Mechanism and kinetics of tyrosinase inhibition by glycolic acid: a study using conventional spectroscopy methods and hydrogen/deuterium exchange coupling with mass spectrometry†
Da
Ma
a,
Zong-Cai
Tu
*ab,
Hui
Wang
a,
Lu
Zhang
b,
Na
He
a and
David Julian
McClements
 *c
*c
aState Key Laboratory of Food Science and Technology, Nanchang University, Nanchang, Jiangxi 330047, China. E-mail: tuzc_mail@aliyun.com; Fax: +86-791-8830-5938; Tel: +86-791-88121868
bCollege of Life Science, Jiangxi Normal University, Nanchang, Jiangxi 330022, China
cDepartment of Food Science, University of Massachusetts, Amherst, MA 01060, USA. E-mail: mcclements@foodsci.umass.edu; Fax: +413-545-1262; Tel: +413-545-2275
First published on 2nd December 2016
Abstract
Tyrosinase is an enzyme that promotes enzymatic browning of fruits and vegetables, thereby reducing product quality. A variety of analytical tools were used to characterize the interactions between tyrosinase and a natural tyrosinase inhibitor (glycolic acid). Hydrogen/deuterium exchange coupling with mass spectrometry (HDX-MS) was used to elucidate the interaction mechanism between glycolic acid and tyrosinase. UV-visible, fluorescence and circular dichroism spectroscopy analysis indicated that glycolic acid inhibited tyrosinase activity in a mixed-type manner with an IC50 of 83 ± 14 μM. The results of these techniques suggested that glycolic acid bound to tyrosinase through hydrophobic attraction, and this interaction led to a pronounced conformational change of the enzyme molecules. HDX-MS analysis showed that the activity of tyrosinase was primarily inhibited by a structural perturbation of its active site (His 263). This study provides a comprehensive understanding of the interaction between glycolic acid and tyrosinase, which could lead to new approaches to control tyrosinase activity in foods and other products.
1. Introduction
A common browning reaction observed in many plants, animals, and microorganisms is catalyzed by the copper-containing enzyme tyrosinase, which is also known as catecholase or diphenol oxidase.1 Tyrosinase catalyzes the hydroxylation of monophenols to O-diphenols (monophenolase activity) and the oxidation of O-diphenols to O-quinones (diphenolase activity).2O-Quinone is a highly reactive substance that can rapidly polymerize to form melanin, which is a brown pigment.3 The browning of food products through this mechanism is undesirable because it leads to detrimental changes in color, flavor, and nutritional profile,1 thereby reducing their market value and shelf life.4 Therefore, identifying natural or synthetic inhibitors of tyrosinase to prevent browning of food products is commercially important.5An improved understanding of the interaction between tyrosinases and their inhibitors would facilitate the rational discovery of new potential candidates that could be used as inhibitors. In general, a wide variety of analytical tools and approaches are needed to provide detailed information about the interactions of small molecules with enzymes. In this study, hydrogen/deuterium exchange coupling with high-resolution mass spectrometry (HDX-MS) was used to provide fundamental information about inhibitor–tyrosinase interactions. HDX-MS can provide detailed information about the nature of the binding site on the tyrosinase molecule via analysis of the location of deuterium probe labels. In particular, we focused on the utilization of HDX-MS to characterize the interactions between tyrosinase and glycolic acid. This technique has previously been shown to be particularly powerful method for studying the interactions between proteins and small molecules.6–8
Glycolic acid is found in nature as a trace component in sugarcane, beets, grapes, and other fruits.9 It has the lowest molecular weight of all alpha-hydroxy acids because it only has two carbon atoms: one carbon atom has a carboxyl group attached while the other has a hydroxyl group attached. Studies using human skin models have shown that glycolic acid and its derivatives can inhibit tyrosinase activity and browning in melanoma cells.10,11 These studies suggest that glycolic acid may also be able to inhibit tyrosinase activity in food applications, such as the inhibition of browning in fruits and vegetables. However, there is still a relatively poor understanding of the mechanism of GA inhibition of tyrosinase activity. The purpose of this study was therefore to carry out a detailed analysis of the interaction of glycolic acid with tyrosinase using a number of spectroscopic methods in combination with HDX-MS.
The kinetics of tyrosinase inhibition by glycolic acid was characterized using UV-visible absorption spectroscopy. The nature of the interaction between glycolic acid and tyrosinase was determined using fluorescence quenching. Changes in the conformation of the tyrosinase molecules resulting from their interactions with glycolic acid were determined by synchronous fluorescence and circular dichroism (CD) spectroscopy. Detailed information about the nature of the glycolic acid binding site on the tyrosine molecule was determined by HDX-MS. The results of this research enhance our understanding of the molecular basis of glycolic acid–tyrosinase interactions, as well as providing information that will be useful for identifying new tyrosinase inhibitors that could be used as anti-browning agents in food products.
2. Materials and methods
2.1. Chemicals and material
Tyrosinase (EC 1.14.18.1, 128 kDa, 25 kU) from mushroom, pepsin, citric acid, and formic acid (FA) were obtained from the Sigma Chemical Co. (St Louis, MO). Tyrosinase stock solution (10 mM) was prepared using Phosphate Buffered Saline (PBS, pH 6.8, 0.05 M), and then diluted to the required concentrations just before use. Deuterated water (D2O) and glycolic acid (GA) were purchased from Aladdin Industrial Co. (Shanghai, China). Stock solutions were prepared by dissolving 2% (v/v) DMSO (containing GA) into PBS (pH 7.4) to form a final solution containing 2 mM GA. The maximum concentration of DMSO in the solutions used to carry out the UV, fluorescence, CD and HDX-MS studies was 0.2% (v/v), which is much lower than the level required to have an impact on these measurements.12 All other reagents were analytical grade, and ultrapure water from a pure water purification system (Millipore, Billerica, MA) was used throughout this study.2.2. Inhibitory kinetics
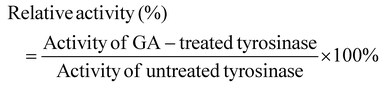 | (1) |
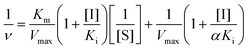 | (2) |
The slope of this equation when 1/v is plotted against 1/[S] is given by
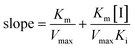 | (3) |
The values of α, Ki, Km and Vmax can be derived from these equations. Here, ν is the enzyme reaction rate in the absence and presence of GA, α is the ratio of the uncompetitive inhibition constant to competitive inhibition constant. The interaction is considered to be a noncompetitive inhibition when α is equal to 1. Ki and Km are the inhibition constant and Michaelis–Menten constant, respectively. [I] and [S] are the concentrations of inhibitor and substrate, respectively.
2.3. Fluorescence titration assay
A fluorescence titration assay was performed using a Hitachi Fluorescence spectrofluorimeter (Model F-7000, Hitachi, Japan) attached to a thermostatic water bath. The widths of the excitation and emission slits were both set at 2.5 nm, and an excitation wavelength of 280 nm was used. The reaction solution of tyrosinase (2.4 μM) was titrated by successive addition of 1.0 mM GA to achieved a concentration range from 0 to 22 μM. Fluorescence emission spectra were measured at wavelengths from 300 to 500 nm at three temperatures (298, 304 and 310 K).The fluorescence quenching data were analyzed by the well-known Stern–Volmer equation:
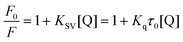 | (4) |
The association constant (ka) was estimated using the modified Stern–Volmer equation:
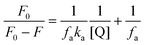 | (5) |
Here, ka and fa are the modified Stern–Volmer association constant and the fraction respectively.
2.4. Thermodynamic analysis
Thermodynamic parameters (enthalpy change ΔH and entropy change ΔS) of binding forces were determined from the van't Hoff equation: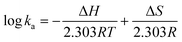 | (6) |
| ΔG = ΔH − TΔS | (7) |
The ka and R are the binding constant and gas constant (8.314 J mol−1 K−1) respectively, and T values were set at 298, 304, and 310 K, respectively.
2.5. Circular dichroism (CD) spectra
The CD spectra (200 to 250 nm) of untreated tyrosinase and GA-treated tyrosinase were performed on a Bio-Logic MOS 450 CD spectrometer (Bio-Logic, Claix, France) at room temperature using a 1.0 mm path length quartz cuvette. The tyrosinase concentration was kept at 2.4 μM, while the [GA]/[tyrosinase] ratios tested were 0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8 and 1
8 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 in 50 mM, pH 6.8 PBS. All observed CD spectra were subtracted from the PBS signal (blank). CD spectra of tyrosinase were analyzed using the online SELCON3 program.
4 in 50 mM, pH 6.8 PBS. All observed CD spectra were subtracted from the PBS signal (blank). CD spectra of tyrosinase were analyzed using the online SELCON3 program.
2.6. Hydrogen–deuterium exchange and mass spectrometry analysis
The hydrogen–deuterium exchange (HDX) and mass spectrometry (MS) analysis of tyrosinase was carried out according to Zhang et al.13 The untreated and GA-treated tyrosinase ([GA]/[tyrosinase] of 0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 1
1 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8) were prepared with a final tyrosinase concentration of 2.4 μM, and diluted 10-fold with D2O in an ice-water bath. After 10 min, the deuterium exchange was quenched with an equal volume of pre-chilled quenching buffer (0.5 M citric acid, 0 °C, pH 2.2). Pepsin (5 μL, 1 mg mL−1, 0.05 mol L−1 HCl) was added immediately to digest the GA–tyrosinase mixture at 25 °C for 10 min. The solution (45 μL) was then injected into a C18 column (50 × 1.00 mm, Phenomenex) on a Shimadzu HPLC with LC-10AD pumps. Solvent A was 5% acetonitrile with 0.1% FA, and solvent B was 95% acetonitrile with 0.1% FA. After desalting for 5 min with 5% solvent B, the samples were eluted with a 5–15% gradient for 1 min and 15–50% gradient for 10 min at 50 μL min−1. The eluent was directly delivered into a Fourier transform ion cyclotron mass spectrometry (FTICR MS) (Thermo Fisher Scientific) for mass-spectrometry analysis. For peptic peptide identification, the mass spectrometer was operated under a data-dependent MS–MS mode. Charge-state screening was enabled, and precursors with unknown charge state were excluded. All HDX experiments were performed in triplicate.
8) were prepared with a final tyrosinase concentration of 2.4 μM, and diluted 10-fold with D2O in an ice-water bath. After 10 min, the deuterium exchange was quenched with an equal volume of pre-chilled quenching buffer (0.5 M citric acid, 0 °C, pH 2.2). Pepsin (5 μL, 1 mg mL−1, 0.05 mol L−1 HCl) was added immediately to digest the GA–tyrosinase mixture at 25 °C for 10 min. The solution (45 μL) was then injected into a C18 column (50 × 1.00 mm, Phenomenex) on a Shimadzu HPLC with LC-10AD pumps. Solvent A was 5% acetonitrile with 0.1% FA, and solvent B was 95% acetonitrile with 0.1% FA. After desalting for 5 min with 5% solvent B, the samples were eluted with a 5–15% gradient for 1 min and 15–50% gradient for 10 min at 50 μL min−1. The eluent was directly delivered into a Fourier transform ion cyclotron mass spectrometry (FTICR MS) (Thermo Fisher Scientific) for mass-spectrometry analysis. For peptic peptide identification, the mass spectrometer was operated under a data-dependent MS–MS mode. Charge-state screening was enabled, and precursors with unknown charge state were excluded. All HDX experiments were performed in triplicate.
2.7. Data analysis
The peptic peptides were identified by Sequest searching against the tyrosinase (from mushroom). The HDX-MS data were analyzed by the EX-MS software.14 Average changes in deuterium incorporation (ΔHDX) ± S.D. were determined from three separate experiments. The changes in deuterium incorporation (ΔHDX) into tyrosinase were defined as the difference between the average values in the absence and presence of GA. The difference in deuteration in each domain was calculated by summing the ΔHDX values for all included peptides and then dividing by the total number of amino-acid residues.2.8. Statistical analysis
The experimental results, including tyrosinase activity assay and kinetic analysis for mix-type inhibition, were expressed as mean values ± S.D. (n = 3) and analyzed using statistical software (version 16.0, SPSS, lnc., Chicago, USA).3. Results
3.1. Effect of GA concentration on tyrosinase activity
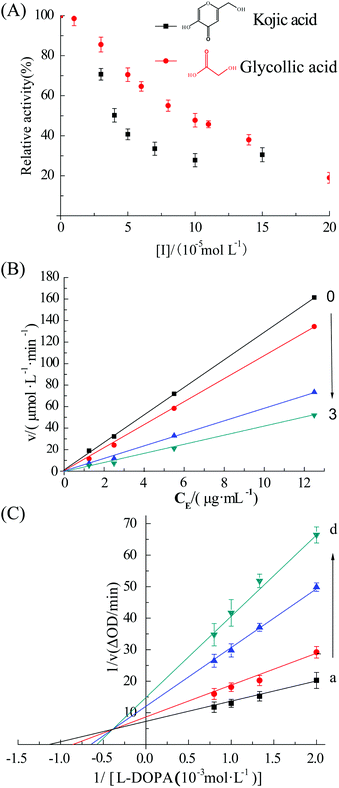
The UV absorption spectra of tyrosinase (2.4 μM) in the presence of different levels of GA are shown in Fig. 1A. The tyrosinase activity was significantly inhibited upon addition of GA (0 to 0.2 mM). The concentrations of GA and kojic acid (positive control) leading to a loss of 50% enzyme activity (IC50) were 83 ± 14 and 39 ± 10 μM, respectively. Previous studies have reported that the IC50 value of kojic acid leading for tyrosinase activity was 22 ± 8 μM,15 which is in reasonable agreement with the value obtained in this study. The relative activity of additional concentrations 10, 50, 100 and 200 μM GA were 98.5%, 70.5%, 47.7% and 19.0%, respectively. These results indicate that GA is a good potential inhibitor of tyrosinase when used at levels of 0.2 mM or more.Plots of remaining activity vs. [tyrosinase] at different GA concentrations are shown in Fig. 1B. These results indicate that all the plots were linear and passed through the origin, and that the slope of the lines decreased with increasing inhibitor concentration, indicating that GA reversibly inhibited tyrosinase.16 This finding is in agreement with earlier studies.17
3.2. Nature of inhibition
The nature of the inhibition of tyrosinase by GA was evaluated using the Lineweaver–Burk double reciprocal plots.18 Changes in both Vmax and Km values are shown in Fig. 1C. The apparent Km increased while the Vmax decreased, indicating that GA was a mixed inhibitor with the substrate L-DOPA during catalysis.19 The values of Ki and Km were determined to be 3.03 ± 0.16 mM and 0.57 mM by eqn (2) and (3), respectively.3.3. Fluorescence quenching assay
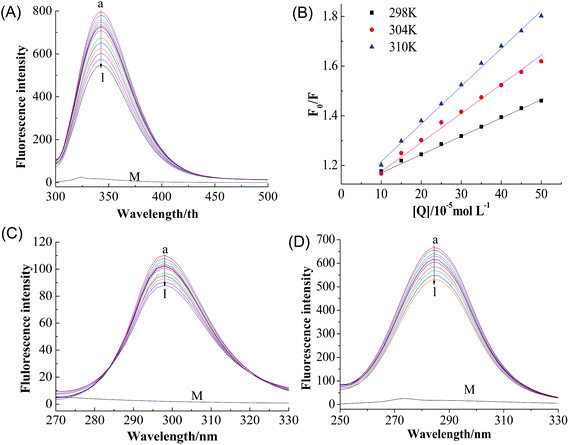
A fluorescence-quenching assay was carried to further investigate the interaction mechanism between GA and tyrosinase. Minimal changes in the local environment of the fluorophore were detected, which is helpful for understanding the nature of the binding mechanism and interactions.20 Fluorescence quenching of proteins is usually attributed to a variety of molecular interactions including excited-state reactions, energy transfer, ground-state complex formation, and collisional quenching.21 The quenching mechanism can be classified as either dynamic or static quenching, which can often be distinguished based on their characteristic temperature dependence.22As shown in Fig. 2A, tyrosinase displayed a strong emission peak at 342 nm when excited at 280 nm, the peak intensity of tyrosinase decreased gradually with increasing GA concentration (0 → 22 μM) without any significant shift of the wavelength of the peak maximum, indicating that GA interacts with tyrosinase and quenches its intrinsic fluorescence.11
The Stern–Volmer plots for the quenching of tyrosinase by GA (Fig. 2B) showed that it exhibited a good linear relationship from 100 to 500 μM [Q], suggesting the quenching by one of the two classic modes (static or dynamic) occurred in the formation of the GA–tyrosinase complex.23 The calculated KSV value in Table 1 positively correlated with temperature. Moreover, the corresponding Kq values at 298, 304 and 310 K were calculated to be (1.65 ± 0.05) × 109, (1.94 ± 0.06) × 109 and (2.71 ± 0.11) × 109 L mol−1 s−1, respectively, which were considerably lower than the maximum scatter collision quenching constant of various quenchers with biopolymers (2.0 × 1010 L mol−1 s−1)24 and the Kq value increased with increasing temperature. These results indicated that dynamic or collisional quenching was dominant for the GA–tyrosinase interaction.25
| T(K) | K SV (L mol−1) | R | K a (×106 L mol−1) | n | R | ΔH (kJ mol−1) | ΔG (kJ mol−1) | ΔS (J mol−1 K−1) |
|---|---|---|---|---|---|---|---|---|
| a R is the correlation coefficient for the KSV values. b R is the correlation coefficient for the Ka values. | ||||||||
| 298 | 16.53 | 0.99295 | 1.07 | 2.137 | 0.9915 | 20.309 | −51.52 | 854.4 |
| 304 | 19.44 | 0.99361 | 5.51 | 2.284 | 0.9920 | −56.7 | ||
| 310 | 27.05 | 0.98926 | 25.53 | 2.406 | 0.9865 | −61.77 | ||
It is also important to study whether GA alters the tyrosinase microenvironment around the tyrosine (Tyr) and tryptophan (Trp) groups. Fig. 2C and D show the synchronous fluorescence intensities of tyrosinase at different GA concentrations (0 → 22 μM) when the scanning interval Δλ was fixed at 15 and 60 nm, respectively. With increasing GA concentration, the maximum emission wavelength of Tyr residue only changed slightly (Δλ = 15 nm) (Fig. 2C), which indicated that GA had little effect on the microenvironment of the Tyr residues. This is consistent with the results for the Trp residue (Δλ = 60 nm) (Fig. 2D).
3.4. Thermodynamic parameters
Generally, the interaction between proteins and small molecules involves several forces, including hydrogen bonding, van der Waals forces, electrostatic interactions, and hydrophobic forces. Thermodynamic parameters are closely connected to changes in interaction forces: a negative ΔH and positive ΔS are characteristic of electrostatic interactions; negative ΔH and ΔS are characteristic of van der Waals forces and hydrogen bonding; positive ΔH and ΔS are characteristic of hydrophobic interactions.The values of ΔH, ΔS and ΔG for the GA–tyrosinase interaction are presented in Table 1. The negative values of ΔG indicate that the binding process was spontaneous. The values of ΔH and ΔS were +20.3 kJ mol−1 and +0.85 kJ mol−1 K−1, respectively, suggesting that the interaction was exothermic and mainly enthalpy favored. These thermodynamic values suggest that hydrophobic forces play an important role in the interaction between GA and tyrosinase.26
3.5. Conformational studies of tyrosinase
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 1
1 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4), the content of α-helix decreased from 33.9% to 18.6%, while the content of β-sheet and random coil increased from 18.6% to 26.2% and from 25.1% to 32.7%, respectively. These results suggest that the addition of GA caused an alteration in the secondary structure of tyrosinase. Previously, it has been reported that citric acid treatment decreased the α-helix content, increased the β-sheet content, but did not alter the β-turn.27 This change in the secondary structure of tyrosinase would be expected to alter its enzymatic activity as reported previously.28
4), the content of α-helix decreased from 33.9% to 18.6%, while the content of β-sheet and random coil increased from 18.6% to 26.2% and from 25.1% to 32.7%, respectively. These results suggest that the addition of GA caused an alteration in the secondary structure of tyrosinase. Previously, it has been reported that citric acid treatment decreased the α-helix content, increased the β-sheet content, but did not alter the β-turn.27 This change in the secondary structure of tyrosinase would be expected to alter its enzymatic activity as reported previously.28
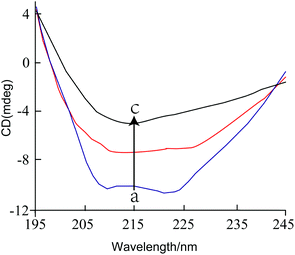 | ||
Fig. 3 The CD spectra of tyrosinase in the presence of increasing amounts of GA. c (tyrosinase) = 2.4 μM, the molar ratios of GA to tyrosinase were 0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (a), 1 1 (a), 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8 (b) and 1 8 (b) and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 (c), respectively. 4 (c), respectively. | ||
Molar ratio [GA]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [tyrosinase] [tyrosinase] |
α-Helix (%) | β-Sheet (%) | β-Turn (%) | Random coil (%) |
|---|---|---|---|---|
0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
33.9 | 18.6 | 22.4 | 25.1 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8 8 |
26.7 | 21.4 | 23.2 | 28.7 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 4 |
18.6 | 26.2 | 22.5 | 32.7 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
19.1 | 25.4 | 21.7 | 33.8 |
8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
20.5 | 24.7 | 22.4 | 32.4 |
32![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
19.7 | 23.4 | 22.5 | 34.4 |
| Peptide location | m/z | [M + H]+ | Sequence | ||
|---|---|---|---|---|---|
| Observed | Theory | Error (ppm) | |||
| 2–8 | 808.4255 | 808.4255 | 808.4240 | 0.0015 | (M)SDKKSLM(P) |
| 17–31 | 1830.0617 | 1830.0617 | 1830.0596 | 0.0021 | (E) IKNRLNILDFVKNDK(F) |
| 47–71 | 1409.1634+2 | 2817.3195 | 2817.3155 | 0.004 | (R) DQSDYSSFFQLGGIHGLPYTEWAKA(Q) |
| 54–83 | 1165.2562+3 | 3493.754 | 3493.7514 | 0.0026 | (S)FFQLGGIHGLPYTEWAKAQPQLHLYKANYC(T) |
| 86–90 | 536.3092 | 536.3092 | 536.3079 | 0.0013 | (H) GTVLF(P) |
| 90–100 | 1394.6483 | 1394.6483 | 1394.6488 | −0.0005 | (L) FPTWHRAYEST(W) |
| 97–116 | 1187.0607+2 | 2373.1141 | 2373.1146 | −0.0005 | (L) FPTWHRAYEST(W) |
| 104–113 | 1018.5180 | 1018.5180 | 1018.5204 | −0.0024 | (Q) TLWEAAGTVA(Q) |
| 119–124 | 735.2955 | 735.2955 | 735.2944 | 0.0011 | (T) SDQAEW(I) |
| 124–130 | 876.4654 | 876.4654 | 876.4668 | −0.0012 | (E) WIQAAKD(L) |
| 145–163 | 1114.6025+2 | 2228.1977 | 2228.2001 | −0.0024 | (D) PDFIGLPDQVIRDKQVEIT(D) |
| 172–176 | 571.3096 | 571.3096 | 571.3086 | 0.001 | (E) VENPI(L) |
| 185–199 | 894.3828+2 | 1787.7583 | 1787.7581 | 0.0002 | (I)EPTFEGDFAQWQTTM(R) |
| 195–206 | 799.3980+2 | 1597.7887 | 1597.7871 | 0.0016 | (Q) WQTTMRYPDVQK(Q) |
| 206–211 | 760.3834 | 760.3834 | 760.3836 | −0.0002 | (Q) KQENIE(G) |
| 216–256 | 907.8537+5 | 4535.2394 | 4535.2353 | 0.0041 | (A) GIKAAAPGFREWTFNMLTKNYTWELFSNHGAVVGAHANSLE(M) |
| 256–263 | 966.4443 | 966.4443 | 966.4462 | −0.0019 | (L) EMVHNTVH(F) |
| 262–279 | 661.6978+3 | 1983.0788 | 1983.0811 | −0.0023 | (T)VHFLIGRDPTLDPLVPGH(M) |
| 307–319 | 768.8197+2 | 1536.6321 | 1536.6345 | −0.0024 | (W) QTMNYDVYVSEGM(N) |
| 330–335 | 646.3411 | 646.3411 | 646.3406 | 0.0005 | (P) GQVLTE(D) |
| 361–366 | 683.3416 | 683.3416 | 683.3399 | 0.0017 | (T) LGFSYP(D) |
| 366–378 | 738.3725+2 | 1475.7377 | 1475.7377 | 0 | (Y) PDFDPVKGKSKEE(K) |
| 375–390 | 1000.0034+2 | 1998.9995 | 1999.0032 | −0.0037 | (K) SKEEKSVYINDWVHKH(Y) |
| 392–421 | 1132.5691+3 | 3395.6927 | 3395.6979 | −0.0052 | (Y)GFVTTQTENPALRLLSSFQRAKSDHETQYA(L) |
| 398–405 | 913.5121 | 913.5121 | 913.5102 | 0.0019 | (Q) TENPALRL(L) |
| 405–431 | 1076.2021+3 | 3226.5917 | 3226.5963 | −0.0046 | (R) LLSSFQRAKSDHETQYALYDWVIHATF(R) |
| 428–445 | 1143.0573+2 | 2285.1073 | 2285.1026 | 0.0047 | (I) HATFRYYELNNSFSIIFY(F) |
| 478–483 | 646.3421 | 646.3421 | 646.3406 | 0.0015 | (R) SQDLIA(E) |
| 484–497 | 821.8834+2 | 1642.7595 | 1642.7570 | 0.0025 | (A) EGFVHLNYYIGCDI(G) |
| 496–523 | 1108.2313+3 | 3322.6793 | 3322.6815 | −0.0022 | (C) DIGQHADHEDDAVPLYEPTRVKEYLKKR(K) |
| 529–534 | 561.2891 | 561.2891 | 561.2879 | 0.0012 | (K) VVSAEG(E) |
| 531–536 | 605.2789 | 605.2789 | 605.2777 | 0.0012 | (V) SAEGEL(T) |
| 545–550 | 683.3416 | 683.3416 | 683.3399 | 0.0017 | (K) GAPYYL(P) |
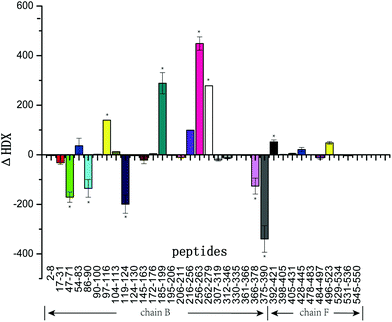
Fig. 4 summarizes the ΔHDX values for all peptides corresponding to the mass differences of tyrosinase peptides in the presence and absence of GA. As illustrated in Fig. 4, peptides 47–71, 86–90, 119–124, 366–378, and 375–390 were protected from deuterium exchange in the presence of GA with decreased deuterium incorporation. Peptides 97–116, 185–199, 216–256, 256–263, and 262–279 exhibited increased deuterium incorporation in the presence of GA. Peptides 2–8, 90–100, 307–319, 361–366, 398–405, 405–431, 478–483 were not affected by the addition of this inhibitor, showing little changes in deuterium incorporation. Tyrosinase comprises four subunits (chain A, E; chain B, F; chain C, G; chain D, H), however, every subunit has similar active sites that contain two copper atoms and some specific amino acids. In this work, chain B and F (Fig. 5) were selected for study. Interestingly, the changes in deuterium incorporation were observed for most peptides in chain B, while little in chain F, suggesting GA disturbs the structure of chain B much greater than chain F. Peptides 97–116, 121–124, 191–194, 216–256, 256–263, 262–279, 366–378 and 375–390 are within α-helix structures. The amount of α-helix present was greatly changed by GA, which is consistent with the results from CD analysis.
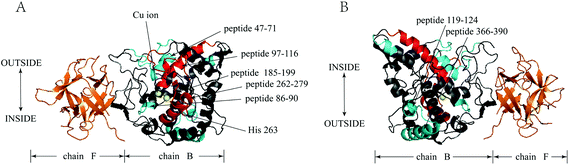 | ||
| Fig. 5 Tyrosinase (PDB code 2Y9X): map of local HDX alterations in the presence of GA front (A) & side (B). Peptides are color-coded as follows: orange & blank = insignificant change in deuterium incorporation; cyan = the values of ΔHDX was negative; red = the values of ΔHDX was positive; blue = the peptide 262–279 (His 263). | ||
4. Discussion
The browning reactions that occur in foods during processing and storage are often induced by tyrosinase-catalyzed oxidation of phenolic compounds. These enzymatic browning reactions adversely affect the quality and sensory properties of fruits and vegetables, and so there is considerable interest in developing effective strategies to inhibit them. In particular, there is interest in identifying cost-effective food-grade inhibitors that can reduce the activity of tyrosinase. Studies in the cosmetics area have shown that glycolic acid can inhibit melanin formation by inhibiting tyrosinase activity, and that this natural organic acid has little toxicity to humans. Consequently, GA may be an effective inhibitor that can be used to inhibit the browning of foods.In this study, GA was shown to inhibit the catalytic activity of tyrosinase with an IC50 value of 83 ± 14 μM (determined spectrophotometrically). Data on the IC50 values of various tyrosinase inhibitors published in the literature or determined in this study can be ranked in the following order: cinnamic acid (IC50 = 7.5 mM)29 > morin (IC50 = 81 mM)15 ≈ GA (IC50 = 83 mM) > kojic acid (IC50 = 39 μM). These values indicate that GA has a similar or better inhibitory effect as many known tyrosinase inhibitors. Glycolic acid has the lowest molecular weight of the alpha-hydroxy acids and would therefore be expected to easily penetrate into the tissues of fruit and vegetables. Moreover, it can be isolated from natural plants or chemically synthesized, which means that it is commercially viable. Thus, glycolic acid would appear to be an extremely promising candidate for preventing the tyrosinase-catalyzed browning of foods.
The CD spectra suggest that there was an appreciable change in the secondary structure of the tyrosinase when the GA-to-tyrosinase ratio was increased from 0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 1
1 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 (i.e., from 0 to 0.6 μM GA), but that there was little further change in secondary structure when the GA level was increased higher (Fig. 3 and Table 2). In particular, there was a substantial decrease in the percentage of α-helix structure, and a corresponding increase in random coil structure, with little change in the percentages of β-sheet or β-turn structures. The GA levels where these changes in secondary structure were observed are appreciably lower than the IC50 value calculated for the inhibition of the catalytic activity of this enzyme, i.e., 83 μM. A possible explanation of this phenomenon is that the CD results only reflect changes in the secondary structure of the proteins (α-helix, β-sheet, β-turn, and random coil), rather than changes in their overall tertiary or quaternary structures. Thus, the addition of low levels of inhibitor (0 to 0.6 μM GA) could have caused appreciable changes in the local (α-helix) structure in certain regions of tyrosinase, without causing appreciable changes in the active site. Other studies have also reported that there may not be a strong correlation between changes in secondary structure and loss of tyrosinase activity.30 Overall, our results suggests there may have been some changes in the secondary structure of the enzyme at certain locations when low GA levels were added, without altering the characteristics of the active site.
4 (i.e., from 0 to 0.6 μM GA), but that there was little further change in secondary structure when the GA level was increased higher (Fig. 3 and Table 2). In particular, there was a substantial decrease in the percentage of α-helix structure, and a corresponding increase in random coil structure, with little change in the percentages of β-sheet or β-turn structures. The GA levels where these changes in secondary structure were observed are appreciably lower than the IC50 value calculated for the inhibition of the catalytic activity of this enzyme, i.e., 83 μM. A possible explanation of this phenomenon is that the CD results only reflect changes in the secondary structure of the proteins (α-helix, β-sheet, β-turn, and random coil), rather than changes in their overall tertiary or quaternary structures. Thus, the addition of low levels of inhibitor (0 to 0.6 μM GA) could have caused appreciable changes in the local (α-helix) structure in certain regions of tyrosinase, without causing appreciable changes in the active site. Other studies have also reported that there may not be a strong correlation between changes in secondary structure and loss of tyrosinase activity.30 Overall, our results suggests there may have been some changes in the secondary structure of the enzyme at certain locations when low GA levels were added, without altering the characteristics of the active site.
Further insights into the conformational changes that occurred when GA interacted with tyrosinase were therefore obtained using HDX-MS. HDX-MS relies on the exchange of protein backbone amide hydrogen atoms with deuterium in solution. Backbone amide hydrogen atoms involved in weak hydrogen bonds or located at the surface of a protein can typically exchange rapidly, whereas those involved in strong hydrogen bonds or buried in the interior of the protein exchange more slowly. In this research, HDX-MS was first applied to elucidate the GA inhibition mechanism of tyrosinase. The binuclear copper binding site of tyrosinase is located at the heart of two pairs of antiparallel α-helices (α3/α4 and α10/α11, respectively), which make an angle of nearly 90° with each other.32 The ligands of the first copper ion, Cu-A, are the Nε2 atoms of His61 (end of helix α3), His85 (in the loop connecting α3 and α4), and His94 (beginning of α4). The ligands of the second copper ion, Cu-B, are the Nε2 atoms of His259, His263 (α10), and His296 (α11). The results from our HDX-MS study indicated that deuterium incorporation into tyrosinase occurred at peptides 256–263 (His263) and 262–279 (His263), which suggested that the tyrosinase active sites were disturbed by GA (Fig. 5). Since copper is known to play an important role in maintaining the activity of tyrosinase,31 it would be useful to measure its potential release from the enzyme in the presence of GA in future studies.
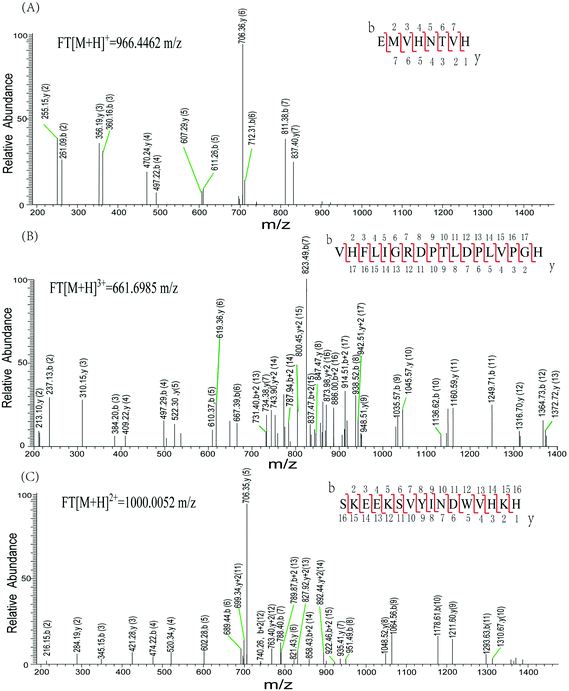
Characteristic peaks (peptides 256–263 and peptides 262–279) from tyrosinase were identified using a LTQ mass spectrometer (LTQ-MS) (Fig. 6). Both peptides were found in tyrosinase untreated or treated by GA. His263 is identified in peptides 256EMVHNTVH263 and 262VHFLIGRDPTLDPLVPGH279. The LTQ-MS of the peptides generated a series of b and y ions (b2–b7 and y1–y7, b2–b17 and y2–y17), which unambiguously confirmed the sequence of the peptide. Similarly, peptides 375–390 were identified by the LTQ-MS with m/z of 1000.0039+. The consecutive b and y ions ensure the positive identification of the peptide sequence. The peptides 86–90 were observed with significantly increased deuterium incorporation, suggesting that this area was disturbed by GA to a more open conformation. Phe90 is wedged between His94, His263, and His296, and so its conformational change may impact histidine side-chain conformations, therefore affecting the integrity of the copper binding sites.33 Our results indicate that dynamic or collisional quenching was the dominant mechanism for the GA–tyrosinase interaction, and the calculated thermodynamic parameters suggest that hydrophobic forces played an important role in the GA–tyrosinase interaction. The catalytic pocket of tyrosinase is known to be hydrophobic,34 and therefore GA may have interacted with tyrosinase primarily through hydrophobic attraction. It should be noted that GA is a highly hydrophilic molecule, although it does have some hydrocarbon regions that give it some amphiphilic character (Fig. 1A). The inhibition mechanism of GA on tyrosinase activity was investigated by various spectroscopic methods including ultraviolet-visible, fluorescence, and CD spectroscopy coupled with kinetic analysis and FTICR-MS. The results of our study demonstrated that (i) GA had a significant inhibitory activity on tyrosinase; (ii) GA was a mixed inhibitor that reversibly inhibited tyrosinase; (iii) GA bound to tyrosinase and caused a major change in the conformation of the enzyme; and, (iv) the tyrosinase active sites were disturbed due to GA binding. Our findings are supported by the results of molecular modeling simulations on related systems. For example, Wang et al. predicted that morin bound to the active site of tyrosinase using molecular modeling, which led to a conformational change of the tyrosinase molecule.15
Our study has shown that HDX-MS is a powerful analytical method that provides valuable information that can be used to elucidate the mechanism of interaction between tyrosinase and its inhibitors. This pratical technology could be used to screen different biological molecules to determine their effectiveness as tyrosinase inhibitors.
5. Conclusion
In this study, it was shown that glycolic acid had a strong inhibitory activity against tyrosinase, which was attributed to the binding of glycolic acid molecules to the tyrosinase and to a subsequent conformational change of the enzyme. These results suggest that glycolic acid could be used as a natural preservative in foods to prevent enzymatic browning reactions. It was also shown that HDX-MS is a powerful new tool for providing insights into the molecular basis of the binding interaction. In particular, we were able to show that glycolic acid molecules bound to the active size of the tyrosinase. HDX-MS may be particularly useful in the identification of new sources of tyrosinase inhibitors, especially when used in conjunction with other analytical methods. Future studies show examine the effectiveness of glycolic acid as an anti-browning agent in actual foods and beverages.Abbreviations
| GA | Glycolic acid |
| LA | Lactic acid |
| HDX-MS | Hydrogen/deuterium exchange with mass spectrometry |
| GA | Glycolic acid |
| UV-vis | Ultraviolet visible spectrometer |
| FL | Fluorescence spectrum |
| CD | Circular dichroism spectrum |
| FA | Formic acid |
| PBS | Sodium phosphate buffer |
| DMSO | Dimethyl sulfoxide |
| FTICR MS | Fourier transform ion cyclotron mass spectrometry |
| Tyr | Tyrosine |
| Trp | Tryptophan |
Acknowledgements
This work was supported by National High Technology Research and Development Program of China (863 Program, no. 2013AA102205), National Natural Science Foundation of China (NSFC) (no. 21276118).References
- M. V. Martinez and J. R. Whitaker, The biochemistry and control of enzymatic browning, Trends Food Sci. Technol., 1995, 6, 195–200 CrossRef CAS.
- S. J. Brooks, J. Nikodinovic, L. Martin, E. M. Doyle, T. O'Sullivan, P. J. Guiry, L. Coulombel, Z. Li and K. E. O'Connor, Production of a chiral alcohol, 1-(3, 4-dihydroxyphenyl) ethanol, by mushroom tyrosinase, Biotechnol. Lett., 2013, 35, 779–783 CrossRef CAS PubMed.
- S.-Y. Seo, V. K. Sharma and N. Sharma, Mushroom tyrosinase: recent prospects, J. Agric. Food Chem., 2003, 51, 2837–2853 CrossRef CAS PubMed.
- M. Jiménez, S. Chazarra, J. Escribano, J. Cabanes and F. García-Carmona, Competitive inhibition of mushroom tyrosinase by 4-substituted benzaldehydes, J. Agric. Food Chem., 2001, 49, 4060–4063 CrossRef.
- Z. Yang, Y. Wang, Y. Wang and Y. Zhang, Bioassay-guided screening and isolation of α-glucosidase and tyrosinase inhibitors from leaves of Morus alba, Food Chem., 2012, 131, 617–625 CrossRef CAS.
- B. Deng, C. Lento and D. J. Wilson, Hydrogen deuterium exchange mass spectrometry in biopharmaceutical discovery and development - A review, Anal. Chim. Acta, 2016, 940, 8–20 CrossRef CAS PubMed.
- L. Konermann, J. X. Pan and Y. H. Liu, Hydrogen exchange mass spectrometry for studying protein structure and dynamics, Chem. Soc. Rev., 2011, 40, 1224–1234 RSC.
- J. J. Lee, Y. S. Park and K. J. Lee, Hydrogen-deuterium exchange mass spectrometry for determining protein structural changes in drug discovery, Arch. Pharmacal Res., 2015, 38, 1737–1745 CrossRef CAS PubMed.
- D. Datta and S. Kumar, Modeling using response surface methodology and optimization using differential evolution of reactive extraction of glycolic acid, Chem. Eng. Commun., 2015, 202, 59–69 CrossRef CAS.
- A. Usuki, A. Ohashi, H. Sato, Y. Ochiai, M. Ichihashi and Y. Funasaka, The inhibitory effect of glycolic acid and lactic acid on melanin synthesis in melanoma cells, Exp. Dermatol., 2003, 12, 43–50 CrossRef CAS PubMed.
- A. Papadopoulou, R. J. Green and R. A. Frazier, Interaction of flavonoids with bovine serum albumin: a fluorescence quenching study, J. Agric. Food Chem., 2005, 53, 158–163 CrossRef CAS PubMed.
- L. Qiu, Q.-H. Chen, J.-X. Zhuang, X. Zhong, J.-J. Zhou, Y.-J. Guo and Q.-X. Chen, Inhibitory effects of α-cyano-4-hydroxycinnamic acid on the activity of mushroom tyrosinase, Food Chem., 2009, 112, 609–613 CrossRef CAS.
- Q. Zhang, Z. Tu, H. Wang, X. Huang, X. Sha and H. Xiao, Structural changes of ultrasonicated bovine serum albumin revealed by hydrogen–deuterium exchange and mass spectrometry, Anal. Bioanal. Chem., 2014, 406, 7243–7251 CrossRef CAS PubMed.
- T. E. Wales and J. R. Engen, Hydrogen exchange mass spectrometry for the analysis of protein dynamics, Mass Spectrom. Rev., 2006, 25, 158–170 CrossRef CAS PubMed.
- Y. Wang, G. Zhang, J. Yan and D. Gong, Inhibitory effect of morin on tyrosinase: Insights from spectroscopic and molecular docking studies, Food Chem., 2014, 163, 226–233 CrossRef CAS PubMed.
- Q.-X. Chen, K.-K. Song, L. Qiu, X.-D. Liu, H. Huang and H.-Y. Guo, Inhibitory effects on mushroom tyrosinase by p-alkoxybenzoic acids, Food Chem., 2005, 91, 269–274 CrossRef CAS.
- J. Yan, G. Zhang, Y. Hu and Y. Ma, Effect of luteolin on xanthine oxidase: Inhibition kinetics and interaction mechanism merging with docking simulation, Food Chem., 2013, 141, 3766–3773 CrossRef CAS PubMed.
- O. Nerya, J. Vaya, R. Musa, S. Izrael, R. Ben-Arie and S. Tamir, Glabrene and isoliquiritigenin as tyrosinase inhibitors from licorice roots, J. Agric. Food Chem., 2003, 51, 1201–1207 CrossRef CAS PubMed.
- J. S. Chen, C.-i. Wei, R. S. Rolle, W. S. Otwell, M. O. Balaban and M. R. Marshall, Inhibitory effect of kojic acid on some plant and crustacean polyphenol oxidases, J. Agric. Food Chem., 1991, 39, 1396–1401 CrossRef CAS.
- F. Ge, L. Jiang, D. Liu and C. Chen, Interaction between alizarin and human serum albumin by fluorescence spectroscopy, Anal. Sci., 2011, 27, 79–84 CrossRef CAS PubMed.
- Y. Wang, G. Zhang and L. Wang, Potential toxicity of phthalic acid esters plasticizer: Interaction of dimethyl phthalate with trypsin in vitro, J. Agric. Food Chem., 2014, 75–84 Search PubMed.
- Z. Cheng, Studies on the interaction between scopoletin and two serum albumins by spectroscopic methods, J. Lumin., 2012, 132, 2719–2729 CrossRef CAS.
- Y. Li, W. He, J. Liu, F. Sheng, Z. Hu and X. Chen, Binding of the bioactive component jatrorrhizine to human serum albumin, Biochim. Biophys. Acta, Gen. Subj., 2005, 1722, 15–21 CrossRef CAS PubMed.
- J. R. Lakowicz, Principles of fluorescence spectroscopy, Springer Science & Business Media, 2007 Search PubMed.
- E.-H. Liu, L.-W. Qi and P. Li, Structural relationship and binding mechanisms of five flavonoids with bovine serum albumin, Molecules, 2010, 15, 9092–9103 CrossRef CAS PubMed.
- P. D. Ross and S. Subramanian, Thermodynamics of protein association reactions: forces contributing to stability, Biochemistry, 1981, 20, 3096–3102 CrossRef CAS PubMed.
- W. Liu, L.-q. Zou, J.-p. Liu, Z.-q. Zhang, C.-m. Liu and R.-h. Liang, The effect of citric acid on the activity, thermodynamics and conformation of mushroom polyphenoloxidase, Food Chem., 2013, 140, 289–295 CrossRef CAS PubMed.
- J. Yan, G. Zhang, J. Pan and Y. Wang, α-Glucosidase inhibition by luteolin: Kinetics, interaction and molecular docking, Int. J. Biol. Macromol., 2014, 64, 213–223 CrossRef CAS PubMed.
- L. Zhou, W. Liu, Z. Xiong, L. Zou, J. Chen, J. Liu and J. Zhong, Different modes of inhibition for organic acids on polyphenoloxidase, Food Chem., 2016, 199, 439–446 CrossRef CAS PubMed.
- N. Gheibi, A. Saboury, K. Haghbeen and A. Moosavi-Movahedi, Activity and structural changes of mushroom tyrosinase induced by n-alkyl sulfates, Colloids Surf., B, 2005, 45, 104–107 CrossRef CAS PubMed.
- E. J. Land, C. A. Ramsden and P. A. Riley, The mechanism of suicide-inactivation of tyrosinase: a substrate structure investigation, Tohoku J. Exp. Med., 2007, 212, 341–348 CrossRef CAS PubMed.
- W. T. Ismaya, H. J. Rozeboom, A. Weijn, J. J. Mes, F. Fusetti, H. J. Wichers and B. W. Dijkstra, Crystal structure of Agaricus bisporus mushroom tyrosinase: identity of the tetramer subunits and interaction with tropolone, Biochemistry, 2011, 50, 5477–5486 CrossRef CAS PubMed.
- B. Hazes, K. H. Kalk, W. Hol, K. A. Magnus, C. Bonaventura, J. Bonaventura and Z. Dauter, Crystal structure of deoxygenated Limulus polyphemus subunit II hemocyanin at 2.18 Å resolution: clues for a mechanism for allosteric regulation, Protein Sci., 1993, 2, 597–619 CrossRef CAS PubMed.
- S. Radhakrishnan, R. Shimmon, C. Conn and A. Baker, Integrated kinetic studies and computational analysis on naphthyl chalcones as mushroom tyrosinase inhibitors, Bioorg. Med. Chem. Lett., 2015, 25, 4085–4091 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c6fo01384h |
| This journal is © The Royal Society of Chemistry 2017 |