Theoretical insight into the binding affinity enhancement of serine with the uranyl ion through phosphorylation†
Qun-Yan Wua,
Cong-Zhi Wanga,
Jian-Hui Lana,
Zhi-Fang Chaiab and
Wei-Qun Shi*a
aLaboratory of Nuclear Energy Chemistry and Key Laboratory for Biomedical Effects of Nanomaterials and Nanosafety, Institute of High Energy Physics, Chinese Academy of Sciences, Beijing, 100049, China. E-mail: shiwq@ihep.ac.cn
bSchool of Radiological and Interdisciplinary Sciences (RAD-X), Collaborative Innovation Center of Radiation Medicine of Jiangsu Higher Education Institutions, Soochow University, Suzhou 215123, China
First published on 18th July 2016
Abstract
To study the coordination modes and the binding affinities of uranyl ions with serine and phosphoserine, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 and 1
2 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes of a uranyl ion with these ligands were optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in an aqueous solution. The analyses of the electronic energies show that the uranyl ion tends to adopt a penta-coordination mode and the binding affinity of the uranyl ion toward three functional groups follows the order of –PO42− > –COO− > –PO4H−. The changes of the Gibbs free energy (ΔGsol) for the studied reactions suggest that the uranyl ion prefers to form a 1
3 type complexes of a uranyl ion with these ligands were optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in an aqueous solution. The analyses of the electronic energies show that the uranyl ion tends to adopt a penta-coordination mode and the binding affinity of the uranyl ion toward three functional groups follows the order of –PO42− > –COO− > –PO4H−. The changes of the Gibbs free energy (ΔGsol) for the studied reactions suggest that the uranyl ion prefers to form a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complex with these ligands. Moreover, the absolute values of ΔGsol increase for the phosphorylated serine with the same type of reaction. Based on the thermodynamic results, a higher solution pH is more favorable for uranyl ion coordination with phosphoserine. This work could render theoretical insights into the specific coordination modes of uranyl ions with serine/phosphoserine under different conditions and provide useful information for further study on the interactions between actinide cations with peptides and proteins.
3 type complex with these ligands. Moreover, the absolute values of ΔGsol increase for the phosphorylated serine with the same type of reaction. Based on the thermodynamic results, a higher solution pH is more favorable for uranyl ion coordination with phosphoserine. This work could render theoretical insights into the specific coordination modes of uranyl ions with serine/phosphoserine under different conditions and provide useful information for further study on the interactions between actinide cations with peptides and proteins.
Introduction
With the growing human demand for nuclear energy, significant amounts of uranium have been occasionally released into the environment which could lead to serious problems in ecological environments and bio-organisms throughout the world due to its high chemical and radioactive toxicity.1 To better evaluate the risk to living organisms, it is of extreme importance to study the interaction mechanisms of the uranyl ion with various biomolecules, especially peptides and proteins, which can give useful information for understanding the trafficking, absorption and deposition pathways of uranium in vivo.2,3 Recently, there are increasing numbers of studies on the interactions of uranyl ion with peptides or proteins with specific binding motifs.4–13 These studies usually concerned the binding parameters, such as stoichiometry, stability constant and thermodynamic property of U(VI)–peptide/protein complexes. However, the coordination structures and bonding nature of uranium complexes in proteins have not been well understood due to the combined intricacy of metallobiochemistry and actinide coordination chemistry.14There are lots of peptides and proteins containing different species amino acids, which have suitable coordination sites and good binding affinity with uranyl ions. Serine is a hydroxyl chiral amino acid with high hydrophilicity and can be found in the outer regions of soluble proteins,15 which makes serine a good simple model to simulate peptide properties. Moreover, protein phosphorylation, one of the most frequent post-translational modifications (PTMs) of proteins, can occur at the hydroxyl side chain of serine, threonine and tyrosine.16–18 Phosphoserine accounts for about 90% of the phosphorylated residues in the human proteome.19 In contrast, phospho-threonine and tyrosine only account for about 10% and 0.05%, respectively.19 In addition, the phosphate or phosphonate group is a well-known efficient binding moiety for hard Lewis acids such as uranyl cation. Therefore, here we selected serine and phosphoserine as typical representatives to study the effect of phosphorylation on peptide coordination with the uranyl ion. Actually, some investigations on phosphopeptides which focused on mimicking the metal binding sites of proteins have been already reported in recent years.20–27 For instance, Vidaud and co-workers reported a phosphorylated hexapeptide which is one of the possible metal binding sites in osteopontin, and confirmed that uranyl ion can bind this motif through phosphate and carboxyl functional groups.22,24 Very recently, Delangle and co-workers reported that cyclic peptides with two phosphoserines and two glutamic acids could mimic high affinity binding sites for uranyl ion in proteins, and revealed that these cyclic peptides are efficient uranyl chelating agents due to the large contribution from the phosphorylated serine.25 It has been demonstrated that in most cases phosphate groups of proteins were directly involved in uranyl coordination. Nevertheless, the specific coordination pattern and bonding nature of the phosphorylated protein motif with the uranyl ion still need further clarifications.
Fortunately, quantum chemical calculation provides a useful approach on this issue. For instance, Hillier and co-workers investigated the coordination and reduction of the uranyl ion with Fe(II) containing proteins using density functional theory (DFT) calculations.28–30 Li et al. theoretically reported the coordination structures, thermodynamic stabilities and luminescence spectra of uranyl–glycine complexes in aqueous solution.31 Recently, in order to understand the binding feature of a super uranyl-binding protein (SUP), Odoh et al. theoretically examined the coordination structures of the uranyl ion with one engineered protein and found that uranyl ion is coordinated to five carboxylate oxygen atoms from four amino acid residues.11 Additionally, Dixon and co-workers studied the complexation of the uranyl ion with anionic phosphate ligands in the presence or absence of water molecules using DFT calculations.32 However, as far as we know, the interactions between serine/phosphoserine and the uranyl ion have not been theoretically addressed yet. Therefore, in this work, we have concentrated on the coordination structures and thermodynamic properties of the uranyl complexes with serine/phosphoserine by using quasi-relativistic DFT calculations, with emphasis on the coordination differences between carboxyl and phosphate groups.
Computational details
There have been many DFT studies on uranyl complexes. To test the effect of the different functionals and basis sets on the geometries and vibrational frequencies, we have investigated complex [UO2(pSerH2)(H2O)3]+-1 (Fig. 1) as representative applying with the generalized gradient approximation (GGA, BP86![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 33 and PBE34), hybrid GGA (B3LYP35,36 and B3LYP-D3
33 and PBE34), hybrid GGA (B3LYP35,36 and B3LYP-D3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 37,38) and hybrid meta GGA (M06
37,38) and hybrid meta GGA (M06![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 39) as well as 6-311+G(d) and 6-31+G(d) basis sets, respectively. The U–O bond distances and harmonic vibrational frequencies at the different level of theory are listed in Table S1 of the ESI,† it is found that the bond distances and vibrational frequencies obtained at the different level of theory have somewhat variations. According to the previous reports, B3LYP functional has been widely used for uranyl complexes,40–47 which can provide reliable geometries and energetics with experimental data. Therefore, the geometries of serine, phosphoserine and the corresponding uranyl complexes were optimized using the hybrid B3LYP functional. The geometrical optimizations and frequency calculations were performed with the quasi-relativistic small-core pseudo-potential ECP60MWB48–50 along with the corresponding ECP60MWB-SEG valence basis sets49,50 for uranium atom and the 6-311+G(d) basis sets for H, C, N, O and P atoms. The small-core pseudopotential replaces 60 core electrons for uranium atom, whereas the remaining 32 electrons were described by the associated valence basis set, which have been shown to provide reliable results for uranium complexes.51–53 In order to simulate the environmental condition, all the structures were optimized using the Solvation Model based on density (SMD model)54 in aqueous solution (H2O, ε = 78.36) with the Gaussian 09 program.55 SMD model can provide the reliable Gibbs free energy in the aqueous solution. Harmonic vibrational frequencies obtained at the optimized structures in the aqueous solution are all positive values. Moreover, to estimate the dispersion effects, we have optimized the structures of 1
39) as well as 6-311+G(d) and 6-31+G(d) basis sets, respectively. The U–O bond distances and harmonic vibrational frequencies at the different level of theory are listed in Table S1 of the ESI,† it is found that the bond distances and vibrational frequencies obtained at the different level of theory have somewhat variations. According to the previous reports, B3LYP functional has been widely used for uranyl complexes,40–47 which can provide reliable geometries and energetics with experimental data. Therefore, the geometries of serine, phosphoserine and the corresponding uranyl complexes were optimized using the hybrid B3LYP functional. The geometrical optimizations and frequency calculations were performed with the quasi-relativistic small-core pseudo-potential ECP60MWB48–50 along with the corresponding ECP60MWB-SEG valence basis sets49,50 for uranium atom and the 6-311+G(d) basis sets for H, C, N, O and P atoms. The small-core pseudopotential replaces 60 core electrons for uranium atom, whereas the remaining 32 electrons were described by the associated valence basis set, which have been shown to provide reliable results for uranium complexes.51–53 In order to simulate the environmental condition, all the structures were optimized using the Solvation Model based on density (SMD model)54 in aqueous solution (H2O, ε = 78.36) with the Gaussian 09 program.55 SMD model can provide the reliable Gibbs free energy in the aqueous solution. Harmonic vibrational frequencies obtained at the optimized structures in the aqueous solution are all positive values. Moreover, to estimate the dispersion effects, we have optimized the structures of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type uranyl complexes applying the B3LYP-D3
1 type uranyl complexes applying the B3LYP-D3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 37,38 functional at the same basis set in aqueous solution using SMD model. The spin–orbital coupling effect has been neglected for all calculations in this work. To evaluate the bonding nature of the U–O bonds for the equatorial plane of uranyl ion, the topological analyses of the electron density for the U–O bonds were performed by employing quantum theory of atoms in molecules (QTAIM) with Multiwfn program.56 In addition, the natural bond orbital (NBO) analysis of the complexes was also performed at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory.
37,38 functional at the same basis set in aqueous solution using SMD model. The spin–orbital coupling effect has been neglected for all calculations in this work. To evaluate the bonding nature of the U–O bonds for the equatorial plane of uranyl ion, the topological analyses of the electron density for the U–O bonds were performed by employing quantum theory of atoms in molecules (QTAIM) with Multiwfn program.56 In addition, the natural bond orbital (NBO) analysis of the complexes was also performed at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory.
Results and discussion
Structures of serine and phosphoserine
As it is known, amino acids appear as ionic species in aqueous solution which are dependent on the solution pH. Amino acids are prone to adopt an anionic form at high pH, whilst they appear predominantly as a cationic species at low pH and turn to zwitterionic ions at medium pH. According to the experimental results available, several possible species at different pH for serine and phosphoserine are presented in Scheme 1.57,58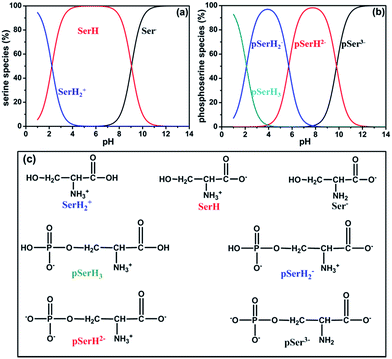 | ||
| Scheme 1 The species at different pH for serine (a) and phosphoserine (b) as well as their structures (c). | ||
Although we paid close attention to the interactions of the uranyl ion with serine and phosphoserine, the study about the most stable conformers of serine and phosphoserine is also an interesting topic. Hence, the structures of serine and phosphoserine at different pH were first investigated. Gronert and O'Hair have reported 51 conformers of serine within an energy gap of about 12 kcal mol−1 at the MP2/6-31+G(d)//HF/6-31G(d) level of theory.59 Herein, according to available experimental results,58 the conformers of the protonated (cation, SerH2+), zwitterionic (neutral, SerH) and deprotonated (anion, Ser−) serine have been optimized at the B3LYP/6-311+G(d) level of theory. Here, we selected six most stable conformations for each species as shown in Fig. S1, ESI.† In addition, the relative energies to the most stable conformer for each species are provided in Fig. S1.† Obviously, six conformers of the zwitterionic serine have similar energies, the maximum energy gap between them is only 0.41 kcal mol−1. However, for the protonated serine, the conformers with the cis carboxyl group are about 2.0 kcal mol−1 higher in energy than those with the trans one due to the intramolecular hydrogen bonding of the latter. The impact of phosphorylation on protein functions is related to the protein conformational changes in many cases. Basically, compared to serine, phosphoserine has four species (pSer3−, pSerH2−, pSerH2− and pSerH3), depending on the pH-induced protonation state of the phosphate, carboxyl and amine groups, as shown in Scheme 1.57 Tekely et al. revealed the successive steps of deprotonation of phosphoserine by solid-state nuclear magnetic resonance (NMR) spectroscopy and ab initio calculations.57 Gaigeot and co-workers investigated vibrational properties of deprotonated phosphoserine (pSerH2−) at room temperature using DFT-based Car–Parrinello molecular dynamics simulations.17 They obtained eight conformers of pSerH2− and classified them into three groups according to the different deprotonated moieties. Although there are a few theoretical studies on the conformers of phosphoserine,16–18,57,60 only limited types of phosphoserine, such as deprotonated mono-anionic phosphoserine has been considered. In this work, the four possible species of phosphoserine have been optimized in the aqueous solution at the B3LYP/6-311+G(d) level of theory. The six most stable conformers for each species of phosphoserine and the corresponding relative energies are provided in Fig. S2.† It is conspicuous that the six conformers of tri-anionic phosphoserine (pSer3−) are relatively close in energy with the maximum energy gap of 1.67 kcal mol−1. While the energy gaps between the conformers for other three species (pSerH2−, pSerH2− and pSerH3) are relatively larger due to the deprotonated carboxyl and phosphate groups. For instance, the maximum energy gap for the pSerH2− species is 7.15 kcal mol−1. In this work, the uranyl complexes studied consist of zwitterionic serine and two phosphorylated serine molecules, pSerH2− and pSerH2−, which are the predominant species at medium pH.
Complexes of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) 1 uranyl ion and serine/phosphoserine
1 uranyl ion and serine/phosphoserine
The structures of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
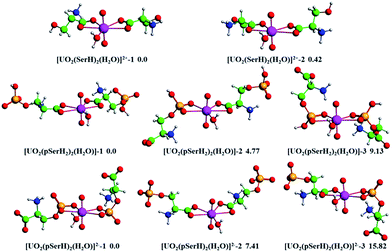
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type complexes of uranyl ion with serine/phosphoserine were optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution. For simplification, we just selected the most stable conformers of each type of uranyl complex as representative cases studied. As shown in Fig. 1, for all the complex structures, the uranyl ion is in penta-coordination mode with one serine/phosphoserine and three water molecules involved as bi- and mono-dentate ligands, respectively. Additionally, as for the two phosphoserine complexes, [UO2(pSerH2)(H2O)3]+ and [UO2(pSerH)(H2O)3], the carboxyl and phosphate groups binding with the uranyl ion were also considered, respectively.
1 type complexes of uranyl ion with serine/phosphoserine were optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution. For simplification, we just selected the most stable conformers of each type of uranyl complex as representative cases studied. As shown in Fig. 1, for all the complex structures, the uranyl ion is in penta-coordination mode with one serine/phosphoserine and three water molecules involved as bi- and mono-dentate ligands, respectively. Additionally, as for the two phosphoserine complexes, [UO2(pSerH2)(H2O)3]+ and [UO2(pSerH)(H2O)3], the carboxyl and phosphate groups binding with the uranyl ion were also considered, respectively.
In order to estimate the dispersion effects on the geometries and energies, the optimizations of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type uranyl complexes have been carried using the B3LYP and B3LYP-D3 functionals, respectively. The bond distances, bond angles and harmonic vibrational frequencies for the 1
1 type uranyl complexes have been carried using the B3LYP and B3LYP-D3 functionals, respectively. The bond distances, bond angles and harmonic vibrational frequencies for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type complexes with both the functionals are provided in Table 1. It is found that the U–O bond distances and harmonic vibrational frequencies are almost same for the same complex using with both the B3LYP and B3LYP-D3 functionals. It is clearly seen from Fig. 1 that the dispersion corrections have a certain effect on the relative energies of the complexes studied here due to non-bonding interactions. Moreover, considering the dispersion correction, the relative energies for mono-anionic phosphoserine complexes decrease, while those for dianionic phosphoserine complexes increase. However, the relative energies obtained using with B3LYP and B3LYP-D3 functionals keep similar trends. Therefore, the geometrical parameters and the energies obtained at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory are used below.
1 type complexes with both the functionals are provided in Table 1. It is found that the U–O bond distances and harmonic vibrational frequencies are almost same for the same complex using with both the B3LYP and B3LYP-D3 functionals. It is clearly seen from Fig. 1 that the dispersion corrections have a certain effect on the relative energies of the complexes studied here due to non-bonding interactions. Moreover, considering the dispersion correction, the relative energies for mono-anionic phosphoserine complexes decrease, while those for dianionic phosphoserine complexes increase. However, the relative energies obtained using with B3LYP and B3LYP-D3 functionals keep similar trends. Therefore, the geometrical parameters and the energies obtained at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory are used below.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles (degree) and the UO22+ harmonic vibrational frequencies (νs and νas, cm−1) for the 1
O angles (degree) and the UO22+ harmonic vibrational frequencies (νs and νas, cm−1) for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type complexes optimized using B3LYP and B3LYP-D3 methods with 6-311+G(d) basis set in the aqueous solution
1 type complexes optimized using B3LYP and B3LYP-D3 methods with 6-311+G(d) basis set in the aqueous solution
| Complexes | Modesa | Methods | U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O O |
U–OCb | U–OPb | U–OWb | ∠O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O O |
νs | νas |
|---|---|---|---|---|---|---|---|---|---|
| a The bi-dentate carboxyl (Cbi) and phosphate (Pbi) groups.b The OC, OP and OW denote the oxygen atom of the carboxyl and phosphate groups and water molecule, respectively. | |||||||||
| [UO2(SerH)(H2O)3]2+-1 | 1Cbi3W | B3LYP | 1.768 | 2.571 | — | 2.498 | 178.64 | 868 | 910 |
| B3LYP-D3 | 1.768 | 2.551 | 2.498 | 178.66 | 870 | 911 | |||
| [UO2(SerH)(H2O)3]2+-2 | 1Cbi3W | B3LYP | 1.768 | 2.567 | — | 2.494 | 178.79 | 858 | 902 |
| B3LYP-D3 | 1.767 | 2.556 | 2.493 | 178.74 | 859 | 904 | |||
| [UO2(pSerH2)(H2O)3]+-1 | 1Cbi3W | B3LYP | 1.770 | 2.561 | — | 2.490 | 178.75 | 865 | 906 |
| B3LYP-D3 | 1.769 | 2.544 | 2.491 | 178.74 | 867 | 908 | |||
| [UO2(pSerH2)(H2O)3]+-2 | 1Pbi3W | B3LYP | 1.771 | — | 2.543 | 2.505 | 177.14 | 855 | 897 |
| B3LYP-D3 | 1.771 | 2.522 | 2.501 | 176.45 | 852 | 892 | |||
| [UO2(pSerH)(H2O)3]-1 | 1Pbi3W | B3LYP | 1.778 | — | 2.456 | 2.525 | 176.01 | 847 | 885 |
| B3LYP-D3 | 1.777 | 2.446 | 2.523 | 176.08 | 848 | 886 | |||
| [UO2(pSerH)(H2O)3]-2 | 1Cbi3W | B3LYP | 1.769 | 2.573 | — | 2.484 | 179.01 | 867 | 907 |
| B3LYP-D3 | 1.769 | 2.546 | 2.486 | 179.12 | 869 | 909 | |||
The relative energies of the two most stable conformers for each type of uranyl complex are provided in Fig. 1. It is clearly seen that the two serine conformers, [UO2(SerH)(H2O)3]2+, have small energy gap with the value of 1.06 kcal mol−1 due to the rotation of the hydroxyl group. However, the relative energies of different conformers of the uranyl/phosphoserine complexes have obvious disparity with respect to the coordination of carboxyl or phosphate groups. For instance, the conformers [UO2(pSerH2)(H2O)3]+-2 with a bidentate phosphate group are 3.33 kcal mol−1 higher in energy compared to conformer [UO2(pSerH2)(H2O)3]+-1 with a bidentate carboxyl group. In particular, the energy of conformer [UO2(pSerH)(H2O)3]-2 with a bidentate carboxyl group is 8.89 kcal mol−1 higher than that of conformer [UO2(pSerH)(H2O)3]-1 with a bidentate phosphate groups. These results suggest that the anionic number of phosphoserine has obvious effect on the binding affinity of the uranyl ion toward phosphoserine. Based on the relative energies of the two types of phosphoserine complexes, it can be concluded that the uranyl ion like to bind with the carboxyl group (–COO−) rather than the –PO4H− group for the mono-anionic phosphoserine complexes [UO2(pSerH2)(H2O)3]+, whilst the uranyl ion prefers to bind with –PO42− group rather than –COO− for the dianionic phosphoserine complexes [UO2(pSerH)(H2O)3] due to the electrostatic interactions. Therefore, the uranyl ion binds toward the three groups mentioned above follows the order of –PO42− > –COO− > –PO4H−.
The selected U–O bond distances and O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond angles for the uranyl–serine/phosphoserine complexes with a chemical stoichiometry of 1
O bond angles for the uranyl–serine/phosphoserine complexes with a chemical stoichiometry of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 are listed in Table 1. It is obvious that phosphorylation makes the U
1 are listed in Table 1. It is obvious that phosphorylation makes the U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond distances slightly longer. The distances (U–OC) between the uranium atom and the oxygen atom of the carboxyl group are almost comparable for all the 1
O bond distances slightly longer. The distances (U–OC) between the uranium atom and the oxygen atom of the carboxyl group are almost comparable for all the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 complexes, in the range of 2.56–2.57 Å. For the two types of phosphoserine complexes, [UO2(pSerH2)(H2O)3]+ and [UO2(pSerH)(H2O)3], the U–OP bonds between the uranium atom and oxygen atom of the phosphate group are shorter compared to the U–OC bonds in length. For instance, the U–OP bond distances in conformer [UO2(pSerH)(H2O)3]-1 is 2.456 Å, which is 0.117 Å shorter than that of the U–OC bond in conformer [UO2(pSerH)(H2O)3]-2. On the contrary, the U–OW distances between the uranium atom and oxygen atom of water molecule in the conformers with a bidentate phosphate group are longer than those with a bidentate carboxyl group. For instance, the U–OW bond distances are about 2.505 Å in conformer [UO2(pSerH2)(H2O)3]+-2, whilst the corresponding values decrease to 2.490 Å in conformer [UO2(pSerH2)(H2O)3]+-1. Moreover, compared to the two types of phosphoserine complexes with a bidentate phosphate group, the U–OP bond distances in conformer [UO2(pSerH2)(H2O)3]+-1 (2.543 Å) are somewhat longer than those in [UO2(pSerH)(H2O)3]-1 (2.456 Å), while the trends of the corresponding U–OW bond distances are converse. Besides, the O
1 complexes, in the range of 2.56–2.57 Å. For the two types of phosphoserine complexes, [UO2(pSerH2)(H2O)3]+ and [UO2(pSerH)(H2O)3], the U–OP bonds between the uranium atom and oxygen atom of the phosphate group are shorter compared to the U–OC bonds in length. For instance, the U–OP bond distances in conformer [UO2(pSerH)(H2O)3]-1 is 2.456 Å, which is 0.117 Å shorter than that of the U–OC bond in conformer [UO2(pSerH)(H2O)3]-2. On the contrary, the U–OW distances between the uranium atom and oxygen atom of water molecule in the conformers with a bidentate phosphate group are longer than those with a bidentate carboxyl group. For instance, the U–OW bond distances are about 2.505 Å in conformer [UO2(pSerH2)(H2O)3]+-2, whilst the corresponding values decrease to 2.490 Å in conformer [UO2(pSerH2)(H2O)3]+-1. Moreover, compared to the two types of phosphoserine complexes with a bidentate phosphate group, the U–OP bond distances in conformer [UO2(pSerH2)(H2O)3]+-1 (2.543 Å) are somewhat longer than those in [UO2(pSerH)(H2O)3]-1 (2.456 Å), while the trends of the corresponding U–OW bond distances are converse. Besides, the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles are nearly close to 180° for all the 1
O angles are nearly close to 180° for all the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 complexes with the maximum deviation of ∼4° in conformer [UO2(pSerH)(H2O)3]-1. Moreover, the O
1 complexes with the maximum deviation of ∼4° in conformer [UO2(pSerH)(H2O)3]-1. Moreover, the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles are larger in the conformers with a bidentate carboxyl group compared to those with a bidentate phosphate group.
O angles are larger in the conformers with a bidentate carboxyl group compared to those with a bidentate phosphate group.
To study the bonding nature of the U–O bonds, we also perform the QTAIM analysis using Multiwfn program.56 QTAIM has been used for evaluating the bonding properties of actinide compounds and could provide valuable information about the nature of chemical bonds.61–63 Based on the electron density (ρ(r)) and its Laplacian (∇2ρ(r)) of the bond critical point (BCP), the bonding interactions can be classified into the covalent or ionic character. Generally speaking, ρ(r) > 0.20 a.u. and ∇2ρ(r) < 0 is a covalent bond, whereas ρ(r) < 0.10 a.u. and ∇2ρ(r) > 0 show an ionic bond. As shown in Table 2, the ρ(r) values at U–O BCP are about 0.05 a.u. and the corresponding values of the Laplacian are all positive, which suggests that the U–O bonds show ionic character. Moreover, the ρ(r) value at the U–OP BCP of [UO2(pSerH)(H2O)3]-1 is larger than those of U–O BCP for other 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type uranyl complexes. This result is consistent with the analysis of the relative energies of uranyl complexes. Wiberg bond indices (WBIs) are related the bond strengths of the similar bonding situation. All the WBIs values of the U–O bonds are about 0.4–0.5 as provided in Table 2 which also suggest that the U–O bonds have ionic characters.
1 type uranyl complexes. This result is consistent with the analysis of the relative energies of uranyl complexes. Wiberg bond indices (WBIs) are related the bond strengths of the similar bonding situation. All the WBIs values of the U–O bonds are about 0.4–0.5 as provided in Table 2 which also suggest that the U–O bonds have ionic characters.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type complexes at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theorya
1 type complexes at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theorya
| Complexes | U–OC | U–OP | U–OW | ||||||
|---|---|---|---|---|---|---|---|---|---|
| ρ | ∇2ρ | WBIs | ρ | ∇2ρ | WBIs | ρ | ∇2ρ | WBIs | |
| a The OC, OP and OW denote the oxygen atom of the carboxyl and phosphate groups and water molecule, respectively. | |||||||||
| [UO2(SerH)(H2O)3]2+-1 | 0.046 | 0.149 | 0.361 | — | — | — | 0.050 | 0.188 | 0.384 |
| [UO2(SerH)(H2O)3]2+-2 | 0.046 | 0.150 | 0.355 | — | — | — | 0.050 | 0.190 | 0.387 |
| [UO2(pSerH2)(H2O)3]+-1 | 0.047 | 0.153 | 0.366 | — | — | — | 0.051 | 0.192 | 0.390 |
| [UO2(pSerH2)(H2O)3]+-2 | — | — | — | 0.049 | 0.161 | 0.432 | 0.052 | 0.205 | 0.386 |
| [UO2(pSerH)(H2O)3]-1 | — | — | — | 0.060 | 0.196 | 0.519 | 0.048 | 0.177 | 0.377 |
| [UO2(pSerH)(H2O)3]-2 | 0.046 | 0.147 | 0.364 | — | — | — | 0.051 | 0.195 | 0.393 |
The harmonic vibrational frequencies corresponding to the symmetric (νs) and asymmetric (νas) stretching of the UO22+ for all the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 complexes in the aqueous solution are also listed in Table 1. It is obvious that the νs and νas for all complexes are red-shifted compared to the uranyl hydrate [UO2(H2O)5]2+ (872 and 914 cm−1). The red-shift extent of νs and νas in the conformers depends on the coordinated functional group with uranyl ion. For instance, the scopes of νs and νas for the conformers with a bidentate phosphate group are 847–855 cm−1 and 885–897 cm−1, respectively, whereas the corresponding values are 858–868 cm−1 and 902–910 cm−1 for the conformers with a bidentate carboxyl group. Moreover, compared to the two types of phosphoserine complexes, the values of νs and νas are larger in conformer [UO2(pSerH2)(H2O)3]+-2 than that of [UO2(pSerH)(H2O)3]-1. These harmonic frequencies could also provide the quantitative pictures for the experimental observation.
1 complexes in the aqueous solution are also listed in Table 1. It is obvious that the νs and νas for all complexes are red-shifted compared to the uranyl hydrate [UO2(H2O)5]2+ (872 and 914 cm−1). The red-shift extent of νs and νas in the conformers depends on the coordinated functional group with uranyl ion. For instance, the scopes of νs and νas for the conformers with a bidentate phosphate group are 847–855 cm−1 and 885–897 cm−1, respectively, whereas the corresponding values are 858–868 cm−1 and 902–910 cm−1 for the conformers with a bidentate carboxyl group. Moreover, compared to the two types of phosphoserine complexes, the values of νs and νas are larger in conformer [UO2(pSerH2)(H2O)3]+-2 than that of [UO2(pSerH)(H2O)3]-1. These harmonic frequencies could also provide the quantitative pictures for the experimental observation.
Complexes of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) 2 uranyl ion and serine/phosphoserine
2 uranyl ion and serine/phosphoserine
The complexes with a chemical stoichiometry of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 (metal/ligand) were optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution as provided in Fig. 2. It is clearly noticed that the two conformers of [UO2(SerH)2(H2O)]2+ complex are very close in energy with an energy gap of 0.42 kcal mol−1, indicating that the different conformer of serine has little effect on the stability of the uranyl–serine complexes. However, for the two types of phosphoserine complexes, the relative energies have obvious difference due to the different coordinated functional group. The mono-anionic phosphoserine conformer [UO2(pSerH2)2(H2O)]-1 has lower energies compared to conformer [UO2(pSerH2)2(H2O)]-2. Furthermore, conformer [UO2(pSerH2)2(H2O)]-3 with two bidentate phosphate groups has the highest energies with the relative energy of 9.13 kcal mol−1. According to the relative energies, the binding affinity of the –COO− group with the uranyl ion is stronger compared to the –PO4H− group. Unlike the mono-phosphoserine complexes, the dianionic phosphoserine complexes with two bidentate phosphate groups have the lowest energies (Fig. 2). While conformer [UO2(pSerH)2(H2O)]2−-3 with two bidentate carboxyl groups has the highest energies. Hence, the binding affinity of the –PO42− group with uranyl ion is stronger compared to the –COO− group. According to the relative energies in Fig. 2, the trend of the binding affinity of the three functional groups with the uranyl ion follows the order of: –PO42− > –COO− > –PO4H−, which is in good agreement with the result of the 1
2 (metal/ligand) were optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution as provided in Fig. 2. It is clearly noticed that the two conformers of [UO2(SerH)2(H2O)]2+ complex are very close in energy with an energy gap of 0.42 kcal mol−1, indicating that the different conformer of serine has little effect on the stability of the uranyl–serine complexes. However, for the two types of phosphoserine complexes, the relative energies have obvious difference due to the different coordinated functional group. The mono-anionic phosphoserine conformer [UO2(pSerH2)2(H2O)]-1 has lower energies compared to conformer [UO2(pSerH2)2(H2O)]-2. Furthermore, conformer [UO2(pSerH2)2(H2O)]-3 with two bidentate phosphate groups has the highest energies with the relative energy of 9.13 kcal mol−1. According to the relative energies, the binding affinity of the –COO− group with the uranyl ion is stronger compared to the –PO4H− group. Unlike the mono-phosphoserine complexes, the dianionic phosphoserine complexes with two bidentate phosphate groups have the lowest energies (Fig. 2). While conformer [UO2(pSerH)2(H2O)]2−-3 with two bidentate carboxyl groups has the highest energies. Hence, the binding affinity of the –PO42− group with uranyl ion is stronger compared to the –COO− group. According to the relative energies in Fig. 2, the trend of the binding affinity of the three functional groups with the uranyl ion follows the order of: –PO42− > –COO− > –PO4H−, which is in good agreement with the result of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 type uranyl complexes. Thus, it is deductive that the charge of the coordinated functional groups rather than the total charge of the whole ligand plays an important role in the studied complexes.
1 type uranyl complexes. Thus, it is deductive that the charge of the coordinated functional groups rather than the total charge of the whole ligand plays an important role in the studied complexes.
The selected U–O bond lengths and O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond angles for 1
O bond angles for 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 type complexes are provided in Table 3. The U
2 type complexes are provided in Table 3. The U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond distances in all the conformers with two bidentate carboxyl groups are about 1.77 Å. However, it should be mentioned that the U
O bond distances in all the conformers with two bidentate carboxyl groups are about 1.77 Å. However, it should be mentioned that the U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond distances in the complexes [UO2(pSerH)2(H2O)]2− with two bidentate phosphate groups are about 0.02 Å longer than those in the complexes [UO2(pSerH)2(H2O)]2− with two bidentate carboxyl groups. This result indicates that the –PO42− group has a stronger binding affinity, which weakens the U
O bond distances in the complexes [UO2(pSerH)2(H2O)]2− with two bidentate phosphate groups are about 0.02 Å longer than those in the complexes [UO2(pSerH)2(H2O)]2− with two bidentate carboxyl groups. This result indicates that the –PO42− group has a stronger binding affinity, which weakens the U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond strength. The U–OC bond distances in all the complexes with two bidentate carboxyl groups are in the range of 2.54–2.55 Å. The U–OP bond distances have obvious difference between the mono- and di-anionic phosphoserine complexes. It is important to highlight that the distances of U–OC bonds are shorter than those of the U–OP bonds in the complexes [UO2(pSerH2)2(H2O)] with a bidentate carboxyl and phosphate group. In addition, the U–OW bond distances are somewhat different in the three types of 1
O bond strength. The U–OC bond distances in all the complexes with two bidentate carboxyl groups are in the range of 2.54–2.55 Å. The U–OP bond distances have obvious difference between the mono- and di-anionic phosphoserine complexes. It is important to highlight that the distances of U–OC bonds are shorter than those of the U–OP bonds in the complexes [UO2(pSerH2)2(H2O)] with a bidentate carboxyl and phosphate group. In addition, the U–OW bond distances are somewhat different in the three types of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 uranyl complexes. Like the 1
2 uranyl complexes. Like the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 complexes, in the 1
1 complexes, in the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 complexes, the O
2 complexes, the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles are close to 180° with the maximum deviation of about 3° in the conformer [UO2(pSerH)2(H2O)]2−-2. Besides, the O
O angles are close to 180° with the maximum deviation of about 3° in the conformer [UO2(pSerH)2(H2O)]2−-2. Besides, the O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles are larger in the complexes with a bidentate carboxyl group compared to those with a bidentate phosphate group. In addition, the values of ρ(r) are about 0.05 and the corresponding Laplacian are positive for the 1
O angles are larger in the complexes with a bidentate carboxyl group compared to those with a bidentate phosphate group. In addition, the values of ρ(r) are about 0.05 and the corresponding Laplacian are positive for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 type complexes (Table S2, ESI†), which suggest that their U–O bonds have ionic character. The values of ρ(r) at U–OP BCP in the complexes with two bidentate phosphate groups are the largest among all at the U–O bonds. And the WBIs of U–O bonds also support the results.
2 type complexes (Table S2, ESI†), which suggest that their U–O bonds have ionic character. The values of ρ(r) at U–OP BCP in the complexes with two bidentate phosphate groups are the largest among all at the U–O bonds. And the WBIs of U–O bonds also support the results.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U
U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O angles (degree) and the UO22+ harmonic vibrational frequencies (νs and νas, cm−1) for the 1
O angles (degree) and the UO22+ harmonic vibrational frequencies (νs and νas, cm−1) for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 type complexes optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution
2 type complexes optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution
| Complexes | Modesa | U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O O |
U–OCb | U–OPb | U–OWb | ∠O![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) U U![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O O |
νs | νas |
|---|---|---|---|---|---|---|---|---|
| a The bi-dentate carboxyl (Cbi) and phosphate (Pbi) group.b The OC, OP and OW denote the oxygen atom of the carboxyl and phosphate groups and water molecule, respectively. | ||||||||
| [UO2(SerH)2(H2O)]2+-1 | 2Cbi1W | 1.771 | 2.547 | — | 2.470 | 178.87 | 853 | 897 |
| [UO2(SerH)2(H2O)]2+-2 | 2Cbi1W | 1.771 | 2.550 | — | 2.453 | 178.48 | 856 | 902 |
| [UO2(pSerH2)2(H2O)]-1 | 2Cbi1W | 1.771 | 2.540 | — | 2.468 | 179.43 | 859 | 900 |
| [UO2(pSerH2)2(H2O)]-2 | 1Cbi1Pbi1W | 1.768 | 2.549 | 2.588 | 2.476 | 178.02 | 861 | 899 |
| [UO2(pSerH2)2(H2O)]-3 | 2Pbi1W | 1.769 | — | 2.572 | 2.451 | 177.18 | 852 | 895 |
| [UO2(pSerH)2(H2O)]2−-1 | 2Pbi1W | 1.789 | — | 2.469 | 2.509 | 177.28 | 818 | 858 |
| [UO2(pSerH)2(H2O)]2−-2 | 1Cbi1Pbi1W | 1.779 | 2.575 | 2.449 | 2.495 | 177.09 | 842 | 882 |
| [UO2(pSerH)2(H2O)]2−-3 | 2Cbi1W | 1.771 | 2.548 | — | 2.480 | 178.80 | 856 | 897 |
The νs and νas harmonic vibrational frequencies of the UO22+ for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 type complexes are obtained in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory, as listed in Table 2. The values of νs and νas for the complexes with two bidentate carboxyl groups are in the range of 853–859 cm−1 and 897–902 cm−1, respectively. It is worthy to notice that the νs and νas values for the complexes [UO2(pSerH)2(H2O)]2− with two bidentate phosphate groups are the smallest among the 1
2 type complexes are obtained in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory, as listed in Table 2. The values of νs and νas for the complexes with two bidentate carboxyl groups are in the range of 853–859 cm−1 and 897–902 cm−1, respectively. It is worthy to notice that the νs and νas values for the complexes [UO2(pSerH)2(H2O)]2− with two bidentate phosphate groups are the smallest among the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 complexes, with the red-shifted values of 54 and 56 cm−1, respectively, compared to the uranyl hydrate [UO2(H2O)5]2+.
2 complexes, with the red-shifted values of 54 and 56 cm−1, respectively, compared to the uranyl hydrate [UO2(H2O)5]2+.
Complexes of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) 3 uranyl ion and serine/phosphoserine
3 uranyl ion and serine/phosphoserine
The structures of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type uranyl complexes were also optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution. Here, we considered two types of complexes with uranyl equatorial coordination numbers of five and six, respectively. One species contains the two bidentate ligands and a mono-dentate ligand, the other has the three ligands as a bidentate fashion. Like the 1
3 type uranyl complexes were also optimized at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory in the aqueous solution. Here, we considered two types of complexes with uranyl equatorial coordination numbers of five and six, respectively. One species contains the two bidentate ligands and a mono-dentate ligand, the other has the three ligands as a bidentate fashion. Like the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 1
1 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 type serine complexes, the energy gaps of the two conformers between penta- and hexa-coordinated is only 0.79 kcal mol−1 as shown in Fig. S3, ESI.†
2 type serine complexes, the energy gaps of the two conformers between penta- and hexa-coordinated is only 0.79 kcal mol−1 as shown in Fig. S3, ESI.†
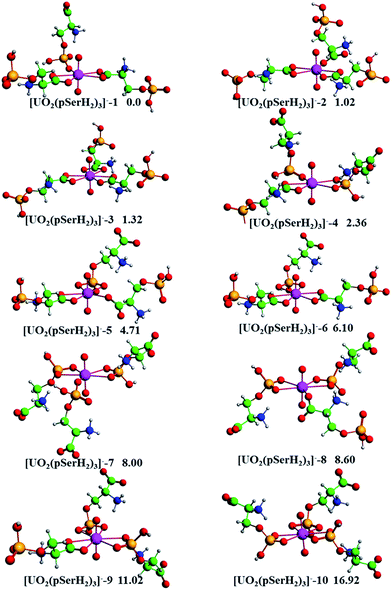
The ten conformers of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complex [UO2(pSerH2)3]− and their relative energies are provided in Fig. 3. It can be clearly seen that the most stable structure contains two bidentate carboxyl and one mono-dentate phosphate groups. According to the coordination modes and the relative energies, we can qualitatively obtain that the binding affinity of the mono-dentate –PO4H− group toward uranyl ion is stronger compared to the mono-dentate –COO− group. In contrast, the binding affinity of bidentate –PO4H− group toward the uranyl ion is weaker than that of bidentate –COO− group. Obviously, the trend of the relative energies of four hexa-coordinated conformers [UO2(pSerH2)3]− follows the order of: [UO2(pSerH2)3]−-3 with three carboxyl groups < [UO2(pSerH2)3]−-6 with two carboxyl and one phosphate groups < [UO2(pSerH2)3]−-9 with one carboxyl and two phosphate groups < [UO2(pSerH2)3]−-10 with three phosphate groups. It is worthy to notice that the penta-coordinated conformers have lower energies compared to the hexa-coordinated ones. These results reveal that the uranyl ion tends to adopt a penta-coordination mode in the complexes [UO2(pSerH2)3]−, that is, the carboxyl and phosphate groups prefer to act as mono-dentate ligand in [UO2(pSerH2)3]− when the equatorial coordination number of uranyl ion gets to five.
3 type complex [UO2(pSerH2)3]− and their relative energies are provided in Fig. 3. It can be clearly seen that the most stable structure contains two bidentate carboxyl and one mono-dentate phosphate groups. According to the coordination modes and the relative energies, we can qualitatively obtain that the binding affinity of the mono-dentate –PO4H− group toward uranyl ion is stronger compared to the mono-dentate –COO− group. In contrast, the binding affinity of bidentate –PO4H− group toward the uranyl ion is weaker than that of bidentate –COO− group. Obviously, the trend of the relative energies of four hexa-coordinated conformers [UO2(pSerH2)3]− follows the order of: [UO2(pSerH2)3]−-3 with three carboxyl groups < [UO2(pSerH2)3]−-6 with two carboxyl and one phosphate groups < [UO2(pSerH2)3]−-9 with one carboxyl and two phosphate groups < [UO2(pSerH2)3]−-10 with three phosphate groups. It is worthy to notice that the penta-coordinated conformers have lower energies compared to the hexa-coordinated ones. These results reveal that the uranyl ion tends to adopt a penta-coordination mode in the complexes [UO2(pSerH2)3]−, that is, the carboxyl and phosphate groups prefer to act as mono-dentate ligand in [UO2(pSerH2)3]− when the equatorial coordination number of uranyl ion gets to five.
The ten conformers of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type dianionic phosphoserine complexes, [UO2(pSerH)3]4−, and the corresponding relative energies are shown in Fig. S4, ESI.† Unlike the mono-anionic phosphoserine complexes, the most stable structure of the complexes [UO2(pSerH)3]4− has three ligands acted as two bidentate and one mono-dentate phosphate groups (–PO42−). Similar to the complexes [UO2(pSerH2)3]−, in the complexes [UO2(pSerH)3]4−, the binding affinity of the mono-dentate –PO42− group toward uranyl ion is stronger compared to the mono-dentate –COO− group. The binding affinity of the bidentate phosphate group toward uranyl ion is also higher than that of the bidentate carboxyl group, which shows a converse trend to that in the complexes [UO2(pSerH2)3]−. Besides, the uranyl ions also tend to adopt a penta-coordination mode rather than hexa-coordination in the complexes [UO2(pSerH2)3]−.
3 type dianionic phosphoserine complexes, [UO2(pSerH)3]4−, and the corresponding relative energies are shown in Fig. S4, ESI.† Unlike the mono-anionic phosphoserine complexes, the most stable structure of the complexes [UO2(pSerH)3]4− has three ligands acted as two bidentate and one mono-dentate phosphate groups (–PO42−). Similar to the complexes [UO2(pSerH2)3]−, in the complexes [UO2(pSerH)3]4−, the binding affinity of the mono-dentate –PO42− group toward uranyl ion is stronger compared to the mono-dentate –COO− group. The binding affinity of the bidentate phosphate group toward uranyl ion is also higher than that of the bidentate carboxyl group, which shows a converse trend to that in the complexes [UO2(pSerH2)3]−. Besides, the uranyl ions also tend to adopt a penta-coordination mode rather than hexa-coordination in the complexes [UO2(pSerH2)3]−.
The trends of U–O bond lengths for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes (Table S3, ESI†) are similar to those for the 1
3 type complexes (Table S3, ESI†) are similar to those for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 1
1 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 type complexes. The detailed discussions were provided in ESI.† The νs and νas harmonic vibrational frequencies of the UO22+ for the 1
2 type complexes. The detailed discussions were provided in ESI.† The νs and νas harmonic vibrational frequencies of the UO22+ for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes are predicted at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory in the aqueous solution, as provided in Table S3.† The scopes of the νs and νas values are small for the 1
3 type complexes are predicted at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory in the aqueous solution, as provided in Table S3.† The scopes of the νs and νas values are small for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type mono-anionic phosphoserine complexes (849–856 cm−1 and 890–898 cm−1, respectively). However, there are significant differences of the νs and νas values for the 1
3 type mono-anionic phosphoserine complexes (849–856 cm−1 and 890–898 cm−1, respectively). However, there are significant differences of the νs and νas values for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type dianionic phosphoserine complexes. Besides, compared to the uranyl hydrate [UO2(H2O)5]2+, the maximum red-shifted value is 72 and 69 cm−1 for νs and νas in the most stable conformer [UO2(pSerH)3]4−-1, respectively.
3 type dianionic phosphoserine complexes. Besides, compared to the uranyl hydrate [UO2(H2O)5]2+, the maximum red-shifted value is 72 and 69 cm−1 for νs and νas in the most stable conformer [UO2(pSerH)3]4−-1, respectively.
In addition, we have considered the structures of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 and 1
4 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 type uranyl–serine complexes optimized in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory, as provided in Fig. S5 and Table S4, ESI.† However, the absolute changes of the Gibbs free energy (ΔGsol) of forming the 1
5 type uranyl–serine complexes optimized in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory, as provided in Fig. S5 and Table S4, ESI.† However, the absolute changes of the Gibbs free energy (ΔGsol) of forming the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 and 1
4 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 type uranyl–serine complexes gradually decrease compared to that of the 1
5 type uranyl–serine complexes gradually decrease compared to that of the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complex as discussed the section “Energies” below. Therefore, we only discussed the 1
3 type complex as discussed the section “Energies” below. Therefore, we only discussed the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 and 1
2 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type uranyl complexes in this work.
3 type uranyl complexes in this work.
Energies
To explore the thermodynamic properties of the uranyl ion with serine/phosphoserine, ΔGsol of the related reactions are obtained in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d)/SMD level of theory, as listed in Table 4. Here, the Gibbs free energies of the most stable conformer for each type of uranyl complex were used to calculate ΔGsol of the reactions. Previous studies reported that the penta-coordinated [UO2(H2O)5]2+ is the dominant species in aqueous solution.64–67 Therefore, [UO2(H2O)5]2+ was used to calculate the changes of the Gibbs free energies of the reactions. To study the trend of ΔGsol with the number of the coordinated ligand, we have investigated the ΔGsol of the reactions between the uranyl and 1–5 serine molecules as provided in Table S5, ESI.† It is clearly seen that the absolute values of ΔGsol firstly increase and then decrease with increasing the number of ligand with the maximum value of 20.24 kcal mol−1 for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type uranyl–serine complex. This result may be caused by the steric effect of the serine molecules. Therefore, we mainly focus on the 1
3 type uranyl–serine complex. This result may be caused by the steric effect of the serine molecules. Therefore, we mainly focus on the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 and 1
2 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type uranyl complexes with serine/phosphoserine. It is obvious from Table 4 that the values of ΔGsol for the reactions are all negative, and the absolute values of ΔGsol for the reactions with the same ligand gradually increase with the increasing number of ligand. For instance, the absolute value of ΔGsol for the reaction (3) is 23.42 kcal mol−1, while the corresponding value for the reaction (6) increases to 38.61 kcal mol−1. It reaches to 45.93 kcal mol−1 for the reaction (9). This result shows that the uranyl ion with serine and phosphoserine tends to form the 1
3 type uranyl complexes with serine/phosphoserine. It is obvious from Table 4 that the values of ΔGsol for the reactions are all negative, and the absolute values of ΔGsol for the reactions with the same ligand gradually increase with the increasing number of ligand. For instance, the absolute value of ΔGsol for the reaction (3) is 23.42 kcal mol−1, while the corresponding value for the reaction (6) increases to 38.61 kcal mol−1. It reaches to 45.93 kcal mol−1 for the reaction (9). This result shows that the uranyl ion with serine and phosphoserine tends to form the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes. It is important to highlight that the absolute values of ΔGsol for the same type reaction increases when serine is phosphorylated. However, the extent of the increase is remarkably different between the mono- and dianionic phosphoserine complexes. For instance, the absolute value of ΔGsol for the reaction (7) is 20.24 kcal mol−1, and the corresponding value increases by 2.5 kcal mol−1 for the reaction (8). While it sharply increases to 45.93 kcal mol−1 for the reaction (9). Based on the values of ΔGsol, the uranyl ion can be easily bound with the di-anionic phosphoserine, which indicates that a high solution pH is more favorable for uranyl ion coordination with phosphoserine.
3 type complexes. It is important to highlight that the absolute values of ΔGsol for the same type reaction increases when serine is phosphorylated. However, the extent of the increase is remarkably different between the mono- and dianionic phosphoserine complexes. For instance, the absolute value of ΔGsol for the reaction (7) is 20.24 kcal mol−1, and the corresponding value increases by 2.5 kcal mol−1 for the reaction (8). While it sharply increases to 45.93 kcal mol−1 for the reaction (9). Based on the values of ΔGsol, the uranyl ion can be easily bound with the di-anionic phosphoserine, which indicates that a high solution pH is more favorable for uranyl ion coordination with phosphoserine.
| Reactions | ΔGsol |
|---|---|
| (1) SerH + [UO2(H2O)5]2+ = [UO2(SerH)(H2O)3]2+ + 2H2O | −10.99 |
| (2) [pSerH2]− + [UO2(H2O)5]2+ = [UO2(pSerH2)(H2O)3]+ + 2H2O | −12.34 |
| (3) [pSerH]2− + [UO2(H2O)5]2+ = [UO2(pSerH)(H2O)3] + 2H2O | −23.42 |
| (4) 2SerH + [UO2(H2O)5]2+ = [UO2(SerH)2(H2O)]2+ + 4H2O | −18.78 |
| (5) 2[pSerH2]− + [UO2(H2O)5]2+ = [UO2(pSerH2)2(H2O)] + 4H2O | −20.02 |
| (6) 2[pSerH]2− + [UO2(H2O)5]2+ = [UO2(pSerH)2(H2O)]2− + 4H2O | −38.61 |
| (7) 3SerH + [UO2(H2O)5]2+ = [UO2(SerH)3]2+ + 5H2O | −20.24 |
| (8) 3[pSerH2]− + [UO2(H2O)5]2+ = [UO2(pSerH2)3]− + 5H2O | −22.74 |
| (9) 3[pSerH]2− + [UO2(H2O)5]2+ = [UO2(pSerH)3]4− + 5H2O | −45.93 |
Conclusions
In summary, the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 and 1
2 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes of the uranyl ion with the zwitterionic serine, mono- and dianionic phosphoserine were optimized in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory using SMD. Various conformers for each type of complex were considered for structure optimization and the corresponding relative energies were compared. As a result, the binding affinity of uranyl ion toward three functional groups was found to follow the order of –PO42− > –COO− > –PO4H−. Moreover, the relative energies for the 1
3 type complexes of the uranyl ion with the zwitterionic serine, mono- and dianionic phosphoserine were optimized in the aqueous solution at the B3LYP/ECP60MWB-SEG/6-311+G(d) level of theory using SMD. Various conformers for each type of complex were considered for structure optimization and the corresponding relative energies were compared. As a result, the binding affinity of uranyl ion toward three functional groups was found to follow the order of –PO42− > –COO− > –PO4H−. Moreover, the relative energies for the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes indicates that the uranyl ion prefers to adopt a penta-coordination mode, that is, the carboxyl and phosphate groups are in a mono-dentate fashion when the equatorial coordination number of the uranyl ion reaches five. The analyses of the NBO and QTAIM suggest that the equatorial U–O bonds of these complexes have ionic characters. The changes of Gibbs free energy for the reactions suggest that the uranyl ion with serine/phosphoserine easily forms the 1
3 type complexes indicates that the uranyl ion prefers to adopt a penta-coordination mode, that is, the carboxyl and phosphate groups are in a mono-dentate fashion when the equatorial coordination number of the uranyl ion reaches five. The analyses of the NBO and QTAIM suggest that the equatorial U–O bonds of these complexes have ionic characters. The changes of Gibbs free energy for the reactions suggest that the uranyl ion with serine/phosphoserine easily forms the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type complexes. Obviously, the absolute values of ΔGsol increase after serine is phosphorylated for the same type reaction. Based on the ΔGsol values, the uranyl ion can be easily bound with the dianionic phosphoserine, which indicates that a higher solution pH (about pH = 8) is more favorable for the uranyl ion coordination with phosphoserine. This work could render insights into the coordination nature of the uranyl ion with serine/phosphoserine under different conditions and provide theoretical basis and guide for study on the interactions between actinide cations with peptides and proteins.
3 type complexes. Obviously, the absolute values of ΔGsol increase after serine is phosphorylated for the same type reaction. Based on the ΔGsol values, the uranyl ion can be easily bound with the dianionic phosphoserine, which indicates that a higher solution pH (about pH = 8) is more favorable for the uranyl ion coordination with phosphoserine. This work could render insights into the coordination nature of the uranyl ion with serine/phosphoserine under different conditions and provide theoretical basis and guide for study on the interactions between actinide cations with peptides and proteins.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 11275219, 21477130, 91426302, 91326202) and the “Strategic Priority Research Program” of the Chinese Academy of Sciences (Grant No. XDA030104). The results described in this work were obtained on the ScGrid of Supercomputing Center, Computer Network Information Center of Chinese Academy of Sciences.Notes and references
- T. S. Anirudhan and P. G. Radhakrishnan, J. Environ. Radioact., 2009, 100, 250–257 CrossRef CAS PubMed.
- J. D. Van Horn and H. Huang, Coord. Chem. Rev., 2006, 250, 765–775 CrossRef CAS.
- C. Bresson, E. Ansoborlo and C. Vidaud, J. Anal. At. Spectrom., 2011, 26, 593–601 RSC.
- C. Vidaud, A. Dedieu, C. Basset, S. Plantevin, I. Dany, O. Pible and E. Quéméneur, Chem. Res. Toxicol., 2005, 18, 946–953 CrossRef CAS PubMed.
- C. Vidaud, S. Gourion-Arsiquaud, F. Rollin-Genetet, C. Torne-Celer, S. Plantevin, O. Pible, C. Berthomieu and E. Quéméneur, Biochemistry, 2007, 46, 2215–2226 CrossRef CAS PubMed.
- M. G. Benavides-Garcia and K. Balasubramanian, Chem. Res. Toxicol., 2009, 22, 1613–1621 CrossRef CAS PubMed.
- G. Montavon, C. Apostolidis, F. Bruchertseifer, U. Repinc and A. Morgenstern, J. Inorg. Biochem., 2009, 103, 1609–1616 CrossRef CAS PubMed.
- C. Basset, O. Averseng, P.-J. Ferron, N. Richaud, A. Hagège, O. Pible and C. Vidaud, Chem. Res. Toxicol., 2013, 26, 645–653 CrossRef CAS PubMed.
- C. Lebrun, M. Starck, V. Gathu, Y. Chenavier and P. Delangle, Chem.–Eur. J., 2014, 20, 16566–16573 CrossRef CAS PubMed.
- L. Zhou, M. Bosscher, C. Zhang, S. Özçubukçu, L. Zhang, W. Zhang, C. J. Li, J. Liu, M. P. Jensen, L. Lai and C. He, Nat. Chem., 2014, 6, 236–241 CrossRef CAS PubMed.
- S. O. Odoh, G. D. Bondarevsky, J. Karpus, Q. Cui, C. He, R. Spezia and L. Gagliardi, J. Am. Chem. Soc., 2014, 136, 17484–17494 CrossRef CAS PubMed.
- S. Safi, A. Jeanson, J. Roques, P. L. Solari, F. Charnay-Pouget, C. Den Auwer, G. Creff, D. J. Aitken and E. Simoni, Inorg. Chem., 2016, 55, 877–886 CrossRef CAS PubMed.
- C. Acharya and C. A. Blindauer, Inorg. Chem., 2016, 55, 1505–1515 CrossRef CAS PubMed.
- P. W. Durbin, B. Kullgren, J. Xu and K. N. Raymond, Radiat. Prot. Dosim., 1998, 79, 433–443 CrossRef CAS.
- G. J. Lesser and G. D. Rose, Proteins: Struct., Funct., Genet., 1990, 8, 6–13 CrossRef CAS PubMed.
- C. F. Correia, P. O. Balaj, D. Scuderi, P. Maitre and G. Ohanessian, J. Am. Chem. Soc., 2008, 130, 3359–3370 CrossRef CAS PubMed.
- A. Cimas, P. Maitre, G. Ohanessian and M. P. Gaigeot, J. Chem. Theory Comput., 2009, 5, 2388–2400 CrossRef CAS PubMed.
- M. L. Stover, C. E. Plummer, S. R. Miller, C. J. Cassady and D. A. Dixon, J. Phys. Chem. B, 2015, 119, 14604–14621 CrossRef CAS PubMed.
- K. Engholm-Keller and M. R. Larsen, Proteomics, 2013, 13, 910–931 CrossRef CAS PubMed.
- R. Pardoux, S. Sauge-Merle, D. Lemaire, P. Delangle, L. Guilloreau, J.-M. Adriano and C. Berthomieu, PLoS One, 2012, 7, e41922 CAS.
- B. Li, J. Raff, A. Barkleit, G. Bernhard and H. Foerstendorf, J. Inorg. Biochem., 2010, 104, 718–725 CrossRef CAS PubMed.
- S. Safi, G. Creff, A. Jeanson, L. Qi, C. Basset, J. Roques, P. L. Solari, E. Simoni, C. Vidaud and C. Den Auwer, Chem.–Eur. J., 2013, 19, 11261–11269 CrossRef CAS PubMed.
- Y. Liu, C. Xiao, L. Mei, G. Qu and W. Shi, J. Nucl. Radiochem. Sci., 2014, 36, 129–139 CAS.
- L. Qi, C. Basset, O. Averseng, E. Quemeneur, A. Hagege and C. Vidaud, Metallomics, 2014, 6, 166–176 RSC.
- M. Starck, N. Sisommay, F. A. Laporte, S. Oros, C. Lebrun and P. Delangle, Inorg. Chem., 2015, 54, 11557–11562 CrossRef CAS PubMed.
- C. T. Yang, J. Han, M. Gu, J. Liu, Y. Li, Z. Huang, H. Z. Yu, S. Hu and X. L. Wang, Chem. Commun., 2015, 51, 11769–11772 RSC.
- G. Creff, S. Safi, J. Roques, H. Michel, A. Jeanson, P.-L. Solari, C. Basset, E. Simoni, C. Vidaud and C. Den Auwer, Inorg. Chem., 2016, 55, 29–36 CrossRef CAS PubMed.
- M. Sundararajan, A. J. Campbell and I. H. Hillier, J. Phys. Chem. A, 2008, 112, 4451–4457 CrossRef CAS PubMed.
- M. Sundararajan, R. S. Assary, I. H. Hillier and D. J. Vaughan, Dalton Trans., 2011, 40, 11156–11163 RSC.
- M. Sundararajan, A. J. Campbell and I. H. Hillier, Faraday Discuss., 2011, 148, 195–205 RSC.
- J. Su, K. Zhang, W. H. E. Schwarz and J. Li, Inorg. Chem., 2011, 50, 2082–2093 CrossRef CAS PubMed.
- V. E. Jackson, K. E. Gutowski and D. A. Dixon, J. Phys. Chem. A, 2013, 117, 8939–8957 CrossRef CAS PubMed.
- J. P. Perdew, Phys. Rev. B: Condens. Matter Mater. Phys., 1986, 33, 8822 CrossRef.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865–3868 CrossRef CAS PubMed.
- C. T. Lee, W. T. Yang and R. G. Parr, Phys. Rev. B: Condens. Matter Mater. Phys., 1988, 37, 785–789 CrossRef CAS.
- A. D. Becke, Phys. Rev. A, 1988, 38, 3098–3100 CrossRef CAS.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- S. Grimme, S. Ehrlich and L. Goerigk, J. Comput. Chem., 2011, 32, 1456–1465 CrossRef CAS PubMed.
- Y. Zhao and D. Truhlar, Theor. Chem. Acc., 2008, 120, 215–241 CrossRef CAS.
- G. Schreckenbach, P. J. Hay and R. L. Martin, Inorg. Chem., 1998, 37, 4442–4451 CrossRef CAS PubMed.
- G. Schreckenbach, P. J. Hay and R. L. Martin, J. Comput. Chem., 1999, 20, 70–90 CrossRef CAS.
- J. L. Sonnenberg, P. J. Hay, R. L. Martin and B. E. Bursten, Inorg. Chem., 2005, 44, 2255–2262 CrossRef CAS PubMed.
- G. A. Shamov and G. Schreckenbach, J. Am. Chem. Soc., 2008, 130, 13735–13744 CrossRef CAS PubMed.
- P. Wåhlin, C. Danilo, V. Vallet, F. Réal, J.-P. Flament and U. Wahlgren, J. Chem. Theory Comput., 2008, 4, 569–577 CrossRef PubMed.
- S. Tsushima, Inorg. Chem., 2012, 51, 1434–1439 CrossRef CAS PubMed.
- S. Vukovic, B. P. Hay and V. S. Bryantsev, Inorg. Chem., 2015, 54, 3995–4001 CrossRef CAS PubMed.
- B. Averkiev, M. Mantina, R. Valero, I. Infante, A. Kovacs, D. Truhlar and L. Gagliardi, Theor. Chem. Acc., 2011, 129, 657–666 CrossRef CAS.
- W. Küchle, M. Dolg, H. Stoll and H. Preuss, J. Chem. Phys., 1994, 100, 7535–7542 CrossRef.
- X. Cao, M. Dolg and H. Stoll, J. Chem. Phys., 2003, 118, 487–496 CrossRef CAS.
- X. Cao and M. Dolg, J. Mol. Struct.: THEOCHEM, 2004, 673, 203–209 CrossRef CAS.
- J.-H. Lan, W.-Q. Shi, L.-Y. Yuan, Y.-L. Zhao, J. Li and Z.-F. Chai, Inorg. Chem., 2011, 50, 9230–9237 CrossRef CAS PubMed.
- Q.-Y. Wu, C.-Z. Wang, J.-H. Lan, C.-L. Xiao, X.-K. Wang, Y.-L. Zhao, Z.-F. Chai and W.-Q. Shi, Inorg. Chem., 2014, 53, 9607–9614 CrossRef CAS PubMed.
- Q.-Y. Wu, J.-H. Lan, C.-Z. Wang, C.-L. Xiao, Y.-L. Zhao, Y.-Z. Wei, Z.-F. Chai and W.-Q. Shi, J. Phys. Chem. A, 2014, 118, 2149–2158 CrossRef CAS PubMed.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Phys. Chem. B, 2009, 113, 6378–6396 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, N. J. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, Ö. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian 09, Gaussian, Inc., Wallingford, CT, USA, 2009 Search PubMed.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- C. Gardiennet-Doucet, X. Assfeld, B. Henry and P. Tekely, J. Phys. Chem. A, 2006, 110, 9137–9144 CrossRef CAS PubMed.
- M. M. Quesada-Moreno, J. R. Avilés-Moreno, A. Á. Márquez-García, F. Partal-Ureña and J. J. López González, J. Mol. Struct., 2013, 1046, 136–146 CrossRef CAS.
- S. Gronert and R. A. J. O'Hair, J. Am. Chem. Soc., 1995, 117, 2071–2081 CrossRef CAS.
- D. Scuderi, C. F. Correia, O. P. Balaj, G. Ohanessian, J. Lemaire and P. Maitre, ChemPhysChem, 2009, 10, 1630–1641 CrossRef CAS PubMed.
- P. L. Arnold, Z. R. Turner, N. Kaltsoyannis, P. Pelekanaki, R. M. Bellabarba and R. P. Tooze, Chem.–Eur. J., 2010, 16, 9623–9629 CrossRef CAS PubMed.
- A. Kerridge, Dalton Trans., 2013, 42, 16428–16436 RSC.
- B. M. Gardner, G. Balázs, M. Scheer, F. Tuna, E. J. L. McInnes, J. McMaster, W. Lewis, A. J. Blake and S. T. Liddle, Angew. Chem., Int. Ed., 2014, 53, 4484–4488 CrossRef CAS PubMed.
- D. Hagberg, G. Karlström, B. O. Roos and L. Gagliardi, J. Am. Chem. Soc., 2005, 127, 14250–14256 CrossRef CAS PubMed.
- K. E. Gutowski and D. A. Dixon, J. Phys. Chem. A, 2006, 110, 8840–8856 CrossRef CAS PubMed.
- F. Réal, A. S. P. Gomes, L. Visscher, V. r. Vallet and E. Eliav, J. Phys. Chem. A, 2009, 113, 12504–12511 CrossRef PubMed.
- S. Kerisit and C. Liu, J. Phys. Chem. A, 2013, 117, 6421–6432 CrossRef CAS PubMed.
Footnote |
† Electronic supplementary information (ESI) available: Various conformers of three types of serine and four types of phosphoserine as well as their relative energies, optimized 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, 1 2, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, 1 3, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 and 1 4 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 type structures of the uranyl ion with zwitterionic serine and the corresponding relative energies, optimized 1 5 type structures of the uranyl ion with zwitterionic serine and the corresponding relative energies, optimized 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 type structures of the uranyl ion with dianionic phosphoserine and the corresponding relative energies and bonding parameters are provided. See DOI: 10.1039/c6ra14906e 3 type structures of the uranyl ion with dianionic phosphoserine and the corresponding relative energies and bonding parameters are provided. See DOI: 10.1039/c6ra14906e |
| This journal is © The Royal Society of Chemistry 2016 |