Open Access Article
Open Access ArticleHigh-throughput assay for measuring target occupancy of covalent compounds: a case study with MK2
Julian L.
Wong
 *a,
Mari
Manuia
a,
Sandra
Gao
a,
Natacha
Stoehr
b,
Theresa
Boersig
b,
Krystine
Vuong
a,
Tao
Jiang
a,
Jian
Cao
a,
Yong
Jia
a,
C. C.
King
a,
John
Joslin
a,
Leslie
Ofori
a,
Jon
Loren
a and
Zuni I.
Bassi
*a,
Mari
Manuia
a,
Sandra
Gao
a,
Natacha
Stoehr
b,
Theresa
Boersig
b,
Krystine
Vuong
a,
Tao
Jiang
a,
Jian
Cao
a,
Yong
Jia
a,
C. C.
King
a,
John
Joslin
a,
Leslie
Ofori
a,
Jon
Loren
a and
Zuni I.
Bassi
 b
b
aNovartis Biomedical Research, San Diego, California, USA
bNovartis Biomedical Research, Basel, Switzerland
First published on 4th February 2026
Abstract
Target engagement metrics provide predictive value for in vivo efficacy of low-molecular-weight compounds. While direct observation of compound binding to its intended target builds the most confidence, the throughput of these measurements tends to be low. Indirect competition approaches that ask if an unlabeled compound can displace a tracer offer a higher throughput option for compound profiling. Most indirect target engagement assays employ reversible tracers; by contrast, programs focused on developing irreversible drugs can benefit from covalent tracers whose properties better match the mechanism of action of the test compounds. We demonstrate how covalent tracers could be employed in high-throughput assays to indirectly measure target occupancy of either endogenous or exogenously overexpressed MK2 and how these cellular assays could be adapted to monitor the kinetics of compound-target binding and bioavailability in the presence of human serum.
Introduction
Covalent compounds irreversibly bind to their target proteins. These molecules typically contain an electrophilic moiety positioned to react rapidly with a proximate nucleophilic residue—often cysteine—at the target site to form a covalent bond. As drugs, covalent compounds provide many pharmacological advantages over their reversible counterparts, including increased potency, selectivity, and sustained duration of action; conversely, balancing reactivity and selectivity of covalent compounds poses a liability, especially when considering the covalent modification of off-target proteins.1Reliably measuring target occupancy is crucial to advance a drug discovery program as it can demonstrate the participation of a specific target in pathology and project what level of inhibition is needed to achieve a clinical effect.2 Two metrics that enable the quantification of target occupancy are the fraction of compound-bound sites and the abundance of total protein. Such values are often obtained by directly probing the available target population, for example using immunoprecipitation followed by mass spectrometry. An alternative approach is quantifying the fraction of available or unoccupied binding sites (“free” sites) after treatment with a test compound. In such competitive applications, a reactive tracer molecule that binds and bonds to the same pocket as the test compound(s) and contains an observable tag or handle (e.g. a fluorophore or biotin) is used to measure the number of free sites. When paired with a method to identify the targeted protein (e.g. a fusion protein or a specific antibody), the irreversible tracer enables labeling of the unoccupied target proteins after pre-treatment with a test compound. The total number of available binding sites is estimated by measuring the abundance of target protein in the same sample, using either direct quantification or calculation of the sum of free plus compound-occupied fractions in the sample. Together, measurement of the free and compound-bound populations enables calculation of the fraction of bound target protein; when plotted, higher target occupancy of a test compound results in fewer free sites within a protein population, which is reported as lower tracer signal.
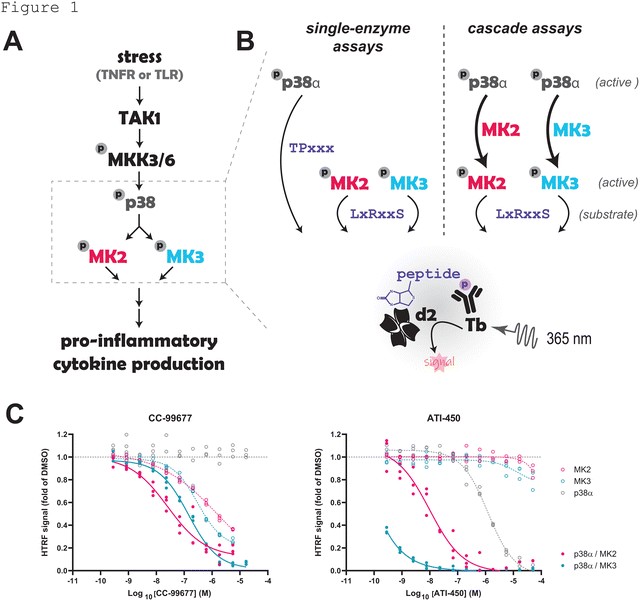
We sought to develop high-throughput, plate-based assays to measure engagement of MAPKAPK2 (MK2) with covalent inhibitors. MK2 and the structurally related protein MK3 are serine/threonine kinases phosphorylated and activated by the mitogen-activated protein kinase (MAPK) p38α/β in response to stress and inflammatory stimuli. Activated MK2 and MK3 phosphorylate and inhibit the mRNA destabilizing protein tristetraprolin (TTP), leading to increased stability and translation of mRNA of key pro-inflammatory cytokines such as tumor necrosis factor (TNF) and interleukins 6 (IL-6) and 1-beta (IL-1β).3 Given their central role in driving inflammation, MK2 and MK3 are considered promising therapeutic targets for the treatment of multiple autoimmune diseases. Initial efforts focused on the discovery of reversible MK2 kinase inhibitors; however, most of these compounds suffered from poor cellular potency and physicochemical properties.4,5 Two MK2 inhibitors that reached the clinic utilize distinct mechanisms of action: CC-99677 is a covalent MK2 inhibitor that directly binds and irreversibly inhibits the catalytic site of MK26,7 whereas ATI-450 selectively stabilizes the association between MK2 and p38, thereby inhibiting downstream MK2 signaling.8–10 We selected these two compounds for evaluating our target engagement assays because they should exhibit distinct occupancy profiles using tracers designed to bind to the catalytic site of MK2.
Experimental
Reagents
Cell culture
All cell lines were cultured at 37 °C, 5% CO2, under saturating humidity. HeLa and RAW264.7 cells were grown in Dulbecco's Minimum Essential Media containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic. HCC1428 cells were grown in RPMI 1640 medium containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic. THP-1 cells were grown in RPMI medium containing 1× Glutamax, 25 mM HEPES, 10% fetal bovine serum, 1 mM sodium pyruvate, 0.05 mM β-mercaptoethanol, 100 µg mL−1 normocin, and 1% penicillin/streptomycin. Cells were passaged regularly, using trypsinization for adherent cells. Adherent cells used for the plate-based experiments were resuspended in Opti-MEM containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic.Biochemical assays
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000. Data were then normalized to mean background signal (no-enzyme wells) and 100% signal (dimethyl sulfoxide control, no inhibitor wells) for non-linear regression curve fitting, using Prism software version 10 (GraphPad, La Jolla, CA, U.S.A.), to determine 50% inhibitor concentration (IC50) values.
000. Data were then normalized to mean background signal (no-enzyme wells) and 100% signal (dimethyl sulfoxide control, no inhibitor wells) for non-linear regression curve fitting, using Prism software version 10 (GraphPad, La Jolla, CA, U.S.A.), to determine 50% inhibitor concentration (IC50) values.
Assay development
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 ratio of DNA
3 ratio of DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) FuGENE HD in Opti-MEM containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic. This cell resuspension was plated to a sterile, solid white tissue-culture-treated 1536-well plate (Greiner Bio-One, Monroe, NC, U.S.A.) using an HD WASHER (GNF Systems, San Diego, CA, U.S.A.) and then returned to culture conditions for 24 hours. An equal volume of dimethyl sulfoxide or tracer diluted in Opti-MEM containing 1× Glutamax, and 1× antibiotic–antimycotic were then added to each well. Cells were allowed to equilibrate for 1 hour under culture conditions or were processed immediately to measure tracer kinetics. Nano-Glo substrate plus NanoLuciferase inhibitor, with or without a supplement of Endurazine, diluted in Opti-MEM were then added to wells for a final concentration of 1× each. Full emission spectra or kinetics were measured on a CLARIOstar Plus (BMG Labtech, Car, NC, U.S.A.) using a 0.5-s exposure and 3600 gain over a continuous series of 5-nm windows or 442–470 nm and 575–625 nm bandpasses every 2 minutes, respectively. Alternatively, a luminescence plate reader (GNF Systems, San Diego, CA, U.S.A.) with custom 452 ± 22.5 nm and 607 ± 35 nm bandpass filters was used to capture signal with a 10-second exposure per filter set.
FuGENE HD in Opti-MEM containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic. This cell resuspension was plated to a sterile, solid white tissue-culture-treated 1536-well plate (Greiner Bio-One, Monroe, NC, U.S.A.) using an HD WASHER (GNF Systems, San Diego, CA, U.S.A.) and then returned to culture conditions for 24 hours. An equal volume of dimethyl sulfoxide or tracer diluted in Opti-MEM containing 1× Glutamax, and 1× antibiotic–antimycotic were then added to each well. Cells were allowed to equilibrate for 1 hour under culture conditions or were processed immediately to measure tracer kinetics. Nano-Glo substrate plus NanoLuciferase inhibitor, with or without a supplement of Endurazine, diluted in Opti-MEM were then added to wells for a final concentration of 1× each. Full emission spectra or kinetics were measured on a CLARIOstar Plus (BMG Labtech, Car, NC, U.S.A.) using a 0.5-s exposure and 3600 gain over a continuous series of 5-nm windows or 442–470 nm and 575–625 nm bandpasses every 2 minutes, respectively. Alternatively, a luminescence plate reader (GNF Systems, San Diego, CA, U.S.A.) with custom 452 ± 22.5 nm and 607 ± 35 nm bandpass filters was used to capture signal with a 10-second exposure per filter set.
Cell plating/serum exchange/compound treatment for target engagement assays
Similar protocols were used for the high-throughput, 1536-well format assays described, including washing of target cells after compound treatment. Briefly, single-cell suspensions were made in Opti-MEM containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic for plating to a sterile, solid white tissue-culture-treated 1536-well plate (Greiner Bio-One, Monroe, NC, U.S.A.) using an HD WASHER (GNF Systems, San Diego, CA, U.S.A.). Plated cells, in 4 µL per well, were then returned to culture conditions for 24 hours. When testing the effect of different serum concentrations or sources on target occupancy, this incubation period was followed by five washes with 2.5-volumes of Opti-MEM containing 1× Glutamax, 1× antibiotic–antimycotic, leaving ∼2.5 µL of fluid per wash cycle, using an HD WASHER. A final aspiration was performed to leave ∼2 µL fluid, to which an equal volume of Opti-MEM containing 20%, 40%, or neat (100%) serum was added to achieve the same 4 µL final volume per well.Test compounds were transferred to the washed cells using an ECHO 555 acoustic dispenser (Beckman Coulter, Brea, CA, U.S.A.) for a final concentration of 0.5% dimethyl sulfoxide. Three hours after compound addition, treated cells were washed five times with 2.5-volumes, leaving ∼2.5 µL of fluid per cycle, using an HD WASHER. Wash buffers used for the post-treatment phase were Opti-MEM containing 1× Glutamax, and 1× antibiotic–antimycotic, for the NanoBRET assays, or Dulbecco's phosphate-buffered saline, for the multiplex HTRF assays. A final aspiration was performed to leave ∼2 µL fluid, followed by assay-specific methods for detection.
Dual homogenous time-resolved fluorescence (HTRF) target engagement assay
Single-cell suspensions of trypsinized RAW264.7 or HCC 1428 cells were made in Opti-MEM containing 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic for plating at a density of 4000 cells per well, as described in the cell plating/serum exchange/compound treatment section; wells reserved for recombinant protein standards were loaded with culture medium only. Treated cells were washed three times with Dulbecco's phosphate-buffered saline, followed immediately by lysis with two microliters of IP Lysis Buffer and incubation on ice for at least 30 minutes. Recombinant murine or human MK2 standard, diluted in IP Lysis Buffer, was added to respective cell-free wells. One microliter of tracer diluted in detection buffer was added per well and incubated at room temperature for 30 minutes, followed by addition of detection antibodies, diluted in detection buffer, for a final per-well concentration of 50 nM tracer, 0.5× anti-MK2 [D1E11] terbium-cryptate, 1 ng µL−1 anti-MK2 [7H4.2]-Alexa Fluor 633, and 0.4 ng µL−1 streptavidin-Alexa Fluor 488. Reactions were allowed to equilibrate at 4 °C overnight before reading on a PHERAstar FSX (BMG Labtech, Car, NC, U.S.A.) using HTRF modules (337 nm excitation/520 nm and 490 nm emission or 337 nm excitation/665 nm and 620 nm emission) with 2-pulse laser excitation and reading over 400 microseconds after a 50-microsecond delay.The HTRF ratio was calculated by dividing the acceptor emission signal (620 nm or 665 nm) by the terbium-donor emission signal (490 nm or 620 nm, respectively), followed by subtraction of the mean background signal (no-cell and no-MK2 wells). Target engagement was then calculated by dividing the tracer HTRF ratio (520/490 nm) by the MK2 protein HTRF ratio (665/620 nm). The fraction of free target was calculated by normalizing the target engagement values to the dimethyl sulfoxide-control conditions.
NanoLuciferase bioluminescence resonance energy transfer (NanoBRET) target engagement assay
Trypsinized HeLa cells were transfected with NanoLuciferase expression constructs at 5 µg DNA (0.5 µg pD649 plasmid + 4.5 µg carrier DNA) per million cells using a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 ratio of DNA
3 ratio of DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) FuGENE HD in Opti-MEM 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic prior to plating at a density of 1000 cells per well, as described in the cell plating/serum exchange/compound treatment section. Two microliters of dimethyl sulfoxide (background control) or tracer diluted in Opti-MEM containing 1× Glutamax, and 1× antibiotic–antimycotic were added to each well of the treated-and-Opti-MEM-washed plates, for a final concentration of 2 µM tracer (0.5% dimethyl sulfoxide), and then returned to culture conditions for 1 hour. Nano-Glo substrate plus NanoLuciferase inhibitor diluted in Opti-MEM were then added to wells for a final concentration of 1× each. Plates were allowed to equilibrate to room temperature for about 5 minutes before measuring luminescence signal on a luminescence plate reader using 452 ± 22.5 nm and 607 ± 35 nm bandpass filters and a 10-second exposure per filter set.
FuGENE HD in Opti-MEM 10% fetal bovine serum, 1× Glutamax, and 1× antibiotic–antimycotic prior to plating at a density of 1000 cells per well, as described in the cell plating/serum exchange/compound treatment section. Two microliters of dimethyl sulfoxide (background control) or tracer diluted in Opti-MEM containing 1× Glutamax, and 1× antibiotic–antimycotic were added to each well of the treated-and-Opti-MEM-washed plates, for a final concentration of 2 µM tracer (0.5% dimethyl sulfoxide), and then returned to culture conditions for 1 hour. Nano-Glo substrate plus NanoLuciferase inhibitor diluted in Opti-MEM were then added to wells for a final concentration of 1× each. Plates were allowed to equilibrate to room temperature for about 5 minutes before measuring luminescence signal on a luminescence plate reader using 452 ± 22.5 nm and 607 ± 35 nm bandpass filters and a 10-second exposure per filter set.
The NanoBRET ratio was calculated by dividing the 607 ± 35 nm signal by the 452 ± 22.5 nm signal, followed by subtraction of the mean NanoBRET ratio from the background wells (no tracer addition). The fraction of free target was calculated by normalizing the corrected NanoBRET ratios to the dimethyl sulfoxide-control conditions. k-on and k-off kinetics were calculated using Prism software version 10 (GraphPad, La Jolla, CA, U.S.A.).
Streptavidin mass shift (SMaSh) assay
One million THP-1 cells were plated per well of a 24-deep-well plate in a final volume of 450 µL. Compound was serially diluted in dimethyl sulfoxide and incubated with cells for 3 hours at 37 °C, 5% CO2. Cell suspensions were collected by centrifugation at 350 × g for 5 minutes and then washed once with phosphate-buffered saline. Washed cell pellets were then lysed in 60 µL RIPA buffer with 1× HALT protease and phosphatase inhibitor, vortexed, and incubated overnight at −80 °C to complete the lysis.Fresh human whole blood was collected in sodium-heparin tubes from healthy donors from which no additional details are available, due to anonymity, with informed consent (Sante’ med Gesundheitszentrum AG Basel, Switzerland; all experiments were performed in accordance with the Swiss Human Research Act and collection of human whole blood (TRI0128) was approved by the Ethikkommission Nordwest- und Zentralschweiz). Blood was diluted in an equal volume of assay medium (RPMI 1640 with 5% fetal bovine serum, 10 mM HEPES, and 50 µM β-mercaptoethanol), resulting in 50% whole blood, and aliquoted to 96-deep-well plates with 250 µL per well. Compound was serially diluted in dimethyl sulfoxide and incubated with blood for 18 hours at 37 °C, 5% CO2 (plates covered in a cell culture incubator), after which pure lipopolysaccharide was added to a final concentration of 1 µg mL−1 for 3 hours. Plates were centrifuged at 500 × g for 10 minutes and supernatants were removed. Red blood cells were lysed with fresh diluted 1× ACK lysis buffer for 10 minutes on ice, then samples were washed twice with cold phosphate-buffered saline. The white blood cell pellets were lysed in 100 µL RIPA buffer containing 1× HALT protease and phosphatase and 5 µM diisopropylfluorophosphate, vortexed, and incubated overnight at −80 °C to complete the lysis.
Frozen blood or THP-1 cell lysates were thawed on ice and centrifuged at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g for 30 minutes at 4 °C. Aliquots of supernatants were incubated for 1 hour at room temperature with 25 µM biotinylated tracer for the SMaSh assay (Fig. S1A). Each sample was then mixed with 0.3 mg mL−1 streptavidin together with fluorescent Master Mix for an additional 10 minutes at room temperature before denaturation at 95 °C for 10 minutes. The samples were loaded for electrophoresis and detection on the WES/Jess platform, according to the manufacturer's instructions, using anti-MK2 clone E341 diluted 1
000 × g for 30 minutes at 4 °C. Aliquots of supernatants were incubated for 1 hour at room temperature with 25 µM biotinylated tracer for the SMaSh assay (Fig. S1A). Each sample was then mixed with 0.3 mg mL−1 streptavidin together with fluorescent Master Mix for an additional 10 minutes at room temperature before denaturation at 95 °C for 10 minutes. The samples were loaded for electrophoresis and detection on the WES/Jess platform, according to the manufacturer's instructions, using anti-MK2 clone E341 diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 250. Electropherograms were analyzed using Protein Simple's Compass software version 6.2.0 (San Jose, CA, U.S.A.), with high dynamic range exposure time and peak fit analysis settings. The amount of free target was calculated using the free MK2 peak area (MK2 bound by the probe and streptavidin) divided by the total MK2 area, calculated as the sum of the bound (short plus long bound MK2 peaks)12 and free MK2 peak areas.
250. Electropherograms were analyzed using Protein Simple's Compass software version 6.2.0 (San Jose, CA, U.S.A.), with high dynamic range exposure time and peak fit analysis settings. The amount of free target was calculated using the free MK2 peak area (MK2 bound by the probe and streptavidin) divided by the total MK2 area, calculated as the sum of the bound (short plus long bound MK2 peaks)12 and free MK2 peak areas.
Results and discussion
Target engagement assays require the quantification of compound-occupied versus total target protein in a sample, often employing tagged tracer molecules that compete for the same site as the unlabeled test compound(s). The abundance of each fraction of target protein is optimally measured within the same sample to reduce variance due to sample loading and to account for changes in de novo protein expression or toxicity of the test compound.13Two published approaches employ different methods for measuring target occupancy. The streptavidin mass shift (SMaSh) assay quantifies the free versus compound-bound target fractions in the same loaded samples.13 This approach employs a covalent tracer with a biotin handle to identify the fraction of free endogenous target protein within a population of cells or in tissue following exposure to a naïve compound of interest. Streptavidin is then added to the homogenate to form a tri-molecular tracer/protein/streptavidin complex that exhibits slower electrophoretic mobility under denaturing conditions, thus distinguishing compound-bound versus free (tracer-bound) populations for quantification via the same immunodetection method (see Fig. S1). While the SMaSh assay represents an elegant approach to measure target occupancy with covalent compounds, electrophoresis and immunodetection steps are major bottlenecks to throughput. NanoLuciferase bioluminescent resonance energy transfer (NanoBRET) is a plate-based assay developed for tracking compound target engagement and residence time in live cells.14 This approach employs cell-permeable, often reversible fluorophore-conjugated tracers that serve as energy acceptors when located within nanometers of the energy donor NanoLuciferase, which is directly fused to the target protein.15,16 This plate-based approach is amenable to at least 384-well format17 but is limited to cell or animal models expressing the target protein fused to NanoLuciferase.
We sought to develop high-throughput target engagement assays for programs focused on developing covalent inhibitors and asked whether irreversible covalent tracers could be used in such assays. We used MK2 as a test case, given the distinct proposed mechanisms of action for the two MK2 inhibitors (CC-99677 and ATI-450) we chose as benchmarks for these tracer-competitor assay builds.
MK2 interactions differ for CC-99677 and ATI-450
We initially tested if CC-99677 and ATI-450 indeed differentially inhibit MK2 activity. CC-99677 and ATI-450 were tested in a biochemical cascade assay where recombinant, active p38α is used to phosphorylate and activate unactive MK2. Phosphorylation of a generic serine/threonine kinase (STK) peptide was then measured as a readout for MK2 activity. Since these enzyme-linked assays capture both inhibitors of individual proteins (p38α or MK2) as well as molecules that target the interaction between p38α and MK2, we compared results of the cascade assays to biochemical kinase assays for active versions of each enzyme run in parallel (Fig. 1A and B). Given the 75% sequence identity between MK2 and MK3,18 we also asked whether MK3 could be inhibited by these two compounds.Both CC-99677 and ATI-450 inhibited downstream phosphorylation of STK peptide in the p38α/MK2 and p38α/MK3 cascade assays (Fig. 1C). CC-99677 also affected individual MK2 and MK3 enzyme assays, with no inhibition in the p38α assay; by contrast, ATI-450 only minorly impacted the individual MK2, MK3, and p38α assays. These data support the claim that CC-99677 and ATI-450 have different mechanisms of action: CC-99677 interaction with MK2 and MK3 is independent of p38α whereas the effect of ATI-450 on MK2 and MK3 requires the presence of and binding with p38α.
Development of a dual homogenous time-resolved assay for measuring endogenous MK2
The SMaSh assay in THP-1 cells and human whole blood allowed us to quantify CC-99677 binding to endogenous MK2 using a covalent tracer with a biotin handle (Fig. S1). Nevertheless, this method's reliance on electrophoretic separation of streptavidin-associated free MK2 from MK2 bound to the test compound13 severely limited its throughput and utility when running concentration-response profiles for each test compound. We therefore sought to develop a plate-based method that could quantify endogenous MK2 target engagement in cells or tissue, measuring both tracer-bound and total protein in the same sample–a feat the SMaSh assay excelled at compared to standard immunosorbent or immuno-pulldown assays.We focused on measuring target engagement in cell and tissue lysates with homogenous time-resolved fluorescence (HTRF) because it can report distance relationships of 1 to 10 nanometers, due to dipole coupling between donor and acceptor that limits efficient energy transfer beyond this range.16 Excitation of terbium with 337-nm light results in emission peaks at 490, 544, 585, and 622 nm19 that can excite fluorophores in discrete families (e.g. respectively Alexa Fluor 488, Alexa Fluor 568, and Alexa Fluor 633). Thus, energy transfer between terbium cryptate and up to three distinct entities is quantifiable in a single well for multiplexing readouts.20
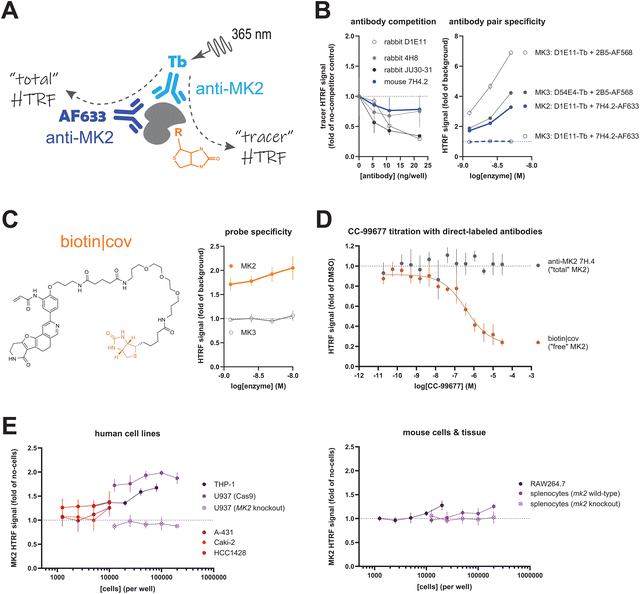
The primary obstacle for a multiplex HTRF assay was identification of an antibody pair that could simultaneously bind MK2 at non-overlapping epitopes as a signal donor and an acceptor (for measuring “total” MK2) without interfering with tracer binding (Fig. 2A). A survey of three rabbit monoclonal antibodies (clones 4H8, D1E11, and JU30-31) revealed anti-MK2 D1E11 as the ideal candidate for direct conjugation to terbium cryptate due to its overall assay window at lower concentrations (data not shown). A competition assay identified mouse anti-MK2 7H4.2, which bound a non-overlapping site to D1E11, for measuring total MK2 abundance (Fig. 2B). Pairing anti-MK2 D1E11 conjugated to terbium cryptate with anti-MK2 7H4.2 conjugated to Alexa Fluor 633 produced an HTRF signal only in the presence of recombinant human MK2; of note, pairing of these two anti-MK2 antibodies proved fortuitous as anti-MK2 D1E11-terbium cryptate also recognizes recombinant MK3, as demonstrated by the signal obtained when paired with anti-MK3 2B5. Thus, the choice of antibody pairs for the same target protein can improve signal specificity for measuring “total” protein abundance. The assay window provided by the combination of anti-MK2 D1E11-terbium cryptate, anti-MK2 7H4.2-Alexa Fluor 633, and biotin|cov, the biotinylated tracer used in this assay (Fig. 2C), yielded a competition profile for CC-99677 target engagement against recombinant human MK2, with the expected concentration-dependent signal loss in biotin|cov signal (Fig. 2D).
This dual HTRF assay was also able to detect endogenous MK2 in cell lines and mouse tissue in a 1536-well format. Immortalized human cell lines representing two lineages were evaluated (Fig. 2E): myeloid cells, which are generally semi-adherent (THP-1 and U937), and those of epithelial origin, which are adherent and possess a tenfold range in MK2 transcript abundance (Caki-2, kidney < A-431, epidermis ≪ HCC1428, mammary gland) (https://www.proteinatlas.org). The strong signal from U937 cells was eliminated following MK2 CRISPR knockout, supporting the specificity of the D1E11-terbium cryptate/7H4.2-Alexa Fluor 633 antibody pairing in human cell lysates. THP-1 cells possess similar levels of endogenous MK2, with signal overlapping that from HCC1428 cells at lower sample concentrations. Among the murine samples, immortalized myeloid RAW264.7 cells yielded signal on par with human THP-1 and HCC1428 cells at lower cell densities; nearly tenfold more splenocytes isolated from a wild-type mouse were required to achieve similar signal. Genetic knockout of mk2, however, eliminated the murine splenocyte signal as well as signal from skeletal muscle and dermis (Fig. S2), further supporting use of the chosen detection antibody pair.
This dual HTRF assay enables high-throughput quantification of endogenous MK2 target engagement in cells or tissues. The quality and reliability of these measurements, however, greatly depends on the abundance of target protein in the chosen cell line or tissue lineage versus per-well load: cells of the monocyte lineage often yielded the lowest variance among technical replicates compared to those of epithelial origin, even at the same cell plating densities (Fig. 2E). Only a few immortalized, adherent human lines maintain high MK2 abundance under standard culture conditions, limiting our ability to screen compounds with this assay due to the requisite wash step to remove excess compound prior to lysis. We therefore sought to develop a complementary, NanoBRET target engagement assay14 in adherent cells overexpressing a NanoLuciferase-MK2 fusion protein.
A high-throughput NanoBRET assay with a covalent tracer for MK2
Commercial NanoBRET tracers were not available for MK2, so we designed two tracers that would engage human and mouse isoforms. The success we had with a covalent tracer in the SMaSh assay (Fig. S1) led us to evaluate both covalent (BODIPY585|cov) and reversible (BODIPY585|rev) tracers with similar MK2-binding moieties for the CC-99677 binding site and with structures based on the characteristics of published and commercially available tracers.15 Both tracers were conjugated to the red-shifted BODIPY585 fluorophore, whose excitation and emission spectra overlap with the tail of NanoLuciferase emission (Fig. 3A). No difference in cell permeability was predicted or observed between these two tracers, yet BODIPY585|cov yielded a stronger emission peak at ∼600 nm that translated into a concentration-dependent increase in NanoBRET signal from HeLa cells transiently overexpressing either amino- or carboxy-terminal NanoLuciferase-human MK2 fusion proteins (Fig. 3B), whether or not the cell membrane was permeabilized with 0.17 mM Triton X-100;21 this signal was specific to the MK2 paralog (Fig. S3). Kinetic monitoring of cells exposed to 2 µM BODIPY585|cov revealed rapid accumulation and no plateau of NanoBRET signal within 1 hour (Fig. 3C). We suspect that the bond created between the covalent tracer and MK2 provided greater detection sensitivity via accumulation of labeled-protein moiety, as opposed to measuring at equilibrium between the bound and unbound BODIPY|rev. Identifying a non-covalent tracer chemotype with a higher affinity may overcome the minimal binding observed with BODIPY|rev as the phenotypic difference between this pair of tracers resides primarily with reactivity of the parent molecule; we did not pursue this path because the covalent tracer provided such a strong assay window. NanoBRET profiles were indistinguishable between the two fusion proteins after transient transfection, implying that either would be suitable for further evaluation; we arbitrarily chose the amino-terminal fusion for further assay development. Note that attempts to generate stable cell lines failed, suggesting that chronic overexpression of either fusion protein is cytotoxic to HeLa cells.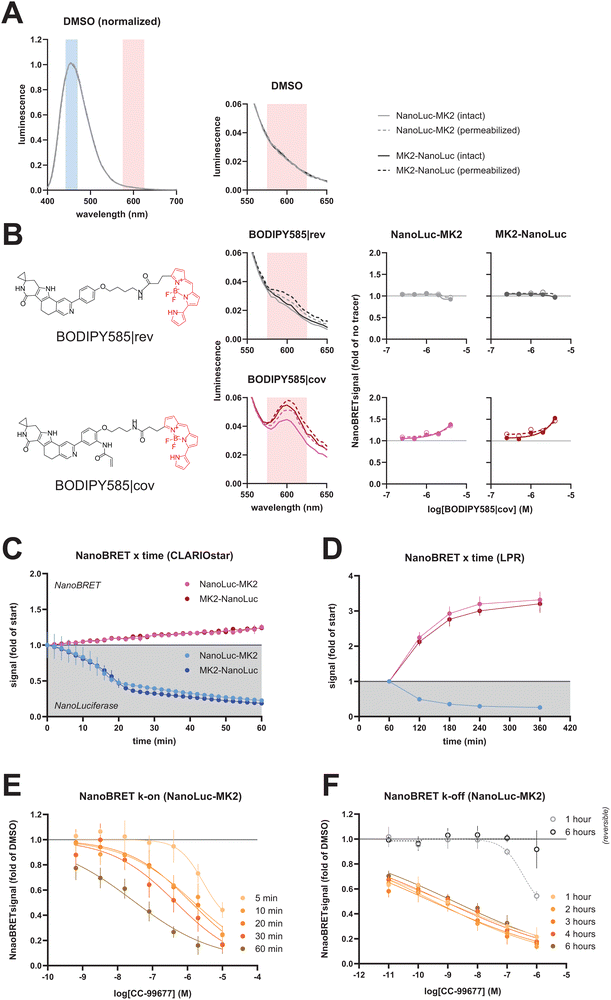 | ||
| Fig. 3 NanoBRET assay development. (A) Emission spectrum of NanoLuciferase measured live in dimethyl sulfoxide-treated HeLa cells over-expressing either amino- or carboxy-terminal MK2 fusions, with or without 0.17 mM (0.004%) Triton X-100 permeabilization (dashed or solid lines, respectively).21 The filter band passes used to measure NanoBRET signal are highlighted (centered at 452 nm for NanoLuciferase and 600 nm for BODIPY585). Magnified portion of the spectrum (right) shows the fraction of light from the NanoLuciferase donor that overlaps with –and should be subtracted from– the BODIPY acceptor emission signal. (B) Characterization of BODIPY|rev and BODIPY585|cov tracers, which compete for the same binding pocket as CC-99677. Structures of tracers (left) are shown alongside magnified emission spectra when added to HeLa cells transfected with plasmids encoding either amino- or carboxy-terminal MK2 fusions (middle) as well as the ratiometric NanoBRET signal following a titration of each tracer (right). Signal was measured 1 hour after tracer addition to the live intact (solid) or permeabilized (open/dashed line) cells using a CLARIOstar Plus (average of 5 replicates with standard deviation is shown). Line weight and style follow the legend in panel (A). (C) and (D) NanoBRET signal kinetics in the presence of BODIPY585|cov tracer, using well-by-well acquisitions (C) on the CLARIOstar Plus (CLARIOstar) or full-plate imaging (D) with a GNF Systems luminescence plate reader (LPR). NanoLuciferase signal (blue profiles in shaded background) decayed over time whereas the NanoBRET ratio (red profiles in white background) increased with exposure time to tracer. Continuous measurements for longer than 1 hour were achieved by supplementing with Endurazine substrate. (E) and (F) Measurement of k-on (E) and k-off (F) for CC-99677 using NanoBRET with BODIPY585|cov on a luminescence plate reader. The time points denote when CC-99677-containing medium was exchanged for medium containing excess BODIPY585|cov, which irreversibly quenched the system by saturating free binding sites. The profile for a reversible compound that occupies the same pocket as the covalent CC-99677 (gray, open circles) is included in the k-off plots (F). | ||
One major obstacle to the throughput of this NanoBRET assay was the time required to acquire data for a single plate. Reading NanoLuciferase luminescence (∼450 nm peak) and BODIPY585|cov (∼600 nm peak) emission sequentially on a per-well basis required about 45 minutes for a 1536-well plate on a CLARIOstar Plus. As a consequence, light production from the first read would be threefold stronger than the last read (Fig. 3B)—a loss that negatively impacts assay sensitivity. By contrast, imaging the entire plate at once, via a GNF Systems luminescence plate reader (LPR), captured similar levels of NanoLuciferase activity across all wells over a given exposure time. Adding specific bandpass filters between the plate and charge-coupled device camera enabled us to quantify luminescence at the respective wavelengths across the entire 1536-well plate in about 5 minutes for both wavelengths. Measurement of kinetics beyond 1 hour was achieved by including Endurazine as a NanoLuciferase substrate, albeit NanoBRET variance consistently increased as NanoLuciferase activity declined due to substrate depletion (Fig. 3D). The assay configurations presented herein are also amenable to lower well-density plate formats (e.g. 96- and 384-well), assuming the plate readers can acquire data for two wavelengths of light before substrate is depleted (conservatively 30 minutes after substrate addition). In these cases, effects of substrate depletion across the plate are minimized by matching the sequence of substrate dispensation with the well-read order. Utilizing luminescence plate readers with greater detector sensitivity would also improve assay window, assuming appropriate bandpass filters are used.
A fast-reacting covalent tracer such as BODIPY585|cov provided a convenient ‘stop’ reagent that permitted capturing snapshots of free MK2 abundance at a given time, thus enabling estimates of compound kinetics with the NanoBRET format. This was demonstrated by measuring k-on and k-off rates of CC-99677 against the amino-terminal NanoLuciferase-MK2 fusion protein. We first asked if BODIPY585|cov would allow for the rapid ‘quenching’ of free binding sites in k-on studies. Transfected cells were treated with CC-99677, followed by a series of quenching steps, providing a time course of compound binding on the same 1536-well plate. Each time point was captured by initiating three media exchanges to remove extracellular compound, followed immediately by addition of BODIPY585|cov to react with all the free MK2 binding sites at that time. NanoBRET signal was measured at least one hour after initiating the final time point on the plate, resulting in a set of concentration-response profiles that revealed the time-dependent binding of CC-99677 (Fig. 3E); replotting and curve fitting using the kinetics curves for the top four concentrations of compound yielded a shared k-on of 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 M−1 min−1. A complementary evaluation was designed to measure k-off using the extended-duration format (see Fig. 3F), that is by longitudinally tracking NanoBRET signal. Transfected cells were preincubated with CC-99677 for 1 hour, followed by compound washout and addition of BODIPY585|cov. About 50 minutes after tracer addition, Nano-Glo plus Endurazine substrates were added to the plate and NanoBRET signal was measured hourly. Consistent with a covalent compound such as CC-99677, no change was observed in target occupancy up to 6 hours after the wash step; conversely, the titration profiles of a fast-off reversible compound shifted over time such that the data points from the 6-hour profile were indistinguishable from the dimethyl sulfoxide control (Fig. 3F).
000 M−1 min−1. A complementary evaluation was designed to measure k-off using the extended-duration format (see Fig. 3F), that is by longitudinally tracking NanoBRET signal. Transfected cells were preincubated with CC-99677 for 1 hour, followed by compound washout and addition of BODIPY585|cov. About 50 minutes after tracer addition, Nano-Glo plus Endurazine substrates were added to the plate and NanoBRET signal was measured hourly. Consistent with a covalent compound such as CC-99677, no change was observed in target occupancy up to 6 hours after the wash step; conversely, the titration profiles of a fast-off reversible compound shifted over time such that the data points from the 6-hour profile were indistinguishable from the dimethyl sulfoxide control (Fig. 3F).
MK2 target engagement assays distinguish ATI-450 and CC-99677 target occupancy profiles
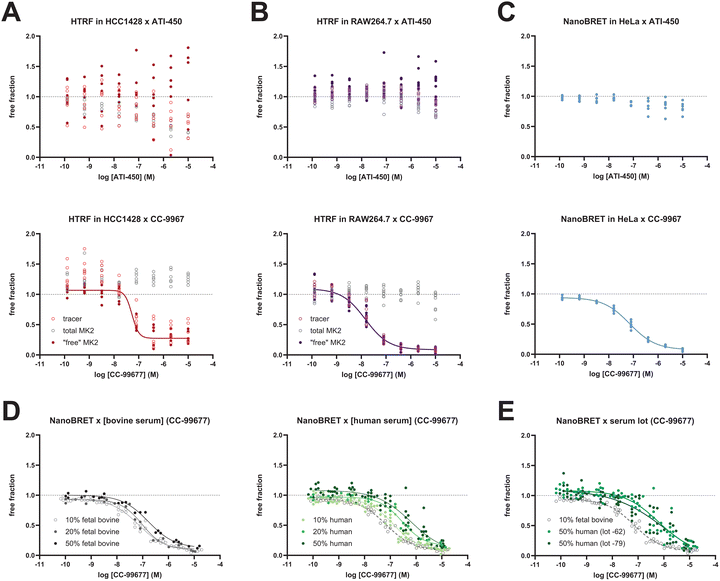
A comparison of the two high-throughput target engagement assays was conducted to determine the performance of the mechanistically independent compounds ATI-450 and CC-99677. We used human HCC1428 and murine RAW264.7 cells for the dual HTRF assay because of their tenacious adherence to standard culture plates, which enabled removal of excess compound via multiple wash cycles prior to cell lysis. The expected reciprocal relationship between target abundance in the tested cell type and variance manifested as tighter replicate datasets when MK2 abundance was greatest (transfected HeLa cells ≫ RAW264.7 cells > HCC1428 cells) (Fig. 4A–C).Both assays revealed concentration-dependent MK2 engagement for CC-99677, which competes for the tracer-binding pocket, whereas the MK2-p38-interface binder, ATI-450, did not show any tracer competition following a 3-hour incubation period (Fig. 4A–C). The importance of measuring both free and total MK2 quantities in target engagement assays was exemplified in the dual HTRF dataset from HCC1428 cells treated with ATI-450, for which high compound concentrations reduced both the tracer and total MK2 signal to the same degree, yielding no apparent change in target occupancy across the titration range; by contrast, total MK2 levels were not impacted by CC-99677 titration in the same cells (Fig. 4A and B). The 3-hour treatment with CC-99677 demonstrated similar MK2 50% occupancy (OC50) across all target engagement assays: 14.5 nM CC-99677 in RAW264.7 cells; 53.2 nM in HCC1248 (both via dual HTRF); 70.3 nM in HeLa cells (via NanoBRET); and 38.7 nM in THP-1 cells (via SMaSh; Fig. S1B). Note that the maximum target occupancy of endogenous MK2 by this compound varied, with partial occupancy in human epithelial HCC1428 cells (by dual HTRF, after standard background subtraction) versus complete occupancy in the myeloid murine RAW264.7 cells (dual HTRF assay) and human THP-1 cells (SMaSh assay). Given that full target occupancy was observed with other covalent MK2 inhibitors using the HCC1428-based dual HTRF assay (data not shown), we attribute the incomplete occupancy phenotype to the performance of CC-99677 in this epithelial cell line.
The range of CC-99677 OC50 values (40–70 nM) obtained for human MK2 in the different 3-hour target engagement assays was lower than the 384 nM OC50 measured in the overnight ex vivo human whole blood using the SMaSh assay (Fig. S1C). We investigated the source of this discrepancy by asking how the presence of human serum and/or the concentration of serum in the assay (∼50%) impacted these estimates. The NanoBRET method was modified to simulate compound exposure in the presence of human serum by including a serum-exchange step, which entailed washing the cells with Opti-MEM followed by addition of culture medium containing a blend of human sera from multiple healthy donors, prior to compound addition (see Experimental). Increasing concentrations of human serum progressively right-shifted the CC-99677 binding profiles, reducing the OC50 value by at least an order of magnitude with 50% human serum compared to 10% fetal bovine serum (Fig. 4D and Table 1). A proportional increase in bovine serum concentration did not exhibit the same shift, suggesting that this concentration-dependent loss in binding is due to a lower free fraction of compound in human serum. Including 50% human serum indeed right-shifted CC-99677 OC50, independent of serum donor pool, in both the NanoBRET and dual HTRF assays (Fig. 4E and Table 1).
| NanoBRET OC50 (nM) | HCC1428 dual HTRF OC50 (nM) | |
|---|---|---|
| Fetal bovine | ||
| 10% serum | 70.3 | 53.2 |
| 20% serum | 95.3 | 31.3 |
| 50% serum | 165.6 | 67.0 |
| Human | ||
| 10% serum (lot-62) | 197.7 | 112.7 |
| 20% serum (lot-62) | 285.1 | 254.1 |
| 50% serum (lot-62) | 588.8 | 929.0 |
| 50% serum (lot-79) | 1472.3 | 304.1 |
Conclusion
We developed two high-throughput target engagement assays using covalent tracers (Fig. S4). Similar to the SMaSh assay, our dual HTRF assay allowed for simultaneous detection of endogenous free MK2 and total MK2 in the same well, with the added advantage of considerably increased throughput. While the assay provides an excellent option for testing target engagement in cells and tissues expressing high levels of MK2, we were unable to employ this assay with samples possessing low endogenous MK2 abundance in the 1536-well plate format. The complementary NanoBRET approach overcomes such limitations through overexpression of a NanoLuciferase fusion protein. Empirical comparison of two physicochemically equivalent tracers for MK2 favored the use of a covalent tracer due to the accumulation of irreversible tracer-fusion protein complexes over time, as opposed to a lower equilibrium concentration of this moiety using a likely fast k-off reversible tracer.Assuming a tracer is available or can be derived from a compound that occupies the site of interest on the target protein, which assay—SMaSh, dual HTRF, or NanoBRET—will best address a program's needs? The following summarizes key considerations to help with the decision:
• What are the chemical properties of the tracer? Most tracers available for indirect measurement of target engagement are reversible molecules.15 The ability to dissociate, however, limits their utility compared to covalent tracers. While both types can report target occupancy through competition for the same pocket, fast-reacting irreversible tracers provide an added benefit as “stop” reagents, making measurement of compound association and dissociation kinetics simpler since k-off rates are null.
• What handle does the tracer contain? Biotin is best suited for the SMaSh and dual HTRF assays whereas red fluorophores excited by ∼545–585 nm light can be used for the dual HTRF or NanoBRET, albeit most tracers can be synthesized with a varienty of handles. One essential characteristic of the fluorescent tracers is cell permeability for the NanoBRET method.
• What is the endogenous abundance of the target protein in the cell line or tissue of interest? If low, but essential to monitor, then the signal amplification offered by SMaSh would provide the greatest sensitivity. If exogenous expression is not objectionable, then any assay will work; the choice is then limited to the cell or tissue of interest and how efficiently the expression construct can be delivered, either transiently or stably and/or with constitutive or conditional expression.
• For in vitro assays, what are the growth characteristics of the cultured cells used? Target engagement assays are usually conducted with wash steps to remove excess extracellular compound prior to the addition of tracer. Adherent cells are preferred for plate-based assays requiring washes, but centrifugation steps could be substituted for suspension or semi-adherent cells plated to compatible lower density (96- or 384-well) plates, as performed for flow cytometry. We recommend using adherent cell lines for high-density plate formats, such as the 1536-well approach presented here for MK2.
The two plate-based methods presented were developed for 1536-well format to improve throughput for screening campaigns, but the protocol is scalable to lower density formats. We recommend incorporation of automation for the wash cycles but acknowledge that availability of such equipment may limit the plate format that can be used for these assays, so a compromise must be made between throughput and precision.
Many programs seek to understand compound bioavailability in human blood, which often requires developing assays involving whole blood from health volunteers. Adsorption of compound to plasma proteins impacts bioavailability and is often measured using dialysis methods.22 By modifying our NanoBRET assay to present compound in the presence of up to 50% human serum, we established a simplified model of a whole-blood assay that allowed us to track the effect of plasma proteins on compound potency. While this configuration does not fully mimic a human whole blood assay, it serves as a practical proxy for assessing target engagement in the presence of human serum in a high-throughput format—an approach that is particularly valuable for targets enriched in blood cells. Such assays exemplify the versatility of these high-throughput target engagement assays for drug discovery programs.
Author contributions
Julian L. Wong – conception, investigation, methodology, visualization, and writing. Mari Manuia – investigation, analysis, and methodology. Sandra Gao – investigation and analysis. Natacha Stoehr – investigation and analysis. Theresa Boersig – investigation, analysis, and visualization. Krystine Vuong – investigation. Tao Jiang – investigation. Jian Cao – investigation. Yong Jia – supervision and review. C. C. King – investigation. John Joslin – resources. Leslie Ofori – supervision. Jon Loren – supervision, analysis, and review. Zuni I. Bassi – supervision, analysis, and writing.Conflicts of interest
All authors were employees and shareholders of Novartis at the time this work was performed.Abbreviations
| biotin|cov | Covalent tracer with a biotin handle |
| BODIPY585|cov | Covalent tracer with BODIPY 585 |
| HTRF | Homogenous time-resolved fluorescence |
| NanoBRET | Nanoluciferase bioluminescence resonance energy transfer |
| OC50 | Concentration at which 50% target occupancy is observed |
| SMaSh | Streptavidin mass shift |
Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Fig. S1. SMaSh target occupancy of CC-99677. Fig. S2. Measurement of endogenous MK2 in mouse tissue by dual HTRF. Fig. S3. Paralog specificity of BODIPY585|cov tracer in the NanoBRET assay. Fig. S4. Schematic comparison of the three target engagement assays. See DOI: https://doi.org/10.1039/d5cb00224a.Acknowledgements
We thank Danielle Boesch for contributing to the development of the SMaSh target occupancy assay as well as Nicole Von Burg, Melanie Vogelsanger, and Michel Schoenboerner for providing mouse tissues.References
- J. Singh, The Ascension of Targeted Covalent Inhibitors, J. Med. Chem., 2022, 65(8), 5886–5901, DOI:10.1021/acs.jmedchem.1c02134 From NLM Medline.
- S. St John-Campbell and G. Bhalay, Target Engagement Assays in Early Drug Discovery, J. Med. Chem., 2025, 68(12), 12331–12368, DOI:10.1021/acs.jmedchem.4c03115 From NLM Medline.
- N. Ronkina and M. Gaestel, MAPK-Activated Protein Kinases: Servant or Partner?, Annu. Rev. Biochem., 2022, 91, 505–540, DOI:10.1146/annurev-biochem-081720-114505.
- M. Fiore, S. Forli and F. Manetti, Targeting Mitogen-Activated Protein Kinase-Activated Protein Kinase 2 (MAPKAPK2, MK2): Medicinal Chemistry Efforts To Lead Small Molecule Inhibitors to Clinical Trials, J. Med. Chem., 2016, 59(8), 3609–3634, DOI:10.1021/acs.jmedchem.5b01457 From NLM Medline.
- A. Schlapbach and C. Huppertz, Low-molecular-weight MK2 inhibitors: a tough nut to crack!, Future Med. Chem., 2009, 1(7), 1243–1257, DOI:10.4155/fmc.09.98 From NLM Medline.
- R. Gaur, K. A. Mensah, J. Stricker, M. Adams, A. Parton, D. Cedzik, J. Connarn, M. Thomas, G. Horan and P. Schafer, et al., CC-99677, a novel, oral, selective covalent MK2 inhibitor, sustainably reduces pro-inflammatory cytokine production, Arthritis Res. Ther., 2022, 24(1), 199, DOI:10.1186/s13075-022-02850-6 From NLM Medline.
- J. Malona, C. Chuaqui, B. M. Seletsky, L. Beebe, S. Cantin, D. V. Kalken, K. Fahnoe, Z. Wang, B. Browning and H. Szabo, et al., Discovery of CC-99677, a selective targeted covalent MAPKAPK2 (MK2) inhibitor for autoimmune disorders, Transl. Res., 2022, 249, 49–73, DOI:10.1016/j.trsl.2022.06.005 From NLM Medline.
- D. Gordon, E. T. Hellriegel, H. R. Hope, D. Burt and J. B. Monahan, Safety, Tolerability, Pharmacokinetics, and Pharmacodynamics of the MK2 Inhibitor ATI-450 in Healthy Subjects: A Placebo-Controlled, Randomized Phase 1 Study, Clin. Pharmacol., 2021, 13, 123–134, DOI:10.2147/CPAA.S305308 From NLM PubMed-not-MEDLINE.
- D. Gordon, A. Kivitz, A. Singhal, D. Burt, M. C. Bangs, E. E. Huff, H. R. Hope and J. B. Monahan, Selective Inhibition of the MK2 Pathway: Data From a Phase IIa Randomized Clinical Trial in Rheumatoid Arthritis, ACR Open Rheumatol., 2023, 5(2), 63–70, DOI:10.1002/acr2.11517 From NLM PubMed-not-MEDLINE.
- C. Wang, S. Hockerman, E. J. Jacobsen, Y. Alippe, S. R. Selness, H. R. Hope, J. L. Hirsch, S. J. Mnich, M. J. Saabye and W. F. Hood, et al., Selective inhibition of the p38alpha MAPK-MK2 axis inhibits inflammatory cues including inflammasome priming signals, J. Exp. Med., 2018, 215(5), 1315–1325, DOI:10.1084/jem.20172063 From NLM Medline.
- Y. Jia, C. M. Quinn, A. Clabbers, R. Talanian, Y. Xu, N. Wishart and H. Allen, Comparative analysis of various in vitro COT kinase assay formats and their applications in inhibitor identification and characterization, Anal. Biochem., 2006, 350(2), 268–276, DOI:10.1016/j.ab.2005.11.010 From NLM Medline.
- D. Stokoe, D. G. Campbell, S. Nakielny, H. Hidaka, S. J. Leevers, C. Marshall and P. Cohen, MAPKAP kinase-2; a novel protein kinase activated by mitogen-activated protein kinase, EMBO J., 1992, 11(11), 3985–3994, DOI:10.1002/j.1460-2075.1992.tb05492.x From NLM Medline.
- M. T. Labenski, L. A. Bateman, L. T. Voortman, G. Giammo, S. Cantin, L. Qiao and A. F. Corin, SMaSh: A Streptavidin Mass Shift Assay for Rapidly Quantifying Target Occupancy by Irreversible Inhibitors, Biochemistry, 2021, 60(39), 2915–2924, DOI:10.1021/acs.biochem.1c00422 From NLM Medline.
- M. B. Robers, M. L. Dart, C. C. Woodroofe, C. A. Zimprich, T. A. Kirkland, T. Machleidt, K. R. Kupcho, S. Levin, J. R. Hartnett and K. Zimmerman, et al., Target engagement and drug residence time can be observed in living cells with BRET, Nat. Commun., 2015, 6, 10091, DOI:10.1038/ncomms10091 From NLM Medline.
- J. Dopfer, J. D. Vasta, S. Muller, S. Knapp, M. B. Robers and M. P. Schwalm, tracerDB: a crowdsourced fluorescent tracer database for target engagement analysis, Nat. Commun., 2024, 15(1), 5646, DOI:10.1038/s41467-024-49896-5 From NLM Medline.
- H. Sahoo, Förster resonance energy transfer - A spectroscopic nanoruler: Principle and applications, J. Photochem. Photobiol., C, 2011, 12(1), 20–30, DOI:10.1016/j.jphotochemrev.2011.05.001.
- H. Y. Jin, Y. Tudor, K. Choi, Z. Shao, B. A. Sparling, J. G. McGivern and A. Symons, High-Throughput Implementation of the NanoBRET Target Engagement Intracellular Kinase Assay to Reveal Differential Compound Engagement by SIK2/3 Isoforms, SLAS Discovery, 2020, 25(2), 215–222, DOI:10.1177/2472555219893277 From NLM Medline.
- N. Ronkina and M. Gaestel, MAPK-Activated Protein Kinases: Servant or Partner?, Annu. Rev. Biochem., 2022, 91, 505–540, DOI:10.1146/annurev-biochem-081720-114505 From NLM Medline.
- C. W. Li and G. V. Korshin, Studies of metal-binding sites in natural organic matter and their role in the generation of disinfection by-products using lanthanide ion probes, Chemosphere, 2002, 49(6), 629–636, DOI:10.1016/s0045-6535(02)00353-3 From NLM Medline.
- W. Ouyang, Q. Niu, M. Qui, H. Fu, Y. Du and X. Mo, A multiplexed time-resolved fluorescence resonance energy transfer ultrahigh-throughput screening assay for targeting SMAD4-SMAD3-DNA complex, bioRxiv, 2023 DOI:10.1101/2023.07.15.549169 From NLM PubMed-not-MEDLINE.
- D. Koley and A. J. Bard, Triton X-100 concentration effects on membrane permeability of a single HeLa cell by scanning electrochemical microscopy (SECM), Proc. Natl. Acad. Sci. U. S. A., 2010, 107(39), 16783–16787, DOI:10.1073/pnas.1011614107 From NLM Medline.
- L. Di, An update on the importance of plasma protein binding in drug discovery and development, Expert Opin. Drug Discovery, 2021, 16(12), 1453–1465, DOI:10.1080/17460441.2021.1961741 From NLM Medline.
| This journal is © The Royal Society of Chemistry 2026 |