Interaction of solid lipid nanoparticles with bovine serum albumin: physicochemical mechanistic insights†
Jyoti
Rathee
and
Nand
Kishore
 *
*
Department of Chemistry, Indian Institute of Technology Bombay, Powai, Mumbai 400076, India. E-mail: nandk@chem.iitb.ac.in
First published on 18th February 2025
Abstract
This study investigates the interaction of solid lipid nanoparticles (SLNs) with the transport protein bovine serum albumin (BSA) in terms of thermodynamic signatures, employing both spectroscopic and calorimetric techniques. When nanoparticles are exposed to biological media, proteins are adsorbed on their surfaces, leading to protein corona formation. Therefore, controlling the formation of the protein corona is essential for in vivo therapeutic efficacy. Although SLNs have previously been explored solely as potential nano-carriers for drug delivery, no prior efforts have been made to study their interactions with biomolecules from a biophysical and mechanistic perspective. SLNs are colloidal dispersions of the solid lipid in an aqueous solution stabilized by surfactants. Herein, a hot emulsification methodology was employed to formulate SLNs, and their interactions with BSA were analyzed. The SLNs were characterized using transmission electron microscopy (TEM) and dynamic light scattering (DLS) techniques to obtain information on their size, zeta potential, and shape. Fluorescence data suggested the presence of weak interactions between the SLNs and BSA. Static quenching is confirmed using time-correlated single-photon counting (TCSPC) experiments. Differential scanning calorimetric (DSC) and fluorescence spectroscopic experiments suggest the thermal stabilization of BSA by the SLNs. This stabilization results from the enhancement of the secondary structure of the protein without significantly altering the tertiary structure. Isothermal calorimetry (ITC) results suggest weak interactions between the SLNs and BSA, although not in a site-specific manner. Overall, mechanistic insights into lipid nanoparticle–protein interactions obtained from such studies efficiently overcome the hurdles associated with targeted drug delivery.
1. Introduction
Advancement in nanotechnology has opened up a new arena of research based on drug-delivery modules. It has been reported that a variety of nano-assemblies, such as liposomes,1 lipid nanoparticles,2 polymeric nanoparticles,3 and metal–organic frameworks,4 can offer enhanced therapeutics and biomedical applications. Among the diverse range of NPs employed in biomedical applications, solid lipid nanoparticles (SLNs) have received significant attention owing to their biocompatibility, stability, higher encapsulation efficiency, and desirable drug delivery properties.5 SLNs are lipid-based nano-assemblies that consist of core–shell structures with a hydrophobic lipid core surrounded by a hydrophilic surfactant shell. SLNs have been utilized in the treatment of cancer, cystic fibrosis, and bacterial infections and have shown promising results in terms of stability, encapsulation efficiency, and targeted delivery.6–9The leading interest in the biomedical and therapeutic applications of smart nanomaterials underscores the need for comprehensive analysis of their interactions with biologically important systems, such as proteins.10 To investigate the complex nature of interfacial processes occurring in living systems, a multi-analytical approach is required to characterize the associated interactions.11,12 The lack of adequate data limits the efficacy of biomaterials in diagnostic and targeted therapeutics, including molecular, cellular, and tissue targeting. Therefore, it is crucial to understand the characteristics of bio-interfaces that contribute to the design of nanostructured materials with enhanced efficacy and fewer side effects.13,14
After administration of NPs, a diverse range of biomolecules, such as proteins, can adsorb onto their surface, leading to protein corona formation.15,16 Initially, NPs under physiological conditions are coated with a firmly attached layer of proteins known as a hard corona. Thereafter, the assembly is further covered by weakly bound proteins, which are in dynamic equilibrium and form a soft corona.17,18 The formed protein corona is influenced by several factors, such as the physicochemical properties of the nano modules, the composition of the protein corona and temperature.
With few exceptions, protein corona formation occurs in almost all known NPs. The interaction of various molecules with the protein corona leads to changes in size, surface potential, and conformational changes in the protein.19 Thus far, only a few NPs have passed clinical trials and are commercially available. The nature and composition of the biomolecular corona surrounding the NPs can influence their biological fate, immune response, toxicity, and therapeutic efficacy.20 The internalization and excretion process of the NPs is also influenced by the protein layer, which affects the biological activity of the nanoparticles.21 Before clinical applications, an in-depth understanding of the interactions of the NPs with plasma protein is crucial for assessing the potential side effects of such systems.22
Serum albumin is the most abundant protein in plasma accounting for about 60% of the protein content. It acts as a transport agent owing to its important role in most of the physiological functions helping in the distribution and transportation of exogenous and endogenous molecules.23 Albumin has already been employed in the formulation of albumin-based nanoparticles for drug delivery applications. BSA-conjugated silver nanoparticles loaded with hyaluronic acid (HA)–polyethylene glycol (PEG) have been reported to show excellent antibacterial activity against Gram-positive/Gram-negative bactericides. Release studies showed that a significant amount of the nanoparticles remained in the matrix even after 14 days owing to the presence of the interactions between the nanoparticles and hydrogel.24
The biological environment surrounding NPs varies with different administration routes depending on the specific diseases. Moreover, NPs with a protein corona are cleared more rapidly by the phagocytic system. Additionally, the adsorption might lessen the NP cellular absorption by reducing their cell membrane adherence. The drug release profile, bio-distribution, and biodegradation are impacted by the formation of protein corona.25 Thus, an understanding of the development of the protein corona in terms of interactions between protein and SLNs is considered one of the keystones in filling the gap between the in vitro design and in vivo therapeutic effects.
Therefore, to regulate the formation of the protein corona, it is imperative to understand the interactions between proteins and NPs. Numerous studies have been conducted to demonstrate how protein corona formation occurs in silica,26 gold,27 and polymeric nanoparticles.20
Recently, Maity et al.28 reported the significant role of the lipid coronas (hard and soft) in the aggregation of the amino acid-functionalised gold NPs in the presence of the lysozyme protein. It was found that human serum albumin (HSA) conjugated amino acid functionalised gold NPs and citrate functionalized gold NPs are more effective in preventing the lysozyme-induced aggregation of the gold NPs compared to the BSA-conjugated gold NPs. Further, the detailed mechanism of the BSA protein interaction with the silver NPs was studied by Dasgupta et al.29 Side-on or end-on interaction of the 1.5 molecules of the BSA with silver nanoparticles was reported. Similarly, the interactions of the ZnO/Ag NPs with the BSA protein with the help of spectroscopic techniques have been studied. The obtained thermodynamic parameters indicated that the binding of the BSA with the ZnO/Ag was spontaneous and enthalpy driven. van der Waals forces and hydrogen bonding play a major role in the interaction between BSA and ZnO/Ag NPs.30 Later on, the binding interactions of 7-(N,N′-diethylamino) courmarin-3-carboxylic acid (7-DCCA) with BSA in the presence and absence of graphene oxide (GO) were explored. The addition of GO restricted the environment surrounding the 7-DCCA molecule in the presence of low and high concentrations of the BSA. ITC results showed higher binding interactions in the presence of GO.31 In a recent report by Maity et al.,32 the diverse role of the buffer molecules of different charges (at particular pH) in the interactions of the phenyl alanine functionalised gold NPs with the lysozyme protein, particularly in protein-induced NPs aggregation, were studied. The aggregation of both phenyl alanine functionalised gold NPs and lysozyme was reported in the negatively charged buffer (citrate and phosphate), whereas the positively charged buffer (HEPES, MOPS and Tris) and zwitterionic buffer only caused the aggregation of the protein.32
The quantitative mechanistic insights into the interaction of the SLNs and protein are largely missing. Analyzing the effect on the transport system can pave the way for further development and advancement of nanosystems in biomedical applications.
Herein, thermodynamic characterization of the protein corona formation of SLNs with BSA is attempted. SLNs in the size range of 100–150 nm were prepared using the emulsification method and subjected to interaction with BSA at pH 7.4. Further, transmission electron microscopy (TEM) and dynamic light scattering (DLS) techniques are employed to obtain the shape and size of the prepared systems. The interaction of BSA with the lipid nanoparticles is studied using UV-visible spectroscopy, fluorescence spectroscopy, time-correlated single photon counting (TCSPC), and circular dichroism (CD) spectroscopy in amalgamation with calorimetric analysis to explore the development and applications of SLNs.
2. Experimental section
2.1. Materials
Potassium phosphate (>99%), pluronic F68 (>99%), and fatty acid-free bovine serum albumin (>99%) were purchased from Sigma-Aldrich. Glyceryl monostearate (GMS) (>98%) was procured from TCI chemicals. Phosphate buffer (20 × 10−3) mol dm−3 was prepared in double distilled deionised water. Before the preparation of the solutions, degassing of the buffer was performed to remove the dissolved gases. A Sartorius BP 211 digital balance with ±0.01 mg readability was used for the mass measurements. The concentration of BSA was determined using a Jasco V550 double-beam spectrophotometer based on its extinction coefficient corresponding to E1%280 = 6.8.332.2. Preparation of solid lipid nanoparticles (SLNs)
The hot emulsification method was employed for the preparation of the SLNs with slight modifications.34,35 Briefly, glyceryl monostearate (GMS, 4% w/w) was heated at 75 °C for 15 min. After complete melting, a 0.05% w/w aqueous solution of pluronic 68 was added to the lipid phase and stirred for 20 min. Thereafter, the obtained hot dispersion was added to the cold buffer dropwise and stirred at 500 rpm for 20 min. Furthermore, centrifugation was performed at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 15 min. The cooling step promotes the formation of SLNs. The resulting nanoparticle suspension was stored at 298 K for further experiments.
000 rpm for 15 min. The cooling step promotes the formation of SLNs. The resulting nanoparticle suspension was stored at 298 K for further experiments.
2.3. Dynamic light scattering, zeta potential, and transmission electron microscopy
The hydrodynamic diameter and zeta potential of the fabricated SLNs in the absence and presence of the protein were determined using a Brookhaven DLS particle size analyser at 298 K. A 12 μM BSA solution prepared in the phosphate buffer at pH 7.4 was incubated with the SLNs for a period of 24 h. All the samples were filtered using a 0.45 μm pre-size microfilter. The hydrodynamic diameter of the SLNs was analyzed based on the concept of Brownian motion.36 The correlation between the Brownian motion and hydrodynamic diameter (dH) is given by the Stokes–Einstein relation (eqn (1)). Each measurement was repeated three times with a laser wavelength of 633 nm, medium viscosity of 0.8872 cP, and refractive index of 1.33.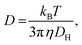 | (1) |
The surface charge on the SLNs is determined by the zeta potential (ζ) measurements using the Nano Brook Omni instrument, which measures the zeta potential based on electrophoretic mobility. The correlation between the electrophoretic mobility (μe) and zeta potential (ζ) is given by the following equation (eqn (2)):37
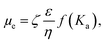 | (2) |
To examine the shape of the prepared systems, TEM images were taken using an electron microscope (FEI Tecnai G2, F30). The accelerating voltage was set at 300 kV. For sample preparation, the stock solution of the lipid nanoparticles with and without BSA was diluted and drop-cast over mesh copper grids. Later on, the samples were air-dried in a desiccator for a minimum of 24 h before being subjected to analysis.
2.4. Fluorescence spectroscopy
Steady-state fluorescence measurements were performed using a Cary Eclipse spectrophotometer with a quartz cell of path length 1 cm at 298 K. The excitation and emission slit widths were fixed at 5 nm each. The excitation wavelength was fixed at 295 nm, and the emission spectra were recorded in the wavelength range of 310–550 nm. The BSA concentration was kept at 4 μM, while that of SLNs varied during the experiment. The SLNs concentration was determined based on the lipid concentration in the solution. The fluorescence emission spectrum of the lipid was also recorded as a control experiment. The quenching of BSA using SLNs was examined to decipher the interaction of the lipid nanoparticles with the protein. Each experiment was carried out at least three times to establish the reproducibility of the measurements.A Horiba (Fluoromax + (RM 360)) spectrofluorometer was employed to study the thermal denaturation behavior of BSA. The concentration of BSA was fixed at 4 μM and excited at 295 nm. Further, the emission fluorescence spectra were recorded in the temperature range of 25–95 °C. The experiments were performed at molar ratios of the BSA to SLNs at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0, 1
0, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, and 1
5, and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20. The EXAM software of Kirchhoff39 was employed for extracting thermodynamic parameters such as transition midpoint (Tm) and enthalpy change (ΔHm) accompanying unfolding of BSA.
20. The EXAM software of Kirchhoff39 was employed for extracting thermodynamic parameters such as transition midpoint (Tm) and enthalpy change (ΔHm) accompanying unfolding of BSA.
2.5. Time correlation single photon counting (TCSPC)
A Horiba Jobin Yvon (Fluorocube) was employed to carry out the time-resolved fluorescence experiments. The excitation and emission wavelengths of the BSA were fixed at 295 nm and 350 nm, respectively. The value of FWHM was 68.8 ps. The fluorescence decay data were fitted bi-exponentially by employing an iterative reconvolution methodology with the DAS 6.0 software from IBH, UK. The goodness of the fit was denoted in terms of χ2 parameters.The experiment was performed at BSA to SLNs molar ratios of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0, 1
0, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, and 1
5, and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 with a protein concentration of 4 μM at 298 K. TCSPC experiments were performed to assess the static or dynamic nature of quenching in complex formation.40
20 with a protein concentration of 4 μM at 298 K. TCSPC experiments were performed to assess the static or dynamic nature of quenching in complex formation.40
The bi-exponential model is employed to analyze the intensity of decay as the sum of individual single exponential decays expressed in the following equation (eqn (3)):
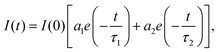 | (3) |
The average lifetime (τ) was calculated using the following equation (eqn (4)):
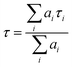 | (4) |
2.6. UV-visible spectroscopy
A double-beam spectrophotometer was used to carry out absorbance measurements of the protein in the absence and presence of the SLNs. The spectra were recorded in the wavelength range of 200–600 nm at 298 K with a scan rate fixed at 400 nm min−1. Each experiment was carried out at least thrice to assess for reproducibility of the results.2.7. Circular dichroism (CD) spectroscopy
To elucidate changes in the secondary and tertiary structures of the protein, CD spectroscopic experiments were performed. A Jasco-810 spectropolarimeter was employed to perform circular dichroism (CD) experiments. To scan in the far UV (260–190) and near UV (320–260 nm) CD regions, the cells with path lengths of 0.2 cm and 1 cm, respectively, were used. The concentrations of BSA in far UV CD and near UV CD measurements were 4 μM and 15 μM, respectively. Purging of the nitrogen gas was done thoroughly during the experiments. The CD spectra were measured at a scan rate of 100 nm min−1 with 1 second response time. The baseline correction was performed for each spectrum, and the final spectra reported are an average of the three accumulations. The measured ellipticity (θ) was converted into molar ellipticity [θ] using the following equation (eqn (5)):41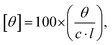 | (5) |
2.8. Differential scanning calorimetry (DSC)
A Nano DSC procured from a TA instrument with a cell volume of 300 μL was employed to study the thermal behaviour of the BSA upon the addition of SLNs. The concentration of BSA was fixed, corresponding to 3 mg mL−1, while that of SLNs varied. A scan rate of 1 °C min−1 was fixed, and heating was performed in the temperature range of 25–95 °C. For the baseline, the reference cell and the sample cell were filled with buffer and scanned at the same rate in the temperature range used in the other experiments. After subtraction of the buffer versus buffer scan from the protein versus buffer scans, the excess heat capacity of the protein versus the temperature plot was obtained. The baseline-corrected results were analyzed using Nano Analyse software with a Gaussian model to extract the thermodynamic parameters accompanying the protein unfolding.2.9. Isothermal titration calorimetry (ITC)
A Nano ITC procured from TA instruments with a cell volume of 300 μL and syringe volume of 50 μL was used to measure heat changes accompanying the interaction of BSA with SLNs. BSA (0.03 mM) was taken in the sample cells and titrated with subsequent injections of SLNs (0.8 mM) from the syringe. The experiments were designed for a total of 24 injections: 2 μL each spaced at an interval of 300 s. The reference cell was filled with buffer, and the solution in the sample cell was stirred at a rate of 350 rpm. The titration did not result in a typical sigmoidal binding isotherm. Based on the trends observed in the values of the heat of interaction (q), limiting standard molar enthalpies of the interaction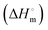 of SLNs with the BSA were obtained by fitting second-order polynomials to the data points in the ORIGIN (eqn (6)):
of SLNs with the BSA were obtained by fitting second-order polynomials to the data points in the ORIGIN (eqn (6)):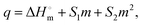 | (6) |
2.10. Statistical analysis
Data are presented as mean ± standard deviation (SD). All experiments were carried out at least three times. Statistical analysis was performed using a one-way analysis of variance (ANOVA) in Excel. A p-value of less than 0.05 was considered to indicate statistical significance.3. Results and discussion
The SLNs were successfully prepared using a hot emulsification methodology by employing biocompatible excipients such as glyceryl monostearate (GMS) as the lipid phase and PF-68 (surfactant). GMS was employed as a core material for the SLNs. To formulate SLNs, GMS was selected owing to its less ordered crystal structure, higher loading efficiency, and improved stability.42 The melting point of the GMS is approximately 75 °C. Therefore, during the preparation of the SLNs, the temperature was maintained at 80 °C to achieve a molten state of the lipid. Further, the prepared systems were stabilized using Pluronic F-68 (PF-68) with a hydrophilic lipophilic balance value of 29. PF 68 is listed as an “inactive ingredient” by the U.S. Food and Drug Administration (FDA) in its GRAS (generally regarded as safe) ingredients.43 The surface hydrophobicity or hydrophilicity of the SLNs depends on the nature and composition of the employed excipients. Here, in our formulation, glyceryl monostearate (GMS) lipid and PF 68 are used to prepare the SLNs. Consequently, owing to the hydrophobic nature of the GMS lipid, it is anticipated that SLNs are primarily hydrophobic. Furthermore, PF 68 incorporated in formulation contributes to the colloidal stability of the SLNs although it imparts hydrophilic character owing to several –O– and –OH groups. Density affects the dynamic properties of the formulation, including stability, release behaviour, and viscosity. Density is defined as the quotient of the weight-to-volume ratio of the system. Herein, the density of the SLNs was found to be (0.99823 ± 0.0001) g cm−3 close to the density of the deionized water/buffer (pH 7.4). High-density differences between the SLNs formulation and the dispersing medium can accelerate sedimentation, which influences the stability of the formulation.443.1. Hydrodynamic and zeta potential analysis
The values of the hydrodynamic diameter and zeta potential were determined using DLS measurements. A comparison of the hydrodynamic diameter of SLNs can shed light on the protein corona formation. The hydrodynamic diameter of the SLNs was found to be 350 ± 10 nm, and that of the SLNs–BSA complex was found to be 362 ± 14 nm (Fig. 1). Herein, the hydrodynamic diameter of the SLNs was found to be nearly the same irrespective of whether or not they were incubated in the protein solution. The results signify no change in the SLNs structure or dynamics after exposure to the plasma protein. Moreover, the results indicate that there is no major adsorption of BSA on SLNs. Similar results are reported by Alberg and coworkers45 in which the size of the cross-linked polymeric nanoparticles after incubation with plasma protein was found to be almost similar to that of bare SLNs, suggesting that the nanoparticles remain intact and there is no significant adsorption of protein on their surface. Wang et al.15 reported that the surface hydrophobicity of the NPs significantly influences their interaction with proteins, with high hydrophilicity being a fundamental trait of materials possessing strong anti-protein adsorption capabilities. It was proposed that this anti-protein adsorption ability is closely linked to the hydration layer near the surface, which acts as both a physical and energetic barrier, preventing protein adsorption.The zeta potential of SLNs was found to be −(25 ± 2) mV (Fig. S1, ESI†). The value of the zeta potential of the SLNs was found to decrease after incubation with BSA compared to SLNs only, which indicates the interaction of BSA with SLNs.
3.2. Transmission electron microscopic (TEM) analysis
The spherical shape morphology of the nanoparticles was depicted by TEM results in the absence and presence of BSA. The average size of the prepared SLNs was found to be ∼(100 ± 4 nm) (Fig. 2(A)). After incubation of the SLNs with BSA, the size was found to be ∼(105 ± 4 nm) (Fig. 2(B)). The size of SLNs after incubation with the BSA is found to be almost similar to pristine SLNs, which suggests that there is less possibility of the protein corona layer on the surface of the nanoparticles determined using Image J software.46 The size of the SLNs obtained using DLS was not consistent with that obtained by TEM, which might be owing to the different mechanisms of the two methods.47 The DLS analysis was conducted in an aqueous state, where the particles were highly hydrated. Consequently, the measured diameter corresponds to the hydrodynamic diameter, which is normally larger than its actual particle size (obtained using TEM imaging).48 The particle sizes of SLNs and SLNs–BSA complex are determined using the resulting histograms shown in Fig. S2 (ESI†).3.3. Fluorescence spectroscopy
Fluorescence spectroscopy provides valuable information on conformational changes in the protein under different environmental conditions.49 To analyze the interaction of BSA with SLNs, fluorescence studies were performed. Upon excitation at 295 nm, BSA exhibited strong and stable intrinsic fluorescence owing to its two tryptophan residues (Trp-212 and Trp-134). Trp-212 is present in the hydrophobic pocket of subdomain IB near the surface of the BSA and Trp-212 is in the subdomain IIA.31,50 The fluorescence emission profiles of 4 μM BSA in the presence of SLNs are shown in Fig. 3(A). The black line corresponds to the emission spectra of the native BSA. The wavelength maxima (λmax) for native BSA was obtained at 346 nm, which is consistent with that reported in the literature.51 Ligand binding and associated conformational changes in the protein lead to changes in the fluorescence emission profile.52 A reduction in the fluorescence intensity of BSA was observed upon the addition of SLNs (0 to 75 μM). Quenching of the intrinsic fluorescence of the proteins upon the addition of ligands has a direct relation to the extent of their interactions.53 The observed changes in the fluorescence emission profile of BSA upon the addition of SLNs can result from interaction, including conformational change, if any. Additionally, the effect of only GMS on the BSA is depicted in Fig. S3 (ESI†). The values of the association constant and stoichiometry of binding (n) between SLNs and BSA were evaluated using the following logarithmic form Stern Volmer equation (eqn (7)):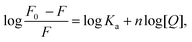 | (7) |
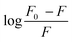 as a function of log[Q] yielded a straight line with a slope of n and intercept of log
as a function of log[Q] yielded a straight line with a slope of n and intercept of log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ka (Fig. 3(B)). The value of Ka was found to be (4.5 ± 0.1)× 103 M−1 with n approximately 1, suggesting a 1
Ka (Fig. 3(B)). The value of Ka was found to be (4.5 ± 0.1)× 103 M−1 with n approximately 1, suggesting a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type of interaction between the protein and SLNs.54 The value of Ka of the order of 103 indicates a weak affinity between SLNs and protein.52 Lipid–BSA complexes have been reported with similar low-affinity constant values. Cholesterol and dioleoylphosphatidylethanolamine (DOPE) lipids interact with human serum albumin with Ka values of the order of 102.52
1 type of interaction between the protein and SLNs.54 The value of Ka of the order of 103 indicates a weak affinity between SLNs and protein.52 Lipid–BSA complexes have been reported with similar low-affinity constant values. Cholesterol and dioleoylphosphatidylethanolamine (DOPE) lipids interact with human serum albumin with Ka values of the order of 102.52
 | ||
Fig. 3 (A) Fluorescence emission spectra of bovine serum albumin (BSA) with increasing concentrations of SLNs at 298 K and pH 7.4 and (B) plot of 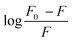 as a function of SLNs concentration [Q]. as a function of SLNs concentration [Q]. | ||
A decrease in the fluorescence intensity of the BSA in the presence of the SLNs cannot be attributed to the formation of the protein corona on the surface of the SLNs. A recent work by Cedervall and co-workers55 mentioned that for the formation of the protein corona, there must be adsorption of at least from 10 to 100 proteins on the surface of the ligand. These results were also supported by Casals and co-workers.56 Therefore, as per the literature, for the formation of protein corona, the expected protein/nanoparticles ratio of n must always be greater than 1.57 Therefore, the formation of the protein corona cannot be established based only on a decrease in the fluorescence intensity.
The decrease in the fluorescence intensity of the BSA after the addition of the SLNs is shown in Fig. 4. A blue shift of 18 nm in the intrinsic fluorescence emission profile of BSA upon the addition of SLNs indicates a change in the microenvironment around the tryptophan residues. This blue shift indicates the internalization of the tryptophan residues in BSA to a more nonpolar environment upon adding the SLNs.58,59 To ascertain the nature of the quenching process, fluorescence lifetime experiments were performed, which confirmed the occurrence of static quenching by SLNs.
3.4. Time-correlated single-photon counting (TCSPC)
Lifetime studies were carried out to monitor the nature of the fluorescence quenching of BSA in the presence of SLNs. After fitting a bi-exponential equation to the decay profiles, two lifetimes for the fluorescence decay were obtained, one at τ1 = 1.52 ± 0.07 ns and the other at τ2 = 6.22 ± 0.03 ns, which are in good agreement with those reported in the literature.60 The component of 6.22 ns refers to Trp-134, while that at 1.52 ns corresponds to the Trp-212 moiety of the protein. Trp-134 is located in a hydrophilic environment, which is responsible for a longer lifetime than Trp-212, which lies in the hydrophobic core.61The local environment of a fluorophore can be examined by employing fluorescence lifetime measurements. The static or dynamic nature of quenching can be differentiated based on the change in the lifetime of the fluorophore upon the addition of the quencher. The decay profiles for the lifetime measurement of BSA in the presence of the SLNs at different mole ratios are shown in Fig. 5 and presented in Table 1.
| [BSA]/[SLNs] | Lifetime (ns) | Amplitude | Average lifetime (ns) | χ 2 | ||
|---|---|---|---|---|---|---|
| τ 1 | τ 2 | a 1 | a 2 | τ | ||
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 0 |
1.52 ± 0.07 | 6.22 ± 0.03 | 0.41 ± 0.01 | 0.58 ± 0.01 | 4.27 ± 0.09 | 1.09 ± 0.02 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
1.50 ± 0.06 | 5.55 ± 0.04 | 0.51 ± 0.02 | 0.48 ± 0.02 | 3.46 ± 0.08 | 1.15 ± 0.02 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 5 |
1.50 ± 0.06 | 5.13 ± 0.04 | 0.54 ± 0.01 | 0.45 ± 0.02 | 3.14 ± 0.07 | 1.12 ± 0.01 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
1.50 ± 0.06 | 5.19 ± 0.04 | 0.53 ± 0.02 | 0.46 ± 0.01 | 3.20 ± 0.07 | 1.12 ± 0.02 |
Static quenching is characterized by τ0/τ = 1, where the lifetime remains almost the same with the increase in concentration of the quencher. In contrast, dynamic quenching is accompanied by F0/F = τ0/τ.62 It is observed that the average lifetime of the tryptophan residue varies in a small range of 3.02–4.27 ns. This slight change in the lifetime suggests that the contribution of the excited state in the overall complex formation is not appreciable. Here, τ0/τ is observed to be close to 1 at different mole ratios of the SLNs to BSA. These results indicate the predominance of the static quenching in the interactions of the SLNs with the protein.
3.5. UV-visible spectroscopy
To further understand SLNs–BSA interaction and conformational change in the protein, UV-visible spectra of BSA in the presence of SLNs were examined. Two absorption peaks of BSA are obtained in the UV-vis spectrum (Fig. 6(A)). The more intense peak at 210 nm (far UV region) is due to its polypeptide backbone (π → π* transition), while the less intense peak at 280 nm (near UV region) corresponds to aromatic amino acid residues (Trp, Phe, and Try) and disulfide bonds of the paired cysteine residues of BSA.63 The addition of SLNs leads to a decrease in the peak intensity at 210 nm, along with the redshift. The presence of redshift denotes a decrease in polarity around the microenvironment of the Trp residues owing to the compaction of the structure or driving away of locally interacting water molecules and moving towards a more hydrophobic environment. Thus, the red shift observed in the UV-visible spectra of BSA upon the addition of SLNs results from the conformational change in the protein.64 | ||
Fig. 6 (A) Ultraviolet-visible absorption spectra and (B) plot of 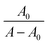 vs. 1/[Q] of BSA with an increase in the concentration of SLNs at 298 K. vs. 1/[Q] of BSA with an increase in the concentration of SLNs at 298 K. | ||
Okada and co-workers65 also reported that redshift around 220 nm denotes the changing microenvironment of the Trp residue towards a more hydrophobic environment. The Benesi–Hildebrand equation (eqn (8)) was applied to determine the value of the association constant accompanying the SLNs–BSA interaction:
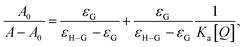 | (8) |
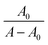 versus 1/[Q] is found to be linear, yielding a Ka value of (1.54 ± 1.1) × 104 M−1 (statistical significance: p < 0.05) from the ratio of intercept to the slope (Fig. 6(B)).
versus 1/[Q] is found to be linear, yielding a Ka value of (1.54 ± 1.1) × 104 M−1 (statistical significance: p < 0.05) from the ratio of intercept to the slope (Fig. 6(B)).
3.6. Thermal studies
 | ||
| Fig. 7 DSC profiles of 4 μM BSA and in the absence and presence of the SLNs heated at a scan rate of 1 °C min−1. | ||
| [BSA]/[SLNs] | T m/(°C) | ΔcalH/(kJ mol−1) |
|---|---|---|
| Native BSA | 53.0 ± 1.2 | 268 ± 5.0 |
| SLNs | 58.1 ± 1.0 | 938 ± 9.1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(a) 58.7 ± 0.1 | 1026 ± 9.0 |
| (b) 81.1 ± 0.1 | 461 ± 9.5 | |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 5 |
(a) 57.9 ± 0.2 | 1449 ± 30.0 |
| (b) 80.4 ± 0.3 | 434 ± 3.1 | |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
(a) 57.4 ± 0.1 | 1047 ± 21.0 |
| (b) 79.9 ± 0.8 | 547 ± 4.1 |
The α form of GMS has lower dense packing than its β form. Melting of the SLNs in our measurements corresponds to melting of α form of glycerol monostearate as its transition temperature is around (58.1 ± 1.0) °C. In the presence of the 4 μM SLNs, Tm of BSA was found to be at (81.0 ± 0.1) °C. Therefore, the protein is stabilized by (28.0 ± 1.0) °C with an enthalpy change (ΔΔH) of (461 ± 9.5) kJ mol−1. Similarly, upon increasing the concentration of lipid nanoparticles to 80 μM, BSA is stabilized by (28.0 ± 1.2) °C with an enthalpy change of (547 ± 4.1) kJ mol−1.68 This stabilization could result from an enhancement in the secondary structure of the protein upon interaction with the SLNs, leading to more compactness of the protein.
Foggia et al.69 reported that, at pH 7, upon the addition of lysozyme (up to 2% w/w), and BSA to the lecithin (DMPC) and cephalin (DMPE) liposomes, a decrease in the Tm of BSA was observed at lower lysozyme concentrations (up to 2% w/w). In contrast, an increase in the Tm of lysozyme was noted as its concentration increased from 2% to 15% w/w in the presence of liposomes. Simultaneously, an increase of approximately 20% in the enthalpy change (ΔH) of the transition was observed. A slight increase in the Tm of the BSA was also noted in the presence of liposomes, accompanied by an approximately 8% increase in the associated ΔH. The data indicate that the interaction between BSA, and the liposome occurs solely on the outer surface of the bilayer, without any penetration into the hydrophobic core of the liposomes. In contrast, with lysozyme present, there is evidence, suggesting partial penetration into the bilayer.
The values of the Tm and ΔvHHm of BSA in the presence of SLNs are illustrated in Table 3. The value of the Tm of BSA is found to increase from (54.3 ± 1.0) °C to (80.5 ± 1.2) °C with the change in ΔvHHm from (203 ± 4.1) kJ mol−1 to (266 ± 4.0) kJ mol−1 upon adding 4 μM SLNs. In general, the values of Tm and ΔvHHm are observed to increase with an increase in [BSA]/[SLNs] molar ratio. After increasing the concentration of the SLNs to 80 μM, the Tm was found to be (82.5 ± 1.2) and (84.2 ± 2.5) °C, and the heat changed from (271 ± 5.1) kJ mol−1 and (533 ± 7.5) kJ mol−1. These results demonstrate that SLNs impart thermal stability to BSA.
| [BSA]/[SLNs] | T m (°C) | ΔvHHm (kJ mol−1) |
|---|---|---|
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 0 |
54.3 ± 1.0 | 203 ± 4.1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
80.5 ± 1.2 | 266 ± 4.0 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 5 |
82.5 ± 1.3 | 271 ± 5.1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
84.2 ± 2.5 | 533 ± 7.5 |
3.7. Circular dichroism (CD) spectroscopy
Circular dichroism (CD) spectroscopy is a commonly utilized method to examine the conformational stability of proteins in aqueous solutions.The CD signatures signifying the secondary structure of proteins arise mainly owing to asymmetry around the peptide bond.72,73 Here, BSA exhibits two negative characteristic peaks at 208 nm and 222 nm, which are the signature peaks for α helix (Fig. 8). The far-UV CD spectra arise owing to n–π* transitions. The tertiary structure of a protein can be assessed from a near UV-CD spectrum based on a positive peak around 280 nm. This band arises owing to asymmetry around disulphide and aromatic chromophores, thereby holding valuable conformational information about the protein.74 Upon the addition of SLNs, a slight enhancement in the secondary structure of BSA (Fig. 8(A)) shows conformational change in the protein. The value of the percent α-helical structure in BSA with and without lipid nanoparticles was calculated using the following equation (eqn (9)):
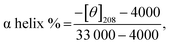 | (9) |
Native BSA has a (50.1 ± 0.8)% helical structure, which slightly increased to (55.4 ± 0.9)% with the addition of SLNs (Table 4). This suggests that the addition of the SLNs induces a slight conformational change in the protein, which leads to a change in the value of Tm from the DSC measurements. This suggests that the interaction of the SLNs with BSA leads to the stabilization of the BSA. However, the tertiary structure remains intact (Fig. 8(B)). Wang et al.15 also reported a slight increment in the BSA secondary structure in the presence of SLNs. The increment in the ellipticity values at 208 nm and 220 nm of BSA in the CD spectra was observed in the presence of SLNs composed of cetyl palmitate. Similarly, Kishore et al.75 reported that niosomes in the presence of ketoprofen provide extra thermal stability to protein. However, no major alterations were observed in the secondary and tertiary structures of the protein.
| [BSA]/[SLNs] | % α-helix | Change (%) |
|---|---|---|
| BSA only | 50.1 ± 0.8 | — |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
52.3 ± 0.4 | 4 ± 0.08 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 5 |
53.5 ± 0.6 | 5 ± 0.05 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
55.4 ± 0.9 | 10 ± 0.06 |
The secondary structure of the BSA changed upon the addition of the SLNs possibly via the exposure of photon absorbing amino acid residues or by the increment in the α-helical content of protein. The ellipticity of the bands in the near UV-CD region did not exhibit significant changes upon adding the lipid nanoparticles. Overall, it is observed that the native state of BSA is stabilized by SLNs, which suggests their biocompatibility and hence can safely be employed for drug delivery applications.
3.8. Isothermal titration calorimetry (ITC) analysis
The binding of a ligand to a protein can be driven mainly by hydrogen bonding, van der Waals, and electrostatic and hydrophobic interactions. Therefore, to understand SLNs–BSA interactions quantitatively, the accompanying thermodynamic signatures were obtained by ITC measurements. ITC is one of the best and most sensitive techniques that permits direct measurements of the thermodynamic signatures involved in the binding of a ligand to a protein.76,77The syringe of ITC was filled with 0.03 mM BSA and titrated with the 0.8 mM SLNs taken in the cell. The experiments were performed at pH 7.4 and 298 K. Fig. 9 shows a typical ITC profile under these conditions. As illustrated in Fig. 9, there is no distinct typical binding profile obtained for the SLNs–BSA interaction. In such a case, the only reliable extractable thermodynamic signature is the value of the enthalpy of interaction 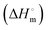 The data were analyzed based on the best-fitting polynomial approach (eqn (6) and Fig. 9). Relevant dilution control experiments were also performed, and corrections were made to the main titrations. Analysis of the dilution-corrected ITC profiles provided an interaction enthalpy
The data were analyzed based on the best-fitting polynomial approach (eqn (6) and Fig. 9). Relevant dilution control experiments were also performed, and corrections were made to the main titrations. Analysis of the dilution-corrected ITC profiles provided an interaction enthalpy 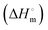 of (9.8 ± 0.2) kJ mol−1 (Fig. S4, ESI†). The p value was found to be less than 0.05. The value of Ka was obtained from fluorescence quenching experiments, which was used to obtain the value of
of (9.8 ± 0.2) kJ mol−1 (Fig. S4, ESI†). The p value was found to be less than 0.05. The value of Ka was obtained from fluorescence quenching experiments, which was used to obtain the value of 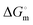 and
and 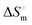 at 298 K (Table 5).
at 298 K (Table 5).
 | ||
| Fig. 9 ITC profile for the interaction of 0.8 mM SLNs with 0.03 mM of BSA at 298 K and pH 7.0. The mole ratio in the horizontal axis represents that of [SLNs] to [protein]. | ||
The interaction between the SLNs and BSA is found to be endothermic, indicating the dominance of hydrophobic interactions. The non-polar patches on the BSA surface can interact with the hydrophobic parts of the lipid nanoparticles.69 Further, the blue shift observed in the fluorescence spectra also suggests internalization of the tryptophan residue strengthening hydrophobic interactions within BSA upon interaction with SLNs.
Prashantan et al.70 reported the thermodynamic signatures accompanying the binding of pluronic F127 and F68 with BSA. The endothermic enthalpy of the interaction was attributed to the dominance of hydrophobic interactions. However, this can also result from the interaction of the hydrophobic chains of the surfactants with the non-polar patches on the surface of the BSA. Similarly, Thoppil et al.78 observed that the interaction of ionic liquids with BSA results in the loss of the integrity of the binding sites on BSA. It was concluded that both the cationic and anionic parts of the ionic liquids interacted with the positive and negative residues on the surface of BSA.
3.9. Mechanistic insights
The interaction of solid lipid nanoparticles (SLNs) with BSA is examined using an amalgamation of spectroscopic and calorimetric methods. SLNs are successfully formulated by employing a hot emulsification methodology using GMS (lipid phase) and stabilized with PF 68 (surfactant). The binding mode of the SLNs with BSA is examined using fluorescence spectroscopy. The value of the association constant (Ka) is found to be of the order of 103 M−1 with a blue shift of 18 nm in the fluorescence emission, indicating the weak interactions among lipid nanoparticles and protein through significant internalization of the tryptophan residues. Further, the occurrence of static quenching is confirmed by the TCSPC results. The average lifetime of the tryptophan is observed to be in the range of 3.02–4.27 ns. This suggests that the contribution of the complex formation in the excited state is not significant to fluorescence quenching upon the addition of SLNs. Further, the Benesi–Hildebrand equation provided the value of the association constant found to be in the order of 104, indicating weak interactions between the SLNs and BSA. Stabilization of BSA is observed in the presence of the SLNs, which suggests a conformational change to a more compact form supported by the blue shift observed in the fluorescence spectra. In the presence of 4 μM SLNs, the protein is stabilized by as much as (28.0 ± 1.0) °C in the presence of SLNs with an enthalpy change of (461 ± 9.5) kJ mol−1. To further compare the changes associated with the thermal unfolding of BSA upon the addition of SLNs, fluorescence thermal studies were also carried out. It was found that Tm increases from (54.3 ± 1.0) to (80.5 ± 1.2) °C with ΔvHH from (203 ± 4.1) to (266 ± 4) kJ mol−1 upon the addition of 4 μM SLNs to the 4 μM BSA. After increasing the concentration of the SLNs to 20 and 80 μM, the Tm of BSA is found to be (82.5 ± 1.2) and (84 ± 2.5) °C, and the value of ΔvHHm from (271 ± 5.1) and (533 ± 7.2) kJ mol−1, respectively. An increase in α-helicity in BSA in the presence of the SLNs is consistent with the DSC results. Thus, the interaction of the SLNs with BSA leads to a conformation change in the protein, which results in a more compact form, leading to the thermal stabilization of the protein.Because the interactions of the SLNs and BSA are weak, no distinct ITC binding profile is observed in ITC. The value of enthalpy of interactions is found to be (9.8 ± 0.2) kJ mol−1 and that of entropy is found to be (7.89 ± 0.5) J K−1 mol−1. These results suggest a predominance of hydrophobic interactions. Nonpolar patches interact with the hydrophobic portion of the lipid nanoparticles, resulting in desolvation and an increase in the randomness of the system. Overall, it can be concluded that there is a presence of weak interactions between the BSA and SLNs. However, no significant binding is observed between the SLNs and BSA, which means that there is less possibility of protein corona formation in the system (Fig. 10).
4. Conclusions
In this work, SLNs are successfully formed by employing GMS (as a solid lipid) and pluronic F68 (as a surfactant phase) using the hot emulsification methodology. The average sizes of the SLNs and SLN–BSA complex are about (100 ± 4) nm and (105 ± 4) nm, respectively. The size of SLNs after incubation with the BSA is found to be almost similar to pristine SLNs, which suggests that there is no significant adsorption of the protein on the SLNs.The energetics of the interactions of BSA with the SLNs were examined using spectroscopic and calorimetry techniques. The value of association constant of the order of 103 M−1 with a blue shift of 18 nm in the fluorescence emission profile of the BSA suggests that the weak interactions among SLNs and the protein lead to internalization of the tryptophan residue in BSA towards a nonpolar environment after the addition of the SLNs. The weak nature of the SLNs–BSA interaction is also inferred from UV-visible spectroscopic measurements.
Thermal stabilization of BSA in the presence of the SLNs suggests a conformational change of protein, leading to a more compact form, as indicated by the blue shift observed in the fluorescence spectra. Further, CD spectroscopy showed an increase in α-helicity in BSA upon the addition of SLNs. The ITC results demonstrated the predominance of hydrophobic interactions in the SLNs–BSA complex. Nonpolar patches of the BSA interact with the hydrophobic portion of SLNs, resulting in desolvation and an increase in the randomness in the system.
Overall, the results demonstrate that there is less possibility of protein corona formation on the surface of the lipid nanoparticles. It is known that the development of the protein corona on the surface of the nanoparticles poses a significant challenge to the bench-to-bedside approach of the targeted drug delivery systems both in terms of unfavorable distribution and by interfering with the targeting of the desired site. To design a targeted drug delivery system, the present work provides new insights into the mechanical aspects of comprehending the interactions of lipid nanoparticles with proteins.
Author contributions
Jyoti Rathee: methodology, conceptualization, investigation, data curation, formal analysis, writing – original draft and writing – review and editing; Nand Kishore: supervision, project administration, methodology, conceptualization, investigation, data curation, formal analysis, and writing – review and editing.Data availability
The data supporting the article have been included in ESI.†Conflicts of interest
The authors declare no conflicts of interest.Acknowledgements
J. R. acknowledges IIT Bombay for a post-doctoral fellowship. The authors acknowledge Indian Institute of Technology, Bombay for financial support. The authors are also thankful to the Central Facility of the Department of Chemistry at the Indian Institute of Technology Bombay, Mumbai, for providing instrument facilities.References
- J. F. Stefanick, D. T. Omstead, T. Kiziltepe and B. Bilgicer, Nanoscale, 2019, 11, 4414–4427 RSC.
- R. Tenchov, R. Bird, A. E. Curtze and Q. Zhou, ACS Nano, 2021, 15, 16982–17015 Search PubMed.
- A. Zielinska, F. Carreiró, A. M. Oliveira, A. Neves, B. Pires, D. Nagasamy Venkatesh, A. Durazzo, M. Lucarini, P. Eder, A. M. Silva, A. Santini and E. B. Souto, Molecules, 2020, 25, 3731 Search PubMed.
- S. Wan, Q. Cheng, X. Zeng and X. Zhang, ACS Nano, 2019, 13, 6561–6571 Search PubMed.
- S. S. Montoto, G. Muraca and M. E. Ruiz, Front Mol Biosci., 2020, 7, 1–24 CrossRef PubMed.
- H. L. Wong, R. Bendayan, A. M. Rauth, Y. Li and X. Y. Wu, Adv. Drug Delivery Rev., 2007, 59, 491–504 CrossRef CAS PubMed.
- M. Pastor, M. Moreno-sastre, A. Esquisabel, E. Sans, M. Viñas, D. Bachiller, V. José, Á. Del, E. Gainza and J. Luis, Int. J. Pharm., 2014, 477, 485–494 Search PubMed.
- S. M. Hosseini, R. Abbasalipourkabir, F. A. Jalilian, S. S. Asl and A. Farmany, Antimicrob. Resist. Infect. Control., 2019, 8, 1–12 CrossRef PubMed.
- S. Ghanbar, M. Fumakia, E. A. Ho and S. Liu, Nanomedicine, 2018, 14, 471–481 Search PubMed.
- S. Schwaminger, S. A. Blank-shim, M. Borkowska-panek, P. Anand, P. Fraga-garc, K. Fink, W. Wenzel and S. Berensmeier, ACS Omega, 2019, 4, 3790–3799 CrossRef CAS PubMed.
- M. M. Sozarukova, E. M. Kochneva, E. V. Proskurnina, I. V. Mikheev, D. O. Novikov, M. A. Proskurnin and V. K. Ivanov, ACS Biomater. Sci. Eng, 2023, 9(12), 6759–6772 CrossRef CAS PubMed.
- G. Yin, Z. Liu, J. Zhan, F. Ding and N. Yuan, Chem. Eng. J., 2002, 87, 181–186 CrossRef CAS.
- Y. Zhang, J. L. Y. Wu, J. Lazarovits and W. C. W. Chan, J. Am. Chem. Soc., 2020, 142, 8827–8836 CrossRef PubMed.
- A. E. Nel, L. Mädler, D. Velegol, T. Xia, E. M. V. Hoek, P. Somasundaran, F. Klaessig, V. Castranova and M. Thompson, Nat. Mater., 2009, 8, 543–557 CrossRef CAS PubMed.
- W. Wang, Z. Huang, Y. Li, W. Wang, J. Shi, F. Fu, Y. Huang, X. Pan and C. Wu, Acta Pharm. Sin. B., 2021, 11, 1030–1046 Search PubMed.
- M. B. Flanagan, M. Lundqvist, J. Stigler, T. Cedervall and T. Bergga, Int. J. Biochem. Cell Biol., 2011, 75, 7503–7509 Search PubMed.
- S. Milani, F. B. Bombelli, A. S. Pitek and K. A. Dawson, ACS Nano, 2012, 6, 2532–2541 CrossRef CAS PubMed.
- D. Baimanov, J. Wang, J. Zhang, K. Liu, Y. Cong, X. Shi, X. Zhang, Y. Li, X. Li, R. Qiao, Y. Zhao, Y. Zhou, L. Wang and C. Chen, Nat. Commun., 2022, 13, 5389, DOI:10.1038/s41467-022-33044-y.
- S. J. Park, Int. J. Nanomed., 2020, 15, 5783–5802 CrossRef CAS PubMed.
- K. Partikel, R. Korte, N. C. Stein, D. Mulac, F. C. Herrmann, H. Humpf and K. Langer, Beilstein J. Nanotechnol., 2019, 141, 70–80 Search PubMed.
- P. Del Pino, B. Pelaz, Q. Zhang, P. Maffre, G. U. Nienhaus and W. J. Parak, Mater. Horiz., 2014, 1, 301–313 Search PubMed.
- L. Treuel, S. Brandholt, P. Maffre, S. Wiegele, L. Shang, U. Nienhaus, A. Physics, F. Nanostructures, P. Chemistry, F. Ict-imm and U. States, ACS Nano, 2014, 8, 503–513 Search PubMed.
- J. C. Y. Kah, J. Chen, A. Zubieta and K. Hamad-Schifferli, ACS Nano, 2012, 6730–6740 Search PubMed.
- B. Zewde, O. Atoyebi, A. Gugssa, K. J. Gaskell and D. Raghavan, ACS Omega, 2024, 17, 11614–11627 Search PubMed.
- P. Corona, G. Nanorods and T. Release, Exploiting the Protein Corona around Gold Nanorods for Loading and Triggered Release, ACS Nano, 2012, 6730–6740 Search PubMed.
- A. Balmori, R. Sandu, D. Gheorghe, A. Botea-Petcu, A. Precupas, S. Tanasescu, D. Sánchez-García and S. Borrós, Front. Bioeng. Biotechnol., 2021, 9, 1–9 Search PubMed.
- M. De, C. C. You, S. Srivastava and V. M. Rotello, J. Am. Chem. Soc., 2007, 129, 10747–10753 Search PubMed.
- A. Maity, D. Bagchi, S. K. De and A. Chakraborty, Langmuir, 2023, 39, 4881–4894 CrossRef CAS PubMed.
- N. Dasgupta, S. Ranjan, D. Patra, P. Srivastava, A. Kumar and C. Ramalingam, Chem. – Biol. Interact., 2016, 253, 100–111 Search PubMed.
- A. Udnoor, M. Lokolkar, B. C. Yallur, R. Kale, M. N. Kalasad, U. Katrahalli and D. H. Manjunatha, J. Biomol. Struct. Dyn., 2023, 1, 352–365 Search PubMed.
- A. Bapli, A. Chatterjee, R. K. Gautam, S. Pandit, R. Jana and D. Seth, J. Mol. Liq., 2021, 323, 114618 Search PubMed.
- A. Maity, D. Bagchi, H. Tabassum, P. Nath, S. Sinha and A. Chakraborty, J. Phys. Chem., 2024, 43, 10625–10635 Search PubMed.
- D. M. Kirschenbaum, Anal. Biochem., 1973, 56, 237–263 Search PubMed.
- N. S. El-salamouni, R. M. Farid, A. H. El-kamel and S. S. El-gamal, Int. J. Pharm., 2015, 496, 976–983 CrossRef CAS PubMed.
- R. Kanwar, M. Gradzielski and S. K. Mehta, J. Phys. Chem. B, 2018, 122, 6837–6845 CrossRef CAS PubMed.
- D. Chicea, Optoelectron. Adv. Mater., Rapid Commun., 2010, 4, 1310–1315 CAS.
- Y. Kuo, J. Phys. Chem. B, 2005, 23, 11727–11734 CrossRef PubMed.
- A. Sze, D. Erickson, L. Ren and D. Li, J. Chem. Inf. Model., 2003, 2, 402–410 Search PubMed.
- K. Kar, B. Alex and N. Kishore, J. Chem. Thermodyn., 2002, 34, 319–336 CrossRef CAS.
- C. A. M. Seidel, A. Schulz and M. H. M. Sauer, J. Phys. Chem., 1996, 13, 5541–5553 CrossRef.
- A. Jain, M. Lopus and N. Kishore, Langmuir, 2025, 41, 473–495 CrossRef CAS PubMed.
- N. K. Garg, R. K. Tyagi, B. Singh, G. Sharma, P. Nirbhavane, V. Kushwah, S. Jain and O. P. Katare, Int. J. Pharm., 2016, 499, 301–320 Search PubMed.
- T. Das, M. P. Venkatesh, T. M. P. Kumar and M. Koland, J. Drug Delivery Sci. Technol., 2010, 55, 101415 CrossRef.
- F. A. Burki, K. U. Shah, G. Razaque, S. U. Shah, A. Nawaz, M. D. Saeed, M. U. Rehman, H. Bibi, M. Alfatama and T. M. Elsayed, ACS Omega, 2023, 8, 19302–19310 Search PubMed.
- I. Alberg, S. Kramer, M. Schinnerer, Q. Hu, C. Seidl, C. Leps, N. Drude, D. Möckel, C. Rijcken, T. Lammers, M. Diken, M. Maskos, S. Morsbach, K. Landfester, S. Tenzer, M. Barz and R. Zentel, Small, 2020, 16, 1907574 CrossRef CAS PubMed.
- M. Bin, S. Hossain, F. Chowdhury and S. Ahmed, Arabian J. Chem., 2022, 15, 104117 CrossRef.
- C. Liu, S. Zhang, D. J. McClements, D. Wang and Y. Xu, J. Agric. Food Chem., 2019, 18, 5113–5121 CrossRef PubMed.
- J. Rathee, A. Kaur, R. Kanwar, D. Kaushik, R. Kumar, D. B. Salunke and S. K. Mehta, Colloids Surf., A, 2022, 632, 127764 Search PubMed.
- C. Hao, G. Xu, Y. Feng, L. Lu, W. Sun and R. Sun, Spectrochim. Acta, Part A, 2017, 184, 191–197 Search PubMed.
- D. Rout, S. Sharma, P. Agarwala, A. K. Upadhyaya, A. Sharma and D. K. Sasmal, ACS Omega, 2023, 8, 3114–3128 CrossRef CAS PubMed.
- J. Lazniewska, M. Agostino, S. M. Hickey, E. Parkinson-Lawrence, S. Stagni, M. Massi, D. A. Brooks and S. E. Plush, Chem. – Eur. J., 2021, 11406–11417 Search PubMed.
- D. M. Charbonneau and H. A. Tajmir-Riahi, J. Phys. Chem. B, 2010, 114, 1148–1155 CrossRef CAS PubMed.
- L. Y. Cheng, M. Fang, A. M. Bai, Y. Ouyang and Y. J. Hu, Luminescence, 2017, 32, 873–879 CrossRef CAS PubMed.
- S. Gandhi and I. Roy, J. Mol. Liq., 2019, 296, 111871 Search PubMed.
- T. Cedervall, I. Lynch, S. Lindman, T. Berggård, E. Thulin, H. Nilsson, K. A. Dawson and S. Linse, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 2050–2055 Search PubMed.
- E. Casals, T. Pfaller, A. Duschl, G. J. Oostingh and V. Puntes, ACS Nano, 2010, 4, 3623–3632 Search PubMed.
- A. Akhuli, D. Chakraborty, A. K. Agrawal and M. Sarkar, Langmuir, 2021, 37, 1823–1837 Search PubMed.
- B. Bhattacharya, S. Nakka, L. Guruprasad and A. Samanta, J. Phys. Chem. B, 2009, 113, 2143–2150 Search PubMed.
- A. Bapli, A. Chatterjee, R. K. Gautam, R. Jana and D. Seth, Langmuir, 2021, 16, 5034–5048 CrossRef PubMed.
- P. Meena and N. Kishore, J. Mol. Liq., 2021, 339, 116718 CrossRef CAS.
- S. Karmakar, T. K. Das, S. Kundu, S. Maiti and A. Saha, ACS Appl. Bio Mater., 2020, 3, 8820–8829 CrossRef CAS PubMed.
- A. Jain, E. Judy and N. Kishore, J. Phys. Chem. B, 2024, 128, 5344–5362 CrossRef CAS PubMed.
- S. M. Kelly and N. C. Price, Curr. Protein Pept. Sci., 2000, 4, 349–384 CrossRef PubMed.
- Y. Ni, S. Su and S. Kokot, Spectrochim. Acta, Part A, 2010, 75, 547–552 Search PubMed.
- A. Okada, T. Miura and H. Takeuchi, Biochemistry, 2001, 40, 6053–6060 CrossRef CAS PubMed.
- A. Michnik, Anal. Calorim., 2003, 71, 509–519 Search PubMed.
- B. Goderis, J. A. Putseys, J. Gommes, G. M. Bosmans and J. A. Delcour, Cryst. Growth Des., 2014, 14, 3221–3233 Search PubMed.
- S. Gumpen and P. E. R. O. Hegg, Biochim. Biophys. Acta, 1979, 574, 189–196 CrossRef CAS PubMed.
- M. Di Foggia, S. Bonora and V. Tugnoli, J. Therm. Anal. Calorim., 2013, 111, 1871–1880 Search PubMed.
- P. Prasanthan and N. Kishore, RSC Adv., 2021, 11, 22057–22069 Search PubMed.
- A. Maheshwari and N. Kishore, J. Mol. Liq., 2022, 368, 120599 Search PubMed.
- F. A. Husband, M. J. Garrood, A. R. Mackie, G. R. Burnett and P. J. Wilde, J. Agric. Food Chem., 2001, 49, 859–866 Search PubMed.
- W. Gospodarczyk, K. Szutkowski and M. Kozak, J. Phys. Chem. B, 2014, 29, 8652–8661 CrossRef PubMed.
- Y. L. Jeyachandran, E. Mielczarski, B. Rai and J. A. Mielczarski, Langmuir, 2009, 19, 11614–11620 Search PubMed.
- E. Judy and N. Kishore, RSC Adv., 2021, 11, 35110–35126 Search PubMed.
- P. J. Huang, F. Wang and J. Liu, Langmuir, 2016, 32, 2458–2463 CrossRef CAS PubMed.
- M. J. Swamy, R. Sankhala and B. P. Singh, PloS one, 2011, 6(10), e25993, DOI:10.1007/978-1-4939-9512-7.
- A. A. Thoppil, B. K. Chennuri and R. L. Gardas, J. Mol. Liq., 2019, 290, 111229 Search PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d4cp04737k |
| This journal is © the Owner Societies 2025 |