results | • Nucleic acid structure prediction51,61,75 |
| • Dimensionality reduction (linear: principal component analysis – PCA; nonlinear: t-distributed stochastic neighbor embedding – t-SNE)122 | • Operates effectively without human supervision | • Lack of standardized evaluation metrics | • Aptamer optimization124,125 | |
| • Deep representation learning (e.g., autoencoders – AEs)123 | • Capable of significant feature dimensionality reduction to simplify data | • High parameter sensitivity | • Functional nucleic acid sequence design59,96,126 | |
| • Deep generative models (e.g., generative adversarial networks – GANs, VAEs)36 | • High versatility across domains like anomaly detection and data visualization | • Susceptibility to noise interference | • Gene editing efficiency prediction80 | |
| Semi-supervised learning | • Self-training (e.g., pseudo-labeling techniques)127 | • Capable of learning high-dimensional features from limited data | • Stringent requirements for data annotation quality | • High-affinity nucleic acid polymer sequence design58 |
| • Consistency regularization (e.g., in models like mean teacher for image classification)128 | • Reduces dependency on expensive labeled samples | • Heightened sensitivity to noise in unlabeled data | • RNA-targeted drug activity prediction83 | |
| • Graph-based methods (e.g., label propagation, label spreading)129 | • Enhances generalization performance | • Elevated algorithmic complexity requiring more computational resources | ||
| • Generative models (e.g., semi-supervised GANs)130 | • Exhibits flexible applicability across scenarios | • Fundamental reliance on distributional consistency assumptions | ||
| • Co-training (e.g., using multiple views of data for mutual supervision)131 | ||||
| Reinforcement learning | • Markov decision process (as foundational framework)132 | • Sequential decision optimization | • Challenging reward engineering | • RNA inverse folding sequence design100 |
| • Q-Learning133 | • Adaptive to environmental dynamics | • Low sample efficiency | ||
| • Policy gradient134 | • Label-free operation | • Difficult exploration-exploitation tradeoff | ||
| • Superior exploration capability | • Unstable convergence properties |
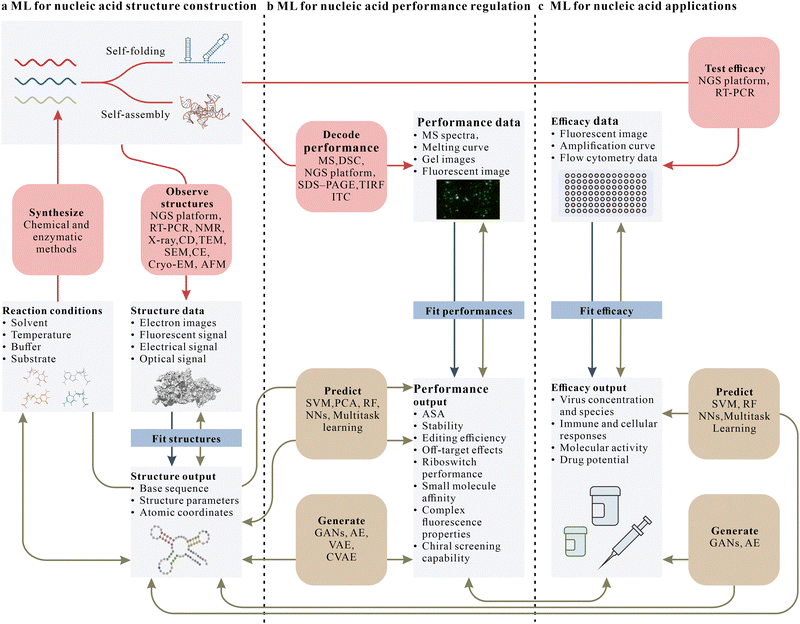 | ||
| Fig. 2 ML-assisted workflow for molecular engineering of nucleic acids. (a)–(c), Researchers employ ML for nucleic acid structure synthesis (a), nucleic acid performance modulation (b), and nucleic acid-based applications (c). Red arrows denote experimental process, dark blue arrows represent experimental data fitting, while brown arrows indicate current ML applications for prediction and generation tasks, as well as future opportunities for ML in molecular engineering of nucleic acids. RT-PCR, reverse transcription polymerase chain reaction; NMR, nuclear magnetic resonance; X-ray, X-radiation; CD, circular dichroism spectrometer; TEM, transmission electron microscopy; SEM, scanning electron microscopy; CE, capillary electrophoresis; Cryo-EM, cryogenic electron microscopy; AFM, atomic force microscopy; MS, mass spectrometry; DSC, differential scanning calorimeter; SDS-PAGE, sodium dodecyl sulfate-polyacrylamide gel electrophoresis; TIRF, single-molecule fluorescence microscopy; ITC, isothermal titration calorimetry; CVAE, conditional variational autoencoder; ASA, accessible surface area. Panels adapted with permission from: a, ref. 70 under a Creative Commons licence CC BY 4.0; ref. 76, Springer Nature Ltd; ref. 45 under a Creative Commons licence CC BY 4.0; c, ref. 116, Copyright 2025, Elsevier Ltd; ref. 37, Springer Nature Ltd. | ||
ML-guided nucleic acid structure construction
Nucleic acid structures form the foundation for their functions. Precise prediction and design of these structures are essential for unlocking their potential in biomedicine and information sciences.64–66 However, their complexity and diversity pose severe challenges to traditional experimental and computational methods.67,68 ML, by learning complex associations between sequences, structures, and functions from vast datasets, enables efficient and accurate prediction and design of nucleic acid molecules with specific structures. From primary sequence design to secondary structure prediction and complex three-dimensional structure construction, ML methods not only enhance prediction accuracy but also significantly shorten design cycles, providing robust support for nucleic acid structure engineering.51,69,70 In practice, these models are most reliable when used to analyze and integrate structural data, prioritize candidate designs, and narrow the search space; fully predictive end-to-end pipelines, especially for three-dimensional structures, still depend heavily on data availability and careful experimental validation.Primary structure
The primary structure of nucleic acids, namely the base sequence, determines their higher-order structures and functions. In fields such as synthetic biology, nanomaterials, and biosensing, precise design of nucleic acid sequences with specific functions is central to engineering applications. However, the vastness of sequence space poses significant challenges to traditional empirical approaches and trial-and-error methods, which are often inefficient for screening,86,87 lack systematic rules,88 and face difficulties in balancing authenticity with customizability.89 These issues underscore the urgent need for efficient, systematic sequence design strategies. ML, with its superior pattern recognition and predictive capabilities, extracts sequence-structure–function relationships from limited experimental data, enabling efficient navigation through vast sequence spaces to generate and optimize nucleic acid sequences with target structures and functions (Fig. 3a).90
Box 2: Toolkit for ML-driven nucleic acid molecular engineeringThis box provides a curated selection of open-source software, models, and platforms discussed in this review, aiming to assist researchers in selecting appropriate tools for specific engineering tasks. These tools cover the entire workflow from structure construction to performance modulation and application-driven engineering.Structure construction NucleoBench. A large-scale benchmark suite for evaluating nucleic acid sequence design algorithms.71 (https://github.com/move37-labs/nucleobench) AdaBeam. An adaptive beam search algorithm for optimizing nucleic acid sequences.71 (https://github.com/move37-labs/nucleobench) GARDN-SANDSTORM. DL framework integrating sequence and structure features for functional RNA design.72 (https://github.com/AlexGreenLab/GARDN-SANDSTORM) GEMORNA. Transformer-based generative model for optimizing mRNA translation and stability.73 (https://github.com/RainaBio/GEMORNA) MXfold2. Integrates thermodynamic parameters with DL for robust RNA folding prediction.70 (https://github.com/keio-bioinformatics/mxfold2/) trRosettaRNA. Transformer-network pipeline for automated RNA 3D structure prediction.45 (https://yanglab.qd.sdu.edu.cn/trRosettaRNA/) RoseTTAFoldNA. End-to-end prediction of protein–nucleic-acid complexes via multi-track neural architectures.74 (https://github.com/uw-ipd/RoseTTAFold2NA) HORNET. Deep neural networks for identifying RNA topologies in solution.75 (https://zenodo.org/records/10637777) CryoREAD. De novo nucleic-acid model building from cryo-EM maps using deep learning.76 (https://github.com/kiharalab/CryoREAD) YOLOv5. Used for fast detection/classification of DNA nanostructures in AFM images.77 (https://github.com/ultralytics/yolov5) Performance modulation RNAsnap2. Predict RNA solvent accessibility from sequence-derived features.78 (https://github.com/jaswindersingh2/RNAsnap2) RNAsol. LSTM-based RNA solvent accessibility prediction.79 (https://yanglab.nankai.edu.cn/RNAsol/) DeepCRISPR. DL model for CRISPR guide activity prediction and optimization.80 (https://github.com/bm2-lab/DeepCRISPR) CRISPR-GPT. Automates the design of gene-editing experiments using large language models.81 (https://github.com/cong-lab/crispr-gpt-pub) RhoDesign. Structure-to-sequence generative design platform for RNA aptamers.57 (https://github.com/ml4bio/RhoDesign) Application-driven engineering DeepMod2. DL frameworks for DNA methylation/base modification calling from nanopore sequencing.82 (https://github.com/WGLab/DeepMod2) sChemNET. DL framework for predicting small molecules targeting microRNA function.83 (https://github.com/diegogalpy/sChemNET/) DNAformer. DL reconstruction model for robust DNA-storage read recovery.84 (https://github.com/itaiorr/Deep-DNA-based-storage.git) 2DDNA. ML-enabled reconstruction in rewritable two-dimensional DNA-based data storage.85 (https://doi.org/10.5281/zenodo.5774385) |
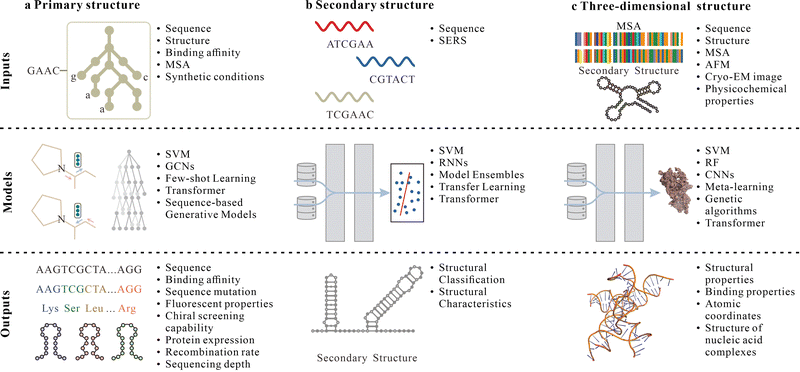 | ||
| Fig. 3 ML-assisted nucleic acid structural construction. (a)–(c) Examples of ML model inputs, architectures, and outputs related to the synthesis of nucleic acid primary (a), secondary (b), and three-dimensional (c) structures. In panel b, ‘structural classification’ refers to assigning sequences to discrete secondary-structure classes, whereas ‘structural characteristics’ denotes residue-level or base-pairing properties. MSA, multiple sequence alignment; GCN, graph convolutional network; SERS, surface-enhanced Raman spectroscopy; RNN, recurrent neural network. Panels adapted with permission from: a, ref. 59, Springer Nature Ltd; ref. 37, Springer Nature Ltd; ref. 97 under a Creative Commons licence CC BY 4.0; b, ref. 37, Springer Nature Ltd; ref. 70 under a Creative Commons licence CC BY 4.0; c, ref. 45 under a Creative Commons licence CC BY 4.0; ref. 76, Springer Nature Ltd; ref. 37, Springer Nature Ltd. | ||
ML applications in nucleic acid primary structure design primarily focus on reverse design based on DNA sequence performance. In carbon nanotube chirality separation, researchers analyzed 12-mer C/T base sequences using ML methods, elevating the prediction efficiency of DNA sequences recognizing specific chiral single-walled carbon nanotubes (SWNTs) to over 50%, and revealing significant contributions of sequence terminal structures and “CCC” motifs to classification.86 Further, employing algorithms such as RF, NNs, and SVMs to analyze DNA sequence features (e.g., position-specific vectors, word frequency vectors), the discovery of resolved sequences was increased from ∼10 to ∼103, corresponding to a 3 orders of magnitude expansion in the accessible design space, with screening success rates rising from 10% to >90%, and elucidating G/C base combinations as a universal rule for super-resolution sequence patterns.87 In DNA-templated silver nanoclusters (DNA–AgNs) design, an ensemble of SVM classifiers was employed to analyze 2661 ten-base DNA sequences, resulting in a 12.3-fold increase in the success rate of designing near-infrared fluorescent DNA–AgN complexes.91 To address the challenges associated with high-dimensional sequence space, one model focused on local sequence motifs rather than full-length sequences. This approach enabled cross-length design of DNA templates ranging from 8 to 16 bases, leading to a 99–154% enhancement in the proportion of sequences exhibiting red fluorescence.92 Additionally, a bidirectional gated recurrent unit-based DL model successfully predicted the number of fluorescence emission peaks in hairpin DNA–AgNs with 81.4% accuracy.93 ML has also demonstrated significant advantages in DNA synthesis feasibility prediction,94 DNA-SWNT composite sensor optimization,95 protein function optimization,96–98 and DNA probe sequence generation and design.89,99
Beyond DNA, ML has achieved breakthroughs in RNA and functional nucleic acid polymer design. For the RNA inverse folding problem, which involves designing RNA sequences given target secondary structures, researchers trained agents using reinforcement learning to generate RNA sequences that fold into specific secondary structures;100 in the design of RNA sequences with tailored properties, the DL framework SANDSTORM, which integrates both sequence and structural information, and the GAN GARDN have significantly enhanced the prediction accuracy and design efficiency of functional RNA molecules. Experimental validation showed that toehold switches designed using these approaches achieved ON/OFF ratios 11.9 times higher than those generated with conventional tools.72 Employing the advanced transformer architecture, the GEMORNA deep generative model significantly enhances mRNA sequence translation efficiency and stability, overcoming the conventional difficulty of simultaneously achieving multiple optimization objectives;73 additionally, GNN and GPT-like language model-based RNA generation models successfully predicted and validated RNA mutations enhancing Escherichia coli ribosome thermal stability;69 addressing data sparsity challenges, the RfamGen deep generative model, by combining covariance models and VAEs, generates high-activity RNA family sequences with only hundreds of training data.59 In functional nucleic acid polymers, CVAE models successfully generated novel polymers unrelated to experimental sequences, yielding new sequences with 9–26 nM high affinities.58 In parallel, tools such as AdaBeam (adaptive beam-search optimization) and NucleoBench (standardized benchmarking of nucleic acid sequence design algorithms) help lower the barrier for end users by enabling efficient sequence optimization and fair model comparison.71 These studies not only markedly improved nucleic acid design efficiency and success rates but also revealed sequence patterns and design rules elusive to traditional methods, providing theoretical guidance and practical tools for nucleic acid molecular engineering.
Secondary structure
Nucleic acid secondary structures are core elements in elucidating biomolecular functions and regulatory mechanisms, exerting profound impacts in gene expression regulation, drug design, and biosensor development.135,136 However, nucleic acids can form diverse secondary conformations, such as DNA G-quadruplexes (G4) and i-motifs (iM), and RNA stem-loops and pseudoknots, whose formation is regulated by multiple factors including sequence features, environmental conditions, and intermolecular interactions.137,138 Traditional physics-based thermodynamic models, exemplified by Mfold,139 ViennaRNA140 and NUPACK,141 rely on energy minimization and dynamic programming algorithms to provide rigorous predictions grounded in biophysical principles. However, these approaches are often computationally prohibitive for large-scale designs owing to their cubic scaling and show reduced accuracy for non-canonical structures where thermodynamic parameters are incomplete; experimental determination provides high-resolution information but faces challenges like high costs, low throughput, and resolution limitations.70 ML, with its superior pattern recognition and nonlinear relationship mining capabilities, extracts conformation-sensitive features from massive data, compensating for traditional methods' deficiencies in atomic-level interactions and enabling real-time monitoring of structural dynamics, thus opening new paths for precise prediction and detection of nucleic acid secondary structures (Fig. 3b), both at the level of discrete structural classification and residue-level structural characteristics.70,142Accurate detection and prediction of DNA secondary structures are crucial for revealing gene regulatory mechanisms and developing biosensors, but traditional methods struggle to capture conformational diversity and lack efficient detection strategies. To address these challenges, researchers have developed a series of innovative ML methods. In structural detection, the integration of SERS with ML has enabled the construction of high-throughput screening platforms. By analyzing spectral features of 54 oligonucleotides with defined conformations, highly accurate classification models were developed. These models can predict the conformations of unseen sequences and identify dominant structural states under different pH conditions. Notably, linear discriminant analysis (LDA) achieved 100% accuracy in three-class classification tasks.136 For G4 prediction, CNN-based models trained on nearly 400 million human genome data from G4-seq enabled precise genome-wide G4 mismatch score evaluation and demonstrated cross-species generalization.143 To fill gaps in G4 topology prediction, G4ShapePredictor integrated 1005 experimentally validated sequence-topology pairs using various ensemble learning models, elevating average test accuracy to 0.75 ± 0.02.144 The Quadron model innovatively integrates high-throughput G4-seq data with ML to analyze 209 features derived from 703![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 091 canonical G4 sequences and their flanking regions in the human genome. Using a gradient-boosted tree (GBT) algorithm for regression training, the model achieved a prediction accuracy of Pearson correlation coefficient (PCC) of 0.80. Furthermore, feature importance analysis revealed that additional G-triplets within flanking regions critically contribute to structural stability.145
091 canonical G4 sequences and their flanking regions in the human genome. Using a gradient-boosted tree (GBT) algorithm for regression training, the model achieved a prediction accuracy of Pearson correlation coefficient (PCC) of 0.80. Furthermore, feature importance analysis revealed that additional G-triplets within flanking regions critically contribute to structural stability.145
RNA secondary structure prediction is foundational for parsing non-coding RNA functions and guiding drug design, but traditional methods, reliant on energy minimization and dynamic programming, struggle with complex structures and heavy computational burdens. ML, through advanced model architectures and algorithmic optimizations, has significantly enhanced the accuracy and efficiency of RNA secondary structure prediction.146 The E2Efold model pioneered unrolling constraint optimization algorithms into differentiable NN modules, constructing an end-to-end DL framework that integrates transformer-encoded deep scoring networks with unfolding optimization-based post-processing modules, achieving an F1 score of 0.821 on RNAStralign and ArchiveII datasets, which represents a 29.7% improvement over traditional methods.135 To address overparameterization and improve generalization, MXfold2 integrates DL with a thermodynamic framework. It computes four folding scores using NNs and incorporates Turner nearest-neighbor parameters, while adding thermodynamic regularization constraints. These design choices enhance model robustness. In family-level cross-validation, MXfold2 achieved state-of-the-art performance, with an overall F1 score of 0.601 for structure prediction and a Spearman correlation of 0.833 between folding scores and experimental free energies.70 The SPOT-RNA integrates deep residual networks with bidirectional LSTMs, augmented by a transfer learning strategy, to address the challenges of predicting non-canonical base pairs and pseudoknots. The model was first pre-trained on large-scale, low-precision structural data and subsequently fine-tuned on a smaller high-precision dataset, leading to a marked improvement in performance. For example, this approach increased the F1-score for non-nested base pairs by 53%.142
Three-dimensional structure
The three-dimensional structures of nucleic acid molecules are core elements in understanding their biological functions, exerting profound influences on gene regulation, viral replication, drug design, and synthetic biology. DNA and RNA not only serve as genetic information carriers but also perform catalytic reactions, gene expression regulation, and molecular recognition through specific three-dimensional conformations.147,148 However, nucleic acid three-dimensional structure research faces challenges such as high experimental resolution costs and time consumption, difficulty in capturing molecular flexible conformations, and reliance on manual expertise for nucleic acid nanostructure characterization, with inefficient yield assessment in complex environments.149,150 ML, particularly DL techniques, provides innovative paths to overcome these barriers by integrating multilayer information and constructing end-to-end predictive models (Fig. 3c).45,151Prediction and modeling of nucleic acid three-dimensional structures are essential for parsing biological mechanisms and guiding drug development, but existing experimental methods are costly and limited in addressing structural flexibility and diversity. DL has achieved key breakthroughs in this domain, encompassing three primary directions. First, in RNA structure prediction, end-to-end architectures excel: NuFold,53 DRfold,152 and RhoFold+44 integrate innovative representation learning and geometric constraints for flexible sugar ring conformation modeling, accuracy enhancements in ab initio predictions, and evolutionary information mining via language models, respectively outperforming traditional methods in benchmarks; ARES,52 as a geometric DL exemplar, employs rotation-translation equivariant architectures to automatically extract three-dimensional features, achieving high-precision structure scoring under small-sample conditions. Second, in DNA structure prediction, Deep DNAshape153 systematically models long-range flanking sequence effects using RNNs for the first time; DGNN61 fuses GNNs with physical information for sub-second prediction of DNA origami three-dimensional conformations. Third, in protein-nucleic acid complex prediction, diverse DL methods offer complementary solutions: RoseTTAFoldNA74 and DeepPBS106 employ three-track NNs and geometric DL to enable end-to-end complex prediction and evaluate binding specificity across protein families; EPBDxDNABERT-2154 integrates DNA structural dynamics with a pre-trained language model to enhance the prediction accuracy of human transcription factor binding sites. DNAffinity,155 DeepCLIP,156 and NucleicNet104 address binding affinity prediction from varied perspectives. These methods highlight DL's advantages in handling high-dimensional data, capturing long-range dependencies, and fusing multiscale features, injecting new vitality into nucleic acid structure prediction. Nevertheless, their accuracy and robustness still depend strongly on the availability and quality of experimentally determined structures, the coverage of non-canonical motifs, and the use of physics-based refinement, so they should currently be viewed as powerful aids rather than fully autonomous predictors for nucleic acid three-dimensional design.
Characterization and analysis of nucleic acid three-dimensional structures bridge experimental observations and theoretical models, but traditional methods are inefficient and imprecise for large-scale heterogeneous data. DL, with its powerful feature extraction and pattern recognition capabilities, achieves breakthroughs in three key areas. First, in experimental characterization techniques, ML enhances imaging efficiency and data quality: HORNET51,75 combines AFM with DNNs for direct measurement and dynamic conformation visualization of RNA tertiary topologies in solution, revealing functional balances between core structural stability and peripheral flexibility; DL frameworks based on YOLOv5,77 YoloX,77 and RNAN112 address rapid detection-classification of DNA nanostructures and super-resolution reconstruction of AFM images, substantially improving characterization efficiency. Second, in structure parsing and reconstruction, diverse DL methods synergistically advance automation and precision: EM2NA,113 Emap2sec+,111 NucleoFind,157 and CryoREAD76 develop dedicated NNs for varying resolution electron density maps, enabling high-precision automated construction from density maps to atomic models and accelerating structure parsing workflows. Third, in structure-binding properties analysis, ML exhibits strong predictive and explanatory capabilities: multimodal DL frameworks158 integrate multilayer RNA structural information to systematically parse tertiary structure influences on protein binding preferences; extreme gradient boosting (XGBoost),159 LASSO regression,160 and L2-regularized linear models105 apply to DNA nanostructure protein corona prediction, RNA-binding chemical space feature extraction, and quantification of DNA shape contributions to protein binding specificity, providing quantitative bases for understanding structure–function relationships. The common feature of these methods is transforming traditional empirical-driven approaches into data-driven automated workflows, not only improving efficiency and accuracy but also revealing complex patterns and rules elusive to conventional methods, thereby offering new design principles and tools for nucleic acid molecular engineering.
ML-assisted nucleic acid performance modulation
As multifunctional biomacromolecules, nucleic acids require performance modulation that is crucial for biomedical research and applications. However, nucleic acid performance modulation faces multifaceted challenges, including system complexity, numerous parameters, and unclear mechanisms, with traditional empirical-driven methods often relying on extensive trial-and-error experiments, resulting in low efficiency and difficulty in achieving precise control. ML can mine complex nonlinear relationships from high-throughput experimental data. This reveals key factors that shape nucleic acid performance.110,124 By integrating multidimensional features, including sequence, structure, and thermodynamic parameters, comprehensive predictive models enable precise evaluation.72,161,162 Guided by these models, reverse-design strategies substantially enhance functional performance.80,163 Because such models are typically trained on data generated under specific experimental conditions, their outputs are best interpreted as context-dependent scoring or ranking functions that accelerate design cycles within a given regime, rather than as universally transferable predictors across arbitrary cell types, delivery systems, or assay formats. These advantages position ML as a powerful tool for addressing bottlenecks in nucleic acid performance modulation.Nucleic acid properties
Nucleic acid molecular properties are foundational for parsing and manipulating their functions, directly determining their diverse roles in organisms, including gene expression regulation, protein interactions, and catalytic activity.164,165 These properties encompass solvent accessibility, stability, self-assembly capability, and mechanical characteristics,166–168 regulated multidimensionally by sequences, structures, and environmental factors, exhibiting highly nonlinear complex relationships.153 Traditional computational methods struggle to capture these patterns, while experimental approaches are limited by low throughput, high costs, and insufficient precision.62 ML, with its superior pattern recognition and feature extraction capabilities, learns high-dimensional regularities from massive data to achieve high-precision prediction of nucleic acid molecular properties, providing theoretical support for nucleic acid molecular engineering (Fig. 4a).169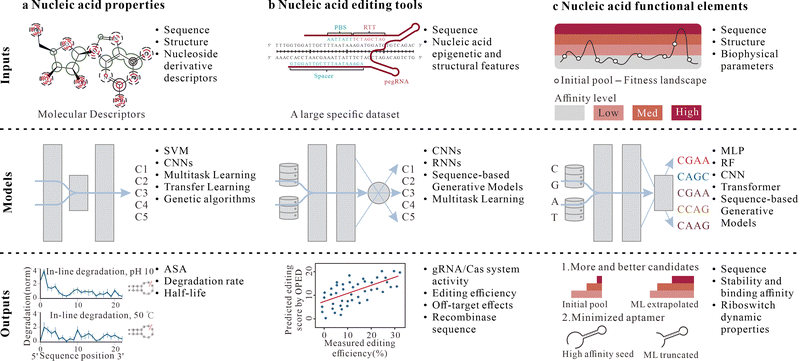 | ||
| Fig. 4 ML-assisted performance regulation of nucleic acids. (a)–(c) Examples of inputs, architectures, and output of ML models related to regulating the performances of nucleic acid molecules (a), nucleic acid editing tools (b), and functional nucleic acid elements (c). C1–C5 denote Class 1 to Class 5. MLP, multilayer perceptron. Panels adapted with permission from: a, ref. 49 under a Creative Commons licence CC BY 4.0; ref. 37, Springer Nature Ltd; ref. 118 under a Creative Commons licence CC BY 4.0; b, ref. 107 under a Creative Commons licence CC BY 4.0; ref. 37, Springer Nature Ltd; c, ref. 110 under a Creative Commons licence CC BY 4.0; ref. 37, Springer Nature Ltd. | ||
RNA solvent accessibility, as a key parameter characterizing RNA tertiary structures, is vital for understanding RNA-protein interactions, functional site localization, and structural feature analysis, but traditional experimental methods like X-ray crystallography and NMR imaging suffer from low throughput, with chemical probes only approximating accessibility, and existing computational methods failing to capture tertiary structural features. To address this, researchers have developed a series of ML tools. RNAsnap was the earliest method, using SVMs trained on protein-bound RNA structures and accepting either sequence spectra or single-sequence features as input. In benchmarking, it achieved PCCs of 0.66 in cross-validation and 0.63 on an independent test set. These predictions were further supported by significant correlations with dimethyl sulfate probing data and with population genetic variation frequencies, reinforcing the method's biological relevance.170 Subsequently, RNAsol enhanced capture of RNA sequence long-range dependencies via improved sequence profile alignment and LSTMs.79 RNAsnap2 further innovatively adopted dilated CNNs combined with LinearPartition171 predicted base-pairing probabilities as new features, elevating median PCCs by 11–22% on benchmark datasets.78 The latest M2pred multiscale DL framework, by integrating base-pairing probabilities, position-specific frequency matrices, and one-hot encoding as three feature types to construct multiscale contextual pyramid features, and designing multibranch NNs with residual attention modules, achieved superior performance on multiple test sets with a PCC of 0.58 and mean absolute error of 31.07.172
Nucleic acid stability and mechanical properties represent another critical dimension influencing function and applications, with accurate prediction holding immense value for synthetic biology, genetic engineering, and biomedicine, yet diverse influencing factors render traditional methods unable to establish precise predictive models. To tackle this challenge, researchers have developed various ML models. In mRNA stability prediction, a study integrated large-scale parallel experiments, biophysical modeling, and gradient boosting algorithm to construct a high-accuracy predictive model by analyzing 62![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 120 5′ untranslated region variants. The study revealed quantitative regulatory roles of RNA pyrophosphohydrolase (RppH) binding sites, translation rates, and single-stranded RNA length in determining mRNA stability.117 Another study adopted a “dual crowdsourcing” strategy, crowdsourcing RNA sequence datasets via the Eterna platform and combining multitask NNs to achieve single-nucleotide-level degradation prediction accuracy of 41%, providing new tools for RNA vaccine thermal stability design.118 For nucleoside derivative hydrogel stability prediction, ML combined with feature engineering screened 24 core features from 4175 molecular descriptors to build a predictive model with 71% accuracy, successfully validating novel hydrogel formation.49 In DNA mechanical property studies, the DNAcycP DL tool, via a hybrid Inception-ResNet and LSTM architecture, precisely predicted DNA cyclization capability, revealing contributions of periodic dinucleotide motifs to DNA bending and providing theoretical foundations for nucleosome assembly and DNA nanotechnology.173
120 5′ untranslated region variants. The study revealed quantitative regulatory roles of RNA pyrophosphohydrolase (RppH) binding sites, translation rates, and single-stranded RNA length in determining mRNA stability.117 Another study adopted a “dual crowdsourcing” strategy, crowdsourcing RNA sequence datasets via the Eterna platform and combining multitask NNs to achieve single-nucleotide-level degradation prediction accuracy of 41%, providing new tools for RNA vaccine thermal stability design.118 For nucleoside derivative hydrogel stability prediction, ML combined with feature engineering screened 24 core features from 4175 molecular descriptors to build a predictive model with 71% accuracy, successfully validating novel hydrogel formation.49 In DNA mechanical property studies, the DNAcycP DL tool, via a hybrid Inception-ResNet and LSTM architecture, precisely predicted DNA cyclization capability, revealing contributions of periodic dinucleotide motifs to DNA bending and providing theoretical foundations for nucleosome assembly and DNA nanotechnology.173
Nucleic acid editing tools
Gene editing tools, as core pillars of modern biotechnology, play indispensable roles in basic research, disease therapy, and bioengineering.174,175 With the rapid development of CRISPR-Cas systems and derivative technologies, precise genome regulation has become feasible;176 however, gene editing tools face challenges like unstable editing efficiency, unpredictable off-target effects, significant cell-type-specific differences, and lengthy optimization cycles for novel editing systems.177,178 Traditional experimental design methods rely on empirical rules and limited sequence feature analysis, failing to comprehensively capture complex sequence-function relationships, leading to inaccurate performance predictions and time-consuming, costly design processes.177 ML methods, by integrating large-scale experimental data, multimodal biological information, and advanced algorithms, promise precise performance prediction and systematic optimization of gene editing tools, thereby accelerating their development and applications (Fig. 4b).179,180 Even with ML, predicted activities and off-target profiles frequently vary across cell types and assay systems, so current models are typically used to rank or filter candidate guides and editor variants that must still be validated experimentally in the target context.Gene editing efficiency and specificity are crucial for overcoming safety and efficacy barriers in clinical applications of CRISPR systems. Traditional methods struggle to accurately predict editing effects across variants and cellular environments. To address these challenges, the field has undergone a paradigm shift from traditional ML to DL, forming three key technical routes. First, large-scale data-driven feature learning has supplanted manual feature engineering, with multiple studies constructing high-throughput screening datasets covering tens to hundreds of thousands of sequences, combined with CNNs, RNNs, and temporal convolutional networks (TCN) for high-precision efficiency prediction.5,80,163,181,182 Second, multimodal data integration strategies markedly improve predictive accuracy. For instance, DeepCRISPR incorporates both genomic sequences and epigenetic features,80 while CRISPRon integrates sequence features with thermodynamic parameters.180 Mixed-effects ML models further enhance predictions by including transcriptomic data such as gene expression levels.183 More recently, CRISPR-GPT has combined sequence scoring, domain-specific knowledge, and experimental validation.81 Together, these approaches underscore the critical importance of multisource data fusion in boosting model performance. Third, interpretable ML methods reveal molecular mechanisms of gene editing: through SHAP analysis, integrated gradients, and feature importance evaluation, researchers uncovered proximal PAM cytosine preferences,80 distal PAM 3–5 position base regulation of high-fidelity Cas9 variant specificity,181 and core regions (positions 15–24) with GC preference motifs in RNA guides.184 These technical routes have not only achieved breakthroughs in DNA editing (Cas9 and variants) but also extended to RNA editing (Cas13d),56,184 CRISPR-Cpf1,54 and bacterial CRISPRi systems,179,183 forming universal predictive frameworks across species and systems, providing reliable computational tools for precise gene editing.
Novel gene editing technologies are vital for expanding the gene editing toolbox and enhancing precision and applicability. Traditional development of novel editing tools typically relies on time-consuming directed evolution and screening, lacking systematic design principles and predictive capabilities, resulting in long cycles, high costs, and low success rates. To address these challenges, the field has formed two complementary ML paths. On one hand, DL-driven sequence optimization strategies have achieved breakthroughs in prime editing: PRIDICT and OPED models, via distinct network architectures (attention-based bidirectional RNNs and deep transfer learning nucleotide language models), enabled precise pegRNA efficiency prediction.55,107 Though employing different algorithmic frameworks, both studies revealed intrinsic associations between key nucleotide positions and editing efficiency, such as negative correlations in the first 7 nucleotides of PBS and positive in positions 8–13,107 establishing universal principles for prime editing design through large-scale dataset construction and multicell-type validation.55 On the other hand, generative DL models open new avenues for protein engineering: the RecGen algorithm, via CVAE, enabled intelligent prediction of novel DNA target-specific recombinases,126 first demonstrating ML's ability to parse complex recombinase-target affinity relationships and substantially shortening recombinase development cycles. These paths collectively embody three core advantages of ML in novel editing tool design: automatic extraction of sequence-function associations from data, generalization across cell types and experimental conditions, and experimental guidance via interpretable analysis. These research outcomes not only accelerate novel gene editing tool development but also lay methodological foundations for future integration of multiomics data, structural prediction tools (e.g., AlphaFold3185), and quantitative activity data to further enhance model performance, providing powerful technological support for gene therapy and precision medicine. At the same time, systematic cross-cell-type and cross-protocol benchmarking of these models remains limited, so retraining or fine-tuning on system-specific data is advisable before deploying them as decision-support tools in new experimental contexts.
Nucleic acid functional elements
Nucleic acid functional elements are key components in synthetic biology and molecular diagnostics, including riboregulators, aptamers, and ribozymes. They are structural units with specific functions capable of executing regulation, recognition, and catalysis, and are widely used in gene expression control, biosensing, and drug development.186,187 However, traditional nucleic acid functional element design primarily relies on trial-and-error experiments or thermodynamics-based rational design,188 while achieving some success, these methods are inefficient and have limited success rates when facing complex functional demands and multivariable optimization. Particularly when simultaneously considering sequence composition, secondary structure stability, tertiary conformational changes, and target interactions, traditional methods struggle to effectively explore vast sequence spaces and predict element performance.189 ML technologies, with their powerful data mining and pattern recognition capabilities, offer new solutions for nucleic acid functional element design and optimization, learning sequence–structure–function relationships from extensive experimental data to predict performance and guide optimization, thereby substantially improving design efficiency and success rates (Fig. 4c).110,190,191Riboregulators, including riboswitches and toehold switches, are indispensable functional elements in synthetic biology due to their ability to precisely control gene expression at post-transcriptional and translational levels. However, their design has long been limited by thermodynamic modeling and low-throughput experimental methods, making it difficult to accurately predict and optimize performance in complex cellular environments. To enhance riboregulator design efficiency and functional prediction accuracy, researchers have developed various ML strategies. In riboswitch optimization, combining RF and CNN extracted biophysical features from riboswitch sequences and secondary structures (e.g., P1 stem melting temperature Tm, free energy, GC content, hydrogen bonding patterns), successfully elevating tandem tetracycline riboswitch dynamic ranges to 40-fold, far surpassing traditional methods.162 These models exhibit notable interpretability, for instance, RF variable importance analysis revealed Tm and GC content as key factors, while hydrogen bond scoring quantified stem-end strong base pairing's promotion of dynamic ranges, providing actionable rules for riboswitch design. In toehold switches, researchers constructed a large-scale dataset of 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 534 toehold switches via high-throughput DNA synthesis and flow-seq, employing MLPs, CNNs, and LSTMs to extract features directly from sequences, achieving functional prediction accuracy enhancements 10-fold over traditional thermodynamic models.191 Model interpretability was augmented via VIS4Map technology, visualizing key secondary structures (e.g., stem-loop competing conformations) through saliency mapping and revealing leakage expression associations with kinetic intermediate states. Further, the STORM and NuSpeak DL frameworks, based on CNN sequence optimization and quasi-RNN RNA “grammar” modeling, enhanced interpretability via transfer learning and attention maps, elevating optimized sensor ON/OFF values up to 28.4-fold and demonstrating significant potential in SARS-CoV-2 pathogen detection.50
534 toehold switches via high-throughput DNA synthesis and flow-seq, employing MLPs, CNNs, and LSTMs to extract features directly from sequences, achieving functional prediction accuracy enhancements 10-fold over traditional thermodynamic models.191 Model interpretability was augmented via VIS4Map technology, visualizing key secondary structures (e.g., stem-loop competing conformations) through saliency mapping and revealing leakage expression associations with kinetic intermediate states. Further, the STORM and NuSpeak DL frameworks, based on CNN sequence optimization and quasi-RNN RNA “grammar” modeling, enhanced interpretability via transfer learning and attention maps, elevating optimized sensor ON/OFF values up to 28.4-fold and demonstrating significant potential in SARS-CoV-2 pathogen detection.50
Aptamers and ribozymes, as nucleic acid elements with molecular recognition and catalytic functions, offer substantial potential in biosensing, molecular diagnostics and targeted therapy. However, traditional screening methods such as SELEX are time-intensive, inefficient and susceptible to high false-positive rates, hindering the identification of optimal candidates from vast sequence spaces. ML has driven notable breakthroughs in their design and optimization. For aptamer screening, the conserved primary/secondary structure clustered pattern searching (CPS2) algorithm integrates sequence abundance, thermodynamic stability and secondary structures (such as hairpin loops) into a three-dimensional scoring system, enabling prediction of binding-active aptamers from single-round sequencing data and substantially reducing screening cycles.125 Similarly, the SMART-Aptamer framework uses a high-dimensional scoring system to evaluate SELEX-derived aptamer families, yielding high-affinity candidates with dissociation constants of 8–80 nM and establishing a new paradigm for ligand discovery in biomedicine.190 Furthermore, an ML-guided particle display approach, combining particle display with three NN architectures, predicts aptamer affinities, achieving 11-fold higher binding than conventional methods while shortening sequences by 70% without activity loss.110 Additionally, in generative models, RaptGen utilizes VAE and profile hidden Markov models to expand aptamer discovery via low-dimensional latent space embedding and optimization.124 Meanwhile, the RhoDesign platform employs geometric vector perceptrons (GVP) and transformers to enable reverse design from three-dimensional structures to RNA sequences, generating novel RNAs that mimic the structures of known fluorescent aptamers but feature distinct sequences, thereby supporting efficient diagnostic and therapeutic applications.57 For ribozyme design, high-throughput screening paired with DL-guided evolutionary algorithms has iteratively evolved eight ribozyme populations (>120![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 sequences), mapping neutral pathway networks between active ribozymes separated by 16 mutations; this reveals that low-order interactions suffice for predicting network topologies, offering frameworks for molecular engineering and viral evolution.192
000 sequences), mapping neutral pathway networks between active ribozymes separated by 16 mutations; this reveals that low-order interactions suffice for predicting network topologies, offering frameworks for molecular engineering and viral evolution.192
ML-enhanced nucleic acid applications
Owing to their specific recognition, programmability, and biocompatibility, nucleic acid molecules hold broad applications in disease diagnosis, drug development, and information processing.165,193,194 However, traditional nucleic acid application development methods often face challenges like low efficiency, insufficient specificity, and scalability difficulties.195 The introduction of ML provides new development opportunities for nucleic acid applications. By learning patterns from extensive experimental data, these algorithms can optimize the performance of nucleic acids across various application fields. This includes enhancing the sensitivity and specificity of diagnostics, improving the delivery efficiency and therapeutic efficacy of drugs, as well as increasing the capacity and speed of information storage and computation. Advanced ML methods like DL, reinforcement learning, and generative models handle complex nonlinear relationships in nucleic acid applications,196 while ensemble learning and transfer learning effectively utilize limited experimental data.197,198 Moreover, ML methods integrate multisource data, such as sequences, structures, functions, and clinical data, enabling more comprehensive application optimization.199 With a few notable exceptions, however, most ML-assisted nucleic acid applications are still at the proof-of-concept stage, typically demonstrated in controlled laboratory experiments or preclinical models and their robustness, scalability, and safety in real-world or clinical settings remain to be established through larger-scale validation.Nucleic acid diagnostics
With the advent of the precision medicine era, nucleic acid molecules, as carriers of life information, play crucial roles in disease diagnosis.195 Traditional nucleic acid detection methods, while widely applied in clinical practice, still face challenges such as signal overlap limiting single-molecule nucleic acid identification, difficulties in modification recognition, and deficiencies in specificity, sensitivity, and multiplexing for pathogen detection.200–202 ML, as a core branch of AI, with its powerful pattern recognition and data mining capabilities, provides new technological paths to address these challenges, propelling qualitative leaps in nucleic acid molecular engineering for disease diagnostics (Fig. 5a).203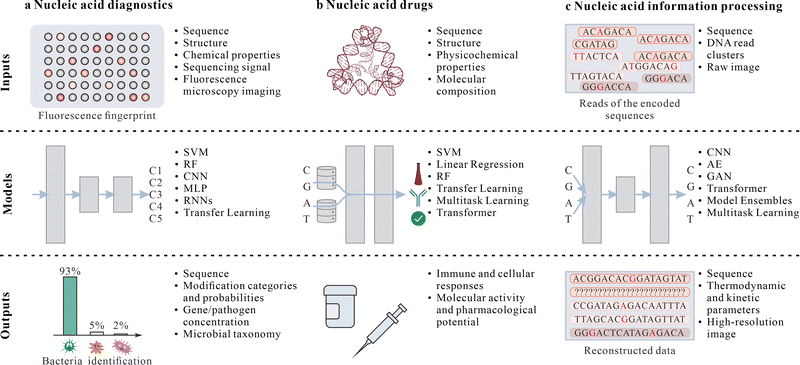 | ||
| Fig. 5 ML-assisted nucleic acid applications. (a)–(c) Examples of ML model inputs, architectures, and outputs related to nucleic acid-based diagnostics (a), drugs (b), and information processing (c). Panels adapted with permission from: a, ref. 116, Copyright 2025, Elsevier Ltd; ref. 37, Springer Nature Ltd; b, ref. 37, Springer Nature Ltd; c, ref. 84, Springer Nature Ltd; ref. 37, Springer Nature Ltd. | ||
Single-molecule-level nucleic acid recognition is fundamental for understanding life processes and disease mechanisms, yet traditional sequencing technologies encounter barriers in single-nucleotide identification, modification detection, and data processing. To tackle these, researchers have developed innovative solutions combining nanotechnology with ML. In nucleic acid molecule recognition, nanostructure construction based on various two-dimensional materials, such as graphene nanopores,204 hybrid graphene/hexagonal boron nitride nanopores,205 germanene nanogaps,206 and MXene nanochannels,207 combined with density functional theory and non-equilibrium Green's function-generated nucleotide transmission function datasets, and ML algorithms like XGBoost, k-NN, SVM, and RF, resolved signal overlap issues for high-precision DNA nucleotide identification.206,208 Notably, SHAP interpretability analysis revealed key contributions of transmission function features to model decisions, providing theoretical bases for algorithmic optimization.208 In RNA modification recognition, recognition tunneling combined with SVM algorithms extracted features from time, frequency, and cepstral domains for efficient modified RNA nucleotide identification,209 while engineered Mycobacterium smegmatis porin A nanopores extended this to high-resolution detection of 11 nucleoside monophosphates.108 In epigenetic modification detection, DeepMod210 and DeepMod2 frameworks82 addressed methylation detection in Oxford nanopore sequencing via bidirectional LSTM and transformer architectures, while the TandemMod framework, integrating one-dimensional CNNs, bidirectional LSTM modules, and attention mechanisms, enabled high-precision single-base resolution detection of 7 RNA modifications.109 These technological advances collectively propel nucleic acid molecule recognition from qualitative to quantitative and from single to multiple modification detection.
Rapid, sensitive pathogen detection is vital for disease prevention and public health security, yet traditional methods exhibit significant limitations in cost, time, and specificity.211,212 To counter these, researchers have developed various nucleic acid sensor-based diagnostic platforms integrating ML. The iFluor-RFA platform, via multiscale network architecture multiscale CNNs for DL analysis of fluorescent ring images, achieved specific, sensitive detection of nucleic acid targets at sub-micromolar levels with over 94% accuracy.213 Similarly, modular DNA origami nanorod-constructed monochromatic fluorescent barcode systems, via XGBoost and visual geometry group architecture CNNs for automatic recognition and classification, demonstrated ML's potential in multiplex biomolecular detection.115 Nonspecific nanosensor arrays based on two-dimensional nanoparticles (e.g., nGO, MoS2, WS2) complexed with single-stranded DNA, combined with models like partial least square discriminant analysis (PLSDA), logistic regression, and SVM, successfully differentiated complex bacterial matrices.214 This approach extended to food safety, where optical sensor arrays built from two-dimensional nanomaterials and single-stranded DNA, combined with MLPs and other ML algorithms, achieved 93.8% bacterial identification accuracy within 30 minutes, rising to 98.4% at 120 minutes.116 Through these ML algorithms, direct mapping from raw signals to quantitative analysis results was realized, providing robust technological support for disease prevention, while also highlighting the need for evaluation on larger, clinically representative cohorts to establish real-world performance and generalizability.
Nucleic acid therapeutics
Nucleic acid therapeutics, as a frontier in precision medicine, offer innovative strategies for various diseases by targeting gene expression networks or directly delivering therapeutic nucleic acid sequences.195 From microRNA modulators and mRNA therapies to structured nucleic acid nanoparticles, nucleic acid therapeutics exhibit broad potential in cancer, metabolic diseases, infectious diseases, and cardiovascular disorders.215,216 However, due to nucleic acid molecules' structural complexity, dynamic conformational changes, and multilayer interactions with biological systems, nucleic acid drug development faces multidimensional challenges like complex nonlinear structure–activity relationships, inefficient delivery systems, insufficient targeting specificity, difficult immune response prediction, and unclear chemical mechanisms.165,217 Thus, integrating high-throughput experiments with advanced computational methods, particularly ML, is essential for systematically parsing nucleic acid drug design rules, predicting biological effects, and accelerating optimization iterations (Fig. 5b).83,120 At the same time, translation of these design rules to human therapies will require extensive in vivo validation of safety, biodistribution, and immune responses beyond the regimes represented in the training data.Understanding nucleic acid drug structure–activity relationships is prerequisite for rational design, yet nucleic acid drug design involves multidimensional parameter spaces, with traditional experimental methods struggling to systematically reveal nonlinear interactions among parameters. To address this, researchers developed methods combining high-throughput experiments with ML to systematically parse SNA nanodrug structure–activity relationships; via picomolar-scale high-throughput synthesis and mass spectrometry, a library of 960 SNAs was constructed, with XGBoost and other ML models revealing nonlinear interactions among 11 design parameters, showing that only 16% random screening suffices for full-library activity prediction, significantly reducing experimental costs.120 Similarly, in mRNA therapies, researchers innovatively combined ML with four-component combinatorial chemistry high-throughput synthesis platforms to build an experimental library of 584 ionizable lipids and screen mRNA transfection efficacies; the trained XGBoost model (AUC-ROC 0.983) successfully screened efficient candidates from 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 virtual lipids, with 119-23 lipid transfection efficiencies in muscle and immune cells surpassing commercial benchmarks, and the model revealing key descriptors like head group hydrophobicity, tail unsaturation, and linker chain steric hindrance contributions to transfection efficacy.119 These studies indicate that ML not only accelerates virtual screening and reduces experimental costs for nucleic acid drugs but also provides interpretable guidance for structural optimization.
000 virtual lipids, with 119-23 lipid transfection efficiencies in muscle and immune cells surpassing commercial benchmarks, and the model revealing key descriptors like head group hydrophobicity, tail unsaturation, and linker chain steric hindrance contributions to transfection efficacy.119 These studies indicate that ML not only accelerates virtual screening and reduces experimental costs for nucleic acid drugs but also provides interpretable guidance for structural optimization.
Precise target identification and immune response prediction directly determine efficacy and safety in nucleic acid drug applications, yet traditional methods rely on time- and labor-intensive blind screening, failing to comprehensively predict complex off-target effects and immunogenicity risks. ML overcomes these barriers via innovative data integration and model construction. In targeted RNA drug discovery, the DL framework sChemNET, by integrating small-molecule chemical structures with miRNA sequence similarity constraints, successfully predicted small molecules targeting miRNA functions, validated in zebrafish embryos and human-derived cells for vitamin D receptor agonists' regulation of miR-181 family and miR-451;83 the DRLiPS method, via multistategy negative sampling and key feature screening, significantly enhanced RNA-binding pocket prediction accuracy, providing efficient tools for RNA-targeted drug design.218 In immune response prediction, systematic analysis of 58 nucleic acid nanoparticles' (NANPs) physicochemical and immune features developed the transformer-based predictive model AI-cell, with experimental validation showing accurate prediction of peripheral blood mononuclear cell interferon responses to NANPs, outperforming traditional RF and providing key tools for rapid design of NANPs with specific immune modulatory functions.219 These studies collectively advance ML's deep applications in nucleic acid therapeutics, accelerating personalized nucleic acid treatment strategy development.
Nucleic acid information processing
Nucleic acid molecules, particularly DNA, as natural information carriers in organisms, possess unique advantages like ultrahigh storage density, ultralong preservation lifespan, and extremely low energy consumption, offering revolutionary solutions to capacity bottlenecks and data aging issues in traditional storage media amid the data explosion era.11 However, nucleic acid information storage and computing systems face challenges in information writing and readout,197,220 information retrieva,114,221,222 and reaction parameter prediction.197 These complex issues often exceed traditional methods' capabilities, necessitating ML technologies for more efficient, precise solutions to fully harness nucleic acid molecules' potential in information storage and computing (Fig. 5c).Optimization and error correction in DNA storage systems are crucial for commercializing nucleic acid information storage, with core challenges in ensuring data integrity and reliability under high-noise environments. Traditional error correction relies on redundant coding, reducing storage efficiency and struggling with complex error patterns in DNA synthesis, PCR, and sequencing, particularly in clustering and reconstruction algorithm efficiency and precision during high-throughput sequencing. ML, with its powerful pattern recognition and data processing capabilities, has brought paradigm shifts to this field. DL models excel in DNA sequence reconstruction, fundamentally altering noisy data handling: on one hand, hybrid transformer and CNN architectures (e.g., DNAformer84) and GAN models (e.g., DNA-GAN197) transform sequence reconstruction into visualization tasks for precise recovery under high error rates; on the other, AE and U-Net network combinations pioneer direct information reconstruction from noisy data,220 reducing reliance on traditional error correction codes. These methods collectively point to a core breakthrough: using ML algorithms to replace or augment traditional error correction codes, significantly enhancing information retrieval speed (up to 3200-fold84) and accuracy (40% improvement84). More innovatively, two-dimensional DNA storage systems (2DDNA85), by encoding information at sequence and structural levels and using DL models for automatic error repair, provide novel approaches to reducing redundancy overhead. These technological advances address noise and error issues in DNA storage from diverse angles, clearing key obstacles for commercialization.
Expanding nucleic acid molecules' computing capabilities and functional applications is significant for building next-generation bio-electronic hybrid information systems, with keys in fully utilizing DNA's parallel computing potential and biological specificity. Traditional nucleic acid computing methods are limited to simple logic operations and sequence matching, lacking precise prediction of complex reaction parameters and advanced functions like content similarity search, severely restricting application scenarios. ML applications in this domain center on two core directions. First, in molecular behavior prediction, quantum chemical computations combined with CNNs223 break traditional nearest-neighbor model limitations by capturing synergistic effects among polynucleotides, establishing more precise DNA reaction parameter prediction frameworks and laying theoretical foundations for high-precision DNA nanodevice design. Second, in functional applications, DNA hybridization-based parallel computing frameworks,114 by establishing continuous feature-sequence encoding spaces, enable seamless transitions from electronic to molecular computing, pioneering content similarity search using DNA molecules. These directions embody ML's unique value in bridging theoretical models and practical applications, collectively propelling nucleic acid computing toward more complex, practical directions. This fusion trend not only expands nucleic acid molecules' application scenarios in information processing but also provides feasible paths for future efficient, low-energy bio-electronic hybrid computing systems.
Opportunities and challenges
ML-driven nucleic acid molecular engineering has achieved remarkable progress. It demonstrates immense potential in this interdisciplinary field, from structure prediction to performance optimization and practical applications. However, the domain still faces multiple technical challenges. These include bottlenecks in data quality, model interpretability, and experimental validation. These challenges not only constrain model generalization and practical application value but also highlight urgent needs for algorithmic innovation, data infrastructure development, and experimental technology integration. Concurrently, with synergistic advancements in computational power, data resources, and experimental platforms, this field harbors broad development opportunities. The following systematically analyzes current major challenges and explores potential solutions to provide insights for advancing ML's deeper applications in nucleic acid molecular engineering.Data quality challenges
Acquiring high-quality nucleic acid structure-function data is a core bottleneck in ML model development. Nucleic acid molecules' sequence spaces are extraordinarily vast, with highly complex structure–function relationships, while experimental characterization is often costly and time-consuming, resulting in limited training dataset scales and insufficient diversity, thereby impacting model generalization and prediction accuracy. Moreover, if models are evaluated only with random train–test splits on such narrow datasets, standard metrics (e.g., accuracy, PCC) can substantially overestimate performance in truly prospective or out-of-distribution scenarios. For example, RNA three-dimensional structure prediction models like NuFold53 and DRfold152 underperform on long sequences or non-canonical structures, while secondary structure prediction models like MXfold270 face similar limitations; nucleic acid switch optimization frameworks like STORM and NuSpeak,50 though advancing based on toehold sequence datasets, remain restricted in predicting novel structures. To overcome this, multilayer innovative strategies are needed. At the data acquisition level, self-supervised learning can leverage massive unlabeled data for pretraining, while physics-informed data augmentation generates synthetic samples conforming to known rules, enhancing dataset diversity. At the knowledge transfer level, transfer learning applies knowledge from related tasks to specific design problems, and active learning optimizes data annotation resource allocation by intelligently selecting maximally informative samples. Additionally, establishing open-shared nucleic acid structure-function databases and standardizing data processing workflows will provide solid infrastructure for the field. Synergistic application of these strategies, together with task-appropriate, standardized evaluation metrics and benchmark protocols that mimic realistic deployment settings, is poised to significantly alleviate data bottlenecks, propelling models toward higher precision and robustness.Model interpretability challenges
The “black box” nature of ML models contrasts sharply with nucleic acid molecular engineering's need for molecular mechanism understanding, severely limiting models' applications in rational design. Though high-performance models provide accurate predictions, their complex nonlinear mappings often obscure decision bases, failing to translate into explicit guidance rules. For instance, XGBoost-based DNA nanostructure protein corona prediction models, despite high accuracy, have feature importance analyses difficult to directly guide design;159 CRISPR-Cas9 optimization models like DeepHF reveal partial sequence feature contributions via SHAP but struggle to convert into universal criteria for novel gRNA design.182 Enhancing model interpretability requires a methodological system combining theory and practice. At the model architecture level, explainable AI techniques like attention mechanisms, gradient class activation mapping, and SHAP value analysis reveal key feature contributions, while mechanism-driven DL balances prediction and explanation by incorporating physical rules. At the analysis tool level, visualization techniques intuitively map high-dimensional feature spaces, and model distillation transfers complex model knowledge to more transparent simple models. Additionally, “human–machine collaborative” interactive frameworks integrate domain expert knowledge with ML outputs to enhance decision credibility. These methods collectively construct interpretive chains from intrinsic mechanisms to practical applications, providing reliable rational design support for nucleic acid molecular engineering. From a tutorial perspective, such interpretable analyses are crucial to help practitioners understand where a given model is expected to perform well and to treat ML as a context-dependent, complementary tool that augments mechanistic insight and experimental design rather than as an opaque oracle.Experimental validation efficiency challenges
The efficiency gap between ML predictions and experimental validation constitutes a key barrier from computational design to practical application. Computational predictions are rapid and low-cost, but experimental validation often involves long cycles and intensive resources, leading to slow feedback loops that hinder innovative iterations and translational applications. For example, DNA-encoded library screening tools like DEL-Dock accelerate virtual screening via ML but require optimization in experimental platform integration;224 DNA storage encoding85 and nucleic acid computing system114 optimizations are similarly limited by validation bottlenecks. Building efficient validation systems demands multidimensional technological innovation and system integration. At the platform level, closed-loop design-build-test-learn systems integrate automated equipment with feedback algorithms for rapid iterations. At the technical level, high-throughput microfluidic platforms support parallel testing, while in situ sequencing and real-time imaging provide immediate data feedback. At the strategic level, Bayesian optimization and active learning intelligently plan experimental paths to maximize information efficiency. Additionally, standardized validation protocols and data-sharing mechanisms will promote cross-institutional collaboration and accelerate translational applications. Fusion of these innovative elements is poised to bridge computational-experimental gaps, advancing nucleic acid molecular engineering toward efficient, reliable directions, provided that model development is tightly coupled to prospective, hypothesis-driven experiments to move beyond purely correlative predictions and toward causal understanding of nucleic acid structure–function relationships, while necessitating a broader examination of ethical and social implications alongside emerging future trajectories.Ethical and social implications
The rapid integration of ML into nucleic acid molecular engineering raises profound ethical and social considerations that must be addressed to ensure equitable benefits and minimize unintended harms. Central to these concerns is the potential for biased datasets, which often underrepresent diverse populations in genomic and structural data, leading to models that perpetuate health disparities in applications like CRISPR-based therapies or mRNA vaccines.6,14 For instance, if training data skew toward certain ethnic groups, predictive models for nucleic acid structure–function relationships could inadvertently exacerbate inequalities in precision medicine, as seen in broader AI-driven clinical care.225 Moreover, the interpretability challenges highlighted in nucleic acid performance modulation and application expansion pose risks in clinical deployment; opaque “black box” models may hinder accountability in gene editing tools, where off-target effects could have irreversible consequences.226 Socially, the dual-use nature of nucleic acid technologies demands robust governance frameworks because it enables both therapeutic advancements and potential biosecurity threats, such as engineered pathogens.227 To mitigate these, interdisciplinary collaborations involving ethicists, policymakers, and diverse stakeholders are essential, alongside initiatives for open-access data repositories to democratize nucleic acid engineering.228 Ultimately, proactive ethical oversight will be crucial to align this field's transformative potential with societal values, fostering trust and inclusivity in biomedicine and information sciences.Future directions
Looking ahead, ML-driven nucleic acid molecular engineering is poised for paradigm-shifting advancements through deeper integration of emerging technologies and multidisciplinary approaches, building on current progress in structure construction, performance modulation, and applications. A key trajectory involves hybrid models combining ML with quantum computing to simulate dynamic nucleic acid behaviors at atomic scales, addressing limitations in three-dimensional structure prediction and enabling real-time optimization of complex systems like DNA origami networks.229,230 In performance modulation, reinforcement learning could evolve toward adaptive frameworks that incorporate real-world feedback loops, enhancing gene editing precision across diverse cellular contexts and accelerating nucleic acid therapeutics from bench to bedside.81 For application expansion, generative AI models, inspired by large language models, hold promise for designing multifunctional nucleic acid platforms that integrate diagnostics, drug delivery, and computing in single systems, potentially revolutionizing point-of-care devices and bioelectronics.39To overcome data scarcity, federated learning paradigms could facilitate collaborative, privacy-preserving datasets across global institutions, while physics-informed NNs improve generalization for underrepresented nucleic acid motifs.231 In federated workflows, models are trained locally on institutional datasets and only parameter updates are shared, enabling multi-centre learning without transferring raw genomic or clinical records. Such approaches have already been demonstrated in medical imaging and electronic health records and could analogously support multi-cohort nucleic acid datasets while respecting regulatory constraints.232 At the same time, physics-informed neural networks embed known thermodynamic, kinetic, or structural constraints directly into the loss function, which can regularize training on small, biased datasets and improve extrapolation to rare sequence or structural contexts.231 Together, these strategies could substantially mitigate current data bottlenecks in ML-driven nucleic acid engineering.
Furthermore, expanding to non-canonical nucleic acids, such as xeno-nucleic acids, could unlock novel biomaterials with enhanced stability for environmental and space applications.233 Synthetic genetic polymers with modified backbones, sugars, or bases have already been shown to support heredity and Darwinian evolution while exhibiting markedly improved resistance to nucleases and chemical degradation.234,235 Such xeno-nucleic acids therefore provide an attractive substrate for designing information-bearing materials that function in extreme pH, temperature, or radiation environments where natural DNA and RNA would rapidly fail. ML-guided generative and predictive models could accelerate the discovery of xeno-nucleic acid aptamers, catalysts, and nanostructures with tailored stability and binding properties, extending nucleic acid molecular engineering beyond the canonical chemical space.
Realizing these directions will require investment in high-throughput experimental platforms and ethical AI guidelines, ultimately propelling nucleic acid engineering toward sustainable innovations that address global challenges in health, sustainability, and data management. On the experimental side, automated design–build–test–learn pipelines, microfluidic screening systems, and single-cell readouts will be essential to generate the large, high-quality datasets needed to close the loop between models and measurements.236,237 On the governance side, emerging frameworks for trustworthy and accountable AI in medicine emphasize transparency, bias assessment, and stakeholder engagement, which are equally relevant for ML-guided gene editing and therapeutics.238 Embedding these principles into nucleic acid engineering workflows will help ensure that resulting technologies are not only powerful but also safe, equitable, and socially acceptable.
Conclusions
ML-driven nucleic acid molecular engineering is profoundly reshaping paradigms in biomedicine and information sciences, as highlighted by the ethical considerations and future directions that underscore its potential for responsible advancement. At the same time, across structure construction, performance modulation, and application expansion, our survey makes clear that ML currently functions primarily as a powerful, context-dependent complement to biophysical modeling and carefully designed experiments, rather than a replacement for them. This review systematically surveys the latest progress in ML across three major domains: nucleic acid structure construction, performance modulation, and application expansion, highlighting the transformative potential of this interdisciplinary field. In structure design, DL models53,70,93,142,152 have realized full-chain innovations. These span from primary sequence design to secondary structure prediction and three-dimensional structure reconstruction. They significantly enhance design precision and efficiency. In performance modulation, ML algorithms use multifactor predictive models.50,117,182 These have successfully decoupled complex variables influencing nucleic acid functions. As a result, they achieve precise optimization of molecular properties, gene editing tools, and functional elements. In application expansion, ML-assisted nucleic acid molecular engineering has achieved breakthroughs in disease diagnosis, therapeutic drugs, and information storage and computing,119,213,220 propelling transitions from laboratory prototypes to clinical and industrial applications. These advancements not only mark a paradigm shift from empirical dependency to data-driven approaches but also provide solid foundations for systemic innovations in nucleic acid molecular engineering, provided that ethical safeguards and forward-looking strategies are integrated to mitigate risks and maximize societal benefits.Despite challenges like insufficient data quality and scale, limited model interpretability, and low experimental validation efficiency, ML-driven nucleic acid molecular engineering exhibits broad prospects. Self-supervised learning combined with data augmentation, explainable AI integrated with mechanism-driven models, and closed-loop design-build-test-learn platform construction will synergistically propel the field from static structure prediction toward dynamic behavior simulation, and from single-molecule design to complex system engineering. Looking ahead, ML will transcend traditional method limitations by constructing integrated sequence-structure-function models, ushering in a new era of nucleic acid design and applications. This interdisciplinary development will not only accelerate biomedical innovations but also provide key solutions for global challenges in human health, environmental monitoring, and information technology. Guided by ethical frameworks and innovative future directions, the continued fusion of algorithms, data resources, and experimental technologies will drive deeper advancements in ML-driven nucleic acid molecular engineering and contribute greater value to human well-being.
Author contributions
Q. S., M. L. and C. F. wrote the manuscript draft. All authors reviewed and edited the manuscript before submission.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Supplementary information (SI) contains Table S1 with references for each milestone in Fig. 1. See DOI: https://doi.org/10.1039/d5cs01091h.
Acknowledgements
Part of the research work described in this review was supported by the National Key R&D Program of China (2021YFF1200300), Shanghai Pilot Program for Basic Research, the National Natural Science Foundation of China (22574100, U24A20497, T2188102, 22322704, 21991134, 22122406), the Science Foundation of Shanghai Municipal Science and Technology Commission (23QA1404800), the Shanghai Science and Technology Innovation Action Plan (24ZR1433100), the Shanghai Key Technology R&D Program (25JC3201403), and the New Cornerstone Science Foundation.References
- J. A. Doudna and E. Charpentier, Science, 2014, 346, 1258096 CrossRef PubMed.
- H. Yan, Science, 2004, 306, 2048–2049 CrossRef CAS PubMed.
- N. C. Seeman and H. F. Sleiman, Nat. Rev. Mater., 2017, 3, 17068 CrossRef.
- P. Guo, Nat. Nanotechnol., 2010, 5, 833–842 CrossRef CAS PubMed.
- H. C. Metsky, N. L. Welch, P. P. Pillai, N. J. Haradhvala, L. Rumker, S. Mantena, Y. B. Zhang, D. K. Yang, C. M. Ackerman, J. Weller, P. C. Blainey, C. Myhrvold, M. Mitzenmacher and P. C. Sabeti, Nat. Biotechnol., 2022, 40, 1123–1131 CrossRef CAS PubMed.
- K. Karikó, H. Muramatsu, F. A. Welsh, J. Ludwig, H. Kato, S. Akira and D. Weissman, Mol. Ther., 2008, 16, 1833–1840 CrossRef PubMed.
- F. P. Polack, S. J. Thomas, N. Kitchin, J. Absalon, A. Gurtman, S. Lockhart, J. L. Perez, G. P. Marc, E. D. Moreira, C. Zerbini, R. Bailey, K. A. Swanson, S. Roychoudhury, K. Koury, P. Li, W. V. Kalina, D. Cooper, R. W. Frenck, L. L. Hammitt, Ö. Türeci, H. Nell, A. Schaefer, S. Ünal, D. B. Tresnan, S. Mather, P. R. Dormitzer, U. Şahin, K. U. Jansen and W. C. Gruber, N. Engl. J. Med., 2020, 383, 2603–2615 CrossRef CAS PubMed.
- U. Sahin, A. Muik, E. Derhovanessian, I. Vogler, L. M. Kranz, M. Vormehr, A. Baum, K. Pascal, J. Quandt, D. Maurus, S. Brachtendorf, V. Lörks, J. Sikorski, R. Hilker, D. Becker, A.-K. Eller, J. Grützner, C. Boesler, C. Rosenbaum, M.-C. Kühnle, U. Luxemburger, A. Kemmer-Brück, D. Langer, M. Bexon, S. Bolte, K. Karikó, T. Palanche, B. Fischer, A. Schultz, P.-Y. Shi, C. Fontes-Garfias, J. L. Perez, K. A. Swanson, J. Loschko, I. L. Scully, M. Cutler, W. Kalina, C. A. Kyratsous, D. Cooper, P. R. Dormitzer, K. U. Jansen and Ö. Türeci, Nature, 2020, 586, 594–599 CrossRef CAS PubMed.
- K. Pardee, A. A. Green, T. Ferrante, D. E. Cameron, A. DaleyKeyser, P. Yin and J. J. Collins, Cell, 2014, 159, 940–954 CrossRef CAS PubMed.
- A. A. Green, P. A. Silver, J. J. Collins and P. Yin, Cell, 2014, 159, 925–939 CrossRef CAS PubMed.
- G. M. Church, Y. Gao and S. Kosuri, Science, 2012, 337, 1628 CrossRef CAS PubMed.
- P. Mali, L. Yang, K. M. Esvelt, J. Aach, M. Guell, J. E. DiCarlo, J. E. Norville and G. M. Church, Science, 2013, 339, 823–826 CrossRef CAS PubMed.
- L. Cong, F. A. Ran, D. Cox, S. Lin, R. Barretto, N. Habib, P. D. Hsu, X. Wu, W. Jiang, L. A. Marraffini and F. Zhang, Science, 2013, 339, 819–823 CrossRef CAS PubMed.
- M. Jinek, K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna and E. Charpentier, Science, 2012, 337, 816–821 CrossRef CAS PubMed.
- W. Tan, H. Wang, Y. Chen, X. Zhang, H. Zhu, C. Yang, R. Yang and C. Liu, Trends Biotechnol., 2011, 29, 634–640 CrossRef CAS PubMed.
- C. Tuerk and L. Gold, Science, 1990, 249, 505–510 CrossRef CAS PubMed.
- N. Goldman, P. Bertone, S. Chen, C. Dessimoz, E. M. LeProust, B. Sipos and E. Birney, Nature, 2013, 494, 77–80 CrossRef CAS PubMed.
- L. Qian and E. Winfree, Science, 2011, 332, 1196–1201 CrossRef CAS PubMed.
- J. Bath and A. J. Turberfield, Nat. Nanotechnol., 2007, 2, 275–284 CrossRef CAS PubMed.
- P. W. K. Rothemund, Nature, 2006, 440, 297–302 CrossRef CAS PubMed.
- E. Shapiro and B. Gil, Science, 2008, 322, 387–388 CrossRef CAS PubMed.
- S. M. Douglas, H. Dietz, T. Liedl, B. Högberg, F. Graf and W. M. Shih, Nature, 2009, 459, 414–418 CrossRef CAS PubMed.
- J. N. Zadeh, C. D. Steenberg, J. S. Bois, B. R. Wolfe, M. B. Pierce, A. R. Khan, R. M. Dirks and N. A. Pierce, J. Comput. Chem., 2011, 32, 170–173 CrossRef CAS PubMed.
- R. M. Dirks, Nucleic Acids Res., 2004, 32, 1392–1403 CrossRef CAS PubMed.
- J. Zhang, Y. Fei, L. Sun and Q. C. Zhang, Nat. Methods, 2022, 19, 1193–1207 CrossRef CAS PubMed.
- T. K. Lu, A. S. Khalil and J. J. Collins, Nat. Biotechnol., 2009, 27, 1139–1150 CrossRef CAS PubMed.
- N. Pardi, M. J. Hogan, F. W. Porter and D. Weissman, Nat. Rev. Drug Discovery, 2018, 17, 261–279 CrossRef CAS PubMed.
- A. S. Khalil and J. J. Collins, Nat. Rev. Genet., 2010, 11, 367–379 CrossRef CAS PubMed.
- K. T. Butler, D. W. Davies, H. Cartwright, O. Isayev and A. Walsh, Nature, 2018, 559, 547–555 CrossRef CAS PubMed.
- Y. LeCun, Y. Bengio and G. Hinton, Nature, 2015, 521, 436–444 CrossRef CAS PubMed.
- M. I. Jordan and T. M. Mitchell, Science, 2015, 349, 255–260 CrossRef CAS PubMed.
- M. H. S. Segler, M. Preuss and M. P. Waller, Nature, 2018, 555, 604–610 CrossRef CAS PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O’Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- M. Ziatdinov, A. Ghosh, C. Y. (Tommy) Wong and S. V. Kalinin, Nat. Mach. Intell., 2022, 4, 1101–1112 CrossRef.
- S. Hochreiter and J. Schmidhuber, Neural Comput., 1997, 9, 1735–1780 CrossRef CAS PubMed.
- D. P. Kingma and M. Welling, arXiv, 2013, preprint, arXiv:1312.6114 DOI:10.48550/arXiv.1312.6114.
- L. Rao, Y. Yuan, X. Shen, G. Yu and X. Chen, Nat. Nanotechnol., 2024, 1–13 Search PubMed.
- J. Devlin, M.-W. Chang, K. Lee and K. Toutanova, Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, (long and Short Papers), Association for Computational Linguistics, Minneapolis, Minnesota, 2019, vol. 1, pp. 4171–4186 Search PubMed.
- T. Brown, B. Mann, N. Ryder, M. Subbiah, J. D. Kaplan, P. Dhariwal, A. Neelakantan, P. Shyam, G. Sastry, A. Askell, S. Agarwal, A. Herbert-Voss, G. Krueger, T. Henighan, R. Child, A. Ramesh, D. Ziegler, J. Wu, C. Winter, C. Hesse, M. Chen, E. Sigler, M. Litwin, S. Gray, B. Chess, J. Clark, C. Berner, S. McCandlish, A. Radford, I. Sutskever and D. Amodei, Adv. Neural Information Process. Syst., 2020, 33, 1877–1901 Search PubMed.
- A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, Ł. Ukasz Kaiser and I. Polosukhin, Advances in Neural Information Processing Systems, Curran Associates, Inc., 2017, vol. 30, pp. 5998–6008 Search PubMed.
- G. Biau and E. Scornet, Test, 2016, 25, 197–227 CrossRef.
- D. Bzdok, N. Altman and M. Krzywinski, Nat. Methods, 2018, 15, 233–234 CrossRef CAS PubMed.
- C. Cortes and V. Vapnik, Mach. Learn., 1995, 20, 273–297 CrossRef.
- T. Shen, Z. Hu, S. Sun, D. Liu, F. Wong, J. Wang, J. Chen, Y. Wang, L. Hong, J. Xiao, L. Zheng, T. Krishnamoorthi, I. King, S. Wang, P. Yin, J. J. Collins and Y. Li, Nat. Methods, 2024, 21, 2287–2298 CrossRef CAS PubMed.
- W. Wang, C. Feng, R. Han, Z. Wang, L. Ye, Z. Du, H. Wei, F. Zhang, Z. Peng and J. Yang, Nat. Commun., 2023, 14, 7266 CrossRef CAS PubMed.
- A. W. Senior, R. Evans, J. Jumper, J. Kirkpatrick, L. Sifre, T. Green, C. Qin, A. Žídek, A. W. R. Nelson, A. Bridgland, H. Penedones, S. Petersen, K. Simonyan, S. Crossan, P. Kohli, D. T. Jones, D. Silver, K. Kavukcuoglu and D. Hassabis, Nature, 2020, 577, 706–710 CrossRef CAS PubMed.
- J. G. Greener, S. M. Kandathil and D. T. Jones, Nat. Commun., 2019, 10, 3977 CrossRef PubMed.
- J. M. Stokes, K. Yang, K. Swanson, W. Jin, A. Cubillos-Ruiz, N. M. Donghia, C. R. MacNair, S. French, L. A. Carfrae, Z. Bloom-Ackermann, V. M. Tran, A. Chiappino-Pepe, A. H. Badran, I. W. Andrews, E. J. Chory, G. M. Church, E. D. Brown, T. S. Jaakkola, R. Barzilay and J. J. Collins, Cell, 2020, 180, 688–702.e13 CrossRef CAS PubMed.
- W. Li, Y. Wen, K. Wang, Z. Ding, L. Wang, Q. Chen, L. Xie, H. Xu and H. Zhao, Nat. Commun., 2024, 15, 2603 CrossRef CAS PubMed.
- J. A. Valeri, K. M. Collins, P. Ramesh, M. A. Alcantar, B. A. Lepe, T. K. Lu and D. M. Camacho, Nat. Commun., 2020, 11, 5058 CrossRef CAS PubMed.
- Y.-T. Lee, M. F. S. Degenhardt, I. Skeparnias, H. F. Degenhardt, Y. R. Bhandari, P. Yu, J. R. Stagno, L. Fan, J. Zhang and Y.-X. Wang, Nature, 2025, 637, 1244–1251 CrossRef CAS PubMed.
- R. J. L. Townshend, S. Eismann, A. M. Watkins, R. Rangan, M. Karelina, R. Das and R. O. Dror, Science, 2021, 373, 1047–1051 CrossRef CAS PubMed.
- Y. Kagaya, Z. Zhang, N. Ibtehaz, X. Wang, T. Nakamura, P. D. Punuru and D. Kihara, Nat. Commun., 2025, 16, 881 CrossRef CAS PubMed.
- H. K. Kim, S. Min, M. Song, S. Jung, J. W. Choi, Y. Kim, S. Lee, S. Yoon and H. (Henry) Kim, Nat. Biotechnol., 2018, 36, 239 CrossRef CAS PubMed.
- N. Mathis, A. Allam, L. Kissling, K. F. Marquart, L. Schmidheini, C. Solari, Z. Balázs, M. Krauthammer and G. Schwank, Nat. Biotechnol., 2023, 41, 1151–1159 CrossRef CAS PubMed.
- H.-H. Wessels, A. Stirn, A. Méndez-Mancilla, E. J. Kim, S. K. Hart, D. A. Knowles and N. E. Sanjana, Nat. Biotechnol., 2024, 42, 628–637 CrossRef CAS PubMed.
- F. Wong, D. He, A. Krishnan, L. Hong, A. Z. Wang, J. Wang, Z. Hu, S. Omori, A. Li, J. Rao, Q. Yu, W. Jin, T. Zhang, K. Ilia, J. X. Chen, S. Zheng, I. King, Y. Li and J. J. Collins, Nat. Comput. Sci., 2024, 4, 829–839 CrossRef CAS PubMed.
- J. C. Chen, J. P. Chen, M. W. Shen, M. Wornow, M. Bae, W.-H. Yeh, A. Hsu and D. R. Liu, Nat. Commun., 2022, 13, 4541 CrossRef CAS PubMed.
- S. Sumi, M. Hamada and H. Saito, Nat. Methods, 2024, 21, 435–443 CrossRef CAS PubMed.
- G. Eraslan, Ž. Avsec, J. Gagneur and F. J. Theis, Nat. Rev. Genet., 2019, 20, 389–403 CrossRef CAS PubMed.
- C. Truong-Quoc, J. Y. Lee, K. S. Kim and D.-N. Kim, Nat. Mater., 2024, 23, 984–992 CrossRef CAS PubMed.
- N. Sapoval, A. Aghazadeh, M. G. Nute, D. A. Antunes, A. Balaji, R. Baraniuk, C. J. Barberan, R. Dannenfelser, C. Dun, M. Edrisi, R. A. L. Elworth, B. Kille, A. Kyrillidis, L. Nakhleh, C. R. Wolfe, Z. Yan, V. Yao and T. J. Treangen, Nat. Commun., 2022, 13, 1728 CrossRef CAS PubMed.
- K. M. Boehm, P. Khosravi, R. Vanguri, J. Gao and S. P. Shah, Nat. Rev. Cancer, 2022, 22, 114–126 CrossRef CAS PubMed.
- H. Lv, N. Xie, M. Li, M. Dong, C. Sun, Q. Zhang, L. Zhao, J. Li, X. Zuo, H. Chen, F. Wang and C. Fan, Nature, 2023, 622, 292–300 CrossRef CAS PubMed.
- J. Yin, S. Wang, J. Wang, Y. Zhang, C. Fan, J. Chao, Y. Gao and L. Wang, Nat. Mater., 2024, 23, 854–862 CrossRef CAS PubMed.
- L. Li, J. Yin, W. Ma, L. Tang, J. Zou, L. Yang, T. Du, Y. Zhao, L. Wang, Z. Yang, C. Fan, J. Chao and X. Chen, Nat. Mater., 2024, 23, 993–1001 CrossRef CAS PubMed.
- X. Liu, F. Zhang, X. Jing, M. Pan, P. Liu, W. Li, B. Zhu, J. Li, H. Chen, L. Wang, J. Lin, Y. Liu, D. Zhao, H. Yan and C. Fan, Nature, 2018, 559, 593–598 CrossRef CAS PubMed.
- F. Praetorius, B. Kick, K. L. Behler, M. N. Honemann, D. Weuster-Botz and H. Dietz, Nature, 2017, 552, 84–87 CrossRef CAS PubMed.
- Y. Shulgina, M. I. Trinidad, C. J. Langeberg, H. Nisonoff, S. Chithrananda, P. Skopintsev, A. J. Nissley, J. Patel, R. S. Boger, H. Shi, P. H. Yoon, E. E. Doherty, T. Pande, A. M. Iyer, J. A. Doudna and J. H. D. Cate, Nat. Commun., 2024, 15, 10627 CrossRef CAS PubMed.
- K. Sato, M. Akiyama and Y. Sakakibara, Nat. Commun., 2021, 12, 941 CrossRef CAS PubMed.
- J. Shor, E. Strand and C. Y. McLean, bioRxiv, 2025, preprint DOI:10.1101/2025.06.20.660785.
- A. T. Riley, J. M. Robson, A. Ulanova and A. A. Green, Nat. Commun., 2025, 16, 4155 CrossRef CAS PubMed.
- H. Zhang, H. Liu, Y. Xu, H. Huang, Y. Liu, J. Wang, Y. Qin, H. Wang, L. Ma, Z. Xun, X. Hou, T. K. Lu and J. Cao, Science, 2025, 0, eadr8470 CrossRef CAS PubMed.
- M. Baek, R. Mchugh, I. Anishchenko, H. Jiang, D. Baker and F. DiMaio, Nat. Methods, 2024, 21, 117–121 CrossRef CAS PubMed.
- M. F. S. Degenhardt, H. F. Degenhardt, Y. R. Bhandari, Y.-T. Lee, J. Ding, P. Yu, W. F. Heinz, J. R. Stagno, C. D. Schwieters, N. R. Watts, P. T. Wingfield, A. Rein, J. Zhang and Y.-X. Wang, Nature, 2024, 637, 1234–1243 CrossRef PubMed.
- X. Wang, G. Terashi and D. Kihara, Nat. Methods, 2023, 20, 1739–1747 CrossRef CAS PubMed.
- M. Chiriboga, C. M. Green, D. A. Hastman, D. Mathur, Q. Wei, S. A. Díaz, I. L. Medintz and R. Veneziano, Sci. Rep., 2022, 12, 3871 CrossRef CAS PubMed.
- A. K. Hanumanthappa, J. Singh, K. Paliwal, J. Singh and Y. Zhou, Bioinformatics, 2020, 36, 5169–5176 CrossRef CAS PubMed.
- S. Sun, Q. Wu, Z. Peng and J. Yang, Bioinformatics, 2019, 35, 1686–1691 CrossRef CAS PubMed.
- G. Chuai, H. Ma, J. Yan, M. Chen, N. Hong, D. Xue, C. Zhou, C. Zhu, K. Chen, B. Duan, F. Gu, S. Qu, D. Huang, J. Wei and Q. Liu, Genome Biol., 2018, 19, 80 CrossRef PubMed.
- Y. Qu, K. Huang, M. Yin, K. Zhan, D. Liu, D. Yin, H. C. Cousins, W. A. Johnson, X. Wang, M. Shah, R. B. Altman, D. Zhou, M. Wang and L. Cong, Nat. Biomed. Eng., 2025, 1–14 Search PubMed.
- M. U. Ahsan, A. Gouru, J. Chan, W. Zhou and K. Wang, Nat. Commun., 2024, 15, 1448 CrossRef CAS PubMed.
- D. Galeano, Imrat, J. Haltom, C. Andolino, A. Yousey, V. Zaksas, S. Das, S. B. Baylin, D. C. Wallace, F. J. Slack, F. J. Enguita, E. S. Wurtele, D. Teegarden, R. Meller, D. Cifuentes and A. Beheshti, Nat. Commun., 2024, 15, 9149 CrossRef CAS PubMed.
- D. Bar-Lev, I. Orr, O. Sabary, T. Etzion and E. Yaakobi, Nat. Mach. Intell., 2025, 7, 639–649 CrossRef.
- C. Pan, S. K. Tabatabaei, S. M. H. Tabatabaei Yazdi, A. G. Hernandez, C. M. Schroeder and O. Milenkovic, Nat. Commun., 2022, 13, 2984 CrossRef CAS PubMed.
- Y. Yang, M. Zheng and A. Jagota, npj Comput. Mater., 2019, 5, 3 CrossRef.
- Z. Lin, Y. Yang, A. Jagota and M. Zheng, ACS Nano, 2022, 16, 4705–4713 CrossRef CAS PubMed.
- S. M. Copp, P. Bogdanov, M. Debord, A. Singh and E. Gwinn, Adv. Mater., 2014, 26, 5839–5845 CrossRef CAS PubMed.
- N. Killoran, L. J. Lee, A. Delong, D. Duvenaud and B. J. Frey, arXiv, 2017, preprint, arXiv:1712.06148 DOI:10.48550/arXiv.1712.06148.
- T. Yang, M. Han, X. Wen and Y. Zheng, J. Artif. Intell. Bioinform., 2025, 1, 12 CAS.
- P. Mastracco, A. Gonzàlez-Rosell, J. Evans, P. Bogdanov and S. M. Copp, ACS Nano, 2022, 16, 16322–16331 CrossRef CAS PubMed.
- S. M. Copp, S. M. Swasey, A. Gorovits, P. Bogdanov and E. G. Gwinn, Chem. Mater., 2020, 32, 430–437 CrossRef CAS.
- F. Zhai, Y. Guan, Y. Li, S. Chen and R. He, ACS Appl. Nano Mater., 2022, 5, 9615–9624 CrossRef CAS.
- S. M. Halper, A. Hossain and H. M. Salis, ACS Synth. Biol., 2020, 9, 1563–1571 CrossRef CAS PubMed.
- P. Kelich, S. Jeong, N. Navarro, J. Adams, X. Sun, H. Zhao, M. P. Landry and L. Vuković, ACS Nano, 2022, 16, 736–745 CrossRef CAS PubMed.
- A. Gupta and J. Zou, Nat. Mach. Intell., 2019, 1, 105–111 CrossRef.
- E.-M. Nikolados, A. Wongprommoon, O. M. Aodha, G. Cambray and D. A. Oyarzún, Nat. Commun., 2022, 13, 7755 CrossRef CAS PubMed.
- Q. Zhang, S. M. Azarin and C. A. Sarkar, Nat. Commun., 2022, 13, 4152 CrossRef CAS PubMed.
- J. X. Zhang, B. Yordanov, A. Gaunt, M. X. Wang, P. Dai, Y.-J. Chen, K. Zhang, J. Z. Fang, N. Dalchau, J. Li, A. Phillips and D. Y. Zhang, Nat. Commun., 2021, 12, 4387 CrossRef CAS PubMed.
- P. Eastman, J. Shi, B. Ramsundar and V. S. Pande, PLoS Comput. Biol., 2018, 14, e1006176 CrossRef PubMed.
- J. A. Nelder and R. W. M. Wedderburn, R. Stat. Soc., J., A: Gen., 1972, 135, 371–384 Search PubMed.
- T. Cover and P. Hart, IEEE Trans. Inf. Theory, 1967, 13, 21–27 Search PubMed.
- W. S. McCulloch and W. Pitts, Bull. Math. Biophys., 1943, 5, 115–133 CrossRef.
- J. H. Lam, Y. Li, L. Zhu, R. Umarov, H. Jiang, A. Héliou, F. K. Sheong, T. Liu, Y. Long, Y. Li, L. Fang, R. B. Altman, W. Chen, X. Huang and X. Gao, Nat. Commun., 2019, 10, 4941 CrossRef PubMed.
- N. Abe, I. Dror, L. Yang, M. Slattery, T. Zhou, H. J. Bussemaker, R. Rohs and R. S. Mann, Cell, 2015, 161, 307–318 CrossRef CAS PubMed.
- R. Mitra, J. Li, J. M. Sagendorf, Y. Jiang, A. S. Cohen, T.-P. Chiu, C. J. Glasscock and R. Rohs, Nat. Methods, 2024, 21, 1674–1683 CrossRef CAS PubMed.
- F. Liu, S. Huang, J. Hu, X. Chen, Z. Song, J. Dong, Y. Liu, X. Huang, S. Wang, X. Wang and W. Shu, Nat. Mach. Intell., 2023, 5, 1261–1274 CrossRef.
- Y. Wang, S. Zhang, W. Jia, P. Fan, L. Wang, X. Li, J. Chen, Z. Cao, X. Du, Y. Liu, K. Wang, C. Hu, J. Zhang, J. Hu, P. Zhang, H.-Y. Chen and S. Huang, Nat. Nanotechnol., 2022, 17, 976–983 CrossRef CAS PubMed.
- Y. Wu, W. Shao, M. Yan, Y. Wang, P. Xu, G. Huang, X. Li, B. D. Gregory, J. Yang, H. Wang and X. Yu, Nat. Commun., 2024, 15, 4049 CrossRef CAS PubMed.
- A. Bashir, Q. Yang, J. Wang, S. Hoyer, W. Chou, C. McLean, G. Davis, Q. Gong, Z. Armstrong, J. Jang, H. Kang, A. Pawlosky, A. Scott, G. E. Dahl, M. Berndl, M. Dimon and B. S. Ferguson, Nat. Commun., 2021, 12, 2366 CrossRef CAS PubMed.
- X. Wang, E. Alnabati, T. W. Aderinwale, S. R. Maddhuri Venkata Subramaniya, G. Terashi and D. Kihara, Nat. Commun., 2021, 12, 2302 CrossRef CAS PubMed.
- Y.-J. Kim, J. Lim and D.-N. Kim, Small, 2022, 18, 2103779 CrossRef CAS PubMed.
- T. Li, H. Cao, J. He and S.-Y. Huang, Nat. Commun., 2024, 15, 9367 CrossRef CAS PubMed.
- C. Bee, Y.-J. Chen, M. Queen, D. Ward, X. Liu, L. Organick, G. Seelig, K. Strauss and L. Ceze, Nat. Commun., 2021, 12, 4764 CrossRef CAS PubMed.
- V. Pan, W. Wang, I. Heaven, T. Bai, Y. Cheng, C. Chen, Y. Ke and B. Wei, ACS Nano, 2021, 15, 15892–15901 CrossRef CAS PubMed.
- Y. Wang, Y. Feng, Z. Xiao and Y. Luo, Food Chem., 2025, 463, 141115 CrossRef CAS PubMed.
- D. P. Cetnar, A. Hossain, G. E. Vezeau and H. M. Salis, Nat. Commun., 2024, 15, 9601 CrossRef CAS PubMed.
- H. K. Wayment-Steele, W. Kladwang, A. M. Watkins, D. S. Kim, B. Tunguz, W. Reade, M. Demkin, J. Romano, R. Wellington-Oguri, J. J. Nicol, J. Gao, K. Onodera, K. Fujikawa, H. Mao, G. Vandewiele, M. Tinti, B. Steenwinckel, T. Ito, T. Noumi, S. He, K. Ishi, Y. Lee, F. Öztürk, K. Y. Chiu, E. Öztürk, K. Amer, M. Fares, Eterna Participants and R. Das, Nat. Mach. Intell., 2022, 4, 1174–1184 CrossRef PubMed.
- B. Li, I. O. Raji, A. G. R. Gordon, L. Sun, T. M. Raimondo, F. A. Oladimeji, A. Y. Jiang, A. Varley, R. S. Langer and D. G. Anderson, Nat. Mater., 2024, 23, 1002–1008 CrossRef CAS PubMed.
- G. Yamankurt, E. J. Berns, A. Xue, A. Lee, N. Bagheri, M. Mrksich and C. A. Mirkin, Nat. Biomed. Eng., 2019, 3, 318–327 CrossRef CAS PubMed.
- S. Pitafi, T. Anwar and Z. Sharif, Appl. Sci., 2023, 13, 3529 CrossRef CAS.
- A. A. Wani, PeerJ Comput. Sci., 2025, 11, e3025 CrossRef PubMed.
- I. D. Mienye and T. G. Swart, Arch. Comput. Methods Eng., 2025, 32, 3981–4000 CrossRef.
- N. Iwano, T. Adachi, K. Aoki, Y. Nakamura and M. Hamada, Nat. Comput. Sci., 2022, 2, 378–386 CrossRef PubMed.
- J. Perez Tobia, P.-J. J. Huang, Y. Ding, R. Saran Narayan, A. Narayan and J. Liu, ACS Synth. Biol., 2023, 12, 186–195 CrossRef CAS PubMed.
- L. T. Schmitt, M. Paszkowski-Rogacz, F. Jug and F. Buchholz, Nat. Commun., 2022, 13, 7966 CrossRef CAS PubMed.
- M.-R. Amini, V. Feofanov, L. Pauletto, L. Hadjadj, É. Devijver and Y. Maximov, Neurocomputing, 2025, 616, 128904 CrossRef.
- X. Yang, Z. Song, I. King and Z. Xu, IEEE Trans. Knowl. Data Eng., 2023, 35, 8934–8954 Search PubMed.
- Y. Chong, Y. Ding, Q. Yan and S. Pan, Neurocomputing, 2020, 408, 216–230 CrossRef.
- Y. Ma, Y. Zheng, W. Zhang, B. Wei, Z. Lin, W. Liu and Z. Li, Int. J. Intell. Comput. Cybern, 2024, 17, 705–736 CrossRef.
- A. Blum and T. Mitchell, in Proceedings of the eleventh annual conference on Computational learning theory, Association for Computing Machinery, New York, NY, USA, 1998, pp. 92–100 Search PubMed.
- F. Garcia and E. Rachelson, Markov decision processes in artificial intelligence, John Wiley & Sons, Ltd, 2013, pp. 1–38 Search PubMed.
- C. J. C. H. Watkins, Learning from delayed rewards, King’s College, 1989 Search PubMed.
- E. H. Sumiea, S. J. Abdulkadir, H. S. Alhussian, S. M. Al-Selwi, A. Alqushaibi, M. G. Ragab and S. M. Fati, Heliyon, 2024, 10, e30697 CrossRef PubMed.
- X. Chen, Y. Li, R. Umarov, X. Gao and L. Song, arXiv, 2020, preprint, arXiv:2002.05810 DOI:10.48550/arXiv.2002.05810.
- G. Xu, Y. Bao, Y. Zhang, X. Xiang, H. Luo and X. Guo, Anal. Chem., 2024, 96, 17109–17117 CrossRef CAS PubMed.
- M. Zeraati, D. B. Langley, P. Schofield, A. L. Moye, R. Rouet, W. E. Hughes, T. M. Bryan, M. E. Dinger and D. Christ, Nat. Chem., 2018, 10, 631–637 CrossRef CAS PubMed.
- K. Sato, J. Lyu, J. van den Berg, D. Braat, V. M. Cruz, C. Navarro Luzón, J. Schimmel, C. Esteban-Jurado, M. Alemany, J. Dreyer, A. Hendrikx, F. Mattiroli, A. van Oudenaarden, M. Tijsterman, S. J. Elsässer and P. Knipscheer, Science, 2025, 388, 1225–1231 CrossRef CAS PubMed.
- M. Zuker, Nucleic Acids Res., 2003, 31, 3406–3415 CrossRef CAS PubMed.
- R. Lorenz, S. H. Bernhart, C. Höner zu Siederdissen, H. Tafer, C. Flamm, P. F. Stadler and I. L. Hofacker, Algorithms Mol. Biol., 2011, 6, 26 CrossRef PubMed.
- M. E. Fornace, J. Huang, C. T. Newman, N. J. Porubsky, M. B. Pierce and N. A. Pierce, ChemRxiv, 2022, preprint DOI:10.26434/chemrxiv-2022-xv98l.
- J. Singh, J. Hanson, K. Paliwal and Y. Zhou, Nat. Commun., 2019, 10, 5407 CrossRef PubMed.
- M. Barshai, B. Engel, I. Haim and Y. Orenstein, PLoS Comput. Biol., 2023, 19, e1010948 CrossRef CAS PubMed.
- D. Liew, Z. W. Lim and E. H. Yong, Sci. Rep., 2024, 14, 24238 CrossRef CAS PubMed.
- A. B. Sahakyan, V. S. Chambers, G. Marsico, T. Santner, M. Di Antonio and S. Balasubramanian, Sci. Rep., 2017, 7, 14535 CrossRef PubMed.
- K. Sato and M. Hamada, Briefings Bioinf., 2023, 24, bbad186 CrossRef PubMed.
- S. Li, F. Dong, Y. Wu, S. Zhang, C. Zhang, X. Liu, T. Jiang and J. Zeng, Nucleic Acids Res., 2017, 45, e129–e129 CrossRef CAS PubMed.
- W. Zeng, Y. Dou, L. Pan, L. Xu and S. Peng, Nat. Commun., 2024, 15, 7838 CrossRef CAS PubMed.
- C. Nithin, S. Kmiecik, R. Błaszczyk, J. Nowicka and I. Tuszyńska, Nucleic Acids Res., 2024, 52, 7465–7486 CrossRef CAS PubMed.
- J. Li and R. Rohs, Nucleic Acids Res., 2024, 52, W7–W12 CrossRef PubMed.
- N. Katz, E. Tripto, N. Granik, S. Goldberg, O. Atar, Z. Yakhini, Y. Orenstein and R. Amit, Nat. Commun., 2021, 12, 1576 CrossRef CAS PubMed.
- Y. Li, C. Zhang, C. Feng, R. Pearce, P. Lydia Freddolino and Y. Zhang, Nat. Commun., 2023, 14, 5745 CrossRef CAS PubMed.
- J. Li, T.-P. Chiu and R. Rohs, Nat. Commun., 2024, 15, 1243 CrossRef CAS PubMed.
- A. Kabir, M. Bhattarai, S. Peterson, Y. Najman-Licht, K. Ø. Rasmussen, A. Shehu, A. R. Bishop, B. Alexandrov and A. Usheva, Nucleic Acids Res., 2024, 52, e91–e91 CrossRef CAS PubMed.
- S. Barissi, A. Sala, M. Wieczór, F. Battistini and M. Orozco, Nucleic Acids Res., 2022, 50, 9105–9114 CrossRef CAS PubMed.
- A. G. B. Grønning, T. K. Doktor, S. J. Larsen, U. S. S. Petersen, L. L. Holm, G. H. Bruun, M. B. Hansen, A.-M. Hartung, J. Baumbach and B. S. Andresen, Nucleic Acids Res., 2020, 48, 7099–7118 Search PubMed.
- J. S. Dialpuri, J. Agirre, K. D. Cowtan and P. S. Bond, Nucleic Acids Res., 2024, 52, e84–e84 CrossRef CAS PubMed.
- S. Zhang, J. Zhou, H. Hu, H. Gong, L. Chen, C. Cheng and J. Zeng, Nucleic Acids Res., 2016, 44, e32–e32 CrossRef PubMed.
- J. Huzar, R. Coreas, M. P. Landry and G. Tikhomirov, ACS Nano, 2025, 19, 4333–4345 CrossRef CAS PubMed.
- K. Yazdani, D. Jordan, M. Yang, C. R. Fullenkamp, D. R. Calabrese, R. Boer, T. Hilimire, T. E. H. Allen, R. T. Khan and J. S. Schneekloth Jr., Angew. Chem., Int. Ed., 2023, 62, e202211358 CrossRef CAS PubMed.
- R. J. Penić, T. Vlašić, R. G. Huber, Y. Wan and M. Šikić, Nat. Commun., 2025, 16, 5671 CrossRef PubMed.
- A.-C. Groher, S. Jager, C. Schneider, F. Groher, K. Hamacher and B. Suess, ACS Synth. Biol., 2019, 8, 34–44 CrossRef CAS PubMed.
- H. K. Kim, Y. Kim, S. Lee, S. Min, J. Y. Bae, J. W. Choi, J. Park, D. Jung, S. Yoon and H. H. Kim, Sci. Adv., 2019, 5, eaax9249 CrossRef CAS PubMed.
- P. Picchetti, S. Volpi, M. Sancho-Albero, M. Rossetti, M. D. Dore, T. Trinh, F. Biedermann, M. Neri, A. Bertucci, A. Porchetta, R. Corradini, H. Sleiman and L. De Cola, J. Am. Chem. Soc., 2023, 145, 22903–22912 CrossRef CAS PubMed.
- B. B. Mendes, J. Conniot, A. Avital, D. Yao, X. Jiang, X. Zhou, N. Sharf-Pauker, Y. Xiao, O. Adir, H. Liang, J. Shi, A. Schroeder and J. Conde, Nat. Rev. Methods Primers, 2022, 2, 24 CrossRef CAS PubMed.
- Y.-H. Peng, S.-K. Hsiao, K. Gupta, A. Ruland, G. K. Auernhammer, M. F. Maitz, S. Boye, J. Lattner, C. Gerri, A. Honigmann, C. Werner and E. Krieg, Nat. Nanotechnol., 2023, 18, 1463–1473 CrossRef CAS PubMed.
- H. Tateishi-Karimata and N. Sugimoto, Nucleic Acids Res., 2014, 42, 8831–8844 CrossRef CAS PubMed.
- S. Kumar, A. Pearse, Y. Liu and R. E. Taylor, Nat. Commun., 2020, 11, 2960 CrossRef CAS PubMed.
- D. M. Camacho, K. M. Collins, R. K. Powers, J. C. Costello and J. J. Collins, Cell, 2018, 173, 1581–1592 CrossRef CAS PubMed.
- Y. Yang, X. Li, H. Zhao, J. Zhan, J. Wang and Y. Zhou, RNA, 2017, 23, 14–22 CrossRef CAS PubMed.
- H. Zhang, L. Zhang, D. H. Mathews and L. Huang, Bioinformatics, 2020, 36, i258–i267 CrossRef CAS PubMed.
- X.-Q. Fan, J. Hu, Y.-X. Tang, N.-X. Jia, D.-J. Yu and G.-J. Zhang, Anal. Biochem., 2022, 654, 114802 CrossRef CAS PubMed.
- K. Li, M. Carroll, R. Vafabakhsh, X. A. Wang and J.-P. Wang, Nucleic Acids Res., 2022, 50, 3142–3154 CrossRef CAS PubMed.
- T. Li, Y. Yang, H. Qi, W. Cui, L. Zhang, X. Fu, X. He, M. Liu, P. Li and T. Yu, Signal Transduction Targeted Ther., 2023, 8, 36 CrossRef PubMed.
- J. A. Ruffolo, S. Nayfach, J. Gallagher, A. Bhatnagar, J. Beazer, R. Hussain, J. Russ, J. Yip, E. Hill, M. Pacesa, A. J. Meeske, P. Cameron and A. Madani, Nature, 2025, 1, 1–8 Search PubMed.
- M. Pacesa, O. Pelea and M. Jinek, Cell, 2024, 187, 1076–1100 CrossRef CAS PubMed.
- Y. Zheng, Y. Li, K. Zhou, T. Li, N. J. VanDusen and Y. Hua, Signal Transduction Targeted Ther., 2024, 9, 47 CrossRef PubMed.
- Q. Chen, G. Chuai, H. Zhang, J. Tang, L. Duan, H. Guan, W. Li, W. Li, J. Wen, E. Zuo, Q. Zhang and Q. Liu, Nat. Commun., 2023, 14, 7521 CrossRef CAS PubMed.
- D. T. Ham, T. S. Browne, P. N. Banglorewala, T. L. Wilson, R. K. Michael, G. B. Gloor and D. R. Edgell, Nat. Commun., 2023, 14, 5514 CrossRef CAS PubMed.
- X. Xiang, G. I. Corsi, C. Anthon, K. Qu, X. Pan, X. Liang, P. Han, Z. Dong, L. Liu, J. Zhong, T. Ma, J. Wang, X. Zhang, H. Jiang, F. Xu, X. Liu, X. Xu, J. Wang, H. Yang, L. Bolund, G. M. Church, L. Lin, J. Gorodkin and Y. Luo, Nat. Commun., 2021, 12, 3238 CrossRef CAS PubMed.
- J. Li, P. Wu, Z. Cao, G. Huang, Z. Lu, J. Yan, H. Zhang, Y. Zhou, R. Liu, H. Chen, L. Ma and M. Luo, Cell Rep., 2024, 43, 113765 CrossRef CAS PubMed.
- D. Wang, C. Zhang, B. Wang, B. Li, Q. Wang, D. Liu, H. Wang, Y. Zhou, L. Shi, F. Lan and Y. Wang, Nat. Commun., 2019, 10, 4284 CrossRef PubMed.
- Y. Yu, S. Gawlitt, L. B. De Andrade, E. Sousa, E. Merdivan, M. Piraud, C. L. Beisel and L. Barquist, Genome Biol., 2024, 25, 13 CrossRef PubMed.
- J. Wei, P. Lotfy, K. Faizi, S. Baungaard, E. Gibson, E. Wang, H. Slabodkin, E. Kinnaman, S. Chandrasekaran, H. Kitano, M. G. Durrant, C. V. Duffy, A. Pawluk, P. D. Hsu and S. Konermann, Cell Syst., 2023, 14, 1087–1102.e13 CrossRef CAS PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O’Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- J. Chen, M. Chen and T. F. Zhu, Nat. Biotechnol., 2022, 40, 1601–1609 CrossRef CAS PubMed.
- Y. Gao, L. Wang and B. Wang, Nat. Commun., 2023, 14, 8415 CrossRef CAS PubMed.
- S. D. Castle, M. Stock and T. E. Gorochowski, Nat. Commun., 2024, 15, 3640 CrossRef CAS PubMed.
- A. M. Yoshikawa, A. E. Rangel, L. Zheng, L. Wan, L. A. Hein, A. A. Hariri, M. Eisenstein and H. T. Soh, Nat. Commun., 2023, 14, 2336 CrossRef CAS PubMed.
- J. Song, Y. Zheng, M. Huang, L. Wu, W. Wang, Z. Zhu, Y. Song and C. Yang, Anal. Chem., 2020, 92, 3307–3314 CrossRef CAS PubMed.
- N. M. Angenent-Mari, A. S. Garruss, L. R. Soenksen, G. Church and J. J. Collins, Nat. Commun., 2020, 11, 5057 CrossRef CAS PubMed.
- R. Rotrattanadumrong and Y. Yokobayashi, Nat. Commun., 2022, 13, 4847 CrossRef CAS PubMed.
- J. S. Gootenberg, O. O. Abudayyeh, M. J. Kellner, J. Joung, J. J. Collins and F. Zhang, Science, 2018, 360, 439–444 CrossRef CAS PubMed.
- K. Pardee, A. A. Green, M. K. Takahashi, D. Braff, G. Lambert, J. W. Lee, T. Ferrante, D. Ma, N. Donghia, M. Fan, N. M. Daringer, I. Bosch, D. M. Dudley, D. H. O’Connor, L. Gehrke and J. J. Collins, Cell, 2016, 165, 1255–1266 CrossRef CAS PubMed.
- J. A. Kulkarni, D. Witzigmann, S. B. Thomson, S. Chen, B. R. Leavitt, P. R. Cullis and R. van der Meel, Nat. Nanotechnol., 2021, 16, 630–643 CrossRef CAS PubMed.
- M. Popova, O. Isayev and A. Tropsha, Sci. Adv., 2018, 4, eaap7885 CrossRef CAS PubMed.
- X. Zheng, R. Xie, X. Yao, Y. Su, L. Chu, P. Xu and W. Liu, Sci. Rep., 2024, 14, 32071 CrossRef PubMed.
- C. V. Theodoris, L. Xiao, A. Chopra, M. D. Chaffin, Z. R. Al Sayed, M. C. Hill, H. Mantineo, E. M. Brydon, Z. Zeng, X. S. Liu and P. T. Ellinor, Nature, 2023, 618, 616–624 CrossRef CAS PubMed.
- J. N. Acosta, G. J. Falcone, P. Rajpurkar and E. J. Topol, Nat. Med., 2022, 28, 1773–1784 CrossRef CAS PubMed.
- K. K. Narayanasamy, J. V. Rahm, S. Tourani and M. Heilemann, Nat. Commun., 2022, 13, 5047 CrossRef CAS PubMed.
- R. R. Wick, L. M. Judd and K. E. Holt, Genome Biol., 2019, 20, 129 CrossRef PubMed.
- Z. Wang, Y. Fang, Z. Liu, N. Hao, H. H. Zhang, X. Sun, J. Que and H. Ding, Nat. Commun., 2024, 15, 7148 CrossRef CAS PubMed.
- L. J. Fahrner, E. Chen, E. Topol and P. Rajpurkar, Cell, 2025, 188, 3648–3660 CrossRef CAS PubMed.
- M. K. Jena and B. Pathak, Nano Lett., 2023, 23, 2511–2521 CrossRef CAS PubMed.
- S. Pandit, M. K. Jena, S. Mittal and B. Pathak, ACS Appl. Nano Mater., 2024, 7, 17120–17132 CrossRef CAS.
- M. K. Jena, D. Roy, S. Mittal and B. Pathak, ACS Mater. Lett., 2023, 5, 2488–2498 CrossRef CAS.
- S. Mittal, S. Manna, M. K. Jena and B. Pathak, ACS Mater. Lett., 2023, 5, 1570–1580 CrossRef CAS.
- M. K. Jena, S. Mittal and B. Pathak, ACS Appl. Mater. Interfaces, 2024, 16, 29891–29901 CrossRef CAS PubMed.
- J. Im, S. Sen, S. Lindsay and P. Zhang, ACS Nano, 2018, 12, 7067–7075 CrossRef CAS PubMed.
- Q. Liu, L. Fang, G. Yu, D. Wang, C.-L. Xiao and K. Wang, Nat. Commun., 2019, 10, 2449 CrossRef PubMed.
- J. Wen, M. Han, N. Feng, G. Chen, F. Jiang, J. Lin and Y. Chen, Chem. Eng. J., 2024, 482, 148845 CrossRef CAS.
- Z. Zhao, R. Wang, X. Yang, J. Jia, Q. Zhang, S. Ye, S. Man and L. Ma, ACS Nano, 2024, 18, 33505–33519 CrossRef CAS PubMed.
- J. Lee, T. Lee, H. N. Lee, H. Kim, Y. K. Kang, S. Ryu and H. J. Chung, ACS Appl. Mater. Interfaces, 2023, 15, 54335–54345 CrossRef CAS PubMed.
- N. Nandu, C. W. Smith, T. B. Uyar, Y.-S. Chen, M. J. Kachwala, M. He and M. V. Yigit, ACS Appl. Nano Mater., 2020, 3, 11709–11714 CrossRef CAS PubMed.
- S. R. J. Hofstraat, T. Anbergen, R. Zwolsman, J. Deckers, Y. van Elsas, M. M. Trines, I. Versteeg, D. Hoorn, G. W. B. Ros, B. M. Bartelet, M. M. A. Hendrikx, Y. B. Darwish, T. Kleuskens, F. Borges, R. J. F. Maas, L. M. Verhalle, W. Tielemans, P. Vader, O. G. de Jong, T. Tabaglio, D. K. B. Wee, A. J. P. Teunissen, E. Brechbühl, H. M. Janssen, P. M. Fransen, A. de Dreu, D. P. Schrijver, B. Priem, Y. C. Toner, T. J. Beldman, M. G. Netea, W. J. M. Mulder, E. Kluza and R. van der Meel, Nat. Nanotechnol., 2025, 20, 532–542 CrossRef CAS PubMed.
- L. D. Nguyen, Z. Wei, M. C. Silva, S. Barberán-Soler, J. Zhang, R. Rabinovsky, C. R. Muratore, J. M. S. Stricker, C. Hortman, T. L. Young-Pearse, S. J. Haggarty and A. M. Krichevsky, Nat. Commun., 2023, 14, 7575 CrossRef CAS PubMed.
- K. Paunovska, D. Loughrey and J. E. Dahlman, Nat. Rev. Genet., 2022, 23, 265–280 CrossRef CAS PubMed.
- S. R. Krishnan, A. Roy, L. Wong and M. M. Gromiha, Nucleic Acids Res., 2025, 53, gkaf239 CrossRef CAS PubMed.
- M. Chandler, S. Jain, J. Halman, E. Hong, M. A. Dobrovolskaia, A. V. Zakharov and K. A. Afonin, Small, 2022, 18, 2204941 CrossRef CAS PubMed.
- Y. Su, L. Chu, W. Lin, X. Yao, P. Xu and W. Liu, Small Methods, 2025, 9, 2400959 CrossRef CAS PubMed.
- C. Imburgia, L. Organick, K. Zhang, N. Cardozo, J. McBride, C. Bee, D. Wilde, G. Roote, S. Jorgensen, D. Ward, C. Anderson, K. Strauss, L. Ceze and J. Nivala, Nat. Commun., 2025, 16, 6388 CrossRef CAS PubMed.
- D. Bar-Lev, I. Orr, O. Sabary, T. Etzion and E. Yaakobi, arXiv, 2021, preprint, arXiv:2109.00031 DOI:10.48550/arXiv.2109.00031.
- L. Wang, N. Li, M. Cao, Y. Zhu, X. Xiong, L. Li, T. Zhu and H. Pei, Adv. Sci., 2024, 11, 2409880 CrossRef CAS PubMed.
- R. Hou, C. Xie, Y. Gui, G. Li and X. Li, ACS Omega, 2023, 8, 19057–19071 CrossRef CAS PubMed.
- A. Rajkomar, M. Hardt, M. D. Howell, G. Corrado and M. H. Chin, Ann. Intern. Med., 2018, 169, 866–872 CrossRef PubMed.
- J. A. Doudna, Nature, 2020, 578, 229–236 CrossRef CAS PubMed.
- K. M. Esvelt, A. L. Smidler, F. Catteruccia and G. M. Church, eLife, 2014, 3, e03401 CrossRef PubMed.
- M. D. Wilkinson, M. Dumontier, Ij. J. Aalbersberg, G. Appleton, M. Axton, A. Baak, N. Blomberg, J.-W. Boiten, L. B. da Silva Santos, P. E. Bourne, J. Bouwman, A. J. Brookes, T. Clark, M. Crosas, I. Dillo, O. Dumon, S. Edmunds, C. T. Evelo, R. Finkers, A. Gonzalez-Beltran, A. J. G. Gray, P. Groth, C. Goble, J. S. Grethe, J. Heringa, P. A. C.’t Hoen, R. Hooft, T. Kuhn, R. Kok, J. Kok, S. J. Lusher, M. E. Martone, A. Mons, A. L. Packer, B. Persson, P. Rocca-Serra, M. Roos, R. van Schaik, S.-A. Sansone, E. Schultes, T. Sengstag, T. Slater, G. Strawn, M. A. Swertz, M. Thompson, J. van der Lei, E. van Mulligen, J. Velterop, A. Waagmeester, P. Wittenburg, K. Wolstencroft, J. Zhao and B. Mons, Sci. Data, 2016, 3, 160018 CrossRef PubMed.
- A. M. Childs, D. Maslov, Y. Nam, N. J. Ross and Y. Su, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, 9456–9461 CrossRef CAS PubMed.
- F. Arute, K. Arya, R. Babbush, D. Bacon, J. C. Bardin, R. Barends, R. Biswas, S. Boixo, F. G. S. L. Brandao, D. A. Buell, B. Burkett, Y. Chen, Z. Chen, B. Chiaro, R. Collins, W. Courtney, A. Dunsworth, E. Farhi, B. Foxen, A. Fowler, C. Gidney, M. Giustina, R. Graff, K. Guerin, S. Habegger, M. P. Harrigan, M. J. Hartmann, A. Ho, M. Hoffmann, T. Huang, T. S. Humble, S. V. Isakov, E. Jeffrey, Z. Jiang, D. Kafri, K. Kechedzhi, J. Kelly, P. V. Klimov, S. Knysh, A. Korotkov, F. Kostritsa, D. Landhuis, M. Lindmark, E. Lucero, D. Lyakh, S. Mandrà, J. R. McClean, M. McEwen, A. Megrant, X. Mi, K. Michielsen, M. Mohseni, J. Mutus, O. Naaman, M. Neeley, C. Neill, M. Y. Niu, E. Ostby, A. Petukhov, J. C. Platt, C. Quintana, E. G. Rieffel, P. Roushan, N. C. Rubin, D. Sank, K. J. Satzinger, V. Smelyanskiy, K. J. Sung, M. D. Trevithick, A. Vainsencher, B. Villalonga, T. White, Z. J. Yao, P. Yeh, A. Zalcman, H. Neven and J. M. Martinis, Nature, 2019, 574, 505–510 CrossRef CAS PubMed.
- G. E. Karniadakis, I. G. Kevrekidis, L. Lu, P. Perdikaris, S. Wang and L. Yang, Nat. Rev. Phys., 2021, 3, 422–440 CrossRef.
- N. Rieke, J. Hancox, W. Li, F. Milletarì, H. R. Roth, S. Albarqouni, S. Bakas, M. N. Galtier, B. A. Landman, K. Maier-Hein, S. Ourselin, M. Sheller, R. M. Summers, A. Trask, D. Xu, M. Baust and M. J. Cardoso, npj Digital Med., 2020, 3, 119 CrossRef PubMed.
- E. M. Lee, N. A. Setterholm, M. Hajjar, B. Barpuzary and J. C. Chaput, Nucleic Acids Res., 2023, 51, 9542–9551 CrossRef CAS PubMed.
- S. Hoshika, N. A. Leal, M.-J. Kim, M.-S. Kim, N. B. Karalkar, H.-J. Kim, A. M. Bates, N. E. Watkins, H. A. SantaLucia, A. J. Meyer, S. DasGupta, J. A. Piccirilli, A. D. Ellington, J. SantaLucia, M. M. Georgiadis and S. A. Benner, Science, 2019, 363, 884–887 CrossRef CAS PubMed.
- V. B. Pinheiro, A. I. Taylor, C. Cozens, M. Abramov, M. Renders, S. Zhang, J. C. Chaput, J. Wengel, S.-Y. Peak-Chew, S. H. McLaughlin, P. Herdewijn and P. Holliger, Science, 2012, 336, 341–344 CrossRef CAS PubMed.
- N. J. Szymanski, B. Rendy, Y. Fei, R. E. Kumar, T. He, D. Milsted, M. J. McDermott, M. Gallant, E. D. Cubuk, A. Merchant, H. Kim, A. Jain, C. J. Bartel, K. Persson, Y. Zeng and G. Ceder, Nature, 2023, 1–6 CAS.
- Y. Jia, R. Frydrych, Y. I. Sobolev, W.-S. Wong, B. Prajapati, D. Matuszczyk, Y. Bilgi, L. Gadina, J. C. Ahumada, G. Moldagulov, N. Kim, E. S. Larsen, M. Deschamps, Y. Jiang and B. A. Grzybowski, Nature, 2025, 645, 922–931 CrossRef CAS PubMed.
- A. Jobin, M. Ienca and E. Vayena, Nat. Mach. Intell., 2019, 1, 389–399 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |