Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Synthetic fluorescent receptor arrays for sensing in complex matrices: from robust chemistry to data-driven readouts
Rossella
Santonocito
 ,
Alessia
Cavallaro
,
Alessia
Cavallaro
 ,
Nunzio
Tuccitto
,
Nunzio
Tuccitto
 and
Giuseppe
Trusso Sfrazzetto
and
Giuseppe
Trusso Sfrazzetto
 *
*
Department of Chemical Sciences, University of Catania, viale A. Doria 6, 95125 Catania, Italy. E-mail: giuseppe.trusso@unict.it
First published on 9th April 2026
Abstract
Sensing is moving from specific lock-and-key biorecognition toward chemically programmable, pattern-based approaches. Synthetic fluorescent receptor arrays, composed of rationally designed organic probes (e.g., rhodamines, BODIPYs, naphthalimides, and carbon dots) that engage analytes through non-covalent interactions, offer a robust and versatile platform for multidimensional detection. Rather than relying on a single high-affinity interaction, these arrays exploit highly selective responses across many probes. Multivariate analysis (PCA/PLS/PLS-DA) or modern machine learning converts the complex optical pattern into identity and concentration. This Feature Article highlights that chemical stability, operational versatility, and compatibility with unprocessed, real samples are the decisive advantages of synthetic arrays over classical biosensing. Synthetic receptors tolerate wider pH/ionic strength/solvent windows, resist denaturation, support long shelf-life, and can be mass-produced with consistent performance. Their modularity enables coverage from gases/VOCs to small metabolites and biomarkers, while their signal transduction (fluorescence/colorimetry) integrates seamlessly with smartphone cameras (fieldable readout), optical fibers (real-time, in situ monitoring), and hyperspectral imaging (spatially resolved analytics). These attributes open opportunities in food safety, environmental surveillance, and point-of-care diagnostics where pretreatment is impractical. In addition, pre-analytical handling (centrifugation, filtration, and storage) reshapes sample composition and erodes both qualitative identities and quantitative readouts. This Feature Article emphasizes that analyzing as-collected specimens preserves the native chemical fingerprint that cross-reactive arrays exploit. We outline practical designs and drift-aware workflows that enable minimal handling while maintaining selectivity, sensitivity, and wide linear ranges in complex matrices. A review of selected recent case studies applied to human health, environmental, and agri-food fields demonstrates the approach: ppb or pM limits of detection, minute-scale analyses, and multi-decade linear ranges (up to nine orders) with reliable performance in untreated matrices such as saliva, fruit headspace, environmental waters, and simulated synovial fluid.
1. Introduction
1.1 What is an array?
A chemical-based optical array is a cross-reactive sensing platform that replaces specific single lock-and-key receptors with a panel of non-specific, chemically responsive dyes/fluorophores. These devices are also called optoelectronic “noses/tongues”, depending on the possibility to detect gas (nose) or liquid (tongue), respectively.1 Each probe exploits different classes of intermolecular interactions (e.g., Brønsted/Lewis acid–base, redox, hydrogen bonding, solvatochromism, and non-covalent interactions) or covalent reactions,2 so exposure to an analyte or a mixture generates a composite optical pattern—a high-dimensional fingerprint—rather than a single scalar signal. This olfaction-inspired strategy takes inspiration from mechanisms that have long been optimized by nature: discrimination arises from pattern recognition across many receptors rather than from one-to-one binding.3 Practically, disposable colorimetric/fluorometric arrays are printed on porous, hydrophobic matrices and read by digital imaging or fiber-optic bundles; chemometric methods, such as hierarchical cluster analysis (HCA), linear discriminant analysis (LDA), and support vector machines (SVM), are then used to classify unknowns and enable qualitative identification and quantitative calibration.4 By sampling broad “chemical property space,” optical arrays achieve very high sensitivity and robust discrimination among closely related analytes and reliable analysis of complex mixtures in gases and liquids.Single-receptor sensing relies on a lock-and-key paradigm: one analyte engages one selective probe via a specific reaction5 or non-covalent interaction,2 yielding a single scalar signal. This approach works when high affinity and exclusivity are chemically achievable, but it struggles with near-neighbors, isobars, and changing matrices; any off-target interaction or matrix effect directly corrupts the readout. In contrast, optical arrays are deliberately cross-reactive collections of dyes/fluorophores that span diverse interactions. Exposure to a sample produces a multiplexed pattern of intensity/color changes—a complex but characteristic fingerprint—whose information content is extracted by chemometrics (e.g., HCA, LDA, SVM, and PLS) for both classification and calibration. A recent comprehensive review summarizes combinational and activatable systems based on the sensor-array and lock-and-key mechanisms.6 Selectivity is thus emergent: it derives from the orthogonality of partial responses across many sensors rather than from the perfect specificity of any single element. The probability that two different analytes lead to the same fingerprint is very low: the higher the number of probes present on an array, the greater the efficiency in terms of selectivity. Furthermore, one of the main goals of the “array philosophy” is the possibility to detect multiple analytes within one single analysis. When properly trained, an array can detect the presence of more than one single target analyte in a mixture, paving the way for the multiple analyte detection and leading to a significant reduction in terms of costs and time.
Because the decision is made in pattern space, array methods naturally tolerate interferences and enable the analysis of complex mixtures (gases or liquids) that defeat one-probe assays. The same fingerprint logic underpins biological olfaction and gustation—discrimination arises from distributed, partially overlapping receptors—providing a conceptual and practical framework for detecting closely related analytes, resolving mixtures, and operating across wide dynamic ranges, using simple imaging or emission spectra readouts.
We highlight that arrays are not a universal replacement for highly engineered single-receptor formats; rather, they become advantageous under specific constraints (matrix complexity, multiplexing, uncertain interferents, and the need for robustness without extensive sample handling).
In recent years, several non-array platforms have progressed substantially. MOF-based sensors can deliver impressive sensitivity through high porosity, tuneable adsorption sites, and engineered host–guest interactions, enabling selective responses toward gases and VOCs across multiple transduction modes.7 Likewise, MIPs act as “synthetic antibodies” and can provide strong affinity/selectivity for a defined target while offering mechanical robustness and attractive shelf-life;8 however, MIPs often face challenges associated with binding-site heterogeneity, diffusion/kinetic limitations in thicker films, and matrix-dependent nonspecific adsorption that can degrade selectivity in complex real samples unless carefully engineered and validated. Structure-switching aptamers and next-generation aptamer biosensors can achieve excellent selectivity and enable signal transduction via conformational changes, and their modularity makes them highly attractive for portable formats and integrated electronics;9 nonetheless, in real biological matrices, aptamers can suffer from variable stability due to nuclease-mediated degradation and matrix fouling, often requiring chemical modifications and protective assay conditions to preserve performance.
Against this background, array-based sensing wins primarily when selectivity must be achieved in the presence of unknown or variable interferents, or when mixtures must be analysed without fractionation. Arrays intentionally use partially selective elements spanning diverse interaction chemistries. Selectivity is therefore emergent from the multivariate fingerprint rather than dependent on a single binding event. This makes arrays particularly competitive for untreated or minimally treated matrices (e.g., saliva, headspace, wash water, and air) where (i) relevant information is distributed across a complex mixture and (ii) pre-analytical handling risks losing volatile or labile components.
At the same time, the array approach has clear limitations that should be acknowledged. Arrays typically require important training datasets, careful control of drift (humidity/temperature, aging, illumination), and calibration transfer across devices. Accordingly, the most realistic view is that the array approach is best positioned as a complementary strategy: it provides robust, mixture-capable sensing when the sample is complex and the “answer” is a fingerprint, while advanced single-receptor platforms remain optimal when the analytical problem is narrowly defined, and the operating conditions can be tightly controlled.
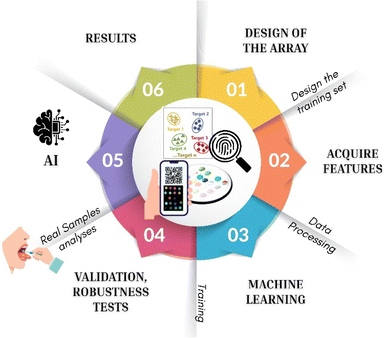
An optical array must be trained to recognize its target(s) before it can be used reliably. Training converts raw multichannel responses into a statistical model that maps patterns to identities (classification) and/or concentrations (regression). This training consists of different subsequent steps (Fig. 1):
(1) Design of the array: the probe panel is engineered backward from the following analytical questions: (i) who the targets are, (ii) in which matrix, (iii) at what concentrations, (iv) and against which interferents. Probes are selected jointly to maximize response orthogonality so that each analyte perturbs the ensemble with a distinct fingerprint. This balance centers on photophysics and recognition chemistry. On the photophysical side, the panel should cover near-UV to red/NIR chromophores, depending on the detection method, with high quantum yields, low self-absorption, and limited crosstalk, complemented by ratiometric pairs to stabilize readout. On the recognition side, expected analyte properties (acid/base character, polarity, hydrogen-bonding capacity, π-systems, redox behaviour, and metal affinity) are mapped to complementary supramolecular motifs.10 Reversible, moderately strong interactions are preferred to preserve the dynamic range and enable mixture deconvolution; covalent or irreversible chemistries are reserved for specific anchors only when necessary. Manufacturability and robustness are built in from the outset. Probes are vetted for stability within the immobilization matrix and over shelf-life, including resistance to leaching, photobleaching, and variations in pH, ionic strength, and organic content. Redundancy is introduced—multiple partially correlated probes per interaction class—so performance degrades gracefully if individual spots drift. Spatial layout is planned for uniform illumination, and internal controls (inert colour patches or intensity standards) are embedded to monitor optical drift and lot-to-lot variation. The array size is an explicit design parameter and, importantly, it can be determined empirically rather than chosen a priori. After an initial screening with an “overspecified” panel, the response matrix allows the minimum number of sensing elements required for a given application to be estimated mathematically. In practice, probe contributions are ranked using multivariate metrics (e.g., VIP scores and regression coefficients in PLS/PLS-DA, feature importance in ensemble models, or orthogonality/variance contributions in PCA), and probes that are redundant, low-informative, or drift-dominated are removed. The array is then progressively reduced while monitoring cross-validated performance (accuracy, sensitivity/specificity, RMSE, LOD/LOQ, and linear range). The point at which performance begins to degrade defines the “minimal sufficient array” for that matrix and task, providing a quantitative link between array size and application requirements. When the target analyte is well defined, the matrix is stable, and quantification is the main goal, a smaller array (often a handful of spots) can be sufficient if the chosen probes provide complementary interaction chemistries and a stable calibration. Conversely, complex matrices, unknown interferents, mixture deconvolution, and multi-class identification generally demand larger panels, because selectivity emerges from the orthogonality of multiple partial responses and because redundancy improves fault tolerance to aging and environmental drift. In other words, the required number of sensing elements scales with the dimensionality of the analytical problem (the number of classes/analytes, expected confounders, and matrix variability), and the optimal array size can be objectively defined by data-driven down-selection rather than subjective design.
(2) Design the training set: the first step is the calibration, preparing standard(s) spanning the expected concentration range. Then, interferents must be taken into account, in order to create a representative matrix (real or matrix-matched) and include mixtures if mixture analysis is required. The run order should be randomized and technical/biological replicates should be acquired to estimate variance.
(3) Acquire features: From elaboration of the images (RGB/HSV, grey channel) or absorption/emission spectra (band intensities, integrals, and principal components). Record environmental covariates for drift compensation. To keep an array honest in the field, temperature (T), humidity (RH), and time must be considered. In fact, fluorescence and color can be affected by T/RH, support aging, and lamp warm-up.11 Ignoring these factors, the model quietly learns the environment alongside the chemistry and then fails when conditions change. The fix starts with rigorous metadata logging for every acquisition—temperature, relative humidity, timestamp, device ID, exposure/integration, and lamp state—so that the drift is measurable. With these records, simple, physics-plausible corrections (for example, subtracting linear terms in T and RH from raw intensities) and, more importantly, feed T/RH/time directly into the learning algorithm can be included. In multivariate models such as PLS, SVR, or LDA, the environmental covariates help separate the analytical signal from environmental bias.12 Internal reference spots or colour patches provide a frame-by-frame estimate of optical drift and enable on-the-fly recalibration. When hardware changes—new smartphone,13 fiber,14 or spectrometer—domain adaptation or calibration transfer methods align the new device to the old response space without requiring retraining from scratch. Daily quality control with a blank and a known standard sets the guardrail.
(4) Data processing: data processing begins with rigorous preprocessing to convert raw images or spectra into stable, comparable features. The background is removed to isolate probe signals; intensities are then normalized at both the spot and the card level to correct illumination and loading differences. Feature scaling brings heterogeneous channels to comparable variance, while batch correction suppresses inter-day and lot effects without erasing genuine analytical structure. Modelling proceeds in a staged fashion. Unsupervised analysis (e.g., PCA, t-SNE, and UMAP) is first used to visualize structure, detect outliers, and diagnose batch artifacts, guiding feature refinement. Supervised models are then trained according to the task. For presence/identity decisions, linear discriminants (LDA/QDA) provide transparent baselines; PLS-DA handles collinearity common in arrays; margin-based learners (SVM) and ensemble methods (random forests and gradient boosting) capture nonlinearity and higher-order interactions among probes. A rejection or out-of-distribution class is included so the system can abstain when patterns deviate from the trained domain. For quantification, regression models exploit the same feature space: PLS-R is robust with correlated predictors; ridge regression or elastic net regularize noisy channels; SVR captures nonlinear calibration curves without overfitting small datasets. Multi-analyte problems are addressed either with multi-output PLS-R, which models shared covariance among targets, or with one-vs.-rest decompositions that maintain interpretability and independent limits of detection. Throughout, hyperparameters are selected with nested cross-validation and stratification across cards and days to ensure that gains reflect chemical signals rather than batch idiosyncrasies, and final performance is reported with uncertainty estimates suitable for validation.
(5) Validation: validation must demonstrate that performance reflects chemistry rather than chance, batch quirks, or analyst bias. A nested cross-validation scheme separates hyperparameter tuning from generalization testing; folds are stratified across cards, days, operators, and matrices to prevent leakage. Limits of detection/quantification (LOD/LOQ) are estimated from blank or low-level replicates and the calibration slope (IUPAC or alternative error-in-variables methods), and the linear range is defined by lack-of-fit testing rather than eyeballing R2.
(6) Robustness tests: arrays must tolerate realistic perturbations without collapsing. Environmental challenges sweep temperature and humidity across the intended operating envelope, with illumination intensity and spectral content varied to probe optical sensitivity. Chemical robustness is assessed by spiking confounders at supra-physiological levels (salts, proteins, and VOC background) and by matrix swaps (artificial vs. real) to reveal hidden interactions. Mechanical robustness includes accelerated aging (elevated T/RH and light exposure) to model shelf-life and transport. Device variability is exercised by swapping smartphones, cameras, or spectrometers. Out-of-distribution detection is tested using unseen analytes and off-range concentrations to verify that the model abstains rather than misclassifies.
In summary, (i) selectivity is emergent from the multivariate fingerprint and machine learning (ML)15 extracts the discriminative structure that no single probe provides; (ii) to analyze complex mixtures, include mixtures in training and use models that can deconvolute overlapping patterns (PLS-R/SVR) or perform multi-label classification; (iii) for real matrices, prefer matrix-matched calibration and (when no reference exists) standard addition to anchor quantitative predictions.
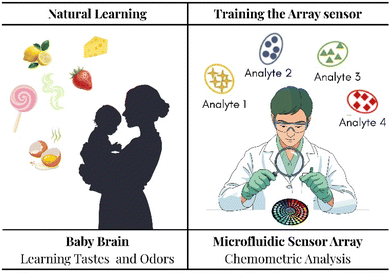
When properly trained, an array delivers reliable detection and quantification across targets, matrices, and devices, with built-in safeguards against drift and unknowns. We can assume that an array can be compared to a newborn. In particular, training an optical array is like teaching a newborn to distinguish between different tastes and smells (Fig. 2).16
(1) Single, simple tastes vs. single analytes in simple media (standard solution). Parents begin with plain flavours (rice cereal and apple purée), presented one at a time, so the infant learns a clear association. Likewise, an array first “meets” each target in a standard solution (ideal buffer); replicate responses are collected, normalized, and a basic model (PCA/LDA or PLS-R) is fitted to anchor the fingerprint of each analyte.
(2) New flavours and pairings vs. binary/ternary mixtures: as tolerance grows, parents introduce combinations (apple + pear), helping the child in separating overlapping notes. The same concept is applied with controlled mixtures to teach the model how patterns superimpose and how to deconvolute partial responses (multi-output PLS-R, SVR), adding a rejection class for unknowns.
(3) Family meals vs. real matrices: eventually the infant eats complex dishes where spices, fats, and textures interact; recognition becomes robust despite background “noise.” Arrays graduate to saliva, air, food extracts—matrix-matched and spiked via standard addition—to learn through confounders (pH, salts, proteins, and humidity).
(4) Repetition and growth vs. drift and recalibration: children refine preferences through repeated exposure; arrays maintain performance with QC samples, domain adaptation (camera/fiber changes), and periodic incremental learning.
Just as a child learns to recognize flavors reliably across meals, a properly trained array identifies and quantifies targets across mixtures and complex matrices, not by perfect specificity of a single “taste receptor,” but by the stable, learned pattern of many partial cues.
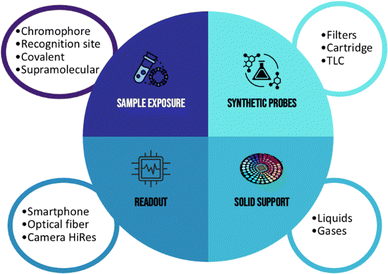
1.2 Chemical and hardware anatomy of the array
(i) Covalent recognition (reactive/probe–analyte chemistry):5 it is useful for irreversible tagging or “turn-on” specificity (e.g., nucleophiles with activated electrophiles). This approach shows very high sensitivity but suffers some limitations: (a) kinetics and selectivity are matrix-dependent (pH, water, and nucleophiles); (b) risk of side reactions, over-tagging, and drift due to probe depletion; (c) lack of selectivity, in particular considering the possibility to perform analyses with real non-treated samples.
(ii) Supramolecular recognition (non-covalent): based on the principles of supramolecular chemistry, exploiting H-bonding, ion–dipole, electrostatics, π–π/CH–π, Lewis acid–base, and hydrophobic/solvophobic and polarity/solvatochromic effects can lead to reversible interactions.2 The advantages of this approach are (a) a reversible binding, thus leading to the possibility to reuse the sensing elements; (b) a dynamic range tuning, by modulating the chemical structure of the probe, and in particular the nature of the recognition site; (c) works for mixtures: each analyte perturbs the ensemble differently, yielding rich fingerprints; (d) lack of false-positive responses: due to the formation of equilibria between the probe and the analyte, the presence of interferents is not a problem because the supramolecular complex between the probe and interferents can be suppressed by the presence of the target analyte.
(1) Chemical and mechanical stability: hydrophobic/low-energy porous films (e.g., polyamide, PVDF, and PTFE-laminates), silica/zeolite/MOF coatings with passivated surfaces, RP-TLC plates with suitable binders, stable to humidity swings, solvents of interest, and repeated illumination are key aspects that an ideal solid support should possess.23
(2) Ease of handling: the support should accept microdispensing/printing, dry quickly without coffee-ring artifacts, and retain spot geometry.24 In addition, the compatibility with adhesives/carriers, cutters, and housings for cartridge assembly can improve the performances.
(3) Portability: the main features of the PoC device are the portability and the easy-to-use system. In this context, thin, flexible cards or small cartridges, minimal outgassing, low background fluorescence, compatibility with sealed chambers or flow cells are important characteristics.
Across both gas and liquid nature, environmental parameters are recorded and, when necessary, stabilized: in particular, temperature and humidity are monitored continuously. These measures ensure that the signal reaching the probes reflects the analyte, not the sampling hardware or handling history, preserving both qualitative identity and quantitative fidelity.
Fiber-optic spectroscopy provides a complementary route where full emission spectra are needed. It delivers high signal-to-noise in milliseconds, enables ratiometric analysis, and interfaces naturally with solid-state arrays26 or flow cells.27 Performance depends on a stable lamp or laser, precise control of the integration time, suppression of stray light, and rigid positioning of the probe. Validation includes wavelength and intensity standards, explicit transfer between instruments, and long-term drift monitoring to ensure that spectral features remain comparable across sessions and hardware.
Multispectral and hyperspectral cameras are employed when spatial heterogeneity carries information or when process monitoring demands both imaging and spectral discrimination.28 Here, calibrated illumination and rigorous flat-field and dark corrections are mandatory. Dataset dimensions are quite large, and they require dedicated pipelines for dimensionality reduction and modelling, typically starting with PCA or PLS and extending to SVMs, random forests, or convolutional networks when spatial patterns matter.15 Robustness is demonstrated by maintaining accuracy across days and batches and by validating on external false unknown test sets resulting in cross-validation and confusion matrixes.
Being independent of the readout process, the analysis chain follows the same principles. Background is subtracted and signals are normalized per spot and per card to remove illumination and loading differences; features are scaled to stabilize variance. Exploratory models map the structure of the data and expose outliers, while supervised models are chosen according to task: discriminants and PLS-DA for classification under collinearity, margin-based or ensemble methods for nonlinearity, and PLS-R or SVR for quantitative predictions, with explicit out-of-distribution rejection, so the system can abstain from unfamiliar patterns.
1.3 The problem of complex matrices: operating without pretreatment
Each manipulation of the matrix to be analysed leads to a loss of analytical information. For this reason, the importance of the analytical validation step is crucial, in order to demonstrate that the target analyte is maintained after the manipulation of the sample. However, if multi-analyte analysis is the target, this process is expensive. In addition, manipulation of the sample requires time, costs and, in most cases, specialized personnel, thus limiting the application for PoC and/or in-house analyses.In clinical analysis, the pre-analytical phase is the primary driver of information loss: it accounts for ∼60–70% of total laboratory errors, and within that, sample integrity issues explain 80–90% of rejections or unreliable results.29 These phenomena are not mere technicalities but actively alter the matrix before analysis, changing both, which analytes we measure (qualitative composition) and how much (quantification).
Concrete examples show how routine operations introduce a systematic bias: haemolysis releases K+/LDH/ALT/AST and changes haemoglobin absorbance, distorting spectrophotometric readouts;30 icterus produces artifactually low glucose/lipids;31 lipemia causes pseudo-hyponatremia with indirect ISE and variability in many measurements.32 Logistical choices—order of draw, additive carryover (e.g., EDTA), patient posture, and the timing of centrifugation/separation—shift matrix chemistry with measurable effects on protein binding and electrolytes. Handling and storage are equally critical: defined time/temperature windows (e.g., separation within 2 h; stability ∼8 h at room temperature/48 h at 2–4 °C, then freezing) and automated HIL indices are required to prevent errors invisible to visual inspection. Without these controls, degradation or matrix interference leads to worse LOD/LOQ, biased results, and poor inter-laboratory comparability.
Since human error predominates (∼82.6%), mitigation requires standardization (EFLM guidance, ISO 15189 requirements), continuous training, and automation.33 The lesson for sensor and rapid-test developers is clear: the more the sample preparation perturbs the matrix's chemical composition, the greater the risk of information loss. Conversely, minimal-manipulation protocols—or measurements on as-collected specimens—better preserve the system's chemical reality, reducing qualitative/quantitative bias and improving the transferability of results.
These problems have also been found in chromatographic analyses.34 Classical preparation strategies, such as dilute-and-shoot, liquid–liquid extraction (LLE), solid-phase extraction (SPE), quick, easy, cheap, effective, rugged, and safe (QuEChERS), introduce trade-offs: dilution attenuates interferences but weakens the analyte signal and does not guarantee the disappearance of the matrix effect; LLE and QuEChERS can co-extract interferents or introduce salts/additives that, if not removed, distort ionization and increase instrument maintenance; SPE reduces the matrix effect only if the sorbent chemistry is well matched to the analytes, otherwise it concentrates the matrix along with analytes.
Instrumentation matters as well: electrospray ionization or chemical ionization (ESI or CI) is more prone to suppression than atmospheric pressure photoionization (APPI). In gas chromatography (GC), active sites in the inlet and injection conditions/liner materials drive adsorption/degradation that translate into a matrix effect. In liquid chromatography (LC), RPLC can exhibit co-elution of matrix components (e.g., phospholipids), requiring gradient redesign, mobile-phase additives, or additional clean-up steps.
For human sampling, saliva is probably the lowest-burden option, minimizing stress and enabling self-collection. The pre-analytical phase directly affects the quality and quantity of measurable analytes in saliva.35
Centrifugation is a critical source of compositional alteration: 1000 × g is most commonly used to remove turbidity, but increasing the speed significantly reduces total protein levels, with selective loss of larger proteins. Furthermore, the sample's rheology (viscosity and gel-like properties) can amplify this loss. Even when helpful for clarification, it must be optimized to balance removal of impurities with preservation of analytes, because the impact on the limit of detection can be substantial.
The device choice alters the measurable composition: absorbent pad systems can retain a high proportion of the interfering mucinous materials while analytes remain in the filtered fraction, functionally equivalent to centrifugation; this may save time but changes the matrix. Even more critical, cotton devices (e.g., Salivette®) can bind specific analytes (Aβ and hormones), preventing accurate quantification; inert polymeric materials are preferable. Concrete examples show that using Salivette led to non-detection of Aβ42/40 relative to passive drool, underscoring how the collection method can overturn analytical outcomes.36,37
Other important parameters are processing timing and storage: the sample should be processed/centrifuged within 1 hour of collection to avoid protein degradation. Inspection for contaminants (blood and pigments) is recommended and, if needed, testing for occult blood to avoid false biomarker increases. After aliquoting, −80 °C storage is indicated to preserve integrity and pH, whereas −20 °C can induce spectral changes already reported in the literature.38 Saliva specimens are not robust to multiple freeze–thaw cycles: a maximum of 2–3 cycles is suggested, because cell lysis releases degradative enzymes.36
In summary, every manipulation, from device selection to centrifugation parameters, from processing timelines to storage schemes, can alter what and how much we measure. Standardizing or, when possible, avoiding sample treatment (measurements on as-collected saliva) reduces the risk of informational loss and increases comparability across studies.
Metabolite profiling vividly shows that “pre-analytical chemistry” rewrites biology: routine quenching, extraction, drying, and platform-specific handling can interconvert species, bias ionization, and fabricate false positives.39 Incomplete quenching allows enzymatic carry-over to transform high-energy metabolites (e.g., from 3-phosphoglycerate to phosphoenolpyruvate; from ATP to ADP), unless strong acidic mixes (e.g., 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40
40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 acetonitrile
20 acetonitrile![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) methanol
methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water + 0.1 M formic acid) are used and promptly neutralized.40 After extraction, drying steps disproportionately oxidize redox pairs (NADPH/NADP+, GSH/GSSG), shifting biologically meaningful ratios even when most other metabolites remain stable.41 On LC-MS, the matrix and source chemistry generate further distortions: isomers and in-source fragments masquerade as genuine metabolites, while ion suppression and adduct competition skew sensitivity and linearity, making quantitative claims precarious unless standards, dilution, and rigorous chromatography are used.42 GC-MS improves separation for many small volatiles but demands derivatization and hot injection, introducing decomposition routes that also alter the matrix. NMR avoids ionization bias yet suffers from crowding, pH/osmolality-dependent chemical shifts, and resolution limits that inflate error without 2D methods and careful calibration. The common lesson is blunt: every extra manipulation perturbs chemical equilibria, degrades labile species, or invents analytes; therefore, assays that either eliminate or tightly standardize sample treatment—and, when possible, interrogate specimens as collected—better preserve qualitative composition and quantitative fidelity.
water + 0.1 M formic acid) are used and promptly neutralized.40 After extraction, drying steps disproportionately oxidize redox pairs (NADPH/NADP+, GSH/GSSG), shifting biologically meaningful ratios even when most other metabolites remain stable.41 On LC-MS, the matrix and source chemistry generate further distortions: isomers and in-source fragments masquerade as genuine metabolites, while ion suppression and adduct competition skew sensitivity and linearity, making quantitative claims precarious unless standards, dilution, and rigorous chromatography are used.42 GC-MS improves separation for many small volatiles but demands derivatization and hot injection, introducing decomposition routes that also alter the matrix. NMR avoids ionization bias yet suffers from crowding, pH/osmolality-dependent chemical shifts, and resolution limits that inflate error without 2D methods and careful calibration. The common lesson is blunt: every extra manipulation perturbs chemical equilibria, degrades labile species, or invents analytes; therefore, assays that either eliminate or tightly standardize sample treatment—and, when possible, interrogate specimens as collected—better preserve qualitative composition and quantitative fidelity.
Very recent and comprehensive reviews on the construction strategies and principles of sensing elements, as well as the applications of sensor arrays for identification and detection of target analytes in a wide range of fields, have been published.6,43,44 However, this Feature Article summarizes the literature relative to untreated sample analyses using optical arrays as sensing devices.
2. Human health application
Clinical samples (saliva, urine, and breath condensate) are highly dynamic and enzyme-rich. Centrifugation, filtration, storage, and freeze–thaw cycles can deplete proteins, adsorb small molecules to collection devices, shift pH/ionic strength, and trigger degradation or interconversion. These pre-analytical steps distort both qualitative profiles and quantitative readouts, undermining comparability and LOD/LOQ. Measuring as-collected specimens with cross-reactive optical arrays preserves native composition, reduces handling bias, and enables direct pattern recognition in real matrices.Hof and co-workers reported abiotic calixarene hosts that self-assemble into dimeric and heterodimeric capsules (“dimer-dyes”).45 This is, to our knowledge, the first array study employing abiotic macrocycles such as calixarenes. The authors target drug sensing in buffered solution and in saliva, diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in water, performing titrations that yield limits of detection around ∼100 µM. The array comprises five selected dimer-dyes and is used to discriminate methamphetamine, amphetamine, MDMA (3,4-methylenedioxymethamphetamine), MDA (3,4-methylenedioxyamphetamine), nicotine, cocaine, oxycodone, and 6-MAM (6-acetylmorphine); array measurements are conducted at 100 µM analyte concentrations. Real saliva samples are diluted to alleviate viscosity issues. The readout uses a simple fluorimeter: probes are plated, and emission spectra are recorded directly. Crucially, the study includes real patients administered with a clinical dose of cocaine. In particular, saliva is sampled between 0 and 4 hours post-administration to determine cocaine specifically by the array. This work establishes a clear, pioneering precedent for macrocycle-based, abiotic array sensing in complex biofluids using straightforward fluorescence plate measurements.
1 in water, performing titrations that yield limits of detection around ∼100 µM. The array comprises five selected dimer-dyes and is used to discriminate methamphetamine, amphetamine, MDMA (3,4-methylenedioxymethamphetamine), MDA (3,4-methylenedioxyamphetamine), nicotine, cocaine, oxycodone, and 6-MAM (6-acetylmorphine); array measurements are conducted at 100 µM analyte concentrations. Real saliva samples are diluted to alleviate viscosity issues. The readout uses a simple fluorimeter: probes are plated, and emission spectra are recorded directly. Crucially, the study includes real patients administered with a clinical dose of cocaine. In particular, saliva is sampled between 0 and 4 hours post-administration to determine cocaine specifically by the array. This work establishes a clear, pioneering precedent for macrocycle-based, abiotic array sensing in complex biofluids using straightforward fluorescence plate measurements.
In 2021, Hormozi-Nezhad and co-workers demonstrated the use of a nanoparticle-based array, combining fluorescent cadmium telluride quantum dots and carbon dots, to detect dopaminergic molecules in human urine samples.46 The dual-emission probe approach used by combining the two types of nanostructures merges both inner filter effects and electron transfer mechanisms to detect molecules like levodopa, carbidopa, benserazide, and entacapone. The array was set up in a 96-well microplate and photographed via a smartphone for convenient readout. In solution studies, the system showed excellent classification of analytes at concentrations ranging from 1 to 100 micromolar. In real human urine samples (diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 with water), the array successfully quantified dopaminergic compounds in the 10–50 micromolar range using standard addition methods, though only a single urine sample was tested, limiting statistical robustness. Nonetheless, this study illustrates the potential of combining multiple nanomaterials for practical, smartphone-readable sensing of neurological drugs in biological fluids.
10 with water), the array successfully quantified dopaminergic compounds in the 10–50 micromolar range using standard addition methods, though only a single urine sample was tested, limiting statistical robustness. Nonetheless, this study illustrates the potential of combining multiple nanomaterials for practical, smartphone-readable sensing of neurological drugs in biological fluids.
In the same year, Minami and co-workers developed a mini-array with coumarin-based probes whose fluorescence was quenched by Zn2+ and restored upon binding sulfur-containing amino acids.47 In solution, the array detected cysteine, homocysteine, and glutathione disulfide and asparagine. When applied to diluted human serum, the system quantified glutathione and cysteine with linear ranges up to 60 parts per billion (ppb) for glutathione and 25 ppb for cysteine, with detection limits of 6 ppb and 2.5 ppb, respectively. Notably, the analytes were spiked into the serum and quantified using a standard addition method without comparison to routine clinical assays, demonstrating the array's potential for selective detection in complex biological samples.
Han and co-workers reported a dual-channel fluorescence array for identifying pathogenic bacteria in urine.48 The platform comprises six probes formed by blending a fluorescent polymer with differently functionalized ammonium species that modulate the polymer's emission (see Fig. 4). Mechanistically, the ammonium components quench the polymer fluorescence; upon exposure to bacteria—whose envelopes present abundant anionic groups—the ammonium moieties bind preferentially to the cell surface, releasing the polymer from quenching and restoring its emission. Additional probe functionalities are tailored to engage lipopolysaccharide and peptidoglycan, broadening the interaction space. The readout exploits two independent channels: one tracking the polymer emission and the other the ammonium-probe emission, thereby increasing discriminative power. Training in buffered solutions against approximately twenty bacterial species produced clear clustering, with especially distinct separation among Gram-positive organisms. Quantitative behavior was demonstrated over optical densities (OD) of ∼0.2 to 0.001. In targeted assays relevant to urinary tract infections, the array identified Escherichia coli and Pseudomonas aeruginosa with good linearity and low detection limits within the same OD window. When challenged with artificial urine, it distinguished mixed cultures—such as E. coli with Enterococcus faecalis—while maintaining dual-channel discrimination. Clinical feasibility was evaluated across three cohorts: healthy controls, E. coli UTI, and E. faecalis UTI. Urine samples were diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 in PBS, combined with the probe solution, and further diluted eightfold prior to measurement. Plate-based fluorescence collected on a benchtop fluorimeter yielded an overall correct identification rate of ∼70.5%, which was benchmarked against standard laboratory diagnostics. Although experiments were conducted in microplate wells without a microfluidic or solid-state point-of-care format, the results constitute a strong proof-of-concept for urine diagnostics using a chemically diverse, dual-channel array.
4 in PBS, combined with the probe solution, and further diluted eightfold prior to measurement. Plate-based fluorescence collected on a benchtop fluorimeter yielded an overall correct identification rate of ∼70.5%, which was benchmarked against standard laboratory diagnostics. Although experiments were conducted in microplate wells without a microfluidic or solid-state point-of-care format, the results constitute a strong proof-of-concept for urine diagnostics using a chemically diverse, dual-channel array.
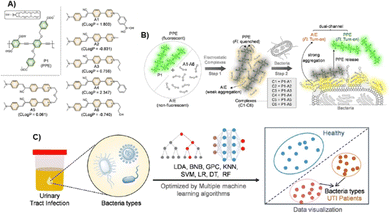 | ||
| Fig. 4 (A) Chemical structures of polymer and ammonium species; (B) proposed mechanism of bacterial recognition; (C) real sample analyses. Adapted with permission from ref. 48. Copyright 2024 Wiley. | ||
Wang and co-workers developed an array for detecting volatile organic compounds (VOCs), both in laboratory settings and in a real-world application as a wearable mask sensor.49 The goal was to detect lung cancer-related metabolites in exhaled breath. The array is built on a porous zeolite framework loaded with various commercial organic dyes (such as bromocresol purple, methyl red, phenol red, and rhodamine B). Detection is based on capturing RGB image data before and after exposure to VOCs. In the laboratory, the array was tested for sensing ethylenediamine at 50% humidity using nitrogen as a carrier gas. The system achieved a detection limit of 0.17 ppm and was tested mainly against air and CO2 as potential interferents. The notable real-world application involved integrating the array into a wearable face mask designed to screen for lung cancer metabolites in breath samples (see Fig. 5). The mask was structured to prevent particulates from reaching the array, ensuring that only relevant gases were detected. The authors found that the best sampling time was early in the morning before breakfast, to avoid interference from food-related VOCs. They tested 15 healthy individuals and 20 lung cancer patients. The system effectively discriminated between healthy and cancer-affected individuals, leveraging the zeolite's porosity to enhance gas penetration and improve sensitivity and selectivity of the sensor array.
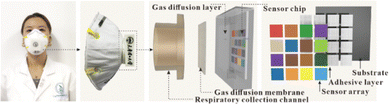 | ||
| Fig. 5 Real image and section of the portable colorimetric sensing device. Reprinted with permission from ref. 49. Copyright 2024 Wiley. | ||
Recently, our group presented an array used for the detection and quantification of creatinine in human saliva.50 This is the first point-of-care system capable of measuring creatinine in saliva without any pretreatment, demonstrating how synthetic probes can be used to quantify a wide concentration range in aqueous samples. Monitoring creatinine levels is crucial because creatinine is involved in key metabolic processes and is an important marker of kidney health. Various renal conditions can be correlated with altered creatinine levels. The array was calibrated for creatinine detection across a concentration range from 10 mM down to 10 nM, covering six orders of magnitude, a much broader range than typical creatinine assays. The selectivity was tested using artificial saliva containing common analytes found in human saliva, as well as potential interferents, such as dopamine, cortisol, adrenaline, noradrenaline, glucose, uric acid, urea, and inorganic salts. An optical fiber detector was used to record the full emission spectrum of all probes onto the solid support of the array, covering wavelengths from 400 to 700 nanometers. A recovery test was performed by spiking 4 µM creatinine into artificial saliva, yielding excellent recovery results. The array was also tested on real saliva samples. Due to the lack of established analytical methods for quantitative creatinine determination in human saliva, the array was validated using a standard addition method, which determined a creatinine concentration of 3.98 mM in the tested sample. This study is particularly significant as it represents the first point-of-care diagnostic device for creatinine detection in human saliva.
In the same year, we introduced the first point-of-care device capable of simultaneously determining cortisol, dopamine, and adrenaline in untreated human saliva in a single analysis.51 The array technology, based on synthetic fluorescent probes, allowed calibration of these three stress biomarkers over a concentration range from 1 mM down to 1 pM. This large limit of detection was achieved using an optical fiber to acquire the full emission spectrum of the probes directly from the array. The calibration, selectivity, and recovery tests were all performed in artificial saliva, yielding excellent results for all three biomarkers at concentrations as low as 100 pM. Real saliva studies were conducted on four subjects. The cortisol values obtained from the array were compared with laboratory ELISA methods, showing excellent agreement in all cases. For dopamine and adrenaline, the standard addition method was used for quantification, as no existing point-of-care kits for these analytes in saliva are currently available. Key highlights of this work include the broad analytical range from millimolar to picomolar levels, the ability to analyse untreated saliva directly, and the capability to detect three stress biomarkers (cortisol, dopamine, and adrenaline) simultaneously in a single test.
More recently, we modified the chemical composition of the array, in terms of synthetic fluorescent probes, obtaining an optical array device capable of detecting Staphylococcus aureus directly from complex matrix secretions.52 The system requires no sample pretreatment or signal amplification, minimizing false positives and making it an ideal candidate for detecting peri-prosthetic infections. The array's probes detect bacterial metabolites, which are complex mixtures of aromatic compounds and other components. The selected fluorescent probes are designed to interact via hydrogen bonds, CH–π, π–π, ion–dipole, and dipole–dipole interactions. Detection is performed using a low-cost, flexible, and chemically resistant optical fiber. The array was trained to detect S. aureus over a concentration range from 102 to 107 CFU per mL, achieving excellent linearity. It was also trained to identify Staphylococcus epidermidis, Escherichia coli, Pseudomonas aeruginosa, and Enterococcus faecalis, showing high selectivity for S. aureus (Fig. 6). The system could also distinguish S. aureus from S. epidermidis in complex mixed samples. Simulated real samples were used, specifically synovial fluid infected with S. aureus in a range from 102 to 105 CFU per mL, achieving excellent linearity (R2 = 0.9997) even in the simulated matrix. The system can effectively cluster and differentiate between synovial samples contaminated with S. aureus and those that are not, demonstrating both qualitative and quantitative diagnostic capabilities. The analysis takes only a few tens of minutes, making it suitable for real-time application in operating rooms. This can help detect infections promptly, reducing hospitalization times, treatment costs, excessive antibiotic use, and potentially lowering post-operative mortality rates.
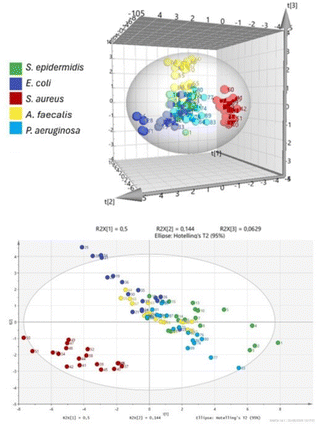 | ||
| Fig. 6 Discrimination of bacteria by the selectivity test using the PLS-DA model. Adapted with permission from ref. 52. Copyright 2026 Elsevier. | ||
Anzenbacher and co-workers reported a boronic acid used to coordinate and recognize sugars, yielding a fluorescent sensor based on the FRET phenomenon.53 This sensor is employed to detect monosaccharides and disaccharides in urine samples. The sensor includes two distinct boron-linked fluorophores: a tryptophenol and a coumarin. Upon sugar detection, coumarin units are displaced, turning off FRET and resulting in tryptophenol emission. Binding constants, in the range 106–107 M−1, are calculated for various sugars by monitoring the fluorescence changes of both fluorophores. Sugar concentrations tested range from 10 to 400 µM. The sensor is then validated in urine samples, where glucose typically falls in the millimolar range. It detects nine different sugars in urine with excellent classification and selectivity. Glucose is tested from 0 to 60 mM and compared with a commercial enzymatic kit, achieving a limit of detection of 0.94 mM.
3. Environmental application
Air and water monitoring often relies on trapping, solvent extraction, or solid-phase cleanup that fractionates complex mixtures, introduces blanks/artefacts, and selectively loses volatiles, semi volatiles, or reactive species. Such processing reshapes fingerprints and biases concentration estimates across time and sites. In situ or minimally handled measurements maintain the original chemical ensemble, yielding more faithful spatiotemporal signatures.Recently, Jiang and co-workers developed an array incorporating four fluorescent probes, each selective for a different metal ion: mercury, chromium, copper, and lead.54 These probes are arranged in a microfluidic chip, where a sample is loaded at the centre and then moves by capillarity to the circularly arranged probes (see Fig. 7). Real samples analysed in the study included lake and river water, which were only pretreated to remove insoluble impurities, thus effectively representing real-world conditions. The presence of each metal ion leads to an increase in the fluorescence emission of its corresponding probe. For mercury, the system shows linearity up to 1.2 µM with a detection limit of 0.89 nM. For chromium, linearity extends to 12 µM with a detection limit of 5.45 nM, for lead, the major linear range was observed (up to 36 µM) with a LOD of 9.60 nM, and for copper, linearity reached 2.40 µM with a detection limit of 1.77 nM. These are relatively narrow linear ranges at low concentrations.
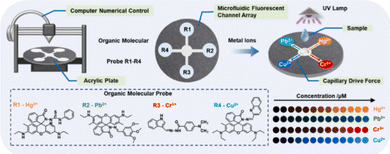 | ||
| Fig. 7 Microfluidic fluorescent sensor array workflow for heavy metal ion detection. Reprinted with permission from ref. 54. Copyright 2024. Elsevier. | ||
The microfluidic chip can detect all ions simultaneously within about one minute of image acquisition. The concentration range for real samples extends up to 20 µM. High-resolution digital imaging is used to capture the RGB channels of the array's fluorescence, which are then processed to quantify the metal content.
Recently, Du and co-workers reported on an array based on four fluorescent carbon dots capable of recognizing rifamycin antibiotics in water.55 The carbon dots were synthesized from boric acid and serine and then complexed with Cr(III), which induces a quenching effect on the fluorescence. This approach was chosen to leverage multiple detection mechanisms—including covalent reactions, hydrogen bonding, Lewis acid–base interactions, and electrostatic interactions—to achieve a broad fingerprint response to the target antibiotics. The sensing is completed in about one minute with analyte concentrations in the micromolar range at a buffered pH of 10. The system can also discriminate against mixtures of antibiotics. Quantitative analysis was specifically performed for rifampicin and rifapentine, with a linear range from 2.5 to 25 µM for rifampicin and 2.5 to 40 µM for rifapentine. The limits of detection were 45 nM and 87 nM, respectively. For real-sample analysis, lake water and BSA (bovine serum albumin) samples were used as matrices. Both were filtered through 0.22 µm filters, and the BSA samples were diluted in water at different dilutions. Since no antibiotics were naturally present in these samples, a standard addition method was used, yielding excellent recovery results. A crucial point is the use of relatively high concentrations of Cr(III) (1–2 mM) in the array, the need for a high operating pH of 10, and the requirement of a solution-phase fluorimeter, meaning that the system is not portable or point-of-care.
Very recently, our research group reported an array system capable of detecting methylamine, an irritant at very low concentrations (ppb) and toxic at higher levels.56 The array achieves a limit of detection of 100 ppb, which is lower than that typically reported in the literature and operates using an optical fiber as a detector over a concentration range of 100 to 1000 ppb, yielding excellent linearity (R2 = 0.9998). The synthetic organic probes in the array utilize non-covalent interactions, including hydrogen bonds, ion–dipole, and dipole–dipole interactions.
A gas permeator is used to precisely dose the methylamine released, allowing the array to perform flow-based measurements and achieve a stable response in about one hour, due to the time required for the analysis environment to saturate (Fig. 8). Methylamine detection was performed using air as the matrix, ensuring selectivity against common atmospheric interferents. The system maintained strong performance even at varying humidity levels and showed excellent selectivity when tested against other gaseous analytes like ammonia, ethyl mercaptan, and hydrogen sulphide.
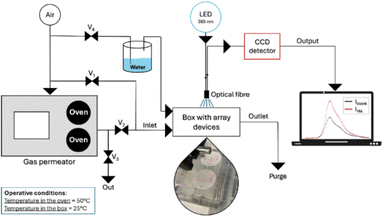 | ||
| Fig. 8 Scheme of the setup for methylamine detection using a fluorescent array. Reprinted with permission from ref. 56. Copyright 2026. Elsevier. | ||
New and co-workers reported a six-probe array for sensing nine different heavy metals.57 All probes contained in the array are coumarin-based with various functionalizations. The array achieved excellent selectivity with 100% classification. The authors later reduced the array to just three probes to minimize synthetic effort and analysis costs, choosing the most chemically representative probes for metal recognition. Heavy metal concentrations tested were 10 µM in solution, with probe concentrations also at 10 µM. The array was then tested in real-world tin-plating water, with results compared to ICP-MS analysis. In this context, some metals exhibited greater fluorescence quenching compared to solution measurements, likely due to existing metal content in the tin-plating water matrix. Quantitative analysis was performed solely for Pb2+, achieving good results at a minimum of 20 µM. Notably, the array operates on microplates examined via fluorescence, requiring two hours of incubation per analysis.
4. Agri-food application
Food and post-harvest samples contain emulsions, proteins, sugars, and volatile bouquets that are readily perturbed by dilution, filtration, or solvent extraction, suppressing key notes and enriching others. Pre-treatment can mask early spoilage VOCs or alter metabolite ratios relevant to freshness and contamination. Direct analysis of untreated headspace or micro-aliquots on solid-state arrays captures the intact multidimensional fingerprint, improving early detection of pathogens, ripening state, and authenticity without sacrificing selectivity or the dynamic range.In 2023, our group presented the first work on fluorescent synthetic probe arrays deposited on a polyamide filter able to detect the presence of pathogens in orange washing water.58 The fluorescence emission was recorded using a smartphone that captured JPEG images, and grayscale channel analysis provided quantitative information on the presence of pathogens. Six different pathogens were studied in untreated orange wash water samples. The array showed excellent selectivity, particularly for Penicillium italicum, and the studied concentration range spanned five orders of magnitude. This demonstrates the potential for smartphone-assisted, array-based detection of pathogens in real-world agricultural samples. In addition, this approach enabled detection over a concentration range between five orders of magnitude, demonstrating an exceptionally broad linearity and detection limit range. This highlights how fluorescent chemical probes can achieve a wider analyte concentration range compared to many existing sensors in the literature.
Recently, we improved the performance of the array, developing a fluorescent synthetic probe array designed to monitor the presence of VOCs inside sealed containers of oranges.59 The array was specifically trained to distinguish VOCs emitted by healthy oranges from those released by oranges contaminated with Penicillium digitatum. In the case of healthy fruit, the VOC profile changes over time, allowing the array to estimate the duration of storage (see Fig. 9). For contaminated oranges, the array detects characteristic VOCs from P. digitatum even before visible signs of infection appear. The chemical detection mechanism is based on non-covalent interactions between the VOCs and the synthetic fluorescent probes in the array. Specifically, the authors attribute the sensing response to π–π stacking and CH–π interactions formed between the VOCs and the probes. This non-covalent recognition allows the array to achieve a broad detection range and a sensitive readout. Optical detection is performed via optical fiber, enabling emission spectra to be obtained in seconds without opening the packaging. This real-time, non-invasive monitoring method can be applied to track both the storage duration and potential fungal contamination inside sealed packaging, offering a practical tool for ensuring product safety and quality.
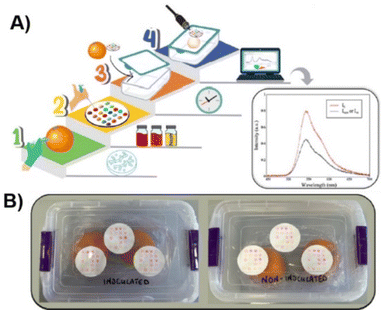 | ||
| Fig. 9 (A) Workflow of sensing by the fluorescent array; (B) photos of the packaging used to detect orange storage and contamination. Reprinted with permission from ref. 59. Copyright 2024. Royal Society of Chemistry. | ||
In the same year, Wang and co-workers developed an optical array using eight commercial probes and two palladium-based probes embedded in porous MOFs to detect VOCs.60 The goal is to sense VOCs emitted by fruits—specifically bananas, mangoes, and kiwis. The study highlights the importance of detecting complex VOC mixtures in ambient air at low concentrations and the need for flexible, surface-attachable devices for real-time analysis. The array is supported on a microchannel structure made of PDMS, which guides VOCs toward the sensing elements. The MOF particles, containing different commercial gas-sensitive dyes, are deposited onto a polyacrylonitrile membrane via vacuum filtration. The entire system is then attached to the fruit's surface or, in the case of human subjects, designed to be flexible for wearable use. The array relies on a digital camera or smartphone for on-site image capture and analysis. In the laboratory, the authors tested model VOCs like benzaldehyde, trans-2-hexenal, 3-carene, ethanol, and ethyl acetate at ppm levels. For example, the detection limit for trans-2-hexenal was around 5 ppm with a response time of up to 10 minutes, and concentrations up to 1500 ppm were tested. The array's real-sample monitoring focused on bananas, capturing VOC-induced changes with a camera, although specific VOC identities and exact analysis times were not detailed. Banana samples can be monitored during a storage time of 1–9 days. Overall, this work demonstrates a portable VOC sensing array that can be easily applied to different surfaces and read out using common digital devices, addressing challenges like detecting low-concentration VOCs and handling complex mixtures in ambient air.
Zor and co-workers reported an array of 16 colorimetric probes for monitoring fish quality, stored both at room temperature and at 4 °C.61 All probes are commercial. Samples are sealed in airtight containers, with the array placed inside to capture and analyse the volatile organic compounds released. Array analysis is conducted by simple scanning and colour analysis. Among the volatiles, the authors likely identify trimethylamine, dimethylamine, cadaverine, and putrescine. The array also monitors physicochemical parameters such as TVB-N, TBA, and pH. Additionally, since the array is placed inside the container, the method is non-destructive and non-contaminating to the food sample.
Table 1 summarizes some important features of the examples previously reported:
| Application field | Target analyte(s) | Matrix | Nature of sensors and detection mechanism | LOD | Linear range | Resp. time | Ref. |
|---|---|---|---|---|---|---|---|
| Health | MA, MDA, MDMA, nicotine, cocaine, oxycodone, and 6-MAM | Water; saliva 50% | “DimerDyes” fluorescent array + chemometric analysis | Nicotina: 3.4 µM (H2O), 18.6 µM (saliva); MDMA: 2.7 µM (H2O), 41.2 µM (saliva); cocaine: 2.7 µM (H2O and saliva) | n.d. | n.d. | 45 |
| Levodopa, carbidopa, benserazide, entacapone | Human urine (spiked) | Fluorescent tongue (QDs + CDs) + smartphone; LDA | n.d. | Bens 1–100 µM; Levo 20–80 µM; Carb 10–100 µM; Enta 1–100 µM | ∼6 min | 46 | |
| Cys, GSH, GSSG, HCy, CySS | Human serum (spiked) | Array chemosensitive (cumarine–Zn2+) UV/vis + FL; LDA/SVM | GSH: 1.0 ppb; Cys: 2.5 ppb | For GSH from 0 to 60 ppm, for Cys from 0 to 25 ppm. | 2–5 min | 47 | |
| E. coli and E. faecalis | Artificial and real urine samples | Array AIE-luminogens (PPE-AIE) | 2.95 × 10−4 OD600 and 3.29 × 10−4 OD600 | From 0.001 to 0.2 OD600 | n.d. | 48 | |
| Ethylenediamine | Gas flow, exhaled breath (real, wearable-mask sampling) | Array colorimetric H-ZIF-8/dye micro-free; read RGB | 0.17 ppm | n.d. | 10 min | 49 | |
| Creatinine | Artificial and real saliva samples | Fluorescent tongue + optical fiber | 10 nM | 10 nM–10 mM | 40 min | 50 | |
| Stress biomarkers (cortisol, dopamine, and adrenaline) | Artificial and real saliva samples | Fluorescent tongue + optical fiber | 1 pM | 1 pM–1 mM | 40 min | 51 | |
| S. aureus | Synovial fluid | Fluorescent tongue + optical fiber | 102 CFU per mL | 102–107 CFU per mL | 40 min | 52 | |
| Sugars | Urine | Fluorescent probe by a FRET mechanism | 0.94 mM | n.d. | n.d. | 53 | |
| Environmental | Transition metals | Lake and river waters | Fluorescent array + smartphone | Hg2+: 0.89 nM; Pb2+: 9.60 nM; Cr3+: 5.45 nM; Cu2+: 1.77 nM | Hg2+: 0–1.2 µM; Pb2+: 0–36 µM; Cr3+: 0–12 µM; Cu2+: 0–2.40 µM | <1 min | 54 |
| Rifampicin (RFC), rifapentine (RFP), rifabutin (RFB), rifaximin (RFX), rifamycin S (RFS) | Lake waters (spiked) and BSA | Fluorescent array based on CDs/Cr3+; LDA/PLS-DA | RFC: 45 nM | RFC: 2.5–25 µM | 1 min | 55 | |
| RFP: 87 nM | RFP: 2.5–40 µM | ||||||
| Methylamine | Air | Fluorescent tongue + optical fiber | 100 ppb | 100–1000 ppb | 1 h | 56 | |
| Transition metals | Tin-plating water | Fluorescent array coumarin-based | 20 µM | n.d. | 2 h | 57 | |
| Agri-food | Penicillium italicum | Oranges | Fluorescent tongue + smartphone | 103 conidia per mL | 103–108 conidia per mL | 3 min | 58 |
| Penicillium digitatum | Oranges | Fluorescent tongue + optical fiber | n.d. | n.d. | 10 min | 59 | |
| VOCs | Headspace fruit | Colorimetric array + smartphone | 5 ppm | 5–500 ppm | 10 min | 60 | |
| VOCs | Fish | Colorimetric array + scanner | n.d. | n.d. | 10 min | 61 |
LODs: the best performance in detection limits is achieved in the saliva-based stress-biomarker array (cortisol, dopamine, and adrenaline), with an LOD of 1.0 pM and a calibration range from 1 pM to 1 m.51 In thiol sensing, the dual-probe system in serum reaches ppb-level detection—1.0 ppb (GSH) and 2.5 ppb (Cys)—with 2–5 min readout.47 In gases, the methylamine array attains 100 ppb,56 while the ethylenediamine VOC sensor reports 0.17 ppm with ∼3 min response.49 For microbial targets in agri-food, the smartphone array detects P. italicum at 103 conidia mL−1.58
Fastest analysis: the transition-metal microfluidic array delivers ∼1 min readout in lake/river waters.54 Rapid kinetics are also seen for thiol sensing (2–5 min) and for both ethylenediamine and P. italicum (∼3 min).
Widest linear ranges: the broadest span is again the stress-biomarker POC, covering nine orders of magnitude (1 pM–1 mM). Creatinine in saliva follows with six orders (10 nM–10 mM). In microbiology, the agri-food array for P. italicum covers 103–108 conidia mL−1 (five orders), while S. aureus detection spans 102–107 CFU mL−1 with excellent linearity in simulated synovial fluid.52 We would like to stress that these wide ranges of linearity are larger with respect to the classical biomolecule-based PoC.
5. Challenges and potential solutions
Despite their conceptual elegance and proven versatility, optical sensor arrays (colorimetric and fluorescent) still face a set of recurring limitations that often separate compelling demonstrations from deployable analytical tools. A critical view is essential because most failures are not due to insufficient sensitivity under ideal conditions, but due to reproducibility, domain shift, and the mismatch between laboratory workflows and real sampling constraints.A first bottleneck is reproducibility of the sensing layer. Spot-to-spot variability, coffee-ring deposition, local differences in film thickness, and lot-to-lot changes in probe purity or loading can alter the multivariate fingerprint enough to degrade classification, even when each individual probe appears stable. This problem is amplified in solid-state formats, where the microenvironment (polarity, water activity, and porosity) controls photophysics and binding equilibria. Practical remedies include standardized deposition (printing/microdispensing with validated drying conditions), internal reference spots, acceptance criteria for spot uniformity, and accelerated-aging protocols that qualify shelf-life before clinical or field testing.
A second bottleneck is matrix complexity. Real biological samples introduce proteins, lipids, salts, surfactants, particulates, and endogenous chromophores that can quench fluorescence, shift apparent colour, or adsorb onto supports, thereby reshaping the fingerprint. Attempts to “clean up” samples can solve one problem while creating another by selectively removing informative components or changing equilibrium composition. For the analysis of untreated matrices, the solution should be surface and device engineering, in particular low-adsorption substrates, antifouling coatings, controlled wicking/flow paths, and matrix-matched calibration (or standard addition) coupled to explicit interference testing at physiologically or environmentally relevant extremes.
A third bottleneck is time dependence. If acquisition time is not fixed, different laboratories (or even different operators) may measure different fingerprints at the same concentration. Robust practice requires that timing be treated as a controlled variable: assays should define a standardized timepoint and report exposure conditions (flow rate, residence time, sample volume, humidity, and temperature).
A fourth bottleneck is readout variability and calibration transfer. Smartphone cameras, low-cost spectrometers, and different illumination geometries can generate systematic shifts in RGB channels or spectral intensity that are unrelated to chemistry. Even within the same device class, automatic exposure, white balance, sensor aging, and lamp warm-up can introduce drift. The path to transferability is metrology: fixed geometry enclosures, stable illumination, raw acquisition when possible, embedded colour/spectral references, and explicit calibration-transfer strategies (e.g., device-specific correction matrices for imaging and piecewise direct standardization for spectra). Without these measures, multicenter studies are easily confounded by hardware effects masquerading as analytical performance.
Finally, regulatory and operational translation remains a major barrier for diagnostic applications. An array-based diagnostic is not only a chemical formulation but a coupled system of materials, hardware, and software. Regulatory-grade deployment demands a “locked” assay (fixed chemistry and readout conditions), version-controlled models with change management, traceable metadata, cybersecurity and privacy considerations for connected devices, and method comparison against accepted reference standards. These requirements are achievable, but they force the field to move from exploratory demonstrations toward standardized manufacturing, quality systems, and multicentre validation designs.
Conclusions and future perspectives
In this Feature Article we highlight that cross-reactive optical arrays— “noses/tongues”—outperform single lock-and-key sensors whenever real samples and mixtures are involved. Selectivity is emergent: a diverse panel of synthetic chromophores and supramolecular motifs translate analyte exposure into a high-dimensional fingerprint that chemometrics converts into identification and calibration. Efficient performance depends on deliberate panel design (chromophore photophysics + recognition site), rigorous training on standards, interferents, mixtures, and real matrices, and disciplined data handling that logs and corrects environmental covariates (T/RH/time) to prevent the drift from masquerading as chemistry. Readout can be smartphone imaging, fiber-optic spectroscopy, or hyperspectral cameras, provided that the geometry, illumination, and calibration transfer are controlled. Since each pre-analytical manipulation reshapes composition, arrays are most powerful when they are used to investigate minimally treated, or as-collected, specimens, preserving qualitative identity and quantitative fidelity while enabling wide linear ranges and concurrent multi-analyte detection. Finally, validation with stratified cross-validation and external holdouts, robustness testing for environment/hardware/aging, and routine quality control ensure that observed gains reflect true chemical signals and are portable across devices, operators, and sites.Across the studies summarized in this Feature Article, three main features emerge.
(1) Sensitivity: solid-state arrays read by fiber-optic spectroscopy deliver low LODs (pM or ppb) and the broadest linear ranges—up to nine orders of magnitude for salivary stress biomarkers and six for creatinine—demonstrating that cross-reactive panels can be calibrated from trace to high physiological levels.
(2) Speed: when arrays are integrated with microfluidics or thin films, minute-scale analyses are routine (1–5 min for metals and thiols), while gas sensing (e.g., methylamine) achieves sub-ppm operation with excellent linearity, albeit with longer equilibration when a controlled flow/saturation step is required.
(3) Real-matrix readiness: several entries quantify directly in untreated saliva, fruit headspace, environmental waters, or simulated synovial fluid using standard addition or matrix-matched calibration, confirming that pattern-based selectivity survives complex backgrounds.
This paper also highlights practical readouts—smartphone imaging for portability, fiber optics for quantitative spectra, and, where needed, hyperspectral imaging for heterogeneous samples—and shows that selectivity arises from the ensemble fingerprint, not from a single lock-and-key event.
Remaining gaps are methodological rather than conceptual: some studies use small clinical cases, a few platforms still rely on benchtop fluorimeters, and calibration transfer across devices needs formalization. Performance must be established on larger, multi-site cohorts with pre-organized protocols, matrix-matched calibrations, and external holdouts collected on different days, operators, and devices. Prospective studies should focus on the definition of the production budgets, on the determination of LOD/LOQ by validated procedures, and on the calibration stability over time.
Most importantly, platforms need to move beyond benchtop fluorimeters toward fully integrated, solid-state/flow-cell devices with fiber-optic or smartphone readout, standardized illumination geometry, and internal reference spots for on-the-fly drift correction.
Over the past few decades, optical arrays have evolved from proof-of-concept “spot panels” into engineered sensing platforms in which materials design is used to control drift, matrix effects, and information density. A clear developmental trend is the shift from simple dye printing on inert substrates toward structured hosts and interfaces (porous polymers, sol–gel matrices, MOFs/zeolites, and microfluidic architectures) that tune analyte transport, preconcentrate targets, and stabilize photophysics in the solid state. In parallel, chromophore selection has moved toward broader spectral coverage, higher photostability, and ratiometric designs to decouple signals from illumination and aging. The main challenges remain reproducibility (spot-to-spot and lot-to-lot loading), time dependence of non-covalent responses, and domain shift caused by humidity/temperature, camera/spectrometer differences, and real-matrix variability. Materials have progressively addressed these limitations through antifouling and low-adsorption supports, internal reference spots and calibration patches, barrier layers to manage water activity, and cartridge-style formats that fix geometry and exposure conditions. Finally, data processing has matured from qualitative pattern recognition to validation-oriented workflows—cross-validation, external holdouts, drift correction, and calibration transfer—so that performance is increasingly governed by standardized materials and metrology rather than by a single bespoke dataset.
A realistic assessment of array-based sensing must also acknowledge the operational risks that accompany its strengths. First, exceptionally low detection limits reported in some cases (e.g. in saliva) should be interpreted in the context of inter-individual variability, matrix heterogeneity, and the potential for false positives driven by confounders (diet, circadian effects, medication, oral microbiome, viscosity, pH/ionic strength, and sample handling). Although laboratory benchmarking against reference methods (e.g., ELISA for cortisol) supports feasibility, true clinical translation requires larger cohorts, repeated sampling, and multicentre validation to quantify precision, robustness, and decision thresholds under real-world variance. Second, metabolite-driven bacterial arrays—such as the S. aureus platform—provide rapid, amplification-free readouts but inherently measure a phenotypic signature rather than the organism itself. Thus, co-infections, strain-to-strain metabolic variability, antibiotic exposure, or inflammatory backgrounds could yield overlapping fingerprints and ambiguous classifications. This limitation can be mitigated by expanding training sets to include mixed cultures, clinically relevant confounders, and “near-neighbour” pathogens, and by incorporating out-of-distribution rejection so the model can abstain when a sample does not match any trained class. Third, calibration transfer remains a non-trivial barrier to multicenter deployment: differences in illumination geometry, sensor aging, batch-to-batch probe loading, camera/spectrometer spectral responses, and site-specific environmental conditions can shift the feature space enough to invalidate a model trained elsewhere. Overcoming this requires standardized acquisition hardware (or rigorous reference standards), internal controls on every array, harmonized SOPs for sampling and exposure, and formal calibration-transfer/domain-adaptation pipelines validated on external holdouts across sites. Taken together, these considerations do not negate the value of arrays; rather, they define the practical steps—cohort scale-up, confounder-aware training, and robust calibration governance—needed to move from compelling proof-of-concept demonstrations to reproducible, clinically and operationally deployable sensing systems.
Ideally, the community would benefit from reference materials and public datasets for array benchmarking: common probe panels, canonical matrices (saliva, fruit headspace, and environmental water) at agreed concentration ladders, and open pipelines for preprocessing and multivariate modelling. Lifecycle and accelerated-aging studies (shelf-life, humidity/temperature stress, and photostability) should become routine. Finally, to enable clinical and field adoption, arrays should be engineered for maintenance at the edge—automated QC, incremental recalibration, secure on-device analytics—and aligned with regulatory expectations (traceability, cybersecurity, and data privacy). With these steps, cross-reactive optical arrays can mature from compelling demonstrations to reliable, transferable tools for human health, environmental, and agri-food diagnostics.
An emerging interesting approach has been recently proposed by Cho and co-workers, in which the high-resolution camera images can be employed to obtain, from each single fluorescent spot, a set of data from each single pixel, in order to obtain an array of data from each single spot. This approach leads to higher sensitivity and selectivity.11 It requires an important support, from a hardware and software point of view, due to the hard machine learning process.
Over the next 3–5 years, progress should be focused on the concrete engineering and validation milestones rather than additional proof-of-concept targets. Technically, priorities include (i) cartridge-level standardization of substrates, probe loading, and internal reference spots; (ii) drift-resilient acquisition (fixed illumination geometry and temperature/humidity logging) with certified calibration-transfer across smartphones/fiber spectrometers; (iii) confounder-aware training sets that explicitly include mixtures, co-infections, and matrix extremes; and (iv) accelerated-aging and shelf-life qualification to support “no cold chain” deployment.
In terms of near-term translation, applications closest to commercialization are those with clear workflows and minimal regulatory burden—environmental/VOC monitoring and agro-food packaging surveillance—followed by low-risk clinical screening use-cases (e.g., salivary panels). For regulatory submission of an array-based diagnostic tool, the key requirements will be a locked assay, multi-site clinical validation with predefined endpoints, method comparison against gold standards, reproducibility and robustness studies, a traceable uncertainty budget, and software documentation consistent with clinical expectations.
Author contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript. R. S., A. C. and N. T. performed literature research and analysis. G. T. S. mainly worked on the manuscript design and writing, with all the authors contributing to the drafting and correction.Conflicts of interest
There are no conflicts to declare.List of acronyms
| HCA | Hierarchical cluster analysis |
| LDA | Linear discriminant analysis |
| SVM | Support vector machines |
| PLS | Partial least square |
| RGB | Red–green–blue |
| HSV | Hue–saturation–value |
| T | Temperature |
| RH | Relative humidity |
| SVR | Support vector regression |
| PCA | Principal component analysis |
| t-SNE | t-Distributed stochastic neighbor embedding |
| UMAP | Uniform manifold approximation and projection |
| QDA | Quadratic discriminant analysis |
| LOD | Limit of detection |
| LOQ | Limit of quantification |
| VOC | Volatile organic compound |
| ML | Machine learning |
| MOF | Metal–organic framework |
| RP-TLC | Reverse phase thin layer chromatography |
| QC | Quality control |
| QA | Quality assurance |
| PVDF | Poly(vinylidene fluoride) |
| PTFE | Polytetrafluoroethylene |
| PoC | Point of care |
| LDH | Lactate dehydrogenase |
| ALT | Alanine aminotransferase |
| AST | Aspartate aminotransferase |
| ISE | Ion-selective electrode |
| CFU per mL | Colony-forming units per milliliter |
| PDMS | Polydimethylsiloxane |
Data availability
No primary research results or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
G. T. S. and N. T. thank the ERMES project, funded under European Union's Horizon Europe EIC Pathfinder Open program, grant agreement no. 101185661. G. T. S. acknowledges his daughter for inspiring the perspective and the ‘learning’ framework adopted throughout.Notes and references
- L. Zheng and K. S. Suslick, Acc. Chem. Res., 2021, 54, 950–960 CrossRef PubMed.
- E. Butera, A. Zammataro, A. Pappalardo and G. Trusso Sfrazzetto, ChemPlusChem, 2021, 86, 681–695 CrossRef PubMed.
- Z. Li, J. R. Askim and K. S. Suslick, Chem. Rev., 2019, 119, 231–292 CrossRef CAS PubMed.
- X. Wang, T. Yang and J.-H. Wang, Sens. Diagn., 2024, 3, 1176–1197 Search PubMed.
- Y. J. Jang, K. Kim, O. G. Tsay, D. A. Atwood and D. G. Churchill, Chem. Rev., 2015, 115, PR1–PR76 CrossRef CAS PubMed.
- Z. Jin, W. Yim, M. Retout, E. Housel, W. Zhong, J. Zhou, M. S. Strano and J. V. Jokers, Chem. Soc. Rev., 2024, 53, 7681–7741 RSC.
- H.-Y. Li, S.-N. Zhao, S.-Q. Zang and J. Li, Chem. Soc. Rev., 2012, 49, 6364–6401 RSC.
- L. Uzun and A. P. F. Turner, Biosens. Bioelectron., 2016, 76, 131–144 CrossRef CAS.
- H. Yoo, H. Jo and S. S. Oh, Mater. Adv., 2020, 1, 2663–2687 RSC.
- M. R. Sambrook and S. Notman, Chem. Soc. Rev., 2013, 42, 9251–9267 RSC.
- S. S. Ahmed, M. A. Hossain, C.-W. Hsu and C.-H. Yang, Biosensors, 2024, 14, 516 CrossRef CAS PubMed.
- A. Rudnitskaya, Front. Chem., 2018, 6, 433 CrossRef CAS PubMed.
- Z. Li and K. S. Suslick, ACS Sens., 2016, 1, 1330–1335 CrossRef CAS.
- X. Wang, O. S. Wolfbeis and Z. Wang, Anal. Chem., 2019, 91, 623–640 Search PubMed.
- C. Tian, S. Shin, Y. Cho, Y. Song and S. Y. Cho, ACS Sens., 2024, 9, 5489–5499 CrossRef CAS PubMed.
- J. A. Mennella, Annu. Rev. Nutr., 2014, 34, 115–134 Search PubMed.
- M. Beija, C. A. M. Afonso and J. M. G. Martinho, Chem. Soc. Rev., 2009, 38, 2410–2433 RSC.
- T. Kowada, H. Maeda and K. Kikuchi, Chem. Soc. Rev., 2015, 44, 4953–4972 RSC.
- H. Zhu, C. Liu, M. Su, X. Rong, Y. Zhang, X. Wang, K. Wang, X. Li, Y. Yu, X. Zhang and B. Zhu, Coord. Chem. Rev., 2021, 448, 214153 CrossRef CAS.
- Y. Fan, Y. Wu, J. Hou, P. Wang, X. Peng and G. Ge, Coord. Chem. Rev., 2023, 480, 215020 CrossRef CAS.
- R. Santonocito, M. Intravaia, I. M. Caruso, A. Pappalardo, G. Trusso Sfrazzetto and N. Tuccitto, Nanoscale Adv., 2022, 4, 1926–1948 RSC.
- Y. L. Pak, Y. Wang and Q. Xu, Coord. Chem. Rev., 2021, 433, 213745 CrossRef CAS.
- A. K. Yetisen, M. S. Akram and C. R. Lowe, Lab Chip, 2013, 13, 2210–2251 RSC.
- A. J. Summers, J. P. Devadhasan, J. Gu, D. C. Montgomery, B. Fischer, M. A. Gates-Hollingsworth, K. J. Pflughoeft, T. Vo-Dinh, D. P. AuCoin and F. Zenhausern, ACS Omega, 2022, 7, 32262–32271 CrossRef CAS PubMed.
- N. Tuccitto, G. Catania, A. Pappalardo and G. Trusso Sfrazzetto, Chem. – Eur. J., 2021, 27, 13715–13718 CrossRef CAS PubMed.
- R. Puglisi, A. Cavallaro, A. Pappalardo, M. Petroselli, R. Santonocito and G. Trusso Sfrazzetto, Molecules, 2024, 29, 3714 CrossRef CAS.
- L. Zeni, C. Perri, N. Cennamo, F. Arcadio, G. D’Agostino, N. Salmona, M. Beeg and M. Gobbi, Sci. Rep., 2020, 10, 11154 CrossRef CAS PubMed.
- Y. Liu, H. Pu and D.-W. Sun, Trends Food Sci. Technol., 2017, 69, 25–35 CrossRef CAS.
- N. Nordin, S. N. A. Rahim, W. F. A. W. Omar, S. Zulkarnain, S. Sinha, S. Kumar and M. Haque, Cureus, 2024, 16, e57243 Search PubMed.
- A. M. Simundic, G. Baird, J. Cadamuro, S. J. Costelloe and G. Lippi, Crit. Rev. Clin. Lab. Sci., 2020, 57, 1–21 CrossRef CAS PubMed.
- R. Cano-Corres, G. Sole-Enrech and M. I. Aparicio-Calvente, Biochem. Med., 2023, 33, 020702 Search PubMed.
- C. D. Koch, M. A. Vera, J. Messina, N. Price, T. J. Durant and J. M. El-Khoury, Clin. Chim. Acta, 2021, 520, 63–66 CrossRef CAS PubMed.
- P. Vermeersch, G. Frans, A. von Meyer, S. Costelloe, G. Lippi and A. M. Simundic, Clin. Chem. Lab. Med., 2021, 59, 1047–1061 CrossRef CAS PubMed.
- M. A. Williams, A. A. Olomukoro, R. V. Emmons, N. H. Godage and E. Gionfriddo, J. Sep. Sci., 2023, 46, 2300571 CrossRef CAS PubMed.
- T. K. S. Ng, C. Udeh-Momoh, M.-A. Lim, H. S. Gleerup, W. Leifert, C. A. N. Ashton, H. Zetterberg, R. A. Rissman, C. N. Winston, S. O’Bryant, S. Jenkins, E. Carro, G. Orive, S. Tamburin, M. Olvera-Rojas, P. Solis-Urra, I. Esteban-Cornejo, G. A. A. Dos Santos, K. B. Rajan, D. Koh, A. H. Simonsen and P. D. Slowey, Alzheimer’s Dementia, 2025, 21, e14420 CrossRef PubMed.
- M. Shi, Y. T. Sui, E. R. Peskind, G. Li, H. J. Hwang, I. Devic, C. Ginghina, J. S. Edgar, C. Pan, D. R. Goodlett, A. R. Furay, L. F. Gonzalez-Cuyar and J. Zhang, J. Alzheimer's Dis., 2011, 27, 299–305 CAS.
- J. Marksteiner, M. Defrancesco and C. Humpel, Front. Aging Neurosci., 2022, 14, 1014305 CrossRef CAS PubMed.
- R. G. Schipper, E. Silletti and M. H. Vingerhoeds, Arch. Oral Biol., 2007, 52, 1114–1135 CrossRef CAS PubMed.
- W. Lu, X. Su, M. S. Klein, I. A. Lewis, O. Fiehn and J. D. Rabinowitz, Annu. Rev. Biochem., 2017, 86, 277–304 CrossRef CAS PubMed.
- J. D. Rabinowitz and E. Kimball, Anal. Chem., 2007, 79, 6167–6173 CrossRef CAS PubMed.
- D. Vuckovic, Anal. Bioanal. Chem., 2012, 403, 1523–1548 CrossRef CAS PubMed.
- W. Lu, M. F. Clasquin, E. Melamud, D. Amador-Noguez, A. A. Caudy and J. D. Rabinowitz, Anal. Chem., 2010, 82, 3212–3221 CrossRef CAS PubMed.
- T. Li, X. Zhu, X. Hai, S. Bi and X. Zhang, ACS Sens., 2023, 8, 994–1016 CrossRef CAS PubMed.
- X. Chen and Q. Wang, Anal. Chem., 2026, 98, 2633–2679 CrossRef CAS PubMed.
- M. A. Beatty, A. J. Selinger, Y. Li and F. Hof, J. Am. Chem. Soc., 2019, 141, 16763–16771 CrossRef CAS PubMed.
- F. Shahdost-Fard, A. Bigdeli and M. R. Hormozi-Nezhad, ACS Chem. Neurosci., 2021, 12, 3157–3166 CrossRef CAS PubMed.
- Y. Sasaki, X. Lyu, R. Kubota, S. Takizawa and T. Minami, ACS Appl. Bio Mater., 2021, 4, 2113–2119 CrossRef CAS PubMed.
- Y. Yu, W. Ni, Q. Hu, H. Li, Y. Zhang, X. Gao, L. Zhou, S. Zhang, S. Ma, Y. Zhang, H. Huang, F. Li and J. Han, Angew. Chem., Int. Ed., 2024, 63, e202318483 CrossRef CAS PubMed.
- W. Bai, R. Wu, Z. Xue, X. Song, Z. Xia, K. Zhang, K. Shao, Y. Li and T. Wang, Adv. Funct. Mater., 2024, 34, 2401421 CrossRef CAS.
- R. Santonocito, A. Cavallaro, F. Ficili, A. Distefano, G. Grasso, A. Pappalardo, N. Tuccitto and G. Trusso Sfrazzetto, Sens. Diagn., 2025, 4, 690–696 RSC.
- R. Santonocito, A. Cavallaro, A. Pappalardo, R. Puglisi, A. Marano, M. Andolina, N. Tuccitto and G. Trusso Sfrazzetto, Biosens. Bioelectron., 2025, 270, 116986 CrossRef CAS PubMed.
- R. Santonocito, S. Avnet, N. Tuccitto, N. Musso, A. Cavallaro, M. Spinello, M. V. Lipreri, M. Ricceri, I. M. P. Filannino, I. Raimondi, S. Stefani, N. Baldini and G. Trusso Sfrazzetto, Biosens. Bioelectron., 2026, 294, 118199 CrossRef CAS PubMed.
- M. Pushina, A. Penavic, S. Farshbaf and P. Anzenbacher, Jr, ACS Sens., 2021, 6, 4001–4008 CrossRef CAS PubMed.
- L. Li, S. Xu, X. Li, H. Gao, L. Yang and C. Jiang, Chem. Eng. J., 2024, 493, 152636 CrossRef CAS.
- Y. Li, Q. Jiang, Z. Li, S. Du and Y. Du, Microchim. Acta, 2025, 192, 738 CrossRef CAS PubMed.
- A. Cavallaro, R. Santonocito, A. Pappalardo, N. Tuccitto and G. Trusso Sfrazzetto, Sens. Actuators, B, 2026, 448, 138936 CrossRef CAS.
- A. Bowyer, C. Shen and E. J. New, Analyst, 2020, 145, 1195–1201 RSC.
- R. Santonocito, R. Parlascino, A. Cavallaro, R. Puglisi, A. Pappalardo, F. Aloi, A. Licciardello, N. Tuccitto, S. O. Cacciola and G. Trusso Sfrazzetto, Sens. Actuators, B, 2023, 393, 134305 CrossRef CAS.
- A. Cavallaro, R. Santonocito, R. Puglisi, A. Pappalardo, F. La Spada, R. Parlascino, M. Riolo, S. O. Cacciola, N. Tuccitto and G. Trusso Sfrazzetto, Chem. Commun., 2024, 60, 13702–13705 RSC.
- Z. You, M. Zhao, H. Lu, H. Chen and Y. Wang, ACS Appl. Mater. Interfaces, 2024, 16, 19359–19368 CrossRef CAS PubMed.
- M. K. Morsy, K. Zor, N. Kostesha, T. S. Alstrøm, A. Heiskanen, H. El-Tanahi, A. Sharoba, D. Papkovsky, J. Larsen, H. Khalaf, M. H. Jakobsen and J. Emneus, Food Control, 2016, 60, 346–352 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |