Graphene oxide regulates the bacterial community and exhibits property changes in soil†
Junjie Du,
Xiangang Hu* and
Qixing Zhou*
Key Laboratory of Pollution Processes and Environmental Criteria (Ministry of Education), Tianjin Key Laboratory of Environmental Remediation and Pollution Control, College of Environmental Science and Engineering, Nankai University, Tianjin 300071, China. E-mail: huxiangang@nankai.edu.cn; zhouqx@nankai.edu.cn
First published on 11th March 2015
Abstract
The extensive use of pristine graphene oxide (PGO) increases its environmental release. The interactions of PGO with soils, one of the ultimate repositories for discharged nanomaterial, remain unclear. In the present study, a pyrosequencing analysis based on the bacterial 16S rRNA gene showed that the bacterial community in a PGO-soil sample (PGOS) became richer and more diverse compared with a control soil sample (CS). PGO altered the structure of soil bacterial communities, with some nitrogen-fixing and dissimilatory iron reducing bacteria being selectively enriched, especially at the genus level. SGO (soil-modified graphene oxide) exhibited a greater thickness, a higher C/O ratio, a rougher texture, a lower transparency and a smaller size than PGO. Nitrogen-containing groups and the elements including Mg, Al, Si, K, Ca and Fe were detected in SGO. The changes in surface groups were consistent with the formation of organic molecules coating the SGO. SGO, which exhibited fewer negative charges, was more unstable than PGO. In addition, SGO presented higher chemical activity than PGO; for example, SGO exhibited more unpaired electrons and disordered structures. This work highlights the critical interactions of PGO and soil which deserve comprehensive consideration in assessing the risks of nanomaterials.
1. Introduction
Graphene oxide (GO) which presents the advantage of carrying oxygen-containing groups, is being promoted as a useful precursor to graphene-based nanomaterial synthesis in various fields.1–4 The rapidly increasing production and use of GO will increase the possibility of its environmental release.5–7 Soil ecosystems are likely to be the largest recipient of nanomaterial contamination, in comparison with water and air.8–10 The nascent state of the nanoproduct industry calls for critical early assessment of environmental impacts before significant releases occur.11 PGO (pristine graphene oxide) could potentially alter the structure and function of soil microbial communities; at the same time, the properties of PGO might undergo changes as a result of soil interactions, such as biomodification by soil microorganisms, and heteroaggregation with soil minerals.12 Therefore, understanding the interactions between PGO and soil components as well as the formation of soil-modified GO (SGO) is critical for assessing the environmental fate and risks of PGO.Fullerenes (C60) and carbon nanotubes (CNTs) have limited effects on soil microbial communities at low concentrations11,13 but exert observable effects under exposure to very high concentrations, such as 5000 mg kg−1.8,14 However, the effects of PGO on soil microbial communities are still not clear. Thus far, the majority of studies on the impact of PGO on soil microorganisms have been conducted in pure cultures or aquatic environments. For example, researchers observed the antimicrobial activity of PGO in pure cultures and in engineered aquatic systems.2,15–17 It was reported that the number of oxygen-containing functional groups in PGO was decreased by 60% via interaction with Escherichia coli.18 Reduction of PGO by the microbe Shewanella and fungi was also verified.3,19,20 Additionally, it was shown that microorganisms were able to alter the properties of PGO via nitrogen doping by denitrifying bacteria.21 However, the investigation of soil modifications exerted on PGO cannot simply be conducted in an aquatic solution or pure culture because soil is an extremely complex matrix with numerous microorganism communities. In particular, soil microbial communities are a sensitive indicator of the soils' response to environmental stressors, and the effects of PGO on the structure and function of microbial communities are obscure.8,14,22 In addition, changes in microbial communities and modifications of PGO should occur simultaneously in soil. The various soil microorganisms could biomodify GO during its interaction with PGO. Biomodifications driven by single microbes cannot truly reflect the characteristic alterations of PGO in a natural environment. The isolation and characterization of soil-modified GO (SGO) from an actual environmental scenario is therefore necessary to investigate the characteristic alterations of PGO.
To verify the above assumption and fill this research gap, the present study focuses on the interactions between PGO and soil. The changes in the bacterial community were first investigated in soil after aging for 90 days through high-throughput sequencing analysis. Subsequently, the morphological and surface chemical alterations and the organic molecule coating of SGO were characterized. Finally, the size distribution, surface charges, unpaired electrons, disordered structures, UV-vis adsorption and redox potential of PGO and SGO were analyzed to reveal the changes in chemical activity.
2. Materials and methods
2.1. Preparation and collection of samples
2.2. Bacterial community analysis
2.3. Identification of the organic molecules immobilized on SGO
Using 1 mL of solution (with a volume ratio of methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) chloroform
chloroform![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DI water = 2.5
DI water = 2.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1; methanol and chloroform were of chromatographic grade) as a solvent, the organic molecules were extracted from 0.01 g of SGO. The organic molecules were intensively extracted through microwave-assisted extraction (at 40 °C and 400 W for 15 min), followed by centrifugation for 10 min at 11
1; methanol and chloroform were of chromatographic grade) as a solvent, the organic molecules were extracted from 0.01 g of SGO. The organic molecules were intensively extracted through microwave-assisted extraction (at 40 °C and 400 W for 15 min), followed by centrifugation for 10 min at 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g. The supernatant was collected. The sediment was extracted a second time using the same solvent and procedure described above. The second supernatant was mixed with the first supernatant, and water (500 μL) was added to the supernatant mixture, which was then centrifuged at 5000g for 3 min. The lower layer was evaporated under nitrogen flux. For the upper layer, methanol was evaporated under nitrogen flux, and water was then lyophilized. Methoxamine hydrochloride (20 mg mL−1, 50 μL) and N-methyl-N-(trimethylsilyl) trifluoroacetamide (80 μL) were used as derivatives. A sample (1 μL) was injected into a gas chromatography (GC) column in a split mode (1
000g. The supernatant was collected. The sediment was extracted a second time using the same solvent and procedure described above. The second supernatant was mixed with the first supernatant, and water (500 μL) was added to the supernatant mixture, which was then centrifuged at 5000g for 3 min. The lower layer was evaporated under nitrogen flux. For the upper layer, methanol was evaporated under nitrogen flux, and water was then lyophilized. Methoxamine hydrochloride (20 mg mL−1, 50 μL) and N-methyl-N-(trimethylsilyl) trifluoroacetamide (80 μL) were used as derivatives. A sample (1 μL) was injected into a gas chromatography (GC) column in a split mode (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5). GC (Agilent 6890N, Agilent Technologies, USA) combined with quadrupole mass spectrometry (MS) (Agilent 5973, Agilent Technologies, USA) was used to analyze the organic molecules. GC separation was achieved in a DB-5 MS capillary column (30 m, 0.25 mm i.d., 0.1 μm film thickness). The injection temperature was 230 °C, and both the transfer line and the ion source were maintained at 250 °C. The spectrometer was operated in electron-impact mode. The detection voltage was 2100 V. The full scan range was from 60 to 800 amu. Helium, as the carrier gas, was maintained at a constant flow rate of 2 mL min−1. The oven temperature was maintained at 80 °C for 2 min and was then increased to 320 °C at a rate of 15 °C min−1 and held for 6 min. The organic molecules were identified using the NIST 08 library.
5). GC (Agilent 6890N, Agilent Technologies, USA) combined with quadrupole mass spectrometry (MS) (Agilent 5973, Agilent Technologies, USA) was used to analyze the organic molecules. GC separation was achieved in a DB-5 MS capillary column (30 m, 0.25 mm i.d., 0.1 μm film thickness). The injection temperature was 230 °C, and both the transfer line and the ion source were maintained at 250 °C. The spectrometer was operated in electron-impact mode. The detection voltage was 2100 V. The full scan range was from 60 to 800 amu. Helium, as the carrier gas, was maintained at a constant flow rate of 2 mL min−1. The oven temperature was maintained at 80 °C for 2 min and was then increased to 320 °C at a rate of 15 °C min−1 and held for 6 min. The organic molecules were identified using the NIST 08 library.
2.4. Characterization of PGO and SGO
The structures and surface morphologies of PGO and SGO were characterized via scanning electron microscopy (SEM), field-emission transmission electron microscopy (TEM) and atomic force microscopy (AFM). The PGO and SGO were dispersed in absolute ethanol, and SEM, TEM and AFM observations were then performed using a LEO-1530VP instrument (Germany), a JEM-2010 FEF instrument (Japan) and a Veeco Nanoscope 4 (USA), respectively. The size distribution and zeta potential of the nanosheets were measured using a ZETAPALS/BI-200SM instrument equipped with a 30 mW, 635 nm laser (Brookhaven Instruments Corporation, Holtsville, NY, USA). The surface chemistry was analyzed via X-ray photoelectron spectroscopy (XPS) and Fourier transform infrared spectroscopy (FTIR). XPS measurements were performed using an Axis Ultra XPS system (Kratos, Japan) with a monochromatic Al Kα X-ray source (1486.6 eV). The XPS spectra were analyzed using Casa-XPS V2.3.13 software. FTIR spectra were recorded with a Bruker Tensor 27 (Germany) infrared spectrometer with a resolution of 2 cm−1 at 4000–400 cm−1. UV-vis spectra were collected using a TU-1901 spectrophotometer (Beijing Purkinje, China) and were subsequently analyzed using UVWin5 software. Raman spectra were obtained in a Renishaw in Via Raman spectrometer (Renishaw plc, UK) with a 514 nm laser (Thermo Scientific, DXR, USA). Eh values were measured using an oxidation–reduction potentiometer (Sinomeasure, Hang Zhou, China).2.5. EPR measurements
To measure unpaired electrons, which are related to the chemical activity of nanomaterials, electron paramagnetic resonance (EPR) was performed. All X-band EPR spectra were collected at room temperature (296 K) using a Magnettech MiniScope 400 EPR spectrometer (Germany) operated at a microwave frequency of 9.4 GHz and a magnetic field modulation frequency of 100 kHz. The spectrometer was controlled using MiniScope Control software. As an unpaired electron probe, 2,6,6-tetramethyl-1-piperidinyloxy (TEMPO) was employed. PGO and SGO at a concentration of 1 mg L−1 were placed in individual quartz EPR tubes. Before the samples were loaded, the quartz EPR tubes were washed thoroughly using ultrapure water (18.2 Ω cm−1) and then dried.2.6. Data analysis and statistical analysis
To process the sequencing data, we clustered sequences into operational taxonomic units (OTUs) with a 97% similarity cutoff using UPARSE (version 7.1, http://drive5.com/uparse/) and removed chimeric sequences using UCHIME (http://www.mothur.org/wiki/Chimera.uchime). On the basis of cluster information, rarefaction curves, the richness estimators of Chao (http://www.mothur.org/wiki/Chao), the abundance coverage estimator (ACE) (http://www.mothur.org/wiki/Ace), and the Shannon diversity index (http://www.mouthur.org/wiki/Shannon) were obtained in MOTHUR (http://www.mothur.org/) for each sample. The phylogenetic affiliation of each 16S rRNA gene sequence was analyzed using the RDP Classifier (http://rdp.cme.msu.edu/) against the silva (SSU115) 16S rRNA database with a confidence threshold of 70%.24 After taxonomic classification of the sequences down to the phylum, class, order, family, and genus level, the relative abundance of a given phylogenetic group was set as the number of sequences affiliated with that group divided by the total number of sequences per sample. Hierarchical cluster analysis was performed using R Project software (http://www.r-project.org/). A Venn diagram with shared and unique OTUs was drawn to depict the similarities and differences among samples.25All treatments were performed in triplicate, and the error bars presented in the results represent the standard deviation (mean ± SD). The standard deviation values were calculated using IBM Statistics SPSS 19 software.
3. Results and discussion
3.1. Richness and diversity of bacterial communities
Two 16S rRNA gene libraries were constructed via high-throughput sequencing of the bacterial communities from the PGO-soil sample (PGOS) and the control soil sample (CS), generating 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 882 and 15
882 and 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 651 high-quality reads, respectively. There were 702 and 691 operational taxonomic units (OTUs) found in PGOS and CS, respectively, at a 3% distance. As listed in Table S2,† the total numbers of OTUs estimated based on the Chao index were 731 (PGOS) and 713 (CS) with infinite sampling at a 3% distance. Both indicators (OTUs and the Chao index) suggested that the richness of the soil bacterial community slightly increased after the introduction of PGO into the soil, which was confirmed by the rarefaction curves shown in Fig. S2a.† The Shannon index, which indicates the diversity of the bacterial population (such as species richness and species evenness),25 also reflected a slight increase from 5.12 (CS) to 5.15 (PGOS), as shown in Fig. S2b.† Among the 764 OTUs observed in both communities, 629 OTUs were shared by the two communities, while 73 OTUs were unique to PGOS, and 62 OTUs were unique to CS (Fig. S3a†). The Venn diagram that depicted the commonalities and differences between the two sets of OTUs also demonstrated that the bacterial communities in PGOS were richer and more diverse than those in CS. Through taxonomic identification of the unique OTUs at the phylum level (Fig. S3b and c†), it was found that the majority of the unique OTUs were assigned to common phyla, i.e., Proteobacteria, Chloroflexi, Bacteroidetes, Planctomycetes, Gemmatimonadetes, Acidobacteria, Actinobacteria and Verrucomicrobia. The total common phyla in PGOS and CS accounted for 89.04% and 82.26% of the OTUs, respectively.
651 high-quality reads, respectively. There were 702 and 691 operational taxonomic units (OTUs) found in PGOS and CS, respectively, at a 3% distance. As listed in Table S2,† the total numbers of OTUs estimated based on the Chao index were 731 (PGOS) and 713 (CS) with infinite sampling at a 3% distance. Both indicators (OTUs and the Chao index) suggested that the richness of the soil bacterial community slightly increased after the introduction of PGO into the soil, which was confirmed by the rarefaction curves shown in Fig. S2a.† The Shannon index, which indicates the diversity of the bacterial population (such as species richness and species evenness),25 also reflected a slight increase from 5.12 (CS) to 5.15 (PGOS), as shown in Fig. S2b.† Among the 764 OTUs observed in both communities, 629 OTUs were shared by the two communities, while 73 OTUs were unique to PGOS, and 62 OTUs were unique to CS (Fig. S3a†). The Venn diagram that depicted the commonalities and differences between the two sets of OTUs also demonstrated that the bacterial communities in PGOS were richer and more diverse than those in CS. Through taxonomic identification of the unique OTUs at the phylum level (Fig. S3b and c†), it was found that the majority of the unique OTUs were assigned to common phyla, i.e., Proteobacteria, Chloroflexi, Bacteroidetes, Planctomycetes, Gemmatimonadetes, Acidobacteria, Actinobacteria and Verrucomicrobia. The total common phyla in PGOS and CS accounted for 89.04% and 82.26% of the OTUs, respectively.
3.2. Taxonomic composition of bacterial communities
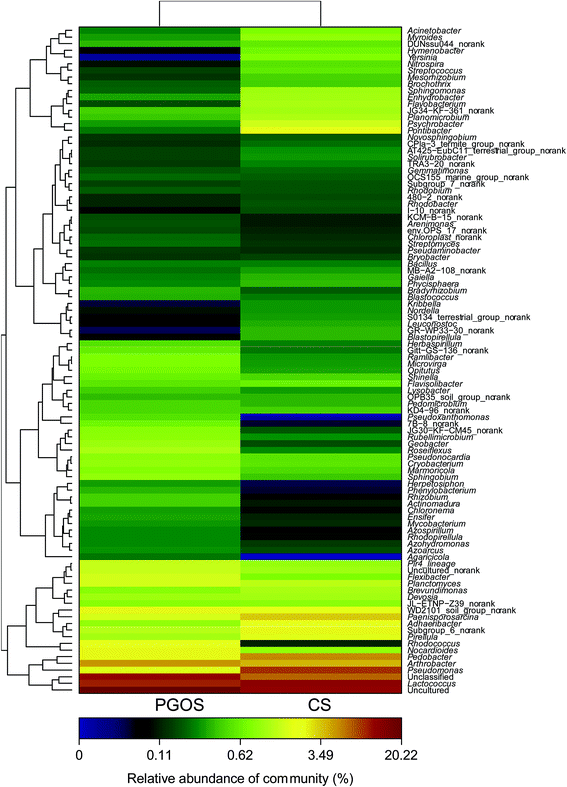
To characterize the changes in the bacterial communities driven by PGO, the bacterial OTUs were assigned to phyla, classes and genera, as shown in Fig. 1. The dominant phyla, classes and genera were shared. At the phylum level, the 8 known phyla out of the 10 total phylotypes (i.e., Proteobacteria, Chloroflexi, Bacteroidetes and Planctomycetes) were common to the PGOS and CS samples, which accounted for 89.40% and 95.41%, respectively. At the class level, the 15 known classes out of the total 18 phylotypes (i.e., Acidobacteria, Actinobacteria, Alphaproteobacteria, Anaerolineae and Bacilli) were common to the PGOS and CS samples and accounted for 83.34% and 89.85%, respectively. At the genus level, the 8 known genera out of the total 19 phylotypes (i.e., Arthrobacter, Devosia and Lactococcus) were common and accounted for 23.03% and 36.00% in the PGOS and CS samples, respectively. There were no large differences found in the bacterial communities between PGOS and CS at the phylum and class levels; however, a clear difference between the two communities was observed at the genus level.Among the bacterial genera that only appeared in PGOS (and not in CS), Nocardioides (2.41%) and Rhodococcus (2.25%) are Gram-positive, and Roseiflexus (1.11%) and Flexibacter (1.57%) are Gram-negative. The bacterial genera that were absent from PGOS (but present in CS) also included both Gram-positive genera (Planomicrobium (1.12%)) and Gram-negative genera (Adhaeribacter (2.26%), Brevundimonas (1.09%), Enhydrobacter (1.02%), Psychrobacter (1.49%), Flavobacterium (1.10%) and Pontibacter (1.83%)). These findings showed that the changes in bacteria after PGO entered the soil were species dependent rather than Gram dependent.
To further investigate the structural and functional changes in the bacterial communities, a more detailed cluster analysis was conducted at the genus level, as listed in Fig. 2. The known PGOS bacterial genera exhibited a higher relative abundance than those in CS and belonged to more diverse phyla. The identified Gram-positive genera, such as Rhodococcus, Nocardioides and Actinomadura, belonged to the phylum Actinobacteria, and the Gram-negative genera were assigned to other phyla (i.e., Proteobacteria, Chloroflexi, Verrucomicrobia, Bacteroidetes and Planctomycetes). Among the PGOS bacterial genera showing a lower abundance than in CS, the Gram-positive genera were assigned to Actinobacteria (Kribbella and Solirubrobacter) and Firmicutes (Brochothrix, Streptococcus and Planomicrobium), and the Gram-negative genera were assigned to Proteobacteria and Planctomycetes. The Gram-positive genera displaying a decreased abundance belonged to Actinobacteria and Firmicutes, while the Gram-positive genera exhibiting an increased abundance only belonged to Actinobacteria. Therefore, the changes in bacterial population abundance were also bacterial species dependent, rather than Gram dependent. A recent report suggested that the antibacterial activity of graphene-family nanomaterials is bacterial species dependent rather than Gram dependent in the aquatic environment as well.26 The present investigation reports the first similar results in the soil environment.
In addition, the genera exhibiting nitrogen-fixing functions (e.g., Azoarcus, Azospirillum, Bradyrhizobium, Herbaspirillum, Ensifer, Microvirga and Rhizobium) all belong to the phylum Proteobacteria, which presented a higher abundance in the PGOS community. Some of the nitrogen-fixing bacteria found in PGOS, such as Azospirillum and Azoarcus, have the capacity to dissimilate nitrate to nitrite or nitrous oxide and nitrogen gas under severe oxygen limitation. The relative abundance of the genus Geobacter also was increased in PGOS; this genus is composed of dissimilatory iron reducing bacteria. Geobacter also belongs to phylum Proteobacteria. Taken together, our results show that PGO can alter the soil bacterial community, which was selectively enriched in the soil after the introduction of PGO, especially at the genus level.
3.3. Changes in surface morphology
The changes in the microbial communities mentioned above and the modifications of PGO mentioned subsequently are expected to occur simultaneously in soil. SEM, TEM and AFM were conducted to study the morphology of PGO before and after the soil modifications. The SEM images presented in Fig. S4a and b† show that both PGO and SGO exhibited a nanosheet morphology; however, SGO exhibited a rougher texture than PGO, in which some particles were distributed on the surface and embedded. In the TEM images presented in Fig. S4c and d,† the transparency of SGO is lower than that of PGO, and black spots with uneven sizes appear on and in the SGO nanosheets. The black spots were further analyzed using EDS as shown in Fig. S4e.† The chemical compositions of the black spots include C and O, which are the essential elements found in PGO, as well as N, Mg, Al, Si, K, Ca and Fe, which originated from the soil. As shown in the AFM image presented in Fig. S5,† PGO exhibited a layer morphology with a thickness of 0.862 nm, consistent with the single-layer GO.27 By contrast, SGO exhibited uneven nanosheets with thicknesses of 3.562 nm and 7.162 nm at the thinner and thicker locations, respectively. For the SGO nanosheets, the thicker locations in the AFM image correspond to the black spots in the TEM images and the particles in the SEM images.3.4. Changes in surface chemical groups
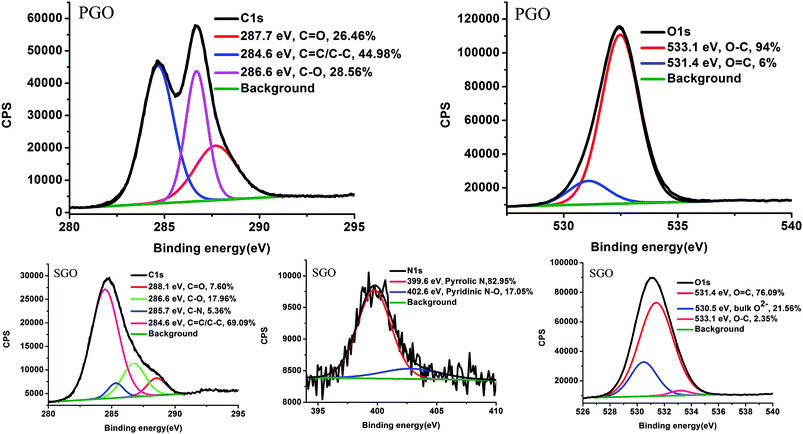
The surface chemistry of PGO and SGO was investigated via XPS. The full spectra of PGO and SGO are presented in Fig. S6.† The spectrum of PGO consisted of 69.73% C1s and 30.27% O1s, whereas the spectrum of SGO consisted of 41.35% C1s, 44.15% O1s and 1.32% N1s as well as a 13.18% contribution of signals from other elements (i.e., Mg, Al, Si, K, Ca and Fe), which is consistent with the results of the analysis of the black spots using EDS shown in Fig. S4e.† These inorganic elements originated from the soil minerals. It was reported that PGO could adsorb minerals by electrostatic attraction.12 Fig. 3 presents the specific groups for the above organic elements C, O and N. The C1s spectrum of PGO revealed that 44.98% of carbon (C–C/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) was not oxidized, while 28.56% presented C–O bonds (representing hydroxyl and epoxide groups), and 26.46% exhibited C
C) was not oxidized, while 28.56% presented C–O bonds (representing hydroxyl and epoxide groups), and 26.46% exhibited C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds. The C1s spectrum of SGO was found to be a composite of C–C/C
O bonds. The C1s spectrum of SGO was found to be a composite of C–C/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C (69.09%), C–O (17.96%), C
C (69.09%), C–O (17.96%), C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O (7.60%) and C–N (5.36%), which was consistent with the groups detected using FTIR spectra presented in Fig. S7.† The numbers of C–O and C
O (7.60%) and C–N (5.36%), which was consistent with the groups detected using FTIR spectra presented in Fig. S7.† The numbers of C–O and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O functional groups in PGO decreased notably (from 28.56% to 17.96% and 26.46% to 7.60%, respectively) during the aging process in soil. In the literature, it has been reported that the oxygen-containing functional groups of PGO are decreased via interaction with the environmental bacteria Escherichia coli18 and Shewanella.3,19,28 The mechanism underlying the bacterial transformation of PGO involves reduction during bacterial respiration for Shewanella and glycolysis for Escherichia coli, and PGO acts as a terminal electron acceptor in the reduction processes.26 In addition, reduction of PGO by other microbes, such as yeast and endophytic microorganisms, have also been reported.20,29 Soil is a potential repository of microorganisms, and soil microorganisms are widely distributed among a variety of species, possessing different metabolic pathways. The reduction of PGO during its interaction with soil microorganisms could be ubiquitous via aging in soil. Additionally, methylene was detected (at 2924 and 2851 cm−1) from the FTIR spectra presented in Fig. S7,† demonstrating the introduction of a carbon chain on SGO, which could increase the ratio of C/O.
O functional groups in PGO decreased notably (from 28.56% to 17.96% and 26.46% to 7.60%, respectively) during the aging process in soil. In the literature, it has been reported that the oxygen-containing functional groups of PGO are decreased via interaction with the environmental bacteria Escherichia coli18 and Shewanella.3,19,28 The mechanism underlying the bacterial transformation of PGO involves reduction during bacterial respiration for Shewanella and glycolysis for Escherichia coli, and PGO acts as a terminal electron acceptor in the reduction processes.26 In addition, reduction of PGO by other microbes, such as yeast and endophytic microorganisms, have also been reported.20,29 Soil is a potential repository of microorganisms, and soil microorganisms are widely distributed among a variety of species, possessing different metabolic pathways. The reduction of PGO during its interaction with soil microorganisms could be ubiquitous via aging in soil. Additionally, methylene was detected (at 2924 and 2851 cm−1) from the FTIR spectra presented in Fig. S7,† demonstrating the introduction of a carbon chain on SGO, which could increase the ratio of C/O.
Compared with PGO, the new observation of N1s signals suggested that nitrogen-containing groups had doped on the SGO surface during the aging process in soil. The bonding configurations of nitrogen atoms of SGO were characterized by N1s spectrum. The N1s spectrum of SGO could be fitted to two peaks at 399.6 and 402.6 eV, corresponding to pyrrolic N and pyridinic N–O bonds, respectively, which were all immobilized on the plane, edges or defects of the graphene networks, rather than having been incorporated into the graphene networks.30,31 For the SGO spectra, the pyridinic N–O bond in the N1s spectrum could have originated from nitrate or nitrite (both of which are widely found in the soil), which was consistent with the results regarding bulk O2− in the O1s spectrum of SGO. The pyrrolic N in the N1s spectrum could have originated from organic nitrogen compounds in the soil and soil microbial metabolic processes, such as denitrification.21
3.5. Identification of organic molecules immobilized on SGO
To investigate the compounds consisting of organic molecules immobilized on SGO, GC-MS/MS was conducted, and 14 organic molecules were identified, as shown in Fig. S8.† These organic molecules included organic acids, phenols, benzene, esters, amides, alkanes and carbonyls, which originated from natural organic molecules or microorganisms in the soil. For example, 8 out of the 14 organic molecules could been detected from the raw soil sample, including phenol, octacosane, hexanoic acid, hexadecanoic acid, undecane, octadecanoic acid, benzene and benzaldehyde, which were presented in Table S3.† PGO contains many oxygen groups, such as –OH, –COOH and –CHO, and specific nanostructures, such as nanopores, active edges, dangling carbons, and sp2 structures.32,33 The aforementioned organic molecules with carboxyl groups, phenolic hydroxy groups, or benzene rings could bind to PGO via hydrogen bonds and π–π bonds. The immobilized organic molecules explained the changes in the C/O ratio. The observation of pentanamide supports the observation of N1s signals through XPS analysis.3.6. Changes in size distribution and surface charges
The size distribution and the surface charges play dominant roles in the environmental behavior of nanomaterials and their associated ecological risks.34–36 As shown in Fig. S9a,† the size distribution of PGO ranged from 342 nm to 532 nm, and the size distribution of SGO ranged from 142 nm to 396 nm. These data demonstrated that the size of PGO decreased during the aging process in soil. It has been reported that PGO can be biodegraded by environmental microorganisms and horseradish peroxidase (HRP).26,37 In soil environments, similar PGO biodegradation may also take place, carried out by a many species of soil microorganisms that secrete various enzymes, resulting in a change in the size distribution. The zeta potential as a function of pH is presented in Fig. S9b.† The surface charges of PGO and SGO were both negative at environmentally relevant pH values (pH 5–9). The results regarding the zeta potential demonstrated that PGO, which presents a greater number of negative charges, was more stable than SGO.3.7. Changes in chemical activity and structures
The unpaired electrons on the surfaces of nanomaterials, which are associated with defects, directly affect the chemical activity and nanotoxicity of nanomaterials.33,36,38 As shown in Fig. 4a, both PGO and SGO exhibited more intense signals than the control without nanomaterials, suggesting that PGO and SGO had the potential to produce unpaired electrons. SGO exhibited slightly more unpaired electrons than PGO, suggesting that the chemical activity of SGO was enhanced following the soil modifications. Unpaired electrons are known to be associated with disordered structures (such as dangling bonds, turns and kinks on edges or inner planes) in carbon-based nanomaterials.34,39 The disordered structures of PGO and SGO were assessed based on the Raman spectra, as shown in Fig. 4b. The D band at 1363 cm−1 is usually related to the vibrations of sp3 carbon atoms, which induces defects and disorders, while the G band at 1593 cm−1 is ascribed to the stretching vibration of sp2 carbon atoms.40,41 The intensity ratio (1.44) of the D band to the G band (D/G) of SGO was higher than that (1.15) of PGO, suggesting that the disordered structure of PGO was enhanced after the soil modifications. This result was consistent with the observed enhancement of unpaired electrons in SGO.The UV-vis absorption spectra were obtained to reflect the photochemical activity of PGO and SGO, as presented in Fig. 4c. The largest peaks in the absorption spectra for PGO and SGO, located at 203 nm and 196 nm, respectively, reflect the n–π* and π–π* transitions (E2 band).42 The largest peak in the SGO spectrum, at 196 nm, was markedly higher than that in the PGO spectrum; this peak corresponds to –COOH, –NH–, unsaturated ketone groups and unpaired electrons from the organic molecules immobilized on SGO. Compared with PGO, the largest peak in the absorption spectra of SGO was shifted from 203 nm to 196 nm, suggesting that the electron concentration had decreased due to an increase in sp3 carbon atoms, which reflects the enhanced structural disorder of SGO.43 Furthermore, compared with PGO, the UV-vis absorption of SGO decreased remarkably in the region between 205 and 800 nm, suggesting that the electronic conjugation within the graphene sheets was disrupted upon the immobilization of the chemical compounds from soil.44
As one of the indicators of the chemical activity of nanomaterials, Eh as a function of pH is presented in Fig. 4d. The Eh values for PGO and SGO both decreased with an increasing pH. The Eh values for SGO were higher than those for PGO in the pH range of 3 to 11. Due to aging in soil, the reduced size of PGO (Fig. S9a†) could enhance its redox potential and chemical activity.34 Furthermore, through the observed soil modifications, the introduction of new chemical groups, such as the N–O bond configuration (Fig. 3, N1s spectrum), could cause SGO to present a higher redox potential than PGO.
4. Conclusion
The soil bacterial communities became richer and more diverse due to the exposure of PGO. Some nitrogen fixing and dissimilatory iron reducing bacteria genus selectively enriched. Simultaneously, the properties of PGO changed in the soil. Organic molecules and nitrogen-containing groups were detected in the soil-modified GO (SGO). SGO, which exhibited fewer negative charges, was more unstable than PGO. In addition, SGO presented higher chemical activity than PGO; for example, SGO exhibited more unpaired electrons and disordered structures.Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (grant nos 31170473 and 21307061), the Ministry of Education of China as an innovative team project (grant no. IRT 13024), Tianjin Natural Science Foundation (grant no. 14JCQNJC08900), the Specialized Research Fund for the Doctoral Program of Higher Education of China (grant no. 2013003112016), the Postdoctoral Science Foundation of China (grant no. 2014M550138) and the Fundamental Research Funds for the Central Universities (grant no. 65121006).Notes and references
- D. A. Dikin, S. Stankovich, E. J. Zimney, R. D. Piner, G. H. Dommett, G. Evmenenko, S. T. Nguyen and R. S. Ruoff, Nature, 2007, 448, 457–460 CrossRef CAS PubMed.
- R. Joshi, P. Carbone, F. Wang, V. Kravets, Y. Su, I. Grigorieva, H. Wu, A. Geim and R. Nair, Science, 2014, 343, 752–754 CrossRef CAS PubMed.
- X. Qu, J. Brame, Q. Li and P. J. Alvarez, Acc. Chem. Res., 2012, 46, 834–843 CrossRef PubMed.
- J. Zhao, Z. Wang, J. C. White and B. Xing, Environ. Sci. Technol., 2014, 48, 9995–10009 CrossRef CAS PubMed.
- X. Qu, P. J. Alvarez and Q. Li, Water Res., 2013, 47, 3931–3946 CrossRef CAS PubMed.
- J. Lee, S. Mahendra and P. J. Alvarez, ACS Nano, 2010, 4, 3580–3590 CrossRef CAS PubMed.
- V. C. Sanchez, A. Jachak, R. H. Hurt and A. B. Kane, Chem. Res. Toxicol., 2011, 25, 15–34 CrossRef PubMed.
- B. Shrestha, V. Acosta-Martinez, S. B. Cox, M. J. Green, S. Li and J. E. Canas-Carrell, J. Hazard. Mater., 2013, 261, 188–197 CrossRef CAS PubMed.
- B. Nowack and T. D. Bucheli, Environ. Pollut., 2007, 150, 5–22 CrossRef CAS PubMed.
- R. Dinesh, M. Anandaraj, V. Srinivasan and S. Hamza, Geoderma, 2012, 173, 19–27 CrossRef.
- Z. H. Tong, M. Bischoff, L. Nies, B. Applegate and R. F. Turco, Environ. Sci. Technol., 2007, 41, 2985–2991 CrossRef CAS PubMed.
- J. Zhao, F. Liu, Z. Wang, X. Cao and B. Xing, Environ. Sci. Technol., 2015, 49, 2849–2857 CrossRef CAS PubMed.
- Z. H. Tong, M. Bischoff, L. F. Nies, P. Myer, B. Applegate and R. F. Turco, Environ. Sci. Technol., 2012, 46, 13471–13479 CrossRef CAS PubMed.
- H. Chung, Y. Son, T. K. Yoon, S. Kim and W. Kim, Ecotoxicol. Environ. Saf., 2011, 74, 569–575 CrossRef CAS PubMed.
- S. Liu, T. H. Zeng, M. Hofmann, E. Burcombe, J. Wei, R. Jiang, J. Kong and Y. Chen, ACS Nano, 2011, 5, 6971–6980 CrossRef CAS PubMed.
- I. E. M. Carpio, C. M. Santos, X. Wei and D. F. Rodrigues, Nanoscale, 2012, 4, 4746–4756 RSC.
- S. Kang, M. S. Mauter and M. Elimelech, Environ. Sci. Technol., 2009, 43, 2648–2653 CrossRef CAS PubMed.
- O. Akhavan and E. Ghaderi, Carbon, 2012, 50, 1853–1860 CrossRef CAS.
- G. Wang, F. Qian, C. W. Saltikov, Y. Jiao and Y. Li, Nano Res., 2011, 4, 563–570 CrossRef CAS.
- P. Khanra, T. Kuila, N. H. Kim, S. H. Bae, D.-S. Yu and J. H. Lee, Chem. Eng. J., 2012, 183, 526–533 CrossRef CAS.
- C. Zhu, Z. Feng, M. Fan, C. Chen, B. Ma, J. Yang and D. Sun, RSC Adv., 2014, 4, 40292–40295 RSC.
- D. F. Rodrigues, D. P. Jaisi and M. Elimelech, Environ. Sci. Technol., 2013, 47, 625–633 CrossRef CAS PubMed.
- J. Zhao, S. Pei, W. Ren, L. Gao and H.-M. Cheng, ACS Nano, 2010, 4, 5245–5252 CrossRef CAS PubMed.
- K. R. Amato, C. J. Yeoman, A. Kent, N. Righini, F. Carbonero, A. Estrada, H. R. Gaskins, R. M. Stumpf, S. Yildirim and M. Torralba, ISME J., 2013, 7, 1344–1353 CrossRef CAS PubMed.
- L. Lu, D. Xing and N. Ren, Water Res., 2012, 46, 2425–2434 CrossRef CAS PubMed.
- O. Akhavan and E. Ghaderi, ACS Nano, 2010, 4, 5731–5736 CrossRef CAS PubMed.
- L. Zhang, X. Li, Y. Huang, Y. Ma, X. Wan and Y. Chen, Carbon, 2010, 48, 2367–2371 CrossRef CAS.
- Y. Jiao, F. Qian, Y. Li, G. Wang, C. W. Saltikov and J. A. Gralnick, J. Bacteriol., 2011, 193, 3662–3665 CrossRef CAS PubMed.
- T. Kuila, S. Bose, P. Khanra, A. K. Mishra, N. H. Kim and J. H. Lee, Carbon, 2012, 50, 914–921 CrossRef CAS.
- Z.-H. Sheng, L. Shao, J.-J. Chen, W.-J. Bao, F.-B. Wang and X.-H. Xia, ACS Nano, 2011, 5, 4350–4358 CrossRef CAS PubMed.
- K. R. Lee, K. U. Lee, J. W. Lee, B. T. Ahn and S. I. Woo, Electrochem. Commun., 2010, 12, 1052–1055 CrossRef CAS.
- S. Kim, S. Zhou, Y. Hu, M. Acik, Y. J. Chabal, C. Berger, W. de Heer, A. Bongiorno and E. Riedo, Nat. Mater., 2012, 11, 544–549 CrossRef CAS PubMed.
- K. Yang, Y. Li, X. Tan, R. Peng and Z. Liu, Small, 2013, 9, 1492–1503 CrossRef CAS PubMed.
- X. Hu and Q. Zhou, Chem. Rev., 2013, 113, 3815–3835 CrossRef CAS PubMed.
- Y. Su, M. Hu, C. Fan, Y. He, Q. Li, W. Li, L.-H. Wang, P. Shen and Q. Huang, Biomaterials, 2010, 31, 4829–4834 CrossRef CAS PubMed.
- K.-H. Liao, Y.-S. Lin, C. W. Macosko and C. L. Haynes, ACS Appl. Mater. Interfaces, 2011, 3, 2607–2615 CAS.
- G. P. Kotchey, B. L. Allen, H. Vedala, N. Yanamala, A. A. Kapralov, Y. Y. Tyurina, J. Klein-Seetharaman, V. E. Kagan and A. Star, ACS Nano, 2011, 5, 2098–2108 CrossRef CAS PubMed.
- S. George, S. Lin, Z. Ji, C. R. Thomas, L. Li, M. Mecklenburg, H. Meng, X. Wang, H. Zhang and T. Xia, ACS Nano, 2012, 6, 3745–3759 CrossRef CAS PubMed.
- X. Hu, L. Mu, K. Lu, J. Kang and Q. Zhou, ACS Appl. Mater. Interfaces, 2014, 6, 9220–9227 CAS.
- X. Hu, L. Mu, J. Kang, K. Lu, R. Zhou and Q. Zhou, Environ. Sci. Technol., 2014, 48, 6919–6927 CrossRef CAS PubMed.
- M. H. Lahiani, E. Dervishi, J. Chen, Z. Nima, A. Gaume, A. S. Biris and M. V. Khodakovskaya, ACS Appl. Mater. Interfaces, 2013, 5, 7965–7973 CAS.
- Z. Luo, Y. Lu, L. A. Somers and A. C. Johnson, J. Am. Chem. Soc., 2009, 131, 898–899 CrossRef CAS PubMed.
- K. Krishnamoorthy, M. Veerapandian, L.-H. Zhang, K. Yun and S. J. Kim, J. Phys. Chem. C, 2012, 116, 17280–17287 CAS.
- C. Zhu, S. Guo, Y. Fang and S. Dong, ACS Nano, 2010, 4, 2429–2437 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available: Table S1 presenting the physicochemical properties of the collected soil. Table S2 listing the data of richness and diversity of bacterial communities in PGO-soil sample (PGOS) and control soil sample (CS) analyzed from pyrosequencing. Table S3 listing the organic molecules extracted from the raw soil sample. Fig. S1 presenting the photographs of separation of SGO (soil-modified graphene oxide) and the photographs of SGO powders. Fig. S2 and S3 showing the characteristics of bacterial communities. Fig. S4 being the SEM, TEM and EDS results. Fig. S5 being the AFM images. Fig. S6 presenting the wide spectra of XPS of PGO and SGO. Fig. S7 presenting FTIR spectra of PGO and SGO. Fig. S8 showing the organic molecules extracted from SGO. Fig. S9 showing the distribution and surface charge of the nanomaterials. See DOI: 10.1039/c5ra01045d |
| This journal is © The Royal Society of Chemistry 2015 |