Bacterial–yeast consortium as an effective biocatalyst for biodegradation of sulphonated azo dye Reactive Red 198
Mayur B. Kuradeab,
Tatoba R. Waghmodecd,
Mital U. Jadhave,
Byong-Hun Jeonb and
Sanjay P. Govindwar*c
aDepartment of Biotechnology, Shivaji University, Kolhapur-416004, India
bDepartment of Natural Resources and Environmental Engineering, Hanyang University, Seoul, South Korea
cDepartment of Biochemistry, Shivaji University, Kolhapur-416004, India. E-mail: govindwarsp@gmail.com; Fax: +91-231-2691533; Tel: +91-231-2609152
dInstitute of Agriculture and Life Sciences, Gyeongsang National University, Jinju 660-701, South Korea
eDepartment of Microbiology, Shivaji University, Kolhapur-416004, India
First published on 19th February 2015
Abstract
A novel bacterial–yeast consortium (Brevibacillus laterosporus and Galactomyces geotrichum) acts as a proficient biocatalyst. It decolorized 92% of sulphonated azo dye Reactive Red 198 (RR 198) within 18 h at a dye concentration of 50 mg L−1 as compared to 58 and 42% decolorization using Brevibacillus laterosporus and Galactomyces geotrichum alone, respectively, in the same experimental conditions (pH 7, 40 °C, in static condition). The cumulative action of enzymes such as veratryl alcohol oxidase, laccase, NADH-DCIP reductase and azoreductase in the consortium culture was responsible for dye degradation. Fourier transform infrared spectroscopy and high performance thin layer chromatography analysis of the dye and its extracted metabolites suggested the biotransformation of RR 198 into simple metabolites; whereas the biotransformation of the same by individual microorganisms was different than by consortial biodegradation. According to gas chromatography-mass spectroscopy studies, RR 198 was biotransformed into much simpler compounds such as (ethylsulfonyl)benzene and 1,3,5-triazine by the bacterial–yeast consortium. This metabolic fate of the dye was entirely different in consortium than when compared to individual microbial treatment. Single microbial species could lead to only partial mineralization of the intact dye molecule; whereas, nearly complete degradation of the dye molecule was achieved using the consortium culture. This study clearly suggests that the consortium has an enormous strength to catalyze RR 198 within a short period as compared to individual microbial cultures.
1. Introduction
Among 12 classes of chromogenic groups of dyes, the azo dyes, including sulphonated and reactive dyes, make up to 70% of all textile dyestuffs produced. This leads to effluent streams containing intense color due to the presence of azo dyes. Normally colors are noticeable at a dye concentration of more than 1 mg L−1 and an average concentration of 300 mg L−1 has been reported in effluents from textile manufacturing processes.1 Azo dyes are of concern in wastewater treatment because of their intense coloration, biorecalcitrance and potential toxicity to animals and humans as they have mutagenic and carcinogenic properties.2,3 Several physicochemical techniques have been used in treating wastewater containing dyes but the methodologies appear to face several technical and economic limitations and lead to the generation of toxic by-products.4–7On the other hand, a number of biotechnological approaches have been suggested by recent research as of potential interest in combating this pollution source in an eco-efficient manner.8,9 Lower cost of treatment and amenability to easy scale up are the merits of biological methods. Numerous studies have been carried out with isolated, selected, mutated and genetically engineered microorganisms for effective bioremediation to achieve enhanced rates of degradation of recalcitrant target compounds.10–14 However, the isolated/single cultures are often specific to the type of dye under consideration. Moreover, pure cultures cannot be scaled up easily and maintained in large-scale operations typical of effluent treatment systems.15 Instead, microbial consortia could be attractive tools to improve the rate of decolorization processes with better suitability in large-scale applications.
Recent studies have suggested that mixed cultures may be more appropriate for decolorization of azo dyes.16–18 Different strains may attack dye molecules at different positions or may use decomposition products produced by another strain for further decomposition.19,20 In recent years, mixed microbial consortia have been believed to have superiority over pure isolates in some biorefractory wastewater treatments. Recently, a bacterial consortium (Enterobacter dissolvens and Pseudomonas aeruginosa) was used for the treatment of diazo dye Acid Maroon V, which showed a 2.15 fold increase in decolorization efficiency compared to the individual microorganisms.21 Fungal species Aspergillus terreus and Rhizopus oryzae in consortium enhanced the removal of metals and dyes.22 Yeast species including Galactomyces geotrichum, Candida tropicalis and Saccharomyces cerevisiae were effectively used for dye degradation;14,23–25 however, as far as our knowledge is concerned, there are still no reports on yeast consortia for dye decolorization.
Although studies carried out in the past have used undefined microbial consortia1,26 or pure cultures for dye decolorization, there is very limited research to unravel the application of bacterial–yeast/fungi consortia in the rehabilitation of our natural resources. In the present study, a bacterial–yeast consortium has been developed using a yeast, Galactomyces geotrichum, with another potential dye-degrading microbial species, Brevibacillus laterosporus, to decolorize a sulphonated azo dye, Reactive Red 198 (RR 198). Brevibacillus laterosporus is an aerobic bacterium and is a potent dye-degrading species. Our previous reports provide strong evidence of the abilities of Brevibacillus laterosporus in the decolorization of several dyes, such as Remazol Red, Rubine GFL, Disperse Brown 118, Golden Yellow HER and mixture of dyes.27–30 In the yeast species, Galactomyces geotrichum is a strong highlight, as it has been used to eliminate different varieties of textile dyes, which include Reactive Yellow 84A, Rubine GFL and Methyl Red.14,23 The presence of a wide range of biocatalytic enzyme systems make these species strong candidates for enhanced dye degradation. High efficiency and biocatalytic performance was expected using the combination of these microbial species. RR 198 is a sulphonated monoazo dye which is toxic and recalcitrant in nature due to the presence of azo and sulphonic groups. It gives intense coloration to water at very low concentrations, hence effluent containing this dye causes deterioration of water bodies and leads to harmful effects on aquatic flora and fauna. It is one of the most used azo dyes in the textile industries in Ichalkaranji, India; hence this dye was used as a model azo dye for the decolorization experiments in the present study. This study puts forward the detailed mechanism of how this bacterial–yeast consortium is superior over an individual microbial culture in eliminating the xenobiotic and toxic azo dye RR 198 with the evidence of enzymatic status, characterization of products extracted after dye decolorization using Fourier transform infrared spectroscopy (FTIR) and high performance thin layer chromatography (HPTLC) along with metabolic fate of RR 198 using gas chromatography-mass spectroscopy (GC-MS) data.
2. Experimental
2.1. Microorganisms and culture conditions
Brevibacillus laterosporus MTCC 2298 was obtained from Microbial Type Culture Collection, Chandigarh, India. The pure culture was maintained at 4 °C on a nutrient agar slant containing (g L−1): NaCl 5.0; bacteriological peptone 5.0; yeast extract 2.0; beef extract 1.0; and agar powder 15.0, at 4 °C. Galactomyces geotrichum MTCC 1360 was obtained from Microbial Type Culture Collection, Chandigarh, India. Pure culture was maintained on malt yeast agar slants at 4 °C; the composition of malt extract medium was (g L−1): malt extract 3; yeast extract 3; peptone 5; and glucose 10.2.2. Dyes and chemicals
All required chemicals were obtained from Sigma Aldrich, USA, Hi-media Laboratories Pvt. Ltd., Mumbai, India, and Sisco Research Laboratory (SRLs), India. All chemicals used were of the highest purity available and of the analytical grade. Textile dye RR 198 was a munificent gift from Manpasand textile processing industry, Ichalkaranji, India.2.3. Development of bacterial–yeast consortium
The consortium was prepared by the earlier reported protocol.31 The surface culture biomass of G. geotrichum (1 cm pieces of mycelium cut from agar slants were used to inoculate 100 mL malt extract medium in 250 mL Erlenmeyer flasks and grown at 30 °C for 24 h) was aseptically transferred to the 24 h old culture of B. laterosporus (grown at 30 °C in 250 mL Erlenmeyer flasks containing 100 mL of nutrient medium) to obtain the bacterial–yeast consortium.2.4. Decolorization experiment
All decolorization experiments were performed in the 250 mL Erlenmeyer flasks. Textile dye RR 198 was added at a concentration of 50 mg L−1 in the flask containing developed consortium with 100 mL nutrient broth. Aliquots were centrifuged (4000 rpm for 20 min) to separate cell mass and clear supernatant was used to determine the decolorization. Decolorization was determined by measuring the absorbance maximum of the dye (λmax of RR 198-530 nm) using a UV visible spectrophotometer (Hitachi U-2800) by using the following formula:To investigate the effect of different dye concentrations on decolorization, the concentrations of RR 198 were varied from 50 to 400 mg L−1 (50, 100, 200, 300 and 400 mg L−1) to determine the amount of dye that could be decolorized by the consortium and decolorization was determined by the formula mentioned above.
All the media used in this study were pre-sterilized before experiments by autoclaving at 121 °C for 20 min. All decolorization experiments were carried out in triplicate. Abiotic controls (without microorganisms) were always included.
2.5. Preparation of cell free extract
The consortium was prepared as mentioned above and individual organisms were grown in their respective medium for 24 h at 30 °C and centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 20 min. This biomass of consortium and individual organisms were separately suspended in 50 mM potassium phosphate buffer (pH 7.4), gently homogenized and used for sonication (sonics-vibracell ultrasonic processor) keeping the sonifier output at 60 amplitude, maintaining the temperature below 4 °C and giving 12 strokes each of 30 s with 1 min intervals. The resulting extracts were then used as enzyme sources. The culture supernatant obtained after centrifugation during the harvesting of cell biomass was directly used as a source of extracellular enzymes. The same procedure was carried out for samples obtained after degradation of the dye.
000 rpm for 20 min. This biomass of consortium and individual organisms were separately suspended in 50 mM potassium phosphate buffer (pH 7.4), gently homogenized and used for sonication (sonics-vibracell ultrasonic processor) keeping the sonifier output at 60 amplitude, maintaining the temperature below 4 °C and giving 12 strokes each of 30 s with 1 min intervals. The resulting extracts were then used as enzyme sources. The culture supernatant obtained after centrifugation during the harvesting of cell biomass was directly used as a source of extracellular enzymes. The same procedure was carried out for samples obtained after degradation of the dye.
2.6. Determination of enzyme activities
All enzyme activities were assayed spectrophotometrically (Hitachi U-2800) in the cell free extract as well as culture media at room temperature (25 °C). Veratryl alcohol oxidase activity was determined by using the procedure reported earlier by Jadhav et al.32 Laccase activity was monitored according to the procedure reported earlier.33 2 mL reaction mixture contained 5 mM 3′,3′-diaminobenzidine tetrahydrate (DAB) in 0.1 M acetate buffer (pH 4.8) and increase in optical density was measured at 410 nm. Tyrosinase activity was determined by the procedure as follows: 3 mL reaction mixture containing 50 mM of catechol and 2.1 mM of ascorbic acid in 50 mM potassium phosphate buffer (pH 6.5) was equilibrated at 25 °C. ΔA265 nm was monitored until constant, and then 0.1 mL of the supernatant from the reaction mixture was added. The formation of o-benzoquinone and dehydroascorbic acid and decrease in optical density was measured at 265 nm. ΔA265 nm was obtained using the maximum linear rate for both the test and control. One unit of tyrosinase activity was equal to a ΔA265 nm of 0.001 per min at pH 6.5 at 25 °C in a 3.0 mL reaction mixture containing L-catechol and L-ascorbic acid.The NADH-DCIP reductase and riboflavin reductase activity were assayed by modifying earlier reported methods by Waghmode et al.33 DCIP reduction was monitored at 590 nm and calculated using an extinction coefficient of 0.019 μM−1 cm−1. The reaction mixture (5.0 mL) prepared contained 25 μM substrate (DCIP) in 50 mM potassium phosphate buffer (pH 7.4) and 0.1 mL enzyme. From this, 2.0 mL reaction mixture was assayed at 590 nm by addition of 250 μM NADH. Riboflavin reductase NAD(P)H:flavin oxidoreductase was measured by monitoring the decrease in absorbance at 340 nm. Cell free extract was added to a solution (final volume 2 mL) containing 100 mM of Tris–HCl (pH 7.5), 25 μM of NADPH and 10 μM of riboflavin. Reaction rates were calculated by using a molar extinction coefficient of 0.0063 μM−1 cm−1. Azo reductase activity was assayed by an earlier reported method.33 The assay mixture (2 mL) contained 4.45 μM of Methyl Red (MR), 50 μM NADH, and 1.2 mL of potassium phosphate buffer (20 mM, pH 7.5). The reaction mixture was pre-incubated for 4 min followed by the addition of NADH and monitored for the decrease in color absorbance (430 nm) at room temperature. The reaction was initiated by addition of 0.2 mL of the enzyme solution. Methyl Red reduction was calculated by using its molar extinction coefficient of 0.023 μM−1 cm−1. One unit of enzyme activity was defined as the amount of enzyme required to reduce 1 μM of substrate per min per mg of protein. All the enzyme assays were run in triplicate.
2.7. Extraction and analysis of products obtained after dye decolorization
Biomass was removed after decolorization by centrifugation (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm at 4 °C for 20 min) and the supernatant was processed for the extraction of degradation metabolites with an equal volume of ethyl acetate. The extracted residues were dried over anhydrous Na2SO4 and evaporated to dryness in a rotary evaporator. The crystals obtained were dissolved in a small volume of high pressure liquid chromatography (HPLC) grade methanol and used for high performance thin layer chromatography (HPTLC), Fourier transform infrared spectroscopy (FTIR) and gas chromatography-mass spectroscopy (GC-MS) analysis. The FTIR analysis of extracted metabolites was carried out using a protocol reported earlier.31 This FTIR analysis of metabolites was carried out on Perkin-Elmer, spectrum one instrument and compared with a control dye in the mid-IR region of 400–4000 cm−1 with 8 scan speed. The samples were mixed with spectroscopically pure KBr in the ratio 5
000 rpm at 4 °C for 20 min) and the supernatant was processed for the extraction of degradation metabolites with an equal volume of ethyl acetate. The extracted residues were dried over anhydrous Na2SO4 and evaporated to dryness in a rotary evaporator. The crystals obtained were dissolved in a small volume of high pressure liquid chromatography (HPLC) grade methanol and used for high performance thin layer chromatography (HPTLC), Fourier transform infrared spectroscopy (FTIR) and gas chromatography-mass spectroscopy (GC-MS) analysis. The FTIR analysis of extracted metabolites was carried out using a protocol reported earlier.31 This FTIR analysis of metabolites was carried out on Perkin-Elmer, spectrum one instrument and compared with a control dye in the mid-IR region of 400–4000 cm−1 with 8 scan speed. The samples were mixed with spectroscopically pure KBr in the ratio 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95, pellets were fixed in a sample holder, and the analysis was carried out. The GC-MS analysis of metabolites was carried out using a Shimadzu 2010 MS Engine, equipped with an integrated gas chromatograph with an HP1 column (60 m long, 0.25 mm id, nonpolar). Helium was used as carrier gas, at a flow rate of 1 mL min−1. The injector temperature was maintained at 280 °C with oven conditions as: 80 °C kept constant for 2 min and increased up to 200 °C by 10 °C min−1, raised up to 280 °C with a 20 °C min−1 rate. The interphase temperature of the MS engine was maintained at 280 °C and the ionization voltage was 70 eV. The minimum and maximum limits of m/z were kept to 40 and 950 respectively. The compounds were identified on the basis of mass spectra using the NIST library. To confirm the biodegradation of dyes, the same metabolites were analyzed by HPTLC using silica gel plates (HPTLC LiChrospher silica gel 60 F254S, Merck). 15 μL of sample was applied on the plate by micro syringe using a sample applicator (Linomat V, Camag, Switzerland). The solvent system used for RR 198 was propanol
95, pellets were fixed in a sample holder, and the analysis was carried out. The GC-MS analysis of metabolites was carried out using a Shimadzu 2010 MS Engine, equipped with an integrated gas chromatograph with an HP1 column (60 m long, 0.25 mm id, nonpolar). Helium was used as carrier gas, at a flow rate of 1 mL min−1. The injector temperature was maintained at 280 °C with oven conditions as: 80 °C kept constant for 2 min and increased up to 200 °C by 10 °C min−1, raised up to 280 °C with a 20 °C min−1 rate. The interphase temperature of the MS engine was maintained at 280 °C and the ionization voltage was 70 eV. The minimum and maximum limits of m/z were kept to 40 and 950 respectively. The compounds were identified on the basis of mass spectra using the NIST library. To confirm the biodegradation of dyes, the same metabolites were analyzed by HPTLC using silica gel plates (HPTLC LiChrospher silica gel 60 F254S, Merck). 15 μL of sample was applied on the plate by micro syringe using a sample applicator (Linomat V, Camag, Switzerland). The solvent system used for RR 198 was propanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate
ethyl acetate![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) methanol
methanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) toluene (3
toluene (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6
6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.5
0.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.5). The chromatogram was analyzed using a scanner (Camag, Switzerland).
0.5). The chromatogram was analyzed using a scanner (Camag, Switzerland).
2.8. Phytotoxicity study
This test was performed to assess the toxicity of the untreated and treated effluent with respect to two kinds of seeds commonly used in Indian agriculture: Sorghum vulgare and Phaseolus mungo, using an earlier reported method.33 Ten seeds of each plant were sown into a plastic sand pot. The sand pot was prepared by adding 15 g of washed sand into the plastic pot. Metabolites were extracted using the procedure described in section 2.7 and the crystals obtained after ethyl acetate extraction were dissolved in distilled water. The toxicity study was carried out by watering the seeds of each plant with untreated textile effluent and extracting the metabolites of effluent (5 mL per day). Control was kept by watering the seeds in distilled water (5 mL per day). Germination (%), lengths of the shoots and roots were recorded after 7 days. The experiment was carried out at room temperature.2.9. Statistical analysis
Data were analyzed by one-way analysis of variance using a Tukey–Kramer multiple comparison test. Values were the mean of three independent experiments. Values were considered significant when P < 0.05.3. Results and discussion
3.1. Decolorization experiment and effect of physicochemical parameters
The decolorization of dyes is directly dependent upon the catalytic potential of a microorganism. Decolorization efficiencies of individual microorganisms and the developed consortium are presented in Fig. 1 in terms of percentage decolorization and time required for decolorization. This developed consortium is nothing but a combination of a bacterium (B. laterosporus) which is known to be a proficient decolorizer27–29 with another yeast species (G. geotrichum). The consortium exhibited 92% decolorization of RR 198 within 18 h with a dye concentration of 50 mg L−1 when optimized conditions (pH 7, 40 °C and static condition) for decolorization were provided to the consortial culture. In contrast, B. laterosporus and G. geotrichum alone exhibited 58 and 42% decolorization respectively in the same experimental conditions. Recently, we have reported the higher decolorization of textile dye Scarlet RR and textile industry effluent using the same consortium of yeast and bacteria.31,33 Decolorization of most of the dyes by microbial cultures is generally carried out in a neutral to slightly alkaline pH range. The consortium of Bacillus sp. and G. geotrichum also showed effective decolorization of Brilliant Blue G at neutral pH,34 while natural microbial consortium required the same temperature range (40 °C) for maximum decolorization of a mixture of dyes and also with faster rate.35 The loss of cell viability or the negligible oxido-reductive enzyme activities might be the cause of reduced decolorization in culture broth which was incubated beyond the optimum temperature range. | ||
| Fig. 1 Decolorization of RR 198 by consortium (♦), B. laterosporus (■) and G. geotrichum (▲) at pH 7, 40 °C and in static condition with a dye concentration of 50 mg L−1. | ||
3.2. Effect of dye concentration on decolorization by consortium
An investigation of the effect of dye concentration on the decolorization ability of a microbial system is very important as it explores the limits of a biological system. To assess this, the consortium culture was provided with higher doses of dye from 50 to 400 mg L−1. It was found that a higher concentration of dye decreased the decolorization performance of the consortium. However, the difference in decolorization was insignificant (P > 0.05) as decolorization in a flask with a dye concentration of 400 mg L−1 was 59% as compared to 78% decolorization in a flask containing the dye at a concentration of 50 mg L−1 within the first 12 h (Fig. 2A). Interestingly, the decolorization of RR 198 using the consortium was found to be much higher with all dye concentrations used with a minor cessation in decolorization with an increase in dye concentration as compared to the decolorization by individual microorganisms. The decolorization of RR 198 by B. laterosporus was significantly reduced from 40% (at a dye concentration of 50 mg L−1) to 7% (at a dye concentration of 400 mg L−1) within the first 12 h; while at the same time, 29 and 12% decolorization were achieved using G. geotrichum at dye concentrations of 50 and 400 mg L−1 respectively (Fig. 2B and C). These results suggest that the consortial culture can tolerate higher dye concentrations with effective decolorization of dyes as compared to the individual strains. It has been found that the higher dye concentrations do not alter the decolorization when bacterial co-culture is used instead of pure culture, and this might be due to the synergistic effect of both microorganisms.31,36 The gradual decrease in the decolorization rate at higher dye concentrations in the case of individual microbial cultures was probably due to the toxic effect of dyes with regard to the individual bacteria and/or inadequate biomass concentration or improper cell to dye ratio.34 Based on previous reports,37 the concentration of dye in the textile industry effluent is generally around 16–32 mg L−1 and as the consortium is unaffected by higher dye concentrations, this can be used as an effective tool in continuous wastewater treatment systems.3.3. Enzyme analysis
A major mechanism of biodegradation by the microbial cells is controlled by the biotransformation enzymes produced by the cell itself. To prove the role of enzymes involved in the biodegradation reaction, various biotransformation enzymes were monitored during the decolorization of RR 198 (18 h) by the consortium and also by the individual microorganisms i.e. B. laterosporus and G. geotrichum, which are summarized in Table 1. In this set of experiments, significant induction of laccase (115%) by the consortium was observed as compared to the activity in the cells of G. geotrichum. NADH-DCIP reductase activity was induced by 43 and 14% in the consortium cells when compared to the activities in the cells of G. geotrichum and B. laterosporus respectively. In addition to that, 22% induction in azo reductase activity was detected in the consortium culture as compared to the activity detected in the cells of B. laterosporus. It should be noted that a few biotransformation enzymes were absent in the individual microbial species during decolorization. In contrast, the same enzymes were in positive action in the consortium system during decolorization of RR 198. This collective action of all the oxido-reductive enzymes in the consortium might be the key factor in rapid decolorization. The communal enzyme activities of mixed microbial cultures are far better for dye decolorization as compared to individual strains.31,36,37 These enzymes have been found in an active role in the biodegradation of various textile dyes such as Acid Violet 19, Amaranth,13,32 Green HE4B,38 Malachite Green39 and mixtures of dyes.35| G. geotrichum | B. laterosporus | Consortium | ||
|---|---|---|---|---|
| a Activity in units per min per mg protein.b Activity in IU.c μg of DCIP reduced per min per mg protein.d μg of riboflavin reduced per min per mg protein.e μM of MR reduced per min per mg protein.f NA – no activity. Values are the mean of three experiments ± SEM. Enzyme activities of the consortium are significantly different from those of G. geotrichum at *P < 0.001 and significantly different from those of B. laterosporus at %P < 0.001 and by one-way ANOVA with a Tukey–Kramer multiple comparisons test. | ||||
| Veratryl alcohol oxidasea | NA | 1.846 ± 0.154 | 1.051 ± 0.106*% | |
| Laccasea | 0.942 ± 0.046 | NA | 1.084 ± 0.087*% | |
| Tyrosinaseb | Intracellular | 1408 ± 151 | 1115 ± 18.57 | NA |
| Extracellular | 1217 ± 111 | 253.4 ± 14.72 | NA | |
| NADH-DCIP reductasec | 183.96 ± 11.22 | 230.94 ± 34.36 | 263.03 ± 23.79*% | |
| Riboflavin reductased | 14.39 ± 0.296 | NA | 4.80 ± 0.126*% | |
| Azoreductasee | 5.546 ± 0.481 | 3.890 ± 0.157 | 4.738 ± 0.227*% | |
3.4. Product characterization
Chromatographic techniques can be used to detect the biotransformation process of a substrate molecule. In the present study, the biotransformation of RR 198 was monitored by FTIR, HPTLC, and GC-MS. Fig. 3A shows the FTIR spectrum of RR 198. The FTIR spectrum of the RR 198 dye shows peaks at 2889 cm−1 for C–H stretching vibrations in alkanes, 1646 and 1490 cm−1 for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching vibrations in cyclic α,β-unsaturated compounds, 1553 cm−1 for C
N stretching vibrations in cyclic α,β-unsaturated compounds, 1553 cm−1 for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching vibrations in pyridines and N–H deformation in acyclic compounds, 1400 cm−1 for O–H deformation in phenols, 1209 cm−1 for C–O stretching in phenols and also for N–H deformation in bonded N–H, 1137 cm−1 for S
N stretching vibrations in pyridines and N–H deformation in acyclic compounds, 1400 cm−1 for O–H deformation in phenols, 1209 cm−1 for C–O stretching in phenols and also for N–H deformation in bonded N–H, 1137 cm−1 for S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O asymmetric stretching in sulphones and C–H deformation in substituted benzenes, 1055 cm−1 for C–O stretching in phenols, 740 cm−1 for C–Cl stretching in halogenated compounds and benzene rings with five adjacent free H atoms, and 621 cm−1 for C–N stretching and N–H deformation in acyclic compounds. On the other hand, significant disappearance of major peaks and the formation of new peaks in the FTIR spectra of metabolites obtained after dye decolorization suggest the biotransformation and more effective mineralization of RR 198 by the consortium than by individual microorganisms (Fig. 3B–D).
O asymmetric stretching in sulphones and C–H deformation in substituted benzenes, 1055 cm−1 for C–O stretching in phenols, 740 cm−1 for C–Cl stretching in halogenated compounds and benzene rings with five adjacent free H atoms, and 621 cm−1 for C–N stretching and N–H deformation in acyclic compounds. On the other hand, significant disappearance of major peaks and the formation of new peaks in the FTIR spectra of metabolites obtained after dye decolorization suggest the biotransformation and more effective mineralization of RR 198 by the consortium than by individual microorganisms (Fig. 3B–D).
HPTLC analysis of a control dye and metabolites obtained after decolorization clearly suggest the biotransformation of RR 198 into simple compounds as multiple bands were obtained in the metabolites lane of the consortium and also in metabolites produced by individual microorganisms (Fig. 4). However, the number of separated compounds and their Rf values were different than those of the control dye (data not shown). In the lane of the metabolites obtained via consortial degradation, eleven bands were detected, while nine and six bands were obtained in the lanes of metabolites extracted after degradation of RR 198 by B. laterosporus and G. geotrichum respectively. These observations suggested the catalytic superiority of the consortium in dye degradation. The change of Rf value of RR 198 (0.90) and presence of multiple products in the metabolites lane provided supplementary substantiation for the biodegradation of dyes as recorded by FTIR.
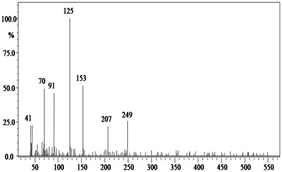
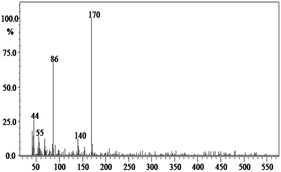
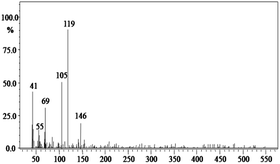
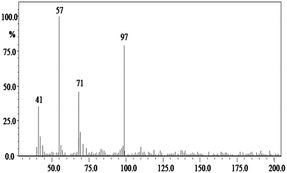
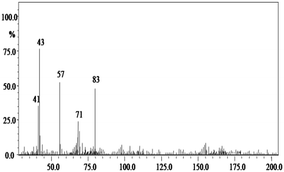
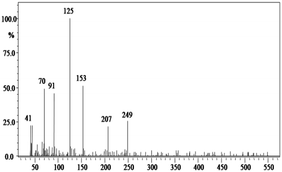
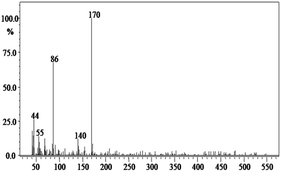
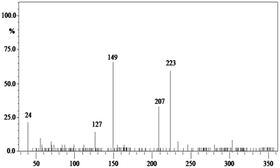
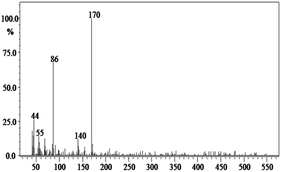
In order to gain insight into the difference between the metabolic fate of RR 198 by consortium and pure cultures during dye degradation, degradative pathways were proposed using GC-MS data of metabolites extracted after the degradation of RR 198, which would help us to obtain an answer as to how the consortium actually works. Fig. 5 illustrates the proposed pathways of RR 198, degraded by consortium and individual organisms on the basis of GC-MS analysis of products formed within 18 h and Table 2 represents mass spectra of the metabolites formed. In all the three possibilities, biodegradation of RR 198 was initiated by reduction of the azo bond. Azo dyes can be cleaved symmetrically and asymmetrically, with an active site available for an enzyme to excite the molecule.27 Accordingly, the first step in degradation of RR 198 by consortium was asymmetric cleavage of the azo bond via azo reductase which led to formation of intermediate [A] 2-(phenylsulfonyl)ethanesulfonate detected at retention time in GC, Rt-24.325, having a molecular weight of MW-249 based on the obtained mass to charge ratio of m/z 247(+2) Fig. 5C. Further desulphonation of this intermediate led to the product [B] (ethylsulfonyl)benzene detected at retention time in GC, Rt-22.417, having a molecular weight of MW-170 and the mass to charge ratio of m/z 170 by the action of veratryl alcohol oxidase. On the other hand, laccase was responsible for dechlorination, desulphonation and deamination of postulated intermediate [1], which yielded product [C] naphthalen-1-ol [(Rt-20.167, MW-144) based on the obtained mass to charge ratio of m/z 146(−2)], and a reactive intermediate [D] 1,3,5-triazin-2-amine [(Rt-21.508, MW-96) based on the mass to charge ratio of m/z 97(−1)] and an unidentified product [2]. The formation of final product [E] 1,3,5-triazine [(Rt-19.500, MW-81) according to the mass to charge ratio obtained at m/z 83(−2)] indicates deamination of reactive intermediate [D] by veratryl alcohol oxidase.
 | ||
| Fig. 5 Proposed metabolic pathways of RR 198 by B. laterosporus (A), G. geotrichum (B) and consortium (C). | ||
| Observations | Sorghum vulgare | Phaseolus mungo | ||||
|---|---|---|---|---|---|---|
| I | II | III | I | II | III | |
| a (I): Seeds germinated in distilled water. (II): Seeds germinated in RR 198. (III): Seeds germinated in metabolites obtained after decolorization of RR 198. Data were analyzed by one way analysis of variance (ANOVA) with a Tukey–Kramer multiple comparison test using mean values of germinated seeds of three experiments. Seeds germinated in RR 198 are significantly different from the seeds germinated in plain distilled water at *P < 0.001 and the seeds germinated in degradation products are significantly different from the seeds germinated in RR 198 at %P < 0.001. | ||||||
| Germination (%) | 100 | 50 | 100 | 100 | 70 | 100 |
| Plumule length (cm) | 3.47 ± 0.180 | 2.32 ± 0.182* | 4.83 ± 0.217% | 12.52 ± 0.271 | 10.31 ± 0.469* | 14.28 ± 0.196% |
| Radicle length (cm) | 5.11 ± 0.133 | 1.38 ± 0.22* | 4.91 ± 0.271% | 5.72 ± 0.116 | 2.35 ± 0.120* | 4.76 ± 0.211% |
In the case of the degradation of RR 198 by individual organisms, different metabolites were obtained. In the case of dye biodegradation by B. laterosporus; formation of products [A] 2-(phenylsulfonyl)ethanesulfonate [detected at retention time of Rt-21.467 having a molecular weight of MW-249 according to the mass to charge ratio of m/z 248(+1)], [B] (ethylsulfonyl)benzene [Rt-20.625, MW-170 based on the mass to charge ratio achieved at m/z 170] and [C] 6-chloro-N-phenyl-1,3,5-triazine-2,4-diamine [detected at Rt-17.350, MW-221 based on the mass to charge ratio of m/z 223(−2)] occurred due to the positive role of azo reductase and veratryl alcohol oxidase [Fig. 5A]. Whereas, only one product i.e. [A] (ethylsulfonyl)benzene [Rt-19.892, MW-170, according to the mass to charge ratio of m/z 170] was obtained in a single step via azo bond cleavage and desulphonation by azo reductase and veratryl alcohol oxidase respectively from G. geotrichum [Fig. 5B]. These results suggested that individual organisms partially mineralized the parent dye molecule; whereas, the consortium totally mineralized the RR 198 into simple forms. Recently, in a different approach, 96% decolorization of RR 198 was attained within 60 h using a plant system,40 but totally different metabolites were obtained which were not completely mineralized as the consortium did in the present study. These observations suggest that, each species has its own way of acting upon a substrate molecule with different metabolic fates.
3.5. Phytotoxicity
Untreated textile effluent released to the environment can exhibit toxicity, when used directly for the agricultural purposes. Hence it is of prime importance to study the toxic effects of treated as well as untreated dye samples on commonly used agricultural plants. Table 3 represents a toxicity analysis of RR 198 and its metabolites obtained after decolorization. In the dye samples, seed germination was inhibited by 50 and 30% in Sorghum vulgare and Phaseolus mungo respectively. But after treatment with the consortium, germination of Sorghum vulgare and Phaseolus mungo was enhanced by 100 and 30% respectively when compared to results with untreated dye. In addition, the metabolites produced after degradation of RR 198 might have played a nutritive role in plant growth regulation as both the plants showed better growth in metabolite solution as compared to plants grown in plain distilled water. Reduction in shoot and root length of both the plants clearly suggested the toxicity of the untreated azo dye. While, reduction in toxicity of the dye after treatment proved the effectiveness of the consortium.4. Conclusions
This study clearly suggests that the bacterial–yeast consortium has an enormous strength to catalyze the degradation of azo dye RR 198 within a short period compared to the individual microbial species. FTIR, HPTLC and GC-MS analysis indicated better mineralization of dye by the consortium with a significant reduction in its toxicity. The tolerance capacity of this developed consortium to high dye concentrations also recommends the application of this consortium into effluent treatment plants.Acknowledgements
M. B. Kurade and T. R. Waghmode wish to thank the Department of Biotechnology, New Delhi, India, for providing funds for this study (DBT Grant Number – BT/PR-10094/BCE/08/599/2007).References
- B. D. Tony, D. Goyal and S. Khanna, Int. Biodeterior. Biodegrad., 2009, 63, 462–469 CrossRef CAS PubMed.
- E. S. Yoo, J. Libra and U. Wiesmann, Water Sci. Technol., 2000, 1, 15–22 Search PubMed.
- J. Lin, X. Zhang, Z. Li and L. Lei, Bioresour. Technol., 2010, 101, 34–40 CrossRef CAS PubMed.
- K. Selvam, K. Swaminathan and K.-S. Chae, World J. Microbiol. Biotechnol., 2003, 19, 591–593 CrossRef CAS.
- O. J. Hao, H. Kim and P.-C. Chiang, Crit. Rev. Environ. Sci. Technol., 2000, 30, 449–505 CrossRef CAS.
- S. Mohanty, N. N. Rao, P. Khare and S. N. Kaul, Water Res., 2005, 39, 5064–5070 CrossRef CAS PubMed.
- N. Rao, G. Bose, P. Khare and S. Kaul, J. Environ. Eng., 2006, 132, 367–376 CrossRef CAS.
- G. McMullan, C. Meehan, A. Conneely, N. Kirby, T. Robinson, P. Nigam, I. M. Banat, R. Marchant and W. F. Smyth, Appl. Microbiol. Biotechnol., 2001, 56, 81–87 CrossRef CAS.
- K. Murugesan, Y. M. Kim, J. R. Jeon and Y. S. Chang, J. Hazard. Mater., 2009, 168, 523–529 CrossRef CAS PubMed.
- R. Renner, Environ. Sci. Technol., 1997, 31, 188A–189A CrossRef CAS PubMed.
- D. H. Pieper and W. Reineke, Curr. Opin. Biotechnol., 2000, 11, 262–270 CrossRef CAS.
- P. S. Patil, U. U. Shedbalkar, D. C. Kalyani and J. P. Jadhav, J. Ind. Microbiol. Biotechnol., 2008, 35, 1181–1190 CrossRef CAS PubMed.
- S. B. Jadhav, N. S. Patil, A. D. Watharkar, O. A. Apine and J. P. Jadhav, Environ. Sci. Pollut. Res., 2013, 20, 2854–2866 CrossRef CAS PubMed.
- S. P. Govindwar, M. B. Kurade, D. P. Tamboli, A. N. Kabra, P. J. Kim and T. R. Waghmode, Chemosphere, 2014, 109, 234–238 CrossRef CAS PubMed.
- N. Dafale, N. N. Rao, S. U. Meshram and S. R. Wate, Bioresour. Technol., 2008, 99, 2552–2558 CrossRef CAS PubMed.
- S. S. Phugare, D. C. Kalyani, S. N. Surwase and J. P. Jadhav, Ecotoxicol. Environ. Saf., 2011, 74, 1288–1296 CrossRef CAS PubMed.
- M. F. Coughlin, B. K. Kinkle and P. L. Bishop, Water Res., 2003, 37, 2757–2763 CrossRef CAS.
- L. Ayed, E. Khelifi, H. B. Jannet, H. Miladi, A. Cheref, S. Achour and A. Bakhrouf, Chem. Eng. J., 2010, 165, 200–208 CrossRef CAS PubMed.
- N. Dafale, S. Wate, S. Meshram and T. Nandy, J. Hazard. Mater., 2008, 159, 319–328 CrossRef CAS PubMed.
- E. Forgacs, T. Cserhati and G. Oros, Environ. Int., 2004, 30, 953–971 CrossRef CAS PubMed.
- Y. Patel, C. Mehta and A. Gupte, Int. Biodeterior. Biodegrad., 2012, 75, 187–193 CrossRef CAS PubMed.
- A. Mishra and A. Malik, Bioresour. Technol., 2014, 171, 217–226 CrossRef CAS PubMed.
- T. R. Waghmode, M. B. Kurade, A. N. Kabra and S. P. Govindwar, Microbiology, 2012, 158, 2344–2352 CrossRef CAS PubMed.
- L. Tan, S. Ning, X. Zhang and S. Shi, Bioresour. Technol., 2013, 138, 307–313 CrossRef CAS PubMed.
- M. S. Mahmoud, HBRC Journal, 2014 DOI:10.1016/j.hbrcj.2014.07.005 , article in press.
- F. He, W. Hu and Y. Li, Chemosphere, 2004, 57, 293–301 CrossRef CAS PubMed.
- M. B. Kurade, T. R. Waghmode, D. P. Tamboli and S. P. Govindwar, J. Basic Microbiol., 2013, 53, 136–146 CrossRef CAS PubMed.
- M. B. Kurade, T. R. Waghmode, A. N. Kabra and S. P. Govindwar, Biotechnol. Lett., 2013, 35, 1593–1598 CrossRef CAS PubMed.
- S. S. Gomare, D. P. Tamboli, A. N. Kagalkar and S. P. Govindwar, Int. Biodeterior. Biodegrad., 2009, 63, 582–586 CrossRef CAS PubMed.
- M. B. Kurade, T. R. Waghmode and S. P. Govindwar, J. Hazard. Mater., 2011, 192, 1746–1755 CrossRef CAS PubMed.
- M. B. Kurade, T. R. Waghmode, A. N. Kagalkar and S. P. Govindwar, Chem. Eng. J., 2012, 184, 33–41 CrossRef CAS PubMed.
- S. B. Jadhav, S. M. Yedurkar, S. S. Phugare and J. P. Jadhav, Clean: Soil, Air, Water, 2012, 40, 551–558 CrossRef CAS.
- T. R. Waghmode, M. B. Kurade, A. N. Kagalkar and S. P. Govindwar, J. Environ. Sci., 2012, 24, 1295–1304 CrossRef CAS.
- S. U. Jadhav, M. U. Jadhav, A. N. Kagalkar and S. P. Govindwar, J. Chin. Inst. Chem. Eng., 2008, 39, 563–570 CrossRef CAS PubMed.
- S. M. Joshi, S. A. Inamdar, A. A. Telke, D. P. Tamboli and S. P. Govindwar, Int. Biodeterior. Biodegrad., 2010, 64, 622–628 CrossRef CAS PubMed.
- S. S. Phugare, D. C. Kalyani, A. V. Patil and J. P. Jadhav, J. Hazard. Mater., 2011, 186, 713–723 CrossRef CAS PubMed.
- R. G. Saratale, G. D. Saratale, D. C. Kalyani, J. S. Chang and S. P. Govindwar, Bioresour. Technol., 2009, 100, 2493–2500 CrossRef CAS PubMed.
- A. N. Kabra, R. V. Khandare, M. B. Kurade and S. P. Govindwar, Environ. Sci. Pollut. Res., 2011, 18, 1360–1373 CrossRef CAS PubMed.
- P. Sathishkumar, T. Palvannan, K. Murugesan and S. Kamala-Kannan, Environ. Technol., 2013, 34, 139–147 CrossRef CAS PubMed.
- R. V. Khandare, A. N. Kabra, D. P. Tamboli and S. P. Govindwar, Chemosphere, 2011, 82, 1147–1154 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2015 |