Liposome-based gene delivery systems containing a steroid derivative: computational and small angle X-ray diffraction study†
R.
Galeazzi
*a,
P.
Bruni
b,
E.
Crucianelli
a,
E.
Laudadio
a,
M.
Marini
b,
L.
Massaccesi
a,
G.
Mobbili
a and
M.
Pisani
*b
aDi.S.V.A. Department, Polytechnic University of Marche, Via Brecce Bianche, Ancona, Italy. E-mail: r.galeazzi@univpm.it
bSIMAU Department, Polytechnic University of Marche, Via Brecce Bianche, Ancona, Italy. E-mail: m.pisani@univpm.it
First published on 15th June 2015
Abstract
In this study, the structural properties and the phase behaviour of mixed composition neutral liposomes containing a functionalized steroid are reported. With the aim to design neutral liposomes able to coordinate cations and to complex DNA, we synthesized cholesteryl-2-(picolinamido)-phenylcarbamate (CHOLp) containing an N-aryl picolinamide group as chelating agent linked to the steroid structure via a carbamate moiety. The phase behaviour of mixtures of the functionalized cholesterol (CHOLp) and dioleoyl-phosphatidylcholine (DOPC) was investigated by means of X-ray diffraction. Simultaneously, atomistic molecular dynamics simulations of DOPC/CHOLp bilayers as a function of CHOLp molar fractions were carried out to investigate the specific effects of the polar steroid on the structural and dynamic properties of these zwitterionic bilayers. The molecular modelling studies have been performed both in absence and in presence of bivalent cations salts in order to assess the CHOLp ability to coordinate metal ions. The results show good stability of the resulting DOPC/CHOLp bilayers which is improved by the presence of salt. This is particularly evident at low amount of CHOLp where a high order of the lipid tails can be observed, suggesting stabilization of the corresponding DOPC liposomes. This feature can be ascribed to the polar nature and structural properties of the ligand. In fact, due to the presence of the aromatic moieties, CHOLp combines two different behaviours, namely a propensity to realize both intermolecular π-stacking interactions and cation–π bonding mainly evident in CaCl2. The last feature confirms for CHOLp a role as a cation-mediated complexation agent for DNA. The X-ray diffraction data on the capability of DOPC/CHOLp liposomes to complex DNA was also reported.
Introduction
The research towards efficient nanocarriers for drug and gene delivery has been continuously carried out to develop safe formulations with a high degree of specificity. The possibility of a successful transfer of genetic material to target cells is closely dependent on the choice of the appropriate delivery system, which can be viral or synthetic. Among synthetic vectors, liposomes are the most studied ones and have been extensively tested because of their ability to encapsulate and to deliver hydrophobic and hydrophilic therapeutic agents into cells.1,2 Cationic liposomes are among the most popular gene delivery systems used since they arrange into supramolecular structures where DNA condenses through electrostatic interactions. However, the toxicity and the low stability of these lipoplexes in serum have overshadowed their application as non-viral vectors. On the other hand, zwitterionic liposomes have not yet received wide attention for gene delivery because of their low capability to interact with nucleic acids.In this area of interest, the self-assembled association of neutral liposomes (L), DNA and divalent metal cations (M2+) in ternary L–DNA–M2+ complexes able to transfect genetic materials into cells have been explored;3–7 in this system the interaction between neutral lipids and DNA is mediated by metal cations that provide stability to the complexes. In order to improve the ability of neutral liposomes to complex DNA, our strategy was to develop liposomal gene delivery systems containing new synthetic lipids lacking in positive charge but capable of acting as effective cationic lipids. This can be achieved by the inclusion in the lipid polar head of a chelating group able to coordinate bivalent metals and to form stable complexes with DNA. The lipids most commonly used as delivery systems have a hydrophobic moiety that consists of long hydrocarbon chains or of a steroid skeleton. Hence, we planned the synthesis of a derivative of functionalized cholesterol with an N-(2-aminophenyl)-picolinamide ligand, a chelating aromatic agent with three donor atoms, able to coordinate cations either as a bidentate ligand, or assessing specific strong cation–π interactions. Indeed, it is well known that steroids are often included in liposome formulations since they may improve in vivo and in vitro the liposome stability.8–12 Many functionalized cholesterol derivatives have been studied in order to improve steroid properties in terms of the ordering and condensing effect.13 In fact, the intermolecular interactions between the steroid and phospholipids are strongly dependent on the matrix lipid structure and on the nature of functionalization.14,15
Since the composition of liposomes is known to have a marked influence on bilayers properties, a structural study was carried out on liposomes and lipid bilayers containing our synthetic compound. Here, we show the results of small angle X-ray diffraction experiments and atomistic molecular dynamics simulations of DOPC-based membrane bilayers containing a variable amount of the functionalized steroid (CHOLp) both in pure water and in presence of bivalent metal salts (MgCl2, CaCl2). Our first aim was to point out the effect of a polar functionalization of cholesterol on the structural and electrostatic properties of neutral bilayers as microscopic models for nanostructured delivery systems. The second aim was to evaluate the capability of CHOLp to improve and form complexes with DNA through a cation coordination pattern. For this purpose mixtures of DOPC/CHOLp in the presence of metal ions and DNA were also investigated by X-ray diffraction.
Experimental section
Chemicals and synthetic protocols
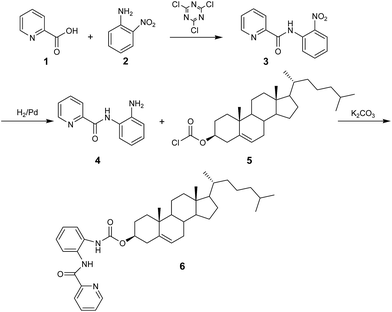
The following lipid 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) was purchased from Avanti Polar Lipids Inc. (Alabaster, AL). The DNA from calf thymus was purchased from Sigma Chemical Co. The DNA aqueous solution was sonicated in order to induce a DNA fragmentation whose length distribution, detected by gel electrophoresis, varied between 500 and 2000 bp. All materials and reagents used in the synthesis and in the samples preparation were purchased from SigmaAldrich Co. (Stenheim, Germany) unless otherwise stated and used without purification. All solvents were analytically pure and dried before use.Column chromatography was performed using silica gel 60 (230–400 mesh). Mass spectra (MS) were obtained by electron impact on a Hewlett-Packard spectrometer 5890, series II. The 1H and 13C NMR spectra were recorded at 400 and 100 MHz, respectively, on a Varian Gemini spectrometer, using CDCl3 as solvent. Chemical shifts (δ) are reported in ppm relative to TMS and coupling constants (J) in Hz. The synthetic routes of the cholesteryl-2-(picolinamido)-phenylcarbamate are reported in Scheme 1. Detailed synthetic procedures are given in the ESI.†
Briefly, starting from 2-picolinic acid and 2-nitroaniline, the corresponding N-(2-nitrophenyl)-picolinamide was obtained by using 2,4,6-trichloro-1,3,5-triazine as suitable activating agent. The subsequent reduction of the nitro group by Pd catalyzed hydrogenation gave the chelating agent N-(2-aminophenyl)-picolinamide which was then linked to the hydroxyl group of cholesterol by reaction with cholesteryl chloroformate.
Sample preparation
Appropriate volumes of stock solutions in chloroform of DOPC (78 mM), CHOLp (15 mM) were mixed to obtain liposomes with different molar fractions X (mol mol−1) of non-zwitterionic lipid (X = non zwitterionic lipid/total lipid = 0.05, 0.15, 0.25, 0.35). Organic solvent was evaporated under vacuum with a Speedvac apparatus to form a dry film. The film was hydrated in 20 mM HEPES buffer, pH = 7.4 to obtain liposomes at 25 mM total lipid concentration for X-ray experiments. The DOPC/CHOLp![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA
DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) M2+ complexes were obtained by mixing the multilamellar liposomal suspension with buffer solution of bivalent metal salt and DNA. The final concentrations in the sample are: 25 mM for DOPC/CHOLp, 62.5 mM for bivalent salt and 12.5 mM for DNA, thus giving molar ratio lipid/DNA/M2+ for the complex formation equal to 2
M2+ complexes were obtained by mixing the multilamellar liposomal suspension with buffer solution of bivalent metal salt and DNA. The final concentrations in the sample are: 25 mM for DOPC/CHOLp, 62.5 mM for bivalent salt and 12.5 mM for DNA, thus giving molar ratio lipid/DNA/M2+ for the complex formation equal to 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10. The samples were incubated at 4 °C for 24 hours.
10. The samples were incubated at 4 °C for 24 hours.
X-ray measurements
XRD measurements were carried out at the high brilliance beamline ID02 of the European Synchrotron Radiation Facility (Grenoble, France). The energy of the incident beam was 12.5 keV (λ = 0.995 Å), the beam size 100 × 100 μm2, and the sample-to-detector distance 1.2 m. The 2D diffraction patterns were collected by a CCD detector. The small angle q range from qmin = 0.1 nm−1 to qmax = 4 nm−1 with a resolution of 5 × 10−3 nm−1 (FWHM) was investigated. The samples were held in a 1 mm sized glass capillary. To avoid radiation damage, each sample was exposed to radiation for a maximum of 3 s per frame.Computational methods and procedures
Setting out CHOLp 3D structure: conformational analysis and DFT calculations
The conformational potential energy surface (PES) of CHOLp was extensively explored in order to localize on it the main stationary points, i.e. lowest energy minimum and the most populated conformers. Molecular mechanics energy calculations were performed using the implementation of AMBER force field included in the Maestro/MacroModel (Schrodinger Inc., Portland, OR, USA) software framework,16,17 and the torsional space of the molecule was randomly varied with the usage-directed Monte Carlo Multiple Minimum (MCMM) conformational search approach.18 For each search, at least 1000 starting structures for each variable torsion angle were generated and minimized until the gradient was less than 0.05 kJ Å−1 mol−1. Duplicate conformations and those with an energy in excess of 6.0 kcal mol−1 above the global minimum were discarded. The solvent effect was included by using the implicit water GB/SA solvation method19 to take into account polar solvent effects. The cluster analysis was performed within the Macromodel package using Xcluster20,21 following a protocol previously reported.22,23 Furthermore, the lowest energy minimum, together with the representative structure of the most populated clusters, were optimized by DFT calculation using G09 suite of Gaussian 09, Revision D.01 software,24 at B3LYP/6-311G** level of theory,25–28 in order to better take into account the electronic effects in the conformer's stabilization and populations.Molecular dynamics of the mixed bilayer
The molecular dynamics simulations of the mixed bilayers containing DOPC lipids and the CHOLp molecule at different concentration were carried out on the isothermal–isobaric (N, P, T) ensemble at 1 atm and 310 K (37 °C). The membrane leaflets are composed by a total of 72 lipid molecules hydrated by 2404 water molecules within an initial simulation box (corresponding to a pre-equilibrated pure DOPC bilayer) that was 7 nm (z) normal to the bilayer and 5 nm long in each of the two dimensions of the bilayer plane (xy). The starting pure DOPC systems were an extension of 35 ns Klauda equilibrated systems by NPT ensemble simulation,29 whilst the mixed composition bilayers considered in this study were built substituting randomly pre-equilibrated DOPC molecules with CHOLp until the corresponding concentration of the mixed bilayer was reached (XCHOLp = 0.05, 0.15, 0.25). All the simulations were conducted both in pure water and in presence of bivalent metal salts (MgCl2, CaCl2). The TIP3P model for solvent has been used and ions were added to reach salt physiological concentration. Water and ions overlapping the membrane bilayer were removed before proceeding to system minimization.30 The chosen system dimensions were based on literature reports concerning the smallest representative size which can be used to accurately reproduce the occurring intermolecular interactions in lipid bilayers.31–33 In particular, MD simulations have been extensively carried out by Klauda and co-workers on different sized systems (72 up to 288 lipids) to examine system size dependence on dynamical properties associated with the Particle Mesh Ewald (PME) treatment of electrostatic interactions.34–37 As a result, it was pointed out that a system size of 72 lipids is sufficiently large for calculating the main structural properties. Consequently, for our purpose, the simulations have been performed in that conditions to rapidly obtain the desired properties.33 The GROMACS 4.6.7 suite of programs38 was used to perform all the simulations using AMBER ff99SB-ILDN force field39 parameter sets; the charges used for simulation were previously calculated quantum mechanically at DFT level of theory using B3LYP/6-311G** method (Gaussian 09, Revision D.01),24–28 and integrated into the bilayer structure.Each bilayer system was hydrated with TIP3P water molecules and the energy minimized under periodic box conditions (starting cell unit in nm, X = 4.87 nm, Y = 4.87 nm, Z = 7.94 nm) applied in all directions using a neighbour searching grid type, and also setting at 1.4 nm the cut-off distance for the short range neighbour list. Electrostatic were taken into account implementing a fast smooth particle-mesh Ewald (SPME) algorithm,34–37 with a 1.4 nm distance for the Coulomb cut-off, since this method is considered to be both efficient and accurate for the evaluation of long-range electrostatic interactions in large macromolecular systems.40,41
Then molecular dynamics equilibration with velocities generated according to a Maxwell distribution at temperature of 310 K using a random seed has been performed. For all systems 2 ns of annealing simulations were carried out to gradually increase the temperature till 310 K and then to generate atom velocities; no water was observed inside the bilayer. At this point a 100 ns dynamics has been set up for each of built system. An accurate leap-frog stochastic dynamics integrator was used as the main run control option. The whole run lasted 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ps with a time step of 0.002 ps, and coordinates were written out every 10 ps, while energy data were collected every 2 ps.
000 ps with a time step of 0.002 ps, and coordinates were written out every 10 ps, while energy data were collected every 2 ps.
The first 2 ns MD simulation for each lipid system was simulated in the NVT ensemble using a Langevin thermostat while the subsequent nanoseconds in the NPT ensemble (T = 310 K, P = 1 atm) using a Berendsen thermostat and semi-isotropic pressure coupling. A time constant for coupling of 0.5 ps and an optimal compressibility for water of 4.5 × 10−5 bar−1, were implemented to obtain the best control on pressure. The MD trajectories were collected until the equilibration period achieved a convergence of the dimensions of the system and the steady state was reached.42 At this point, the dimensions of the lipid bilayer was evaluated over the last 20 ns of the stabilized trajectory in order to define accurately the bilayer properties such as the area per lipid, membrane thickness in an equilibrium state. The analysis of the MD trajectories was performed within the VMD43 and CHIMERA software.44 Computation of each MD trajectory was performed in parallel at a speed of 9 ns per day on a PLX IBM workstation (CINECA-HPC ISCRA project, PRACE DECI project).
Results and discussion
In order to improve the complexation of zwitterionic liposomes with DNA, we planned to synthesize a lipid bearing a chelating group in the polar head capable to coordinate metal cations. We started from cholesterol which was modified introducing an aromatic polar moiety like N-(2-aminophenyl)-picolinamide which could be easily inserted into the leaflets and it can interact with metal ions. This synthetic derivative CHOLp carrying a metal-chelating headgroup was predicted in silico as a promising candidate by means of molecular modeling tools.The conformational preferences asset of CHOLp were assessed performing a Monte Carlo MC/MM extensive search using AMBER force field for energy calculations. The results obtained allowed to evaluate the CHOLp capability to insert into the lipid bilayers and they showed that the most populated conformations belong to the same cluster family differing mainly in the orientation of the aromatic moieties and of the alkyl terminal group with respect to the steroid skeleton (Fig. 1). In the most populated conformer the aromatic rings are oriented through a π–π stacking interactions (“close” conformation, Fig. 1(B)). The optimized lowest energy conformer for CHOLp shows high stability and is perfectly suitable to be inserted into a mixed composition bilayer.
 | ||
| Fig. 1 Superimposition of the first six most populated conformers of CHOLp ΔE < 3.0 kJ mol−1 (A); lowest energy conformation for CHOLp (B). | ||
Thus, CHOLp was then introduced into DOPC liposomes and their structural features were analyzed by means of both full atom molecular dynamics calculation and X-ray scattering experiments.
MD simulations results
Molecular dynamics simulations were performed to study the effect of insertion of the functionalized steroid on the structural properties of pure DOPC matrix bilayers. Indeed, even upon first analysis, the CHOLp molecule clearly appears well inserted into the lipid bilayer, a behaviour which can be ascribed, as already suggested, to its physical chemical properties including shape and conformation. Furthermore, its aromatic polar head orients itself outside the leaflet and almost parallel to the DOPC polar head. This feature immediately places CHOLp in a good light with respect to an eventual metal ion complexation.The bilayer dynamical and structural properties arising from the collected MD trajectories, clearly point out that the CHOLp molecules strongly influence each other throughout all the simulations. This behaviour is particularly evident in absence of salt, while in the presence of MgCl2 and CaCl2 the intermolecular interactions between the steroids are competing with those between CHOLp and metal ions. In the following Section, we will discuss the results obtained in details.
MD simulations in pure water
In pure water, CHOLp intermolecular interactions are smooth at low concentration (XCHOLp = 0.05) but increasing CHOLp molar fraction (XCHOLp = 0.15, 0.25), aggregation of the functionalized polar cholesterol molecules occurs (Fig. 2).More in particular, the intermolecular interactions occurring between the phenyl moieties of CHOLp are responsible for this peculiar behaviour. In fact, from the MD trajectories, after cluster analysis, the most representative structures were identified. Indeed, the CHOLp molecules interact strongly through edge to face, π–π stacking, CH–π, NH–π interactions as shown in Fig. 3. Moreover, in all the simulations a strong stability of the overall bilayers is observed as can be seen in Fig. 4. Moreover, as can be deduced from RMSD values calculated with respect to the starting minimized structure (Fig. 4), the higher the CHOLp concentration the greater is the molecular reorganization of the bilayers. Anyway, the steady state is reached easily in all cases and at early simulation time.
 | ||
| Fig. 3 Principal intermolecular NH–π, π–π interactions between CHOLp molecules in the lipid bilayers in representative MD frames at XCHOLp = 0.15. | ||
 | ||
| Fig. 4 Root mean square deviation (RMSD) evolution for MD trajectories frames for CHOLp through all 100 ns simulations. RMSD is calculated with respect to the starting minimized structure (frame 0). | ||
MD simulations in MgCl2
In presence of magnesium chloride, CHOLp intermolecular interactions are evident even at low concentration (XCHOLp = 0.05) and well organized at higher concentrations. Beside, the steroid polar head is kept outside the hydrophobic lipidic environment, being able in this way to directly interact also with the metal ions through their aromatic moieties. Furthermore, the starting “closed” conformation of CHOLp opens becoming more suitable for a greater number of stabilizing interactions with metal ions (Fig. 5). Electrostatic interactions with magnesium are present and are responsible of the steroid molecules orientation of the head outside the leaflets, but the overall coordination is poor (Fig. 6). The membrane thickness is kept at values very close to the model in pure water at all CHOLp values (Fig. 7, Table 1). At higher steroid concentration, the thickness slightly increases as a result of the kind and number of intermolecular CHOLp interactions. | ||
| Fig. 7 Membrane thickness of MD trajectories frames (calculated on phosphate groups) for CHOLp in water (A), MgCl2 (B) and CaCl2 (C), average values for the last 20 ns. | ||
| System | X = 0.05 | X = 0.15 | X = 0.25 |
|---|---|---|---|
| With water | 4.10 nm ± 0.01 | 4.16 nm ± 0.02 | 4.20 nm ± 0.01 |
| With MgCl2 | 4.14 nm ± 0.01 | 3.99 nm ± 0.01 | 4.34 nm ± 0.01 |
| With CaCl2 | 4.20 nm ± 0.01 | 4.22 nm ± 0.02 | 4.12 nm ± 0.01 |
MD simulations in CaCl2
In presence of calcium chloride, we can observe the existence of two different competing behaviours: the first is calcium coordination through a cation–π interaction with the aromatic moieties of CHOLp, the second is the steroid aggregation. Increasing the CHOLp concentration the first behaviour overcomes the second one and to better realize it, the steroid assumes once again an extended conformation (the “open” conformation, see Fig. 5) which puts the polar head close to metal ion in the water phase (Fig. 8). Overall CHOLp shows an high propensity to coordinate the calcium cation regardless its concentration. The membrane thickness is overall kept at values very close to the model in pure water at all CHOLp concentrations and underlines a conserved behaviour in each supramolecular system (Fig. 7, Table 1). | ||
| Fig. 8 CHOLp cation–π and π–π stacking interactions at XCHOLp = 0.25 in CaCl2; calcium ions are represented in cyan while chlorine ions are represented in orange. | ||
In silico data comparison: area per lipid
Achievement of the steady state during simulation was monitored throughout the RMSD analysis and by average area per lipid calculations. The area occupied by each lipid molecule was obtained by multiplying the X and Y dimensions of the simulation box and dividing it by the number of lipid molecules present in each leaflet.33,42 Then, the area per lipid obtained from the MD simulations was calculated and reported (Table 2) for all the three models considered, considering both DOPC and CHOLp as lipids. The behaviour of CHOLp mixed bilayer in pure water was compared with that containing CHOLp at the same molar fraction in presence of CaCl2 and MgCl2. A significative variation can be reported only at higher concentration for CHOLp in CaCl2, as can be seen from Table 2, where are reported the average values for the last 20 ns. The lower value of average area for lipid can be ascribed to a more ordered steroid reposition into the bilayer, thanks to the increasing number of cation–steroid interactions. Thus this reduction in the average area per lipid is a direct consequence of the tighter packaging of the lipids (Fig. 8).| System | X = 0.05 | X = 0.15 | X = 0.25 |
|---|---|---|---|
| With water | 0.54 ± 0.01 | 0.54 ± 0.03 | 0.52 ± 0.01 |
| With MgCl2 | 0.52 ± 0.02 | 0.52 ± 0.01 | 0.52 ± 0.02 |
| With CaCl2 | 0.55 ± 0.01 | 0.56 ± 0.02 | 0.48 ± 0.02 |
On the contrary, in pure water and in MgCl2, CHOLp is more inserted inside the leaflet since the intermolecular steroid interactions overcomes the cation coordination.
Small angle X-ray diffraction
The DOPC/CHOLp system as a function of the molar fraction of CHOLp was investigated by XRD. It is well known that for fully hydrated samples, DOPC forms a fluid lamellar phase Lα at room temperature, with the typical pattern of a one-dimensional lamellar phase with lipid bilayers separated by aqueous layers. Using the Bragg equation, the repeat period d of lamellar phase was determined from the position of the first-order diffraction peak in the reciprocal space. At room temperature, pure DOPC45 shows d = 6.32 nm.The behaviour of the binary mixture DOPC/CHOLp as a function of the guest lipid concentrations is shown in Fig. 9. The X-ray diffraction patterns of DOPC/CHOLp systems show lamellar Lα phases. The presence of CHOLp in the bilayer induces a peak splitting already observed at low molar fraction of CHOLp and the split becomes more evident at XCHOLp = 0.35.
Moreover, the repeat distances (d1, d2) do not increase with CHOLp concentration as reported in Table 3. Supported by the MD simulation results we can deduce that the split reflections is induced from two different populations of the bilayer: one is the phase (L1α) which has the larger spacing (d1) very close to the reported DOPC lamellar spacing, while the second one (L2α) with bilayer thickness d2, could be attributed to the liquid phase richer in CHOLp (see Table 3).
| X CHOLp | d 1 − d2 (nm) |
|---|---|
| 0 | 6.32 |
| 0.05 | 6.35 − 6.23 |
| 0.15 | 6.36 − 6.21 |
| 0.25 | 6.34 − 6.22 |
| 0.35 | 6.37 − 6.16 |
The lipid aggregation of CHOLp in the bilayer is dependent upon the structural properties of the polar steroid. Indeed, the MD simulations for the mixture DOPC/CHOLp show intermolecular interactions among the phenyl moieties of CHOLp and this behaviour is confirmed by the Bragg peaks split.
Although, the formation of these regions richer in CHOLp could hinder the coordination of metal ions by the chelating agent, the addition of salts (MgCl2, CaCl2) to the mixed liposomes produces changes in the overall supramolecular structure.
The Fig. 10 shows the effect of MgCl2 and CaCl2 addition to the DOPC/CHOLp mixture. The X-ray diffraction patterns correspond to the Lα phase with a lamellar repeat distance about d = 5.73 nm for CaCl2 and d = 5.66 nm for MgCl2 and no peaks split is observed for all concentration. As already reported, metal ions induce a local dehydration of the phospholipid polar headgroup and the consequent removal of coordinated water at the lipid headgroup.46,47
 | ||
Fig. 10 X-ray diffraction patterns of the DOPC/CHOLp (0.25) mixture at DOPC/CHOLp![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CaCl2 (MgCl2) (2 CaCl2 (MgCl2) (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5) molar ratio. 5) molar ratio. | ||
Therefore, the presence of the cations, as reported by MD simulation data, generally decreases the number of intermolecular interaction among CHOLp molecules and the cation coordination in this case compete energetically with respect to the intermolecular steroid interactions which are, instead, prevalent in the absence of salts. As a consequence, the steroid assumes a more extended conformation which put the polar head close to metal ion in the water phase. This behaviour is particularly marked with CaCl2.
The next step was to verify the capability of the investigated liposomes to complex DNA in the presence of CaCl2 or MgCl2 by SAXS experiments. The data show that the presence of the polar steroid in the bilayers promotes the complexation. In particular, the best molar ratio for the complex formation among DOPC/CHOLp![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA
DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CaCl2 and DOPC/CHOLp
CaCl2 and DOPC/CHOLp![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA
DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MgCl2, is 2
MgCl2, is 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 and a comparison between the diffraction patterns of both complexes is reported (Fig. 11).
5 and a comparison between the diffraction patterns of both complexes is reported (Fig. 11).
 | ||
Fig. 11 SAXS patterns of DOPC/CHOLp (0.25)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CaCl2 complexes (A) and DOPC/CHOLp (0.25) CaCl2 complexes (A) and DOPC/CHOLp (0.25)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MgCl2 complexes (B) at molar ratio 2 MgCl2 complexes (B) at molar ratio 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5. 5. | ||
The addition of DNA and CaCl2 or MgCl2 to the lipid mixtures produced significant changes in the structures. The SAXS data show in both systems DOPC/CHOLp (0.25)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA
DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CaCl2 and DOPC/CHOLp (0.25)
CaCl2 and DOPC/CHOLp (0.25)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA
DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MgCl2 the coexistence of two distinct liquid-crystalline multilamellar phases. Two sets of Bragg reflections, shown in Fig. 11, correspond to a multilayer Lcα phase, associated with the complexed structure consisting of smectic-like arrays of stacked bilayers with intercalated DNA monolayers, and to a multilayer Lα phase, associated with uncomplexed structures.
MgCl2 the coexistence of two distinct liquid-crystalline multilamellar phases. Two sets of Bragg reflections, shown in Fig. 11, correspond to a multilayer Lcα phase, associated with the complexed structure consisting of smectic-like arrays of stacked bilayers with intercalated DNA monolayers, and to a multilayer Lα phase, associated with uncomplexed structures.
Concerning the XRD complexes measurements, associated to the complexes with MgCl2 and CaCl2, the lamellar repeat distance for the complexed phase was about dc = 7.39 nm and d = 5.69 nm for the uncomplexed one. From XRD patterns reported in Fig. 11 the formation of the complexes with DNA is confirmed in both systems. In particular, Fig. 11A shows that the addition of CHOLp in the lipid mixtures in presence of CaCl2 improves the formation of the complexed phase with respect to the DOPC/CHOLp (0.25)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DNA
DNA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MgCl2, as deduced from the intensities ratio between the complexed and uncomplexed phase.
MgCl2, as deduced from the intensities ratio between the complexed and uncomplexed phase.
Conclusions
From the combination of experimental methods such as X-ray diffraction and in silico modeling techniques, we characterized the structural properties and phase behaviour of mixed composition zwitterionic liposomes containing DOPC and the newly synthesized lipid, CHOLp, at different molar fraction. The data obtained suggest a good insertion of CHOLp into DOPC bilayers and a stable steady state of the overall bilayers resulting in a stable average lipid area. In pure water, there is a high organization of the CHOLp lipid tails at low molar fractions (XCHOLp = 0.05, 0.15), which confirm a good insertion of the functionalized cholesterol in DOPC based liposomes. At higher molar concentration (XCHOLp = 0.25), the intramolecular aromatic π stacking interactions in CHOLp are dominant. This behaviour is also observed experimentally from X-ray diffraction data that showed an increased splitting of the Bragg peaks as a function of CHOLp molar ratio, which suggests a DOPC bilayer with regions more or less rich in CHOLp. The addition of the metal ions increases the stability of the steroid CHOLp in the bilayer. In fact, the CHOLp polar head is positioned in the aqueous environment like the zwitterionic DOPC head and it assumes an extended conformation in order to maximize the number and the type of the intermolecular interactions both with cations and other steroid molecules. This is particularly evident in CaCl2. Moreover, from the preliminary SAXS experiments, the addition of CHOLp among the phospholipid molecules increases the capability to complex DNA in the presence of metal ions and could provide a basis for employing these systems as DNA carriers. In silico computational simulations on these bilayers in complex with DNA will be carried out to further assess this behaviour.Acknowledgements
This work was supported by Genzyme and Cariverona Foundation (prot.2012.0020). We would like to thank the CINECA Supercomputing Center and PRACE Research Infrastructure for providing computer time on its IBM PLX supercomputer (CINECA-ISCRA C 2012–2013 and PRACE DECI-9 HPC grant 2012–2013).References
- D. A. Balazs and W. T. Godbey, J. Drug Delivery, 2011, 2011, 326497 CrossRef PubMed.
- D. D. Lasic, Liposomes in Gene Delivery, CRC Press, 1997 Search PubMed.
- P. Bruni, M. Pisani, A. Amici, C. Marchini, M. Montani and O. Francescangeli, Appl. Phys. Lett., 2006, 88, 73901 CrossRef.
- O. Francescangeli, M. Pisani, V. Stanic, P. Bruni and T. M. Weiss, Europhys. Lett., 2004, 67, 669 CrossRef CAS.
- P. Bruni, O. Francescangeli, M. Marini, G. Mobbili, M. Pisani and A. Smorlesi, Mini-Rev. Org. Chem., 2011, 8(1), 38 CrossRef CAS.
- M. Pisani, G. Mobbili, I. F. Placentino, A. Smorlesi and P. Bruni, J. Phys. Chem. B, 2011, 115(34), 10198 CrossRef CAS PubMed.
- G. Angelini, M. Pisani, G. Mobbili, M. Marini and C. Gasbarri, Biochim. Biophys. Acta, Biomembr., 2013, 1828(11), 2506 CrossRef CAS PubMed.
- M. Alwarawrah, J. Dia and J. Huang, J. Phys. Chem. B, 2010, 114, 7516 CrossRef CAS PubMed.
- R. Kaur, M. Henriksen-Lacey, J. Wilkhu, A. Devitt, D. Christensen and Y. Perrie, Mol. Pharmaceutics, 2014, 11, 197 CrossRef CAS PubMed.
- G. Khelashvili and D. Harries, J. Phys. Chem. B, 2013, 117, 2411 CrossRef CAS PubMed.
- T. A. Daly, M. Wang and S. L. Regen, Langmuir, 2011, 27, 2159 CrossRef CAS PubMed.
- M. Sugahara Uragami, X. Yan and S. L. Regen, J. Am. Chem. Soc., 2001, 123, 7939 CrossRef.
- A. Zidovska, H. M. Evans, A. Ahmad, K. K. Ewert and C. R. Safinya, J. Phys. Chem. B, 2009, 113(15), 5208 CrossRef CAS PubMed.
- T. Róg, M. Pasenkiewicz-Gierula, I. Vattulainen and M. Karttunen, Biochim. Biophys. Acta, 2009, 1788, 97 CrossRef PubMed , and reference therein.
- F. de Meyer and B. Smit, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 3654 CrossRef CAS PubMed.
- S. J. Weiner, P. A. Kollman, D. T. Nguyen and D. A. Case, J. Comput. Chem., 1986, 7, 230 CrossRef CAS.
- F. Mohamadi, N. G. J. Richards, W. C. Guida, R. Liskamp, M. Lipton, C. Caufield, G. Chang, T. Hendrickson and W. C. Still, J. Comput. Chem., 1990, 11, 440 CrossRef CAS.
- J. M. Goodman and W. C. Still, J. Comput. Chem., 1991, 12, 1110 CrossRef CAS.
- W. C. Still, A. Tempczyk, R. C. Hawley and T. Hendrickson, J. Am. Chem. Soc., 1990, 12, 6127–6129 CrossRef.
- J. V. Ryzin, Classification and clustering, Academic Press, New York, 1977 Search PubMed.
- J. Zupan, Algorithms for Chemists, Wiley-VCH, New York, 1989, ch. 7 Search PubMed.
- R. Galeazzi, G. Martelli, E. Marcucci, G. Mobbili, D. Natali, M. Orena and S. Rinaldi, Eur. J. Org. Chem., 2007, 26, 4402 CrossRef.
- R. Galeazzi, G. Martelli, E. Marcucci, M. Orena, S. Rinaldi, R. Lattanzi and L. Negri, Amino Acids, 2010, 38(4), 1057 CrossRef CAS PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, E. Brothers, K. N. Kudin, V. N. Staroverov, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, N. J. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, Ö. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian, Inc., Wallingford CT, 2009.
- A. D. Becke, J. Chem. Phys., 1993, 98, 5648 CrossRef CAS.
- C. Lee, W. Yang and R. G. Parr, Phys. Rev. B: Condens. Matter Mater. Phys., 1988, 37(2), 37785 CrossRef.
- S. H. Vosko, L. Wilk and M. Nusair, Can. J. Phys., 1980, 58, 1200 CrossRef CAS.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, J. Phys. Chem. B, 1994, 98, 11623 CrossRef CAS.
- J. B. Klauda, R. M. Venable, J. A. Freites, J. W. O'Connor, C. Mondragon-Ramirez, I. Vorobyov, D. J. Tobias, A. D. MacKerell and R. W. Pastor, J. Phys. Chem. B, 2010, 114, 7830 CrossRef CAS PubMed.
- A. K. Soper, F. Bruni and M. A. Ricci, J. Chem. Phys., 1997, 106(1), 247–254 CrossRef CAS.
- D. Y. Villanueva, J. B. Lim and J. B. Klauda, Langmuir, 2013, 29, 14196 CrossRef CAS PubMed.
- S. H. Jo, H. Rui, J. B. Lim, J. B. Klauda and W. Im, J. Phys. Chem. B, 2010, 114, 13342 CrossRef CAS PubMed.
- J. B. Klauda, B. R. Brooks and R. W. Pastor, J. Chem. Phys., 2006, 125, 144710 CrossRef PubMed.
- M. J. S. Dewar, E. G. Zoebisch, E. F. Healy and J. J. P. Stewart, J. Am. Chem. Soc., 1985, 107(13), 3902 CrossRef CAS.
- T. Darden, D. York and L. Pederson, J. Chem. Phys., 1993, 98, 10089 CrossRef CAS.
- P. Ewald, Ann. Phys., 1921, 64, 253 CrossRef.
- R. Hockney and J. Eastwood, Computer simulation using particles, McGraw-Hill, New York, 1981 Search PubMed.
- S. Pronk, S. Pall, R. Schulz, P. Larsson, P. Bjelkmar, R. Apostolov, M. R. Shirts, J. C. Smith, P. M. Kasson, D. van der Spoel, B. Hess and E. Lindahl, Bioinformatics, 2013, 29, 845 CrossRef CAS PubMed.
- J. S. Weiner, P. A. Kollman, D. A. Case, U. C. Singh, C. Ghio, G. Alagona, S. Profeta and P. A. Weiner Jr, J. Am. Chem. Soc., 1984, 106, 765 CrossRef.
- J. Eastwood, R. Hockney and D. Lawrence, Comput. Phys. Commun., 1980, 19, 215 CrossRef CAS.
- R. Galeazzi, Curr. Comput.-Aided Drug Des., 2009, 5, 225 CrossRef CAS.
- R. D. Porasso and J. J. Lopez Cascales, Pap. Phys., 2012, 4, 040005 Search PubMed.
- W. Humphrey, A. Dalke and K. J. Schulten, J. Mol. Graphics, 1996, 14, 33 CrossRef CAS PubMed.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng and T. E. Ferrin, J. Comput. Chem., 2004, 25(13), 1605 CrossRef CAS PubMed.
- S. Tristam-Nagle, H. I. Petrache and J. F. Nagle, Biophys. J., 1998, 75(2), 917 CrossRef.
- L. J. Lis, V. A. Parsegian and R. P. Rand, Biochemistry, 1981, 20, 1761 CrossRef CAS PubMed.
- J. J. Potoff, Z. Issa, C. W. Manke Jr and B. P. Jena, Cell Biol. Int., 2008, 32, 361 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c5ra08439c |
| This journal is © The Royal Society of Chemistry 2015 |