Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Repression of bacterial gene expression by antivitamin B12 binding to a cobalamin riboswitch
Florian J.
Widner
a,
Naziyat I.
Khan
b,
Evelyne
Deery
 b,
Martin J.
Warren
*bc,
Michiko E.
Taga
b,
Martin J.
Warren
*bc,
Michiko E.
Taga
 *d and
Bernhard
Kräutler
*d and
Bernhard
Kräutler
 *a
*a
aInstitute of Organic Chemistry & Center for Molecular Biosciences (CMBI), University of Innsbruck, 6020, Innsbruck, Austria. E-mail: bernhard.kraeutler@uibk.ac.at
bSchool of Biosciences, University of Kent, Canterbury, CT2 7NJ, UK
cQuadram Institute Bioscience, Norwich Research Park, Norwich, NR4 7UA, UK. E-mail: martin.warren@quadram.ac.uk
dDepartment of Plant and Microbial Biology, University of California, Berkeley, California, USA. E-mail: taga@berkeley.edu
First published on 26th January 2026
Abstract
The E. coli btuB riboswitch is a cobalamin-sensing RNA element that selectively binds coenzyme B12 (adenosylcobalamin, AdoCbl) to downregulate the expression of the outer membrane B12-transporter BtuB. Here, we examined adenosylrhodibalamin (AdoRhbl), the isostructural Rh-analogue of AdoCbl, as a surrogate effector ligand for this riboswitch. Two riboswitch-reporter systems were employed: an engineered E. coli strain with a fluorescent reporter for intracellular AdoCbl-sensing, and a plasmid-based construct for analogous in vitro transcription/translation assays. In the in-vitro system AdoRhbl closely mimicked AdoCbl in down-regulating reporter expression with apparent EC50 values of 2.8 µM and 0.8 µM respectively. In contrast, the engineered E. coli strain revealed much higher effective sensitivities, with EC50 values of 1.4 nM for AdoRhbl and of 6.9 nM for AdoCbl, reflecting strong intracellular accumulation of both corrinoids, and comparably efficient uptake. These findings uncover a previously undocumented gene-regulatory activity of an antivitamin, suggesting that AdoRhbl can repress bacterial B12 uptake by binding to the btuB riboswitch. Together with its ability to inhibit AdoCbl-dependent enzymes, the designed antivitamin B12AdoRhbl thus emerges as a multifunctional antibiotic candidate targeting B12-utilizing microorganisms.
Introduction
Genetic regulation by bacterial riboswitches, first discovered at the turn of this century,1,2 has opened a valuable biological toolbox for controlling transcription, translation, and RNA splicing in microorganisms.3 Riboswitches are widespread regulatory mRNA elements that typically consist of two functional domains: an aptamer domain that selectively binds a cognate metabolite, and an expression platform that undergoes a structural rearrangement in response to ligand binding, thereby modulating gene expression.4 The first riboswitch identified was the cobalamin-responsive btuB riboswitch of Escherichia coli (E. coli), which binds coenzyme B12 (adenosylcobalamin, AdoCbl) with high selectivity,1 hence typified as a class-I cobalamin (Cbl) riboswitch.5 AdoCbl-binding triggers a conformational rearrangement that sequesters the ribosomal binding site, thereby repressing translation of the outer membrane B12-uptake protein BtuB.6 In-line probing demonstrated that the 202-nucleotide-long btuB riboswitch undergoes this structural shift not only in the presence of AdoCbl but also with vitamin B12 (cyanocobalamin, CNCbl), albeit with markedly reduced affinity for CNCbl.7 Class-II Cbl-riboswitches, in contrast, preferentially recognise Cbl derivatives with less bulky ‘upper’ axial ligands, such as methylcobalamin (MeCbl).8 Moreover, some riboswitches can also sense cobamides (Cbas) other than the canonical Cbls, thus Cbl-riboswitches may be considered an important subgroup of the wider class of B12 riboswitches7 or corrinoid riboswitches.9Genetic regulation through B12 riboswitches in many bacteria reflects their fundamental dependence on the complex natural cobamides as indispensable biocatalyst molecules.10–14 Only a subset of these microorganisms possess the complete B12-biosynthetic pathway for vitamin B12,15 while others rely on external sources and scavenge corrinoids from the environment.16–19 In order to adjust its metabolism to the availability of extracellular cobalt-corrinoids, E. coli employs a B12-responsive regulatory system20 centred on the btuB riboswitch.1 This riboswitch plays a vital function by controlling expression of the outer membrane transporter BtuB, which mediates B12 uptake.21,22
Our interest in the structural basis of btuB riboswitch selectivity for corrinoid ligands7,9 was been renewed by recent studies on the biological effects of antivitamins B12 (aVitB12s).23–25 These compounds were originally designed as metabolically inert structural mimics of vitamin B12, i.e., as type-I aVitB12s,23 to induce functional B12-deficiency in humans and other mammals, by impairing the cellular supply of active B12-cofactors.26 In fact, we also anticipated type-I aVitB12s to impair a broad spectrum of B12-dependent cellular processes and to act as bacterial growth inhibitors,27,28 like some other B12-antimetabolites.29–31
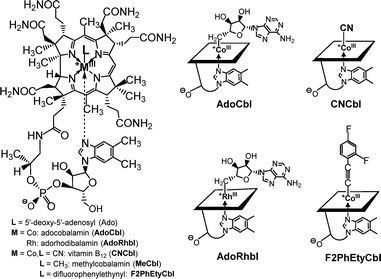
Among the aVitB12s, the Rh-analogue of AdoCbl, adenosylrhodibalamin (AdoRhbl, Fig. 1), stands out, as it is a nearly perfect structural mimic of AdoCbl25 that lacks its essential chemical reactivity.28,32AdoRhbl not only resists the critical tailoring catalysed by the (human) enzyme CblC,32 but also potently inhibits bacterial AdoCbl-dependent diol-dehydratase,25 as well as bacterial and human adenosyltransferases.32 It also acts as a light-stable anti-photoregulatory ligand of the widely occurring bacterial CarH photoreceptors.33 The particular activity of AdoRhbl as an effective growth-repressor of Salmonella typhimurum has raised the question of whether this involves gene regulation via a Cbl-riboswitch,25 a mechanism proposed as a central mode of activity of proper type-I aVitB12s.28 Because AdoRhbl closely reproduces the molecular shape and surface functionalities of AdoCbl, it is expected to match the natural coenzyme in binding very tightly to class-I Cbl-riboswitches, and in inducing the same conformational changes and downstream regulatory effects. In fact, binding of AdoRhbl to the btuB-riboswitch would provide a false-positive signal of intracellular AdoCbl availability, leading to repression of BtuB production, and thereby blocking import of natural B12-derivatives, effectively inducing growth-inhibiting Cbl-deficiency.28 A natural antibiotic, named roseoflavin, plays a related role by binding flavine-mononucleotide (FMN) riboswitches34 and inhibiting bacterial growth.35
Results and discussion
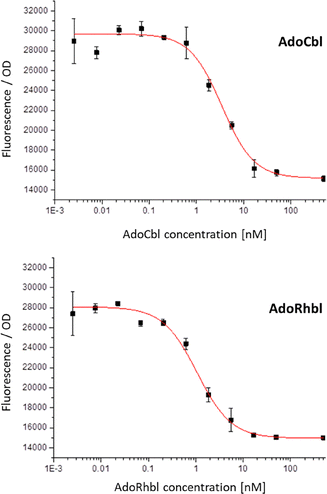
We report here a comparison of the aVitB12AdoRhbl25 and of the natural effector ligand AdoCbl in their capacity of binding to the btuB riboswitch and regulating protein expression. For this investigation, we employed two RNA AdoCbl-reporter constructs derived from the E. coli btuB-riboswitch.1,6 Both constructs (see SI) were designed to provide fluorescence readouts in response to riboswitch-mediated regulation by B12-type ligands. Because the isolated (originally used) 240 nt btuB riboswitch sequence alone proved insufficient to drive reporter gene expression20,36,37 the constructs used here included the promoter, the aptamer, the expression platform and the first 70 codons (210 nucleotides) of the btuB coding sequence. Fluorescent protein reporters, red fluorescent protein (RFP) or enhanced green fluorescent protein (eGFP)38 were fused downstream. The constructs were incorporated either into the genome of an engineered E. coli strain9 for in-vivo sensing of AdoCbl, or within a correspondingly constructed E. coli plasmid, designed for in-vitro assays monitoring fluorescence.The aVitB12AdoRhbl25 was produced for these studies via a one-step synthesis from chlororhodibalamin.39 As shown here, AdoRhbl acts as a highly potent and specific ligand for the AdoCbl-sensitive btuB riboswitch. To investigate intracellular sensing of AdoCbl and AdoRhbl, we employed E.coli MG1655 harboring a pUC19 plasmid containing a BtuB-RFP fusion (pUC19-BtuB-RFP),9 in which RFP expression is regulated by the btuB B12-riboswitch (for details, see SI).40 Bacterial cultures were diluted and exposed to either AdoCbl or AdoRhbl, over a concentration range from 2.54 pM to 0.5 µM. Upon light-protected incubation for 16 h at 37 °C, RFP-fluorescence was determined at 590 nm. In both cases, fluorescence decreased in a concentration-dependent manner, with reproducible results in two independent experimental series. These data indicate efficient binding of the riboswitch by either corrin, with a critical transition observed at low nM concentrations (Fig. 2).
Curve fitting with a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding isotherm model provided effective EC50s of 1.4 nM (+/−0.6 nM) for the Rh-corrin AdoRhbl and 6.9 nM (+/−4 nM) for the homologous Co-corrin AdoCbl. Both the natural B12-cofactor AdoCbl and its isostructural Rh-analogue AdoRhbl efficiently triggered switching of the btuB-riboswitch construct at remarkably low nM concentrations in the medium. The high intracellular sensitivity of this class-I Cbl-riboswitch to either organometallic Ado-corrinoid is consistent with the well-documented extensive accumulation of AdoCbl in bacteria,40,41 and is matched by the designed Rh-corrin AdoRhbl. Notably, the conformational switch induced by AdoRhbl occurs at roughly 5-fold lower extracellular concentrations than with the natural cognate ligand AdoCbl. This apparent difference does not reflect a higher intrinsic binding affinity of the riboswitch for the aVitB12AdoRhbl, which was not observed in in-vitro experiments (see below). Rather, it likely arises because a fraction of the internalized AdoCbl is diverted into metabolism, serving as a precursor for other Cbls, such as methylcobalamin (MeCbl),41 which bind the class-I Cbl- riboswitch btuB with lower affinity.42
1 binding isotherm model provided effective EC50s of 1.4 nM (+/−0.6 nM) for the Rh-corrin AdoRhbl and 6.9 nM (+/−4 nM) for the homologous Co-corrin AdoCbl. Both the natural B12-cofactor AdoCbl and its isostructural Rh-analogue AdoRhbl efficiently triggered switching of the btuB-riboswitch construct at remarkably low nM concentrations in the medium. The high intracellular sensitivity of this class-I Cbl-riboswitch to either organometallic Ado-corrinoid is consistent with the well-documented extensive accumulation of AdoCbl in bacteria,40,41 and is matched by the designed Rh-corrin AdoRhbl. Notably, the conformational switch induced by AdoRhbl occurs at roughly 5-fold lower extracellular concentrations than with the natural cognate ligand AdoCbl. This apparent difference does not reflect a higher intrinsic binding affinity of the riboswitch for the aVitB12AdoRhbl, which was not observed in in-vitro experiments (see below). Rather, it likely arises because a fraction of the internalized AdoCbl is diverted into metabolism, serving as a precursor for other Cbls, such as methylcobalamin (MeCbl),41 which bind the class-I Cbl- riboswitch btuB with lower affinity.42
The strong intracellular sensing of AdoCbl and AdoRhbl by the btuB riboswitch encouraged us to examine binding properties of a more readily accessible and robust Cbl-based aVitB12, the alkynylcobalamin 2(2,4-difluorophenyl)ethynyl-cobalamin (F2PhEtyCbl)43 (see SI). At solution concentrations above 10 nM, F2PhEtyCbl reproducibly reduced fluorescence in E. coli cells carrying the pUC19-BtuB-RFP construct, consistent with riboswitch downregulation of RFP expression. Data fitting over the concentration range of 0.2 nM to >104 nM yielded an apparent EC50 of about 180 nM (see SI, Fig. S1). The roughly 100-fold weaker riboswitch binding affinity of F2PhEtyCbl relative to AdoCbl or AdoRhbl reflects the known sensitivity of class-I Cbl- riboswitches to the structure of the cofactor's ‘upper’ ligand,7 and is in line with the reported lack of evidence for effective binding of structurally related alkynylcobalamins to the E. coli btuB-riboswitch.44
In the alternative ‘in-vitro’ experiments with the E.coli btuB riboswitch a corresponding test system based on an E. coli in-vitro transcription/translation kit was used (see SI). A key advantage of this ‘in-vitro’ setup is that it directly reports on interactions between the riboswitch and the effector corrinoids, thereby allowing determination of the riboswitch response to the solution concentration of the supplied ligand. In the context of this study, these data provided a basis for estimating how intracellular accumulation of corrinoids influences in vivo riboswitch response observed in the engineered E. coli strain.
For the ‘in-vitro’ system, the riboswitch construct was generated within a pET14b plasmid carrying the gene for eGFP under the control of a T7 promoter. Using specific primers, the functional btuB riboswitch segment was PCR amplified and inserted upstream of the eGFP coding sequence, yielding Rib70-eGFP-pET14b (described in the SI, Fig. S2 and Table S1). This design positioned the riboswitch correctly in-frame between the T7 promoter and the eGFP reporter gene, enabling riboswitch-dependent regulation of eGFP expression.
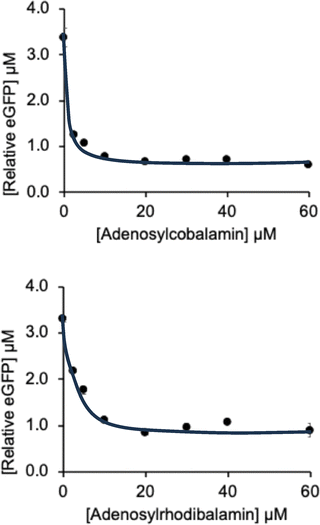
To confirm that Rib70-eGFP-pET14b supported reporter expression, the plasmid was transformed into E. coli BL21*(DE3)-pLysS and cultured in M9 minimal medium. The resulting cell pellets appeared green, demonstrating successful eGFP production. To assess riboswitch control and activity in response to AdoCbl, in vitro protein synthesis experiments were performed using the NEB PURExpress transcription/translation kit (see SI). Riboswitch function was evaluated by quantifying eGFP production, both via fluorescence intensity measurements (Fig. 3), and by densitometry of western blots (SI, Fig. S3). To examine the regulatory response to AdoCbl and AdoRhbl, in vitro reactions were conducted across a ligand concentration range of 0-60 µM. The resulting values were analyzed by curve-fitting using 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 isotherm models (Fig. 3 and SI Fig. S3).
1 isotherm models (Fig. 3 and SI Fig. S3).
Since both AdoCbl and AdoRhbl are colored at higher concentrations, their potential interference with eGFP fluorescence was examined. Calibration curves were generated by recording eGFP fluorescence at known concentrations in the presence of varying amounts of AdoCbl or AdoRhbl. Comparable fluorescence intensities were obtained in samples with and without corrinoids, confirming that neither ligand significantly affected the eGFP fluorescence signal.
Calibration curves generated with defined concentrations of AdoCbl were used to relate fluorescence intensities to relative eGFP levels in each in vitro reaction. To validate these measurements, western blotting followed by densitometric analysis was performed, comparing band intensities of the reaction samples with eGFP standards of known concentration. This dual approach enabled accurate determination of relative eGFP levels across the samples. The combined fluorescence and densitometry datasets were then used to plot the effect of increasing AdoCbl or AdoRhbl concentrations on eGFP production under btuB riboswitch control (Fig. 3 and SI, Fig. S3).
Our findings demonstrate that both AdoCbl and AdoRhbl bind to and repress the btuB riboswitch, as evidenced by a concentration dependent decrease in eGFP production between 2.5 µM to 60 µM. The similar response profiles indicate that AdoCbl and AdoRhbl exhibit comparably strong binding affinities for the riboswitch, which is consistent with their close structural similarity.25 Interestingly, neither ligand completely switches off the riboswitch activity: even at 60 µM ligand concentration, residual eGFP production persisted. Quantitative analysis suggests that both AdoCbl and AdoRhbl reduce translation by about 80% (SI Table S2). This incomplete repression likely reflects suboptimal folding of this structurally complex riboswitch in the course of AdoCbl binding, permitting constitutive low-level protein production, presumably a striking physiological safeguard against Cbl-deficiency.
To confirm that the observed reduction in eGFP production was specifically mediated through riboswitch binding, rather than interference with the in vitro translation system, we employed a control plasmid (eGFP-pET14b) lacking the riboswitch sequence. Under identical conditions, eGFP expression from this construct was unaffected by either AdoCbl or AdoRhbl, confirming that both ligands act solely through riboswitch-dependent regulation.
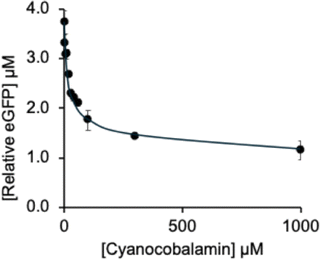
The plasmid Rib70-eGFP-pET14b carrying the “full-length” btuB riboswitch was also employed for in vitro assays with vitamin B12 (CNCbl). These experiments confirmed CNCbl binding to the btuB riboswitch, leading to reduced eGFP expression, as reflected in the concentration-dependent decline in fluorescence (Fig. 4 and SI, Fig. S4). However, CNCbl repressed expression only partially, with fluorescence measurements indicating a maximum deactivation of about 63% (SI, Table S2). Half-maximal deactivation was reached at 31.4 µM according to fluorescence data, and at 77.3 µM based on densitometric analysis. These results demonstrate that CNCbl has a lower affinity for the btuB riboswitch and does not achieve the same level of repression as AdoCbl, in qualitative agreement with earlier in-line probing studies.7
Our results show that the E. coli btuB B12-riboswitch discriminates only minimally between its natural effector ligand AdoCbl and the iso-structural aVitB12AdoRhbl, in which the Co(III)-centre of AdoCbl is replaced by Rh(III).25In vitro fluorescence and densitometry analyses with the eGFP/btuB reporter construct yielded EC50 values of 800 nM and 3.1 µM for AdoCbl, and 2.8 µM and 1.9 µM for AdoRhbl, respectively (Fig. 3 and SI, Fig. S3). The nearly identical regulatory effects of AdoCbl and AdoRhbl reflect their close structural similarity, with AdoCbl being the cognate ligand of the class-I Cbl-riboswitch. These EC50 values are broadly consistent with KD-values obtained by in-line probing experiments of the isolated 202 nucleotide btuB riboswitch, for which the somewhat lower KD values of 300 nM6 and 89 nM7 were reported, possibly reflecting (in part) differences between mere riboswitch binding and its actual effect on gene-expression.
Strikingly, in vivo experiments using an engineered E. coli MG1655 strain carrying the intracellular sensor revealed EC50 values nearly three orders of magnitude lower: 6.9 nM for AdoCbl and 1.4 nM for AdoRhbl (Fig. 2). These values are comparable to previous measurements with related E. coli riboswitch sensors for AdoCbl,40 and confirm the highly effective binding of AdoRhbl to the btuB riboswitch in vivo. The strong intracellular signal induced by externally supplied AdoRhbl indicates that this non-natural B12-mimic is imported into bacterial cells with similar efficiency to AdoCbl. The apparent increase in sensitivity in vivo likely reflects the substantial intracellular accumulation of Cbls in growing E. coli, estimated to reach approximately 1000-fold higher than in the growth medium.40,41 Indeed, the similar BtuB-dependent intracellular accumulation of AdoRhbl and AdoCbl, recently documented in both E.coli and Myxococcus xanthus,33 is consistent with the ratios of their deduced KD-values in vivo versus in vitro.
In contrast to AdoRhbl, which binds the E. coli btuB riboswitch with affinities comparable to the natural ligand AdoCbl, the more readily available Cbl-based aVitB12F2PhEtyCbl43 exhibits roughly 100 times lower affinity with an intracellular EC50 of 180 nM, qualitatively consistent with its ‘in-vitro’ EC50 of about 340 µM (see SI, Fig. S4b). A similar reduction in affinity is expected for other stable Cbl-based aVitB12s carrying alternative ‘upper’ axial ligands. Indeed, a hardly detectable btuB riboswitch binding has been reported for several such non-natural Cbls,44 consistent with the established preference of class-I Cbl-riboswitches for AdoCbl over other natural corrinoids.1,7
The precise structural basis for AdoCbl recognition by the btuB riboswitch remains unresolved, and high-resolution three-dimensional structures of B12-riboswitches are still scarce.5,8,45,46 Nonetheless, the crystal structure of a 172 nucleotide class-I Cbl-riboswitch from Symbiobacterium thermophilum revealed a B12-binding pocket with a well-defined complementarity to the ‘upper’ Ado-ligand, providing a rationale for its strong selectivity towards AdoCbl.45 A comparable architecture has been described for the AdoCbl-selective riboswitch from Thermoanaerobacter tengcongensis.8 By contrast, the env8 riboswitch displays a roughly 104-fold preference for MeCbl over AdoCbl.42 Structural analysis showed that its narrower B12-binding cavity restricts accommodation of the bulky Ado-group while permitting favourable interactions with smaller ‘upper’ axial ligands.8,44 In addition, an ‘atypical’ B12-riboswitch from Bacillus subtilis demonstrates structural flexibility that allows broader corrinoid recognition, consistent with its more promiscuous ligand-bonding profile.9,46
The discovery of natural B12-riboswitches that preferentially bind corrinoids with smaller ‘upper’ axial ligands, such as MeCbl and H2OCbl, now typified as class-II Cbl- riboswitches,8,42,44,46 suggests that analogous ligand selectivity could be exploited in the design of corresponding aVitB12s. Guided by structure-based design, Rh-analogues of key Cbls represent promising aVitB12 candidates.28 Indeed, several structurally characterized rhodibalamins (Rhbls), including methylrhodibalamin (MeRhbl),47 chlororhodibalamin (ClRhbl)39 and acetylrhodibalamin (AcRhbl),48 (see SI), represent potential further tools to probe class-II and other Cbl-riboswitches. In parallel, the broader selectivity of ‘promiscuous’ natural B12-riboswitches for cobamides (Cbas) with different ‘lower’ ligands9 provides an additional design principle for novel aVitB12s, this time mimicking other Cba structures. Such natural preferences could be harnessed by tailoring Rhbls to exploit the structure-selective binding patterns of these riboswitches. Synthetic access to Rh-analogues of ‘incomplete’ natural corrinoids has provided advanced Rhbl-precursors, such as adenosylrhodibyrate (AdoRhby)25,32 and methylrhodibyrate (MeRhby) (see SI).49 Remarkably, key B12-biosynthetic enzymes exhibit sufficient promiscuity to accept Rh-substituted corrinoid substrates in place of their natural cobalt counterparts.25,32 As a result, certain rhodibinamides (Rhbis), such as AdoRhbi,49 may serve as substrates for alternative metabolic or enzymatic avenues to generate ‘complete’ Rhbls and other Rhbas featuring an adenosyl ‘upper’ ligand.
As exemplified by AdoRhbl, carefully designed aVitB12s hold great promise as bacterial growth inhibitors owing to their high affinity for Cbl-riboswitches. Metabolite-sensing riboswitches have recently emerged as novel antibacterial drug targets,50–52 where appropriately tailored ligands may offer valuable new approaches to antibiotic development.53–55 Given the central roles of B12-derivatives in microbial physiology,10–14 B12-dependent riboswitches represent particularly promising additions to the repertoire of riboswitch-based drug targets.50,52,56 In this context, AdoRhbl, a close structural AdoCbl-mimic, illustrates the fundamental potential of rationally designed aVitB12s as drug candidates,28 since AdoCbl-responsive class-I Cbl-riboswitches, such as the btuB riboswitch, are key regulators of corrinoid uptake and metabolism in various bacteria.
Conclusions
Our here described experimental study firmly supports the proposal that a type-I antivitamin can closely mimic the regulatory role as riboswitch ligand of the naturally selected vitamin.28 In fact, the aVitB12AdoRhbl bound the btuB riboswitch very tightly and functioned as a highly effective surrogate for the natural effector ligand AdoCbl of this key Cbl-riboswitch. By repressing expression of the B12-uptake protein BtuB, monitored here via the fluorescence of RFP and eGFP, AdoRhbl can potentially block the external B12-supply required for E. coli growth. Beyond this riboswitch-mediated control, AdoRhbl acts as a potent aVitB1228,32 and as effective inhibitor of AdoCbl-dependent enzymes,25,32 establishing and strengthening its potential as a multifunctional antibiotic that operates through several mutually independent growth-inhibiting mechanisms.23,28 Rhodium-based aVitB12s, designed as precise structural mimics of Cbl-cofactors,28,47 may be broadly useful multifunctional suppressors of the essential activities of natural B12-derivatives and thereby serve as selective and powerful novel inhibitors of the growth of B12-dependent bacteria.Author contributions
F. J. W., M. J. W., M. E. T. and B. K. conceived the project. F. J. W., N. I. K., E. D. and M. J. T. designed and performed experiments. All authors discussed the results. F. J. W., M. J. W., M. E. T. and B. K. wrote the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included in the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5cb00308c.Acknowledgements
This work was supported by the Austrian Science Fund (FWF projects P-28892 and P-33059) to B. K., Biotechnology and Biological Sciences Research Council (BBSRC) grants BB/X001946/1 and BB/Y008456/1 to M. J. W., and National Institutes of Health grant R01GM114535 to M. E. T.Notes and references
- A. Nahvi, N. Sudarsan, M. S. Ebert, X. Zou, K. L. Brown and R. R. Breaker, Chem. Biol., 2002, 9, 1043–1049 CrossRef CAS PubMed.
- W. C. Winkler and R. R. Breaker, ChemBioChem, 2003, 4, 1024–1032 CrossRef CAS PubMed.
- K. Kavita and R. R. Breaker, Trends Biochem. Sci., 2023, 48, 119–141 CrossRef CAS PubMed.
- A. Roth and R. R. Breaker, Annual Rev. Biochem., 2009, 78, 305–334 CrossRef CAS PubMed.
- S. R. Lennon and R. T. Batey, J. Mol. Biol., 2022, 434, 167585 CrossRef CAS PubMed.
- A. Nahvi, J. E. Barrick and R. R. Breaker, Nucl. Acids Res., 2004, 32, 143–150 Search PubMed.
- S. Gallo, M. Oberhuber, R. K. O. Sigel and B. Kräutler, ChemBioChem, 2008, 9, 1408–1414 CrossRef CAS PubMed.
- J. E. Johnson, F. E. Reyes, J. T. Polaski and R. T. Batey, Nature, 2012, 492, 133–137 CrossRef CAS PubMed.
- K. J. Kennedy, F. J. Widner, O. M. Sokolovskaya, L. V. Innocent, R. R. Procknow, K. C. Mok and M. E. Taga, mBio, 2022, 13, e01121–01122 CrossRef PubMed.
- R. Banerjee and S. W. Ragsdale, Ann. Rev. Biochem., 2003, 72, 209–247 CrossRef CAS PubMed.
- T. Toraya, Chem. Rev., 2003, 103, 2095–2127 CrossRef CAS PubMed.
- K. Gruber, B. Puffer and B. Kräutler, Chem. Soc. Rev., 2011, 40, 4346–4363 Search PubMed.
- M. T. Croft, M. J. Warren and A. G. Smith, Eukaryotic Cell, 2006, 5, 1175–1183 CrossRef CAS PubMed.
- Z. I. Alvarez-Aponte, A. M. Govindaraju, Z. F. Hallberg, A. M. Nicolas, M. A. Green, K. C. Mok, C. Fonseca-García, D. Coleman-Derr, E. L. Brodie, H. K. Carlson and M. E. Taga, ISME J., 2024, 18, wrae068 CrossRef CAS PubMed.
- D. A. Bryant, C. N. Hunter and M. J. Warren, J. Biol. Chem., 2020, 295, 6888–6925 Search PubMed.
- E. C. Seth and M. E. Taga, Front. Microbiol., 2014, 5, 350 Search PubMed.
- O. M. Sokolovskaya, A. N. Shelton and M. E. Taga, Science, 2020, 369(6499), eaba0165 Search PubMed.
- M. T. Croft, A. D. Lawrence, E. Raux-Deery, M. J. Warren and A. G. Smith, Nature, 2005, 438, 90–93 CrossRef CAS PubMed.
- Z. I. Alvarez-Aponte, R. R. Procknow and M. E. Taga, Ann. Rev. Microbiol., 2025, 79, 8.1–8.21 CrossRef PubMed.
- X. W. Nou and R. J. Kadner, Proc. Natl. Acad. Sci. U. S. A., 2000, 97, 7190–7195 CrossRef CAS PubMed.
- A. Gudmundsdottir, C. Bradbeer and R. J. Kadner, J. Biol. Chem., 1988, 263, 14224–14230 Search PubMed.
- D. D. Shultis, M. D. Purdy, C. N. Banchs and M. C. Wiener, Science, 2006, 312, 1396–1399 Search PubMed.
- B. Kräutler, Chem. – Eur. J., 2015, 21, 11280–11287 CrossRef PubMed.
- M. Ruetz, C. Gherasim, S. N. Fedosov, K. Gruber, R. Banerjee and B. Kräutler, Angew. Chem., Int. Ed., 2013, 52, 2606–2610 Search PubMed.
- F. J. Widner, A. D. Lawrence, E. Deery, D. Heldt, S. Frank, K. Gruber, K. Wurst, M. J. Warren and B. Kräutler, Angew. Chem., Int. Ed., 2016, 55, 11281–11286 Search PubMed.
- E. Mutti, M. Ruetz, H. Birn, B. Kräutler and E. Nexo, PLoS One, 2013, 8, e75312 Search PubMed.
- B. Kräutler, in Vitamins and Hormones, ed. G. Litwack, Academic Press, Cambridge, Mass., 2022, vol. 119, pp. 221–240 Search PubMed.
- B. Kräutler, Chem. – Eur. J., 2020, 26, 15438–15445 CrossRef PubMed.
- V. B. Koppenhagen, in B12, ed. D. Dolphin, John Wiley & Sons, 1982, vol. 2, pp. 105–150 Search PubMed.
- W. Friedrich, Vitamins, Walter de Gruyter, Berlin, 1988 Search PubMed.
- F. Zelder, M. Sonnay and L. Prieto, ChemBioChem, 2015, 16, 1264–1278 CrossRef CAS PubMed.
- M. Ruetz, R. Mascarenhas, F. Widner, C. Kieninger, M. Koutmos, B. Kräutler and R. Banerjee, Biochemistry, 2024, 63, 1955–1962 Search PubMed.
- R. Perez-Castano, J. Aranda, F. J. Widner, C. Kieninger, E. Deery, M. J. Warren, M. Orozco, M. Elias-Arnanz, S. Padmanabhan and B. Kräutler, Angew. Chem., Int. Ed., 2024, 63, e202401626 Search PubMed.
- M. Mandal and R. R. Breaker, Nat. Rev. Mol. Cell Biol., 2004, 5, 451–463 Search PubMed.
- M. Mansjö and J. Johansson, RNA Biol., 2011, 8, 674–680 CrossRef PubMed.
- C. V. Franklund and R. J. Kadner, J. Bacteriol., 1997, 179, 4039–4042 Search PubMed.
- M. D. Lundrigan, W. Köster and R. J. Kadner, Proc. Natl. Acad. Sci. U. S. A., 1991, 88, 1479–1483 Search PubMed.
- R. Y. Tsien, Angew. Chem., Int. Ed., 2009, 48, 5612–5626 Search PubMed.
- F. J. Widner, C. Kieninger, K. Wurst, E. Deery, M. J. Warren and B. Kräutler, Synthesis, 2021, 332–337 CAS.
- C. C. Fowler, E. D. Brown and Y. F. Li, Chem. & Biol., 2010, 17, 756–765 CrossRef CAS PubMed.
- C. Bradbeer, in Chemistry and Biochemistry of B12, ed. R. Banerjee, John Wiley & Sons, New York, 1999, pp. 489–506 Search PubMed.
- J. T. Polaski, S. M. Webster, J. E. Johnson and R. T. Batey, J. Biol. Chem., 2017, 292, 11650–11658 CrossRef CAS PubMed.
- M. Ruetz, A. Shanmuganathan, C. Gherasim, A. Karasik, R. Salchner, C. Kieninger, K. Wurst, R. Banerjee, M. Koutmos and B. Kräutler, Angew. Chem., Int. Ed., 2017, 56, 7387–7392 CrossRef CAS PubMed.
- S. R. Lennon, A. J. Wierzba, S. H. Siwik, D. Gryko, A. E. Palmer and R. T. Batey, ACS Chem. Biol., 2023, 18, 1136–1147 Search PubMed.
- A. Peselis and A. Serganov, Nat. Struct. Mol. Biol., 2012, 19, 1182–1184 CrossRef CAS PubMed.
- C. W. Chan and A. Mondragón, Nucleic Acids Res., 2020, 48, 7569–7583 CAS.
- F. J. Widner, K. Wurst, M. Ruetz, C. Kieninger, C. Kreutz, M. D. Paxhia, E. Deery, M. J. Warren and B. Kräutler, ChemistryEurope, 2025, 3, e202500157 CrossRef CAS.
- M. Wiedemair, C. Kieninger, K. Wurst, M. Podewitz, E. Deery, M. D. Paxhia, M. J. Warren and B. Kräutler, Helv. Chim. Acta, 2023, 106, e202200158 CrossRef CAS.
- F. J. Widner, C. Kieninger and B. Kräutler, J. Porph. Phthal., 2025, 29, 408–417 Search PubMed.
- K. F. Blount and R. R. Breaker, Nat. Biotechnol., 2006, 24, 1558–1564 CrossRef CAS PubMed.
- E. M. Aghdam, M. S. Hejazi and A. Barzegar, Gene, 2016, 592, 244–259 CrossRef PubMed.
- V. Panchal and R. Brenk, Antibiotics, 2021, 10, 45 Search PubMed.
- R. Penchovsky and M. Traykovska, Expert Opin. Drug Discovery, 2015, 10, 631–650 Search PubMed.
- M. F. Richter, B. S. Drown, A. P. Riley, A. Garcia, T. Shirai, R. L. Svec and P. J. Hergenrother, Nature, 2017, 545, 299–304 Search PubMed.
- V. Mizrahi and D. F. Warner, Science, 2019, 363, 457–458 CrossRef CAS PubMed.
- K. E. Deigan and A. R. Ferre-D'Amare, Acc. Chem. Res., 2011, 44, 1329–1338 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |