Open Access Article
Open Access ArticlePhospholipid asymmetry in biomimetic vesicles alters membrane permeability
Paige
Allard
 ab,
Alex R.
McDonald
ab,
Kaitlyn
Ramsay
ab,
Alex R.
McDonald
ab,
Kaitlyn
Ramsay
 ab and
Katherine S.
Elvira
ab and
Katherine S.
Elvira
 *ab
*ab
aDepartment of Chemistry, University of Victoria, Victoria, BC, Canada. E-mail: kelvira@uvic.ca
bCentre for Advanced Materials and Related Technology (CAMTEC), University of Victoria, Victoria, BC, Canada
First published on 9th April 2026
Abstract
Artificial cell membranes are rarely able to capture a key structural component of biological cells, namely the asymmetric distribution of phospholipids in the inner and outer leaflets. Here we show a straightforward method for generating biomimetic asymmetric vesicles that mimic the composition of human red blood cells using a plug-and-play microfluidic device. Small molecule diffusion was compared between symmetric and asymmetric vesicles using quinine hemisulphate. Asymmetric vesicles showed an 85% lower permeability than symmetric vesicles, suggesting that asymmetry is an important, yet undervalued, consideration in the design of vesicles for a variety of applications from the fundamental study of drug-membrane interactions to the use of vesicles for drug delivery.
Cell membranes are highly discriminatory barriers separating cellular compartments from the outside environment. The mammalian cell membrane is a bilayer composed of a heterogeneous and asymmetric distribution of phospholipids in the inner and outer leaflets, as well as other molecules such as cholesterol and proteins. This asymmetric distribution of phospholipids affects fundamental characteristics such as membrane potential, shape, surface charge and stability, with loss of asymmetry causing significant physiological consequences.1 For example, the externalization of phosphatidylserine (PS), a lipid which is primarily contained in the inner red blood cell leaflet, to the outer leaflet has been linked to blood coagulation and erythrocyte adhesion.1 The loss of asymmetry between bilayers can also be a marker of cancer cells, and we have used another type of artificial cell membrane (droplet interface bilayers, DIBs) to show that this can potentially contribute to chemoresistance.2 While there is some enticing prior work to show that lipid asymmetry in artificial vesicles alters lipid diffusion, packing and fluidity,3 as well as improving mRNA drug delivery,4 the link between lipid asymmetry and membrane permeability in vesicles is not clear.
Artificial vesicles are made of a phospholipid bilayer surrounding an aqueous core.5 They are commonly used as drug delivery vehicles, for example to prolong the release of drugs in the human body,6 and to investigate fundamental biophysical properties of human cell membranes,3 such as mechanisms of protein insertion.7 However, they are typically generated with a symmetric composition of synthetic phospholipids in the bilayer,8 limiting the biomimetic properties of the model membranes. The creation of asymmetric artificial vesicles is challenging because most manual techniques rely on spontaneous bilayer formation from phospholipids in a single solution of phospholipids.9,10 Asymmetric artificial vesicles can be produced in bulk from synthetic lipids using methods such as reverse phase evaporation, calcium ion induced flip-flop, cyclodextrin exchange and the use of enzymes to induce lipid flip-flop.8 However, bulk methods suffer from issues with monodispersity, unilamellarity and often result in only partial asymmetry.10 The use of microfluidic technologies for the formation of vesicles has enabled tighter control over lamellarity and vesicle size.5,11 However, the specialized expertise required to fabricate and operate microfluidic devices, along with the additional steps needed to generate asymmetric lipid bilayers, may explain why only a few studies report the use of microfluidic technologies to produce asymmetric vesicles.12–19
The plasma membrane of red blood cells is well characterized and displays a distinct asymmetric distribution of phosphatidylcholine (PC), phosphatidylethanolamine (PE), sphingomyelin (SM) and PS between the inner and outer leaflets. Although asymmetric vesicles have been used to investigate protein insertion,14 membrane fusion,20 and bending rigidity,21 direct quantitative comparisons of drug permeability between symmetric and asymmetric membranes of identical lipid composition are lacking. This represents a significant gap in our understanding of how membrane structure influences passive diffusion in cell-sized vesicles, with implications for drug delivery and predictive pharmacokinetic modeling.
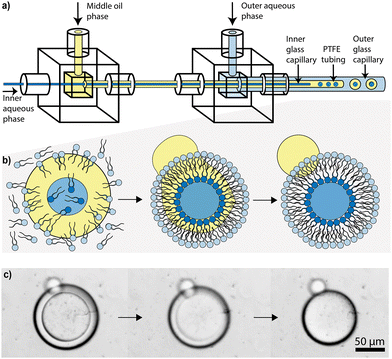
Here, we present a straightforward microfluidic method to make asymmetric vesicles from complex lipid mixtures that mimic the composition of eukaryotic cell membranes. To ensure accessibility for non-expert users, we used a plug-and-play microcapillary platform recently developed in our group22 and repurposed it for vesicle generation. We use parameters that replicate physiological conditions including pH, temperature, and scale, and demonstrate that our method fabricates asymmetric, unilamellar vesicles resembling red blood cell membranes, enabling quantification of passive diffusion. A common microfluidic method to form symmetric vesicles is via double emulsions, where one droplet is created inside another.23 Here, we form double emulsions consisting of three immiscible phases: an inner aqueous phase (dark blue, Fig. 1a and b) containing the lipid(s) for the inner leaflet; a middle oil-based phase (yellow, Fig. 1a and b) that promotes monolayer formation in the correct orientation; and an outer aqueous phase (light blue, Fig. 1a and b) containing the lipid(s) for the outer leaflet. The oil phase then de-wets, allowing the monolayers to assemble into a bilayer surrounding the aqueous core (Fig. 1c). The device consists of two junction boxes cast from flexible polyurethane resin using 3D-printed moulds (Fig. 1a). These boxes hold and align the three capillaries that generate the emulsions. The resin provides an effective seal around the capillaries without adhesives or sealants. The inner capillary is glass (ID = 0.20 mm, dark blue, Fig. 1a), the middle is polytetrafluoroethylene (PTFE) tubing (ID = 0.75 mm, yellow), and the outer is silane-treated glass (ID = 1.5 mm, light blue). The first junction box holds the inner and middle capillaries, enabling the formation of the initial water-in-oil droplets, while the second junction box introduces the outer aqueous phase to create water-in-oil-in-water double emulsions.
The inner and outer aqueous phases (dark and light blue respectively, Fig. 1b) contained the phospholipid mixtures for the inner and outer leaflets, and the middle oil phase was 50:50 v/v chloroform:hexanes (yellow, Fig. 1b). A syringe pump was used to dispense each solution at a flow rate of 50 µL min−1, 100 µL min−1 and 200 µL min−1 for the inner, middle and outer phases, respectively. Double emulsion formation was observed at an average rate of 1.4 ± 0.2 droplets per second. The double emulsions were then collected into 4-(2-hydroxyethyl)-1-piperazineethane sulfonic acid (HEPES) buffer at pH 7.4. The double emulsions de-wetted within 15 min of collection as the oil phase thinned sufficiently to cause a spontaneous rearrangement of the phospholipid molecules in the inner and outer phases (Fig. 1c).9
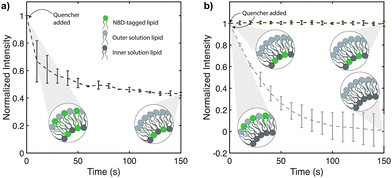
To quantitatively assess whether both unilamellar and asymmetric vesicles were formed, we used fluorescence quenching assays to verify that their structure mimics a natural lipid bilayer in terms of lamellarity and asymmetry.24 For these assays, we used similar synthetic phospholipids as prior work as a proof of concept.25 Vesicles were formed with 100% 1-palmitoyl-2-oleoyl-glycero-3-phosphocholine (POPC) in the outer leaflet and 75% 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) with 25% POPC in the inner leaflet, since this ratio reduces phospholipid flip-flop and loss of asymmetry caused by differing lipid shapes.26 Green-tagged fluorescent phospholipids (NBD-PC, 1 mol%) were added to both leaflets (to prove unilamellarity) or to either the inner or the outer leaflet (to prove asymmetry). In both cases, a quenching solution (100 mM sodium hydrosulphite in 1 M Tris, pH 10) was added to the outside of the vesicles. The quencher reduces NBD-tagged phospholipids, extinguishing the fluorescence. Since the quencher cannot diffuse across the bilayer,24 only NBD-tagged phospholipids in the outer bilayer are extinguished. Vesicles were first imaged 2 h after formation to ensure complete de-wetting. Then ∼75 µL of quencher was added, and images were taken every 10 s for 150 s.
The lamellarity of the vesicles was determined by adding green fluorescent NBD-tagged phospholipids to both the inner and outer leaflets. Following the addition of the quencher to the outside of the vesicles, fluorescence was reduced to 50% (Fig. 2a).27 If more than 50% fluorescence signal had remained following quenching, then the vesicles would be multilamellar because more bilayers containing green-tagged lipids would be present inside the vesicle.18,27 Proof of the asymmetry of the vesicles was determined by adding green fluorescent NBD-tagged phospholipids first only to the inner leaflet. NBD-tagged lipids showed no reduction in fluorescence after adding the quencher (Fig. 2b, dark grey data). This indicates that the inner solution phospholipids were indeed entirely present in the inner leaflet. Texas Red-tagged lipids were added to the outer solution to control for photobleaching and also showed no decrease in fluorescence (Fig. S1, SI). Vesicles were then generated with green fluorescent NBD-tagged phospholipids present only in the outer leaflet. Texas Red-tagged lipids were added to the inner solution to show that both lipid solutions are incorporated into the vesicles (Fig. S2, SI). NBD-tagged lipid fluorescence showed a total reduction in fluorescence after adding the quencher (Fig. 2b, light grey data). In addition, an Annexin V binding assay was performed which confirmed the expected leaflet-specific localization of PS (Fig. S3, SI). Both assays show that the outer solution phospholipids were assembled in the outer leaflet, confirming that the vesicles are asymmetric.
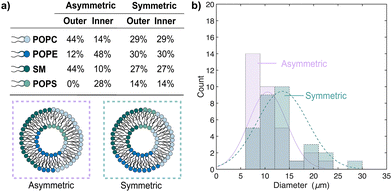
We then generated asymmetric vesicles designed to mimic the membrane of red blood cells using phospholipid mixtures similar to those found in mammalian red blood cells.28 Cholesterol was not included since it would shift permeability in the same manner in both symmetric and asymmetric vesicles. Four lipids (POPC, POPE, SM and 1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-L-serine, POPS) were added to the inner or outer phases depending on the ratio of their distribution in red blood cell membranes (Fig. 3a).29,30 To assess and demonstrate the impact of asymmetry in phospholipid composition on permeability, we also prepared vesicles using the same phospholipids but equally distributed in both the inner and outer leaflets. We defined this as mean mol% of each phospholipid added to the inner and outer leaflets (Fig. 3a).2
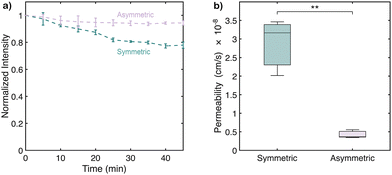
Asymmetric and symmetric vesicles were generated using the microcapillary device with the same conditions described previously. The asymmetric vesicles had an average size (diameter) of 11 ± 4 µm (lilac data, Fig. 3b), while the symmetric vesicle sizes had an average size of 14 ± 5 µm (teal data, Fig. 3b). The observed size difference is significant (p = 0.0103, unpaired t-test) and may arise from differences in membrane curvature stress and lipid packing,2,31 as lipid shape contributes to the spontaneous curvature.32 The larger proportion of conical PE33 in the inner leaflet would drive the formation of a smaller core of the vesicle. To assess the impact of asymmetry on permeability, quinine hemisulphate, a membrane-permeable anti-malaria drug, was loaded into the vesicles by adding it to the inner solution during vesicle formation with an average encapsulation efficiency of 94 ± 4% (Fig. S4 and Table S1, SI). The fluorescence intensity was monitored over time by imaging the vesicles every 5 min for 45 min, with the intrinsic fluorescence of quinine serving as a reporter of the vesicle drug concentration. Normalized fluorescence over time (Fig. 4a) revealed a more rapid signal loss in symmetric vesicles (teal data) compared to asymmetric ones (purple data). We calculated permeability coefficients and found symmetric vesicles exhibited a mean permeability of 2.88 × 10−8 cm s−1, while asymmetric vesicles showed a significantly lower permeability of 4.29 × 10−9 cm s−1 (p = 0.0053, Fig. 4b). While the symmetric vesicles were slightly larger, the calculated permeability values remain directly comparable as the equations account for differences in the bilayer area. This corresponds to an 85% reduction in quinine permeability due to lipid asymmetry, suggesting that passive diffusion of small molecules can be dependent on the location of the lipids within the membrane, even when the overall lipid composition remains constant.
The lower permeability observed in asymmetric vesicles may be due to tighter lipid packing in one or both leaflets, or reduced fluidity caused by interleaflet coupling.32 Additionally, a higher amount of SM in the outer leaflet could allow for more rigid and tightly packed lipids, which may contribute to reduced permeability due to the formation of tightly packed lipid rafts.34 Similar effects have been reported in previous studies, where planar asymmetric membranes exhibited lower passive diffusion rates due to altered lipid dynamics and increased order.35 Recent work has shown that red blood cells which had their lipid asymmetry scrambled to become symmetric were 35% more permeable to a fluorescent dye than naturally occurring asymmetric red blood cells.36 Our data are in line with these findings, suggesting we can recreate the effect asymmetry has on small molecule permeation in natural cells in our artificial vesicles. Further investigation into the exact mechanism is necessary in future work.
Our findings highlight the critical role of membrane structure (asymmetry) in regulating small molecule permeability. This underscores the importance of lipid organisation in designing model membranes that more accurately reflect the structure and function of biological systems. A major limitation in the field has been the technical difficulty of producing asymmetric membranes. Our microcapillary-based method offers a significant advantage as it is accessible to non-specialists, thus lowering the barrier to wider adoption. The ability to easily recreate key features of natural membranes, such as lipid asymmetry, in artificial vesicles will enable their use as more physiologically relevant models for studying membrane dynamics, drug transport, and disease-related membrane dysfunction, as well as introducing an additional consideration in the design of vesicles for drug delivery.
Conflicts of interest
There are no conflicts to declare.Data availability
The data presented in this study are included in the article and supplementary information (SI). Further inquiries can be made to the corresponding author. Supplementary information: photobleaching and asymmetry controls, quinine encapsulation efficiency data, materials and methods, author contributions and acknowledgements. See DOI: https://doi.org/10.1039/d5cc06115f.References
- A. Sathi, V. Viswanad, T. P. Aneesh and B. A. Kumar, J. Pharm. BioAllied Sci., 2014, 6, 81–85 CrossRef PubMed.
- E. B. Stephenson and K. S. Elvira, Chem. Commun., 2021, 57, 6534–6537 RSC.
- S. Chiantia, P. Schwille, A. S. Klymchenko and E. London, Biophys. J., 2011, 100, L1–3 CrossRef CAS PubMed.
- C. Yang, J. Menge, N. Zhvania, M. Yu, H. Yang, D. Chen, Z. Zheng, D. A. Weitz and K. Jahnke, Adv. Funct. Mater., 2025, 35, 2505738 CrossRef CAS.
- D. Carugo, E. Bottaro, J. Owen, E. Stride and C. Nastruzzi, Sci. Rep., 2016, 6, 25876 CrossRef CAS PubMed.
- M. Alavi, N. Karimi and M. Safaei, Adv. Pharm. Bull., 2017, 7, 3–9 CrossRef CAS PubMed.
- V. L. Perillo, D. A. Peñalva, A. J. Vitale, F. J. Barrantes and S. S. Antollini, Arch. Biochem. Biophys., 2016, 591, 76–86 CrossRef CAS PubMed.
- Y. N. Al Badri, C. S. Chaw and A. A. Elkordy, Pharmaceutics, 2023, 15, 294 CrossRef CAS PubMed.
- N.-N. Deng, M. Yelleswarapu and W. T. S. Huck, J. Am. Chem. Soc., 2016, 138, 7584–7591 CrossRef CAS PubMed.
- D. Gardea-Gutiérrez, E. Núñez-García, B. E. Oseguera-Guerra, M. Román-Aguirre and S. L. Montes-Fonseca, Pharmaceuticals, 2023, 16, 777 CrossRef PubMed.
- M. Krompers and H. Heerklotz, Membranes, 2023, 13, 267 CrossRef CAS PubMed.
- M. Doktorova, F. A. Heberle, B. Eicher, R. Standaert, J. Katsaras, E. London, G. Pabst and D. Marquardt, Nat. Protoc., 2018, 13, 2086–2101 CrossRef CAS PubMed.
- P. C. Hu, S. Li and N. Malmstadt, ACS Appl. Mater. Interfaces, 2011, 3, 1434–1440 CrossRef CAS PubMed.
- K. Kamiya, R. Kawano, T. Osaki, K. Akiyoshi and S. Takeuchi, Nat. Chem., 2016, 8, 881–889 CrossRef CAS PubMed.
- K. Karamdad, R. V. Law, J. M. Seddon, N. J. Brooks and O. Ces, Chem. Commun., 2016, 52, 5277–5280 RSC.
- L. Lu, W. J. Doak, J. W. Schertzer and P. R. Chiarot, Soft Matter, 2016, 12, 7521–7528 RSC.
- S. Maktabi, N. Malmstadt, J. W. Schertzer and P. R. Chiarot, Biomicrofluidics, 2021, 15, 024112 CrossRef CAS PubMed.
- S. Matosevic and B. M. Paegel, Nat. Chem., 2013, 5, 958–963 CrossRef CAS PubMed.
- V. Romanov, J. McCullough, B. K. Gale and A. Frost, Adv. Biosyst., 2019, 3, 1900010 CrossRef PubMed.
- M. Arribas Perez and P. A. Beales, Biophys. J., 2023, 122, 1985–1995 CrossRef CAS PubMed.
- Y. Elani, S. Purushothaman, P. J. Booth, J. M. Seddon, N. J. Brooks, R. V. Law and O. Ces, Chem. Commun., 2015, 51, 6976–6979 RSC.
- S. Farley, K. Ramsay and K. S. Elvira, Lab Chip, 2021, 21, 2781–2790 RSC.
- C. Has and P. Sunthar, J. Liposome Res., 2020, 30, 336–365 CrossRef CAS PubMed.
- J. C. McIntyre and R. G. Sleight, Biochemistry, 1991, 30, 11819–11827 CrossRef CAS PubMed.
- L. Lu, J. W. Schertzer and P. R. Chiarot, Lab Chip, 2015, 15, 3591–3599 RSC.
- K. K. Ewert, P. Scodeller, L. Simón-Gracia, V. M. Steffes, E. A. Wonder, T. Teesalu and C. R. Safinya, Pharmaceutics, 2021, 13, 1365 CrossRef CAS PubMed.
- E. C. Heider, M. Barhoum, K. Edwards, K.-H. Gericke and J. M. Harris, Anal. Chem., 2011, 83, 4909–4915 CrossRef CAS PubMed.
- P. van Hoogevest and A. Wendel, Eur. J. Lipid Sci. Technol., 2014, 116, 1088–1107 CrossRef PubMed.
- P. W. Dijck, E. J. Zoelen, R. Seldenrijk, L. L. Deenen and J. Gier, Chem. Phys. Lipids, 1976, 17, 336–343 CrossRef CAS PubMed.
- I. Bernhardt and L. Kaestner, Front. Biosci., 2025, 30, 25331 CrossRef CAS PubMed.
- W. Helfrich, Z. Naturforsch. C, 1973, 28, 693–703 CrossRef CAS PubMed.
- A. Hossein and M. Deserno, Biophys. J., 2020, 118, 624–642 CrossRef CAS PubMed.
- F. M. Goñi, Biochem. Biophys. Res. Commun., 2022, 633, 23–25 CrossRef PubMed.
- B. Ramstedt and J. P. Slotte, FEBS Lett., 2002, 531, 33–37 CrossRef CAS PubMed.
- B. Kollmitzer, P. Heftberger, M. Rappolt and G. Pabst, Soft Matter, 2013, 9, 10877–10884 RSC.
- M. Doktorova, J. L. Symons, X. Zhang, H.-Y. Wang, J. Schlegel, J. H. Lorent, F. A. Heberle, E. Sezgin, E. Lyman, K. R. Levental and I. Levental, Cell, 2025, 188, 2586–2602.e24 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |