Open Access Article
Open Access ArticleControlling the pH of aqueous succinic and maleic acids analyzed by X-ray absorption spectroscopy
Risa
Okada
a,
Rikuya
Adachi
a,
Ryosuke
Yamamura
ab,
Taiga
Suenaga
ab,
Takashi
Tokushima†
 b,
Yuka
Horikawa
b,
Yuka
Horikawa
 bc,
Masaki
Oura
bc,
Masaki
Oura
 b and
Osamu
Takahashi
b and
Osamu
Takahashi
 *bd
*bd
aChemistry Program, Graduate School of Advanced Science and Engineering, Hiroshima University, 1-3-1, Kagamiyama, Higashi-Hiroshima, Hiroshima 739-8526, Japan
bRIKEN SPring-8 Center, Sayo-Cho, Sayo, Hyogo 679-5148, Japan
cGraduate school of Science and Technology for Innovation, Yamaguchi University, Yamaguchi 753-8512, Japan
dResearch Institute for Synchrotron Radiation Science, Hiroshima University, 2-313, Kagamiyama, Higashi-Hiroshima, Hiroshima 739-0046, Japan. E-mail: shu@hiroshima-u.ac.jp
First published on 25th March 2026
Abstract
X-ray absorption spectroscopy (XAS) measurements were performed at the oxygen K-edge using aqueous solutions of succinic and maleic acids. Molecular dynamics simulations were employed to perform structural sampling and calculate the theoretical spectra using the framework of the density functional theory. When the pH was varied, the first peak in the XAS profiles exhibited different behaviors for succinic and maleic acids. This peak behavior was reproduced using theoretical calculations. Maleic acid, which has a double bond between its central carbon atoms, exhibits resonance effects between its two carboxyl groups. Because this resonance effect was not observed for succinic acid, it was considered to account for the different peak shifts. Hydrogen bonding analyses of the water molecules surrounding succinic and maleic acids were performed to elucidate the structure of the water network.
Introduction
Since the beginning of the 21st century, structural research based on the molecular theory of liquids under ambient conditions has progressed considerably.1 The development of technologies such as liquid jets2,3 and liquid flow cells4–7 has made it possible to quantity the X-ray absorption spectroscopy (XAS) profiles of liquid samples. To date, XAS studies have been conducted on liquid samples such as water,2,8 alcohols,9 and aqueous solutions of organic acids.10A typical example of a study in aqueous solutions using soft X-rays is acetic acid. This molecule has two oxygen atoms in the carboxyl group, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and OH, which have different chemical environments and are observed in a site-selective manner.10 In addition, Horikawa et al.11,12 quantified the XAS and X-ray emission spectroscopy (XES) profiles of aqueous acetic acid solutions at various pH levels and reported their pH dependence. The molar fractions of the neutral and anionic species quantitatively analyzed from the XES profiles were consistent with the Henderson–Hasselbalch equation. For oxalic acid, which is a dicarboxylic acid, both XAS and site-selective XES of its aqueous solution were conducted, and the respective profiles were quantified. By combining experimental findings with theoretical calculations, a resonance effect between the two carboxyl groups, which was not observed in monocarboxylic acids, was demonstrated.13 The pH control of aqueous solutions of organic molecules has also been studied for amino acids; for example, Alías-Rodríguez et al.14 confirmed that three protonation states dependent on the pH could be distinguished for proline. Using XPS and XAS, Greenspoon et al.15 also showed that changes in the pH affect the surface composition of ammonium sulfate aerosols. Horikawa et al.16 applied XAS to aqueous glycine as a biomolecular prototype.
O and OH, which have different chemical environments and are observed in a site-selective manner.10 In addition, Horikawa et al.11,12 quantified the XAS and X-ray emission spectroscopy (XES) profiles of aqueous acetic acid solutions at various pH levels and reported their pH dependence. The molar fractions of the neutral and anionic species quantitatively analyzed from the XES profiles were consistent with the Henderson–Hasselbalch equation. For oxalic acid, which is a dicarboxylic acid, both XAS and site-selective XES of its aqueous solution were conducted, and the respective profiles were quantified. By combining experimental findings with theoretical calculations, a resonance effect between the two carboxyl groups, which was not observed in monocarboxylic acids, was demonstrated.13 The pH control of aqueous solutions of organic molecules has also been studied for amino acids; for example, Alías-Rodríguez et al.14 confirmed that three protonation states dependent on the pH could be distinguished for proline. Using XPS and XAS, Greenspoon et al.15 also showed that changes in the pH affect the surface composition of ammonium sulfate aerosols. Horikawa et al.16 applied XAS to aqueous glycine as a biomolecular prototype.
Theoretical calculations based on the electronic structure theory are also powerful tools for studying solutions. Reinholdt et al.17 investigated the coupled cluster modeling of nitrogen K-edge XAS for aqueous ammonia and ammonium based on quantum mechanics/molecular dynamics (MD) and compared their results with published experimental data and transition potential density functional theory (TP-DFT)-based simulations. Odelius et al.18 conducted a theoretical and experimental study on the same system and showed symmetry breaking around the ammonia molecules in an aqueous solution. In addition, they simulated spectra using both ΔSCF (where orbitals were optimized for the final state) and second-order algebraic diagrammatic construction [ADC(2)] (which uses molecular orbitals optimized for the ground state), and interpreted the observed solvatochromism by comparing the XAS profiles in the gas phase and in aqueous solutions.19
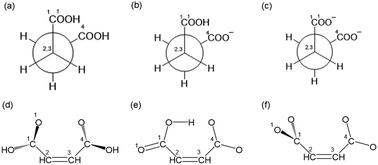
In this study, we focused on succinic and maleic acids as dicarboxylic acids. Dicarboxylic acids with two carboxyl groups play important roles in the metabolic pathways of living organisms. Succinic acid is among the compounds that makes up the citric acid cycle and is oxidized by succinate dehydrogenase to form fumarate. Maleic acid has the same empirical formula as fumaric acid, but refers to the cis-isomer. Maleic acid (pKa = 1.92, 6.23) and succinic acid (pKa = 4.21, 5.64)20 are four-carbon dicarboxylic acids. Maleic acid contains a carbon–carbon double bond, whereas succinic acid does not. Maleic acid has several conformational isomers owing to the rotation of its carboxyl groups and the orientation of the hydrogen atoms on its OH groups, whereas succinic acid has additional conformational isomers owing to the rotation around the C2–C3 axis. It would be interesting to observe the spectral features upon changing the pH and bond order. Furthermore, owing to intra- and intermolecular hydrogen bonding, succinic and maleic acids adopt various conformations. Theoretical analysis is also important to elucidate the features of these compounds at the molecular level.
Previous studies using IR spectroscopy21,22 and NMR23,24 have provided insights into the solvation structures and hydrogen bonding environments of these molecules. Conformational studies of dicarboxylic acids have been performed using IR and Raman spectroscopy by Suzuki and coworkers25 and theoretically.26,27 However, direct experimental information on how differences in hydrogen bonding motifs are reflected in the electronic states of oxygen atoms remains limited. In the present study, we employ soft XAS at the oxygen K-edge to investigate aqueous solutions of succinic and maleic acids, with the aim of elucidating how intra- and intermolecular hydrogen bonding influence their electronic structures.
Methods
Experimental methods
Aqueous 0.3 M succinic acid solutions at pH levels of 2.4, 5.0, and 12.5 was prepared by mixing two reagents [namely succinic acid (99.5% purity) and sodium hydroxide (97% purity), provided by Nacalai Tesque] with water. A 0.2 M maleic acid solution at pH levels of 1.4, 4.2, and 11.7 were prepared by mixing two reagents provided by Nacalai Tesque, maleic anhydride (99% purity) and sodium hydroxide (97% purity), with water.XAS profiles were recorded using the soft X-ray beamline BL17SU at the SPring-8 facility.28,29 All spectra were quantified using a liquid flow cell with a 150 nm-thick Au-coated thin Si3N4 window to separate the liquid flowing at atmospheric pressure from the high-vacuum region.5,30 The energy resolution of the XAS profiles was 0.1 eV. The background consisted of oxygen impurities in the Si3N4 window.
Theoretical methods
Maleic acid has several conformational isomers owing to the rotation of the carboxyl group and the orientation of the H atom of the OH group. The chemical structures of succinic and maleic acids are shown in Scheme 1. Succinic acid and its rotation around the C2–C3 axis should be considered. Conformational searches for succinic and maleic acids were performed using the CONFLEX31 program for neutral, anionic, and dianionic acids. Each conformation was structurally optimized using the molecular orbital program package Gaussian 0932 based on the Møller–Plesset second-order perturbation theory and cc-pVTZ basis sets.33 The polarizable continuum model (PCM)34–36 was also used to account for solvent effects.MD simulations were performed using the simulation program GROMACS.37 The aqueous solution model of maleic acid was created by randomly placing nine maleic acid molecules and 2500 water molecules in a box (0.2 M). The aqueous solution model of succinic acid was prepared by randomly adding 14 molecules of succinate and 2500 molecules of water to a box (0.3 M). Sodium ions were added to the anionic and dianionic acid solutions to achieve a total charge of zero. The water model used was the TIP4P force field.38
The bond and angular force constants of succinic and maleic acids were determined for the most stable succinic and maleic acid conformations. The lists of force constants are compiled in the SI.
We first performed energy minimization, followed by 20 ps simulations in the NVT ensemble (i.e., the number of particles N, volume V, and temperature T of the system were kept constant). Short-range interactions were corrected using the Verlet algorithm39 with a cutoff distance of 0.9 nm. Velocity–rescaled temperature coupling was used.40 Long-range electrostatics were corrected using the particle mesh Ewald (PME) algorithm41,42 during equilibration. The subsequent simulations were followed by 10 ns simulations in the NPT ensemble (i.e., the number of particles N, pressure P, and temperature T of the system were kept constant) with a time step of 0.1 fs, pressure of 1 bar, and a temperature of 300 K.
From several snapshots of the MD simulations, 90 clusters consisting of one succinic and one maleic acid molecule and surrounding 30 water molecules were sampled. XAS calculations were performed using the density functional theory implemented with the deMon2k code.43 The standard Perdew–Burke–Ernzerhof gradient-corrected exchange–correlation functional44,45 was used. In addition, the IGRO-III basis set46 was used to describe core-excited oxygen atoms. Triple-zeta valence plus polarization basis sets were used for hydrogen atoms. The effective core potential was used for the carbon and non-core-excited oxygen atoms. The obtained XAS line spectra were convolved using a Gaussian function (full width at half maximum, FWHM: 1.2 eV at 534 eV and 5.0 eV at 547 eV for maleic acid, 1.0 eV at 535 eV and 5.0 eV at 547 eV for succinic acid) and averaged.
Results and discussion
XAS profiles of succinic and maleic acids
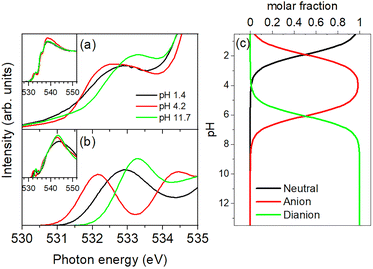
Fig. 1 shows the XAS profile of succinic acid. The insets show a broad energy range (528–548 eV). The first peaks assigned as O(1s) to π*(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) of succinic acid are observed at 532.4, 532.6, and 532.6 eV corresponding to pH levels of 2.4, 5.0, and 12.5. Note that there is no water absorption in this energy region, as also previously observed for acetic acid10 and oxalic acid.13 The positions and intensities of the first peaks increase with increasing pH. The theoretical X-ray absorption spectra are shown in Fig. 1(b). The theoretical spectra reproduce the experimental trend well; the first peaks are at 532.3, 532.3, and 533.4 eV for the neutral, anionic, and dianionic forms, respectively.
O) of succinic acid are observed at 532.4, 532.6, and 532.6 eV corresponding to pH levels of 2.4, 5.0, and 12.5. Note that there is no water absorption in this energy region, as also previously observed for acetic acid10 and oxalic acid.13 The positions and intensities of the first peaks increase with increasing pH. The theoretical X-ray absorption spectra are shown in Fig. 1(b). The theoretical spectra reproduce the experimental trend well; the first peaks are at 532.3, 532.3, and 533.4 eV for the neutral, anionic, and dianionic forms, respectively.
Fig. 2 shows XAS profile of maleic acid. The insets show a broad energy range (528–548 eV). The assignment of the first peak is the same as that of succinic acid. In contrast to succinic acid, the order of the first peaks for different pH levels is different, i.e., the peaks are at 532.8, 532.6, and 533.3 eV corresponding to pH levels of 1.4, 4.2, and 11.7. The neutral, anionic, and dianionic forms, respectively, are theoretically reproduced at 532.9, 532.1, and 533.3 eV.
Conformation analysis of succinic and maleic acids
To elucidate the spectral features of succinic and maleic acids, conformational analyses were performed. Succinic and maleic acids have multiple conformations. To systematically explore all the conformers, a conformational search for a single molecule was performed using CONFLEX. Eight neutral maleic acids, six anionic maleic acids, three dianionic maleic acids, 14 neutral succinic acids, 11 anionic succinic acids, and three dianionic succinic acids were obtained. The complete results are presented in the SI. Geometry optimization was performed for the isolated molecule using the PCM. The most stable conformations are shown in Fig. 3 and are the same as those for the isolated molecule and PCM, except for dianionic succinic acid. For neutral and anionic maleic acids, the most stable conformer was planar owing to intramolecular hydrogen bonding. In dianionic maleic acid, the most stable conformation is nonplanar owing to the repulsion between the negative charges of the oxygen atoms, and the two carboxylic groups are orthogonal to each other. Succinic acid has both gauche and anti-conformations; however, in the PCM, the gauche conformation is the most stable.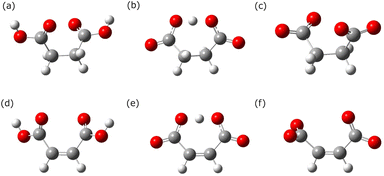 | ||
| Fig. 3 Most stable molecular structures of succinic and maleic acids. The following text was used for atom labeling. (a) and (d) Neutral, (b), (e) anionic, and (c) and (f) dianionic forms. | ||
Fig. 3 shows the conformations of the succinic and maleic acids used in the MD simulations. The most stable conformational isomers of dianionic succinic acid differ between a single molecular model and PCM, with the linear (unfolded) conformation being more stable in the single molecular model and the folded conformation being more stable in the PCM. Wanko et al.47 showed that the dianion of succinic acid adopts a folded conformation in aqueous solutions. Neutral maleic acid was the most stable in the gas phase and the PCM, with intramolecular hydrogen-bonded cyclic structures. However, Hyttinen et al.48 suggested that neutral maleic acid in water forms hydrogen bonds with the surrounding water rather than intramolecular hydrogen bonds. The single molecule and PCM calculations do not fully account for interactions with the surrounding water molecules. Therefore, for neutral maleic acid, we adopted a conformational isomer without intramolecular hydrogen bonds.
In the XAS profile of oxalic acid, the resonance effect between the two COOH functional groups was observed as a high-energy shift in the peak.13 This resonance effect was observed when the two COOH groups of oxalic acid were parallel; when the COOH groups were twisted, the π resonance was destroyed, and this did not occur. To elucidate this effect explicitly, the XAS profile of a single molecule was obtained by rotating the carboxyl group of neutral maleic acid by 10° with respect to the molecular plane. The spectral changes observed in this case are shown in Fig. 4(a). The angle shown in the figure corresponds to the C3–C2–C1–O1 dihedral angle. The first peak, which is assigned as O(1s) → π*(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), blue-shifts, and its intensity increases as the dihedral angle increases. The peak position is at 531.1 eV when the dihedral angle is 0°, but it is at 531.9 eV at 90°. The π* orbitals of the excited destination of neutral maleic acid when the dihedral angles of the carboxylic groups are ϕ = 0°, 30°, 60°, and 90° are shown in Fig. 4(b). When the dihedral angle is 0°, all atoms are on the same plane, and π resonance between the two carboxylic groups is observed. As the dihedral angle increases, the π resonance begins to break, and completely disappears at ϕ = 90°. Yamamura et al.13 showed that it is possible to adjust the π resonance of oxalic acid. The same mechanism was observed in the case of maleic acid.
O), blue-shifts, and its intensity increases as the dihedral angle increases. The peak position is at 531.1 eV when the dihedral angle is 0°, but it is at 531.9 eV at 90°. The π* orbitals of the excited destination of neutral maleic acid when the dihedral angles of the carboxylic groups are ϕ = 0°, 30°, 60°, and 90° are shown in Fig. 4(b). When the dihedral angle is 0°, all atoms are on the same plane, and π resonance between the two carboxylic groups is observed. As the dihedral angle increases, the π resonance begins to break, and completely disappears at ϕ = 90°. Yamamura et al.13 showed that it is possible to adjust the π resonance of oxalic acid. The same mechanism was observed in the case of maleic acid.
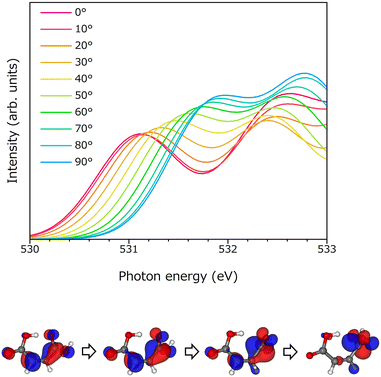 | ||
| Fig. 4 (upper) Dihedral angle C3–C2–C1–O1 dependence of XAS profiles for neutral maleic acid from 0° to 90°. (lower) HOMOs at 0, 30, 60, and 90°. | ||
The π* orbitals of succinic and maleic acids are shown in Fig. 5. Succinic acid has four C atoms that are not in the same plane, and its dihedral angle is approximately 60°. Therefore, π-orbital delocalization through the four C atoms does not occur. In contrast, maleic acid has a dihedral angle of 0° and consists of four C atoms owing to its double bond. In anionic maleic acid, the two carboxyl groups are parallel to a plane consisting of four C atoms, thus preserving resonance, whereas in neutral and dianionic maleic acids, the carboxyl groups are not parallel to the molecular plane and have a twisted structure. This twisting breaks the π resonance and blue-shifts the first peak in the XAS profiles. The peculiar peak behavior of the aqueous maleic acid solutions in the XAS profile was caused by the C atoms in the same plane and the different dihedral angles of the carboxylic groups for each valence.
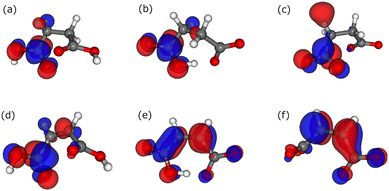 | ||
| Fig. 5 HOMOs for succinic acid in the (a) neutral, (b) anionic, and (c) dianionic forms, and HOMOs for maleic acid in the (d) neutral, (e) anionic, and (f) dianionic forms. | ||
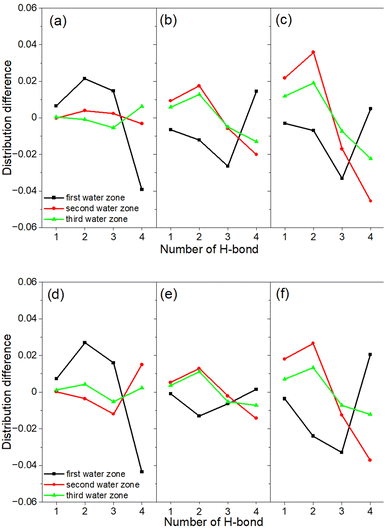
To understand the behavior at the interface between the organic substances and water, it is interesting to investigate the hydrogen bonding behavior of the water surrounding the organic substances. Based on the MD simulation results of the aqueous solutions of succinic and maleic acids, hydrogen bond analysis was conducted. The results are shown in Fig. 6.
The difference between the structures of succinic and maleic acids lies in the bond distance between the C2–C3 atoms and the number of hydrogen atoms attached to the carbons. These differences had only a minor effect on the hydrogen-bonded network of the surrounding water molecules, with the effect being more prominent for maleic acid. This was due to the difference in the C2–C3 bond length. In the first hydration shell, tetra-coordinated water decreased, and bi-coordinated and tri-coordinated water increased, indicating the disruption of the hydrogen-bond network of water caused by succinic and maleic acids. In the first and second hydration shells, which were closer to the solute, the structural differences were greater compared with those in liquid water, suggesting that the surrounding water structure was disturbed by the presence of these acids. In the third hydration shell, the effect of the solute decreased, and the number of hydrogen bonds became closer to that in liquid water. As the valence increased from neutral to monovalent to divalent, the difference from liquid water increased because the water molecules were more strongly attracted to the solute.
Conformations of succinic and maleic acids controlling pH
Dicarboxylic acids can exist in three different states depending on the pH adjustment. In the case of succinic acid, there is no resonance between the carboxyl groups at either end, and each functional group acts independently because the central C2–C3 bond can rotate freely. In such case, succinic acid behaves as if there were two acetic acid molecules. Horikawa et al.11,13 showed that, in acetic acid, as the pH increased, the first peak gradually increased. The behaviors of dicarboxylic acids with longer carbon chains, such as glutaric and adipic acids, can also be predicted.Although maleic acid has the same number of carbon atoms as succinic acid, the behavior of its peaks differs from that of succinic acid. The first peak of neutral maleic acid was larger and shifted to a higher energy compared with the others. This is because the two carboxyl groups cannot lie in the same plane owing to the electrostatic repulsion between the functional groups, even though resonance exists via the double bond between the two carboxyl groups at either end. When the pH is increased and maleic acid becomes a monovalent anion, intramolecular hydrogen bonding allows the two carboxyl groups to remain in the same plane. Therefore, the first peak shifted to a lower energy compared with that of the neutral case. When the pH is further increased and the carboxyl group becomes a divalent anion, the two carboxyl groups twist considerably, as in the case of oxalic acid, shifting the peak to a higher energy.
Summary
In this study, we investigated the structures of succinic and maleic acids in aqueous solutions using XAS. Although succinic and maleic acids have the same number of carbon atoms, their XAS behaviors differ in response to pH. This can be explained based on the electrostatic repulsion between the two carboxyl groups and the presence or absence of resonant structures mediated by double bonds. By combining XAS measurements with theoretical calculations, we were able to discuss the rich chemical information present in the aqueous solutions in detail.While NMR and IR provide averaged information, XAS/XES captures the local electronic state obtained by core-excitation from oxygen 1s electron site-selectively as an extremely fast process occurring on the femtosecond scale. Observing how the concept of resonance actually behaves in the structurally fluctuating environment of an aqueous solution is not straightforward, but soft X-ray spectroscopy has made it possible for the first time. The methodology used in this study can be applied to other organic compounds in aqueous solutions that undergo structural changes depending on the pH, and is expected to be useful for future research on the solution chemistry of aqueous systems.
Author contributions
OT devised the project and prepared the beamtime proposal. TT, YH, and MO designed the experimental setup. RY, TS, TT, and OT performed the experiments and analyzed the data under guidance of TT and YH. RY, RO, RA and OT performed theoretical analysis. RO and OT wrote the original draft. OT is a corresponding author.Conflicts of interest
There are no conflicts to declare.Data availability
The authors confirm that the data supporting the findings of this study are available within the article and/or its supplementary information (SI). Supplementary information: Tables S1–S20. See DOI: https://doi.org/10.1039/d6cp00520a.Acknowledgements
Synchrotron radiation experiments were performed at BL17SU at Spring-8 with the approval of the Japan Synchrotron Radiation Institute (JASRI) (Project number 20160069). The authors thank the Research Center for Computational Science, Okazaki, Japan (Project: 23-IMS-C033, 24-IMS-C033) and the Research Institute for Information Technology at Kyushu University, Fukuoka, for providing computational resources. This work was supported by JSPS KAKENHI (Grant Number JP23K26499), the JSPS Program for Forming Japan's Peak Research Universities (J-PEAKS) (Grant Number JPJS00420230011), and the Salt Science Research Foundation (Grant Numbers 202413 and 202512). This work were performed at Research Institute for Synchrotron Radation Science, Hiroshima University, HiSOR, under Proposal No. 25BU016.Notes and references
- J. W. Smith and R. J. Saykally, Chem. Rev., 2017, 117, 13909–13934 CrossRef CAS PubMed
.
- K. R. Wilson, B. S. Rude, T. Catalano, R. D. Schaller, J. G. Tobin, D. T. Co and R. J. Saykally, J. Phys. Chem. B, 2001, 105, 3346–3349 CrossRef CAS
.
- M. Ekimova, W. Quevedo, M. Faubel, P. Wernet and E. T. J. Nibbering, Struct. Dyn., 2015, 2, 054301 CrossRef PubMed
.
- M. Freiwald, S. Cramm, W. Eberhardt and S. Eisebitt, J. Electron Spectrosc. Relat. Phenom., 2004, 137–140, 413–416 CrossRef CAS
.
- T. Tokushima, Y. Harada, H. Ohashi, Y. Senba and S. Shin, Rev. Sci. Instrum., 2006, 77, 063107 CrossRef
.
- M. Nagasaka, T. Hatsui, T. Horigome, Y. Hamamura and N. Kosugi, J. Electron Spectrosc. Relat. Phenom., 2010, 177, 130–134 CrossRef CAS
.
- Y. Harada, M. Kobayashi, H. Niwa, Y. Senba, H. Ohashi, T. Tokushima, Y. Horikawa, S. Shin and M. Oshima, Rev. Sci. Instrum., 2012, 83, 013116 CrossRef PubMed
.
- P. Wernet, D. Nordlund, U. Bergmann, M. Cavalleri, M. Odelius, H. Ogasawara, L. A. Naslund, T. K. Hirsch, L. Ojamae, P. Glatzel, L. G. M. Pettersson and A. Nilsson, Science, 2004, 304, 995–999 CrossRef CAS PubMed
.
- K. R. Wilson, M. Cavalleri, B. S. Rude, R. D. Schaller, T. Catalano, A. Nilsson, R. J. Saykally and L. G. M. Pettersson, J. Phys. Chem. B, 2005, 109, 10194–10203 CrossRef CAS PubMed
.
- T. Tokushima, Y. Horikawa, Y. Harada, O. Takahashi, A. Hiraya and S. Shin, Phys. Chem. Chem. Phys., 2009, 11, 1679–1682 RSC
.
- Y. Horikawa, T. Tokushima, Y. Harada, O. Takahashi, A. Chainani, Y. Senba, H. Ohashi, A. Hiraya and S. Shin, Phys. Chem. Chem. Phys., 2009, 11, 8676–8679 RSC
.
- Y. Horikawa, T. Tokushima, A. Hiraya and S. Shin, Phys. Chem. Chem. Phys., 2010, 12, 9165–9168 RSC
.
- R. Yamamura, T. Suenaga, M. Oura, T. Tokushima and O. Takahashi, Chem. Phys. Lett., 2020, 738, 136895 CrossRef CAS
.
- M. Alías-Rodríguez, S. Bonfrate, W. Park, N. Ferré, C. H. Choi and M. Huix-Rotllant, J. Phys. Chem. A, 2023, 127, 10382–10392 CrossRef PubMed
.
- E. H. Greenspoon, S. Warkander, P. Kim, A. Jana, J. Qian, K. R. Wilson, M. Ahmed and J. B. Bergner, J. Chem. Phys., 2025, 162, 234307 CrossRef CAS PubMed
.
- Y. Horikawa, T. Tokushima, O. Takahashi, Y. Harada, A. Hiraya and S. Shin, Phys. Chem. Chem. Phys., 2018, 20, 23214–23221 RSC
.
- P. Reinholdt, M. L. Vidal, J. Kongsted, M. Iannuzzi, S. Coriani and M. Odelius, J. Phys. Chem. Lett., 2021, 12, 8865–8871 CrossRef CAS PubMed
.
- M. Odelius, S. D. Folkestad, T. Saisopa, Y. Rattanachai, W. Sailuam, H. Yuzawa, N. Kosugi, A. C. Paul, H. Koch and D. Céolin, J. Phys. Chem. Lett., 2025, 16, 3411–3419 CrossRef CAS PubMed
.
- S. Tsuru and M. Nagasaka, J. Phys. Chem. A, 2025, 129, 3020–3031 CrossRef CAS PubMed
.
-
R. D. Lide, CRC Handbook of Chemistry and Physics, CRC, Taylor and Francis, Boca Raton, FL, 87th edn, 2007 Search PubMed
.
- M. a J. Ayora-Cañada and B. Lendl, Vib. Spectrosc., 2000, 24, 297–306 CrossRef
.
- J. T. Hołaj-Krzak, N. Rekik, N. A. M. Alsaif and G. Lakshminarayana, J. Phys. Chem. A, 2022, 126, 5604–5620 CrossRef PubMed
.
- J. D. Roberts, Acc. Chem. Res., 2006, 39, 889–896 CrossRef CAS PubMed
.
- J. Guo, P. M. Tolstoy, B. Koeppe, G. S. Denisov and H.-H. Limbach, J. Phys. Chem. A, 2011, 115, 9828–9836 CrossRef CAS PubMed
.
- M. Suzuki and T. Shimanouchi, J. Mol. Spectrosc., 1968, 28, 394–410 CrossRef CAS
.
- P. Tarakeshwar and S. Manogaran, THEOCHEM, 1996, 362, 77–99 CrossRef CAS
.
- T. H. Nguyen, D. E. Hibbs and S. T. Howard, J. Comput. Chem., 2005, 26, 1233–1241 CrossRef CAS PubMed
.
- H. Ohashi, Y. Senba, H. Kishimoto, T. Miura, E. Ishiguro, T. Takeuchi, M. Oura, K. Shirasawa, T. Tanaka, M. Takeuchi, K. Takeshita, S. Goto, S. Takahashi, H. Aoyagi, M. Sano, Y. Furukawa, T. Ohata, T. Matsushita, Y. Ishizawa, S. Taniguchi, Y. Asano, Y. Harada, T. Tokushima, K. Horiba, H. Kitamura, T. Ishikawa and S. Shin, AIP Conf. Proc., 2007, 879, 523–526 CrossRef CAS
.
- Y. Senba, H. Ohashi, H. Kishimoto, T. Miura, S. Goto, S. Shin, T. Shintake and T. Ishikawa, AIP Conf. Proc., 2007, 879, 718–721 CrossRef CAS
.
- T. Tokushima, Y. Harada, O. Takahashi, Y. Senba, H. Ohashi, L. G. M. Pettersson, A. Nilsson and S. Shin, Chem. Phys. Lett., 2008, 460, 387–400 CrossRef CAS
.
- H. Goto, S. Obata, N. Nakayama and K. Ohta, CONFLEX Corporation, Tokyo, Japan, 2012.
-
M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. J. A. Montgomery, J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, T. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian, Inc., Wallingford CT, 2010
.
- J. T. H. Dunning, J. Chem. Phys., 1989, 90, 1007–1023 CrossRef
.
- B. Mennucci, E. Cancès and J. Tomasi, J. Phys. Chem. B, 1997, 101, 10506–10517 CrossRef CAS
.
- M. Cossi, V. Barone, B. Mennucci and J. Tomasi, Chem. Phys. Lett., 1998, 286, 253–260 CrossRef CAS
.
- E. Cancès, B. Mennucci and J. Tomasi, J. Chem. Phys., 1997, 107, 3032–3041 CrossRef
.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef
.
- H. W. Horn, W. C. Swope, J. W. Pitera, J. D. Madura, T. J. Dick, G. L. Hura and T. Head-Gordon, J. Chem. Phys., 2004, 120, 9665–9678 CrossRef CAS PubMed
.
- L. Verlet, Phys. Rev., 1967, 159, 98–103 CrossRef CAS
.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 614101 CrossRef PubMed
.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS
.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, J. Chem. Phys., 1995, 103, 8577–8593 CrossRef CAS
.
-
A. M. Köster, G. Geudtner, A. Alvarez-Ibarra, P. Calaminici, M. E. Casida, J. Carmona-Espindola, V. D. Domínguez-Soria, R. Flores-Moreno, G. U. Gamboa, A. Goursot, T. Heine, A. Ipatov, A. de la Lande, F. Janetzko, J. M. del Campo, D. Mejia-Rodriguez, J. U. Reveles, J. Vasquez-Perez, A. Vela, B. Zúñinga-Gutierrez and D. R. Salahub, deMon2k, version 5, The deMon developers, 2018 Search PubMed
.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed
.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1997, 78, 1396 CrossRef CAS
.
-
W. Kutzelnigg, U. Fleischer and M. Schindler, NMR-Basic Principles and Progress, Springer-Verlag, Heidelberg, 1990 Search PubMed
.
- M. Wanko, T. Wende, M. M. Saralegui, L. Jiang, A. Rubio and K. R. Asmis, Phys. Chem. Chem. Phys., 2013, 15, 20463–20472 RSC
.
- N. Hyttinen and N. L. Prisle, J. Phys. Chem. A, 2020, 124, 4801–4812 CrossRef CAS PubMed
.
Footnote |
| † Current address: MAX IV Laboratory, Lund University, Fotongatan 2, 224 84 Lund, Sweden. |
| This journal is © the Owner Societies 2026 |