Open Access Article
Open Access ArticleA DNA small molecular probe with increasing K+ concentration promoted selectivity†
Ya-Ping Gongab,
Jian Yangc,
Ji-Wang Fangac,
Qian Lib,
Zhi-Yong Yu*a,
Aijiao Guan *b and
Han-Yuan Gong
*b and
Han-Yuan Gong *c
*c
aDepartment of Chemistry, Renmin University of China, Beijing, 100872, P. R. China
bInstitute of Chemistry, Chinese Academy of Sciences, Zhongguancunbeiyijie 2, Beijing 100190, P. R. China
cCollege of Chemistry, Beijing Normal University, Xinjiekouwaidajie 19, Beijing 100875, P. R. China. E-mail: hanyuangong@bnu.edu.cn
First published on 22nd April 2021
Abstract
DNA small molecular probe study was considered as a promising approach to achieve DNA related disease diagnosis. Most related reports were performed under specific salinity. Herein, 4-imino-3-(pyridin-2-yl)-4H-quinolizine-1-carbonitrile (IPQC) was generated via a facile procedure with high yield (85%). It is found that IPQC could act as a universal probe for most tested ssDNA, dsDNA and G4 DNA in low [K+] concentration (less than 20 mM). However, IPQC showed highly selective G4 DNA binding via UV-vis and fluorescence response in increasing [K+] (e.g., 150 mM) conditions. The ion atmosphere effects are instructive for DNA probe exploration. This provides guidance for the design, selection and optimization of the probes for target DNA sensing.
Introduction
Nucleic acid is the key genetic material in biological systems.1–3 Its structural integrity determines physiological functions, cell proliferation, and/or differentiation.4,5 DNA small molecular probes were developed to detect specific DNA species,6–9 also their different folding types (e.g. G-quadruplex (G4), considered as a tumor marker species).10–19 Up to now, probe study was considered as a promising approach to realize DNA related disease diagnosis.20–26 However, most reported interactions between DNA and small molecular probes were explored under an environment with specific salinity.27–32 Ion atmosphere effects were considered as a critical factor in DNA recognition. However, only limited examples involving the important effects were reported. These reports included the sodium concentration effect on the complexation between DNA and protein,33 as well as the binding between ligand and double-stranded DNA (dsDNA).34 Understanding and using ion atmosphere effects to promote specific DNA (such as G4) recognition is still under development.Herein, we reported an unusual DNA small molecular probe, 4-imino-3-(pyridin-2-yl)-4H-quinolizine-1-carbonitrile (IPQC) with high K+ concentration dependent selectivity. In low [K+] environment (i.e., [K+] ≤ 20 mM), IPQC shows obvious UV-vis and/or fluorescence responses for most additional DNA species. The tested DNAs include ssDNA (e.g., ssVEGF, ssPS1c-a, ssPS1c-b, ssAf17, ssG-tripl and ssT30), dsDNA (including dsPS1c, ds19AT, dsDx12, ds20, ds22, ds26 and 15GC), and G4 DNA (involving VEGF, bcl-2, H22, P21, c-myc and c-kit). Surprisingly, increasing [K+] improved the DNA recognition selectivity of IPQC. 150 mM [K+] (an intracellular K+ concentration) induces IPQC response only for some additional G4 (Scheme 1). Such findings suggested that ion atmosphere effects (e.g., [K+]) in DNA detection should not be ignored in either exploring new or re-examining previous probes. The ion atmosphere effect study in the DNA detection will provide guidance for the design, selection and optimization of target DNA post-probes.
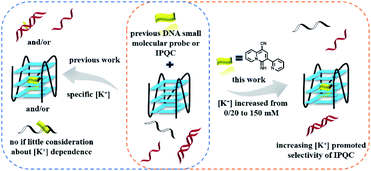 | ||
| Scheme 1 The binding selectivity of previous DNA probe (left, in specific salinity (i.e., [K+])) and IPQC probe (right) with increasing salinity (e.g., [K+]) promoted selectivity for G4 DNA sensing. | ||
Experimental
Reagents and chemicals
All oligonucleotides were synthesized by Invitrogen (Beijing, China) and purified by ultra-polyacrylamide gel electrophoresis (ULTRAPAGE) (purity 95%). Chemicals were purchased from Beijing Chem. Co. (China) and used without further purification. Ultrapure water was prepared by Milli-Q Gradient ultrapure water system (Millipore) and used throughout the experiments.Sample preparation
The stock solutions of the oligonucleotides were prepared by dissolving oligonucleotides into the mixture containing ethanol and Tris–HCl buffer ([Tris] = 20 mM; the ratio between ethanol and aqueous phase as 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 99, v/v, pH = 7.2, containing 0, 20 or 150 mM KCl, respectively) and annealed in a thermocycler (first heating at 90 °C for 5 min, and slowly cooled down to room temperature, then stored at 4 °C before use). The concentration of oligonucleotides was determined based on their absorbance at 260 nm, dsDNA was prepared by heat denaturing and annealing the mixture of ssDNA1 and ssDNA2 (1
99, v/v, pH = 7.2, containing 0, 20 or 150 mM KCl, respectively) and annealed in a thermocycler (first heating at 90 °C for 5 min, and slowly cooled down to room temperature, then stored at 4 °C before use). The concentration of oligonucleotides was determined based on their absorbance at 260 nm, dsDNA was prepared by heat denaturing and annealing the mixture of ssDNA1 and ssDNA2 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). Stock solutions of IPQC were prepared in ethanol (2.00 × 10−3 M).
1). Stock solutions of IPQC were prepared in ethanol (2.00 × 10−3 M).
Synthesis of IPQC
2-Pyridylacetonitrile (1, 0.24 g, 2 mmol), sodium bis(trimethylsilyl)-amide (NaHMDS) (0.55 g, 3 mmol) and anhydrous DMF (5 mL) were added into an oven-dried 25 mL Schleck tube and then heated at 100 °C under air for 3 h. Cooling the mixture down to room temperature and then filtered. Collecting the liquid phase and removing solvent under reduced pressure, the residue was purified by column chromatography on silica gel (200–300 mesh, eluent as the mixture of acetone and petroleum ether (1/3, v/v) to afford the corresponding product as red-brown solid (0.42 g, 85% yield). Its structure was fully characterized via 1H-NMR, 13C-NMR, high resolution mass spectrometry (HRMS) and X-ray single crystal diffraction study.Complexation study between IPQC and DNAs
The absorption spectra were recorded on an Agilent 8453 UV-visible spectrophotometer at the wavelength range 190–1100 nm using a 1 cm quartz cuvette. UV-vis absorbance titrations were carried out by preparing a series of mixture containing IPQC and DNAs. Concentration of IPQC kept as 2.00 × 10−5 M. Additional DNAs concentrations were changed from 0 to 4.0 molar equiv. of IPQC. The solution contained KCl as 0, 20 or 150 mM at room temperature in a special mixture solvent used in all the tests shown below (i.e., ethanol–Tris–HCl buffer; which contained ethanol and Tris–HCl buffer ([Tris] = 20 mM; the ratio between ethanol and aqueous phase as 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 99, v/v, pH = 7.2)). The solvent mixture, each absorption spectrum was carried out after the sample incubation for 2 h.
99, v/v, pH = 7.2)). The solvent mixture, each absorption spectrum was carried out after the sample incubation for 2 h.
Fluorescence spectra were recorded on a HITACHI F-4600 fluorescence spectrophotometer (Hitachi Limited, Japan). Fluorescence titration experiments were carried out with the same samples used in the UV-vis titration study. Fluorescence spectroscopic Job-plot was used to determine the binding stoichiometry. The total concentration of IPQC and DNAs were maintained as 2.00 × 10−5 M, with the molar ratio between IPQC and DNAs as 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0, 5
0, 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 4
1, 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 3
1, 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 2
1, 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1.5
1, 1.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.5, 1
1.5, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, 1
2, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, 1
3, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 and 1
4 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, respectively.
5, respectively.
CD (circular dichroism) spectra were collected on a Jasco 815 spectropolarimeter in a 2 mm path-length quartz cell. CD spectra of DNAs (2.00 × 10−5 M) were measured in the absence or presence of 1.0 molar equiv. of IPQC. 1H-NMR and 13C-NMR spectra were recorded on Bruker AVANCE 400. The single crystals used to obtain the X-ray diffraction structure grew clear light red cube. The .cif documents are available as separate ESI† files, which provide details regarding the specific crystal used for the analysis, along with the structure in question. Diffraction grade crystals were obtained via slow evaporation from solution using a mixture of dichloromethane/n-hexane.
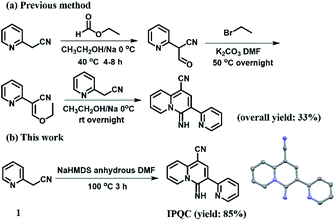
Results and discussion
Although the synthetic method of IPQC was demonstrated in previous studies,35 its properties are still under exploration. The previous synthetic route contained three steps with overall yield of 33%. Low yield, cumbersome steps and the use of hazardous reagents restrict its further exploration. Herein, the improved ‘one-pot’ reaction providing ‘one pot’ facile synthetic steps and high yield (85%). The efficient synthetic route will facilitate the subsequent property study (Fig. 1).The UV-vis and fluorescence spectroscopic properties of IPQC were characterized. The related experiments were carried out in ethanol–Tris–HCl buffer. It is noted that 1% ethanol was added due to the limited solubility of IPQC in pure water phase. Finally, IPQC reveals a peak at λ = 392 nm with ε392 nm = 2.57 × 104 mol−1 L cm−1 in UV-vis absorption. Fluorescence spectrum recorded at λex = 392 nm showed a strong green emission peak of 528 nm, with fluorescence quantum yield Φ = 0.35 and lifetime as 4.84 ns.
Cationic H+·IPQC was believed as the form of IPQC in aqueous phase under pH around 7 (note: the pKb value of IPQC is deduced as around 9.15 from the value of analogic quinoline). The pH dependent fluorescence spectra of 2.00 × 10−5 M IPQC show obvious change with pH value scale as 2 to 12 (cf. ESI Fig. S68 and S69†). It suggests that H+·IPQC is the main form with pH around 7. The planar skeleton and protonation induce the expectation to use IPQC as a potential biological probe of anionic biological macromolecules (e.g., nucleic acids), as the well-known G-quartet ligand, berberine.36–38
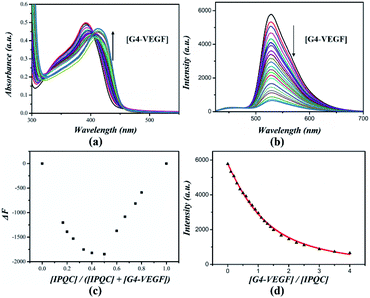
As initial work, we investigated the interaction between IPQC and a series of DNA structures (cf., Table S1†). G-quadruplex VEGF (Vascular Endothelial Growth Factor, a cancer treatment target)39–41 was selected for the first test. UV-vis absorption peak of IPQC at λ = 392 nm red-shifts 19 nm with adding 4.0 molar equiv. of G4-VEGF. Meanwhile, obvious intensity decreasing was shown in fluorescence emission spectrum (carried out in ethanol–Tris–HCl buffer with [KCl] as 20 mM). The decreasing fluorescence intensity attributed mainly to the H+·IPQC binding on the surface of G4-VEGF via π–π stacking interaction via molecular simulation (cf. Fig. 4a–c). Other synergistic effects, such as electrostatic attraction between negative DNA and positive H+·IPQC, suitable shape and volume of H+·IPQC to the groove, also may lead to the fluorescence quenching of H+·IPQC.
Further CD (circular dichroism) experiments showed DNA signals are almost the same before and after adding IPQC, while IPQC has no Cotton effect in the presence or absence of DNA species (cf. ESI Fig. S73†). This phenomenon indicates DNA configuration is kept during the binding process. The stoichiometric ratio between IPQC and DNA were speculated by Job-plot analysis, despite the method is limited in some complicated cases.42–44 Fluorescence Job-plot analysis resulted in a minimum value at [IPQC]/([IPQC] + [G4-VEGF]) ratio as 0.5. The finding suggested a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry (Fig. 2). Further UV-vis titration experiment suggested the association constant Ka1 = (1.3 ± 0.1) × 105 M−1 for the 1
1 binding stoichiometry (Fig. 2). Further UV-vis titration experiment suggested the association constant Ka1 = (1.3 ± 0.1) × 105 M−1 for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 complexation. The result is consistent with calculated Ka1 = (7.9 ± 0.2) × 104 M−1 from fluorescence titration.
1 complexation. The result is consistent with calculated Ka1 = (7.9 ± 0.2) × 104 M−1 from fluorescence titration.
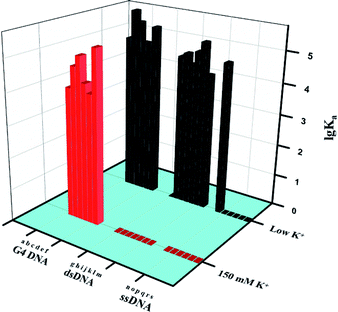
Same methods were also used to study the interactions between IPQC and other tested DNAs. The association constants were summarized in Table 1 and Fig. 3, which exhibit both obvious UV-vis and fluorescence signal response in most cases involving ssDNA, dsDNA, and G-quadruples.
| Substrate (S) | [IPQC]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [S] [S] |
Ka measurement in [K+] as 0 or 20 mM | Ka measurement in [K+] as 150 mM | ||
|---|---|---|---|---|---|
| Method | Method | ||||
| UV-vis | Fluorescence | UV-vis | Fluorescence | ||
a (1) Equations governing the relevant equilibria:  here H indicated IPQC and G for various DNA substrates. (2) ‘—’ indicates titration experiments are unsuccessful because of the weak response of IPQC to additional DNAs. (3) All the tests were carried out in buffer ([Tris] = 20 mM; the ratio between ethanol and aqueous phase as 1 here H indicated IPQC and G for various DNA substrates. (2) ‘—’ indicates titration experiments are unsuccessful because of the weak response of IPQC to additional DNAs. (3) All the tests were carried out in buffer ([Tris] = 20 mM; the ratio between ethanol and aqueous phase as 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 99, v/v, pH = 7.2). 20 or 150 mM KCl was used in G4 DNA tests, 0 or 150 mM KCl was used in the cases of ss/ds DNAs. 99, v/v, pH = 7.2). 20 or 150 mM KCl was used in G4 DNA tests, 0 or 150 mM KCl was used in the cases of ss/ds DNAs. |
|||||
| G4-bcl-2 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(5.0 ± 0.2) × 105 | (2.5 ± 0.1) × 105 | (1.6 ± 0.1) × 105 | (2.5 ± 0.1) × 105 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(7.9 ± 0.4) × 104 | (1.3 ± 0.1) × 105 | (5.0 ± 0.3) × 104 | (2.5 ± 0.1) × 104 | |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(3.2 ± 0.3) × 104 | (2.0 ± 0.2) × 103 | (1.3 ± 0.1) × 104 | (4.0 ± 0.3) × 103 | |
| G4-H22 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(7.9 ± 0.2) × 104 | (1.0 ± 0.1) × 105 | (2.0 ± 0.1) × 104 | (1.3 ± 0.1) × 104 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(6.3 ± 0.3) × 104 | (2.5 ± 0.1) × 104 | — | — | |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(2.0 ± 0.2) × 105 | (7.9 ± 0.6) × 104 | — | — | |
| G4-P21 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(7.9 ± 0.2) × 104 | (7.9 ± 0.2) × 104 | (1.3 ± 0.1) × 104 | (1.0 ± 0.1) × 104 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(1.3 ± 0.1) × 105 | (1.3 ± 0.1) × 105 | — | — | |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(2.5 ± 0.2) × 104 | (2.5 ± 0.2) × 104 | — | — | |
| G4-VEGF | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(1.3 ± 0.1) × 105 | (7.9 ± 0.2) × 104 | (2.5 ± 0.1) × 104 | (2.0 ± 0.1) × 104 |
| G4-c-kit | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(5.0 ± 0.2) × 105 | (3.2 ± 0.1) × 105 | (2.0 ± 0.1) × 105 | (1.3 ± 0.1) × 105 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(1.0 ± 0.1) × 104 | (7.9 ± 0.4) × 103 | (1.6 ± 0.1) × 104 | (1.6 ± 0.1) × 104 | |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(2.5 ± 0.2) × 103 | (5.0 ± 0.4) × 103 | (4.0 ± 0.3) × 104 | (4.0 ± 0.3) × 103 | |
| G4-c-myc | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(6.3 ± 0.2) × 105 | (7.9 ± 0.2) × 105 | (1.6 ± 0.1) × 105 | (5.0 ± 0.2) × 104 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(1.0 ± 0.1) × 105 | (3.2 ± 0.2) × 104 | (6.3 ± 0.3) × 104 | (7.9 ± 0.4) × 104 | |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(5.0 ± 0.4) × 104 | (2.0 ± 0.2) × 105 | (1.0 ± 0.1) × 105 | (5.0 ± 0.4) × 104 | |
| ds20 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(1.6 ± 0.1) × 104 | (2.0 ± 0.1) × 104 | — | — |
| ds22 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(2.0 ± 0.1) × 105 | (2.0 ± 0.1) × 105 | — | — |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(1.6 ± 0.1) × 104 | (1.3 ± 0.1) × 104 | |||
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(1.3 ± 0.1) × 106 | (3.2 ± 0.3) × 105 | |||
| ds26 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(5.0 ± 0.2) × 104 | (8.0 ± 0.2) × 104 | — | — |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
(1.0 ± 0.1) × 104 | (6.3 ± 0.3) × 103 | |||
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 3 |
(7.9 ± 0.6) × 104 | (1.6 ± 0.1) × 104 | |||
| ds19AT | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(3.2 ± 0.1) × 104 | (3.2 ± 0.1) × 104 | — | — |
| dsDx12 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(7.9 ± 0.2) × 104 | (6.3 ± 0.2) × 104 | — | — |
| hairpin15GC | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
— | (5.0 ± 0.2) × 104 | — | — |
| ssVEGF | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
(5.0 ± 0.2) × 104 | (6.3 ± 0.2) × 104 | — | — |
| ssPS1c-a | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
— | — | — | — |
| ssAf17 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
— | — | — | — |
| ssG-tripl | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
— | — | — | — |
| dsPS1c | — | — | — | — | — |
| ssPS1c-b | — | — | — | — | — |
| ssT30 | — | — | — | — | — |
For further in vivo and/or in situ detection, we studied the DNA sensing behaviour of IPQC in simulated physiological environment. Specifically, the interaction between IPQC and former tested DNAs were explored in ethanol–Tris–HCl buffer with 150 mM K+ (an intracellular K+ concentration). Surprisingly, IPQC exhibits both UV-vis and fluorescence response only for G-quadruplex in this condition, while the responses for all the tested ss/ds DNA disappeared. Specifically, the association constant value Ka1 = (2.0 ± 0.1) × 104 M−1 between IPQC and G4-VEGF indicates that excess potassium do not strongly weaken combination between IPQC and G4-VEGF. As Fig. 3 shows, although the affinity between other G-quadruplex and probe IPQC may be slightly weakened with increasing [K+], they all show similar spectral response. The Ka1 value of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding calculated by UV-vis and fluorescence titration has no significant change compared with which in low [K+] system.
1 binding calculated by UV-vis and fluorescence titration has no significant change compared with which in low [K+] system.
20 mM magnesium ions and 10 mM sodium ions were added to ethanol–Tris–HCl buffer with 150 mM K+ to further mimic the intracellular environment (cf. ESI Fig. S74–S79†). The final spectral responses (especially in fluorescence spectrum) are observed as the same as before adding. It implied that the high selectivity of IPQC for G4 remains after adding 20 mM MgCl2 and 10 mM NaCl. The result proves high [K+] (150 mM) promoted high selectivity of IPQC for G-quadruplex binding.
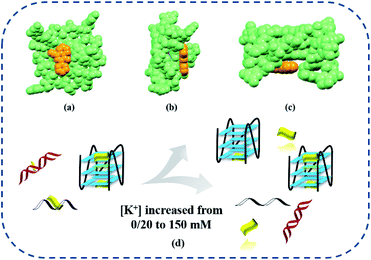
To further explore the increasing selectivity of IPQC, K+ concentrate dependent effect between IPQC and dsDNA was studied in detail via UV-vis and fluorescence methods. Additional dsDNA induced spectral response of IPQC was carried out. The spectroscopic changes decline with adding [K+], until completely disappear with [K+] as 500 mM (cf. Fig. S82–S84†). This experiment interpreted that intermolecular interactions between dsDNA and probe IPQC are acutely sensitive to ion atmosphere effects. To deeper insight of the phenomena, molecular modelling was carried out. It revealed that IPQC can stack on the surface of the terminal quartet of G4 DNA (Fig. 4) or insert into the groove of ss/dsDNA structure in low [K+]. It is suggested that excess cations can compete with protonated IPQC to bind DNA (cf. ESI Fig. S70 and S71†). Furthermore, K+ bind with polyanionic DNA helix to reduce electrostatic repulsion of the negative charges on DNA phosphate backbone. It is unfavourable for IPQC to intercalate to the groove binding sites in ss/dsDNAs. As the result, the binding between IPQC and ss/dsDNA weakens or even disappears with increasing K+. Differently, in the cases of G4 DNAs, the presence of metal cation (e.g., Li+, Na+ or K+) benefit to form a metal-ion-stabilized G4 structures (although Li+ provide inefficient induction for G4 structure formation45–47) so that the electron-rich quinoline plane of IPQC can effectively sit on the planar surface of the G-quadruplex via π–π stacking (cf. ESI Fig. S80 and S81†). Finally, the electrostatic completion of K+ and the enhancement of ion atmosphere effect shown above led a net result as little influence on the binding process between G4 and IPQC sitting on the G-quadruplex. However, the other G4 binding mode(s) of IPQC will also be weaken as the cases of ss/ds DNAs. Actually, the binding stoichiometries between IPQC and some G4 DNAs (e.g., G4-H22 or G4-P21) change from 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 to 1
3 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 corresponding to [K+] as 20 or 150 mM, respectively (cf. ESI Fig. S67†).
1 corresponding to [K+] as 20 or 150 mM, respectively (cf. ESI Fig. S67†).
Conclusions
In conclusion, molecule 4-imino-3-(pyridin-2-yl)-4H-quinolizine-1-carbonitrile (IPQC) was efficiently synthesized via a simple one-pot method. It exhibits high selectivity to G-quadruplex compared to ss/dsDNA in sufficient [K+], it even acts as a universal DNA probe for ss/ds and G4 DNA under low [K+] (0 or 20 mM) environment. Ion atmosphere effects induced selectivity urges us to re-examine the previous DNA probes and provide more perspective for the design of new ones.Conflicts of interest
There are no conflicts to declare.Acknowledgements
This research was supported under the National Natural Science Foundation of China (Grant numbers 21303225, 21675162, 21977096, 21778058, 21672025, 21472014, 21971022), the Beijing Natural Science Foundation (Grant Number 7182189 and 7172248).Notes and references
- I. Ahmad, A. Ahmad and M. Ahmad, Phys. Chem. Chem. Phys., 2016, 18, 6476–6485 RSC.
- A. Mukherjee, S. Mondal and B. Singh, Int. J. Biol. Macromol., 2017, 101, 527–535 CrossRef CAS PubMed.
- K. Gracie, V. Dhamodharan, P. I. Pradeepkumar, K. Faulds and D. Graham, Analyst, 2014, 139, 4458–4465 RSC.
- A. Subastri, C. H. Ramamurthy, A. Suyavaran, R. Mareeswaran, P. Lokeswara Rao, M. Harikrishna, M. Suresh Kumar, V. Sujatha and C. Thirunavukkarasu, Int. J. Biol. Macromol., 2015, 78, 122–129 CrossRef CAS PubMed.
- F. Wu, J. Jin, L. Wang, P. Sun, H. Yuan, Z. Yang, G. Chen, Q.-H. Fan and D. Liu, ACS Appl. Mater. Interfaces, 2015, 7, 7351–7356 CrossRef CAS PubMed.
- M. Jin, X. Liu, X. Zhang, L. Wang, T. Bing, N. Zhang, Y. Zhang and D. Shangguan, ACS Appl. Mater. Interfaces, 2018, 10, 25166–25173 CrossRef CAS PubMed.
- S. N. Journey, S. L. Alden, W. M. Hewitt, M. L. Peach, M. C. Nicklaus and J. S. Schneekloth Jr, Med. Chem. Comm., 2018, 9, 2000–2007 RSC.
- W. Gai, Q. Yang, J. Xiang, W. Jiang, Q. Li, H. Sun, A. Guan, Q. Shang, H. Zhang and Y. Tang, Nucleic Acids Res., 2013, 41, 2709–2722 CrossRef CAS PubMed.
- L. Yu, Q. Yang, J. Xiang, H. Sun, L. Wang, Q. Li, A. Guan and Y. Tang, Analyst, 2015, 140, 1637–1646 RSC.
- M.-H. Hu, S.-B. Chen, R.-J. Guo, T.-M. Ou, Z.-S. Huang and J.-H. Tan, Analyst, 2015, 140, 4616–4625 RSC.
- A. C. Bhasikuttan and J. Mohanty, Chem. Commun., 2015, 51, 7581–7597 RSC.
- A. C. Bhasikuttan, J. Mohanty and H. Pal, Angew. Chem., Int. Ed., 2007, 46, 9305–9307 CrossRef CAS PubMed.
- H. Chen, H. Sun, S. Zhang, W. Yan, Q. Li, A. Guan, J. Xiang, M. Liu and Y. Tang, Chem. Commun., 2019, 55, 5060–5063 RSC.
- M. Liu, Y. Liu, F. Wu, Y. Du and X. Zhou, Bioorg. Med. Chem., 2019, 27, 1962–1965 CrossRef CAS PubMed.
- T. M. Ou, Y. J. Lu, J. H. Tan, Z. S. Huang, K. Y. Wong and L. Q. Gu, ChemMedChem, 2008, 3, 690–713 CrossRef CAS PubMed.
- L. L. Tong, L. Li, Z. Chen, Q. Wang and B. Tang, Biosens. Bioelectron., 2013, 49, 420–425 CrossRef CAS PubMed.
- S. Zhang, H. Sun, L. Wang, Y. Liu, H. Chen, Q. Li, A. Guan, M. Liu and Y. Tang, Nucleic Acids Res., 2018, 46, 7522–7532 CrossRef CAS PubMed.
- D. Wei, A. K. Todd, M. Zloh, M. Gunaratnam, G. N. Parkinson and S. Neidle, J. Am. Chem. Soc., 2013, 135, 19319–19329 CrossRef CAS PubMed.
- G. N. Parkinson and G. W. Collie, G-Quadruplex Nucleic Acids, 2019, 131–155 CAS.
- L. Zhou, J. Yang, C. Estavillo, J. D. Stuart, J. B. Schenkman and J. F. Rusling, J. Am. Chem. Soc., 2003, 125, 1431–1436 CrossRef CAS PubMed.
- D. Gibson, Pharmacogenomics J., 2002, 2, 275–276 CrossRef CAS PubMed.
- L. Xie, X. Qian, J. Cui, Y. Xiao, K. Wang, P. Wu and L. Cong, Bioorg. Med. Chem., 2008, 16, 8713–8718 CrossRef CAS PubMed.
- A. J. Pickard, F. Liu, T. F. Bartenstein, L. G. Haines, K. E. Levine, G. L. Kucera and U. Bierbach, Chem. –Eur. J., 2014, 20, 16174–16187 CrossRef CAS PubMed.
- A. Rajendran and B. U. Nair, Biochim. Biophys. Acta, 2006, 1760, 1794–1801 CrossRef CAS PubMed.
- M. Gunaratnam, G. W. Collie, A. P. Reszka, A. K. Todd, G. N. Parkinson and S. Neidle, Bioorg. Med. Chem., 2018, 26, 2958–2964 CrossRef CAS PubMed.
- M. Sayed and H. Pal, Phys. Chem. Chem. Phys., 2015, 17, 9519–9532 RSC.
- C. C. Chang, J. Y. Wu, C. W. Chien, W. S. Wu, H. Liu, C. C. Kang, L. J. Yu and T. C. Chang, Anal. Chem., 2003, 75, 6177–6183 CrossRef CAS PubMed.
- B. Jin, X. Zhang, W. Zheng, X. Liu, J. Zhou, N. Zhang, F. Wang and D. Shangguan, Anal. Chem., 2014, 86, 7063–7070 CrossRef CAS PubMed.
- M. Nikan, M. D. Antonio, K. Abecassis, K. McLuckie and S. Balasubramanian, Angew. Chem., Int. Ed., 2013, 52, 1428–1431 CrossRef CAS PubMed.
- D. Verga, C. H. N'Guyen, M. Dakir, J. L. Coll, M. P. Teulade-Fichou and A. Molla, J. Med. Chem., 2018, 61, 10502–10518 CrossRef CAS PubMed.
- M.-Q. Wang, Y. Zhang, X.-Y. Zeng, H. Yang, C. Yang, R.-Y. Fu and H.-J. Li, Dyes Pigm., 2019, 168, 334–340 CrossRef CAS.
- X.-f. Zhang, J.-f. Xiang, M.-y. Tian, Q.-f. Yang, H.-x. Sun, S. Yang and Y.-l. Tang, J. Phys. Chem. B, 2009, 113, 7662–7667 CrossRef CAS PubMed.
- B. E. Allred, M. Gebala and D. Herschlag, J. Am. Chem. Soc., 2017, 139, 7540–7548 CrossRef CAS PubMed.
- M. T. Record Jr, T. M. Lohman and P. J. De Haseth, J. Mol. Biol., 1976, 107, 145–158 CrossRef CAS.
- Y.-F. Luo, T. Yang and Y.-Q. Wei, CN 107739375A, G01N21/64, 2018.
- F. Papi, C. Bazzicalupi, M. Ferraroni, G. Ciolli, P. Lombardi, A. Y. Khan, G. S. Kumar and P. Gratteri, ACS Med. Chem. Lett., 2020, 11, 645–650 CrossRef CAS PubMed.
- Y. Ma, T. M. Ou, J. H. Tan, J. Q. Hou, S. L. Huang, L. Q. Gu and Z. S. Huang, Eur. J. Med. Chem., 2011, 46, 1906–1913 CrossRef CAS PubMed.
- I. Bessi, C. Bazzicalupi, C. Richter, H. R. Jonker, K. Saxena, C. Sissi, M. Chioccioli, S. Bianco, A. R. Bilia, H. Schwalbe and P. Gratteri, ACS Chem. Biol., 2012, 7, 1109–1119 CrossRef CAS PubMed.
- G. Martiny-Baron and D. Marmé, Pharm. Technol., 1995, 6, 675–680 CAS.
- A. Bikfalvi and R. Bicknell, Trends Pharmacol. Sci., 2002, 23, 576–582 CrossRef CAS PubMed.
- D. Sun, K. Guo, J. J. Rusche and L. H. Hurley, Nucleic Acids Res., 2005, 33, 6070–6080 CrossRef CAS PubMed.
- F. Ulatowski, K. Dabrowa, T. Balakier and J. Jurczak, J. Org. Chem., 2016, 81, 1746–1756 CrossRef CAS PubMed.
- D. Brynn Hibbert and P. Thordarson, Chem. Commun., 2016, 52, 12792–12805 RSC.
- J. S. Renny, L. L. Tomasevich, E. H. Tallmadge and D. B. Collum, Angew. Chem., Int. Ed., 2013, 52, 11998–12013 CrossRef CAS PubMed.
- T. J. Pinnavaia, C. L. Marshall, C. M. Mettler, C. L. Fisk, H. T. Miles and E. D. Becker, J. Am. Chem. Soc., 1978, 100, 3625–3627 CrossRef CAS.
- C. Bardin and J. L. Leroy, Nucleic Acids Res., 2008, 36, 477–488 CrossRef CAS PubMed.
- J. W. Shim, Q. Tan and L.-Q. Gu, Nucleic Acids Res., 2009, 37, 972–982 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: UV-vis, fluorescence, NMR, HRMS and X-ray single crystal diffraction data. CCDC 1977107. For ESI and crystallographic data in CIF or other electronic format see DOI: 10.1039/d0ra06274j |
| This journal is © The Royal Society of Chemistry 2021 |