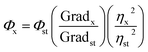Hyaluronic acid functionalized nitrogen-doped carbon quantum dots for targeted specific bioimaging
M. Zhang†
a,
Z. Fang†a,
X. Zhaoa,
Y. Niua,
J. Loua,
L. Zhaoa,
Y. Wua,
S. Zoub,
F. Du*ab and
Q. Shao*a
aSchool of Medicine, Jiangsu University, Zhenjiang, 212013, P. R. China. E-mail: shao_qx@ujs.edu.cn; biodfy@ujs.edu.cn; Tel: +86-511-85038145
bDepartment of Hepatosis, The Third Hospital of Zhenjiang Affiliated to Jiangsu University, Zhenjiang, Jiangsu, P. R. China
First published on 20th October 2016
Abstract
Carbon quantum dots (CQDs), a new inorganic quasi-zero dimensional carbon nanomaterial, are extensively applied in biomedical fields. However, the lack of specific cell targeting ligands on the surface has consistently been a critical problem to be overcome in application of CQD technology. In this study, hyaluronic acid-functionalized CQDs (HA-CQDs) were synthesized via hydrothermal carbonization for tumor-targeted bioimaging. The synthesized HA-CQDs possessed highly hydrophilic surfaces, uniform size distribution (approximately 2.3 nm) and heterogeneous multi-layered crystal structures. Due to the superior fluorescence property of HA-CQDs, the labeled cells could be observed and detected using a confocal microscope and flow cytometry. By virtue of the functional HA ligands, HA-CQDs exhibited excellent colloidal stability and favorable biocompatibility, and readily entered into the cytoplasm of CD44-overexpression cancer cells. HA competitive binding assay demonstrated that the internalization process was regulated by CD44 (HA receptor)-mediated endocytosis. These results validated that the as-prepared HA-CQDs could particularly serve as novel cell-specific fluorescent probes for CD44 high expression in tumor-targeted imaging and labelling.
1. Introduction
Carbon quantum dots (CQDs), in carbon nanomaterials, have attracted extensive attention due to their remarkable optical properties, water solubility, low cytotoxicity, excellent biocompatibility and cost-effectiveness compared with traditional metal-based semiconductor quantum dots. CQDs have a broad range of biomedical applications1–4 in photothermal therapy, drug delivery,5 biosensors,6 bioimaging7 and contrast agents.8 Furthermore, CQDs are eco-friendly nanomaterials that possess stable photoluminescence and excitation wavelength-dependant emission. However, CQDs are limited as fluorescent probes in biomedical applications on account of their low quantum yield.At present, there are two main strategies for the synthesis of CQDs: top-down and bottom-up methods.5 For top-down strategies, CQDs are generally formed by breaking down large carbonaceous materials, followed by polymerization into nanoparticles using physical or chemical methods9 such as electrochemical oxidation and plasma treatment. However, purification and functionalization of CQDs under top-down strategies involve complex and time-consuming procedures. For bottom-up strategies, CQDs are produced by carbonization of small molecules, involving polyhydroxy carbohydrates and passivation agents. Based on numerous reports, it is speculated that the passivation agent containing terminal NH2 groups fills the surface defects of the CQD, and the energy of the surface caves the CQDs' emission of visible light. However, this method requires expensive organic passivation agents and the operation process is very complicated. Therefore, we paid more attention to develop a simple and feasible method to improve the fluorescence yield of carbon quantum dots; an effective and convenient method for this is doping CQDs with nitrogen. Recently, our group has demonstrated that nitrogen-doped carbon dots (NCDs) exhibited heterogeneous multi-layered structures, which corresponded to the characteristic planes between graphite and graphite oxide.10–13 As a fluorescent probe, CQDs have been proven highly effective in cell labeling, which is a technology that holds great potential applications in many biomedical areas, particularly in vivo imaging and early diagnosis.14 However, the application towards targeted tumor administration remains a challenge. Several determining factors are still to be resolved, and several research areas are underdeveloped; for example, the synthesis of novel CQDs excited by near infrared light with deeper tissue penetration, as well as target-specific CQDs for enhanced tumor recognition and ablation.
Application of CQDs to biological imaging was first investigated by Cao et al.2 They found that MCF-7 cells, a type of human breast cancer cell line, emitted brighter light under a two-photon fluorescence microscope upon activation triggered by a pulse of 800 nm when cultured in a CQD-buffer at 37 °C. Yang et al.15 observed a high and stable fluorescence intensity in the urine of the mice after back, lower limb or vein injection of CQDs, all without impairing the status of the mice. Yang et al.16 reported that folic acid molecules were introduced into the carbon dots, and these modified carbon dots accumulated preferentially in FA-receptor-positive cancer cells. In summary, the application of CQDs in biomedicine and bioimaging has a broader prospect compared with that of traditional biological strains and CQDs.
In this study, we developed a facile and effective bottom-up route to synthesize HA-CQDs using hyaluronic acid and glycine as precursors. The amino group was found to be critical for the carbonization,9 polymerization, and formation of HA-CQDs. We proposed a mechanism by which nitrogen was doped into the carbon dots by the chemical and structural information of the HA-CQDs. HA acted as a ligand for CD44, and it was therefore expected that HA-CQDs had the potential for targeted delivery. As a result, the HA-CQDs were successfully tested as a new biocompatible probe with the ability to distinguish CD44 receptor cells from other cells. Therefore, application of HA-CQDs as fluorescence probes for labelling and imaging of living cells was demonstrated.
2. Experimental
2.1 Materials and reagent
Quinine sulfate (98%, suitable for fluorescence), glycine, and hyaluronate were utilized as the precursors for the synthesis of carbon quantum dots. 3-(4,5-Dimethyl-2-thiazoyl)-2,5-diphenyltetrazolium bromide (CCK-8 assay, 98%) was applied to reveal cytotoxicity of carbon quantum dots. Fetal bovine serum (FBS) and Dulbecco's Minimum Essential Medium (DMEM) were purchased from Invitrogen China Limited (China, Shanghai). PCR green master mix was obtained from Thermo (UAS). PCR primer was obtained from Shanghai Sangon Company (Shanghai, China). All chemicals were of analytical grade and were used without further purification. All the aqueous solutions were prepared using double distilled water.2.2 Synthesis of HA-CQDs
First, 0.2 g hyaluronate was dissolved in 20 mL water and mixed with 0.35 g glycine under vigorous stirring to form a transparent homogeneous solution in a common beaker. This solution was then transferred into a 50 mL Teflon-lined stainless steel autoclave and heated at 200 °C for 4 h. The black precipitates were removed by centrifugation at 2000 rpm for 15 min. The supernatant was transferred into a dialysis membrane (MWCO of 1000 Dalton) and dialyzed against pure water for 4 days. Finally, the brown-yellow solution was collected and freeze-dried using a vacuum freeze dryer.2.3 Apparatus and characterization
A Fourier transform infrared (FT-IR) spectrometer (Nicolet Nexus 470, America) was applied to analyse the chemical structures of HA-CQDs. The morphologies of the HA-CQDs were examined by high-resolution transmission electron microscopy (HRTEM) on a JEM-2100 microscope (JEOL, Japan) at an accelerating voltage of 200 kV. UV-Vis absorption spectra were recorded using a UV-2450 UV-Vis spectrophotometer (Shimadzu, Japan). Photoluminescence (PL) emission measurements were obtained using a Cary Eclipse Fluorometer (Varian, America).2.4 Fluorescence quantum yields
The quantum yield (Φ) of the as-prepared carbon nanodots was determined by a comparative method. Quinine sulfate (literature quantum yield: 54%) was dissolved in 0.1 M H2SO4 (refractive index, η = 1.33) and the HA-CQDs were dissolved in distilled water (η = 1.33) at different concentrations. Aqueous HA-CQDs solutions at different concentrations were prepared by dissolving the desired quantity in distilled water. Photoluminescence (PL) emission spectra of all the samples were recorded at an excitation wavelength of 360 nm. The samples were then measured by PL spectrometer in order to get the PL emission intensity at the excitation wavelength with which the C-dots and the reference had the same UV absorbance. Then, the quantum yield was calculated according to the following equation:where st and x referred to standard group and test group, respectively; Φ is the fluorescence quantum yield, grad is the gradient from the plot of integrated fluorescence intensity vs. absorbance, and η is the refractive index of the solvent. In order to minimize the re-absorption effects, absorbance in the 10 mm fluorescence cuvette did not exceed 0.1 at the excitation wavelength.
2.5 Cell viability assay
The cytotoxicity of HA-CQDs was evaluated on Hela and MCF-7 cells using a modified CCK8. Cells were seeded in a 96-well plate at a density of 3 × 104 cells per well at 37 °C and 5% CO2. After overnight incubation, the growth medium was replaced with DMEM containing different concentrations of HA-CQDs. After incubation for 24 h, 20 μL of 2.5 mg mL−1 MTT solution (in 0.01 M sterilized PBS, pH 7.4) was added to each well and the plates were incubated for an additional 4 h at 37 °C and 5% CO2. After the removal of the medium, 150 μL DMSO was added to each well and the mixtures were shaken for 15 min. The absorbance of each well at 490 nm was recorded using a Synergy HT Multi-Mode Microplate Reader (BioTek, USA). Untreated cells (in DMEM) were used as the control and the relative cell viability (mean ± SD, n = 3) was expressed as (Abssample/Abscontrol) × 100%.2.6 Hemolysis assay
Animal experiments were executed according to the protocol approved by Animal Management Rules of the Ministry of Health of the People's Republic of China and the guidelines for the Care and Use of the Jiangsu University Laboratory Animal Center. Fresh blood was sampled from rat eyes and stabilized with heparin. The supernatant of the blood was removed by centrifugation at 1000 rpm for 10 min and purified 5 times with PBS. 100 μL of the diluted blood cells was then mixed with the following: 900 μL DD H2O for the positive control, PBS for the negative control, and 900 μL PBS solutions containing HA-CQDs in different concentrations (50, 100, 200, 300, 400 μg mL−1). The mixture was shaken slightly and rested at room temperature for 2 h. Subsequently, the samples were centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 1 min and the absorbance of the supernatant was recorded from each groups.
000 rpm for 1 min and the absorbance of the supernatant was recorded from each groups.
2.7 Cell imaging and labeling
The newly developed HA-CQDs were evaluated for live cell imaging in Hela cells, which were seeded on a 24-well plate and cultured at 37 °C with 5% CO2 for 24 h in a medium containing 10% FBS. HA-CQDs were added to each well at a concentration of 50 mg mL−1 and the mixtures were incubated for 6 h at 37 °C and 5% CO2. The labeled cells were then fixed with 4% paraformaldehyde/PBS and washed with PBS at room temperature for 10 min. These cells were then observed using a Laser Confocal Scanning Microscope (Zeiss LSM-710) and the fluorescence emission from the carbon dots inside the cells was recorded at different excitation wavelengths using an excitation laser.2.8 Agarose gel electrophoresis
Cells were seeded in a 6-well plate and HA-CQDs solution was added at 0, 100, 200, 300, 400, 500 μg mL−1 concentration. Total RNA of the cells was extracted from different concentrations of HA-CQDs after incubation for 24 h. Following this, 0.1 μg of the RNA from each group was reverse transcribed into cDNA using the reverse transcription system. 2 μL of the cDNA was amplified by Thermo® Green Master Mix. PCR was then performed in a PTC-200 Peltier thermal cycler (MJ Research) with the following program: an initial denaturation at 94 °C for 3 min, followed by 29 cycles (for CD44 gene and GAPDH gene) at 94 °C for 30 s, 56 °C for 30 s, and 72 °C for 30 s. After the last cycle, the program was ended by a final extension at 72 °C for 10 min. The sequences of the PCR primer sets used were as follows: CD44 forward: 5′-CTACAGCATCTCTCGGAC-3′ and reverse: 5′-GTCTCAAATCCGATGCTC-3′; GAPDH forward: 5′-TGGGTGTGAACCATGAGAAG-3' and reverse: 5′-GAGTCCTTCCACGATACCAAAG-3′.2.9 Flow cytometry
For cellular binding analysis, MCF-7 and Hela cells were incubated at 37 °C and 5% CO2 for 24 h. Each cell line (3 × 104) was seeded in three wells of the six-well plates. The first well was the negative control, the second well was the HA-CQDs group, and the third well was HA-CQDs plus free HA group (5 mg mL−1), in order to observe the effect of free HA on cellular uptake. After 1 h, the HA dilution solution was discarded and 400 μg mL−1 HA-CQDs solution was added to the second and third well. To remove unbound HA-CQDs, the cells were washed three times and resuspended in PBS. The cells were analyzed using flow cytometry on a FACScalibur cytometer.3. Results and discussion
3.1 The preparation of HA-CQDs
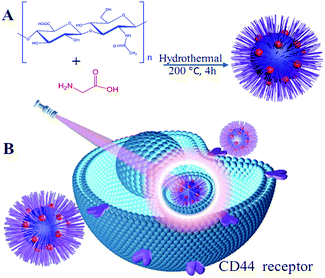
The synthetic process of HA-CQDs is shown in Scheme 1. A mixture of HA and glycine was subjected to a hydrothermal process that included dehydration, polymerization and aromatization. In this process, a certain amount of glycine in the raw solution was necessary to enhance the HA-CQDs formation and was critical for carbonization degree. Finally, the brown solution was dialyzed against distilled water to obtain highly fluorescent HA-CQDs.3.2 The characterization of the HA-CQDs
The as-synthesized HA-CQDs dispersed homogeneously in water without any precipitation under ambient conditions at room temperature. The TEM images displayed in Fig. 1A showed that the HA-CQDs were uniformly spherical and the average diameters were less than 10 nm.17 Fig. 1B displays the diameter distribution of HA-CQDs. The average size of HA-CQDs was about 2.38 nm, which was determined by statistical analysis of more than one hundred particles using the Image J software.18 The HA-CQDs aqueous solution was brown and transparent in daylight but emitted strong blue luminescence under excitation at 365 nm, a change which could be easily observed (Fig. 1C, inset). Furthermore, the optical properties of HA-CQDs were characterized by UV/Vis and photoluminescence (PL) spectroscopy. As indicated in Fig. 1C, the UV/Vis spectrum of HA-CQDs featured an absorption peak located between 300–550 nm, which may give rise to a series of emission traps between n and π* states of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C bond. As shown in Fig. 1D, the emission spectra of the HA-CQDs were broad, ranging from black (310 nm) to brown (380 nm) depending on the excitation wavelengths. Evidently, the strongest fluorescence emission was centered at approximately 470 nm under 360 nm excitation; this may be due to unique surface defect states near the Fermi level and has been widely observed from previously reported luminescent carbon nanomaterials.19
C bond. As shown in Fig. 1D, the emission spectra of the HA-CQDs were broad, ranging from black (310 nm) to brown (380 nm) depending on the excitation wavelengths. Evidently, the strongest fluorescence emission was centered at approximately 470 nm under 360 nm excitation; this may be due to unique surface defect states near the Fermi level and has been widely observed from previously reported luminescent carbon nanomaterials.19
In order to understand the chemical structure and surface compositions of HA-CQDs, FTIR and X-ray photoelectron spectroscopy (XPS) was conducted. The XPS survey spectrum of the as-produced HA-CQDs (Fig. 2A) showed three peaks at 284.0, 400.0, and 530 eV, which indicated that the HA-CQDs were composed of carbon, nitrogen, and oxygen. The high-resolution spectrum of C1s (Fig. 2B) displayed three main peaks at 285.0, 287.8, and 288.9 eV, which could be attributed to C–C, C–O, and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, respectively. Moreover, the binding energy peak at 285 eV confirmed the graphite-like structure (sp2, C–C) of the HA-CQDs. And spectrum of O1s (Fig. 2C) showed two characteristic oxygen states, C
O, respectively. Moreover, the binding energy peak at 285 eV confirmed the graphite-like structure (sp2, C–C) of the HA-CQDs. And spectrum of O1s (Fig. 2C) showed two characteristic oxygen states, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (530.5 eV) and C–O (532.2 eV), which was consistent with the spectrum of C1s. The N1s spectrum (Fig. 2D) revealed two nitrogen species, N–H (402.2 eV) and C–N–C (398.4 eV), which was consistent with the results of FTIR spectral analysis. Among these peaks, the peak at 398.4 eV and 402.2 eV were attributed to the N atoms.
O (530.5 eV) and C–O (532.2 eV), which was consistent with the spectrum of C1s. The N1s spectrum (Fig. 2D) revealed two nitrogen species, N–H (402.2 eV) and C–N–C (398.4 eV), which was consistent with the results of FTIR spectral analysis. Among these peaks, the peak at 398.4 eV and 402.2 eV were attributed to the N atoms.
 | ||
| Fig. 2 XPS spectra of the HA-CQDs. (A) Survey spectrum. (B) C1s spectrum. (C) O1s spectrum. (D) N1s spectrum. | ||
As shown in the FTIR spectrum of HA-CQDs (Fig. 3A), a broad peak centered at 3413 cm−1 and a sharp peak at 2937 cm−1 emerged, revealing O–H and C–H bonding, respectively. Both these peaks were weaker in the spectra of the HA-CQDs, indicating that a drastic reduction in H occurred after the formation of HA-CQDs. Moreover, the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching with a strong vibrational absorption peak was observed for HA-CQDs at 1602 cm−1. Absorption bands at 1405 cm−1 could be attributed to the amide groups (–NHCO–). As shown in Fig. 3B, the X-ray diffraction pattern (XRD) displayed a main broad diffraction peak centered at 12°, which was assigned to highly disordered carbon atoms and was similar to the characteristic lattice spacing {001} and {002} of graphene oxide and graphene, respectively. The third shoulder peak at 2θ = 42.5° (d100 = 0.212 nm) was attributed to in-plane diffraction from the graphene-like structure of the NCDs, which corresponded to the {100} plane of graphite. The two peaks demonstrated that the HA-CQDs have graphene and graphene oxide-like structures.
O stretching with a strong vibrational absorption peak was observed for HA-CQDs at 1602 cm−1. Absorption bands at 1405 cm−1 could be attributed to the amide groups (–NHCO–). As shown in Fig. 3B, the X-ray diffraction pattern (XRD) displayed a main broad diffraction peak centered at 12°, which was assigned to highly disordered carbon atoms and was similar to the characteristic lattice spacing {001} and {002} of graphene oxide and graphene, respectively. The third shoulder peak at 2θ = 42.5° (d100 = 0.212 nm) was attributed to in-plane diffraction from the graphene-like structure of the NCDs, which corresponded to the {100} plane of graphite. The two peaks demonstrated that the HA-CQDs have graphene and graphene oxide-like structures.
3.3 Biocompatibility of HA-CQDs
CCK8 assay was used to evaluate the cytotoxicity of HA-CQDs in Hela and MCF-7 cells. As shown in Fig. 4A, the HA-CQDs showed minimal cytotoxicity compared with the control group. The cell viability remained at approximately 90% after 24 h incubation with HA-CQDs at a concentration of 250 μg mL−1. According to Fig. 4B, the dissolution of blood cells in double distilled water (low osmolality) caused their destruction, which induced an evident increase in absorption value under ultraviolet light.20 Compared with the PBS group, however, the absorption value of HA-CQDs from different concentrations and blood cell showed no prominent change. These results showed that the HA-CQDs had very low cytotoxicity with excellent hemocompatibility.21,223.4 Biomedical applications of HA-CQDs
The potential application of the as-prepared HA-CQDs in imaging and labelling was evaluated by laser confocal scanning microscopy (LCSM). Hela and U251 cells were incubated with the HA-CQDs for 6 h and washed to remove extracellular HA-CQDs. As shown in Fig. 5, the HA-CQDs emitted bright green fluorescence at 496 nm. The bright-field and merged fluorescence images revealed that the HA-CQDs were localized to the cytoplasmic regions, particularly around the nucleus, and the photoluminescence of HA-CQDs was very weak in the cell nucleus. It appeared that a large amount of HA-CQDs were taken up by the cells and spread to the cytoplasm, which may be regulated by receptor-mediated endocytosis within a short time. Therefore, HA-CQDs with high fluorescence and cellular internalization performance were recognized as ideal candidates for real-time cell imaging applications, similar to the previous report.23–26 | ||
| Fig. 5 Fluorescence microscope images of Hela (A) and U251 (B) after incubation of the HA-CQDs (100 μg mL−1) for 6 h. | ||
3.5 Tumor-specific targeting bioimaging and biolabeling by CD44-mediated endocytosis
Natural HA macromolecule, as the main component of extracellular matrix (ECM) and an intercellular substance, can specifically bind to the CD44 receptor, which is a surface molecule that is over-expressed in cancer cells.27 PCR assay could then be conducted to verify whether the functionalized HA-CQDs interacted with CD44. As shown in Fig. 6A, CD44 expression increased with the concentration of HA-CQDs, which demonstrated that as-synthesized HA-CQDs could induced the expression of CD44 at the RNA level. In order to verify their tumor-specific uptake behavior by the receptor-mediated endocytosis, HA-CQDs were incubated with both CD44 high- and low-expression cell lines, Hela and MCF-7. The HA-CQDs taken up by the cells was quantitatively evaluated using flow cytometry after incubation with 200 mg mL−1 for 6 h. Twenty thousand cells were collected and analyzed by flow cytometry. Hela cells (Fig. 6B) effectively took up HA-CQDs, however MCF-7 (Fig. 6C) endocytosed only a very small amount of the nanoparticles. Moreover, this data also confirmed the feasibility of HA-CQDs as effective biolabeling probes for flow cytometry. More importantly, uptake of HA-CQDs in Hela cells was inhibited when they were pre-treated by free HA, however the uptake process in MCD-7 cells was only slightly affected. According to the fluorescence microscopic images (Fig. 6D), it was confirmed that HA-CQDs were readily distributed in cytosols of Hela cells. These results were in a good agreement with the cellular binding behavior of HA-CQDs, and suggested that HA-CQDs could be effectively taken up via HA receptor-mediated endocytosis. Thus, they could serve as cell-specific targeting probes for imaging and labeling of CD44-high expression tumors.4. Conclusions
In summary, we have successfully synthesized HA-CQDs with high fluorescence and superior biocompatibility for targeted specific bioimaging using hyaluronate and glycine as precursors in a facile, simple bottom-up route. Although the mechanism of these HA-CQDs is still unknown, it appears that the nitrogen doping introduces a new internal structure. The two main types of interlayer lattice spaces in the HA-CQDs were similar to graphite and graphite oxide. We further demonstrated that HA-CQDs exhibited good biocompatibility without significant cytotoxicity when incubated with cancer cells. The internalized HA-CQDs were localized in the cytoplasm, particularly around the nucleus, and were used as fluorescent probes to target CD44 high-expression tumor cells. In brief, the HA-CQDs have great application in targeted specific live cell labeling and imaging.Acknowledgements
The research was supported by the National Natural Science Foundation of China (No. 81301316, 31200676), China Postdoctoral Science Foundation (2013M540425, 2014T70487, 2015M571705), Health and family planning commission of Jiangsu province scientific research subject (H201557), Natural Science Foundation of Jiangsu Province (BK20161317), Senior Talents Scientific Research Foundation of Jiangsu University (No. 13JDG022, 11JDG113).References
- Y. P. Sun, B. Zhou, Y. Lin, W. Wang, K. A. S. Fernando, P. Pathak, M. J. Meziani, B. A. Harruff, X. Wang, H. F. Wang, P. G. Luo, H. Yang, M. E. Kose, B. L. Chen, L. M. Veca and S. Y. Xie, J. Am. Chem. Soc., 2006, 128, 7756–7757 CrossRef CAS PubMed.
- L. Cao, X. Wang, M. J. Meziani, F. S. Lu, H. F. Wang, P. G. Luo, Y. Lin, B. A. Harruff, L. M. Veca, D. Murray, S. Y. Xie and Y. P. Sun, J. Am. Chem. Soc., 2007, 129, 11318–11319 CrossRef CAS PubMed.
- Y. Qu, W. Li, Y. L. Zhou, X. F. Liu, L. L. Zhang, L. M. Wang, Y. F. Li, A. Iida, Z. Y. Tang, Y. L. Zhao, Z. F. Chai and C. Y. Chen, Nano Lett., 2011, 11, 3174–3183 CrossRef CAS PubMed.
- L. Ye, K. T. Yong, L. W. Liu, I. Roy, R. Hu, J. Zhu, H. X. Cai, W. C. Law, J. W. Liu, K. Wang, J. Liu, Y. Q. Liu, Y. Z. Hu, X. H. Zhang, M. T. Swihart and P. N. Prasad, Nat. Nanotechnol., 2012, 7, 453–458 CrossRef CAS PubMed.
- C. Liu, P. Zhang, X. Y. Zhai, F. Tian, W. C. Li, J. H. Yang, Y. Liu, H. B. Wang, W. Wang and W. G. Liu, Biomaterials, 2012, 33, 3604–3613 CrossRef CAS PubMed.
- L. F. Tan, X. L. He, D. Chen, X. L. Wu, H. B. Li, X. L. Ren, X. W. Meng and F. Q. Tang, J. Biomed. Nanotechnol., 2013, 9, 53–60 CrossRef CAS PubMed.
- E. J. Goh, K. S. Kim, Y. R. Kim, H. S. Jung, S. Beack, W. H. Kong, G. Scarcelli, S. H. Yun and S. K. Hahn, Biomacromolecules, 2012, 13, 2554–2561 CrossRef CAS PubMed.
- M. Zhang, H. Ju and L. Zhang, Int. J. Nanomed., 2015, 10, 6943–6953 CAS.
- H. Zhu, X. L. Wang, Y. L. Li, Z. J. Wang, F. Yang and X. Yang, Chem. Commun., 2009, 34, 5118–5120 RSC.
- M. Zhang, H. Ju, L. Zhang, M. Sun, Z. Zhou, Z. Dai, L. Zhang, A. Gong, C. Wu and F. Du, Int. J. Nanomed., 2015, 10, 6943–6953 CAS.
- F. Du, M. Zhang, X. Li, J. Li, X. Jiang, Z. Li, Y. Hua, G. Shao, J. Jin, Q. Shao, M. Zhou and A. Gong, Nanotechnology, 2014, 25(31), 315702 CrossRef PubMed.
- F. Du, J. Yuan, M. Zhang, J. Li, Z. Zhou and L. Zhang, RSC Adv., 2014, 4(71), 37536–37541 RSC.
- F. Du, X. Jin, J. Chen, Y. Hua, M. Cao and L. Zhang, J. Nanopart. Res., 2014, 16, 2720 CrossRef.
- X. Michalet, F. F. Pinaud, L. A. Bentolila, J. M. Tsay, S. Doose, J. J. Li, G. Sundaresan, A. M. Wu, S. S. Gambhir and S. Weiss, Science, 2005, 307, 538–544 CrossRef CAS PubMed.
- S. T. Yang, L. Cao and P. Luo, J. Am. Chem. Soc., 2009, 131, 11308 CrossRef CAS PubMed.
- X. D. Yang, X. Yang, Z. Y. Li, S. Y. Li, Y. X. Han and Y. Chen, J. Colloid Interface Sci., 2015, 456, 1–6 CrossRef CAS PubMed.
- P. Miao, K. Han, Y. Tang, B. Wang, T. Lin and W. Cheng, Nanoscale, 2015, 7, 1586–1595 RSC.
- F. Y. Du, J. Yuan, M. M. Zhang, J. A. Li, Z. Zhou and Z. Li, RSC Adv., 2014, 4, 37536–37541 RSC.
- W. Li, Z. Zhang, B. Kong, S. Feng, J. Wang, L. Wang, J. Yang, F. Zhang, P. Wu and D. Zhao, Angew. Chem., Int. Ed. Engl., 2013, 52, 8151–8155 CrossRef CAS PubMed.
- J. C. Li, L. F. Zheng, H. D. Cai, M. W. Shen, G. X. Zhang and X. Y. Shi, Biomaterials, 2013, 34, 8382–8392 CrossRef CAS PubMed.
- H. Cai, X. An, J. Cui, S. Wen and K. Li, ACS Appl. Mater. Interfaces, 2013, 5, 1722–1731 CAS.
- J. C. Li, L. F. Zheng and X. Y. Shi, Biomaterials, 2013, 34, 8382–8392 CrossRef CAS PubMed.
- S. K. Bhunia, A. Saha, A. R. Maity, S. C. Ray and N. R. Jana, Sci. Rep., 2013, 3, 1473 Search PubMed.
- W. Wang, Y. C. Lu, H. Huang, J. J. Feng, J. R. Chen and A. J. Wang, Analyst, 2014, 139, 1692–1696 RSC.
- H. Huang, Y. C. Lu, A. J. Wang, J. H. Liu, J. R. Chen and J. J. Feng, RSC Adv., 2014, 4, 11872–11875 RSC.
- H. Li, F. Q. Shao, S. Y. Zou, Q. J. Yang, H. Huang, J. J. Feng and A. J. Wang, Microchim. Acta, 2016, 183, 821–826 CrossRef CAS.
- G. Y. Lee, J. H. Kim, K. Y. Choi, H. Y. Yoon, K. Kim and I. C. Kwon, Biomaterials, 2015, 53, 341–348 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2016 |